Voltage-Gated Sodium Channel Dysfunction in Epilepsy: Zebrafish Models for Therapeutics
Abstract
1. Voltage-Gated Ion Channels
2. VGSC Subunits
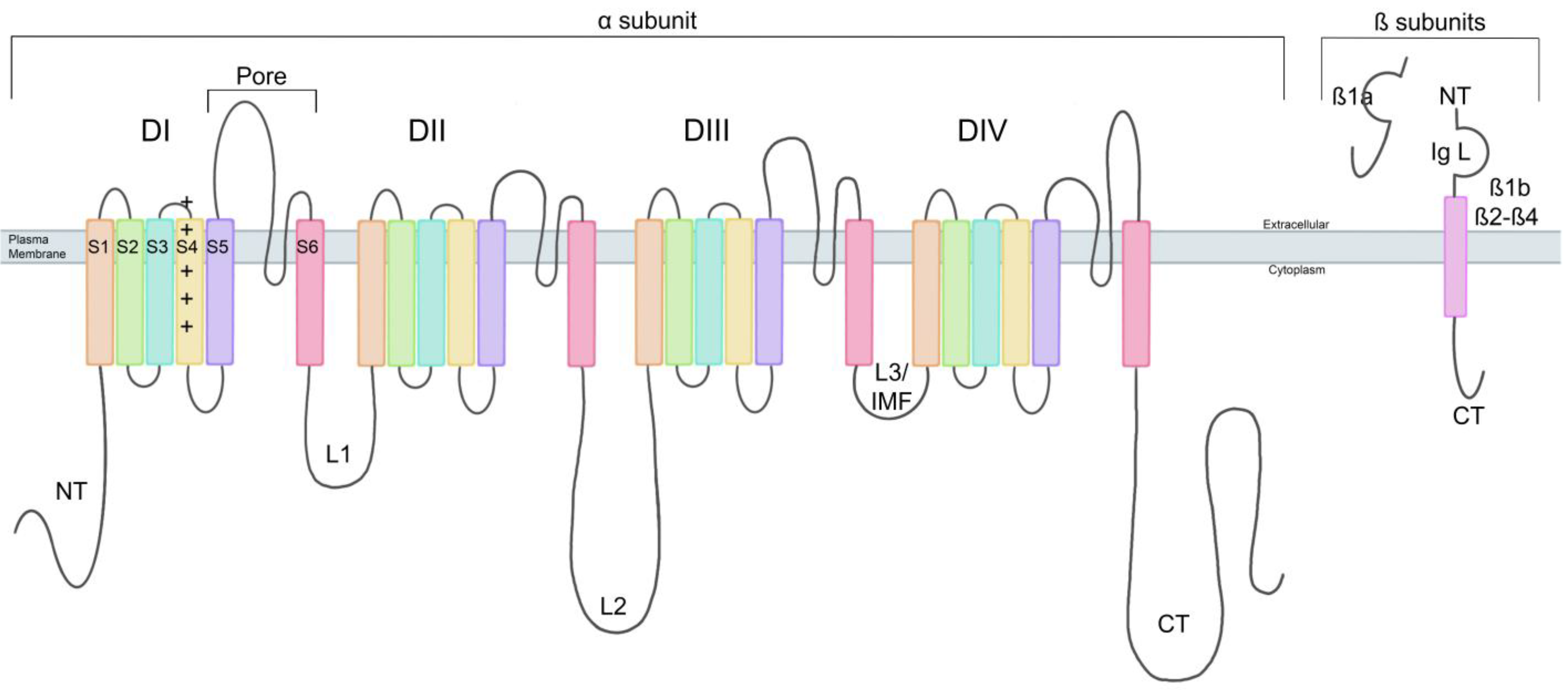
2.1. α Subunit Structure
2.2. α Subunit Function
2.3. β Subunit Structure
2.4. β Subunit Function
3. Sodium Channelopathies
4. Epilepsy: Incidence, Prevalence, and Types
5. Zebrafish as a Model for Epilepsy
Zebrafish SCNA Genes
6. Key SCNA Genes Implicated in Epilepsy
6.1. SCN1A
6.1.1. Dravet Syndrome
6.1.2. Genetic Epilepsy with Febrile Seizures Plus (GEFS+)
6.2. SCN2A
Early- and Late-Onset Infantile Epilepsies
6.3. SCN3A
6.4. SCN8A
7. Conclusion: VGSCs as Therapeutic Targets
8. Future Directions
Author Contributions
Funding
Conflicts of Interest
References
- Barker, B.S.; Young, G.T.; Soubrane, C.H.; Stephens, G.J.; Stevens, E.B.; Patel, M.K. Ion channels. In Conn’s Translational Neuroscience; Elsevier: Amsterdam, The Netherlands, 2017; pp. 11–43. [Google Scholar]
- Subramanyam, P.; Colecraft, H.M. Ion channel engineering: Perspectives and strategies. J. Mol. Biol. 2015, 427, 190–204. [Google Scholar] [CrossRef]
- Alberts, B.; Johnson, A.; Lewis, J.; Raff, M.; Roberts, K.; Walter, P. Ion channels and the electrical properties of membranes. In Molecular Biology of the Cell, 4th ed.; Garland Science: New York, NY, USA, 2002. [Google Scholar]
- Wang, J.; Ou, S.-W.; Wang, Y.-J. Distribution and function of voltage-gated sodium channels in the nervous system. Channels 2017, 11, 534–554. [Google Scholar] [CrossRef]
- Kim, D.M.; Nimigean, C.M. Voltage-gated potassium channels: A structural examination of selectivity and gating. Cold Spring Harb. Perspect. Biol. 2016, 8, a029231. [Google Scholar] [CrossRef]
- DeCoursey, T.E.; Hosler, J. Philosophy of voltage-gated proton channels. J. R. Soc. Interface 2014, 11, 20130799. [Google Scholar] [CrossRef]
- Payandeh, J.; Minor, D.L., Jr. Bacterial voltage-gated sodium channels (BacNaVs) from the soil, sea, and salt lakes enlighten molecular mechanisms of electrical signaling and pharmacology in the brain and heart. J. Mol. Biol. 2015, 427, 3–30. [Google Scholar] [CrossRef]
- Kim, J.-B. Channelopathies. Korean J. Pediatr. 2014, 57, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Bouza, A.A.; Isom, L.L. Voltage-gated sodium channel β subunits and their related diseases. Volt.-Gated Sodium Channels: Struct. Funct. Channelopathies 2018, 246, 423–450. [Google Scholar]
- Wood, J.N.; Iseppon, F. Sodium channels. Brain Neurosci. Adv. 2018, 2, 2398212818810684. [Google Scholar] [CrossRef]
- Lai, H.C.; Jan, L.Y. The distribution and targeting of neuronal voltage-gated ion channels. Nat. Rev. Neurosci. 2006, 7, 548–562. [Google Scholar] [CrossRef] [PubMed]
- Namadurai, S.; Yereddi, N.R.; Cusdin, F.S.; Huang, C.L.-H.; Chirgadze, D.Y.; Jackson, A.P. A new look at sodium channel β subunits. Open Biol. 2015, 5, 140192. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Sandoval, A.L.; Hernández-Plata, E.; Gomora, J.C. Voltage-gated sodium channels: From roles and mechanisms in the metastatic cell behavior to clinical potential as therapeutic targets. Front. Pharmacol. 2023, 14, 1206136. [Google Scholar] [CrossRef] [PubMed]
- Strege, P.R.; Cowan, L.M.; Alcaino, C.; Mazzone, A.; Ahern, C.A.; Milescu, L.S.; Farrugia, G.; Beyder, A. Mechanosensitive pore opening of a prokaryotic voltage-gated sodium channel. Elife 2023, 12, e79271. [Google Scholar] [CrossRef]
- Jiang, D.; Banh, R.; Gamal El-Din, T.M.; Tonggu, L.; Lenaeus, M.J.; Pomès, R.; Zheng, N.; Catterall, W.A. Open-state structure and pore gating mechanism of the cardiac sodium channel. Cell 2021, 184, 5151–5162.e5111. [Google Scholar] [CrossRef] [PubMed]
- Eijkelkamp, N.; Linley, J.E.; Baker, M.D.; Minett, M.S.; Cregg, R.; Werdehausen, R.; Rugiero, F.; Wood, J.N. Neurological perspectives on voltage-gated sodium channels. Brain 2012, 135, 2585–2612. [Google Scholar] [CrossRef] [PubMed]
- Patlak, J. Molecular kinetics of voltage-dependent Na+ channels. Physiol. Rev. 1991, 71, 1047–1080. [Google Scholar] [CrossRef]
- Augustine, G.J.; Groh, J.M.; Huettel, S.A.; LaMantia, A.-S.; White, L.E. Neuroscience; Oxford University Press: Oxford, UK, 2024. [Google Scholar]
- Groome, J.; Lehmann-Horn, F.; Holzherr, B. Open- and closed-state fast inactivation in sodium channels. Channels 2011, 5, 65–78. [Google Scholar] [CrossRef]
- Hodgkin, A.L.; Huxley, A.F. Action Potentials Recorded from Inside a Nerve Fibre. Nature 1939, 144, 710–711. [Google Scholar] [CrossRef]
- Bouza, A.A.; Edokobi, N.; Hodges, S.L.; Pinsky, A.M.; Offord, J.; Piao, L.; Zhao, Y.T.; Lopatin, A.N.; Lopez-Santiago, L.F.; Isom, L.L. Sodium channel β1 subunits participate in regulated intramembrane proteolysis-excitation coupling. JCI Insight 2021, 6, e141776. [Google Scholar] [CrossRef]
- O’Malley, H.A.; Isom, L.L. Sodium channel β subunits: Emerging targets in channelopathies. Annu. Rev. Physiol. 2015, 77, 481–504. [Google Scholar] [CrossRef]
