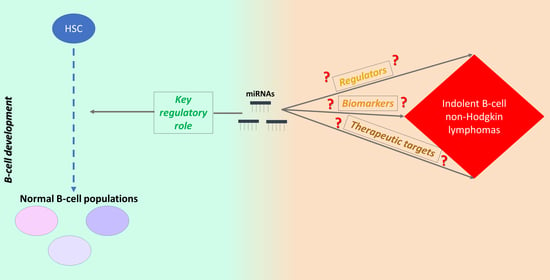
The Multifaceted Role and Utility of MicroRNAs in Indolent B-Cell Non-Hodgkin Lymphomas
Abstract
1. Introduction
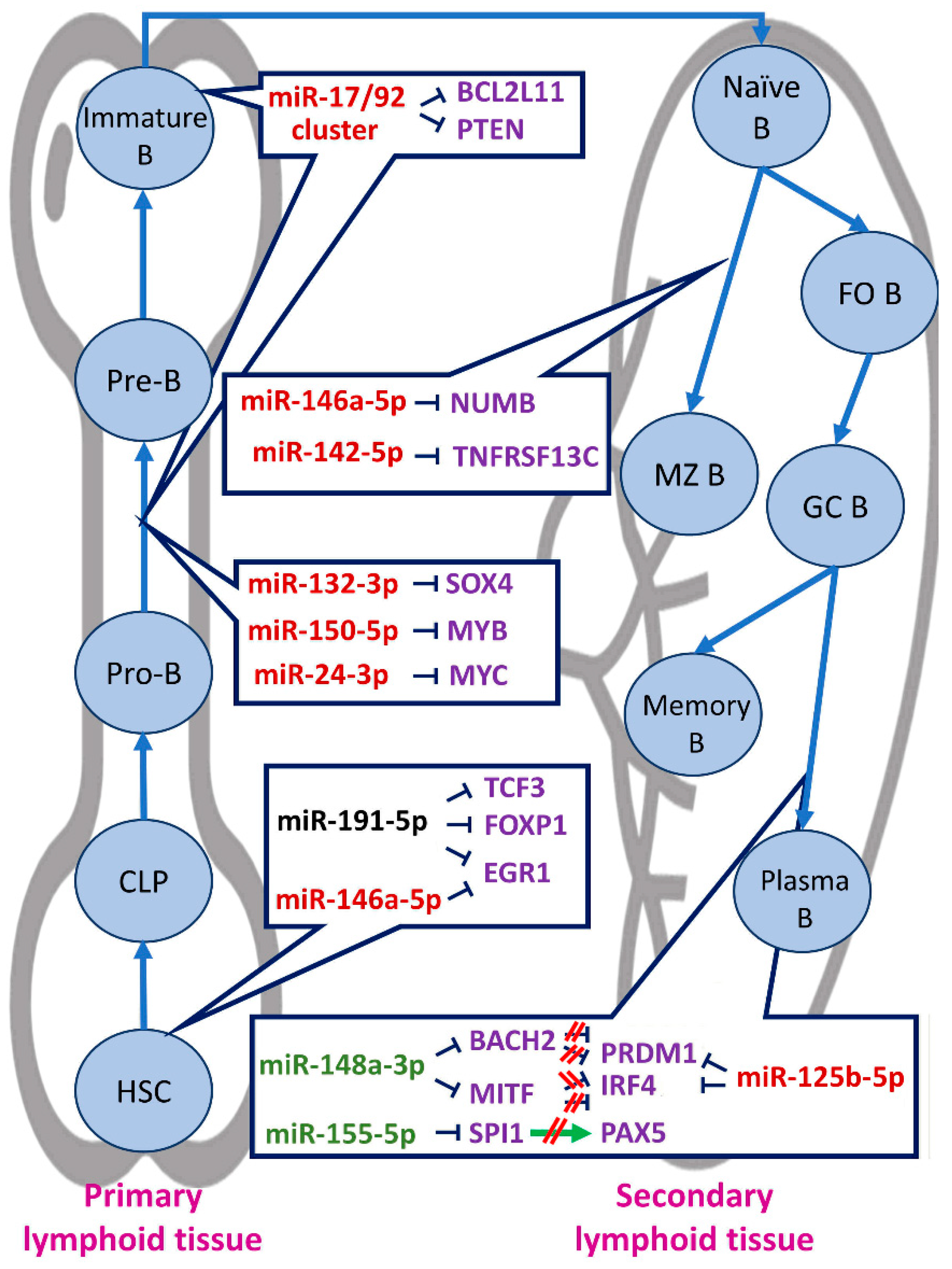
2. miRNAs in Normal B-Cell Development
2.1. miRNAs in Primary Lymphoid Tissue B-Cell Development
2.2. miRNAs in Secondary Lymphoid Tissue B-Cell Development
3. miRNAs in Follicular Lymphoma
3.1. miRNAs as Potential Regulators and Biomarkers in FL
3.2. Genetic Polymorphisms of miRNA Genes in FL
3.3. miRNAs and the Immune System
| miRNAs | Sample Origin | Expression | Potential Biomarker | References |
|---|---|---|---|---|
| miRNAs of miR-17/92 and miR-106a-363 clusters, miR-200c-3p, miR-638, miR-518a-3p | FFPE tissues | Upregulated in DLBCL vs. FL | Differential diagnosis | [53] |
| miR-17-5p | [54] | |||
| miR-217-5p, miR-634, miR-26b-5p | Upregulated in FL vs. DLBCL | [53] | ||
| miR-330-3p, miR-106a-5p, miR-210-3p | Upregulated in FL vs. DLBCL | [54] | ||
| miR-612, miR-188-5p, miR-302c-3p, miR-433-3p, miR-584-5p | Upregulated in BCL2+/BCL6+ and BCL2−/BCL6+ FL vs. BCL2+/BCL6− FL | [51] | ||
| miR-200a-3p, miR-135a-5p, miR-375-3p, miR-138-5p, miR-517 isomiRs | Downregulated in BCL2+/BCL6+ and BCL2−/BCL6+ FL vs. BCL2+/BCL6− FL | |||
| miR-16-5p, miR-138-5p, miR-26a-5p, miR-29c-3p | Downregulated in t(14;18)–negative FL vs. t(14;18)–positive FL | [48] | ||
| miR-193a-5p, miR-345-5p, miR-574-3p, miR-1287-5p, miR-1471 | Enriched FL cells | Upregulated in FL vs. with follicular hyperplasia | [63] | |
| miR-570-3p, miR-205-5p, miR-222-3p, miR-30a-5p, miR-301b-3p, miR-141-3p | Downregulated in FL vs. with follicular hyperplasia | |||
| miR-20b-5p, miR-26a-5p, miR-92b-3p, miR-487b-3p | Cancer cell lines | Upregulated in FL cell lines vs. DLBCL cell lines | [87] | |
| miR-330-3p, miR-106a-5p, miR-210-3p, miR-301 isomiRs, miR-338-5p | FFPE tissues | Upregulated in FL vs. non- neoplastic lymph nodes | Diagnosis | [54] |
| miR-149-5p, miR-139-5p | Downregulated in FL vs. non- neoplastic lymph nodes | |||
| miR-16-5p, miR-17-5p, miR-26a-5p, miR-29a-3p, let-7d-5p, let-7g-5p, let-7i-5p | Cancer cell lines | Downregulated in FL cell lines vs. CD19+ negatively- selected B cells | [87] | |
| miR-144-3p, miR-431-5p | FL biopsies and blood samples | Upregulated in relapsed FL patients | Prognosis, prediction of disease progression | [88] |
| miR-376b-3p | Downregulated in non-relapsed FL patients |
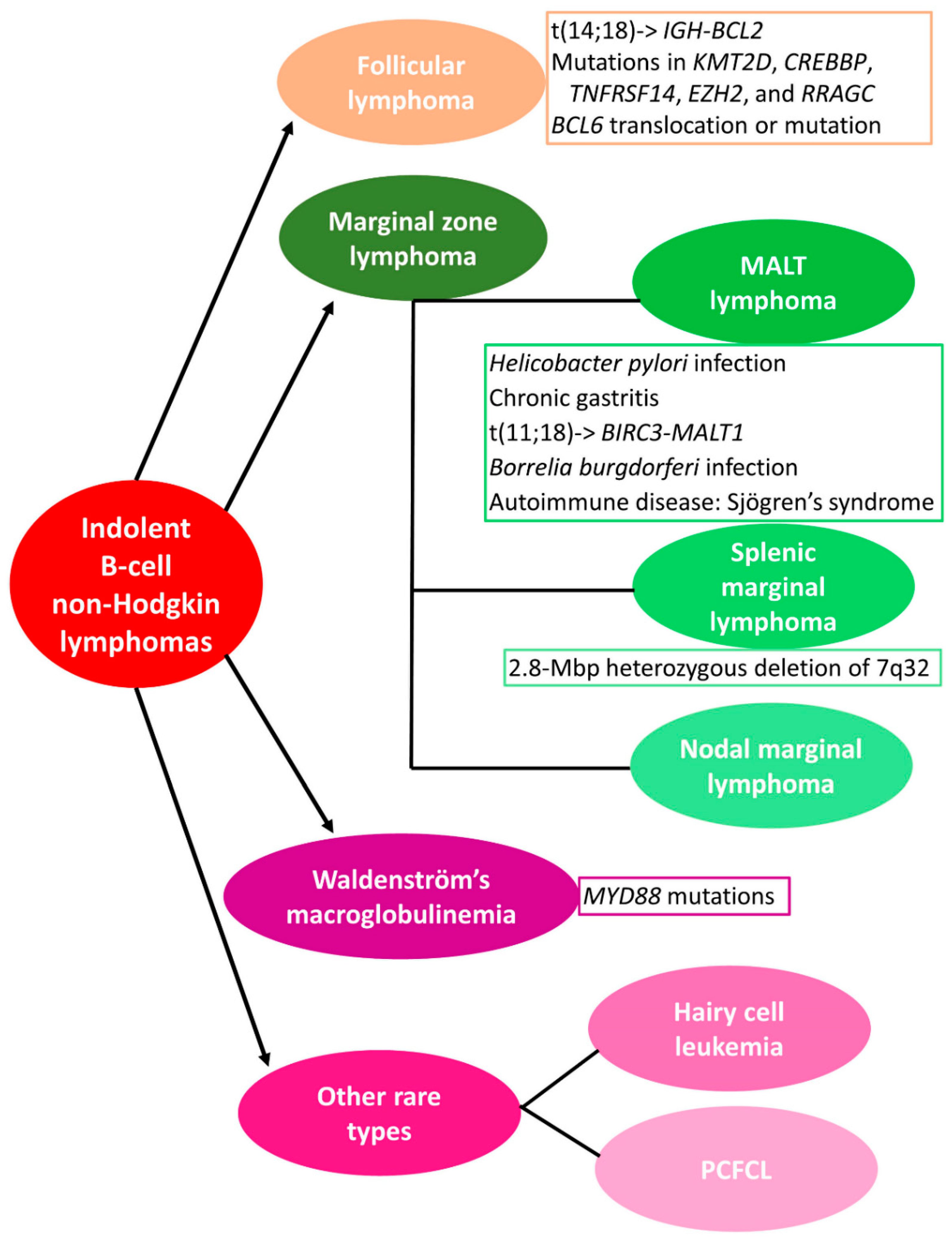
4. miRNAs in Marginal Zone Lymphoma
4.1. Extranodal Marginal Zone Lymphoma or Mucosa-Associated Lymphoid Tissue (MALT) Lymphoma
4.2. Gastric MALT Lymphoma and H. pylori
4.3. From Chronic Gastritis to MALT GL
4.4. From MALT GL to Gastric DLBCL
4.5. Non-Gastric MALT Lymphoma
4.6. Splenic Marginal Zone Lymphoma
4.7. Nodal Marginal Zone Lymphoma
| Disease | miRNAs | Sample Origin | Expression | Potential Biomarker | References |
|---|---|---|---|---|---|
| Gastric MALT lymphoma | miR-142-3p, miR-155-5p | FFPE tissues | Upregulated in gastric MALT lymphoma vs. chronic gastritis | Differential diagnosis | [74] |
| miR-203a-3p | Downregulated in gastric MALT lymphoma vs. chronic gastritis | ||||
| Ocular adnexal lymphoma (OAL) | let-7g-5p, miR-16-5p, miR-29 family, miR-199a-5p, miR-222-3p | FFPE tissues | Upregulated in OAL vs. ocular DLBCL | Differential diagnosis | [118] |
| Sjögren’s syndrome (SS) associated with MALT lymphoma | miR-200b-5p | Minor salivary glands and PBMCs | Downregulated in SS-associated MALT lymphoma vs. SS | Prognosis, prediction of patients’ relapse | [123,124] |
| Splenic MZL (SMZL) | miR-21-5p | Fresh frozen and FFPE tissues | Upregulated in aggressive SMZL vs. indolent SMZL | Prognosis, prediction of patients’ relapse | [133] |
| Nodal MZL (NMZL) | miR-223-3p, let-7f-5p | FFPE tissues | Upregulated in NMZL vs. FL | Differential diagnosis | [139] |
5. miRNAs in Rare Types of Indolent B-Cell NHLs
5.1. Waldenström’s Macroglobulinemia or Lymphoplasmacytic Lymphoma
5.2. Other Rare Types of B-Cell NHLs
| Disease | miRNAs | Sample Origin | Expression | Potential Biomarker | References |
|---|---|---|---|---|---|
| WM | miR-363-3p, miR-206-3p, miR-494-3p, miR-155-5p, miR-184-3p, miR-542-3p | Bone marrow CD19+ B cells | Upregulated in WM vs. normal CD19+ B cells | Diagnosis | [81] |
| miR-9-3p | Downregulated in WM vs. normal CD19+ B cells | ||||
| miR-193b-3p, miR-126-3p, miR-181a-5p, miR-125b-5p, miR-451a | Bone marrow or peripheral blood CD19+ and CD138+ B cells | Upregulated in WM vs. CLL | Differential diagnosis | [149] | |
| miR-92a-3p, miR-223-3p, miR-92b-3p, miR-363-3p | Upregulated in WM vs. MM | ||||
| miR-9-3p, miR-193b-3p, miR-182-5p, miR-152-3p | Downregulated in WM vs. MM | ||||
| miR-21-5p, miR-142-3p | Upregulated in WM vs. normal B-lineage cells | Diagnosis | |||
| miR-182-5p, miR-152-3p, miR-373-5p, miR-575-3p | Downregulated in WM vs. normal B-lineage cells | ||||
| Combination of miR-320a-3p and miR-320b-3p | Serum | Downregulated in WM vs. normal blood donors; downregulated in WM vs. MGUS and MM | Diagnosis, differential diagnosis | [151] | |
| miR-151-5p, let-7a-5p | Downregulated in WM vs. normal blood donors; downregulated in WM vs. MGUS | ||||
| miR-21-5p, miR-192-5p, miR-320b-3p | Exosomes | Increases with disease progression | Prediction of disease progression | [152] | |
| let-7d-5p | Decreases with disease progression | ||||
| HCL | miR-363-3p, miR-708-5p | Peripheral blood B cells | Downregulated in HCL vs. CLL | Differential diagnosis | [146] |
| miR-221-3p, miR-222-3p, miR-22-3p, miR-24-3p, miR-27a-3p, let-7b-5p | Peripheral blood CD19+ B cells | Upregulated in HCL vs. other B-cell malignancies; upregulated in HCL vs. normal B-lineage cells | Diagnosis, differential diagnosis | [147] | |
| PCFCL | miR-150-5p, miR-155-5p | FFPE tissues | Upregulated in PCFCL vs. cutaneous MZL | Differential diagnosis | [117] |
| miR-129-2-3p, miR-214-3p, miR-31-5p, miR-9-5p | Upregulated in PCFCL vs. primary cutaneous DLBCL-leg type | [148] |
6. Interplay between Cytokines and miRNAs in B-Cell Malignancies
7. Limitations
8. Future Perspectives
9. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Swerdlow, S.H.; Campo, E.; Pileri, S.A.; Harris, N.L.; Stein, H.; Siebert, R.; Advani, R.; Ghielmini, M.; Salles, G.A.; Zelenetz, A.D.; et al. The 2016 revision of the World Health Organization classification of lymphoid neoplasms. Blood 2016, 127, 2375–2390. [Google Scholar] [CrossRef]
- Armitage, J.O.; Gascoyne, R.D.; Lunning, M.A.; Cavalli, F. Non-Hodgkin lymphoma. Lancet 2017, 390, 298–310. [Google Scholar] [CrossRef]
- Freedman, A.; Jacobsen, E. Follicular lymphoma: 2020 update on diagnosis and management. Am. J. Hematol. 2020, 95, 316–327. [Google Scholar] [CrossRef]
- Carbone, A.; Roulland, S.; Gloghini, A.; Younes, A.; von Keudell, G.; Lopez-Guillermo, A.; Fitzgibbon, J. Follicular lymphoma. Nat. Rev. Dis. Primers 2019, 5, 83. [Google Scholar] [CrossRef]
