Early Gut Microbiota Profile in Healthy Neonates: Microbiome Analysis of the First-Pass Meconium Using Next-Generation Sequencing Technology
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population and Sampling
2.2. DNA Extraction and Sequencing
2.3. Bioinformatic Analysis
3. Results
3.1. Study Cohort
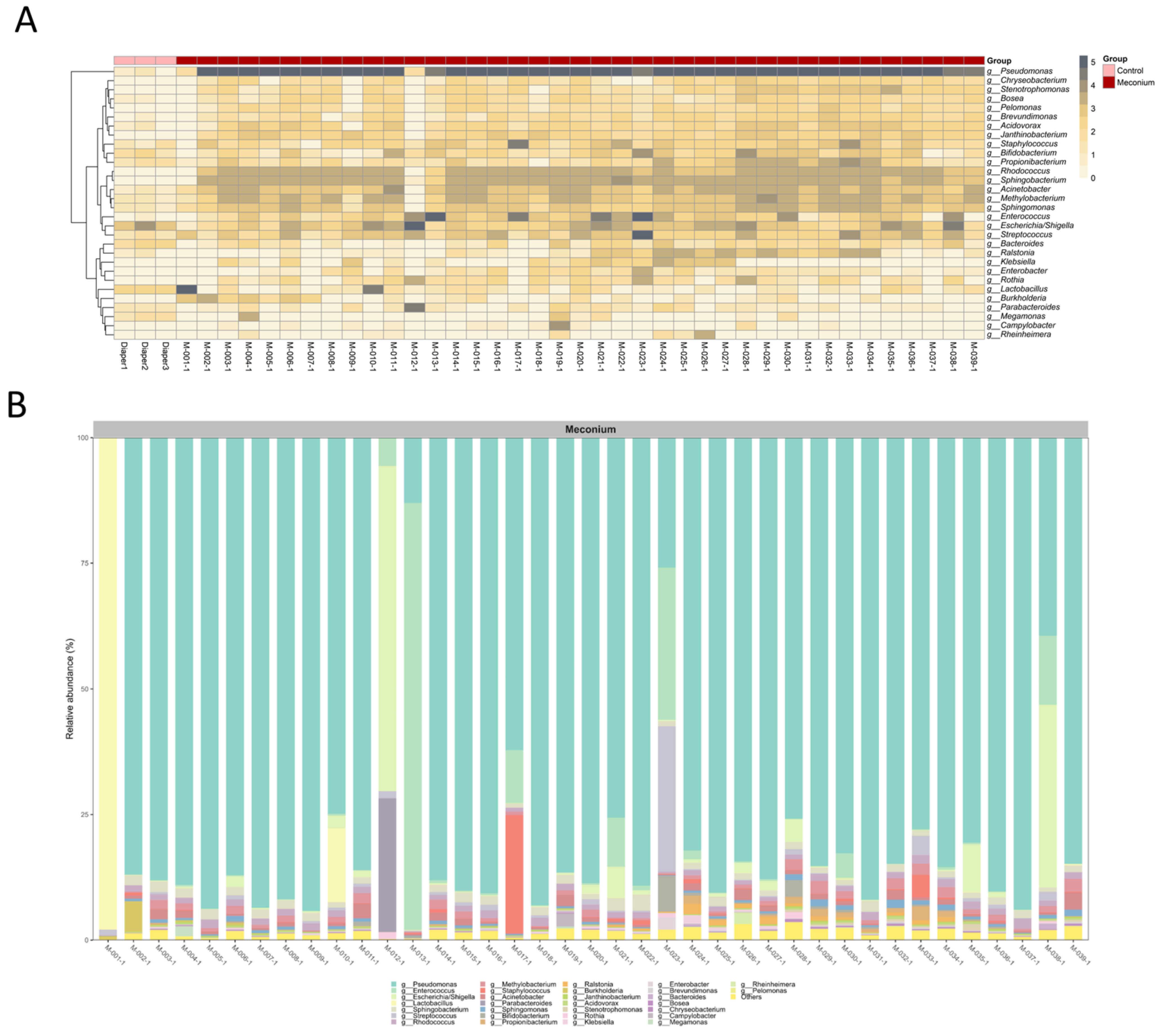
3.2. Distinct Microbiota Community Structures in the Meconium Samples and Control Samples
3.3. A Similar Profile of the Microbiota Composition in the First-Pass Meconium
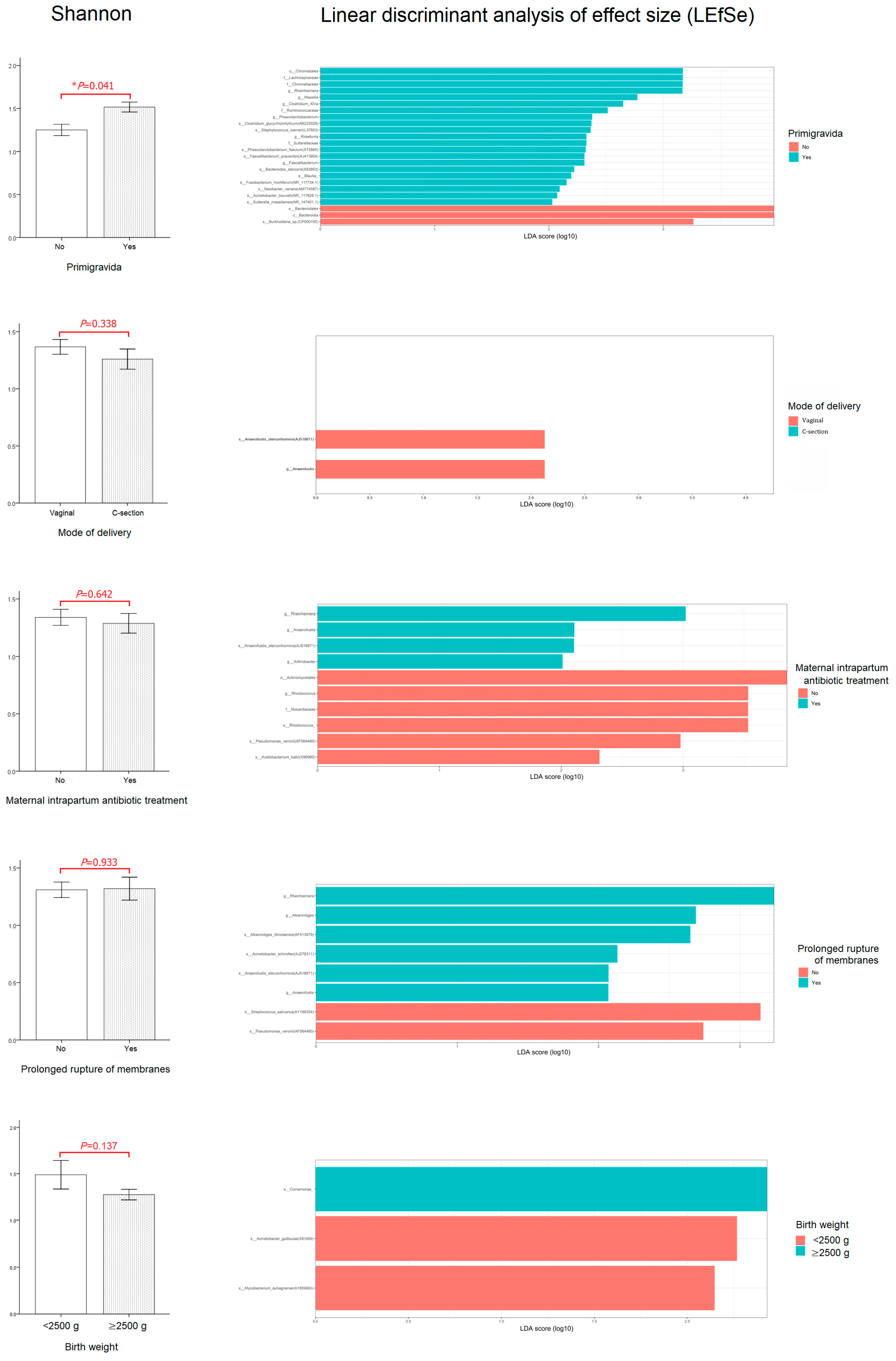
3.4. Relationship between Perinatal Characteristics and the First-Pass Meconium Microbiome
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Del Chierico, F.; Vernocchi, P.; Bonizzi, L.; Carsetti, R.; Castellazzi, A.M.; Dallapiccola, B.; de Vos, W.; Guerzoni, M.E.; Manco, M.; Marseglia, G.L.; et al. Early-life gut microbiota under physiological and pathological conditions: The central role of combined meta-omics-based approaches. J. Proteom. 2012, 75, 4580–4587. [Google Scholar] [CrossRef] [PubMed]
- Tain, Y.L.; Hsu, C.N. Role of the gut microbiota in children with kidney disease. Children 2023, 10, 269. [Google Scholar] [CrossRef]
- Zhang, X.; Zeng, S.; Cheng, G.; He, L.; Chen, M.; Wang, M.; Zhou, W.; Qiu, H.; Wang, Z. Clinical manifestations of neonatal hyperbilirubinemia are related to alterations in the gut microbiota. Children 2022, 9, 764. [Google Scholar] [CrossRef]
- Martin, V.M.; Virkud, Y.V.; Dahan, E.; Seay, H.L.; Itzkovits, D.; Vlamakis, H.; Xavier, R.; Shreffler, W.G.; Yuan, Q.; Yassour, M. Longitudinal disease-associated gut microbiome differences in infants with food protein-induced allergic proctocolitis. Microbiome 2022, 10, 154. [Google Scholar] [CrossRef]
- Girdhar, K.; Huang, Q.; Chow, I.T.; Vatanen, T.; Brady, C.; Raisingani, A.; Autissier, P.; Atkinson, M.A.; Kwok, W.W.; Kahn, C.R.; et al. A gut microbial peptide and molecular mimicry in the pathogenesis of type 1 diabetes. Proc. Natl. Acad. Sci. USA 2022, 119, e2120028119. [Google Scholar] [CrossRef] [PubMed]
