A Deep Learning Approach for Prognostic Evaluation of Lung Adenocarcinoma Based on Cuproptosis-Related Genes
Abstract
1. Introduction
2. Materials and Methods
2.1. Datasets
2.2. Differentially Expressed Gene Screening and DNN Model Construction
2.3. Based on the Determination of Model Grouping and Model Rationality Analysis
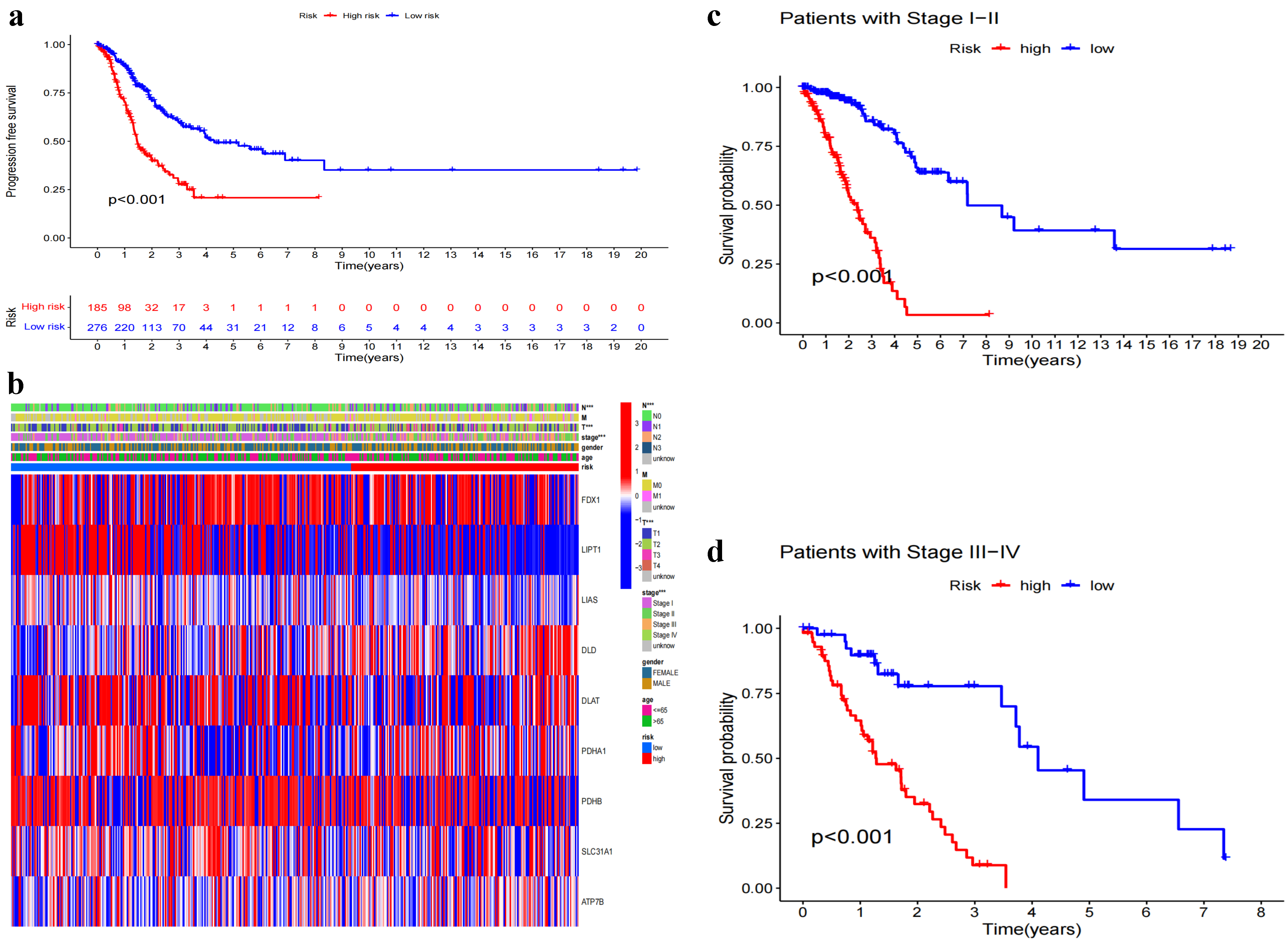
2.4. Progression-Free Survival (PFS) Analysis and Clinical Characteristics Exploration
2.5. Functional Analysis of Differences between Model Subgroups
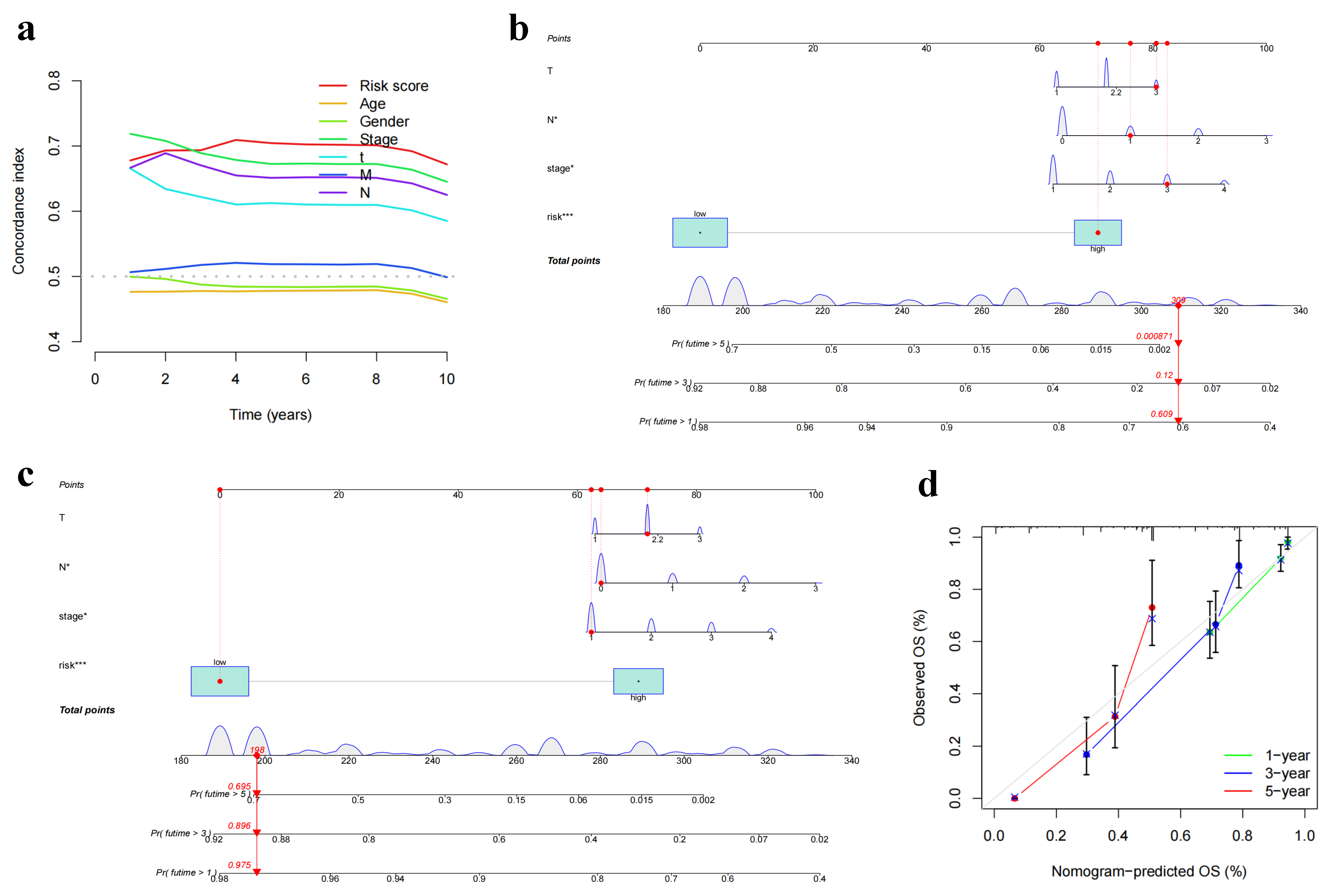
2.6. Development of Nomogram Model for Individualized Clinical Decision Making
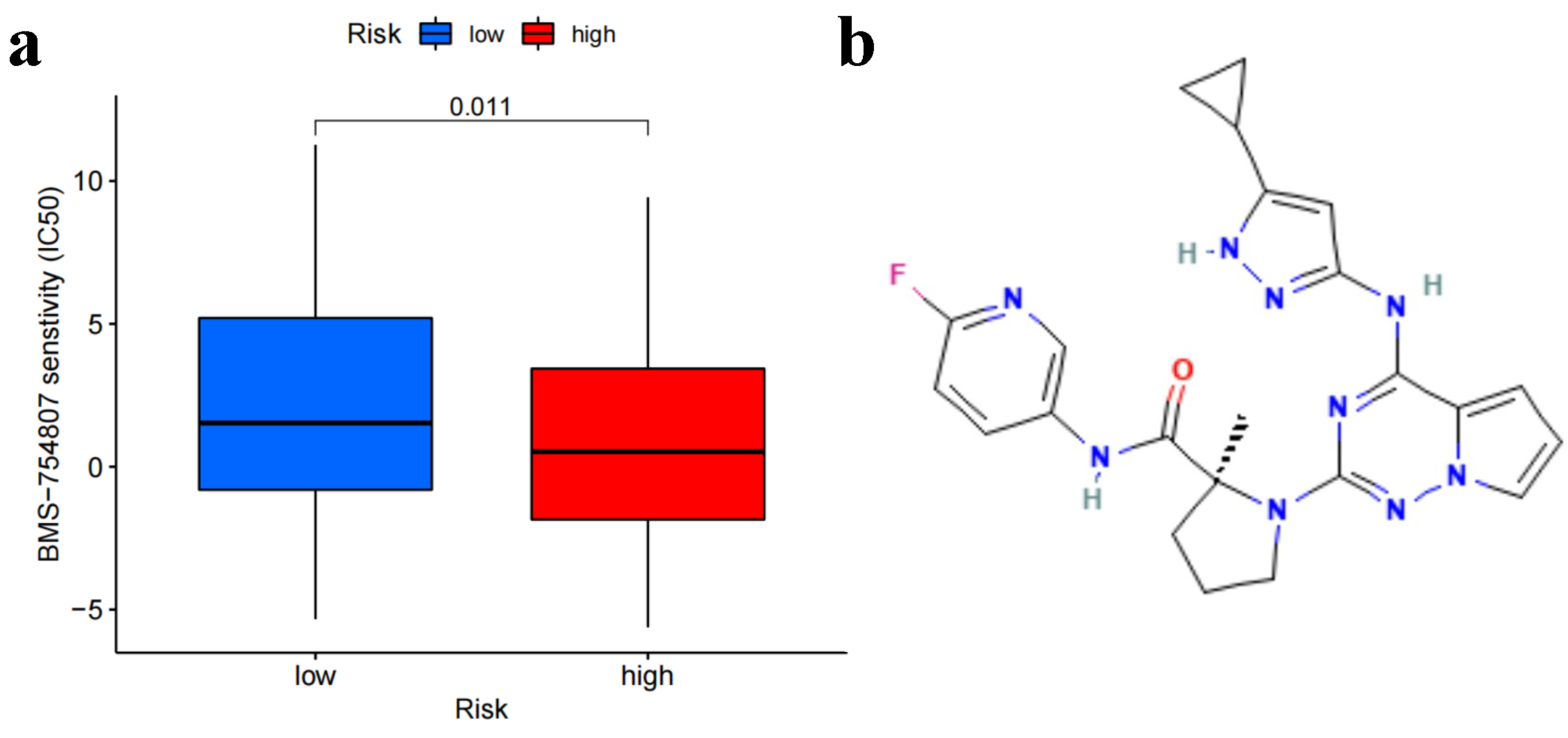
2.7. Screening of Anti-Tumor Sensitive Drugs
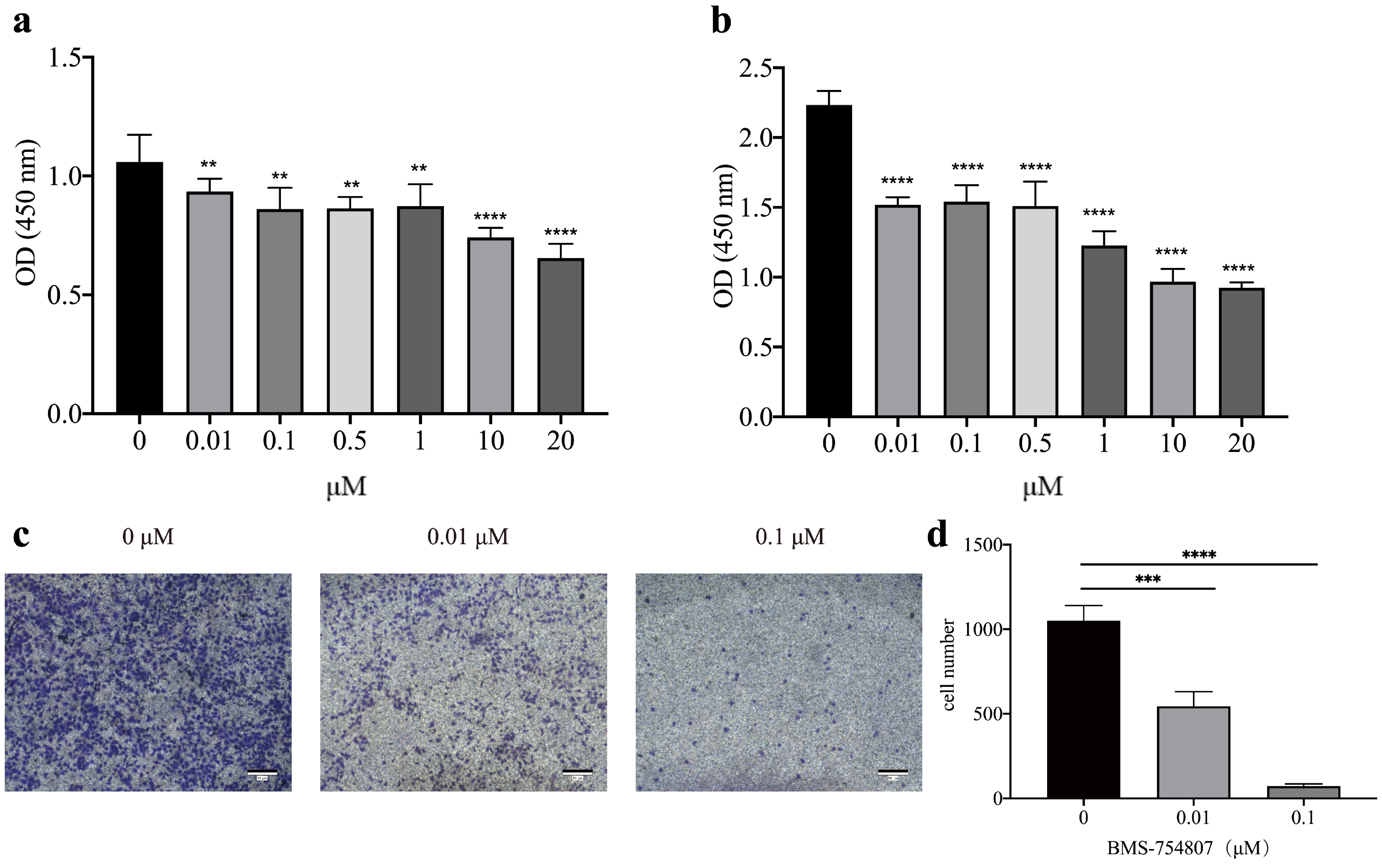
2.8. Effect of Sensitive Drugs on the Activity of LUAD A549 Cell Line
2.9. Statistical Analysis
3. Results