- Sindel, A.; Al-Juhaishi, T.; Yazbeck, V. Marginal Zone Lymphoma: State-of-the-Art Treatment. Curr. Treat. Options Oncol. 2019, 20, 90. [Google Scholar] [CrossRef]
- Ha, M.; Kim, V.N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 2014, 15, 509–524. [Google Scholar] [CrossRef]
- Musilova, K.; Mraz, M. MicroRNAs in B-cell lymphomas: How a complex biology gets more complex. Leukemia 2015, 29, 1004–1017. [Google Scholar] [CrossRef] [PubMed]
- Katsaraki, K.; Karousi, P.; Artemaki, P.I.; Scorilas, A.; Pappa, V.; Kontos, C.K.; Papageorgiou, S.G. MicroRNAs: Tiny Regulators of Gene Expression with Pivotal Roles in Normal B-Cell Development and B-Cell Chronic Lymphocytic Leukemia. Cancers 2021, 13, 593. [Google Scholar] [CrossRef]
- Condrat, C.E.; Thompson, D.C.; Barbu, M.G.; Bugnar, O.L.; Boboc, A.; Cretoiu, D.; Suciu, N.; Cretoiu, S.M.; Voinea, S.C. miRNAs as Biomarkers in Disease: Latest Findings Regarding Their Role in Diagnosis and Prognosis. Cells 2020, 9, 276. [Google Scholar] [CrossRef] [PubMed]
- Melchers, F. Checkpoints that control B cell development. J. Clin. Investig. 2015, 125, 2203–2210. [Google Scholar] [CrossRef]
- Koralov, S.B.; Muljo, S.A.; Galler, G.R.; Krek, A.; Chakraborty, T.; Kanellopoulou, C.; Jensen, K.; Cobb, B.S.; Merkenschlager, M.; Rajewsky, N.; et al. Dicer ablation affects antibody diversity and cell survival in the B lymphocyte lineage. Cell 2008, 132, 860–874. [Google Scholar] [CrossRef]
- Brandl, A.; Daum, P.; Brenner, S.; Schulz, S.R.; Yap, D.Y.; Bosl, M.R.; Wittmann, J.; Schuh, W.; Jack, H.M. The microprocessor component, DGCR8, is essential for early B-cell development in mice. Eur. J. Immunol. 2016, 46, 2710–2718. [Google Scholar] [CrossRef] [PubMed]
- Spierings, D.C.; McGoldrick, D.; Hamilton-Easton, A.M.; Neale, G.; Murchison, E.P.; Hannon, G.J.; Green, D.R.; Withoff, S. Ordered progression of stage-specific miRNA profiles in the mouse B2 B-cell lineage. Blood 2011, 117, 5340–5349. [Google Scholar] [CrossRef]
- Xiao, C.; Calado, D.P.; Galler, G.; Thai, T.H.; Patterson, H.C.; Wang, J.; Rajewsky, N.; Bender, T.P. MiR-150 controls B cell differentiation by targeting the transcription factor c-Myb. Cell 2007, 131, 146–159. [Google Scholar] [CrossRef]
- Okuyama, K.; Ikawa, T.; Gentner, B.; Hozumi, K.; Harnprasopwat, R.; Lu, J.; Yamashita, R.; Ha, D.; Toyoshima, T.; Chanda, B.; et al. MicroRNA-126-mediated control of cell fate in B-cell myeloid progenitors as a potential alternative to transcriptional factors. Proc. Natl. Acad. Sci. USA 2013, 110, 13410–13415. [Google Scholar] [CrossRef]
- Chen, C.Z.; Li, L.; Lodish, H.F.; Bartel, D.P. MicroRNAs modulate hematopoietic lineage differentiation. Science 2004, 303, 83–86. [Google Scholar] [CrossRef]
- Hagman, J.; Lukin, K. Transcription factors drive B cell development. Curr. Opin. Immunol. 2006, 18, 127–134. [Google Scholar] [CrossRef]
- Medvedovic, J.; Ebert, A.; Tagoh, H.; Busslinger, M. Pax5: A master regulator of B cell development and leukemogenesis. Adv. Immunol. 2011, 111, 179–206. [Google Scholar] [CrossRef]
- Kwon, K.; Hutter, C.; Sun, Q.; Bilic, I.; Cobaleda, C.; Malin, S.; Busslinger, M. Instructive role of the transcription factor E2A in early B lymphopoiesis and germinal center B cell development. Immunity 2008, 28, 751–762. [Google Scholar] [CrossRef] [PubMed]
- Hagman, J.; Ramirez, J.; Lukin, K. B lymphocyte lineage specification, commitment and epigenetic control of transcription by early B cell factor 1. Curr. Top. Microbiol. Immunol. 2012, 356, 17–38. [Google Scholar] [CrossRef] [PubMed]
- Nechanitzky, R.; Akbas, D.; Scherer, S.; Gyory, I.; Hoyler, T.; Ramamoorthy, S.; Diefenbach, A.; Grosschedl, R. Transcription factor EBF1 is essential for the maintenance of B cell identity and prevention of alternative fates in committed cells. Nat. Immunol. 2013, 14, 867–875. [Google Scholar] [CrossRef]
- Hu, H.; Wang, B.; Borde, M.; Nardone, J.; Maika, S.; Allred, L.; Tucker, P.W.; Rao, A. Foxp1 is an essential transcriptional regulator of B cell development. Nat. Immunol. 2006, 7, 819–826. [Google Scholar] [CrossRef]
- Contreras, J.R.; Palanichamy, J.K.; Tran, T.M.; Fernando, T.R.; Rodriguez-Malave, N.I.; Goswami, N.; Arboleda, V.A.; Casero, D.; Rao, D.S. MicroRNA-146a modulates B-cell oncogenesis by regulating Egr1. Oncotarget 2015, 6, 11023–11037. [Google Scholar] [CrossRef][Green Version]
- Blume, J.; Zietara, N.; Witzlau, K.; Liu, Y.; Sanchez, O.O.; Puchalka, J.; Winter, S.J.; Kunze-Schumacher, H.; Saran, N.; Duber, S.; et al. miR-191 modulates B-cell development and targets transcription factors E2A, Foxp1, and Egr1. Eur. J. Immunol. 2019, 49, 121–132. [Google Scholar] [CrossRef]
- Mehta, A.; Mann, M.; Zhao, J.L.; Marinov, G.K.; Majumdar, D.; Garcia-Flores, Y.; Du, X.; Erikci, E.; Chowdhury, K.; Baltimore, D. The microRNA-212/132 cluster regulates B cell development by targeting Sox4. J. Exp. Med. 2015, 212, 1679–1692. [Google Scholar] [CrossRef]
- Zhou, B.; Wang, S.; Mayr, C.; Bartel, D.P.; Lodish, H.F. miR-150, a microRNA expressed in mature B and T cells, blocks early B cell development when expressed prematurely. Proc. Natl. Acad. Sci. USA 2007, 104, 7080–7085. [Google Scholar] [CrossRef]
- Kurkewich, J.L.; Bikorimana, E.; Nguyen, T.; Klopfenstein, N.; Zhang, H.; Hallas, W.M.; Stayback, G.; McDowell, M.A.; Dahl, R. The mirn23a microRNA cluster antagonizes B cell development. J. Leukoc. Biol. 2016, 100, 665–677. [Google Scholar] [CrossRef]
- Kong, K.Y.; Owens, K.S.; Rogers, J.H.; Mullenix, J.; Velu, C.S.; Grimes, H.L.; Dahl, R. MIR-23A microRNA cluster inhibits B-cell development. Exp. Hematol. 2010, 38, 629–640.e1. [Google Scholar] [CrossRef]
- Lai, M.; Gonzalez-Martin, A.; Cooper, A.B.; Oda, H.; Jin, H.Y.; Shepherd, J.; He, L.; Zhu, J.; Nemazee, D.; Xiao, C. Regulation of B-cell development and tolerance by different members of the miR-17 approximately 92 family microRNAs. Nat. Commun. 2016, 7, 12207. [Google Scholar] [CrossRef]
- Benhamou, D.; Labi, V.; Getahun, A.; Benchetrit, E.; Dowery, R.; Rajewsky, K.; Cambier, J.C.; Melamed, D. The c-Myc/miR17-92/PTEN Axis Tunes PI3K Activity to Control Expression of Recombination Activating Genes in Early B Cell Development. Front. Immunol. 2018, 9, 2715. [Google Scholar] [CrossRef]
- Xiao, C.; Srinivasan, L.; Calado, D.P.; Patterson, H.C.; Zhang, B.; Wang, J.; Henderson, J.M.; Kutok, J.L.; Rajewsky, K. Lymphoproliferative disease and autoimmunity in mice with increased miR-17-92 expression in lymphocytes. Nat. Immunol. 2008, 9, 405–414. [Google Scholar] [CrossRef]
- Benhamou, D.; Labi, V.; Novak, R.; Dai, I.; Shafir-Alon, S.; Weiss, A.; Gaujoux, R.; Arnold, R.; Shen-Orr, S.S.; Rajewsky, K.; et al. A c-Myc/miR17-92/Pten Axis Controls PI3K-Mediated Positive and Negative Selection in B Cell Development and Reconstitutes CD19 Deficiency. Cell Rep. 2016, 16, 419–431. [Google Scholar] [CrossRef]