- Biscetti, F.; Nardella, E.; Cecchini, A.L.; Landolfi, R.; Flex, A. The role of the microbiota in the diabetic peripheral artery disease. Mediat. Inflamm. 2019, 2019, 4128682. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.; Gao, R.; Zhang, Y.; Pan, D.; Zhu, Y.; Zhang, X.; Yang, R.; Jiang, R.; Xu, Y.; Qin, H. Dysbiosis signatures of gut microbiota in coronary artery disease. Physiol. Genom. 2018, 50, 893–903. [Google Scholar] [CrossRef]
- Pittayanon, R.; Lau, J.T.; Yuan, Y.; Leontiadis, G.I.; Tse, F.; Surette, M.; Moayyedi, P. Gut microbiota in patients with irritable bowel syndrome-a systematic review. Gastroenterology 2019, 157, 97–108. [Google Scholar] [CrossRef]
- Zhu, Z.; Ren, J.; Michail, S.; Sun, F. MicroPro: Using metagenomic unmapped reads to provide insights into human microbiota and disease associations. Genome Biol. 2019, 20, 154. [Google Scholar] [CrossRef]
- Zhang, L.; Wu, Y.-N.; Chen, T.; Ren, C.-H.; Li, X.; Liu, G.-X. Relationship between intestinal microbial dysbiosis and primary liver cancer. Hepatobiliary Pancreat. Dis. Int. 2019, 18, 149–157. [Google Scholar] [CrossRef]
- Komaroff, A.L. The microbiome and risk for atherosclerosis. JAMA 2018, 319, 2381–2382. [Google Scholar] [CrossRef] [PubMed]
- Mady, E.A.; Doghish, A.S.; El-Dakroury, W.A.; Elkhawaga, S.Y.; Ismail, A.; El-Mahdy, H.A.; Elsakka, E.G.E.; El-Husseiny, H.M. Impact of the mother’s gut microbiota on infant microbiome and brain development. Neurosci. Biobehav. Rev. 2023, 150, 105195. [Google Scholar] [CrossRef] [PubMed]
- Gritz, E.C.; Bhandari, V. The human neonatal gut microbiome: A brief review. Front. Pediatr. 2015, 1, 17. [Google Scholar]
- Wernroth, M.L.; Peura, S.; Hedman, A.M.; Hetty, S.; Vicenzi, S.; Kennedy, B.; Fall, K.; Svennblad, B.; Andolf, E.; Pershagen, G.; et al. Development of gut microbiota during the first 2 years of life. Sci. Rep. 2022, 12, 9080. [Google Scholar] [CrossRef]
- Roswall, J.; Olsson, L.M.; Kovatcheva-Datchary, P.; Nilsson, S.; Tremaroli, V.; Simon, M.C.; Kiilerich, P.; Akrami, R.; Kramer, M.; Uhlen, M.; et al. Developmental trajectory of the healthy human gut microbiota during the first 5 years of life. Cell Host Microbe 2021, 29, 765–776. [Google Scholar] [CrossRef] [PubMed]
- Backhed, F.; Roswall, J.; Peng, Y.; Feng, Q.; Jia, H.; Kovatcheva-Datchary, P.; Li, Y.; Xia, Y.; Xie, H.; Zhong, H.; et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 2015, 17, 690–703. [Google Scholar] [CrossRef]
