Research Advances and Disease Modeling in Respiratory Organoids
Abstract
1. Introduction
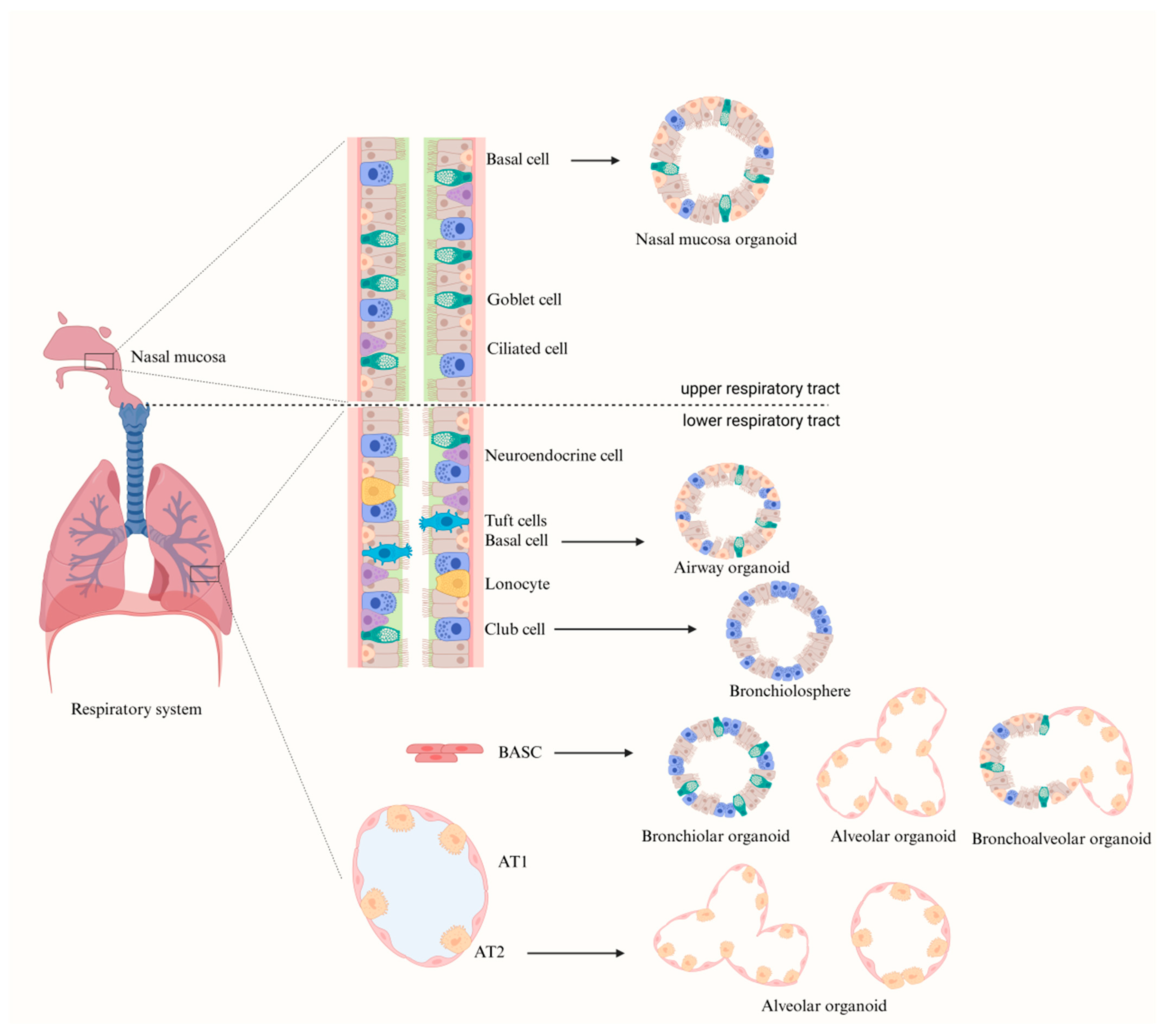
2. Composition of the Respiratory System and Modeling of Respiratory Organoids
2.1. Nasal Mucosa Organoid
2.2. Airway Organoid
2.3. Alveolar Organoid
3. Applications of Respiratory Organoid Models to Study Lung Diseases
3.1. Pulmonary Fibrosis
3.1.1. Idiopathic Pulmonary Fibrosis
3.1.2. Cystic Fibrosis
3.2. Chronic Obstructive Pulmonary Disease
3.3. Respiratory Cancers
3.3.1. Lung Cancer
3.3.2. Nasopharyngeal Carcinoma
4. Conclusions and Future Direction
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| 2D | Two-dimensional |
| 3D | Three-dimensional |
| GEMMs | Genetically engineered mouse models |
| PDXs | Patient-derived xenografts |
| ASCs | Adult stem cells |
| ESCs | Embryonic stem cells |
| iPSCs | Induced pluripotent stem cells |
| AT1 | Type I alveolar epithelial |
| AT2 | Type II alveolar epithelial |
| ALI | Air-liquid interface |
| IPF | Idiopathic pulmonary fibrosis |
| CF | Cystic fibrosis |
| COPD | Chronic obstructive pulmonary disease |
| NPC | Nasopharyngeal carcinoma |
References
- Clevers, H. Modeling Development and Disease with Organoids. Cell 2016, 165, 1586–1597. [Google Scholar] [CrossRef]
- Zhao, Z.; Chen, X.; Dowbaj, A.M.; Sljukic, A.; Bratlie, K.; Lin, L.; Fong, E.L.S.; Balachander, G.M.; Chen, Z.; Soragni, A.; et al. Organoids. Nature reviews. Methods Primers 2022, 2, 94. [Google Scholar] [CrossRef]
- Tang, X.-Y.; Wu, S.; Wang, D.; Chu, C.; Hong, Y.; Tao, M.; Hu, H.; Xu, M.; Guo, X.; Liu, Y. Human organoids in basic research and clinical applications. Signal Transduct. Target. Ther. 2022, 7, 168. [Google Scholar] [CrossRef]
- Saraswathibhatla, A.; Indana, D.; Chaudhuri, O. Cell-extracellular matrix mechanotransduction in 3D. Nat. Rev. Mol. Cell Biol. 2023, 24, 495–516. [Google Scholar] [CrossRef]
- Baker, B.M.; Chen, C.S. Deconstructing the third dimension: How 3D culture microenvironments alter cellular cues. J. Cell Sci. 2012, 125, 3015–3024. [Google Scholar] [CrossRef] [PubMed]
- Scalise, M.; Marino, F.; Salerno, L.; Cianflone, E.; Molinaro, C.; Salerno, N.; De Angelis, A.; Viglietto, G.; Urbanek, K.; Torella, D. From Spheroids to Organoids: The Next Generation of Model Systems of Human Cardiac Regeneration in a Dish. Int. J. Mol. Sci. 2021, 22, 13180. [Google Scholar] [CrossRef] [PubMed]
- Bresnahan, E.; Ramadori, P.; Heikenwalder, M.; Zender, L.; Lujambio, A. Novel patient-derived preclinical models of liver cancer. J. Hepatol. 2020, 72, 239–249. [Google Scholar] [CrossRef] [PubMed]
- Corrò, C.; Novellasdemunt, L.; Li, V.S.W. A brief history of organoids. Am. J. Physiol. Cell Physiol. 2020, 319, C151–C165. [Google Scholar] [CrossRef]
- Brevini, T.; Tysoe, O.C.; Sampaziotis, F. Tissue engineering of the biliary tract and modelling of cholestatic disorders. J. Hepatol. 2020, 73, 918–932. [Google Scholar] [CrossRef]
- Wood, L.D.; Ewald, A.J. Organoids in cancer research: A review for pathologist-scientists. J. Pathol. 2021, 254, 395–404. [Google Scholar] [CrossRef]
- Yang, S.; Hu, H.; Kung, H.; Zou, R.; Dai, Y.; Hu, Y.; Wang, T.; Lv, T.; Yu, J.; Li, F. Organoids: The current status and biomedical applications. Medcomm 2023, 4, e274. [Google Scholar] [CrossRef] [PubMed]
- Thomson, J.A.; Itskovitz-Eldor, J.; Shapiro, S.S.; Waknitz, M.A.; Swiergiel, J.J.; Marshall, V.S.; Jones, J.M. Embryonic Stem Cell Lines Derived from Human Blastocysts. Science 1998, 282, 1145–1147. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef]
- McCracken, K.W.; Aihara, E.; Martin, B.; Crawford, C.M.; Broda, T.; Treguier, J.; Zhang, X.; Shannon, J.M.; Montrose, M.H.; Wells, J.M. Wnt/β-catenin promotes gastric fundus specification in mice and humans. Nature 2017, 541, 182–187. [Google Scholar] [CrossRef]
- Brassard, J.A.; Lutolf, M.P. Engineering Stem Cell Self-organization to Build Better Organoids. Cell Stem Cell 2019, 24, 860–876. [Google Scholar] [CrossRef]
- Schutgens, F.; Clevers, H. Human Organoids: Tools for Understanding Biology and Treating Diseases. Annu. Rev. Pathol. 2020, 15, 211–234. [Google Scholar] [CrossRef]
- Günther, C.; Winner, B.; Neurath, M.F.; Stappenbeck, T.S. Organoids in gastrointestinal diseases: From experimental models to clinical translation. Gut 2022, 71, 1892–1908. [Google Scholar] [CrossRef]
- Sato, T.; Stange, D.E.; Ferrante, M.; Vries, R.G.; van Es, J.H.; Brink, S.v.D.; van Houdt, W.J.; Pronk, A.; van Gorp, J.; Siersema, P.D.; et al. Long-term Expansion of Epithelial Organoids from Human Colon, Adenoma, Adenocarcinoma, and Barrett’s Epithelium. Gastroenterology 2011, 141, 1762–1772. [Google Scholar] [CrossRef]
- Giani, A.M.; Chen, S. Human pluripotent stem cell-based organoids and cell platforms for modelling SARS-CoV-2 infection and drug discovery. Stem Cell Res. 2021, 53, 102207. [Google Scholar] [CrossRef]
- McCracken, K.W.; Catá, E.M.; Crawford, C.M.; Sinagoga, K.L.; Schumacher, M.; Rockich, B.E.; Tsai, Y.-H.; Mayhew, C.N.; Spence, J.R.; Zavros, Y.; et al. Modelling human development and disease in pluripotent stem-cell-derived gastric organoids. Nature 2014, 516, 400–404. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.-C.; Wernig, M.; Duncan, I.D.; Brüstle, O.; Thomson, J.A. In vitro differentiation of transplantable neural precursors from human embryonic stem cells. Nat. Biotechnol. 2001, 19, 1129–1133. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, H.; Sauvey, C.; Yao, L.; Zarnowska, E.D.; Zhang, S.-C. Directed differentiation of forebrain GABA interneurons from human pluripotent stem cells. Nat. Protoc. 2013, 8, 1670–1679. [Google Scholar] [CrossRef] [PubMed]
- Wimmer, R.A.; Leopoldi, A.; Aichinger, M.; Wick, N.; Hantusch, B.; Novatchkova, M.; Taubenschmid, J.; Hämmerle, M.; Esk, C.; Bagley, J.A.; et al. Human blood vessel organoids as a model of diabetic vasculopathy. Nature 2019, 565, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Park, H.J.; Choi, H.; Chang, Y.; Park, H.; Shin, J.; Kim, J.; Lengner, C.J.; Lee, Y.K.; Kim, J. Modeling G2019S-LRRK2 Sporadic Parkinson’s Disease in 3D Midbrain Organoids. Stem Cell Rep. 2019, 12, 518–531. [Google Scholar] [CrossRef] [PubMed]
- Tran, E.; Shi, T.; Li, X.; Chowdhury, A.Y.; Jiang, D.; Liu, Y.; Wang, H.; Yan, C.; Wallace, W.D.; Lu, R.; et al. Development of human alveolar epithelial cell models to study distal lung biology and disease. iScience 2022, 25, 103780. [Google Scholar] [CrossRef]