3.1. Identifying and Modeling Cuproptosis-Related Differentially Expressed Genes
3.2. External Validation Using GEO Dataset
3.3. Association between Model-Based Risk Stratification and Clinical Characteristics
3.4. Functional Analysis of Model Grouping
3.5. Construction of a Nomogram Model for Personalized Clinical Decision-Making
3.6. Anti-Tumor Susceptibility Drug Screening and Sensitivity Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chansky, K.; Sculier, J.P.; Crowley, J.J.; Giroux, D.; Van Meerbeeck, J.; Goldstraw, P. The International Association for the Study of Lung Cancer Staging Project: Prognostic factors and pathologic TNM stage in surgically managed non-small cell lung cancer. J. Thorac. Oncol. 2009, 4, 792–801. [Google Scholar] [CrossRef]
- Li, J.; Ching, T.; Huang, S.; Garmire, L.X. Using epigenomics data to predict gene expression in lung cancer. BMC Bioinform. 2015, 16, S10. [Google Scholar] [CrossRef]
- Huang, C.H.; Chang, P.M.H.; Hsu, C.W.; Huang, C.Y.F.; Ng, K.L. Drug repositioning for non-small cell lung cancer by using machine learning algorithms and topological graph theory. BMC Bioinform. 2016, 17, S2. [Google Scholar] [CrossRef]
- Saygideger, Y.; Avci, A.; Bagir, E.; Saygıdeğer Demir, B.; Sezan, A.; Ekici, M.; Baydar, O.; Erkin, Ö.C. Slug and Vimentin downregulation at the metastatic site is associated with Skip-N2 metastasis of lung adenocarcinoma. Discov. Oncol. 2022, 13, 7. [Google Scholar] [CrossRef]
- Ettinger, D.S.; Wood, D.E.; Akerley, W.; Bazhenova, L.A.; Borghaei, H.; Camidge, D.R.; Cheney, R.T.; Chirieac, L.R.; D’Amico, T.A.; Demmy, T.L.; et al. Non–small cell lung cancer, version 6.2015. J. Natl. Compr. Cancer Netw. 2015, 13, 515–524. [Google Scholar] [CrossRef] [PubMed]