- Stewart, C.J.; Ajami, N.J.; O’Brien, J.L.; Hutchinson, D.S.; Smith, D.P.; Wong, M.C.; Ross, M.C.; Lloyd, R.E.; Doddapaneni, H.; Metcalf, G.A.; et al. Temporal development of the gut microbiome in early childhood from the TEDDY study. Nature 2018, 562, 583–588. [Google Scholar] [CrossRef]
- Nino, G.; Rodriguez-Martinez, C.E.; Gutierrez, M.J. Early microbial-immune interactions and innate immune training of the respiratory system during health and disease. Children 2021, 8, 413. [Google Scholar] [CrossRef]
- Dogra, S.; Sakwinska, O.; Soh, S.E.; Ngom-Bru, C.; Brück, W.M.; Berger, B.; Brüssow, H.; Karnani, N.; Lee, Y.S.; Yap, F.; et al. Rate of establishing the gut microbiota in infancy has consequences for future health. Gut Microbes 2015, 6, 321–325. [Google Scholar] [CrossRef]
- Alderete, T.L.; Jones, R.B.; Shaffer, J.P.; Holzhausen, E.A.; Patterson, W.B.; Kazemian, E.; Chatzi, L.; Knight, R.; Plows, J.F.; Berger, P.K.; et al. Early life gut microbiota is associated with rapid infant growth in Hispanics from Southern California. Gut Microbes 2021, 13, 1961203. [Google Scholar] [CrossRef]
- Tanaka, M.; Nakayama, J. Development of the gut microbiota in infancy and its impact on health in later life. Allergol. Int. 2017, 66, 515–522. [Google Scholar] [CrossRef]
- Turunen, J.; Tejesvi, M.V.; Paalanne, N.; Hekkala, J.; Lindgren, O.; Kaakinen, M.; Pokka, T.; Kaisanlahti, A.; Reunanen, J.; Tapiainen, T. Presence of distinctive microbiome in the first-pass meconium of newborn infants. Sci. Rep. 2021, 11, 19449. [Google Scholar] [CrossRef]
- Hansen, R.; Scott, K.P.; Khan, S.; Martin, J.C.; Berry, S.H.; Stevenson, M.; Okpapi, A.; Munro, M.J.; Hold, G.L. First-pass meconium samples from healthy term vaginally-delivered neonates: An analysis of the microbiota. PLoS ONE 2015, 10, e0133320. [Google Scholar] [CrossRef] [PubMed]
- Ardissone, A.N.; de la Cruz, D.M.; Davis-Richardson, A.G.; Rechcigl, K.T.; Li, N.; Drew, J.C.; Murgas-Torrazza, R.; Sharma, R.; Hudak, M.L.; Triplett, E.W.; et al. Meconium microbiome analysis identifies bacteria correlated with premature birth. PLoS ONE 2014, 9, e90784. [Google Scholar] [CrossRef]