- Kalender, M.; Bulbul, M.V.; Kolbasi, B.; Keskin, I. In 2D and 3D Cell Culture Models, Effects of Endothelial Cells on E-cadherin/β-catenin Expression Levels and Spheroid Sizes in Ishikawa Cells. Asian Pac. J. Cancer Prev. 2022, 23, 39–51. [Google Scholar] [CrossRef]
- Laube, M.; Pietsch, S.; Pannicke, T.; Thome, U.H.; Fabian, C. Development and Functional Characterization of Fetal Lung Organoids. Front. Med. 2021, 8, 678438. [Google Scholar] [CrossRef]
- Chen, S.; Schoen, J. Air-liquid interface cell culture: From airway epithelium to the female reproductive tract. Reprod. Domest. Anim. 2019, 54, 38–45. [Google Scholar] [CrossRef]
- Lacroix, G.; Koch, W.; Ritter, D.; Gutleb, A.C.; Larsen, S.T.; Loret, T.; Zanetti, F.; Constant, S.; Chortarea, S.; Rothen-Rutishauser, B.; et al. Air–Liquid Interface In Vitro Models for Respiratory Toxicology Research: Consensus Workshop and Recommendations. Appl. Vitr. Toxicol. 2018, 4, 91–106. [Google Scholar] [CrossRef]
- Zhou, J.; Li, C.; Sachs, N.; Chiu, M.C.; Wong, B.H.-Y.; Chu, H.; Poon, V.K.-M.; Wang, D.; Zhao, X.; Wen, L.; et al. Differentiated human airway organoids to assess infectivity of emerging influenza virus. Proc. Natl. Acad. Sci. USA 2018, 115, 6822–6827. [Google Scholar] [CrossRef]
- Kühl, L.; Graichen, P.; von Daacke, N.; Mende, A.; Wygrecka, M.; Potaczek, D.P.; Miethe, S.; Garn, H. Human Lung Organoids—A Novel Experimental and Precision Medicine Approach. Cells 2023, 12, 2067. [Google Scholar] [CrossRef]
- Joo, H.; Min, S.; Cho, S. Advanced lung organoids for respiratory system and pulmonary disease modeling. J. Tissue Eng. 2024, 15, 1788722806. [Google Scholar] [CrossRef]
- Velasco, S.; Kedaigle, A.J.; Simmons, S.K.; Nash, A.; Rocha, M.; Quadrato, G.; Paulsen, B.; Nguyen, L.; Adiconis, X.; Regev, A.; et al. Individual brain organoids reproducibly form cell diversity of the human cerebral cortex. Nature 2019, 570, 523–527. [Google Scholar] [CrossRef] [PubMed]
- Man, W.H.; de Steenhuijsen Piters, W.A.A.; Bogaert, D. The microbiota of the respiratory tract: Gatekeeper to respiratory health. Nat. Rev. Microbiol. 2017, 15, 259–270. [Google Scholar] [CrossRef] [PubMed]
- Weibel, E.R. Lung morphometry: The link between structure and function. Cell Tissue Res. 2017, 367, 413–426. [Google Scholar] [CrossRef]
- Hewitt, R.J.; Lloyd, C.M. Regulation of immune responses by the airway epithelial cell landscape. Nat. Rev. Immunol. 2021, 21, 347–362. [Google Scholar] [CrossRef]
- Widdicombe, J.H.; Wine, J.J. Airway Gland Structure and Function. Physiol. Rev. 2015, 95, 1241–1319. [Google Scholar] [CrossRef]
- Rogers, T.D.; Button, B.; Kelada, S.N.P.; Ostrowski, L.E.; Livraghi-Butrico, A.; Gutay, M.I.; Esther, C.R.; Grubb, B.R. Regional Differences in Mucociliary Clearance in the Upper and Lower Airways. Front. Physiol. 2022, 13, 842592. [Google Scholar] [CrossRef]
- Okuda, K.; Dang, H.; Kobayashi, Y.; Carraro, G.; Nakano, S.; Chen, G.; Kato, T.; Asakura, T.; Gilmore, R.C.; Morton, L.C.; et al. Secretory Cells Dominate Airway CFTR Expression and Function in Human Airway Superficial Epithelia. Am. J. Respir. Crit. Care Med. 2021, 203, 1275–1289. [Google Scholar] [CrossRef] [PubMed]
- Gohy, S.T.; Hupin, C.; Pilette, C.; Ladjemi, M.Z. Chronic inflammatory airway diseases: The central role of the epithelium revisited. Clin. Exp. Allergy 2016, 46, 529–542. [Google Scholar] [CrossRef]
- Knudsen, L.; Ochs, M. The micromechanics of lung alveoli: Structure and function of surfactant and tissue components. Histochem. Cell Biol. 2018, 150, 661–676. [Google Scholar] [CrossRef]
- Bluhmki, T.; Traub, S.; Müller, A.-K.; Bitzer, S.; Schruf, E.; Bammert, M.-T.; Leist, M.; Gantner, F.; Garnett, J.P.; Heilker, R. Functional human iPSC-derived alveolar-like cells cultured in a miniaturized 96-Transwell air–liquid interface model. Sci. Rep. 2021, 11, 17028. [Google Scholar] [CrossRef]
- Rindler, T.N.; Stockman, C.A.; Filuta, A.L.; Brown, K.M.; Snowball, J.M.; Zhou, W.; Veldhuizen, R.; Zink, E.M.; Dautel, S.E.; Clair, G.; et al. Alveolar injury and regeneration following deletion of ABCA3. J. Clin. Investig. 2017, 2, e97381. [Google Scholar] [CrossRef]
- Evans, K.V.; Lee, J. Alveolar wars: The rise of in vitro models to understand human lung alveolar maintenance, regeneration, and disease. Stem Cells Transl. Med. 2020, 9, 867–881. [Google Scholar] [CrossRef] [PubMed]
- Jain, K.G.; Zhao, R.; Liu, Y.; Guo, X.; Yi, G.; Ji, H.-L. Wnt5a/β-catenin axis is involved in the downregulation of AT2 lineage by PAI-1. Am. J. Physiol. Cell. Mol. Physiol. 2022, 323, L515–L524. [Google Scholar] [CrossRef]
- Strunz, M.; Simon, L.M.; Ansari, M.; Kathiriya, J.J.; Angelidis, I.; Mayr, C.H.; Tsidiridis, G.; Lange, M.; Mattner, L.F.; Yee, M.; et al. Alveolar regeneration through a Krt8+ transitional stem cell state that persists in human lung fibrosis. Nat. Commun. 2020, 11, 3559. [Google Scholar] [CrossRef]
- Basil, M.C.; Katzen, J.; Engler, A.E.; Guo, M.; Herriges, M.J.; Kathiriya, J.J.; Windmueller, R.; Ysasi, A.B.; Zacharias, W.J.; Chapman, H.A.; et al. The Cellular and Physiological Basis for Lung Repair and Regeneration: Past, Present, and Future. Cell Stem Cell 2020, 26, 482–502. [Google Scholar] [CrossRef] [PubMed]
- Konkimalla, A.; Tata, A.; Tata, P.R. Lung Regeneration: Cells, Models, and Mechanisms. Cold Spring Harb. Perspect. Biol. 2022, 14, a040873. [Google Scholar] [CrossRef]
- Hogan, B.L.; Barkauskas, C.E.; Chapman, H.A.; Epstein, J.A.; Jain, R.; Hsia, C.C.; Niklason, L.; Calle, E.; Le, A.; Randell, S.H.; et al. Repair and Regeneration of the Respiratory System: Complexity, Plasticity, and Mechanisms of Lung Stem Cell Function. Cell Stem Cell 2014, 15, 123–138. [Google Scholar] [CrossRef]
- Hong, K.U.; Reynolds, S.D.; Watkins, S.; Fuchs, E.; Stripp, B.R. Basal Cells Are a Multipotent Progenitor Capable of Renewing the Bronchial Epithelium. Am. J. Pathol. 2004, 164, 577–588. [Google Scholar] [CrossRef] [PubMed]
- Boers, J.E.; Ambergen, A.W.; Thunnissen, F.B. Number and proliferation of basal and parabasal cells in normal human airway epithelium. Am. J. Respir. Crit. Care Med. 1998, 157, 2000–2006. [Google Scholar] [CrossRef]
- Rokicki, W.; Rokicki, M.; Wojtacha, J.; Dżeljijli, A. The role and importance of club cells (Clara cells) in the pathogenesis of some respiratory diseases. Pol. J. Cardio-Thoracic Surg. 2016, 1, 26–30. [Google Scholar] [CrossRef]
- Van Keymeulen, A.; Blanpain, C. Tracing epithelial stem cells during development, homeostasis, and repair. J. Cell Biol. 2012, 197, 575–584. [Google Scholar] [CrossRef] [PubMed]