- Edokobi, N.; Isom, L.L. Voltage-gated sodium channel β1/β1B subunits regulate cardiac physiology and pathophysiology. Front. Physiol. 2018, 9, 351. [Google Scholar] [CrossRef]
- Hull, J.M.; Isom, L.L. Voltage-gated sodium channel β subunits: The power outside the pore in brain development and disease. Neuropharmacology 2018, 132, 43–57. [Google Scholar] [CrossRef]
- Xiao, Y.; Barbosa, C.; Pei, Z.; Xie, W.; Strong, J.A.; Zhang, J.-M.; Cummins, T.R. Increased resurgent sodium currents in Nav1. 8 contribute to nociceptive sensory neuron hyperexcitability associated with peripheral neuropathies. J. Neurosci. 2019, 39, 1539–1550. [Google Scholar] [CrossRef] [PubMed]
- Lewis, A.H.; Raman, I.M. Resurgent current of voltage-gated Na+ channels. J. Physiol. 2014, 592, 4825–4838. [Google Scholar] [CrossRef] [PubMed]
- Zemel, B.M.; Nevue, A.A.; Dagostin, A.; Lovell, P.V.; Mello, C.V.; von Gersdorff, H. Resurgent Na+ currents promote ultrafast spiking in projection neurons that drive fine motor control. Nat. Commun. 2021, 12, 6762. [Google Scholar] [CrossRef]
- George, A.L. Lessons learned from genetic testing for channelopathies. Lancet Neurol. 2014, 13, 1068–1070. [Google Scholar] [CrossRef] [PubMed]
- Bagal, S.K.; Brown, A.D.; Cox, P.J.; Omoto, K.; Owen, R.M.; Pryde, D.C.; Sidders, B.; Skerratt, S.E.; Stevens, E.B.; Storer, R.I. Ion channels as therapeutic targets: A drug discovery perspective. J. Med. Chem. 2013, 56, 593–624. [Google Scholar] [CrossRef]
- Frolov, R.V.; Weckström, M. Ion channels as therapeutic targets, part a. Adv. Protein Chem. Struct. Biol. 2016, 103, 1–386. [Google Scholar]
- Kapur, J. Sodium channel mutations in GEFS+ produce persistent inward current. Epilepsy Curr. 2002, 2, 149–150. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ramakrishna, H.; O’Hare, M.; Mookadam, F.; Gutsche, J.T.; Shah, R.; Augoustides, J.G. Sudden cardiac death and disorders of the QT interval: Anesthetic implications and focus on perioperative management. J. Cardiothorac. Vasc. Anesth. 2015, 29, 1723–1733. [Google Scholar] [CrossRef]
- Spillane, J.; Kullmann, D.; Hanna, M. Genetic neurological channelopathies: Molecular genetics and clinical phenotypes. J. Neurol. Neurosurg. Psychiatry 2016, 87, 37–48. [Google Scholar] [CrossRef]
- Imbrici, P.; Liantonio, A.; Camerino, G.M.; De Bellis, M.; Camerino, C.; Mele, A.; Giustino, A.; Pierno, S.; De Luca, A.; Tricarico, D. Therapeutic approaches to genetic ion channelopathies and perspectives in drug discovery. Front. Pharmacol. 2016, 7, 121. [Google Scholar] [CrossRef]
- Meisler, M.H.; Hill, S.F.; Yu, W. Sodium channelopathies in neurodevelopmental disorders. Nat. Rev. Neurosci. 2021, 22, 152–166. [Google Scholar] [CrossRef]
- Kaplan, D.I.; Isom, L.L.; Petrou, S. Role of sodium channels in epilepsy. Cold Spring Harb. Perspect. Med. 2016, 6, a022814. [Google Scholar] [CrossRef]
- Beghi, E. The epidemiology of epilepsy. Neuroepidemiology 2020, 54, 185–191. [Google Scholar] [CrossRef]
- Fisher, R.S.; Acharya, J.N.; Baumer, F.M.; French, J.A.; Parisi, P.; Solodar, J.H.; Szaflarski, J.P.; Thio, L.L.; Tolchin, B.; Wilkins, A.J. Visually sensitive seizures: An updated review by the Epilepsy Foundation. Epilepsia 2022, 63, 739–768. [Google Scholar] [CrossRef]
- Giourou, E.; Stavropoulou-Deli, A.; Giannakopoulou, A.; Kostopoulos, G.K.; Koutroumanidis, M. Introduction to Epilepsy and Related Brain Disorders. In Cyberphysical Systems for Epilepsy and Related Brain Disorders: Multi-Parametric Monitoring and Analysis for Diagnosis and Optimal Disease Management; Voros, N.S., Antonopoulos, C.P., Eds.; Springer International Publishing: Cham, Swizerland, 2015; pp. 11–38. [Google Scholar]
- Stafstrom, C.E. Epilepsy: A Review of Selected Clinical Syndromes and Advances in Basic Science. J. Cereb. Blood Flow Metab. 2006, 26, 983–1004. [Google Scholar] [CrossRef] [PubMed]
- Scharfman, H.E. The neurobiology of epilepsy. Curr. Neurol. Neurosci. Rep. 2007, 7, 348–354. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Li, H.; Ma, T.; Han, S.; Zhao, Q. Biological function simulation in neuromorphic devices: From synapse and neuron to behavior. Sci. Technol. Adv. Mater. 2023, 24, 2183712. [Google Scholar] [CrossRef]
- Bechi, G.; Scalmani, P.; Schiavon, E.; Rusconi, R.; Franceschetti, S.; Mantegazza, M. Pure haploinsufficiency for Dravet syndrome NaV1. 1 (SCN1A) sodium channel truncating mutations. Epilepsia 2012, 53, 87–100. [Google Scholar] [CrossRef]
- Bayat, A.; Bayat, M.; Rubboli, G.; Møller, R.S. Epilepsy Syndromes in the First Year of Life and Usefulness of Genetic Testing for Precision Therapy. Genes 2021, 12, 1051. [Google Scholar] [CrossRef] [PubMed]
- Ighodaro, E.T.; Maini, K.; Arya, K.; Sharma, S. Focal onset seizure. In StatPearls [Internet]; StatPearls Publishing: Treasure Island, FL, USA, 2023. [Google Scholar]