- Jimenez, E.; Marin, M.L.; Martin, R.; Odriozola, J.M.; Olivares, M.; Xaus, J.; Fernandez, L.; Rodriguez, J.M. Is meconium from healthy newborns actually sterile? Res. Microbiol. 2008, 159, 187–193. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.C.; Guo, H.; Chen, J.; Sun, G.; Ren, R.R.; Guo, M.Z.; Peng, L.H.; Yang, Y.S. Initial meconium microbiome in Chinese neonates delivered naturally or by cesarean section. Sci. Rep. 2018, 8, 3255. [Google Scholar] [CrossRef] [PubMed]
- Scholtens, P.A.; Oozeer, R.; Martin, R.; Amor, K.B.; Knol, J. The early settlers: Intestinal microbiology in early life. Annu. Rev. Food Sci. Technol. 2012, 3, 425–447. [Google Scholar] [CrossRef]
- Kielenniva, K.; Ainonen, S.; Vanni, P.; Paalanne, N.; Renko, M.; Salo, J.; Tejesvi, M.V.; Pokka, T.; Pirttila, A.M.; Tapiainen, T. Microbiota of the first-pass meconium and subsequent atopic and allergic disorders in children. Clin. Exp. Allergy 2022, 52, 684–696. [Google Scholar] [CrossRef]
- Dornelles, L.V.; Procianoy, R.S.; Roesch, L.F.W.; Corso, A.L.; Dobbler, P.T.; Mai, V.; Silveira, R.C. Meconium microbiota predicts clinical early-onset neonatal sepsis in preterm neonates. J. Matern. Fetal Neonatal Med. 2022, 35, 1935–1943. [Google Scholar] [CrossRef]
- Terrazzan Nutricionist, A.C.; Procianoy, R.S.; Roesch, L.F.W.; Corso, A.L.; Dobbler, P.T.; Silveira, R.C. Meconium microbiome and its relation to neonatal growth and head circumference catch-up in preterm infants. PLoS ONE 2020, 15, e0238632. [Google Scholar] [CrossRef]
- Allaband, C.; McDonald, D.; Vázquez-Baeza, Y.; Minich, J.J.; Tripathi, A.; Brenner, D.A.; Loomba, R.; Smarr, L.; Sandborn, W.J.; Schnabl, B.; et al. Microbiome 101: Studying, analyzing, and interpreting gut microbiome data for clinicians. Clin. Gastroenterol. Hepatol. 2019, 17, 218–230. [Google Scholar] [CrossRef] [PubMed]
- Putignani, L.; Del Chierico, F.; Petrucca, A.; Vernocchi, P.; Dallapiccola, B. The human gut microbiota: A dynamic interplay with the host from birth to senescence settled during childhood. Pediatr. Res. 2014, 76, 2–10. [Google Scholar] [CrossRef]
- El Mouzan, M.; Al-Hussaini, A.A.; Al Sarkhy, A.; Assiri, A.; Alasmi, M. Intestinal microbiota profile in healthy Saudi children: The bacterial domain. Saudi J. Gastroenterol. 2022, 28, 312–317. [Google Scholar] [CrossRef] [PubMed]
- Rothschild, D.; Weissbrod, O.; Barkan, E.; Kurilshikov, A.; Korem, T.; Zeevi, D.; Costea, P.I.; Godneva, A.; Kalka, I.N.; Bar, N.; et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 2018, 1, 210–215. [Google Scholar] [CrossRef]