- Rock, J.R.; Onaitis, M.W.; Rawlins, E.L.; Lu, Y.; Clark, C.P.; Xue, Y.; Randell, S.H.; Hogan, B.L.M. Basal cells as stem cells of the mouse trachea and human airway epithelium. Proc. Natl. Acad. Sci. USA 2009, 106, 12771–12775. [Google Scholar] [CrossRef] [PubMed]
- Tata, P.R.; Mou, H.; Pardo-Saganta, A.; Zhao, R.; Prabhu, M.; Law, B.M.; Vinarsky, V.; Cho, J.L.; Breton, S.; Sahay, A.; et al. Dedifferentiation of committed epithelial cells into stem cells in vivo. Nature 2013, 503, 218–223. [Google Scholar] [CrossRef]
- Chen, H.; Matsumoto, K.; Brockway, B.L.; Rackley, C.R.; Liang, J.; Lee, J.-H.; Jiang, D.; Noble, P.W.; Randell, S.H.; Kim, C.F.; et al. Airway Epithelial Progenitors Are Region Specific and Show Differential Responses to Bleomycin-Induced Lung Injury. Stem Cells 2012, 30, 1948–1960. [Google Scholar] [CrossRef] [PubMed]
- Rawlins, E.L.; Okubo, T.; Xue, Y.; Brass, D.M.; Auten, R.L.; Hasegawa, H.; Wang, F.; Hogan, B.L. The Role of Scgb1a1+ Clara Cells in the Long-Term Maintenance and Repair of Lung Airway, but Not Alveolar, Epithelium. Cell Stem Cell 2009, 4, 525–534. [Google Scholar] [CrossRef]
- Butler, C.R.; Hynds, R.E.; Gowers, K.H.C.; Lee, D.D.H.; Brown, J.M.; Crowley, C.; Teixeira, V.H.; Smith, C.M.; Urbani, L.; Hamilton, N.J.; et al. Rapid Expansion of Human Epithelial Stem Cells Suitable for Airway Tissue Engineering. Am. J. Respir. Crit. Care Med. 2016, 194, 156–168. [Google Scholar] [CrossRef]
- Danahay, H.; Pessotti, A.D.; Coote, J.; Montgomery, B.E.; Xia, D.; Wilson, A.; Yang, H.; Wang, Z.; Bevan, L.; Thomas, C.; et al. Notch2 Is Required for Inflammatory Cytokine-Driven Goblet Cell Metaplasia in the Lung. Cell Rep. 2015, 10, 239–252. [Google Scholar] [CrossRef]
- Hild, M.; Jaffe, A.B. Production of 3-D Airway Organoids from Primary Human Airway Basal Cells and Their Use in High-Throughput Screening. Curr. Protoc. Stem Cell Biol. 2016, 37, IE.9.1–IE.9.15. [Google Scholar] [CrossRef]
- Kim, C.F.B.; Jackson, E.L.; Woolfenden, A.E.; Lawrence, S.; Babar, I.; Vogel, S.; Crowley, D.; Bronson, R.T.; Jacks, T. Identification of Bronchioalveolar Stem Cells in Normal Lung and Lung Cancer. Cell 2005, 121, 823–835. [Google Scholar] [CrossRef]
- Salwig, I.; Spitznagel, B.; Vazquez-Armendariz, A.I.; Khalooghi, K.; Guenther, S.; Herold, S.; Szibor, M.; Braun, T. Bronchioalveolar stem cells are a main source for regeneration of distal lung epithelia in vivo. EMBO J. 2019, 38, e102099. [Google Scholar] [CrossRef]
- Kotton, D.N.; Morrisey, E.E. Lung regeneration: Mechanisms, applications and emerging stem cell populations. Nat. Med. 2014, 20, 822–832. [Google Scholar] [CrossRef]
- Song, H.; Yao, E.; Lin, C.; Gacayan, R.; Chen, M.-H.; Chuang, P.-T. Functional characterization of pulmonary neuroendocrine cells in lung development, injury, and tumorigenesis. Proc. Natl. Acad. Sci. USA 2012, 109, 17531–17536. [Google Scholar] [CrossRef]
- Li, K.; Li, M.; Li, W.; Yu, H.; Sun, X.; Zhang, Q.; Li, Y.; Li, X.; Li, Y.; Abel, E.D.; et al. Airway epithelial regeneration requires autophagy and glucose metabolism. Cell Death Dis. 2019, 10, 875. [Google Scholar] [CrossRef]
- Barkauskas, C.E.; Cronce, M.J.; Rackley, C.R.; Bowie, E.J.; Keene, D.R.; Stripp, B.R.; Randell, S.H.; Noble, P.W.; Hogan, B.L. Type 2 alveolar cells are stem cells in adult lung. J. Clin. Investig. 2013, 123, 3025–3036. [Google Scholar] [CrossRef]
- Liu, Z.; Anderson, J.D.; Natt, J.; Guimbellot, J.S. Culture and Imaging of Human Nasal Epithelial Organoids. J. Vis. Exp. JoVE 2021, 178, 10–3791. [Google Scholar]
- Brewington, J.J.; Filbrandt, E.T.; LaRosa, F.; Ostmann, A.J.; Strecker, L.M.; Szczesniak, R.D.; Clancy, J.P. Detection of CFTR function and modulation in primary human nasal cell spheroids. J. Cyst. Fibros. 2018, 17, 26–33. [Google Scholar] [CrossRef]
- Gamage, A.M.; Tan, K.S.; Chan, W.O.Y.; Liu, J.; Tan, C.W.; Ong, Y.K.; Thong, M.; Andiappan, A.K.; Anderson, D.E.; Wang, Y.; et al. Infection of human Nasal Epithelial Cells with SARS-CoV-2 and a 382-nt deletion isolate lacking ORF8 reveals similar viral kinetics and host transcriptional profiles. PLoS Pathog. 2020, 16, e1009130. [Google Scholar] [CrossRef]
- Chiu, M.C.; Li, C.; Liu, X.; Song, W.; Wan, Z.; Yu, Y.; Huang, J.; Xiao, D.; Chu, H.; Cai, J.-P.; et al. Human Nasal Organoids Model SARS-CoV-2 Upper Respiratory Infection and Recapitulate the Differential Infectivity of Emerging Variants. mBio 2022, 13, e0194422. [Google Scholar] [CrossRef]
- Rajan, A.; Weaver, A.M.; Aloisio, G.M.; Jelinski, J.; Johnson, H.L.; Venable, S.F.; McBride, T.; Aideyan, L.; Piedra, F.-A.; Ye, X.; et al. The Human Nose Organoid Respiratory Virus Model: An Ex Vivo Human Challenge Model To Study Respiratory Syncytial Virus (RSV) and Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Pathogenesis and Evaluate Therapeutics. mBio 2022, 13, e0351121. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Yu, Y.; Han, R.; Wang, X.; Zhao, Y.; Tang, H.; Li, G. Establishment of a culture system for human nasal mucosa organoids with controllable differentiation. Nan Fang Yi Ke Da Xue Xue Bao 2022, 42, 868–877. [Google Scholar]
- Wang, X.-W.; Xia, T.-L.; Tang, H.-C.; Liu, X.; Han, R.; Zou, X.; Zhao, Y.-T.; Chen, M.-Y.; Li, G. Establishment of a patient-derived organoid model and living biobank for nasopharyngeal carcinoma. Ann. Transl. Med. 2022, 10, 526. [Google Scholar] [CrossRef]
- Hui, K.P.Y.; Ching, R.H.H.; Chan, S.K.H.; Nicholls, J.M.; Sachs, N.; Clevers, H.; Peiris, J.S.M.; Chan, M.C.W. Tropism, replication competence, and innate immune responses of influenza virus: An analysis of human airway organoids and ex-vivo bronchus cultures. Lancet Respir. Med. 2018, 6, 846–854. [Google Scholar] [CrossRef]
- Li, L.; Jiao, L.; Feng, D.; Yuan, Y.; Yang, X.; Li, J.; Jiang, D.; Chen, H.; Meng, Q.; Chen, R.; et al. Human apical-out nasal organoids reveal an essential role of matrix metalloproteinases in airway epithelial differentiation. Nat. Commun. 2024, 15, 143. [Google Scholar] [CrossRef]
- Kumar, P.A.; Hu, Y.; Yamamoto, Y.; Hoe, N.B.; Wei, T.S.; Mu, D.; Sun, Y.; Joo, L.S.; Dagher, R.; Zielonka, E.M.; et al. Distal Airway Stem Cells Yield Alveoli In Vitro and during Lung Regeneration following H1N1 Influenza Infection. Cell 2011, 147, 525–538. [Google Scholar] [CrossRef]
- Konishi, S.; Gotoh, S.; Tateishi, K.; Yamamoto, Y.; Korogi, Y.; Nagasaki, T.; Matsumoto, H.; Muro, S.; Hirai, T.; Ito, I.; et al. Directed Induction of Functional Multi-ciliated Cells in Proximal Airway Epithelial Spheroids from Human Pluripotent Stem Cells. Stem Cell Rep. 2016, 6, 18–25. [Google Scholar] [CrossRef]