- Milligan, T.A. Epilepsy: A Clinical Overview. Am. J. Med. 2021, 134, 840–847. [Google Scholar] [CrossRef]
- Kumar, A.; Ighodaro, E.; Sharma, S. Focal Impaired Awareness Seizure. In StatPearls [Internet]; StatPearls Publishing: Treasure Island, FL, USA, 2024. [Google Scholar]
- Mullen, S.A.; Berkovic, S.F. Genetic generalized epilepsies. Epilepsia 2018, 59, 1148–1153. [Google Scholar] [CrossRef]
- Thakran, S.; Guin, D.; Singh, P.; Singh, P.; Kukal, S.; Rawat, C.; Yadav, S.; Kushwaha, S.S.; Srivastava, A.K.; Hasija, Y.; et al. Genetic Landscape of Common Epilepsies: Advancing towards Precision in Treatment. Int. J. Mol. Sci. 2020, 21, 7784. [Google Scholar] [CrossRef]
- Rastin, C.; Schenkel, L.C.; Sadikovic, B. Complexity in genetic epilepsies: A comprehensive review. Int. J. Mol. Sci. 2023, 24, 14606. [Google Scholar] [CrossRef]
- Loiseau, J.; Loiseau, P.; Guyot, M.; Duche, B.; Dartigues, J.F.; Aublet, B. Survey of seizure disorders in the French southwest. I. Incidence of epileptic syndromes. Epilepsia 1990, 31, 391–396. [Google Scholar] [CrossRef] [PubMed]
- Stephen, L.J.; Brodie, M.J. Pharmacological Management of the Genetic Generalised Epilepsies in Adolescents and Adults. CNS Drugs 2020, 34, 147–161. [Google Scholar] [CrossRef] [PubMed]
- Atalar, A.C.; Baykan, B. Different prognostic patterns in epilepsies and considerations about the denotations of atypical patterns. Arch. Neuropsychiatry 2022, 59, 68. [Google Scholar] [CrossRef]
- Yesudhas, D.; Anwar, M.A.; Panneerselvam, S.; Kim, H.K.; Choi, S. Evaluation of Sox2 binding affinities for distinct DNA patterns using steered molecular dynamics simulation. FEBS Open Bio 2017, 7, 1750–1767. [Google Scholar] [CrossRef]
- Sills, G.J. Mechanisms of action of antiepileptic drugs. Epilepsy 2011, 36, 295–303. [Google Scholar]
- Hakami, T. Neuropharmacology of antiseizure drugs. Neuropsychopharmacol. Rep. 2021, 41, 336–351. [Google Scholar] [CrossRef]
- Brunklaus, A.; Brünger, T.; Feng, T.; Fons, C.; Lehikoinen, A.; Panagiotakaki, E.; Vintan, M.-A.; Symonds, J.; Andrew, J.; Arzimanoglou, A.; et al. The gain of function SCN1A disorder spectrum: Novel epilepsy phenotypes and therapeutic implications. Brain 2022, 145, 3816–3831. [Google Scholar] [CrossRef]
- Catterall, W.A.; Swanson, T.M. Structural basis for pharmacology of voltage-gated sodium and calcium channels. Mol. Pharmacol. 2015, 88, 141–150. [Google Scholar] [CrossRef]
- Howe, K.; Clark, M.D.; Torroja, C.F.; Torrance, J.; Berthelot, C.; Muffato, M.; Collins, J.E.; Humphray, S.; McLaren, K.; Matthews, L.; et al. Correction: Corrigendum: The zebrafish reference genome sequence and its relationship to the human genome. Nature 2014, 505, 248. [Google Scholar] [CrossRef]
- Kundap, U.P.; Kumari, Y.; Othman, I.; Shaikh, M.F. Zebrafish as a model for epilepsy-induced cognitive dysfunction: A pharmacological, biochemical and behavioral approach. Front. Pharmacol. 2017, 8, 515. [Google Scholar] [CrossRef] [PubMed]
- Kolesnikova, T.O.; Demin, K.A.; Costa, F.V.; Zabegalov, K.N.; de Abreu, M.S.; Gerasimova, E.V.; Kalueff, A.V. Towards Zebrafish Models of CNS Channelopathies. Int. J. Mol. Sci. 2022, 23, 13979. [Google Scholar] [CrossRef]
- Xu, N.; LaGrow, T.J.; Anumba, N.; Lee, A.; Zhang, X.; Yousefi, B.; Bassil, Y.; Clavijo, G.P.; Khalilzad Sharghi, V.; Maltbie, E. Functional connectivity of the brain across rodents and humans. Front. Neurosci. 2022, 16, 816331. [Google Scholar] [CrossRef]
- Kalueff, A.V.; Gebhardt, M.; Stewart, A.M.; Cachat, J.M.; Brimmer, M.; Chawla, J.S.; Craddock, C.; Kyzar, E.J.; Roth, A.; Landsman, S.; et al. Towards a comprehensive catalog of zebrafish behavior 1.0 and beyond. Zebrafish 2013, 10, 70–86. [Google Scholar] [CrossRef]
- Cho, S.J.; Park, E.; Baker, A.; Reid, A.Y. Age Bias in Zebrafish Models of Epilepsy: What Can We Learn From Old Fish? Front. Cell Dev. Biol. 2020, 8, 573303. [Google Scholar] [CrossRef] [PubMed]
- Baraban, S.C.; Taylor, M.; Castro, P.; Baier, H. Pentylenetetrazole induced changes in zebrafish behavior, neural activity and c-fos expression. Neuroscience 2005, 131, 759–768. [Google Scholar] [CrossRef] [PubMed]
- D’Amora, M.; Galgani, A.; Marchese, M.; Tantussi, F.; Faraguna, U.; De Angelis, F.; Giorgi, F.S. Zebrafish as an innovative tool for epilepsy modeling: State of the art and potential future directions. Int. J. Mol. Sci. 2023, 24, 7702. [Google Scholar] [CrossRef]
- Stewart, A.; Wong, K.; Cachat, J.; Gaikwad, S.; Kyzar, E.; Wu, N.; Hart, P.; Piet, V.; Utterback, E.; Elegante, M. Zebrafish models to study drug abuse-related phenotypes. Prog. Neurobiol. 2011, 22, 95–105. [Google Scholar] [CrossRef]
- Gawel, K.; Turski, W.A.; van der Ent, W.; Mathai, B.J.; Kirstein-Smardzewska, K.J.; Simonsen, A.; Esguerra, C.V. Phenotypic characterization of larval zebrafish (Danio rerio) with partial knockdown of the cacna1a gene. Mol. Neurobiol. 2020, 57, 1904–1916. [Google Scholar] [CrossRef]
