RNA-Binding Protein Motifs Predict microRNA Secretion and Cellular Retention in Hypothalamic and Other Cell Types
Abstract
1. Introduction
2. Methods
2.1. Cell Culture and miRNA Microarray
2.2. Classification of Preferentially Secreted and Cellular miRNAs
2.3. Pathway Enrichment Analysis for miRNAs
2.4. Motif Discovery
2.5. Machine Learning Modeling
3. Results
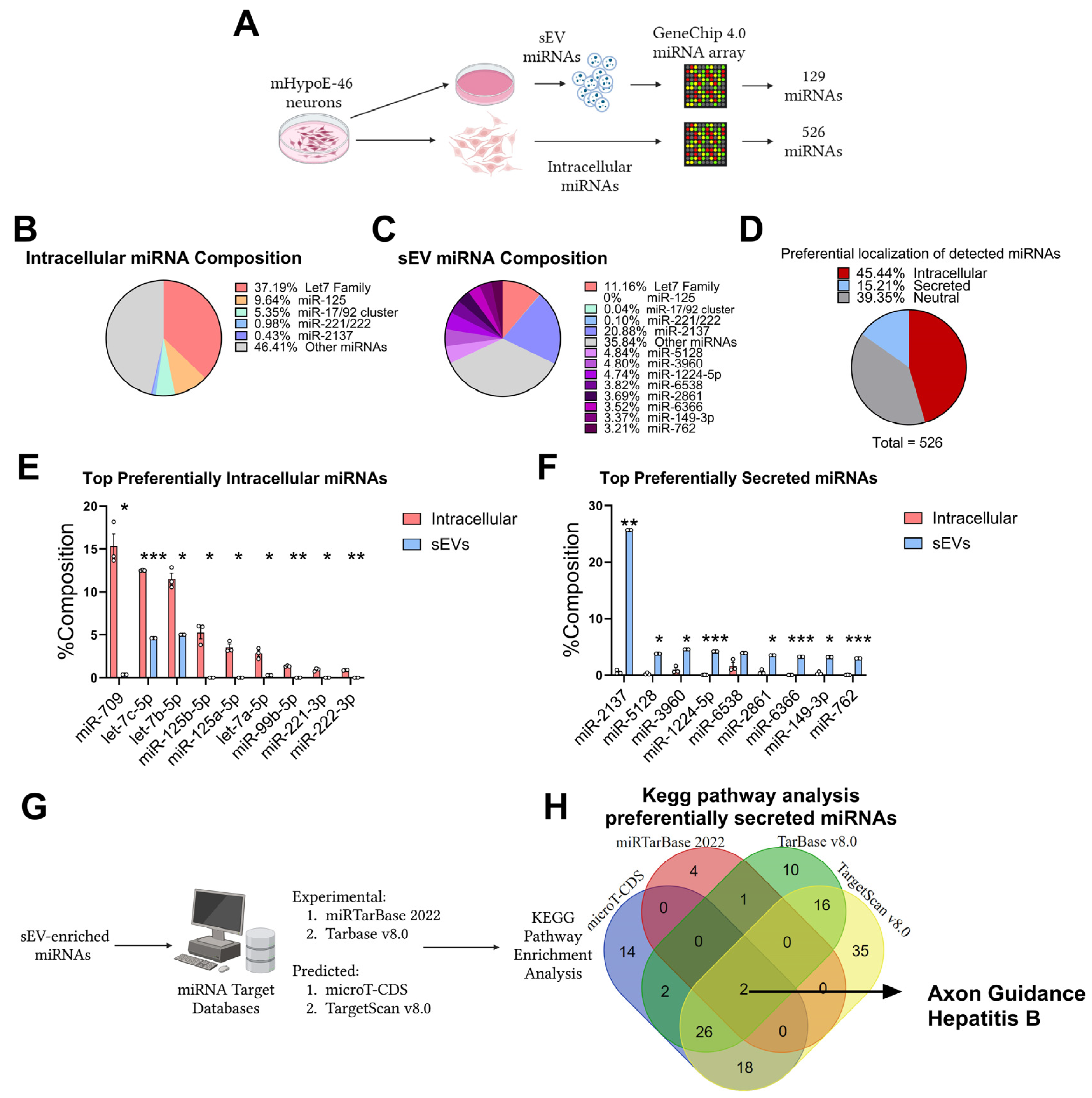
3.1. Preferential Retention and Secretion of Neuronal miRNAs
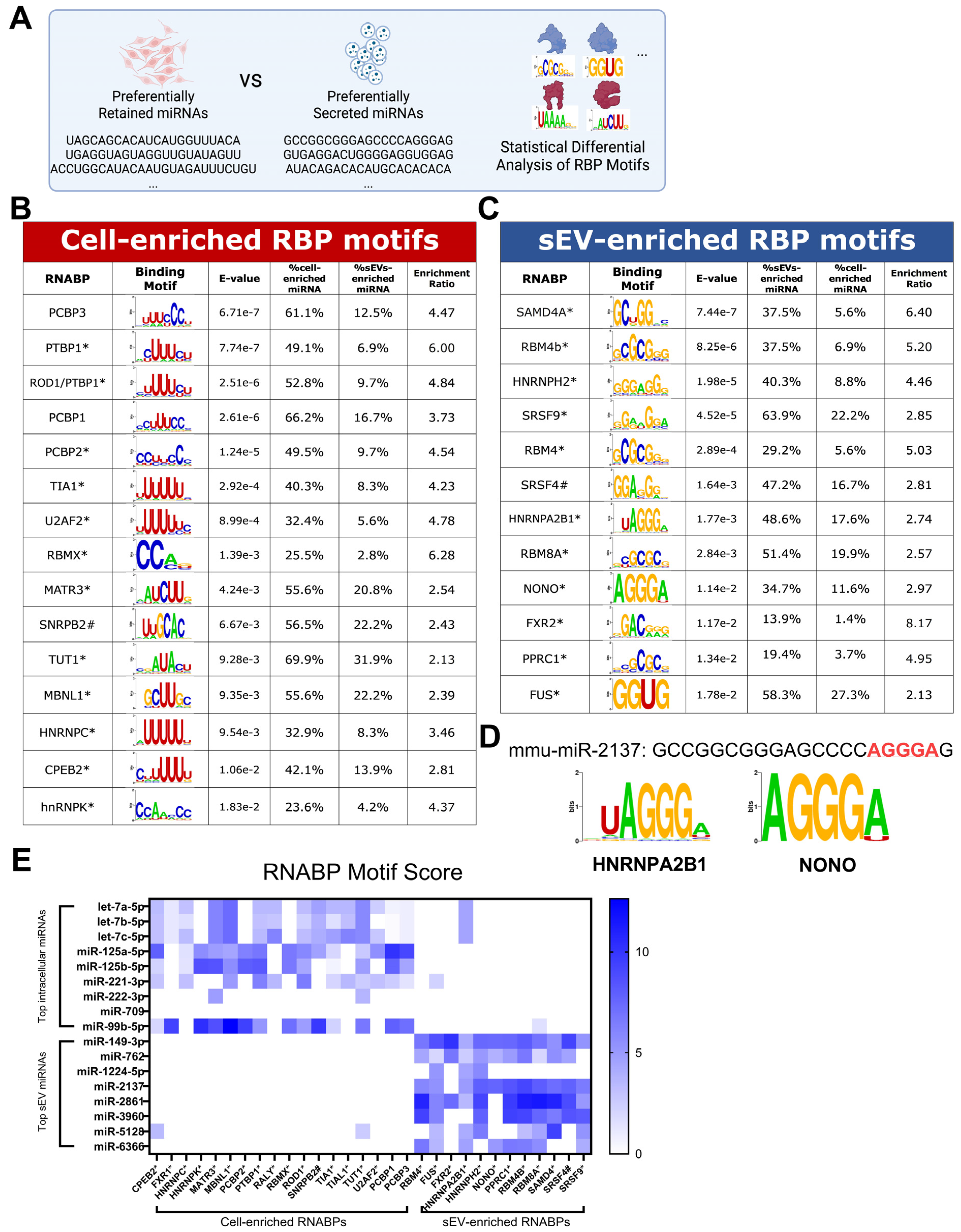
3.2. RBP Motifs Statistically Correlate with the Differential Sorting of Neuronal miRNAs
3.3. The Presence and Strength of RBP Motifs Can Be Used to Accurately Predict miRNA Localization in mHypoE-46 Neurons
3.4. Enrichment of RBP Motifs in Preferentially Sorted miRNAs across Multiple Cell Types
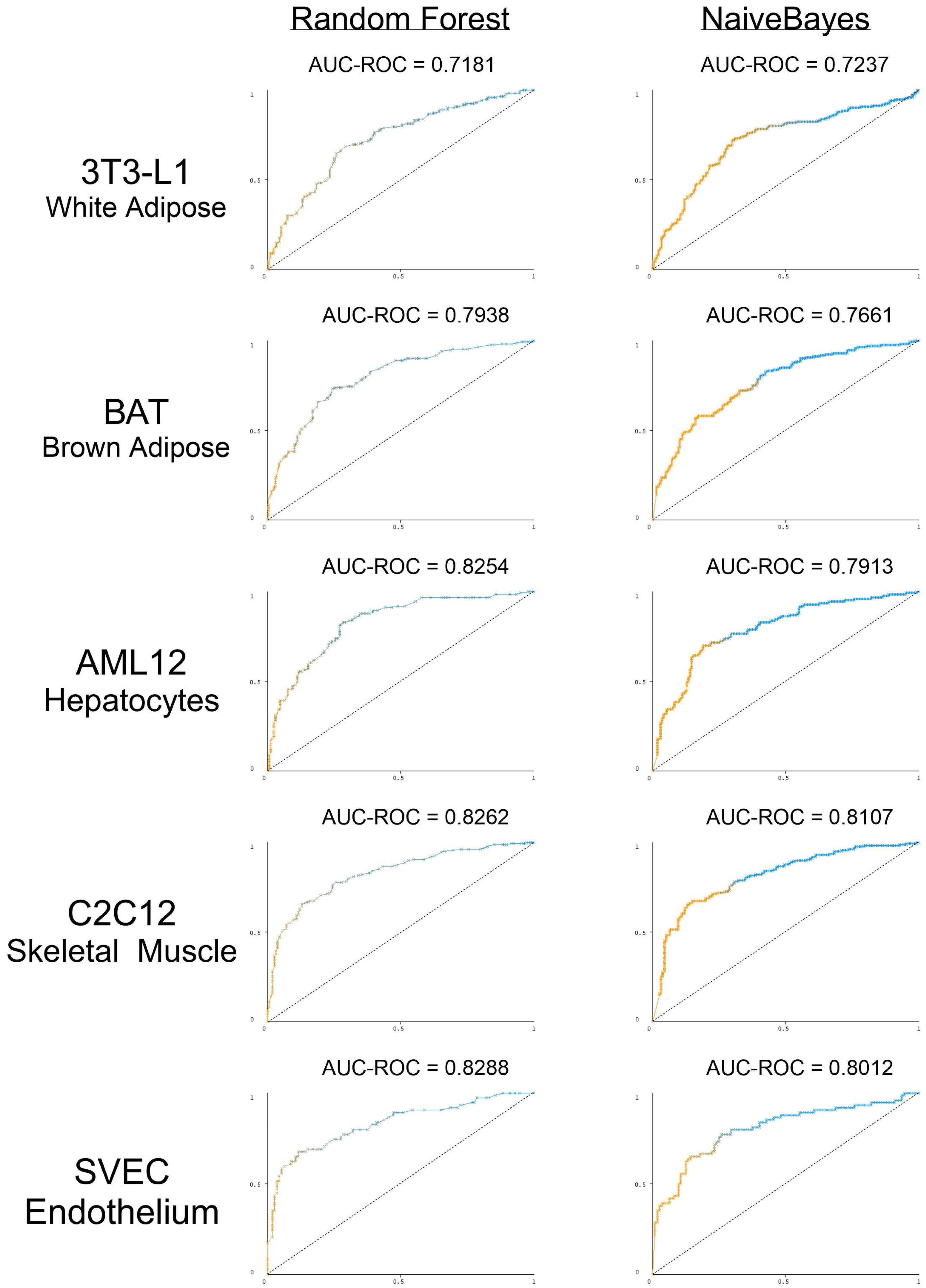
3.5. The Presence and Strength of RBP Motifs Can Be Used to Accurately Predict miRNA Localization in Multiple Peripheral Models
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Valadi, H.; Ekstrom, K.; Bossios, A.; Sjostrand, M.; Lee, J.J.; Lotvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zhao, Y.; Yin, Y.; Jia, X.; Mao, L. Mechanism of cargo sorting into small extracellular vesicles. Bioengineered 2021, 12, 8186–8201. [Google Scholar] [CrossRef] [PubMed]
- Crooke, S.T.; Vickers, T.A.; Liang, X.H. Phosphorothioate modified oligonucleotide-protein interactions. Nucleic Acids Res. 2020, 48, 5235–5253. [Google Scholar] [CrossRef] [PubMed]
- Mori, M.A.; Ludwig, R.G.; Garcia-Martin, R.; Brandao, B.B.; Kahn, C.R. Extracellular miRNAs: From Biomarkers to Mediators of Physiology and Disease. Cell Metab. 2019, 30, 656–673. [Google Scholar] [CrossRef] [PubMed]
