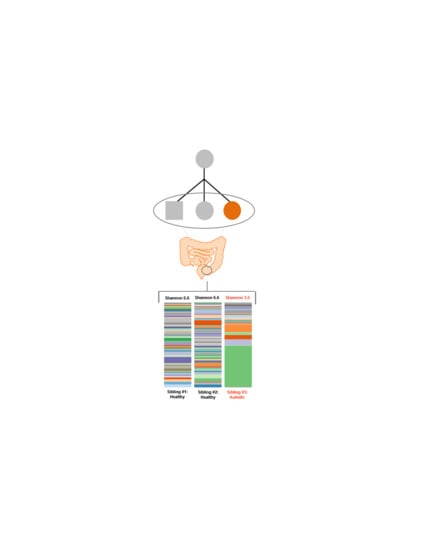
Shotgun Metagenomic Sequencing Identifies Dysbiosis in Triplet Sibling with Gastrointestinal Symptoms and ASD
Abstract
1. Introduction
2. Materials and Methods
3. Results
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Adams, J.B.; Johansen, L.J.; Powell, L.D.; Quig, D.W.; A Rubin, R. Gastrointestinal flora and gastrointestinal status in children with autism—Comparisons to typical children and correlation with autism severity. BMC Gastroenterol. 2011, 11, 22. [Google Scholar] [CrossRef] [PubMed]
- Vuong, H.E.; Hsiao, E.Y. Emerging Roles for the Gut Microbiome in Autism Spectrum Disorder. Biol. Psychiatry 2017, 81, 411–423. [Google Scholar] [CrossRef] [PubMed]
- Hughes, H.K.; Rose, D.; Ashwood, P. The Gut Microbiota and Dysbiosis in Autism Spectrum Disorders. Curr. Neurol. Neurosci. Rep. 2018, 18, 81. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Tian, J.; Yang, B. Targeting gut microbiome: A novel and potential therapy for autism. Life Sci. 2018, 194, 111–119. [Google Scholar] [CrossRef]
- Liu, F.; Li, J.; Wu, F.; Zheng, H.; Peng, Q.; Zhou, H. Altered composition and function of intestinal microbiota in autism spectrum disorders: A systematic review. Transl. Psychiatry 2019, 9, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Ho, L.K.H.; Tong, V.J.W.; Syn, N.; Nagarajan, N.; Tham, E.H.; Tay, S.K.; Shorey, S.; Tambyah, P.A.; Law, E.C.N. Gut microbiota changes in children with autism spectrum disorder: A systematic review. Gut Pathog. 2020, 12, 1–18. [Google Scholar] [CrossRef]
- Cussotto, S.; Strain, C.R.; Fouhy, F.; Strain, R.G.; Peterson, V.L.; Clarke, G.; Stanton, C.; Dinan, T.G.; Cryan, J.F. Differential effects of psychotropic drugs on microbiome composition and gastrointestinal function. Psychopharmacology 2019, 236, 1671–1685. [Google Scholar] [CrossRef] [PubMed]
- Collins, S.M.; Bercik, P. The Relationship Between Intestinal Microbiota and the Central Nervous System in Normal Gastrointestinal Function and Disease. Gastroenterology 2009, 136, 2003–2014. [Google Scholar] [CrossRef] [PubMed]
- Kang, D.W.; Adams, J.B.; Gregory, A.C.; Borody, T.; Chittick, L.; Fasano, A.; Khoruts, A.; Geis, E.; Maldonado, J.; McDonough-Means, S.; et al. Microbiota Transfer Therapy alters gut ecosystem and improves gastrointestinal and autism symptoms: An open-label study. Microbiome 2017, 5, 10. [Google Scholar] [CrossRef]