- Zayat, V.; Szlendak, R.; Hoffman-Zacharska, D. Concise Review: Stem Cell Models of SCN1A-Related Encephalopathies-Current Perspective and Future Therapies. Cells 2022, 11, 3119. [Google Scholar] [CrossRef]
- Office of Laboratory Animal Welfare; FAQs. PHS Policy on Humane Care and Use of Laboratory Animals, FAQ’s. Available online: https://olaw.nih.gov/faqs#/guidance/faqs (accessed on 29 July 2025).
- Guidelines for the Euthanasia of Animals. Available online: https://www.avma.org/resources-tools/avma-policies/avma-guidelines-euthanasia-animals (accessed on 29 July 2025).
- Hardison, R.C. Comparative genomics. PLoS Biol. 2003, 1, e58. [Google Scholar] [CrossRef]
- Liu, J.; Baraban, S.C. Network properties revealed during multi-scale calcium imaging of seizure activity in zebrafish. eNeuro 2019, 6, ENEURO.0041-19.2019. [Google Scholar] [CrossRef]
- Baraban, S.C. Forebrain electrophysiological recording in larval zebrafish. J. Vis. Exp. 2013, 24, 50104. [Google Scholar] [CrossRef]
- Zhang, R.-W.; Du, J.-L. In vivo whole-cell patch-clamp recording in the zebrafish brain. In Zebrafish: Methods and Protocols; Springer: Berlin/Heidelberg, Germany, 2016; pp. 281–291. [Google Scholar]
- Kawatani, M.; Yamashita, T. In Vivo Whole-Cell Recording from the Mouse Brain. In Cerebral Cortex Development: Methods and Protocols; Springer: Berlin/Heidelberg, Germany, 2024; pp. 245–257. [Google Scholar]
- Stephan, M.; Volkmann, P.; Rossner, M.J. Assessing behavior and cognition in rodents, nonhuman primates, and humans: Where are the limits of translation? Dialogues Clin. Neurosci. 2019, 21, 249–259. [Google Scholar] [CrossRef]
- Lee, Y.; Lee, K.J.; Jang, J.-W.; Lee, S.-i.; Kim, S. An EEG system to detect brain signals from multiple adult zebrafish. Biosens. Bioelectron. 2020, 164, 112315. [Google Scholar] [CrossRef] [PubMed]
- Carter, M.; Shieh, J. Chapter 4—Electrophysiology. In Guide to Research Techniques in Neuroscience, 2nd ed.; Carter, M., Shieh, J., Eds.; Academic Press: San Diego, CA, USA, 2015; pp. 89–115. [Google Scholar]
- Pinion, J.; Walsh, C.; Goodfellow, M.; Randall, A.D.; Tyler, C.R.; Winter, M.J. Differential Electrographic Signatures Generated by Mechanistically-Diverse Seizurogenic Compounds in the Larval Zebrafish Brain. eNeuro 2022, 9, ENEURO.0337-21.2022. [Google Scholar] [CrossRef] [PubMed]
- Eimon, P.M.; Ghannad-Rezaie, M.; De Rienzo, G.; Allalou, A.; Wu, Y.; Gao, M.; Roy, A.; Skolnick, J.; Yanik, M.F. Brain activity patterns in high-throughput electrophysiology screen predict both drug efficacies and side effects. Nat. Commun. 2018, 9, 219. [Google Scholar] [CrossRef]
- Kettunen, P. Calcium imaging in the zebrafish. Calcium Signal. 2019, 1131, 901–942. [Google Scholar]
- Cozzolino, O.; Sicca, F.; Paoli, E.; Trovato, F.; Santorelli, F.M.; Ratto, G.M.; Marchese, M. Evolution of Epileptiform Activity in Zebrafish by Statistical-Based Integration of Electrophysiology and 2-Photon Ca(2+) Imaging. Cells 2020, 9, 769. [Google Scholar] [CrossRef]
- Escayg, A.; Goldin, A.L. Sodium channel SCN1A and epilepsy: Mutations and mechanisms. Epilepsia 2010, 51, 1650–1658. [Google Scholar] [CrossRef]
- Butler, K.M.; da Silva, C.; Shafir, Y.; Weisfeld-Adams, J.D.; Alexander, J.J.; Hegde, M.; Escayg, A. De novo and inherited SCN8A epilepsy mutations detected by gene panel analysis. Epilepsy Res. 2017, 129, 17–25. [Google Scholar] [CrossRef]
- Meisler, M.H.; Kearney, J.A. Sodium channel mutations in epilepsy and other neurological disorders. J. Clin. Investig. 2005, 115, 2010–2017. [Google Scholar] [CrossRef]
- Novak, A.E.; Taylor, A.D.; Pineda, R.H.; Lasda, E.L.; Wright, M.A.; Ribera, A.B. Embryonic and larval expression of zebrafish voltage-gated sodium channel α-subunit genes. Dev. Dyn. An. Off. Publ. Am. Assoc. Anat. 2006, 235, 1962–1973. [Google Scholar] [CrossRef] [PubMed]
- Chopra, S.S.; Watanabe, H.; Smith, A.H.; Zhong, T.P.; Roden, D.M. Abstract 1701: Sodium Channel Beta Subunits Modulate Heart Rate, Drug Sensitivity, and Development in Zebrafish Embryos. Circulation 2006, 114, II_332. [Google Scholar] [CrossRef]
- Chopra, S.S.; Stroud, D.M.; Watanabe, H.; Bennett, J.S.; Burns, C.G.; Wells, K.S.; Yang, T.; Zhong, T.P.; Roden, D.M. Voltage-gated sodium channels are required for heart development in zebrafish. Circ. Res. 2010, 106, 1342–1350. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.-H.; Chen, Y.-H.; Huang, F.-L.; Chang, C.-H.; Chang, Y.-F.; Tsay, H.-J. Multiple regulatory elements mediating neuronal-specific expression of zebrafish sodium channel gene, Scn8aa. Dev. Dyn. 2008, 237, 2554–2565. [Google Scholar] [CrossRef]
- Griffin, A.; Carpenter, C.; Liu, J.; Paterno, R.; Grone, B.; Hamling, K.; Moog, M.; Dinday, M.T.; Figueroa, F.; Anvar, M. Phenotypic analysis of catastrophic childhood epilepsy genes. Commun. Biol. 2021, 4, 680. [Google Scholar] [CrossRef]