- King, J.K.; Ung, N.M.; Paing, M.H.; Contreras, J.R.; Alberti, M.O.; Fernando, T.R.; Zhang, K.; Pellegrini, M.; Rao, D.S. Regulation of Marginal Zone B-Cell Differentiation by MicroRNA-146a. Front. Immunol. 2016, 7, 670. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Pear, W.S. Notch signalling in B cells. Semin Cell Dev. Biol. 2003, 14, 135–142. [Google Scholar] [CrossRef]
- Luo, Z.; Mu, L.; Zheng, Y.; Shen, W.; Li, J.; Xu, L.; Zhong, B.; Liu, Y.; Zhou, Y. NUMB enhances Notch signaling by repressing ubiquitination of NOTCH1 intracellular domain. J. Mol. Cell Biol. 2020, 12, 345–358. [Google Scholar] [CrossRef]
- Kramer, N.J.; Wang, W.L.; Reyes, E.Y.; Kumar, B.; Chen, C.C.; Ramakrishna, C.; Cantin, E.M.; Vonderfecht, S.L.; Taganov, K.D.; Chau, N.; et al. Altered lymphopoiesis and immunodeficiency in miR-142 null mice. Blood 2015, 125, 3720–3730. [Google Scholar] [CrossRef]
- De Yebenes, V.G.; Belver, L.; Pisano, D.G.; Gonzalez, S.; Villasante, A.; Croce, C.; He, L.; Ramiro, A.R. miR-181b negatively regulates activation-induced cytidine deaminase in B cells. J. Exp. Med. 2008, 205, 2199–2206. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.; Shan, X.; Wang, T.; Shu, Y.; Liu, P. miR-181b modulates multidrug resistance by targeting BCL2 in human cancer cell lines. Int. J. Cancer 2010, 127, 2520–2529. [Google Scholar] [CrossRef] [PubMed]
- Gururajan, M.; Haga, C.L.; Das, S.; Leu, C.M.; Hodson, D.; Josson, S.; Turner, M.; Cooper, M.D. MicroRNA 125b inhibition of B cell differentiation in germinal centers. Int. Immunol. 2010, 22, 583–592. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; So, A.Y.; Sookram, R.; Wong, S.; Wang, J.K.; Ouyang, Y.; He, P.; Su, Y.; Casellas, R.; Baltimore, D. Epigenetic silencing of miR-125b is required for normal B-cell development. Blood 2018, 131, 1920–1930. [Google Scholar] [CrossRef]
- Enomoto, Y.; Kitaura, J.; Hatakeyama, K.; Watanuki, J.; Akasaka, T.; Kato, N.; Shimanuki, M.; Nishimura, K.; Takahashi, M.; Taniwaki, M.; et al. Emu/miR-125b transgenic mice develop lethal B-cell malignancies. Leukemia 2011, 25, 1849–1856. [Google Scholar] [CrossRef]
- Zhang, J.; Jima, D.D.; Jacobs, C.; Fischer, R.; Gottwein, E.; Huang, G.; Lugar, P.L.; Lagoo, A.S.; Rizzieri, D.A.; Friedman, D.R.; et al. Patterns of microRNA expression characterize stages of human B-cell differentiation. Blood 2009, 113, 4586–4594. [Google Scholar] [CrossRef] [PubMed]
- Porstner, M.; Winkelmann, R.; Daum, P.; Schmid, J.; Pracht, K.; Corte-Real, J.; Schreiber, S.; Haftmann, C.; Brandl, A.; Mashreghi, M.F.; et al. miR-148a promotes plasma cell differentiation and targets the germinal center transcription factors Mitf and Bach2. Eur. J. Immunol. 2015, 45, 1206–1215. [Google Scholar] [CrossRef] [PubMed]
- Lu, D.; Nakagawa, R.; Lazzaro, S.; Staudacher, P.; Abreu-Goodger, C.; Henley, T.; Boiani, S.; Leyland, R.; Galloway, A.; Andrews, S.; et al. The miR-155-PU.1 axis acts on Pax5 to enable efficient terminal B cell differentiation. J. Exp. Med. 2014, 211, 2183–2198. [Google Scholar] [CrossRef] [PubMed]
- Bouska, A.; Zhang, W.; Gong, Q.; Iqbal, J.; Scuto, A.; Vose, J.; Ludvigsen, M.; Fu, K.; Weisenburger, D.D.; Greiner, T.C.; et al. Combined copy number and mutation analysis identifies oncogenic pathways associated with transformation of follicular lymphoma. Leukemia 2017, 31, 83–91. [Google Scholar] [CrossRef]
- Fassina, A.; Marino, F.; Siri, M.; Zambello, R.; Ventura, L.; Fassan, M.; Simonato, F.; Cappellesso, R. The miR-17-92 microRNA cluster: A novel diagnostic tool in large B-cell malignancies. Lab. Invest. 2012, 92, 1574–1582. [Google Scholar] [CrossRef]
- Devan, J.; Janikova, A.; Mraz, M. New concepts in follicular lymphoma biology: From BCL2 to epigenetic regulators and non-coding RNAs. Semin. Oncol. 2018, 45, 291–302. [Google Scholar] [CrossRef]
- Leich, E.; Zamo, A.; Horn, H.; Haralambieva, E.; Puppe, B.; Gascoyne, R.D.; Chan, W.C.; Braziel, R.M.; Rimsza, L.M.; Weisenburger, D.D.; et al. MicroRNA profiles of t(14;18)-negative follicular lymphoma support a late germinal center B-cell phenotype. Blood 2011, 118, 5550–5558. [Google Scholar] [CrossRef]
- Wagner, S.D.; Ahearne, M.; Ko Ferrigno, P. The role of BCL6 in lymphomas and routes to therapy. Br. J. Haematol. 2011, 152, 3–12. [Google Scholar] [CrossRef]
- Braga, T.V.; Evangelista, F.C.G.; Gomes, L.C.; Araujo, S.; Carvalho, M.D.G.; Sabino, A.P. Evaluation of MiR-15a and MiR-16-1 as prognostic biomarkers in chronic lymphocytic leukemia. Biomed. Pharmacother. 2017, 92, 864–869. [Google Scholar] [CrossRef] [PubMed]
- Gebauer, N.; Gollub, W.; Stassek, B.; Bernard, V.; Rades, D.; Feller, A.C.; Thorns, C. MicroRNA signatures in subtypes of follicular lymphoma. Anticancer Res. 2014, 34, 2105–2111. [Google Scholar]
- Lu, J.; Getz, G.; Miska, E.A.; Alvarez-Saavedra, E.; Lamb, J.; Peck, D.; Sweet-Cordero, A.; Ebert, B.L.; Mak, R.H.; Ferrando, A.A.; et al. MicroRNA expression profiles classify human cancers. Nature 2005, 435, 834–838. [Google Scholar] [CrossRef]
- Lawrie, C.H.; Chi, J.; Taylor, S.; Tramonti, D.; Ballabio, E.; Palazzo, S.; Saunders, N.J.; Pezzella, F.; Boultwood, J.; Wainscoat, J.S.; et al. Expression of microRNAs in diffuse large B cell lymphoma is associated with immunophenotype, survival and transformation from follicular lymphoma. J. Cell. Mol. Med. 2009, 13, 1248–1260. [Google Scholar] [CrossRef]
- Roehle, A.; Hoefig, K.P.; Repsilber, D.; Thorns, C.; Ziepert, M.; Wesche, K.O.; Thiere, M.; Loeffler, M.; Klapper, W.; Pfreundschuh, M.; et al. MicroRNA signatures characterize diffuse large B-cell lymphomas and follicular lymphomas. Br. J. Haematol. 2008, 142, 732–744. [Google Scholar] [CrossRef]
- Psathas, J.N.; Doonan, P.J.; Raman, P.; Freedman, B.D.; Minn, A.J.; Thomas-Tikhonenko, A. The Myc-miR-17-92 axis amplifies B-cell receptor signaling via inhibition of ITIM proteins: A novel lymphomagenic feed-forward loop. Blood 2013, 122, 4220–4229. [Google Scholar] [CrossRef]
- Olive, V.; Bennett, M.J.; Walker, J.C.; Ma, C.; Jiang, I.; Cordon-Cardo, C.; Li, Q.J.; Lowe, S.W.; Hannon, G.J.; He, L. miR-19 is a key oncogenic component of mir-17-92. Genes Dev. 2009, 23, 2839–2849. [Google Scholar] [CrossRef]
- Filip, D.; Mraz, M. The role of MYC in the transformation and aggressiveness of ‘indolent’ B-cell malignancies. Leuk. Lymphoma 2020, 61, 510–524. [Google Scholar] [CrossRef]
- Yan, S.; Jia, C.; Quan, L.; Zhao, L.; Tian, Y.; Liu, A. Significance of the microRNA1792 gene cluster expressed in Bcell nonHodgkin’s lymphoma. Mol. Med. Rep. 2019, 20, 2459–2467. [Google Scholar] [CrossRef] [PubMed]
- Musilova, K.; Devan, J.; Cerna, K.; Seda, V.; Pavlasova, G.; Sharma, S.; Oppelt, J.; Pytlik, R.; Prochazka, V.; Prouzova, Z.; et al. miR-150 downregulation contributes to the high-grade transformation of follicular lymphoma by upregulating FOXP1 levels. Blood 2018, 132, 2389–2400. [Google Scholar] [CrossRef]