- Miller, A.J.; Dye, B.R.; Ferrer-Torres, D.; Hill, D.R.; Overeem, A.W.; Shea, L.D.; Spence, J.R. Generation of lung organoids from human pluripotent stem cells in vitro. Nat. Protoc. 2019, 14, 518–540. [Google Scholar] [CrossRef] [PubMed]
- Miller, A.J.; Yu, Q.; Czerwinski, M.; Tsai, Y.-H.; Conway, R.F.; Wu, A.; Holloway, E.M.; Walker, T.; Glass, I.A.; Treutlein, B.; et al. In Vitro and In Vivo Development of the Human Airway at Single-Cell Resolution. Dev. Cell 2020, 53, 117–128. [Google Scholar] [CrossRef]
- Nikolić, M.Z.; Caritg, O.; Jeng, Q.; Johnson, J.-A.; Sun, D.; Howell, K.J.; Brady, J.L.; Laresgoiti, U.; Allen, G.; Butler, R.; et al. Human embryonic lung epithelial tips are multipotent progenitors that can be expanded in vitro as long-term self-renewing organoids. eLife 2017, 6, e26575. [Google Scholar] [CrossRef] [PubMed]
- Hein, R.F.; Wu, J.H.; Holloway, E.M.; Frum, T.; Conchola, A.S.; Tsai, Y.-H.; Wu, A.; Fine, A.S.; Miller, A.J.; Szenker-Ravi, E.; et al. R-SPONDIN2 mesenchymal cells form the bud tip progenitor niche during human lung development. Dev. Cell 2022, 57, 1598–1614. [Google Scholar] [CrossRef]
- Tan, Q.; Choi, K.M.; Sicard, D.; Tschumperlin, D.J. Human airway organoid engineering as a step toward lung regeneration and disease modeling. Biomaterials 2017, 113, 118–132. [Google Scholar] [CrossRef] [PubMed]
- Sachs, N.; Papaspyropoulos, A.; Zomer-van Ommen, D.D.; Heo, I.; Böttinger, L.; Klay, D.; Weeber, F.; Huelsz-Prince, G.; Iakobachvili, N.; Amatngalim, G.D.; et al. Long-term expanding human airway organoids for disease modeling. EMBO J. 2019, 38, e100300. [Google Scholar] [CrossRef]
- Mou, H.; Vinarsky, V.; Tata, P.R.; Brazauskas, K.; Choi, S.; Crooke, A.K.; Zhang, B.; Solomon, G.M.; Turner, B.; Bihler, H.; et al. Dual SMAD Signaling Inhibition Enables Long-Term Expansion of Diverse Epithelial Basal Cells. Cell Stem Cell 2016, 19, 217–231. [Google Scholar] [CrossRef] [PubMed]
- Moiseenko, A.; Vazquez-Armendariz, A.I.; Kheirollahi, V.; Chu, X.; Tata, A.; Rivetti, S.; Günther, S.; Lebrigand, K.; Herold, S.; Braun, T.; et al. Identification of a Repair-Supportive Mesenchymal Cell Population during Airway Epithelial Regeneration. Cell Rep. 2020, 33, 108549. [Google Scholar] [CrossRef]
- Vazquez-Armendariz, A.I.; Seeger, W.; Herold, S.; El Agha, E. Protocol for the generation of murine bronchiolospheres. STAR Protoc. 2021, 2, 100594. [Google Scholar] [CrossRef]
- Lee, J.-H.; Bhang, D.H.; Beede, A.; Huang, T.L.; Stripp, B.R.; Bloch, K.D.; Wagers, A.J.; Tseng, Y.-H.; Ryeom, S.; Kim, C.F. Lung Stem Cell Differentiation in Mice Directed by Endothelial Cells via a BMP4-NFATc1-Thrombospondin-1 Axis. Cell 2014, 156, 440–455. [Google Scholar] [CrossRef]
- Raasch, K.; Maurat, E.; Henrot, P.; Zysman, M.; Prevel, R.; Thumerel, M.; Nassoy, P.; Berger, P.; Andrique, L.; Recher, G.; et al. A novel in vitro tubular model to recapitulate features of distal airways: The bronchioid. Rev. Des Mal. Respir. 2024, 41, 2400562. [Google Scholar] [CrossRef]
- Liu, M.Y.; Chen, B.; Borji, M.; Rivas, C.G.d.A.; Dost, A.F.M.; Moye, A.L.; Abdulla, N.M.; Paschini, M.; Rollins, S.D.; Wang, R.; et al. Human Airway and Alveolar Organoids from BAL Fluid. Am. J. Respir. Crit. Care Med. 2024, 209, 1501–1504. [Google Scholar] [CrossRef]
- Chiu, M.C.; Zhang, S.; Li, C.; Liu, X.; Yu, Y.; Huang, J.; Wan, Z.; Zhu, X.; Zhou, J. Apical-Out Human Airway Organoids Modeling SARS-CoV-2 Infection. Viruses 2023, 15, 1166. [Google Scholar] [CrossRef] [PubMed]
- Zacharias, W.J.; Frank, D.B.; Zepp, J.A.; Morley, M.P.; Alkhaleel, F.A.; Kong, J.; Zhou, S.; Cantu, E.; Morrisey, E.E. Regeneration of the lung alveolus by an evolutionarily conserved epithelial progenitor. Nature 2018, 555, 251–255. [Google Scholar] [CrossRef]
- Leeman, K.T.; Pessina, P.; Lee, J.-H.; Kim, C.F. Mesenchymal Stem Cells Increase Alveolar Differentiation in Lung Progenitor Organoid Cultures. Sci. Rep. 2019, 9, 6479. [Google Scholar] [CrossRef]
- Hoffmann, K.; Obermayer, B.; Hönzke, K.; Fatykhova, D.; Demir, Z.; Löwa, A.; Alves, L.G.T.; Wyler, E.; Lopez-Rodriguez, E.; Mieth, M.; et al. Human alveolar progenitors generate dual lineage bronchioalveolar organoids. Commun. Biol. 2022, 5, 875. [Google Scholar] [CrossRef]
- Wu, D.; Pan, W. GSK3: A multifaceted kinase in Wnt signaling. Trends Biochem. Sci. 2010, 35, 161–168. [Google Scholar] [CrossRef]
- Shiraishi, K.; Nakajima, T.; Shichino, S.; Deshimaru, S.; Matsushima, K.; Ueha, S. In vitro expansion of endogenous human alveolar epithelial type II cells in fibroblast-free spheroid culture. Biochem. Biophys. Res. Commun. 2019, 515, 579–585. [Google Scholar] [CrossRef]
- Liu, K.; Tang, M.; Liu, Q.; Han, X.; Jin, H.; Zhu, H.; Li, Y.; He, L.; Ji, H.; Zhou, B. Bi-directional differentiation of single bronchioalveolar stem cells during lung repair. Cell Discov. 2020, 6, 1. [Google Scholar] [CrossRef] [PubMed]
- Vazquez-Armendariz, A.I.; Heiner, M.; El Agha, E.; Salwig, I.; Hoek, A.; Hessler, M.C.; Shalashova, I.; Shrestha, A.; Carraro, G.; Mengel, J.P.; et al. Multilineage murine stem cells generate complex organoids to model distal lung development and disease. EMBO J. 2020, 39, e103476. [Google Scholar] [CrossRef]
- Salahudeen, A.A.; Choi, S.S.; Rustagi, A.; Zhu, J.; van Unen, V.; de la O, S.M.; Flynn, R.A.; Margalef-Català, M.; Santos, A.J.M.; Ju, J.; et al. Progenitor identification and SARS-CoV-2 infection in human distal lung organoids. Nature 2020, 588, 670–675. [Google Scholar] [CrossRef] [PubMed]
- Dye, B.R.; Hill, D.R.; Ferguson, M.A.; Tsai, Y.-H.; Nagy, M.S.; Dyal, R.; Wells, J.M.; Mayhew, C.N.; Nattiv, R.; Klein, O.D.; et al. In vitro generation of human pluripotent stem cell derived lung organoids. eLife 2015, 4, e05098. [Google Scholar] [CrossRef][Green Version]
- Tamò, L.; Hibaoui, Y.; Kallol, S.; Alves, M.P.; Albrecht, C.; Hostettler, K.E.; Feki, A.; Rougier, J.-S.; Abriel, H.; Knudsen, L.; et al. Generation of an alveolar epithelial type II cell line from induced pluripotent stem cells. Am. J. Physiol. Cell. Mol. Physiol. 2018, 315, L921–L932. [Google Scholar] [CrossRef] [PubMed]
- Ozan, V.B.; Wang, H.; Akshay, A.; Anand, D.; Hibaoui, Y.; Feki, A.; Gote-Schniering, J.; Gheinani, A.H.; Heller, M.; Uldry, A.-C.; et al. Influence of Microenvironmental Orchestration on Multicellular Lung Alveolar Organoid Development from Human Induced Pluripotent Stem Cells. Stem Cell Rev. Rep. 2024, 21, 254–275. [Google Scholar] [CrossRef]