- Musto, E.; Gardella, E.; Møller, R.S. Recent advances in treatment of epilepsy-related sodium channelopathies. Eur. J. Paediatr. Neurol. 2020, 24, 123–128. [Google Scholar] [CrossRef] [PubMed]
- Liang, L.; Fazel Darbandi, S.; Pochareddy, S.; Gulden, F.O.; Gilson, M.C.; Sheppard, B.K.; Sahagun, A.; An, J.-Y.; Werling, D.M.; Rubenstein, J.L.R.; et al. Developmental dynamics of voltage-gated sodium channel isoform expression in the human and mouse brain. Genome Med. 2021, 13, 135. [Google Scholar] [CrossRef]
- Sanders, S.J.; Campbell, A.J.; Cottrell, J.R.; Moller, R.S.; Wagner, F.F.; Auldridge, A.L.; Bernier, R.A.; Catterall, W.A.; Chung, W.K.; Empfield, J.R.; et al. Progress in Understanding and Treating SCN2A-Mediated Disorders. Trends Neurosci. 2018, 41, 442–456. [Google Scholar] [CrossRef]
- Pelkey, K.A.; Chittajallu, R.; Craig, M.T.; Tricoire, L.; Wester, J.C.; McBain, C.J. Hippocampal GABAergic inhibitory interneurons. Physiol. Rev. 2017, 97, 1619–1747. [Google Scholar] [CrossRef]
- Weuring, W.J.; Hoekman, J.W.; Braun, K.P.J.; Koeleman, B.P.C. Genetic and Functional Differences between Duplicated Zebrafish Genes for Human SCN1A. Cells 2022, 11, 454. [Google Scholar] [CrossRef]
- Das, A.; Zhu, B.; Xie, Y.; Zeng, L.; Pham, A.T.; Neumann, J.C.; Safrina, O.; Benavides, D.R.; MacGregor, G.R.; Schutte, S.S.; et al. Interneuron Dysfunction in a New Mouse Model of SCN1A GEFS. eNeuro 2021, 8, ENEURO.0394-20.2021. [Google Scholar] [CrossRef]
- Ding, J.; Wang, L.; Jin, Z.; Qiang, Y.; Li, W.; Wang, Y.; Zhu, C.; Jiang, S.; Xiao, L.; Hao, X.; et al. Do All Roads Lead to Rome? Genes Causing Dravet Syndrome and Dravet Syndrome-Like Phenotypes. Front. Neurol. 2022, 13, 2380. [Google Scholar] [CrossRef]
- Catterall, W.A. Chapter 4—Dravet Syndrome: A Sodium Channel Interneuronopathy. In Ion Channels in Health and Disease; Pitt, G.S., Ed.; Academic Press: Boston, MA, USA, 2016; pp. 85–111. [Google Scholar]
- Chancey, J.H.; Howard, M.A. Synaptic Integration in CA1 Pyramidal Neurons Is Intact despite Deficits in GABAergic Transmission in the Scn1a Haploinsufficiency Mouse Model of Dravet Syndrome. eNeuro 2022, 9, ENEURO.0080-22.2022. [Google Scholar] [CrossRef]
- Baraban, S.C.; Dinday, M.T.; Hortopan, G.A. Drug screening in Scn1a zebrafish mutant identifies clemizole as a potential Dravet syndrome treatment. Nat. Commun. 2013, 4, 2410. [Google Scholar] [CrossRef] [PubMed]
- Han, S.; Tai, C.; Westenbroek, R.E.; Yu, F.H.; Cheah, C.S.; Potter, G.B.; Rubenstein, J.L.; Scheuer, T.; de la Iglesia, H.O.; Catterall, W.A. Autistic-like behaviour in Scn1a+/- mice and rescue by enhanced GABA-mediated neurotransmission. Nature 2012, 489, 385–390. [Google Scholar] [CrossRef] [PubMed]
- Bertuccelli, M.; Verheyen, K.; Hallemans, A.; Sander, J.W.; Ragona, F.; Bisiacchi, P.; Masiero, S.; Del Felice, A. Deconstructing Dravet syndrome neurocognitive development: A scoping review. Epilepsia 2021, 62, 874–887. [Google Scholar] [CrossRef]
- Dyment, D.A.; Schock, S.C.; Deloughery, K.; Tran, M.H.; Ure, K.; Nutter, L.M.J.; Creighton, A.; Yuan, J.; Banderali, U.; Comas, T.; et al. Electrophysiological Alterations of Pyramidal Cells and Interneurons of the CA1 Region of the Hippocampus in a Novel Mouse Model of Dravet Syndrome. Genetics 2020, 215, 1055–1066. [Google Scholar] [CrossRef]
- Weuring, W.J.; Singh, S.; Volkers, L.; Rook, M.B.; van ‘t Slot, R.H.; Bosma, M.; Inserra, M.; Vetter, I.; Verhoeven-Duif, N.M.; Braun, K.P.J.; et al. NaV1.1 and NaV1.6 selective compounds reduce the behavior phenotype and epileptiform activity in a novel zebrafish model for Dravet Syndrome. PLoS ONE 2020, 15, e0219106. [Google Scholar] [CrossRef]
- Yamakawa, K.; Meisler, M.H.; Isom, L.L. Sodium Channelopathies in Human and Animal Models of Epilepsy and Neurodevelopmental Disorders. In Jasper’s Basic Mechanisms of the Epilepsies; Noebels, J.L., Avoli, M., Rogawski, M.A., Vezzani, A., Delgado-Escueta, A.V., Eds.; Oxford University Press: New York, NY, USA, 2024; pp. 881–920. [Google Scholar]
- Richards, K.L.; Milligan, C.J.; Richardson, R.J.; Jancovski, N.; Grunnet, M.; Jacobson, L.H.; Undheim, E.A.B.; Mobli, M.; Chow, C.Y.; Herzig, V.; et al. Selective Na(V)1.1 activation rescues Dravet syndrome mice from seizures and premature death. Proc. Natl. Acad. Sci. USA 2018, 115, E8077–E8085. [Google Scholar] [CrossRef]
- Colasante, G.; Lignani, G.; Brusco, S.; Di Berardino, C.; Carpenter, J.; Giannelli, S.; Valassina, N.; Bido, S.; Ricci, R.; Castoldi, V.; et al. dCas9-Based Scn1a Gene Activation Restores Inhibitory Interneuron Excitability and Attenuates Seizures in Dravet Syndrome Mice. Mol. Ther. 2020, 28, 235–253. [Google Scholar] [CrossRef]
- Sourbron, J.; Smolders, I.; de Witte, P.; Lagae, L. Pharmacological Analysis of the Anti-epileptic Mechanisms of Fenfluramine in scn1a Mutant Zebrafish. Front. Pharmacol. 2017, 8, 191. [Google Scholar] [CrossRef]