- Kang, D.-W.; Adams, J.B.; Coleman, D.M.; Pollard, E.L.; Maldonado, J.; McDonough-Means, S.; Caporaso, J.G.; Krajmalnik-Brown, R. Long-term benefit of Microbiota Transfer Therapy on autism symptoms and gut microbiota. Sci. Rep. 2019, 9, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Nitschke, A.; Deonandan, R.; Konkle, A.T. The link between autism spectrum disorder and gut microbiota: A scoping review. Autism 2020, 24, 1328–1344. [Google Scholar] [CrossRef] [PubMed]
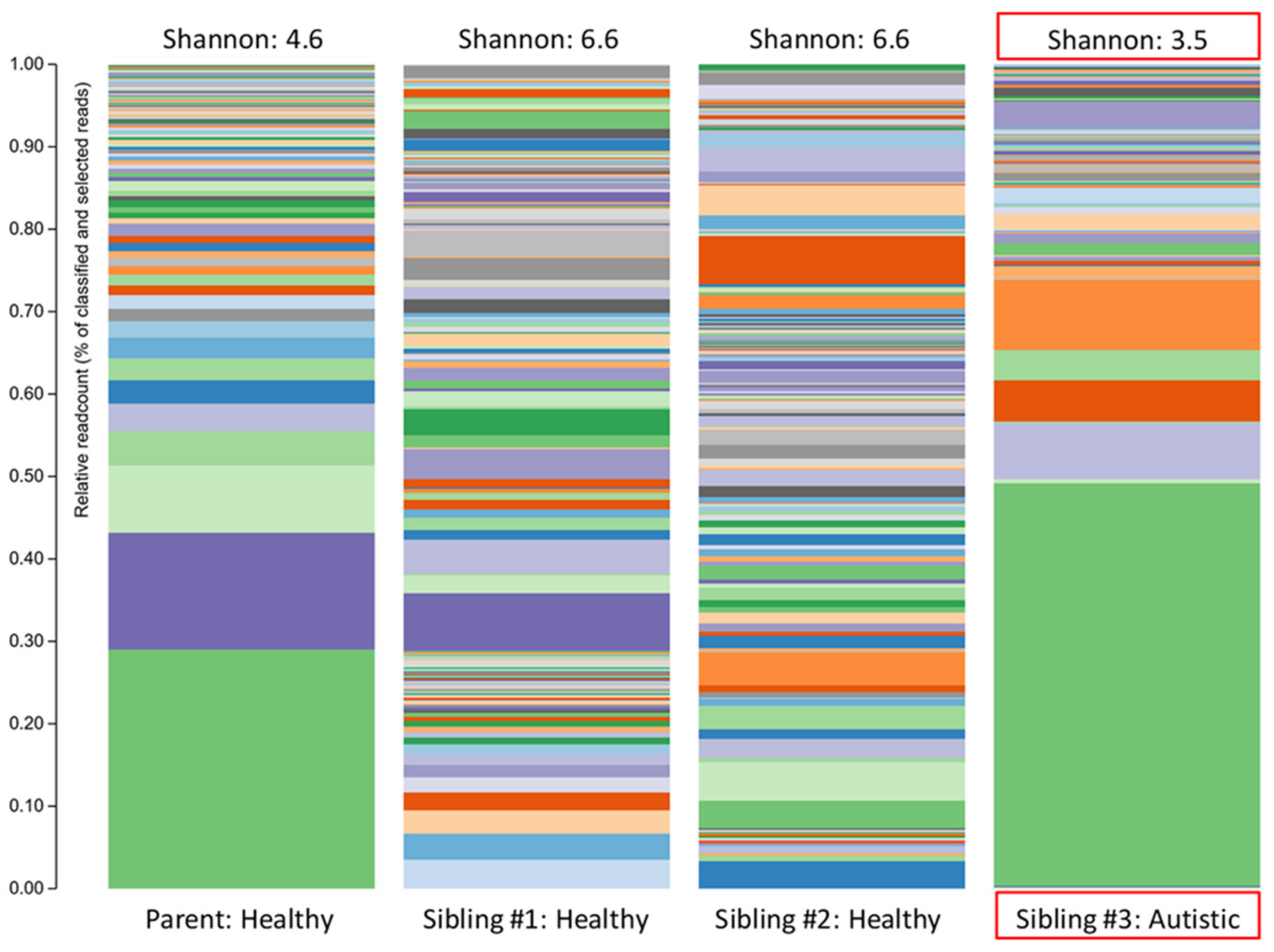
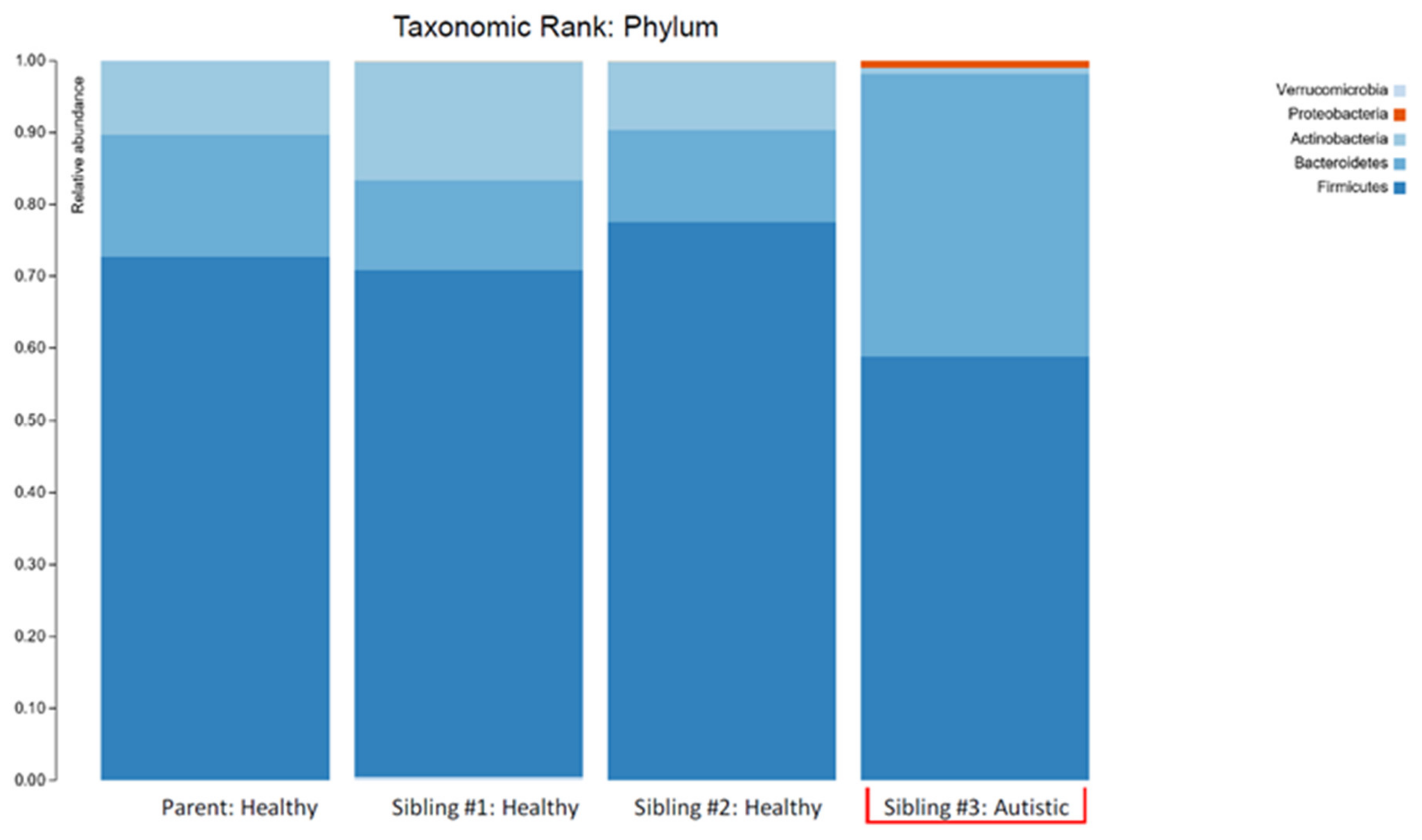
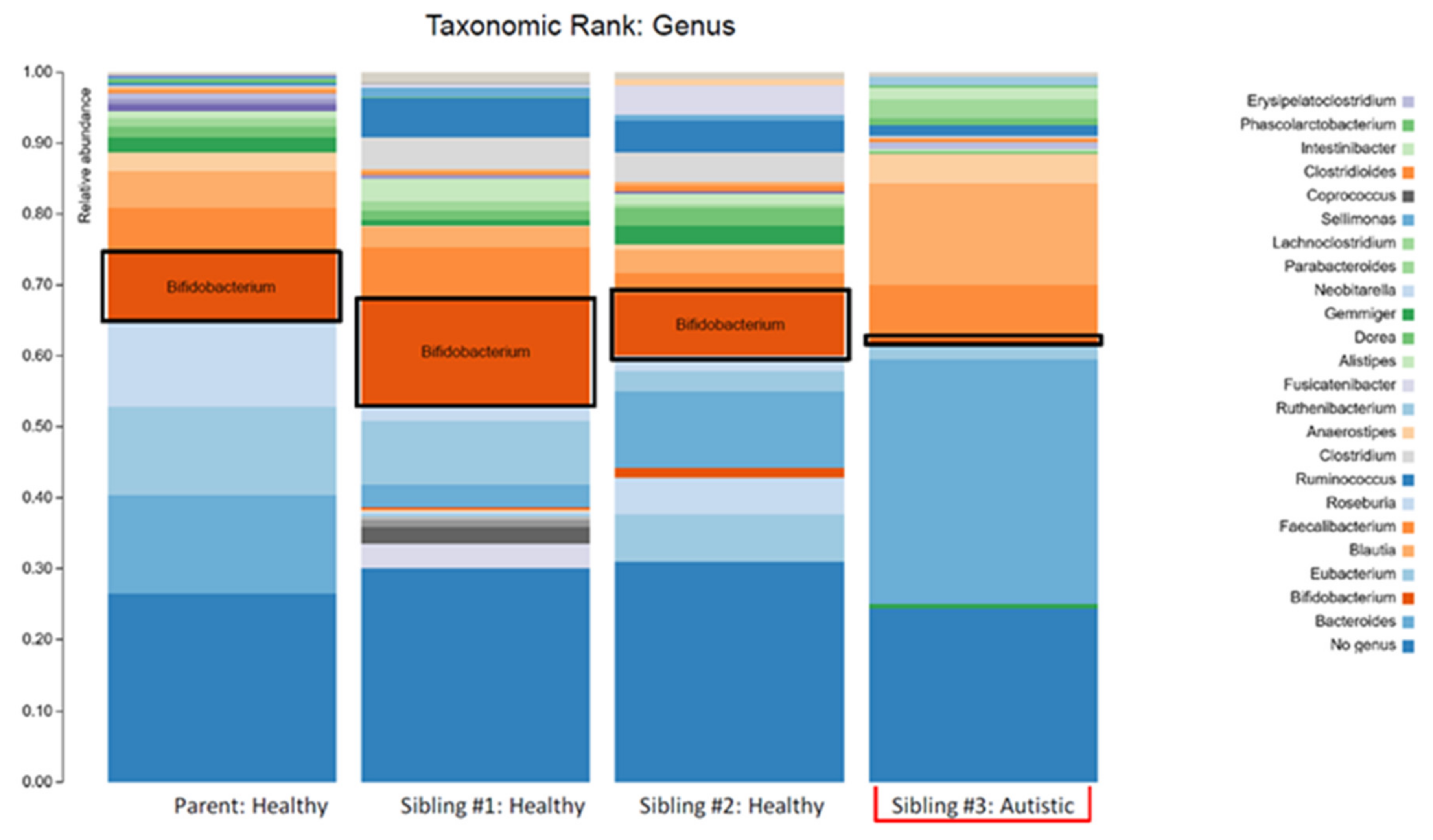
| Relative Abundance (%) | |||||
|---|---|---|---|---|---|
| Mother | Sibling #1 | Sibling #2 | Sibling #3 | ||
| Healthy | Healthy | Healthy | ASD | ||
| Phylum | Actinobacteria | 10.32 | 16.50 | 9.48 | 0.79 |
| Bacteroidetes | 17.07 | 12.39 | 12.90 | 39.33 | |
| Firmicutes | 72.59 | 70.38 | 77.48 | 58.82 | |
| Bacteroidetes/Firmicutes ratio | 0.24 | 0.18 | 0.17 | 0.67 | |
| Genus | Bacteroides | 13.84 | 3.14 | 10.76 | 34.57 |
| Eubacterium | 12.47 | 9.00 | 2.79 | 1.86 | |
| Bifidobacterium | 9.88 | 15.44 | 8.70 | 0.70 | |
| Blautia | 5.33 | 2.68 | 3.38 | 14.30 | |
| Species | Bacteroides uniformis | <0.01 | <0.01 | <0.01 | 16.86 |
| Bacteroides plebius | 11.72 | 0.00 | 1.01 | 17.71 | |
| Bacteroides vulgatus | 0.00 | 0.00 | 6.55 | 0.00 | |
| Roseburia faecis | 11.06 | 1.04 | 1.61 | 0.00 | |