- Lee, J.; Baek, H.; Jang, J.; Park, J.; Cha, S.-R.; Hong, S.-H.; Kim, J.; Lee, J.-H.; Hong, I.-S.; Wang, S.-J.; et al. Establishment of a human induced pluripotent stem cell derived alveolar organoid for toxicity assessment. Toxicol. In Vitro 2023, 89, 105585. [Google Scholar] [CrossRef]
- Seo, H.-R.; Han, H.-J.; Lee, Y.; Noh, Y.-W.; Cho, S.-J.; Kim, J.-H. Human Pluripotent Stem Cell-Derived Alveolar Organoid with Macrophages. Int. J. Mol. Sci. 2022, 23, 9211. [Google Scholar] [CrossRef]
- Yamamoto, Y.; Gotoh, S.; Korogi, Y.; Seki, M.; Konishi, S.; Ikeo, S.; Sone, N.; Nagasaki, T.; Matsumoto, H.; Muro, S.; et al. Long-term expansion of alveolar stem cells derived from human iPS cells in organoids. Nat. Methods 2017, 14, 1097–1106. [Google Scholar] [CrossRef]
- Tamai, K.; Sakai, K.; Yamaki, H.; Moriguchi, K.; Igura, K.; Maehana, S.; Suezawa, T.; Takehara, K.; Hagiwara, M.; Hirai, T.; et al. iPSC-derived mesenchymal cells that support alveolar organoid development. Cell Rep. Methods 2022, 2, 100314. [Google Scholar] [CrossRef]
- Purev, E.; Bahmed, K.; Kosmider, B. Alveolar Organoids in Lung Disease Modeling. Biomolecules 2024, 14, 115. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Na, F. Organoid technology and applications in lung diseases: Models, mechanism research and therapy opportunities. Front. Bioeng. Biotechnol. 2022, 10, 1066869. [Google Scholar] [CrossRef]
- Lee, J.; Kim, J.-H.; Hong, S.-H.; Yang, S.-R. Organoid Model in Idiopathic Pulmonary Fibrosis. Int. J. Stem Cells 2021, 14, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.-H.; An, G.H.; Kim, J.-Y.; Rasaei, R.; Kim, W.J.; Jin, X.; Woo, D.-H.; Han, C.; Yang, S.-R.; Kim, J.-H.; et al. Human pluripotent stem cell-derived alveolar organoids for modeling pulmonary fibrosis and drug testing. Cell Death Discov. 2021, 7, 48. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.-W.; Huang, S.X.; de Carvalho, A.L.R.T.; Ho, S.-H.; Islam, M.N.; Volpi, S.; Notarangelo, L.D.; Ciancanelli, M.; Casanova, J.-L.; Bhattacharya, J.; et al. A three-dimensional model of human lung development and disease from pluripotent stem cells. Nat. Cell Biol. 2017, 19, 542–549. [Google Scholar] [CrossRef]
- Surolia, R.; Li, F.J.; Wang, Z.; Li, H.; Liu, G.; Zhou, Y.; Luckhardt, T.; Bae, S.; Liu, R.-M.; Rangarajan, S.; et al. 3D pulmospheres serve as a personalized and predictive multicellular model for assessment of antifibrotic drugs. J. Clin. Investig. 2017, 2, e91377. [Google Scholar]
- Strikoudis, A.; Cieślak, A.; Loffredo, L.; Chen, Y.-W.; Patel, N.; Saqi, A.; Lederer, D.J.; Snoeck, H.-W. Modeling of Fibrotic Lung Disease Using 3D Organoids Derived from Human Pluripotent Stem Cells. Cell Rep. 2019, 27, 3709–3723. [Google Scholar] [CrossRef]
- Xu, C.; Zhao, J.; Li, Q.; Hou, L.; Wang, Y.; Li, S.; Jiang, F.; Zhu, Z.; Tian, L. Exosomes derived from three-dimensional cultured human umbilical cord mesenchymal stem cells ameliorate pulmonary fibrosis in a mouse silicosis model. Stem Cell Res. Ther. 2020, 11, 503. [Google Scholar] [CrossRef]
- Ptasinski, V.; Monkley, S.J.; Öst, K.; Tammia, M.; Alsafadi, H.N.; Overed-Sayer, C.; Hazon, P.; E Wagner, D.; A Murray, L. Modeling fibrotic alveolar transitional cells with pluripotent stem cell-derived alveolar organoids. Life Sci. Alliance 2023, 6, e202201853. [Google Scholar] [CrossRef]
- Wang, S.; Li, X.; Ma, Q.; Wang, Q.; Wu, J.; Yu, H.; Li, K.; Li, Y.; Wang, J.; Zhang, Q.; et al. Glutamine Metabolism Is Required for Alveolar Regeneration during Lung Injury. Biomolecules 2022, 12, 728. [Google Scholar] [CrossRef]
- Suezawa, T.; Kanagaki, S.; Moriguchi, K.; Masui, A.; Nakao, K.; Toyomoto, M.; Tamai, K.; Mikawa, R.; Hirai, T.; Murakami, K.; et al. Disease modeling of pulmonary fibrosis using human pluripotent stem cell-derived alveolar organoids. Stem Cell Rep. 2021, 16, 2973–2987. [Google Scholar] [CrossRef] [PubMed]
- Sundarakrishnan, A.; Chen, Y.; Black, L.D.; Aldridge, B.B.; Kaplan, D.L. Engineered cell and tissue models of pulmonary fibrosis. Adv. Drug Deliv. Rev. 2018, 129, 78–94. [Google Scholar] [CrossRef]
- Wilkinson, D.C.; Mellody, M.; Meneses, L.K.; Hope, A.C.; Dunn, B.; Gomperts, B.N. Development of a Three-Dimensional Bioengineering Technology to Generate Lung Tissue for Personalized Disease Modeling. Curr. Protoc. Stem Cell Biol. 2018, 46, e56. [Google Scholar] [CrossRef]
- Jaeger, B.; Schupp, J.C.; Plappert, L.; Terwolbeck, O.; Artysh, N.; Kayser, G.; Engelhard, P.; Adams, T.S.; Zweigerdt, R.; Kempf, H.; et al. Airway basal cells show a dedifferentiated KRT17highPhenotype and promote fibrosis in idiopathic pulmonary fibrosis. Nat. Commun. 2022, 13, 5637. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.; Liang, J.; Huang, K.; Liu, X.; Taghavifar, F.; Yao, C.; Parimon, T.; Liu, N.; Dai, K.; Aziz, A.; et al. Basal Cell–derived WNT7A Promotes Fibrogenesis at the Fibrotic Niche in Idiopathic Pulmonary Fibrosis. Am. J. Respir. Cell Mol. Biol. 2023, 68, 302–313. [Google Scholar] [CrossRef] [PubMed]
- Enomoto, Y.; Katsura, H.; Fujimura, T.; Ogata, A.; Baba, S.; Yamaoka, A.; Kihara, M.; Abe, T.; Nishimura, O.; Kadota, M.; et al. Autocrine TGF-β-positive feedback in profibrotic AT2-lineage cells plays a crucial role in non-inflammatory lung fibrogenesis. Nat. Commun. 2023, 14, 4956. [Google Scholar] [CrossRef] [PubMed]
- Sui, J.; Boatz, J.C.; Shi, J.; Hu, Q.; Li, X.; Zhang, Y.; Königshoff, M.; Kliment, C.R. Loss of ANT1 Increases Fibrosis and Epithelial Cell Senescence in Idiopathic Pulmonary Fibrosis. Am. J. Respir. Cell Mol. Biol. 2023, 69, 556–569. [Google Scholar] [CrossRef]
- Cohen, M.L.; Brumwell, A.N.; Ho, T.C.; Garakani, K.; Montas, G.; Leong, D.; Ding, V.W.; Golden, J.A.; Trinh, B.N.; Jablons, D.M.; et al. A fibroblast-dependent TGF-β1/sFRP2 noncanonical Wnt signaling axis promotes epithelial metaplasia in idiopathic pulmonary fibrosis. J. Clin. Investig. 2024, 134, e174598. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Verschut, V.; Woest, M.E.; Ng-Blichfeldt, J.-P.; Matias, A.; Villetti, G.; Accetta, A.; Facchinetti, F.; Gosens, R.; Kistemaker, L.E.M. Rho-Kinase 1/2 Inhibition Prevents Transforming Growth Factor-β-Induced Effects on Pulmonary Remodeling and Repair. Front. Pharmacol. 2021, 11, 609509. [Google Scholar] [CrossRef]
- Wang, H.; Han, Z.; Yang, Y.; Liu, L.; Huang, Y.; Chen, J.; Wang, Y.; Liu, Z.; Xin, L.; Zhao, Y.; et al. Modeling of lung organoid-based fibrosis for testing the sensitivity of anti-fibrotic drugs. Stem Cell Res. Ther. 2025, 16, 132. [Google Scholar] [CrossRef]