- Jardin, F.; Jais, J.P.; Molina, T.J.; Parmentier, F.; Picquenot, J.M.; Ruminy, P.; Tilly, H.; Bastard, C.; Salles, G.A.; Feugier, P.; et al. Diffuse large B-cell lymphomas with CDKN2A deletion have a distinct gene expression signature and a poor prognosis under R-CHOP treatment: A GELA study. Blood 2010, 116, 1092–1104. [Google Scholar] [CrossRef]
- Yu, T.; Ma, P.; Wu, D.; Shu, Y.; Gao, W. Functions and mechanisms of microRNA-31 in human cancers. Biomed. Pharmacother. 2018, 108, 1162–1169. [Google Scholar] [CrossRef] [PubMed]
- Thompson, M.A.; Edmonds, M.D.; Liang, S.; McClintock-Treep, S.; Wang, X.; Li, S.; Eischen, C.M. miR-31 and miR-17-5p levels change during transformation of follicular lymphoma. Hum. Pathol. 2016, 50, 118–126. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Corrigan-Cummins, M.; Hudson, J.; Maric, I.; Simakova, O.; Neelapu, S.S.; Kwak, L.W.; Janik, J.E.; Gause, B.; Jaffe, E.S.; et al. MicroRNA profiling of follicular lymphoma identifies microRNAs related to cell proliferation and tumor response. Haematologica 2012, 97, 586–594. [Google Scholar] [CrossRef] [PubMed]
- Hoffman, A.E.; Liu, R.; Fu, A.; Zheng, T.; Slack, F.; Zhu, Y. Targetome profiling, pathway analysis and genetic association study implicate miR-202 in lymphomagenesis. Cancer Epidemiol. Biomark. Prev. 2013, 22, 327–336. [Google Scholar] [CrossRef] [PubMed]
- Fu, A.; Hoffman, A.E.; Liu, R.; Jacobs, D.I.; Zheng, T.; Zhu, Y. Targetome profiling and functional genetics implicate miR-618 in lymphomagenesis. Epigenetics 2014, 9, 730–737. [Google Scholar] [CrossRef]
- Lin, J.; Lwin, T.; Zhao, J.J.; Tam, W.; Choi, Y.S.; Moscinski, L.C.; Dalton, W.S.; Sotomayor, E.M.; Wright, K.L.; Tao, J. Follicular dendritic cell-induced microRNA-mediated upregulation of PRDM1 and downregulation of BCL-6 in non-Hodgkin’s B-cell lymphomas. Leukemia 2011, 25, 145–152. [Google Scholar] [CrossRef]
- Raju, S.; Kretzmer, L.Z.; Koues, O.I.; Payton, J.E.; Oltz, E.M.; Cashen, A.; Polic, B.; Schreiber, R.D.; Shaw, A.S.; Markiewicz, M.A. NKG2D-NKG2D Ligand Interaction Inhibits the Outgrowth of Naturally Arising Low-Grade B Cell Lymphoma In Vivo. J. Immunol. 2016, 196, 4805–4813. [Google Scholar] [CrossRef]
- Craig, V.J.; Cogliatti, S.B.; Imig, J.; Renner, C.; Neuenschwander, S.; Rehrauer, H.; Schlapbach, R.; Dirnhofer, S.; Tzankov, A.; Muller, A. Myc-mediated repression of microRNA-34a promotes high-grade transformation of B-cell lymphoma by dysregulation of FoxP1. Blood 2011, 117, 6227–6236. [Google Scholar] [CrossRef]
- Chang, T.C.; Yu, D.; Lee, Y.S.; Wentzel, E.A.; Arking, D.E.; West, K.M.; Dang, C.V.; Thomas-Tikhonenko, A.; Mendell, J.T. Widespread microRNA repression by Myc contributes to tumorigenesis. Nat. Genet. 2008, 40, 43–50. [Google Scholar] [CrossRef] [PubMed]
- Sagaert, X.; de Paepe, P.; Libbrecht, L.; Vanhentenrijk, V.; Verhoef, G.; Thomas, J.; Wlodarska, I.; de Wolf-Peeters, C. Forkhead box protein P1 expression in mucosa-associated lymphoid tissue lymphomas predicts poor prognosis and transformation to diffuse large B-cell lymphoma. J. Clin. Oncol. 2006, 24, 2490–2497. [Google Scholar] [CrossRef]
- He, M.; Gao, L.; Zhang, S.; Tao, L.; Wang, J.; Yang, J.; Zhu, M. Prognostic significance of miR-34a and its target proteins of FOXP1, p53, and BCL2 in gastric MALT lymphoma and DLBCL. Gastric Cancer 2014, 17, 431–441. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Jiang, K.; Su, S.; Wang, B.; Chen, G. Clinical manifestations and epigenetic mechanisms of gastric mucosa associated lymphoid tissue lymphoma and long-term follow-up following Helicobacter pylori eradication. Exp. Ther. Med. 2018, 15, 553–561. [Google Scholar] [CrossRef] [PubMed]
- Craig, V.J.; Cogliatti, S.B.; Rehrauer, H.; Wundisch, T.; Muller, A. Epigenetic silencing of microRNA-203 dysregulates ABL1 expression and drives Helicobacter-associated gastric lymphomagenesis. Cancer Res. 2011, 71, 3616–3624. [Google Scholar] [CrossRef]
- Fernandez, C.; Bellosillo, B.; Ferraro, M.; Seoane, A.; Sanchez-Gonzalez, B.; Pairet, S.; Pons, A.; Barranco, L.; Vela, M.C.; Gimeno, E.; et al. MicroRNAs 142-3p, miR-155 and miR-203 are Deregulated in Gastric MALT Lymphomas Compared to Chronic Gastritis. Cancer Genom. Proteom. 2017, 14, 75–82. [Google Scholar] [CrossRef][Green Version]
- Saito, Y.; Suzuki, H.; Tsugawa, H.; Imaeda, H.; Matsuzaki, J.; Hirata, K.; Hosoe, N.; Nakamura, M.; Mukai, M.; Saito, H.; et al. Overexpression of miR-142-5p and miR-155 in gastric mucosa-associated lymphoid tissue (MALT) lymphoma resistant to Helicobacter pylori eradication. PLoS ONE 2012, 7, e47396. [Google Scholar] [CrossRef]
- Blosse, A.; Levy, M.; Robe, C.; Staedel, C.; Copie-Bergman, C.; Lehours, P. Deregulation of miRNA in Helicobacter pylori-Induced Gastric MALT Lymphoma: From Mice to Human. J. Clin. Med. 2019, 8, 845. [Google Scholar] [CrossRef]
- Peveling-Oberhag, J.; Crisman, G.; Schmidt, A.; Doring, C.; Lucioni, M.; Arcaini, L.; Rattotti, S.; Hartmann, S.; Piiper, A.; Hofmann, W.P.; et al. Dysregulation of global microRNA expression in splenic marginal zone lymphoma and influence of chronic hepatitis C virus infection. Leukemia 2012, 26, 1654–1662. [Google Scholar] [CrossRef]
- Fry, A.M.; O’Regan, L.; Sabir, S.R.; Bayliss, R. Cell cycle regulation by the NEK family of protein kinases. J. Cell Sci. 2012, 125, 4423–4433. [Google Scholar] [CrossRef] [PubMed]
- Roccaro, A.M.; Sacco, A.; Jia, X.; Azab, A.K.; Maiso, P.; Ngo, H.T.; Azab, F.; Runnels, J.; Quang, P.; Ghobrial, I.M. microRNA-dependent modulation of histone acetylation in Waldenstrom macroglobulinemia. Blood 2010, 116, 1506–1514. [Google Scholar] [CrossRef] [PubMed]
- Fulciniti, M.; Amodio, N.; Bandi, R.L.; Cagnetta, A.; Samur, M.K.; Acharya, C.; Prabhala, R.; D’Aquila, P.; Bellizzi, D.; Passarino, G.; et al. miR-23b/SP1/c-myc forms a feed-forward loop supporting multiple myeloma cell growth. Blood Cancer J. 2016, 6, e380. [Google Scholar] [CrossRef]
- Roccaro, A.M.; Sacco, A.; Chen, C.; Runnels, J.; Leleu, X.; Azab, F.; Azab, A.K.; Jia, X.; Ngo, H.T.; Melhem, M.R.; et al. microRNA expression in the biology, prognosis, and therapy of Waldenstrom macroglobulinemia. Blood 2009, 113, 4391–4402. [Google Scholar] [CrossRef]
- Roccaro, A.M.; Sacco, A.; Jia, X.; Banwait, R.; Maiso, P.; Azab, F.; Flores, L.; Manier, S.; Azab, A.K.; Ghobrial, I.M. Mechanisms of activity of the TORC1 inhibitor everolimus in Waldenstrom macroglobulinemia. Clin. Cancer Res. 2012, 18, 6609–6622. [Google Scholar] [CrossRef]
- Gaudette, B.T.; Dwivedi, B.; Chitta, K.S.; Poulain, S.; Powell, D.; Vertino, P.; Leleu, X.; Lonial, S.; Chanan-Khan, A.A.; Kowalski, J.; et al. Low expression of pro-apoptotic Bcl-2 family proteins sets the apoptotic threshold in Waldenstrom macroglobulinemia. Oncogene 2016, 35, 479–490. [Google Scholar] [CrossRef] [PubMed]