- Gogou, M.; Cross, J.H. Fenfluramine as antiseizure medication for epilepsy. Dev. Med. Child Neurol. 2021, 63, 899–907. [Google Scholar] [CrossRef] [PubMed]
- Sourbron, J.; Lagae, L. Serotonin receptors in epilepsy: Novel treatment targets? Epilepsia Open 2022, 7, 231–246. [Google Scholar] [CrossRef] [PubMed]
- Samanta, D. Fenfluramine: A Review of Pharmacology, Clinical Efficacy, and Safety in Epilepsy. Children 2022, 9, 1159. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Kecskés, A.; Copmans, D.; Langlois, M.; Crawford, A.D.; Ceulemans, B.; Lagae, L.; de Witte, P.A.; Esguerra, C.V. Pharmacological characterization of an antisense knockdown zebrafish model of Dravet syndrome: Inhibition of epileptic seizures by the serotonin agonist fenfluramine. PLoS ONE 2015, 10, e0125898. [Google Scholar] [CrossRef]
- Morales-Lázaro, S.L.; González-Ramírez, R.; Rosenbaum, T. Molecular Interplay Between the Sigma-1 Receptor, Steroids, and Ion Channels. Front. Pharmacol. 2019, 10, 419. [Google Scholar] [CrossRef]
- Martin, P.; Reeder, T.; Sourbron, J.; de Witte, P.A.M.; Gammaitoni, A.R.; Galer, B.S. An Emerging Role for Sigma-1 Receptors in the Treatment of Developmental and Epileptic Encephalopathies. Int. J. Mol. Sci. 2021, 22, 8416. [Google Scholar] [CrossRef]
- Su, T.P.; Su, T.C.; Nakamura, Y.; Tsai, S.Y. The Sigma-1 Receptor as a Pluripotent Modulator in Living Systems. Trends Pharmacol. Sci. 2016, 37, 262–278. [Google Scholar] [CrossRef]
- Penke, B.; Fulop, L.; Szucs, M.; Frecska, E. The Role of Sigma-1 Receptor, an Intracellular Chaperone in Neurodegenerative Diseases. Curr. Neuropharmacol. 2018, 16, 97–116. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.; Lucke-Wold, B.P.; Mookerjee, S.A.; Cavendish, J.Z.; Robson, M.J.; Scandinaro, A.L.; Matsumoto, R.R. Role of sigma-1 receptors in neurodegenerative diseases. J. Pharmacol. Sci. 2015, 127, 17–29. [Google Scholar] [CrossRef] [PubMed]
- Hutcheson, J.D.; Setola, V.; Roth, B.L.; Merryman, W.D. Serotonin receptors and heart valve disease—It was meant 2B. Pharmacol. Ther. 2011, 132, 146–157. [Google Scholar] [CrossRef] [PubMed]
- Sourbron, J.; Schneider, H.; Kecskés, A.; Liu, Y.; Buening, E.M.; Lagae, L.; Smolders, I.; de Witte, P. Serotonergic Modulation as Effective Treatment for Dravet Syndrome in a Zebrafish Mutant Model. ACS Chem. Neurosci. 2016, 7, 588–598. [Google Scholar] [CrossRef]
- Volkers, L.; Kahlig, K.M.; Verbeek, N.E.; Das, J.H.; van Kempen, M.J.; Stroink, H.; Augustijn, P.; van Nieuwenhuizen, O.; Lindhout, D.; George, A.L., Jr.; et al. Nav 1.1 dysfunction in genetic epilepsy with febrile seizures-plus or Dravet syndrome. Eur. J. Neurosci. 2011, 34, 1268–1275. [Google Scholar] [CrossRef]
- Tang, B.; Dutt, K.; Papale, L.; Rusconi, R.; Shankar, A.; Hunter, J.; Tufik, S.; Yu, F.H.; Catterall, W.A.; Mantegazza, M.; et al. A BAC transgenic mouse model reveals neuron subtype-specific effects of a Generalized Epilepsy with Febrile Seizures Plus (GEFS+) mutation. Neurobiol. Dis. 2009, 35, 91–102. [Google Scholar] [CrossRef]
- Catterall, W.A.; Dib-Hajj, S.; Meisler, M.H.; Pietrobon, D. Inherited neuronal ion channelopathies: New windows on complex neurological diseases. J. Neurosci. 2008, 28, 11768–11777. [Google Scholar] [CrossRef]
- Zhang, T.; Chen, M.; Zhu, A.; Zhang, X.; Fang, T. Novel mutation of SCN9A gene causing generalized epilepsy with febrile seizures plus in a Chinese family. Neurol. Sci. 2020, 41, 1913–1917. [Google Scholar] [CrossRef] [PubMed]
- Martin, M.S.; Dutt, K.; Papale, L.A.; Dubé, C.M.; Dutton, S.B.; de Haan, G.; Shankar, A.; Tufik, S.; Meisler, M.H.; Baram, T.Z.; et al. Altered function of the SCN1A voltage-gated sodium channel leads to gamma-aminobutyric acid-ergic (GABAergic) interneuron abnormalities. J. Biol. Chem. 2010, 285, 9823–9834. [Google Scholar] [CrossRef]
- Hunt, R.F.; Hortopan, G.A.; Gillespie, A.; Baraban, S.C. A novel zebrafish model of hyperthermia-induced seizures reveals a role for TRPV4 channels and NMDA-type glutamate receptors. Exp. Neurol. 2012, 237, 199–206. [Google Scholar] [CrossRef]
- Ling, Y.; Wang, Y.; Jiang, X.; Yuan, C. Mechanism of the promotion of GEFS+ by the STAT3-mediated expression of interleukin-6. Transl. Pediatr. 2022, 11, 1491–1501. [Google Scholar] [CrossRef]
- Brenet, A.; Somkhit, J.; Csaba, Z.; Ciura, S.; Kabashi, E.; Yanicostas, C.; Soussi-Yanicostas, N. Microglia Mitigate Neuronal Activation in a Zebrafish Model of Dravet Syndrome. Cells 2024, 13, 684. [Google Scholar] [CrossRef]
- Gazina, E.V.; Leaw, B.T.W.; Richards, K.L.; Wimmer, V.C.; Kim, T.H.; Aumann, T.D.; Featherby, T.J.; Churilov, L.; Hammond, V.E.; Reid, C.A.; et al. ‘Neonatal’ Nav1.2 reduces neuronal excitability and affects seizure susceptibility and behaviour. Hum. Mol. Genet. 2014, 24, 1457–1468. [Google Scholar] [CrossRef] [PubMed]