- Isaac, R.; Reis, F.C.G.; Ying, W.; Olefsky, J.M. Exosomes as mediators of intercellular crosstalk in metabolism. Cell Metab. 2021, 33, 1744–1762. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Di, K.; Fan, B.; Wu, J.; Gu, X.; Sun, Y.; Khan, A.; Li, P.; Li, Z. MicroRNAs in extracellular vesicles: Sorting mechanisms, diagnostic value, isolation, and detection technology. Front. Bioeng. Biotechnol. 2022, 10, 948959. [Google Scholar] [CrossRef] [PubMed]
- Tosar, J.P.; Gambaro, F.; Sanguinetti, J.; Bonilla, B.; Witwer, K.W.; Cayota, A. Assessment of small RNA sorting into different extracellular fractions revealed by high-throughput sequencing of breast cell lines. Nucleic Acids Res. 2015, 43, 5601–5616. [Google Scholar] [CrossRef] [PubMed]
- Villarroya-Beltri, C.; Gutierrez-Vazquez, C.; Sanchez-Cabo, F.; Perez-Hernandez, D.; Vazquez, J.; Martin-Cofreces, N.; Martinez-Herrera, D.J.; Pascual-Montano, A.; Mittelbrunn, M.; Sanchez-Madrid, F. Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nat. Commun. 2013, 4, 2980. [Google Scholar] [CrossRef] [PubMed]
- Teng, Y.; Ren, Y.; Hu, X.; Mu, J.; Samykutty, A.; Zhuang, X.; Deng, Z.; Kumar, A.; Zhang, L.; Merchant, M.L.; et al. MVP-mediated exosomal sorting of miR-193a promotes colon cancer progression. Nat. Commun. 2017, 8, 14448. [Google Scholar] [CrossRef]
- Shurtleff, M.J.; Temoche-Diaz, M.M.; Karfilis, K.V.; Ri, S.; Schekman, R. Y-box protein 1 is required to sort microRNAs into exosomes in cells and in a cell-free reaction. eLife 2016, 5, 19276. [Google Scholar] [CrossRef]
- Santangelo, L.; Giurato, G.; Cicchini, C.; Montaldo, C.; Mancone, C.; Tarallo, R.; Battistelli, C.; Alonzi, T.; Weisz, A.; Tripodi, M. The RNA-Binding Protein SYNCRIP Is a Component of the Hepatocyte Exosomal Machinery Controlling MicroRNA Sorting. Cell Rep. 2016, 17, 799–808. [Google Scholar] [CrossRef] [PubMed]