- Marwick, J.A.; Elliott, R.J.R.; Longden, J.; Makda, A.; Hirani, N.; Dhaliwal, K.; Dawson, J.C.; Carragher, N.O. Application of a High-Content Screening Assay Utilizing Primary Human Lung Fibroblasts to Identify Antifibrotic Drugs for Rapid Repurposing in COVID-19 Patients. SLAS Discov. Adv. Sci. Drug Discov. 2022, 26, 1091–1106. [Google Scholar] [CrossRef]
- Calucho, M.; Gartner, S.; Barranco, P.; Fernández-Álvarez, P.; Pérez, R.G.; Tizzano, E.F. Validation of nasospheroids to assay CFTR functionality and modulator responses in cystic fibrosis. Sci. Rep. 2021, 11, 15511. [Google Scholar] [CrossRef]
- McCauley, K.B.; Hawkins, F.; Serra, M.; Thomas, D.C.; Jacob, A.; Kotton, D.N. Efficient Derivation of Functional Human Airway Epithelium from Pluripotent Stem Cells via Temporal Regulation of Wnt Signaling. Cell Stem Cell 2017, 20, 844–857. [Google Scholar] [CrossRef] [PubMed]
- Dekkers, J.F.; Berkers, G.; Kruisselbrink, E.; Vonk, A.; de Jonge, H.R.; Janssens, H.M.; Bronsveld, I.; van de Graaf, E.A.; Nieuwenhuis, E.E.S.; Houwen, R.H.J.; et al. Characterizing responses to CFTR-modulating drugs using rectal organoids derived from subjects with cystic fibrosis. Sci. Transl. Med. 2016, 8, 344ra84. [Google Scholar] [CrossRef]
- Ramalho, A.S.; Fürstová, E.; Vonk, A.M.; Ferrante, M.; Verfaillie, C.; Dupont, L.; Boon, M.; Proesmans, M.; Beekman, J.M.; Sarouk, I.; et al. Correction of CFTR function in intestinal organoids to guide treatment of cystic fibrosis. Eur. Respir. J. 2020, 57, 1902426. [Google Scholar] [CrossRef]
- Guimbellot, J.S.; Leach, J.M.; Chaudhry, I.G.; Quinney, N.L.; Boyles, S.E.; Chua, M.; Aban, I.; Jaspers, I.; Gentzsch, M. Nasospheroids permit measurements of CFTR-dependent fluid transport. J. Clin. Investig. 2017, 2, e95734. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Anderson, J.D.; Deng, L.; Mackay, S.; Bailey, J.; Kersh, L.; Rowe, S.M.; Guimbellot, J.S. Human Nasal Epithelial Organoids for Therapeutic Development in Cystic Fibrosis. Genes 2020, 11, 603. [Google Scholar] [CrossRef]
- Sette, G.; Lo Cicero, S.; Blaconà, G.; Pierandrei, S.; Bruno, S.M.; Salvati, V.; Castelli, G.; Falchi, M.; Fabrizzi, B.; Cimino, G.; et al. Theratyping cystic fibrosis in vitro in ALI culture and organoid models generated from patient-derived nasal epithelial conditionally reprogrammed stem cells. Eur. Respir. J. 2021, 58, 2100908. [Google Scholar] [CrossRef]
- Amatngalim, G.D.; Rodenburg, L.W.; Aalbers, B.L.; Raeven, H.H.; Aarts, E.M.; Sarhane, D.; Spelier, S.; Lefferts, J.W.; AL Silva, I.; Nijenhuis, W.; et al. Measuring cystic fibrosis drug responses in organoids derived from 2D differentiated nasal epithelia. Life Sci. Alliance 2022, 5, e202101320. [Google Scholar] [CrossRef]
- Keating, D.; Marigowda, G.; Burr, L.; Daines, C.; Mall, M.A.; McKone, E.F.; Ramsey, B.W.; Rowe, S.M.; Sass, L.A.; Tullis, E.; et al. VX-445-Tezacaftor-Ivacaftor in Patients with Cystic Fibrosis and One or Two Phe508del Alleles. N. Engl. J. Med. 2018, 379, 1612–1620. [Google Scholar] [CrossRef]
- Anderson, J.D.; Liu, Z.; Odom, L.V.; Kersh, L.; Guimbellot, J.S. CFTR function and clinical response to modulators parallel nasal epithelial organoid swelling. Am. J. Physiol. Cell. Mol. Physiol. 2021, 321, L119–L129. [Google Scholar] [CrossRef] [PubMed]
- Geurts, M.H.; de Poel, E.; Amatngalim, G.D.; Oka, R.; Meijers, F.M.; Kruisselbrink, E.; van Mourik, P.; Berkers, G.; Groot, K.M.d.W.-D.; Michel, S.; et al. CRISPR-Based Adenine Editors Correct Nonsense Mutations in a Cystic Fibrosis Organoid Biobank. Cell Stem Cell 2020, 26, 503–510. [Google Scholar] [CrossRef]
- Wu, X.; Bos, I.S.T.; Conlon, T.M.; Ansari, M.; Verschut, V.; van der Koog, L.; Verkleij, L.A.; D’aMbrosi, A.; Matveyenko, A.; Schiller, H.B.; et al. A transcriptomics-guided drug target discovery strategy identifies receptor ligands for lung regeneration. Sci. Adv. 2022, 8, eabj9949. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Ng-Blichfeldt, J.-P.; Ota, C.; Ciminieri, C.; Ren, W.; Hiemstra, P.S.; Stolk, J.; Gosens, R.; Königshoff, M. Wnt/β-catenin signaling is critical for regenerative potential of distal lung epithelial progenitor cells in homeostasis and emphysema. STEM CELLS 2020, 38, 1467–1478. [Google Scholar] [CrossRef]
- Chan, L.L.Y.; Anderson, D.E.; Cheng, H.S.; Ivan, F.X.; Chen, S.; Kang, A.E.Z.; Foo, R.; Gamage, A.M.; Tiew, P.Y.; Koh, M.S.; et al. The establishment of COPD organoids to study host-pathogen interaction reveals enhanced viral fitness of SARS-CoV-2 in bronchi. Nat. Commun. 2022, 13, 7635. [Google Scholar] [CrossRef]
- Li, Y.; He, Y.; Zheng, Q.; Zhang, J.; Pan, X.; Zhang, X.; Yuan, H.; Wang, G.; Liu, X.; Zhou, X.; et al. Mitochondrial pyruvate carriers control airway basal progenitor cell function through glycolytic-epigenetic reprogramming. Cell Stem Cell 2024, 32, 105–120. [Google Scholar] [CrossRef] [PubMed]
- Bernardo, M.E.; Fibbe, W.E. Mesenchymal Stromal Cells: Sensors and Switchers of Inflammation. Cell Stem Cell 2013, 13, 392–402. [Google Scholar] [CrossRef] [PubMed]
- Weiss, A.R.R.; Dahlke, M.H. Immunomodulation by Mesenchymal Stem Cells (MSCs): Mechanisms of Action of Living, Apoptotic, and Dead MSCs. Front. Immunol. 2019, 10, 1191. [Google Scholar] [CrossRef]
- Reinders, M.E.; de Fijter, J.W.; Roelofs, H.; Bajema, I.M.; de Vries, D.K.; Schaapherder, A.F.; Claas, F.H.; van Miert, P.P.; Roelen, D.L.; van Kooten, C.; et al. Autologous Bone Marrow-Derived Mesenchymal Stromal Cells for the Treatment of Allograft Rejection After Renal Transplantation: Results of a Phase I Study. STEM CELLS Transl. Med. 2013, 2, 107–111. [Google Scholar] [CrossRef]
- Rodrigo, S.F.; van Ramshorst, J.; Hoogslag, G.E.; Boden, H.; Velders, M.A.; Cannegieter, S.C.; Roelofs, H.; Al Younis, I.; Dibbets-Schneider, P.; Fibbe, W.E.; et al. Intramyocardial Injection of Autologous Bone Marrow-Derived Ex Vivo Expanded Mesenchymal Stem Cells in Acute Myocardial Infarction Patients is Feasible and Safe up to 5 Years of Follow-up. J. Cardiovasc. Transl. Res. 2013, 6, 816–825. [Google Scholar] [CrossRef] [PubMed]
- Kruk, D.M.L.W.; Wisman, M.; Noordhoek, J.A.; Nizamoglu, M.; Jonker, M.R.; de Bruin, H.G.; Gomez, K.A.; Hacken, N.H.T.T.; Pouwels, S.D.; Heijink, I.H. Paracrine Regulation of Alveolar Epithelial Damage and Repair Responses by Human Lung-Resident Mesenchymal Stromal Cells. Cells 2021, 10, 2860. [Google Scholar] [CrossRef]