- Zhou, D.; Lambert, S.; Malen, P.L.; Carpenter, S.; Boland, L.M.; Bennett, V. AnkyrinG is required for clustering of voltage-gated Na channels at axon initial segments and for normal action potential firing. J. Cell Biol. 1998, 143, 1295–1304. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.; Piguel, N.H.; Penzes, P. Roles and mechanisms of ankyrin-G in neuropsychiatric disorders. Exp. Mol. Med. 2022, 54, 867–877. [Google Scholar] [CrossRef]
- Ogiwara, I.; Miyamoto, H.; Tatsukawa, T.; Yamagata, T.; Nakayama, T.; Atapour, N.; Miura, E.; Mazaki, E.; Ernst, S.J.; Cao, D.; et al. Nav1.2 haplodeficiency in excitatory neurons causes absence-like seizures in mice. Commun. Biol. 2018, 1, 96. [Google Scholar] [CrossRef]
- Kaczmarek, L.K. Loss of Na(V)1.2-Dependent Backpropagating Action Potentials in Dendrites Contributes to Autism and Intellectual Disability. Neuron 2019, 103, 551–553. [Google Scholar] [CrossRef]
- Wolff, M.; Brunklaus, A.; Zuberi, S.M. Phenotypic spectrum and genetics of SCN2A-related disorders, treatment options, and outcomes in epilepsy and beyond. Epilepsia 2019, 60, S59–S67. [Google Scholar] [CrossRef]
- Menezes, L.F.S.; Sabiá Júnior, E.F.; Tibery, D.V.; Carneiro, L.D.A.; Schwartz, E.F. Epilepsy-Related Voltage-Gated Sodium Channelopathies: A Review. Front. Pharmacol. 2020, 11, 1276. [Google Scholar] [CrossRef]
- Van Wart, A.; Matthews, G. Expression of sodium channels Nav1.2 and Nav1.6 during postnatal development of the retina. Neurosci. Lett. 2006, 403, 315–317. [Google Scholar] [CrossRef] [PubMed]
- Caldwell, J.H.; Schaller, K.L.; Lasher, R.S.; Peles, E.; Levinson, S.R. Sodium channel Na(v)1.6 is localized at nodes of ranvier, dendrites, and synapses. Proc. Natl. Acad. Sci. USA 2000, 97, 5616–5620. [Google Scholar] [CrossRef] [PubMed]
- Black, J.A.; Renganathan, M.; Waxman, S.G. Sodium channel Nav1.6 is expressed along nonmyelinated axons and it contributes to conduction. Mol. Brain Res. 2002, 105, 19–28. [Google Scholar] [CrossRef]
- Dhindsa, R.S.; Lowenstein, D.H.; Goldstein, D.B. Chapter 37—Molecular Architecture and Neurobiology of the Epilepsies. In Genomics, Circuits, and Pathways in Clinical Neuropsychiatry; Lehner, T., Miller, B.L., State, M.W., Eds.; Academic Press: San Diego, MA, USA, 2016; pp. 601–617. [Google Scholar]
- Thompson, C.H.; Ben-Shalom, R.; Bender, K.J.; George, A.L., Jr. Alternative splicing potentiates dysfunction of early-onset epileptic encephalopathy SCN2A variants. J. Gen. Physiol. 2020, 152, e201912442. [Google Scholar] [CrossRef]
- Fletcher, E.V.; Kullmann, D.M.; Schorge, S. Alternative Splicing Modulates Inactivation of Type 1 Voltage-gated Sodium Channels by Toggling an Amino Acid in the First S3-S4 Linker. J. Biol. Chem. 2011, 286, 36700–36708. [Google Scholar] [CrossRef] [PubMed]
- Garrison, J.; Drews, V.; Jones, J.; Dugas, J.; Barres, B.; Meisler, M. Rbfox proteins regulate alternative splicing of neuronal sodium channel SCN8A. Mol. Cell. Neurosci. 2011, 49, 120–126. [Google Scholar] [CrossRef]
- Veldman, M.B.; Lin, S. Zebrafish as a developmental model organism for pediatric research. Pediatr. Res. 2008, 64, 470–476. [Google Scholar] [CrossRef]
- Liao, S.; Liu, T.; Yang, R.; Tan, W.; Gu, J.; Deng, M. Structure and function of sodium channel Nav1. 3 in neurological disorders. Cell. Mol. Neurobiol. 2023, 43, 575–584. [Google Scholar] [CrossRef]
- Smith, R.S.; Kenny, C.J.; Ganesh, V.; Jang, A.; Borges-Monroy, R.; Partlow, J.N.; Hill, R.S.; Shin, T.; Chen, A.Y.; Doan, R.N.; et al. Sodium Channel SCN3A (Na(V)1.3) Regulation of Human Cerebral Cortical Folding and Oral Motor Development. Neuron 2018, 99, 905–913.e907. [Google Scholar] [CrossRef]
- Estacion, M.; Gasser, A.; Dib-Hajj, S.D.; Waxman, S.G. A sodium channel mutation linked to epilepsy increases ramp and persistent current of Nav1.3 and induces hyperexcitability in hippocampal neurons. Exp. Neurol. 2010, 224, 362–368. [Google Scholar] [CrossRef]
- Vanoye, C.G.; Gurnett, C.A.; Holland, K.D.; George, A.L.; Kearney, J.A. Novel SCN3A variants associated with focal epilepsy in children. Neurobiol. Dis. 2014, 62, 313–322. [Google Scholar] [CrossRef] [PubMed]
- Holland, K.D.; Kearney, J.A.; Glauser, T.A.; Buck, G.; Keddache, M.; Blankston, J.R.; Glaaser, I.W.; Kass, R.S.; Meisler, M.H. Mutation of sodium channel SCN3A in a patient with cryptogenic pediatric partial epilepsy. Neurosci. Lett. 2008, 433, 65–70. [Google Scholar] [CrossRef]
- Zaman, T.; Helbig, I.; Božović, I.B.; DeBrosse, S.D.; Bergqvist, A.C.; Wallis, K.; Medne, L.; Maver, A.; Peterlin, B.; Helbig, K.L. Mutations in SCN3A cause early infantile epileptic encephalopathy. Ann. Neurol. 2018, 83, 703–717. [Google Scholar] [CrossRef] [PubMed]
- Lamar, T.; Vanoye, C.G.; Calhoun, J.; Wong, J.C.; Dutton, S.B.B.; Jorge, B.S.; Velinov, M.; Escayg, A.; Kearney, J.A. SCN3A deficiency associated with increased seizure susceptibility. Neurobiol. Dis. 2017, 102, 38–48. [Google Scholar] [CrossRef] [PubMed]