- Luo, X.; Jean-Toussaint, R.; Sacan, A.; Ajit, S.K. Differential RNA packaging into small extracellular vesicles by neurons and astrocytes. Cell Commun. Signal. 2021, 19, 75. [Google Scholar] [CrossRef] [PubMed]
- Krol, J.; Busskamp, V.; Markiewicz, I.; Stadler, M.B.; Ribi, S.; Richter, J.; Duebel, J.; Bicker, S.; Fehling, H.J.; Schubeler, D.; et al. Characterizing light-regulated retinal microRNAs reveals rapid turnover as a common property of neuronal microRNAs. Cell 2010, 141, 618–631. [Google Scholar] [CrossRef] [PubMed]
- La Rocca, G.; Olejniczak, S.H.; Gonzalez, A.J.; Briskin, D.; Vidigal, J.A.; Spraggon, L.; DeMatteo, R.G.; Radler, M.R.; Lindsten, T.; Ventura, A.; et al. In vivo, Argonaute-bound microRNAs exist predominantly in a reservoir of low molecular weight complexes not associated with mRNA. Proc. Natl. Acad. Sci. USA 2015, 112, 767–772. [Google Scholar] [CrossRef] [PubMed]
- Steuernagel, L.; Lam, B.Y.H.; Klemm, P.; Dowsett, G.K.C.; Bauder, C.A.; Tadross, J.A.; Hitschfeld, T.S.; Del Rio Martin, A.; Chen, W.; de Solis, A.J.; et al. HypoMap-a unified single-cell gene expression atlas of the murine hypothalamus. Nat. Metab. 2022, 4, 1402–1419. [Google Scholar] [CrossRef] [PubMed]
- Mayer, C.M.; Belsham, D.D. Insulin directly regulates NPY and AgRP gene expression via the MAPK MEK/ERK signal transduction pathway in mHypoE-46 hypothalamic neurons. Mol. Cell. Endocrinol. 2009, 307, 99–108. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Loganathan, N.; Tran, A.; Belsham, D.D. Npy transcription is regulated by noncanonical STAT3 signaling in hypothalamic neurons: Implication with lipotoxicity and obesity. Mol. Cell. Endocrinol. 2024, 586, 112179. [Google Scholar] [CrossRef]