- Penders, J.; Thijs, C.; Vink, C.; Stelma, F.F.; Snijders, B.; Kummeling, I.; van den Brandt, P.A.; Stobberingh, E.E. Factors influencing the composition of the intestinal microbiota in early infancy. Pediatrics 2006, 118, 511–521. [Google Scholar] [CrossRef] [PubMed]
- Yatsunenko, T.; Rey, F.E.; Manary, M.J.; Trehan, I.; Dominguez-Bello, M.G.; Contreras, M.; Magris, M.; Hidalgo, G.; Baldassano, R.N.; Anokhin, A.P.; et al. Human gut microbiome viewed across age and geography. Nature 2012, 486, 222–227. [Google Scholar] [CrossRef]
- Reyman, M.; van Houten, M.A.; van Baarle, D.; Bosch, A.; Man, W.H.; Chu, M.; Arp, K.; Watson, R.L.; Sanders, E.A.M.; Fuentes, S.; et al. Impact of delivery mode-associated gut microbiota dynamics on health in the first year of life. Nat. Commun. 2019, 10, 4997. [Google Scholar] [CrossRef] [PubMed]
- Bokulich, N.A.; Chung, J.; Battaglia, T.; Henderson, N.; Jay, M.; Li, H.; Lieber, A.D.; Wu, F.; Perez-Perez, G.I.; Chen, Y.; et al. Antibiotics, birth mode, and diet shape microbiome maturation during early life. Sci. Transl. Med. 2016, 8, 343ra382. [Google Scholar] [CrossRef]
- Arumugam, M.; Raes, J.; Pelletier, E.; Le Paslier, D.; Yamada, T.; Mende, D.R.; Fernandes, G.R.; Tap, J.; Bruls, T.; Batto, J.M.; et al. Enterotypes of the human gut microbiome. Nature 2011, 473, 174–180. [Google Scholar] [CrossRef]
- Dos Santos, S.J.; Pakzad, Z.; Elwood, C.N.; Albert, A.Y.K.; Gantt, S.; Manges, A.R.; Dumonceaux, T.J.; Maan, E.J.; Hill, J.E.; Money, D.M. Early neonatal meconium does not have a demonstrable microbiota determined through use of robust negative controls with cpn60-based microbiome profiling. Microbiol. Spectr. 2021, 9, e0006721. [Google Scholar] [CrossRef]


| Characteristics | N = 39 |
|---|---|
| Gender | |
| Male | 19 |
| Female | 20 |
| Gestational age (weeks) | 38.1 (35.2–40.0) |
| Birth weight (g) | 2880 (2020–3620) |
| Mode of delivery | |
| Vaginal | 19 |
| C-section | 20 |
| Primigravida | 9 (23.0%) |
| Mother’s age (years) | 33.8 (23.9–45.9) |
| Maternal intrapartum antibiotic treatment | 21 (53.8%) |
| Prolonged rupture of membranes | 12 (30.7%) |
| Feeding style | |
| Exclusive breastfeeding or formula feeding | 0 |