- Cummins, T.R.; Aglieco, F.; Renganathan, M.; Herzog, R.I.; Dib-Hajj, S.D.; Waxman, S.G. Nav1. 3 sodium channels: Rapid repriming and slow closed-state inactivation display quantitative differences after expression in a mammalian cell line and in spinal sensory neurons. J. Neurosci. 2001, 21, 5952–5961. [Google Scholar] [CrossRef] [PubMed]
- Talwar, D.; Hammer, M.F. SCN8A Epilepsy, Developmental Encephalopathy, and Related Disorders. Pediatr. Neurol. 2021, 122, 76–83. [Google Scholar] [CrossRef]
- Meisler, M.H.; Helman, G.; Hammer, M.F.; Fureman, B.E.; Gaillard, W.D.; Goldin, A.L.; Hirose, S.; Ishii, A.; Kroner, B.L.; Lossin, C.; et al. SCN8A encephalopathy: Research progress and prospects. Epilepsia 2016, 57, 1027–1035. [Google Scholar] [CrossRef]
- Meisler, M.H. SCN8A encephalopathy: Mechanisms and models. Epilepsia 2019, 60 (Suppl. S3), S86–S91. [Google Scholar] [CrossRef]
- King, D.R.; Demirtas, M.; Tarasov, M.; Struckman, H.L.; Meng, X.; Nassal, D.; Moise, N.; Miller, A.; Min, D.; Soltisz, A.M.; et al. Cardiac-Specific Deletion of Scn8a Mitigates Dravet Syndrome-Associated Sudden Death in Adults. JACC Clin. Electrophysiol. 2024, 10, 829–842. [Google Scholar] [CrossRef] [PubMed]


| Characteristics | Zebrafish Models | Rodent Models |
|---|---|---|
| Genetic homology | High sequence homology with human disease genes (84%) [4,56] | Overall high genetic and physiological homology with humans (~90%) [72] |
| Sodium channel isoforms | Different sodium channel isoforms, e.g., zebrafish express scn1lab, an ortholog to SCN1A in humans [73] | Structure and function of VGSCs conserved in mammals |
| Nervous system complexity | Primitive nervous system; limited ability to simulate complex neuronal diseases | Homologous brain networks with primates [62] |
| Electrophysiology | Patch-clamp is challenging due to small cell size, but protocols like in vivo whole-cell and multichannel recordings are established [74,75] | In vivo patch-clamp methods are established but often require anesthesia and invasive electrode insertion, raising ethical concerns [76] |
| Genetic manipulation | External fertilization allows easy genetic manipulation and access to one-cell zygotes for injections [64] | In vitro, ex vitro, and in vivo experimental models, but more resource-intensive |
| Behavioral studies | While many behavior analyses have been established for zebrafish, translating results to human models can be challenging | Rodent models can capture behavioral components that can be translated into human neurological disorders and psychiatry [77] |
| Development | High proliferation, oviparity, and translucence of developing embryos make observation of early development stages easier | In utero development, prolonged growth, and maternal dependence complicate early embryogenesis studies and toxin exposure assessments |
| Ethical considerations | Fewer ethical concerns of embryos and larvae up to 5 dpf [70,71] | Ethical concerns regarding invasive procedures [63] |
| Cost of maintenance | Lower cost of maintenance | Higher cost of maintenance [63] |
| Human Gene | Encoded Protein | Zebrafish Orthologs | Expression Pattern | Identity to Homolog | References |
|---|---|---|---|---|---|
| SCN1A | Nav 1.1 | scn1laa/scn1lab | Central nervous system (throughout the brain, eye, ventral regions of spinal cord) | 77% | [87] |
| SCN2A | Nav 1.2 | ||||
| SCN3A | Nav 1.3 | ||||
| SCN9A | Nav 1.7 | ||||
| SCN4A | Nav 1.4 | scn4aa/scn4ab | Skeletal muscle/mesodermal tissues | NA | [87] |
| SCN5A | Nav 1.5 | scn5Laa/scn5Lab | Cardiac tissues | 60–65% | [87,89] |
| SCN10A | Nav 1.8 | ||||
| SCN11A | Nav 1.9 | ||||
| SCN8A | Nav 1.6 | scn8aa/scn8ab | Rohon–Beard neurons and trigeminal ganglion (16–17 hpf) | 83% | [87,90,91] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gyamfi, A.; Manikandan, P.; Cisneros, W.A.; Cummins, T.R.; Marrs, J.A. Voltage-Gated Sodium Channel Dysfunction in Epilepsy: Zebrafish Models for Therapeutics. Biomedicines 2025, 13, 2078. https://doi.org/10.3390/biomedicines13092078
Gyamfi A, Manikandan P, Cisneros WA, Cummins TR, Marrs JA. Voltage-Gated Sodium Channel Dysfunction in Epilepsy: Zebrafish Models for Therapeutics. Biomedicines. 2025; 13(9):2078. https://doi.org/10.3390/biomedicines13092078
Chicago/Turabian StyleGyamfi, Angela, Priyadharshini Manikandan, William A. Cisneros, Theodore R. Cummins, and James A. Marrs. 2025. "Voltage-Gated Sodium Channel Dysfunction in Epilepsy: Zebrafish Models for Therapeutics" Biomedicines 13, no. 9: 2078. https://doi.org/10.3390/biomedicines13092078
APA StyleGyamfi, A., Manikandan, P., Cisneros, W. A., Cummins, T. R., & Marrs, J. A. (2025). Voltage-Gated Sodium Channel Dysfunction in Epilepsy: Zebrafish Models for Therapeutics. Biomedicines, 13(9), 2078. https://doi.org/10.3390/biomedicines13092078