- Bradshaw, G.; Sutherland, H.G.; Haupt, L.M.; Griffiths, L.R. Dysregulated MicroRNA Expression Profiles and Potential Cellular, Circulating and Polymorphic Biomarkers in Non-Hodgkin Lymphoma. Genes 2016, 7, 130. [Google Scholar] [CrossRef] [PubMed]
- Capuano, C.; Pighi, C.; Maggio, R.; Battella, S.; Morrone, S.; Palmieri, G.; Santoni, A.; Klein, C.; Galandrini, R. CD16 pre-ligation by defucosylated tumor-targeting mAb sensitizes human NK cells to gammac cytokine stimulation via PI3K/mTOR axis. Cancer Immunol. Immunother. 2020, 69, 501–512. [Google Scholar] [CrossRef]
- Battella, S.; Cox, M.C.; Santoni, A.; Palmieri, G. Natural killer (NK) cells and anti-tumor therapeutic mAb: Unexplored interactions. J. Leukoc. Biol. 2016, 99, 87–96. [Google Scholar] [CrossRef]
- Culpin, R.E.; Proctor, S.J.; Angus, B.; Crosier, S.; Anderson, J.J.; Mainou-Fowler, T. A 9 series microRNA signature differentiates between germinal centre and activated B-cell-like diffuse large B-cell lymphoma cell lines. Int. J. Oncol. 2010, 37, 367–376. [Google Scholar] [CrossRef]
- Malpeli, G.; Barbi, S.; Greco, C.; Zupo, S.; Bertolaso, A.; Scupoli, M.T.; Krampera, M.; Kamga, P.T.; Croce, C.M.; Scarpa, A.; et al. MicroRNA signatures and Foxp3(+) cell count correlate with relapse occurrence in follicular lymphoma. Oncotarget 2018, 9, 19961–19979. [Google Scholar] [CrossRef][Green Version]
- Sinkovics, J.G. Molecular biology of oncogenic inflammatory processes. I. Non-oncogenic and oncogenic pathogens, intrinsic inflammatory reactions without pathogens, and microRNA/DNA interactions (Review). Int. J. Oncol. 2012, 40, 305–349. [Google Scholar] [CrossRef]
- Ruskone-Fourmestraux, A.; Lavergne, A.; Aegerter, P.H.; Megraud, F.; Palazzo, L.; de Mascarel, A.; Molina, T.; Rambaud, J.L. Predictive factors for regression of gastric MALT lymphoma after anti-Helicobacter pylori treatment. Gut 2001, 48, 297–303. [Google Scholar] [CrossRef]
- Tzioufas, A.G.; Kapsogeorgou, E.K.; Moutsopoulos, H.M. Pathogenesis of Sjogren’s syndrome: What we know and what we should learn. J. Autoimmun. 2012, 39, 4–8. [Google Scholar] [CrossRef] [PubMed]
- Steinbach, G.; Ford, R.; Glober, G.; Sample, D.; Hagemeister, F.B.; Lynch, P.M.; McLaughlin, P.W.; Rodriguez, M.A.; Romaguera, J.E.; Sarris, A.H.; et al. Antibiotic treatment of gastric lymphoma of mucosa-associated lymphoid tissue. An uncontrolled trial. Ann. Intern. Med. 1999, 131, 88–95. [Google Scholar] [CrossRef]
- Morgner, A.; Miehlke, S.; Fischbach, W.; Schmitt, W.; Muller-Hermelink, H.; Greiner, A.; Thiede, C.; Schetelig, J.; Neubauer, A.; Stolte, M.; et al. Complete remission of primary high-grade B-cell gastric lymphoma after cure of Helicobacter pylori infection. J. Clin. Oncol. 2001, 19, 2041–2048. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Ye, H.; Ruskone-Fourmestraux, A.; de Jong, D.; Pileri, S.; Thiede, C.; Lavergne, A.; Boot, H.; Caletti, G.; Wundisch, T.; et al. T(11;18) is a marker for all stage gastric MALT lymphomas that will not respond to H. pylori eradication. Gastroenterology 2002, 122, 1286–1294. [Google Scholar] [CrossRef] [PubMed]
- Akagi, T.; Motegi, M.; Tamura, A.; Hosokawa, Y.; Suzuki, H.; Ota, H.; Nakamura, S.; Morishima, Y.; Taniwaki, M.; Seto, M. A novel gene, MALT1 at 18q21, is involved in t(11;18) (q21;q21) found in low-grade B-cell lymphoma of mucosa-associated lymphoid tissue. Oncogene 1999, 18, 5785–5794. [Google Scholar] [CrossRef]
- Hosokawa, Y.; Suzuki, H.; Suzuki, Y.; Takahashi, R.; Seto, M. Antiapoptotic function of apoptosis inhibitor 2-MALT1 fusion protein involved in t(11;18)(q21;q21) mucosa-associated lymphoid tissue lymphoma. Cancer Res. 2004, 64, 3452–3457. [Google Scholar] [CrossRef][Green Version]
- Rosebeck, S.; Lucas, P.C.; McAllister-Lucas, L.M. Protease activity of the API2-MALT1 fusion oncoprotein in MALT lymphoma development and treatment. Future Oncol. 2011, 7, 613–617. [Google Scholar] [CrossRef] [PubMed]
- Tomasini, R.; Seux, M.; Nowak, J.; Bontemps, C.; Carrier, A.; Dagorn, J.C.; Pebusque, M.J.; Iovanna, J.L.; Dusetti, N.J. TP53INP1 is a novel p73 target gene that induces cell cycle arrest and cell death by modulating p73 transcriptional activity. Oncogene 2005, 24, 8093–8104. [Google Scholar] [CrossRef]
- Dai, Y.H.; Tang, Y.P.; Zhu, H.Y.; Lv, L.; Chu, Y.; Zhou, Y.Q.; Huo, J.R. ZEB2 promotes the metastasis of gastric cancer and modulates epithelial mesenchymal transition of gastric cancer cells. Dig. Dis. Sci. 2012, 57, 1253–1260. [Google Scholar] [CrossRef] [PubMed]
- Gebauer, N.; Kuba, J.; Senft, A.; Schillert, A.; Bernard, V.; Thorns, C. MicroRNA-150 is up-regulated in extranodal marginal zone lymphoma of MALT type. Cancer Genom. Proteom. 2014, 11, 51–56. [Google Scholar]
- Wu, Q.; Jin, H.; Yang, Z.; Luo, G.; Lu, Y.; Li, K.; Ren, G.; Su, T.; Pan, Y.; Feng, B.; et al. MiR-150 promotes gastric cancer proliferation by negatively regulating the pro-apoptotic gene EGR2. Biochem. Biophys. Res. Commun. 2010, 392, 340–345. [Google Scholar] [CrossRef] [PubMed]
- Young, E.; Noerenberg, D.; Mansouri, L.; Ljungstrom, V.; Frick, M.; Sutton, L.A.; Blakemore, S.J.; Galan-Sousa, J.; Plevova, K.; Baliakas, P.; et al. EGR2 mutations define a new clinically aggressive subgroup of chronic lymphocytic leukemia. Leukemia 2017, 31, 1547–1554. [Google Scholar] [CrossRef] [PubMed]
- Krysiak, K.; Gomez, F.; White, B.S.; Matlock, M.; Miller, C.A.; Trani, L.; Fronick, C.C.; Fulton, R.S.; Kreisel, F.; Cashen, A.F.; et al. Recurrent somatic mutations affecting B-cell receptor signaling pathway genes in follicular lymphoma. Blood 2017, 129, 473–483. [Google Scholar] [CrossRef]
- Sun, M.; Liu, X.H.; Li, J.H.; Yang, J.S.; Zhang, E.B.; Yin, D.D.; Liu, Z.L.; Zhou, J.; Ding, Y.; Li, S.Q.; et al. MiR-196a is upregulated in gastric cancer and promotes cell proliferation by downregulating p27(kip1). Mol. Cancer Ther. 2012, 11, 842–852. [Google Scholar] [CrossRef]
- Yuan, Y.; Du, W.; Wang, Y.; Xu, C.; Wang, J.; Zhang, Y.; Wang, H.; Ju, J.; Zhao, L.; Wang, Z.; et al. Suppression of AKT expression by miR-153 produced anti-tumor activity in lung cancer. Int. J. Cancer 2015, 136, 1333–1340. [Google Scholar] [CrossRef]
- Xie, J.; Chen, M.; Zhou, J.; Mo, M.S.; Zhu, L.H.; Liu, Y.P.; Gui, Q.J.; Zhang, L.; Li, G.Q. miR-7 inhibits the invasion and metastasis of gastric cancer cells by suppressing epidermal growth factor receptor expression. Oncol. Rep. 2014, 31, 1715–1722. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Dou, W.; He, L.; Liang, S.; Tie, J.; Liu, C.; Li, T.; Lu, Y.; Mo, P.; Shi, Y.; et al. MicroRNA-7 functions as an anti-metastatic microRNA in gastric cancer by targeting insulin-like growth factor-1 receptor. Oncogene 2013, 32, 1363–1372. [Google Scholar] [CrossRef] [PubMed]
- Pang, L.; Li, B.; Zheng, B.; Niu, L.; Ge, L. miR-138 inhibits gastric cancer growth by suppressing SOX4. Oncol. Rep. 2017, 38, 1295–1302. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Suo, A.L.; Li, Z.F.; Liu, L.Y.; Tian, T.; Ni, L.; Zhang, W.G.; Nan, K.J.; Song, T.S.; Huang, C. MicroRNA profiling of human gastric cancer. Mol. Med. Rep. 2009, 2, 963–970. [Google Scholar] [CrossRef] [PubMed]