| Mixed (breast + formula feeding) | 39 (100%) |
| Level | Organism | Abundance (%) | Level | Organism | Abundance (%) |
|---|---|---|---|---|---|
| Phylum | Proteobacteria | 85.45 | Family | Pseudomonadaceae | 77.11 |
| Firmicutes | 9.74 | Enterococcaceae | 4.30 | ||
| Bacteroidetes | 2.42 | Enterobacteriaceae | 3.88 | ||
| Actinobacteria | 2.25 | Lactobacillaceae | 2.91 | ||
| Fusobacteria | 0.03 | Sphingobacteriaceae | 1.43 | ||
| Class | Gammaproteobacteria | 82.21 | Genus | Pseudomonas | 77.09 |
| Bacilli | 9.44 | Enterococcus | 4.30 | ||
| Actinobacteria | 2.25 | Escherichia/Shigella | 3.60 | ||
| Alphaproteobacteria | 1.96 | Lactobacillus | 2.91 | ||
| Sphingobacteriia | 1.43 | Sphingobacterium | 1.42 | ||
| Order | Pseudomonadales | 78.06 | Species | Pseudomonas sp. (EU557337) | 17.35 |
| Lactobacillales | 8.49 | Streptococcus salivarius | 0.83 | ||
| Enterobacteriales | 3.88 | Parabacteroides distasonis | 0.69 | ||
| Actinomycetales | 1.80 | Enterococcus faecalis | 0.65 | ||
| Sphingobacteriales | 1.43 | Methylobacterium komagatae | 0.50 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chang, Y.-S.; Li, C.-W.; Chen, L.; Wang, X.-A.; Lee, M.-S.; Chao, Y.-H. Early Gut Microbiota Profile in Healthy Neonates: Microbiome Analysis of the First-Pass Meconium Using Next-Generation Sequencing Technology. Children 2023, 10, 1260. https://doi.org/10.3390/children10071260
Chang Y-S, Li C-W, Chen L, Wang X-A, Lee M-S, Chao Y-H. Early Gut Microbiota Profile in Healthy Neonates: Microbiome Analysis of the First-Pass Meconium Using Next-Generation Sequencing Technology. Children. 2023; 10(7):1260. https://doi.org/10.3390/children10071260
Chicago/Turabian StyleChang, Yi-Sheng, Chang-Wei Li, Ling Chen, Xing-An Wang, Maw-Sheng Lee, and Yu-Hua Chao. 2023. "Early Gut Microbiota Profile in Healthy Neonates: Microbiome Analysis of the First-Pass Meconium Using Next-Generation Sequencing Technology" Children 10, no. 7: 1260. https://doi.org/10.3390/children10071260
APA StyleChang, Y.-S., Li, C.-W., Chen, L., Wang, X.-A., Lee, M.-S., & Chao, Y.-H. (2023). Early Gut Microbiota Profile in Healthy Neonates: Microbiome Analysis of the First-Pass Meconium Using Next-Generation Sequencing Technology. Children, 10(7), 1260. https://doi.org/10.3390/children10071260