- McIlwraith, E.K.; Belsham, D.D. Palmitate alters miRNA content of small extracellular vesicles secreted from NPY/AgRP-expressing hypothalamic neurons. Brain Res. 2023, 1810, 148367. [Google Scholar] [CrossRef]
- Garcia-Martin, R.; Wang, G.; Brandao, B.B.; Zanotto, T.M.; Shah, S.; Kumar Patel, S.; Schilling, B.; Kahn, C.R. MicroRNA sequence codes for small extracellular vesicle release and cellular retention. Nature 2021, 601, 446–451. [Google Scholar] [CrossRef]
- McIlwraith, E.K.; He, W.; Belsham, D.D. Promise and Perils of MicroRNA Discovery Research: Working Toward Quality Over Quantity. Endocrinology 2023, 164, bqad111. [Google Scholar] [CrossRef]
- Tastsoglou, S.; Skoufos, G.; Miliotis, M.; Karagkouni, D.; Koutsoukos, I.; Karavangeli, A.; Kardaras, F.S.; Hatzigeorgiou, A.G. DIANA-miRPath v4.0: Expanding target-based miRNA functional analysis in cell-type and tissue contexts. Nucleic Acids Res. 2023, 51, W154–W159. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.Y.; Lin, Y.C.; Cui, S.; Huang, Y.; Tang, Y.; Xu, J.; Bao, J.; Li, Y.; Wen, J.; Zuo, H.; et al. miRTarBase update 2022: An informative resource for experimentally validated miRNA-target interactions. Nucleic Acids Res. 2022, 50, D222–D230. [Google Scholar] [CrossRef]
- Karagkouni, D.; Paraskevopoulou, M.D.; Chatzopoulos, S.; Vlachos, I.S.; Tastsoglou, S.; Kanellos, I.; Papadimitriou, D.; Kavakiotis, I.; Maniou, S.; Skoufos, G.; et al. DIANA-TarBase v8: A decade-long collection of experimentally supported miRNA-gene interactions. Nucleic Acids Res. 2018, 46, D239–D245. [Google Scholar] [CrossRef] [PubMed]