- Rao, D.S.; O’Connell, R.M.; Chaudhuri, A.A.; Garcia-Flores, Y.; Geiger, T.L.; Baltimore, D. MicroRNA-34a perturbs B lymphocyte development by repressing the forkhead box transcription factor Foxp1. Immunity 2010, 33, 48–59. [Google Scholar] [CrossRef]
- Streubel, B.; Vinatzer, U.; Lamprecht, A.; Raderer, M.; Chott, A. T(3;14)(p14.1;q32) involving IGH and FOXP1 is a novel recurrent chromosomal aberration in MALT lymphoma. Leukemia 2005, 19, 652–658. [Google Scholar] [CrossRef] [PubMed]
- Goatly, A.; Bacon, C.M.; Nakamura, S.; Ye, H.; Kim, I.; Brown, P.J.; Ruskone-Fourmestraux, A.; Cervera, P.; Streubel, B.; Banham, A.H.; et al. FOXP1 abnormalities in lymphoma: Translocation breakpoint mapping reveals insights into deregulated transcriptional control. Mod. Pathol. 2008, 21, 902–911. [Google Scholar] [CrossRef] [PubMed]
- Gu, S.; Wang, X.; Feng, L.; Xie, Y.; Ping, L.; Liu, W.; Li, J.; Liu, Y.; Ding, N.; Song, Y.; et al. Prognostic role of serum miRNA-16 in primary gastric lymphoma. Int. J. Clin. Exp. Pathol. 2018, 11, 5430–5435. [Google Scholar] [PubMed]
- Cimmino, A.; Calin, G.A.; Fabbri, M.; Iorio, M.V.; Ferracin, M.; Shimizu, M.; Wojcik, S.E.; Aqeilan, R.I.; Zupo, S.; Dono, M.; et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc. Natl. Acad. Sci. USA 2005, 102, 13944–13949. [Google Scholar] [CrossRef] [PubMed]
- Cerroni, L.; Zochling, N.; Putz, B.; Kerl, H. Infection by Borrelia burgdorferi and cutaneous B-cell lymphoma. J. Cutan. Pathol. 1997, 24, 457–461. [Google Scholar] [CrossRef]
- Bogle, M.A.; Riddle, C.C.; Triana, E.M.; Jones, D.; Duvic, M. Primary cutaneous B-cell lymphoma. J. Am. Acad. Dermatol. 2005, 53, 479–484. [Google Scholar] [CrossRef] [PubMed]
- Monsalvez, V.; Montes-Moreno, S.; Artiga, M.J.; Rodriguez, M.E.; Sanchez-Espiridion, B.; Lozano, M.; Fernandez-de-Misa, R.; Rodriguez-Peralto, J.L.; Piris, M.A.; Ortiz-Romero, P.L. MicroRNAs as prognostic markers in indolent primary cutaneous B-cell lymphoma. Mod. Pathol. 2013, 26, 171–181. [Google Scholar] [CrossRef]
- Hother, C.; Rasmussen, P.K.; Joshi, T.; Reker, D.; Ralfkiaer, U.; Workman, C.T.; Heegaard, S.; Ralfkiaer, E.; Gronbaek, K. MicroRNA profiling in ocular adnexal lymphoma: A role for MYC and NFKB1 mediated dysregulation of microRNA expression in aggressive disease. Investig. Ophthalmol. Vis. Sci. 2013, 54, 5169–5175. [Google Scholar] [CrossRef] [PubMed]
- Kwon, J.J.; Factora, T.D.; Dey, S.; Kota, J. A Systematic Review of miR-29 in Cancer. Mol. Ther. Oncolytics 2019, 12, 173–194. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Ye, B.; Wang, P.; Yao, F.; Zhang, C.; Yu, G. Overview of microRNA-199a Regulation in Cancer. Cancer Manag. Res. 2019, 11, 10327–10335. [Google Scholar] [CrossRef]
- Troppan, K.; Wenzl, K.; Pichler, M.; Pursche, B.; Schwarzenbacher, D.; Feichtinger, J.; Thallinger, G.G.; Beham-Schmid, C.; Neumeister, P.; Deutsch, A. miR-199a and miR-497 are Associated with Better Overall Survival due to Increased Chemosensitivity in Diffuse Large B-Cell Lymphoma Patients. Int. J. Mol. Sci. 2015, 16, 18077–18095. [Google Scholar] [CrossRef]
- Alunno, A.; Leone, M.C.; Giacomelli, R.; Gerli, R.; Carubbi, F. Lymphoma and Lymphomagenesis in Primary Sjogren’s Syndrome. Front. Med. 2018, 5, 102. [Google Scholar] [CrossRef]
- Gourzi, V.C.; Kapsogeorgou, E.K.; Kyriakidis, N.C.; Tzioufas, A.G. Study of microRNAs (miRNAs) that are predicted to target the autoantigens Ro/SSA and La/SSB in primary Sjogren’s Syndrome. Clin. Exp. Immunol. 2015, 182, 14–22. [Google Scholar] [CrossRef] [PubMed]
- Kapsogeorgou, E.K.; Papageorgiou, A.; Protogerou, A.D.; Voulgarelis, M.; Tzioufas, A.G. Low miR200b-5p levels in minor salivary glands: A novel molecular marker predicting lymphoma development in patients with Sjogren’s syndrome. Ann. Rheum. Dis. 2018, 77, 1200–1207. [Google Scholar] [CrossRef]
- Rhodes, L.V.; Martin, E.C.; Segar, H.C.; Miller, D.F.; Buechlein, A.; Rusch, D.B.; Nephew, K.P.; Burow, M.E.; Collins-Burow, B.M. Dual regulation by microRNA-200b-3p and microRNA-200b-5p in the inhibition of epithelial-to-mesenchymal transition in triple-negative breast cancer. Oncotarget 2015, 6, 16638–16652. [Google Scholar] [CrossRef] [PubMed]
- Robinson, J.E.; Cutucache, C.E. Deciphering splenic marginal zone lymphoma pathogenesis: The proposed role of microRNA. Oncotarget 2018, 9, 30005–30022. [Google Scholar] [CrossRef]
- Mateo, M.; Mollejo, M.; Villuendas, R.; Algara, P.; Sanchez-Beato, M.; Martínez, P.; Piris, M.A. 7q31-32 Allelic Loss is a Frequent Finding in Splenic Marginal Zone Lymphoma. Am. J. Pathol. 1999, 154, 1583–1589. [Google Scholar] [CrossRef]
- Watkins, A.J.; Hamoudi, R.A.; Zeng, N.; Yan, Q.; Huang, Y.; Liu, H.; Zhang, J.; Braggio, E.; Fonseca, R.; de Leval, L.; et al. An integrated genomic and expression analysis of 7q deletion in splenic marginal zone lymphoma. PLoS ONE 2012, 7, e44997. [Google Scholar] [CrossRef]
- Ruiz-Ballesteros, E.; Mollejo, M.; Rodriguez, A.; Camacho, F.I.; Algara, P.; Martinez, N.; Pollan, M.; Sanchez-Aguilera, A.; Menarguez, J.; Campo, E.; et al. Splenic marginal zone lymphoma: Proposal of new diagnostic and prognostic markers identified after tissue and cDNA microarray analysis. Blood 2005, 106, 1831–1838. [Google Scholar] [CrossRef] [PubMed]
- Pekarsky, Y.; Santanam, U.; Cimmino, A.; Palamarchuk, A.; Efanov, A.; Maximov, V.; Volinia, S.; Alder, H.; Liu, C.G.; Rassenti, L.; et al. Tcl1 expression in chronic lymphocytic leukemia is regulated by miR-29 and miR-181. Cancer Res. 2006, 66, 11590–11593. [Google Scholar] [CrossRef] [PubMed]
- Karaayvaz, M.; Zhai, H.; Ju, J. miR-129 promotes apoptosis and enhances chemosensitivity to 5-fluorouracil in colorectal cancer. Cell Death Dis. 2013, 4, e659. [Google Scholar] [CrossRef]
- Suzuki, R.; Amatya, V.J.; Kushitani, K.; Kai, Y.; Kambara, T.; Takeshima, Y. miR-182 and miR-183 Promote Cell Proliferation and Invasion by Targeting FOXO1 in Mesothelioma. Front. Oncol. 2018, 8, 446. [Google Scholar] [CrossRef]
- Bouteloup, M.; Verney, A.; Rachinel, N.; Callet-Bauchu, E.; Ffrench, M.; Coiffier, B.; Magaud, J.P.; Berger, F.; Salles, G.A.; Traverse-Glehen, A. MicroRNA expression profile in splenic marginal zone lymphoma. Br. J. Haematol. 2012, 156, 279–281. [Google Scholar] [CrossRef]
- Fuertes, T.; Ramiro, A.R.; de Yebenes, V.G. miRNA-Based Therapies in B Cell Non-Hodgkin Lymphoma. Trends Immunol. 2020, 41, 932–947. [Google Scholar] [CrossRef]
- Xiong, W.; Lv, R.; Li, H.; Li, Z.; Wang, H.; Liu, W.; Zou, D.; Qiu, L.; Yi, S. Prevalence of hepatitis B and hepatitis C viral infections in various subtypes of B-cell non-Hodgkin lymphoma: Confirmation of the association with splenic marginal zone lymphoma. Blood Cancer J. 2017, 7, e548. [Google Scholar] [CrossRef] [PubMed]