- Skronska-Wasek, W.; Mutze, K.; Baarsma, H.A.; Bracke, K.R.; Alsafadi, H.N.; Lehmann, M.; Costa, R.; Stornaiuolo, M.; Novellino, E.; Brusselle, G.G.; et al. Reduced Frizzled Receptor 4 Expression Prevents WNT/β-Catenin–driven Alveolar Lung Repair in Chronic Obstructive Pulmonary Disease. Am. J. Respir. Crit. Care Med. 2017, 196, 172–185. [Google Scholar] [CrossRef]
- Basil, M.C.; Cardenas-Diaz, F.L.; Kathiriya, J.J.; Morley, M.P.; Carl, J.; Brumwell, A.N.; Katzen, J.; Slovik, K.J.; Babu, A.; Zhou, S.; et al. Human distal airways contain a multipotent secretory cell that can regenerate alveoli. Nature 2022, 604, 120–126. [Google Scholar] [CrossRef]
- Costa, R.; Wagner, D.E.; Doryab, A.; De Santis, M.M.; Schorpp, K.; Rothenaigner, I.; Lehmann, M.; Baarsma, H.A.; Liu, X.; Schmid, O.; et al. A drug screen with approved compounds identifies amlexanox as a novel Wnt/β-catenin activator inducing lung epithelial organoid formation. Br. J. Pharmacol. 2021, 178, 4026–4041. [Google Scholar] [CrossRef]
- Pardal, R.; Clarke, M.F.; Morrison, S.J. Applying the principles of stem-cell biology to cancer. Nat. Rev. Cancer 2003, 3, 895–902. [Google Scholar] [CrossRef]
- Zhang, D.-G.; Jiang, A.-G.; Lu, H.-Y.; Zhang, L.-X.; Gao, X.-Y. Isolation, cultivation and identification of human lung adenocarcinoma stem cells. Oncol. Lett. 2014, 9, 47–54. [Google Scholar] [CrossRef]
- Di Liello, R.; Ciaramella, V.; Barra, G.; Venditti, M.; Della Corte, C.M.; Papaccio, F.; Sparano, F.; Viscardi, G.; Iacovino, M.L.; Minucci, S.; et al. Ex vivo lung cancer spheroids resemble treatment response of a patient with NSCLC to chemotherapy and immunotherapy: Case report and translational study. ESMO Open 2019, 4, e000536. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Wang, H.; Ding, Q.; Xing, Y.; Xu, Z.; Lu, C.; Luo, D.; Xu, L.; Xia, W.; Zhou, C.; et al. Establishment of patient-derived tumor spheroids for non-small cell lung cancer. PLOS ONE 2018, 13, e0194016. [Google Scholar] [CrossRef] [PubMed]
- Weeber, F.; van de Wetering, M.; Hoogstraat, M.; Dijkstra, K.K.; Krijgsman, O.; Kuilman, T.; Gadellaa-van Hooijdonk, C.G.M.; van der Velden, D.L.; Peeper, D.S.; Cuppen, E.P.J.G.; et al. Preserved genetic diversity in organoids cultured from biopsies of human colorectal cancer metastases. Proc. Natl. Acad. Sci. USA 2015, 112, 13308–13311. [Google Scholar] [CrossRef]
- Park, J.W.; Lee, J.K.; Sheu, K.M.; Wang, L.; Balanis, N.G.; Nguyen, K.; Smith, B.A.; Cheng, C.; Tsai, B.L.; Cheng, D.; et al. Reprogramming normal human epithelial tissues to a common, lethal neuroendocrine cancer lineage. Science 2018, 362, 91–95. [Google Scholar] [CrossRef]
- Kim, M.; Mun, H.; Sung, C.O.; Cho, E.J.; Jeon, H.-J.; Chun, S.-M.; Jung, D.J.; Shin, T.H.; Jeong, G.S.; Kim, D.K.; et al. Patient-derived lung cancer organoids as in vitro cancer models for therapeutic screening. Nat. Commun. 2019, 10, 3991. [Google Scholar] [CrossRef] [PubMed]
- Richer, A.L.; Cala, J.M.; O’Brien, K.; Carson, V.M.; Inge, L.J.; Whitsett, T.G. WEE1 Kinase Inhibitor AZD1775 Has Preclinical Efficacy in LKB1-Deficient Non–Small Cell Lung Cancer. Cancer Res. 2017, 77, 4663–4672. [Google Scholar] [CrossRef]
- Zhang, H.; Brainson, C.F.; Koyama, S.; Redig, A.J.; Chen, T.; Li, S.; Gupta, M.; Garcia-De-Alba, C.; Paschini, M.; Herter-Sprie, G.S.; et al. Lkb1 inactivation drives lung cancer lineage switching governed by Polycomb Repressive Complex 2. Nat. Commun. 2017, 8, 14922. [Google Scholar] [CrossRef]
- Lazarus, K.A.; Hadi, F.; Zambon, E.; Bach, K.; Santolla, M.-F.; Watson, J.K.; Correia, L.L.; Das, M.; Ugur, R.; Pensa, S.; et al. BCL11A interacts with SOX2 to control the expression of epigenetic regulators in lung squamous carcinoma. Nat. Commun. 2018, 9, 3327. [Google Scholar] [CrossRef]
- Caponigro, G.; Sellers, W.R. Advances in the preclinical testing of cancer therapeutic hypotheses. Nat. Rev. Drug Discov. 2011, 10, 179–187. [Google Scholar] [CrossRef]
- Barrera-Rodríguez, R.; Fuentes, J.M. Multidrug resistance characterization in multicellular tumour spheroids from two human lung cancer cell lines. Cancer Cell Int. 2015, 15, 47. [Google Scholar] [CrossRef]
- Hai, J.; Zhang, H.; Zhou, J.; Wu, Z.; Chen, T.; Papadopoulos, E.; Dowling, C.M.; Pyon, V.; Pan, Y.; Bin Liu, J.; et al. Generation of Genetically Engineered Mouse Lung Organoid Models for Squamous Cell Lung Cancers Allows for the Study of Combinatorial Immunotherapy. Clin. Cancer Res. 2020, 26, 3431–3442. [Google Scholar] [CrossRef]
- Wang, Y.; Jiang, T.; Qin, Z.; Jiang, J.; Wang, Q.; Yang, S.; Rivard, C.; Gao, G.; Ng, T.; Tu, M.; et al. HER2 exon 20 insertions in non-small-cell lung cancer are sensitive to the irreversible pan-HER receptor tyrosine kinase inhibitor pyrotinib. Ann. Oncol. 2019, 30, 447–455. [Google Scholar] [CrossRef]
- Takahashi, N.; Hoshi, H.; Higa, A.; Hiyama, G.; Tamura, H.; Ogawa, M.; Takagi, K.; Goda, K.; Okabe, N.; Muto, S.; et al. An In Vitro System for Evaluating Molecular Targeted Drugs Using Lung Patient-Derived Tumor Organoids. Cells 2019, 8, 481. [Google Scholar] [CrossRef] [PubMed]
- Dijkstra, K.K.; Cattaneo, C.M.; Weeber, F.; Chalabi, M.; Van De Haar, J.; Fanchi, L.F.; Slagter, M.; Van Der Velden, D.L.; Kaing, S.; Kelderman, S.; et al. Generation of Tumor-Reactive T Cells by Co-culture of Peripheral Blood Lymphocytes and Tumor Organoids. Cell 2018, 174, 1586–1598. [Google Scholar] [CrossRef] [PubMed]
- Sahin, U. Studying Tumor-ReacTive T Cells: A Personalized Organoid Model. Cell Stem Cell 2018, 23, 318–319. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Xu, C. Immune checkpoint signaling and cancer immunotherapy. Cell Res. 2020, 30, 660–669. [Google Scholar] [CrossRef]
- Yui, S.; Nakamura, T.; Sato, T.; Nemoto, Y.; Mizutani, T.; Zheng, X.; Ichinose, S.; Nagaishi, T.; Okamoto, R.; Tsuchiya, K.; et al. Functional engraftment of colon epithelium expanded in vitro from a single adult Lgr5+ stem cell. Nat. Med. 2012, 18, 618–623. [Google Scholar] [CrossRef]
- Yin, X.; Mead, B.E.; Safaee, H.; Langer, R.; Karp, J.M.; Levy, O. Engineering Stem Cell Organoids. Cell Stem Cell 2016, 18, 25–38. [Google Scholar] [CrossRef]
- Miura, Y.; Li, M.-Y.; Birey, F.; Ikeda, K.; Revah, O.; Thete, M.V.; Park, J.-Y.; Puno, A.; Lee, S.H.; Porteus, M.H.; et al. Generation of human striatal organoids and cortico-striatal assembloids from human pluripotent stem cells. Nat. Biotechnol. 2020, 38, 1421–1430. [Google Scholar] [CrossRef]