- Tastsoglou, S.; Alexiou, A.; Karagkouni, D.; Skoufos, G.; Zacharopoulou, E.; Hatzigeorgiou, A.G. DIANA-microT 2023: Including predicted targets of virally encoded miRNAs. Nucleic Acids Res. 2023, 51, W148–W153. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, V.; Bell, G.W.; Nam, J.W.; Bartel, D.P. Predicting effective microRNA target sites in mammalian mRNAs. eLife 2015, 4, e05005. [Google Scholar] [CrossRef]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME Suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef]
- Ray, D.; Kazan, H.; Cook, K.B.; Weirauch, M.T.; Najafabadi, H.S.; Li, X.; Gueroussov, S.; Albu, M.; Zheng, H.; Yang, A.; et al. A compendium of RNA-binding motifs for decoding gene regulation. Nature 2013, 499, 172–177. [Google Scholar] [CrossRef] [PubMed]
- Frank, E.; Hall, M.; Trigg, L.; Holmes, G.; Witten, I.H. Data mining in bioinformatics using Weka. Bioinformatics 2004, 20, 2479–2481. [Google Scholar] [CrossRef] [PubMed]
- Hock, J.; Weinmann, L.; Ender, C.; Rudel, S.; Kremmer, E.; Raabe, M.; Urlaub, H.; Meister, G. Proteomic and functional analysis of Argonaute-containing mRNA-protein complexes in human cells. EMBO Rep. 2007, 8, 1052–1060. [Google Scholar] [CrossRef]
- Lin, J.C.; Tarn, W.Y. RNA-binding motif protein 4 translocates to cytoplasmic granules and suppresses translation via argonaute2 during muscle cell differentiation. J. Biol. Chem. 2009, 284, 34658–34665. [Google Scholar] [CrossRef]
- Pinder, B.D.; Smibert, C.A. microRNA-independent recruitment of Argonaute 1 to nanos mRNA through the Smaug RNA-binding protein. EMBO Rep. 2013, 14, 80–86. [Google Scholar] [CrossRef] [PubMed]
- Leidal, A.M.; Huang, H.H.; Marsh, T.; Solvik, T.; Zhang, D.; Ye, J.; Kai, F.; Goldsmith, J.; Liu, J.Y.; Huang, Y.H.; et al. The LC3-conjugation machinery specifies the loading of RNA-binding proteins into extracellular vesicles. Nat. Cell. Biol. 2020, 22, 187–199. [Google Scholar] [CrossRef] [PubMed]