- Hermine, O.; Lefrere, F.; Bronowicki, J.P.; Mariette, X.; Jondeau, K.; Eclache-Saudreau, V.; Delmas, B.; Valensi, F.; Cacoub, P.; Brechot, C.; et al. Regression of splenic lymphoma with villous lymphocytes after treatment of hepatitis C virus infection. N. Engl. J. Med. 2002, 347, 89–94. [Google Scholar] [CrossRef]
- Arcaini, L.; Vallisa, D.; Rattotti, S.; Ferretti, V.V.; Ferreri, A.J.M.; Bernuzzi, P.; Merli, M.; Varettoni, M.; Chiappella, A.; Ambrosetti, A.; et al. Antiviral treatment in patients with indolent B-cell lymphomas associated with HCV infection: A study of the Fondazione Italiana Linfomi. Ann. Oncol. 2014, 25, 1404–1410. [Google Scholar] [CrossRef] [PubMed]
- Jeon, Y.J.; Lee, K.Y.; Cho, Y.Y.; Pugliese, A.; Kim, H.G.; Jeong, C.H.; Bode, A.M.; Dong, Z. Role of NEK6 in tumor promoter-induced transformation in JB6 C141 mouse skin epidermal cells. J. Biol. Chem. 2010, 285, 28126–28133. [Google Scholar] [CrossRef] [PubMed]
- Arribas, A.J.; Campos-Martin, Y.; Gomez-Abad, C.; Algara, P.; Sanchez-Beato, M.; Rodriguez-Pinilla, M.S.; Montes-Moreno, S.; Martinez, N.; Alves-Ferreira, J.; Piris, M.A.; et al. Nodal marginal zone lymphoma: Gene expression and miRNA profiling identify diagnostic markers and potential therapeutic targets. Blood 2012, 119, e9–e21. [Google Scholar] [CrossRef] [PubMed]
- Jardin, F.; Figeac, M. MicroRNAs in lymphoma, from diagnosis to targeted therapy. Curr. Opin. Oncol. 2013, 25, 480–486. [Google Scholar] [CrossRef] [PubMed]
- Van den Brand, M.; van Krieken, J.H. Recognizing nodal marginal zone lymphoma: Recent advances and pitfalls. A systematic review. Haematologica 2013, 98, 1003–1013. [Google Scholar] [CrossRef]
- Barh, D.; Malhotra, R.; Ravi, B.; Sindhurani, P. MicroRNA let-7: An emerging next-generation cancer therapeutic. Curr. Oncol. 2010, 17, 70–80. [Google Scholar] [CrossRef]
- Gebauer, N.; Thorns, C.; Bernard, V.; Senft, A.; Schillert, A.; Merz, H.; Feller, A.C.; Bernd, H.W. MicroRNA profiling of low-grade and transformed nodal marginal zone lymphoma reveals a similar signature pattern distinct from diffuse large B cell lymphoma. Acta Haematol. 2015, 133, 214–220. [Google Scholar] [CrossRef] [PubMed]
- Angelopoulou, M.K. Transformed nodal marginal zone lymphoma versus diffuse large B cell lymphoma: The microRNA aspect. Acta Haematol. 2015, 133, 212–213. [Google Scholar] [CrossRef] [PubMed]
- Braggio, E.; Keats, J.J.; Leleu, X.; van Wier, S.; Jimenez-Zepeda, V.H.; Valdez, R.; Schop, R.F.; Price-Troska, T.; Henderson, K.; Sacco, A.; et al. Identification of copy number abnormalities and inactivating mutations in two negative regulators of nuclear factor-kappaB signaling pathways in Waldenstrom’s macroglobulinemia. Cancer Res. 2009, 69, 3579–3588. [Google Scholar] [CrossRef]
- Moussay, E.; Wang, K.; Cho, J.H.; van Moer, K.; Pierson, S.; Paggetti, J.; Nazarov, P.V.; Palissot, V.; Hood, L.E.; Berchem, G.; et al. MicroRNA as biomarkers and regulators in B-cell chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 2011, 108, 6573–6578. [Google Scholar] [CrossRef] [PubMed]
- Kitagawa, Y.; Brahmachary, M.; Tiacci, E.; Dalla-Favera, R.; Falini, B.; Basso, K. A microRNA signature specific for hairy cell leukemia and associated with modulation of the MAPK-JNK pathways. Leukemia 2012, 26, 2564–2567. [Google Scholar] [CrossRef]
- Koens, L.; Qin, Y.; Leung, W.Y.; Corver, W.E.; Jansen, P.M.; Willemze, R.; Vermeer, M.H.; Tensen, C.P. MicroRNA profiling of primary cutaneous large B-cell lymphomas. PLoS ONE 2013, 8, e82471. [Google Scholar] [CrossRef]
- Hodge, L.S.; Elsawa, S.F.; Grote, D.M.; Price-Troska, T.L.; Asmann, Y.W.; Fonseca, R.; Gertz, M.A.; Witzig, T.E.; Novak, A.J.; Ansell, S.M. MicroRNA expression in tumor cells from Waldenstrom’s macroglobulinemia reflects both their normal and malignant cell counterparts. Blood Cancer J. 2011, 1, e24. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Qi, L.; Bart, J.; Tan, L.P.; Platteel, I.; Sluis, T.; Huitema, S.; Harms, G.; Fu, L.; Hollema, H.; Berg, A. Expression of miR-21 and its targets (PTEN, PDCD4, TM1) in flat epithelial atypia of the breast in relation to ductal carcinoma in situ and invasive carcinoma. BMC Cancer 2009, 9, 163. [Google Scholar] [CrossRef]
- Kubiczkova Besse, L.; Sedlarikova, L.; Kryukov, F.; Nekvindova, J.; Radova, L.; Almasi, M.; Pelcova, J.; Minarik, J.; Pika, T.; Pikalova, Z.; et al. Combination of serum microRNA-320a and microRNA-320b as a marker for Waldenstrom macroglobulinemia. Am. J. Hematol. 2015, 90, E51–E52. [Google Scholar] [CrossRef]
- Bouyssou, J.M.; Liu, C.J.; Bustoros, M.; Sklavenitis-Pistofidis, R.; Aljawai, Y.; Manier, S.; Yosef, A.; Sacco, A.; Kokubun, K.; Tsukamoto, S.; et al. Profiling of circulating exosomal miRNAs in patients with Waldenstrom Macroglobulinemia. PLoS ONE 2018, 13, e0204589. [Google Scholar] [CrossRef] [PubMed]
- Vazquez, M.I.; Catalan-Dibene, J.; Zlotnik, A. B cells responses and cytokine production are regulated by their immune microenvironment. Cytokine 2015, 74, 318–326. [Google Scholar] [CrossRef] [PubMed]
- De Silva, N.S.; Klein, U. Dynamics of B cells in germinal centres. Nat. Rev. Immunol. 2015, 15, 137–148. [Google Scholar] [CrossRef]
- Garavelli, S.; de Rosa, V.; de Candia, P. The Multifaceted Interface between Cytokines and microRNAs: An Ancient Mechanism to Regulate the Good and the Bad of Inflammation. Front. Immunol. 2018, 9, 3012. [Google Scholar] [CrossRef]
- Barnes, N.A.; Stephenson, S.; Cocco, M.; Tooze, R.M.; Doody, G.M. BLIMP-1 and STAT3 counterregulate microRNA-21 during plasma cell differentiation. J. Immunol. 2012, 189, 253–260. [Google Scholar] [CrossRef]
- Loffler, D.; Brocke-Heidrich, K.; Pfeifer, G.; Stocsits, C.; Hackermuller, J.; Kretzschmar, A.K.; Burger, R.; Gramatzki, M.; Blumert, C.; Bauer, K.; et al. Interleukin-6 dependent survival of multiple myeloma cells involves the Stat3-mediated induction of microRNA-21 through a highly conserved enhancer. Blood 2007, 110, 1330–1333. [Google Scholar] [CrossRef] [PubMed]
- Papanota, A.M.; Karousi, P.; Kontos, C.K.; Ntanasis-Stathopoulos, I.; Scorilas, A.; Terpos, E. Multiple Myeloma Bone Disease: Implication of MicroRNAs in Its Molecular Background. Int. J. Mol. Sci. 2021, 22, 2375. [Google Scholar] [CrossRef]
- Pedersen, I.M.; Otero, D.; Kao, E.; Miletic, A.V.; Hother, C.; Ralfkiaer, E.; Rickert, R.C.; Gronbaek, K.; David, M. Onco-miR-155 targets SHIP1 to promote TNFalpha-dependent growth of B cell lymphomas. EMBO Mol. Med. 2009, 1, 288–295. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Ma, Z.; Kim, B.H.; Wu, W.; Cayting, P.; Boyle, A.P.; Sundaram, V.; Xing, X.; Dogan, N.; Li, J.; et al. Principles of regulatory information conservation between mouse and human. Nature 2014, 515, 371–375. [Google Scholar] [CrossRef] [PubMed]