- Xiang, Y.; Tanaka, Y.; Patterson, B.; Kang, Y.-J.; Govindaiah, G.; Roselaar, N.; Cakir, B.; Kim, K.-Y.; Lombroso, A.P.; Hwang, S.-M.; et al. Fusion of Regionally Specified hPSC-Derived Organoids Models Human Brain Development and Interneuron Migration. Cell Stem Cell 2017, 21, 383–398. [Google Scholar] [CrossRef] [PubMed]
- Bin Ramli, M.N.; Lim, Y.S.; Koe, C.T.; Demircioglu, D.; Tng, W.; Gonzales, K.A.U.; Tan, C.P.; Szczerbinska, I.; Liang, H.; Soe, E.L.; et al. Human Pluripotent Stem Cell-Derived Organoids as Models of Liver Disease. Gastroenterology 2020, 159, 1471–1486. [Google Scholar] [CrossRef] [PubMed]
- Koike, H.; Iwasawa, K.; Ouchi, R.; Maezawa, M.; Giesbrecht, K.; Saiki, N.; Ferguson, A.; Kimura, M.; Thompson, W.L.; Wells, J.M.; et al. Modelling human hepato-biliary-pancreatic organogenesis from the foregut–midgut boundary. Nature 2019, 574, 112–116. [Google Scholar] [CrossRef]
- Shah, V.S.; Chivukula, R.R.; Lin, B.; Waghray, A.; Rajagopal, J. Cystic Fibrosis and the Cells of the Airway Epithelium: What Are Ionocytes and What Do They Do? Annu. Rev. Pathol. Mech. Dis. 2022, 17, 23–46. [Google Scholar] [CrossRef] [PubMed]
- Lamers, M.M.; van der Vaart, J.; Knoops, K.; Riesebosch, S.; Breugem, T.I.; Mykytyn, A.Z.; Beumer, J.; Schipper, D.; Bezstarosti, K.; Koopman, C.D.; et al. An organoid-derived bronchioalveolar model for SARS-CoV-2 infection of human alveolar type II-like cells. EMBO J. 2021, 40, e105912. [Google Scholar] [CrossRef]
- Wang, J.; Li, X.; Chen, H. Organoid models in lung regeneration and cancer. Cancer Lett. 2020, 475, 129–135. [Google Scholar] [CrossRef]
- Zhang, C.; Zhao, Z.; Rahim, N.A.A.; van Noort, D.; Yu, H. Towards a human-on-chip: Culturing multiple cell types on a chip with compartmentalized microenvironments. Lab Chip 2009, 9, 3185–3192. [Google Scholar] [CrossRef]
- Daly, A.C.; Davidson, M.D.; Burdick, J.A. 3D bioprinting of high cell-density heterogeneous tissue models through spheroid fusion within self-healing hydrogels. Nat. Commun. 2021, 12, 753. [Google Scholar] [CrossRef]
- Quadrato, G.; Nguyen, T.; Macosko, E.Z.; Sherwood, J.L.; Min Yang, S.; Berger, D.R.; Maria, N.; Scholvin, J.; Goldman, M.; Kinney, J.P.; et al. Cell diversity and network dynamics in photosensitive human brain organoids. Nature 2017, 545, 48–53. [Google Scholar] [CrossRef]
- Ng, W.H.; Johnston, E.K.; Tan, J.J.; Bliley, J.M.; Feinberg, A.W.; Stolz, D.B.; Sun, M.; Wijesekara, P.; Hawkins, F.; Kotton, D.N.; et al. Recapitulating human cardio-pulmonary co-development using simultaneous multilineage differentiation of pluripotent stem cells. eLife 2022, 11, e67872. [Google Scholar] [CrossRef]
- Qadir, A.S.; Das, S.; Nedunchezian, S.; Masuhara, K.; Desai, T.J.; Rehman, J.; Murthy, P.K.; Tsukasaki, Y.; Shao, L.; Malik, A.B. Physiological Modeling of the Vascularized Human Lung Organoid. Am. J. Respir. Cell Mol. Biol. 2025, 72, 354–363. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Chen, D.; Tian, L.; Pan, Z.; Long, H.; Chu, L.; Kong, W.; Yao, Q.; Ma, X.; Zhao, Y.; et al. Role of BMPR2 Mutation in Lung Organoid Differentiation. Biomedicines 2025, 13, 1623. [Google Scholar] [CrossRef] [PubMed]
- Huskin, G.; Chen, J.; Davis, T.; Jun, H.-W. Tissue-Engineered 3D In Vitro Disease Models for High-Throughput Drug Screening. Tissue Eng. Regen. Med. 2023, 20, 523–538. [Google Scholar] [CrossRef] [PubMed]



| ESC/iPSC-Derived Organoids | ASC-Derived Organoids | References | |
|---|---|---|---|
| Potential for differentiation | Multidirectional differentiation. | Limited ability to differentiate into organoids of specific tissue origin. | [12,13] |
| Cellular components | Complex cellular components, including mesenchymal, epithelial, and even endothelial components. | Single cellular component, mainly epithelial. | [1,14,15] |
| Cultivation conditions | Complex procedure; the procedure involves adding specific growth factor cocktails at each stage to form germ layers (endoderm, mesoderm, ectoderm) and incubating with growth factors, signaling molecules, and cytokines for cell differentiation and maturation. | Can be directly obtained from regenerative human adult tissues using a simpler procedure, but prior knowledge of culturing media from various tissues is required. | [3,15,16,17,18] |
| Incubation time | Generally take several months. | Generally shorter, about 3–4 weeks. | [15,16,17] |
| Technical maturity | The gene knockout and precise gene-editing techniques are better established. | The technology is relatively mature. | [19] |
| Maturity of organoids | Low maturity, most still have a fetal or neonatal component. | High maturity, closer to the adult tissue. | [16,18] |
| Self-renewal capacity | Exhibit a high proliferative capacity; however, they typically lose their potential for further expansion upon reaching the stage of terminal differentiation, ultimately failing to achieve full maturation. | Limited self-renewal capacity for a prolonged period of time. | [16,20] |
| Genetic stability | High genetic stability, but problems can occur with long-term cultures. | High genetic stability maintained during long-term expansion. | [9] |
| Other | Ability to induce neural lineage differentiation and reconstruct neural tube structures in vitro. | ASCs with stemness cannot be obtained from tissues such as the brain, heart, or pancreatic islets. | [15,21,22] |
| Application | ESC/iPSC-derived organoids are typically naïve and resemble fetal tissues, making them excellent models for studying early human organ development. | ASC-derived organoids offer a comprehensive view of adult tissue repair and viral infections, making them ideal for regenerative medicine and disease modeling. | [3,11,23,24] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Chu, L.; Chen, D.; Jiang, S.; Long, H.; Liu, X.; Chen, Y. Research Advances and Disease Modeling in Respiratory Organoids. Biomedicines 2026, 14, 221. https://doi.org/10.3390/biomedicines14010221
Chu L, Chen D, Jiang S, Long H, Liu X, Chen Y. Research Advances and Disease Modeling in Respiratory Organoids. Biomedicines. 2026; 14(1):221. https://doi.org/10.3390/biomedicines14010221
Chicago/Turabian StyleChu, Lanhe, Dian Chen, Simin Jiang, Huanyu Long, Xiaojuan Liu, and Yahong Chen. 2026. "Research Advances and Disease Modeling in Respiratory Organoids" Biomedicines 14, no. 1: 221. https://doi.org/10.3390/biomedicines14010221
APA StyleChu, L., Chen, D., Jiang, S., Long, H., Liu, X., & Chen, Y. (2026). Research Advances and Disease Modeling in Respiratory Organoids. Biomedicines, 14(1), 221. https://doi.org/10.3390/biomedicines14010221