| Bifidobacterium longum | 9.87 | 3.96 | 5.08 | 0.47 | |
| Bifidobacterium adolescentis | 0.00 | 11.48 | 3.62 | <0.01 | |
| Clostridiales bacterium | 2.32 | 2.73 | 11.56 | 0.00 | |
| Escherichia coli | 0.00 | 0.00 | 0.00 | 0.62 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hazan, S.; Spradling-Reeves, K.D.; Papoutsis, A.; Walker, S.J. Shotgun Metagenomic Sequencing Identifies Dysbiosis in Triplet Sibling with Gastrointestinal Symptoms and ASD. Children 2020, 7, 255. https://doi.org/10.3390/children7120255
Hazan S, Spradling-Reeves KD, Papoutsis A, Walker SJ. Shotgun Metagenomic Sequencing Identifies Dysbiosis in Triplet Sibling with Gastrointestinal Symptoms and ASD. Children. 2020; 7(12):255. https://doi.org/10.3390/children7120255
Chicago/Turabian StyleHazan, Sabine, Kimberly D. Spradling-Reeves, Andreas Papoutsis, and Stephen J. Walker. 2020. "Shotgun Metagenomic Sequencing Identifies Dysbiosis in Triplet Sibling with Gastrointestinal Symptoms and ASD" Children 7, no. 12: 255. https://doi.org/10.3390/children7120255
APA StyleHazan, S., Spradling-Reeves, K. D., Papoutsis, A., & Walker, S. J. (2020). Shotgun Metagenomic Sequencing Identifies Dysbiosis in Triplet Sibling with Gastrointestinal Symptoms and ASD. Children, 7(12), 255. https://doi.org/10.3390/children7120255