- Bose, D.; Neumann, A.; Timmermann, B.; Meinke, S.; Heyd, F. Differential Interleukin-2 Transcription Kinetics Render Mouse but Not Human T Cells Vulnerable to Splicing Inhibition Early after Activation. Mol. Cell. Biol. 2019, 39. [Google Scholar] [CrossRef] [PubMed]
- Heykers, A.; Leemans, A.; van der Gucht, W.; de Schryver, M.; Cos, P.; Delputte, P. Differences in Susceptibility of Human and Mouse Macrophage Cell Lines to Respiratory Syncytial Virus Infection. Intervirology 2019, 62, 134–144. [Google Scholar] [CrossRef] [PubMed]
- LeBien, T.W.; Tedder, T.F. B lymphocytes: How they develop and function. Blood 2008, 112, 1570–1580. [Google Scholar] [CrossRef] [PubMed]
- Vijg, J. Somatic mutations, genome mosaicism, cancer and aging. Curr. Opin. Genet. Dev. 2014, 26, 141–149. [Google Scholar] [CrossRef]
- Fernandez, L.C.; Torres, M.; Real, F.X. Somatic mosaicism: On the road to cancer. Nat. Rev. Cancer 2016, 16, 43–55. [Google Scholar] [CrossRef]
- Sato, F.; Tsuchiya, S.; Terasawa, K.; Tsujimoto, G. Intra-platform repeatability and inter-platform comparability of microRNA microarray technology. PLoS ONE 2009, 4, e5540. [Google Scholar] [CrossRef]
- De Tullio, G.; de Fazio, V.; Sgherza, N.; Minoia, C.; Serrati, S.; Merchionne, F.; Loseto, G.; Iacobazzi, A.; Rana, A.; Petrillo, P.; et al. Challenges and opportunities of microRNAs in lymphomas. Molecules 2014, 19, 14723–14781. [Google Scholar] [CrossRef] [PubMed]
- Rupaimoole, R.; Slack, F.J. MicroRNA therapeutics: Towards a new era for the management of cancer and other diseases. Nat. Rev. Drug Discov. 2017, 16, 203–222. [Google Scholar] [CrossRef] [PubMed]
- Hayes, J.; Peruzzi, P.P.; Lawler, S. MicroRNAs in cancer: Biomarkers, functions and therapy. Trends Mol. Med. 2014, 20, 460–469. [Google Scholar] [CrossRef]
- Yao, Q.; Chen, Y.; Zhou, X. The roles of microRNAs in epigenetic regulation. Curr. Opin. Chem. Biol. 2019, 51, 11–17. [Google Scholar] [CrossRef]
- Sohel, M.M.H. Circulating microRNAs as biomarkers in cancer diagnosis. Life Sci. 2020, 248, 117473. [Google Scholar] [CrossRef] [PubMed]
| Disease | miRNAs | Expression in Lymphomas | Targets | Effect | References |
|---|---|---|---|---|---|
| Follicular lymphoma (FL) | miR-150-5p | Decreased | FOXP1 | Inhibition of B-cell survival | [59] |
| miR-31-5p | E2F2, PIK3C2A | Inhibition of cell cycle, survival, and migration | [62] | ||
| miR-202-3p | DICER1 | Regulation of biogenesis of miRNAs | [64] | ||
| SKP2 | Regulation of cell cycle transition | ||||
| miR-618 | HDAC3 | Inhibition of cell cycle | [65] | ||
| CUL4A | Inhibition of DNA damage response | ||||
| miR-155-5p | INPP5D | Promotion of anti-tumor immune responses | [66] | ||
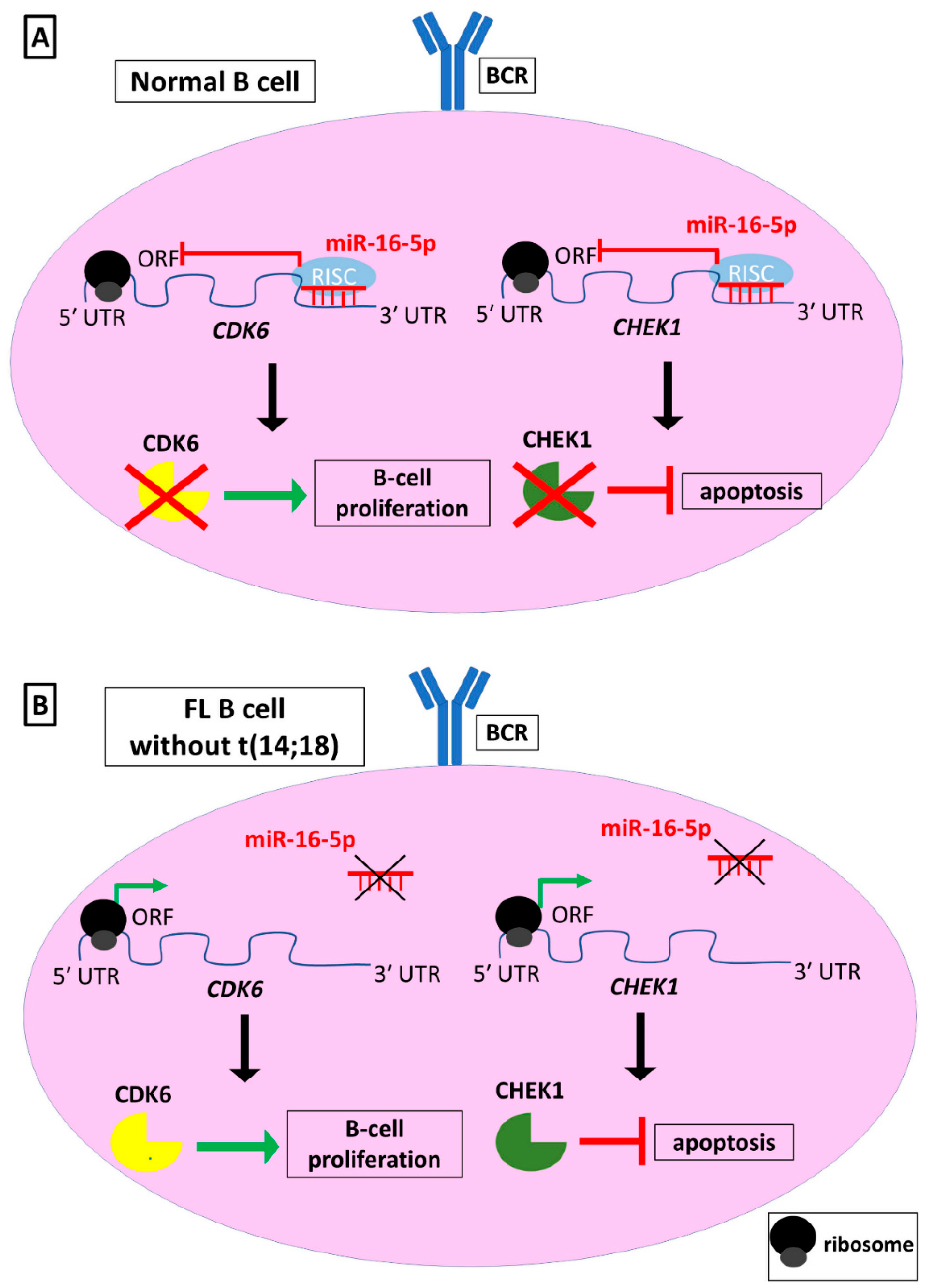
| miR-16-5p | CHEK1 | Promotion of B-cell apoptosis | [48] | ||
| CDK6 | Inhibition of B-cell proliferation | ||||
| miR-20a-5p, miR-20b-5p | Increased | CDKN1A | Promotion of cell cycle | [63] | |
| miR-194-5p | SOCS2 | Promotion of B-cell proliferation and survival | |||
| miR-93-5p | MICA, MICB | Inhibition of B-cell cytotoxicity | [67] | ||
| Gastric MALT lymphoma | miR-34a-5p | Decreased | FOXP1 | Inhibition of B-cell survival | [22,68,69,70,71] |
| miR-383-5p | ZEB2 | Inhibition of epithelial-to-mesenchymal transition (EMT) | [72] | ||
| miR-203a-3p | ABL1 | Inhibition of B-cell proliferation | [73,74] | ||
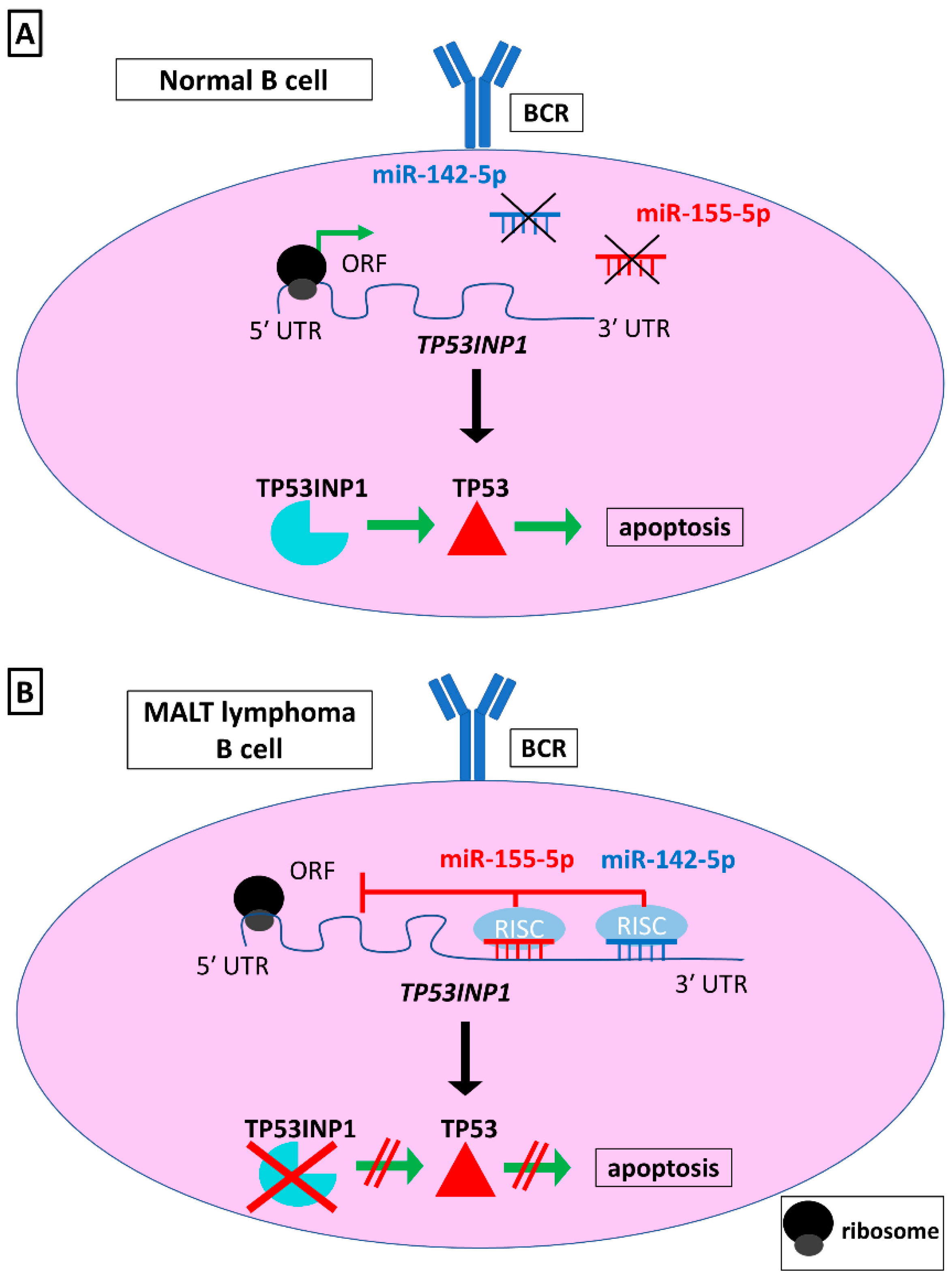
| miR-155-5p, miR-142-5p | Increased | TP53INP1 | Inhibition of apoptosis | [74,75,76] | |
| Splenic MZL | miR-26b-5p | Decreased | NEK6 | Inhibition of mitosis-cell division | [77,78] |
| Waldenström’s macroglobulinemia (WM) | miR-9-3p | Decreased | HDAC4, HDAC5 | Regulation of histone acetylation; Induction of WM cell cytotoxicity; promotion of WM cell autophagy and apoptosis | [79] |
| miR-23b-3p | SP1 | Suppression of NFkB signaling; Inhibition of cell proliferation and survival | [80] | ||
| miR-155-5p | Increased | - | Promotion of MAPK/ERK, PI3K/AKT, and NFkB signaling; promotion of cell proliferation, adhesion, and migration | [81,82] | |
| FOXO3, BCL2L11 | Inhibition of apoptosis | [83] | |||
| miR-206-3p | KAT6A | Regulation of histone acetylation | [79] |
| Disease | miRNAs | Potential Target in Lymphomas | Potential Effect in B Cells | References |
|---|---|---|---|---|
| Gastric MALT lymphoma | miR-150-5p | MYB | Regulation of B-cell development | [14,76,100,101] |
| EGR2 | Inhibition of apoptosis | |||
| miR-196a-5p | CDKN1B | Promotion of cell cycle | [76,104] | |
| miR-153-3p | AKT3 | Inhibition of cell proliferation | [76,105] | |
| miR-7-5p | EGFR and IGF1R | Inhibition of metastasis | [76,106,107] | |
| miR-16-5p | BCL2 | Promotion of apoptosis | [113,114] | |
| Splenic MZL | miR-29a/b1 cluster | TCL1A | Inhibition of cell proliferation | [126,128,129,130,131] |
| miR-129-5p | BCL2 | Promotion of apoptosis | ||
| miR-21-5p | PTEN, FOXO3 | Promotion of cell proliferation, inhibition of apoptosis | [133,134] | |
| Nodal MZL | miR-223-3p | LMO2 | Inhibition of B-cell differentiation | [42,139] |
| OAL | miR-29 family | TCL1A | Inhibition of cell proliferation | [118,119] |
| CDK6 | ||||
| DNMT3B | Inhibition of DNA methylation | |||
| MCL1 | Promotion of apoptosis | |||
| miR-199a-5p | IKBKB | Promotion of apoptosis | [118,120,121] | |
| WM | miR-9-3p | PDRM1 | Inhibition of B-cell differentiation | [149] |
| miR-125b-5p and miR-181a-5p | PRDM1, IRF4 | |||
| let-7a-5p | MYC | Inhibition of cell proliferation | [142] | |
| miR-21-5p | PTEN, PDCD4 | Promotion of cell proliferation, regulation of apoptosis | [150] | |
| PCFCL | miR-9-5p | PRDM1 | Inhibition of B-cell differentiation | [148] |
| miR-129-2-3p | SOX4 | |||
| miR-155-5p | SPI1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Artemaki, P.I.; Letsos, P.A.; Zoupa, I.C.; Katsaraki, K.; Karousi, P.; Papageorgiou, S.G.; Pappa, V.; Scorilas, A.; Kontos, C.K. The Multifaceted Role and Utility of MicroRNAs in Indolent B-Cell Non-Hodgkin Lymphomas. Biomedicines 2021, 9, 333. https://doi.org/10.3390/biomedicines9040333
Artemaki PI, Letsos PA, Zoupa IC, Katsaraki K, Karousi P, Papageorgiou SG, Pappa V, Scorilas A, Kontos CK. The Multifaceted Role and Utility of MicroRNAs in Indolent B-Cell Non-Hodgkin Lymphomas. Biomedicines. 2021; 9(4):333. https://doi.org/10.3390/biomedicines9040333
Chicago/Turabian StyleArtemaki, Pinelopi I., Petros A. Letsos, Ioanna C. Zoupa, Katerina Katsaraki, Paraskevi Karousi, Sotirios G. Papageorgiou, Vasiliki Pappa, Andreas Scorilas, and Christos K. Kontos. 2021. "The Multifaceted Role and Utility of MicroRNAs in Indolent B-Cell Non-Hodgkin Lymphomas" Biomedicines 9, no. 4: 333. https://doi.org/10.3390/biomedicines9040333
APA StyleArtemaki, P. I., Letsos, P. A., Zoupa, I. C., Katsaraki, K., Karousi, P., Papageorgiou, S. G., Pappa, V., Scorilas, A., & Kontos, C. K. (2021). The Multifaceted Role and Utility of MicroRNAs in Indolent B-Cell Non-Hodgkin Lymphomas. Biomedicines, 9(4), 333. https://doi.org/10.3390/biomedicines9040333