- Thomou, T.; Mori, M.A.; Dreyfuss, J.M.; Konishi, M.; Sakaguchi, M.; Wolfrum, C.; Rao, T.N.; Winnay, J.N.; Garcia-Martin, R.; Grinspoon, S.K.; et al. Adipose-derived circulating miRNAs regulate gene expression in other tissues. Nature 2017, 542, 450–455. [Google Scholar] [CrossRef] [PubMed]
- Wei, Z.; Batagov, A.O.; Schinelli, S.; Wang, J.; Wang, Y.; El Fatimy, R.; Rabinovsky, R.; Balaj, L.; Chen, C.C.; Hochberg, F.; et al. Coding and noncoding landscape of extracellular RNA released by human glioma stem cells. Nat. Commun. 2017, 8, 1145. [Google Scholar] [CrossRef]
- Friedrich, M.; Aigner, A. Therapeutic siRNA: State-of-the-Art and Future Perspectives. BioDrugs 2022, 36, 549–571. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, W.; Belsham, D.D. RNA-Binding Protein Motifs Predict microRNA Secretion and Cellular Retention in Hypothalamic and Other Cell Types. Biomedicines 2024, 12, 857. https://doi.org/10.3390/biomedicines12040857
He W, Belsham DD. RNA-Binding Protein Motifs Predict microRNA Secretion and Cellular Retention in Hypothalamic and Other Cell Types. Biomedicines. 2024; 12(4):857. https://doi.org/10.3390/biomedicines12040857
Chicago/Turabian StyleHe, Wenyuan, and Denise D. Belsham. 2024. "RNA-Binding Protein Motifs Predict microRNA Secretion and Cellular Retention in Hypothalamic and Other Cell Types" Biomedicines 12, no. 4: 857. https://doi.org/10.3390/biomedicines12040857
APA StyleHe, W., & Belsham, D. D. (2024). RNA-Binding Protein Motifs Predict microRNA Secretion and Cellular Retention in Hypothalamic and Other Cell Types. Biomedicines, 12(4), 857. https://doi.org/10.3390/biomedicines12040857