- Shukla, S.; Evans, J.R.; Malik, R.; Feng, F.Y.; Dhanasekaran, S.M.; Cao, X.; Chen, G.; Beer, D.G.; Jiang, H.; Chinnaiyan, A.M. Development of a RNA-Seq based prognostic signature in lung adenocarcinoma. JNCI J. Natl. Cancer Inst. 2017, 109, djw200. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Kou, Y.; Zhang, X.; Xiao, X.; Ou, Y.; Cao, L.; Guo, M.; Qi, C.; Wang, Z.; Liu, Y.; et al. Biochanin A inhibits lung adenocarcinoma progression by targeting ZEB1. Discov. Oncol. 2022, 13, 138. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Niu, Y.; Zhao, W.; Yan, C.; Qi, Y. Construction and validation of a prognostic model for lung adenocarcinoma based on endoplasmic reticulum stress-related genes. Sci. Rep. 2022, 12, 19857. [Google Scholar] [CrossRef]
- Lu, F.; Gao, J.; Hou, Y.; Cao, K.; Xia, Y.; Chen, Z.; Yu, H.; Chang, L.; Li, W. Construction of a Novel Prognostic Model in Lung Adenocarcinoma Based on 7-Methylguanosine-Related Gene Signatures. Front. Oncol. 2022, 12, 876360. [Google Scholar] [CrossRef]
- Ren, J.; Yang, Y.; Li, C.; Xie, L.; Hu, R.; Qin, X.; Zhang, M. A novel prognostic model of early-stage lung adenocarcinoma integrating methylation and immune biomarkers. Front. Genet. 2021, 11, 634634. [Google Scholar] [CrossRef]
- Liang, P.; Li, J.; Chen, J.; Lu, J.; Hao, Z.; Shi, J.; Chang, Q.; Zeng, Z. Immunoprognostic model of lung adenocarcinoma and screening of sensitive drugs. Sci. Rep. 2022, 12, 7162. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Zeng, T.; Zhou, C.; Chen, Y.; Yin, W. A prognostic signature model for unveiling tumor progression in lung adenocarcinoma. Front. Oncol. 2022, 12, 1019442. [Google Scholar] [CrossRef]
- He, J.; Li, W.; Li, Y.; Liu, G. Construction of a prognostic model for lung adenocarcinoma based on bioinformatics analysis of metabolic genes. Transl. Cancer Res. 2020, 9, 3518. [Google Scholar] [CrossRef]
- Yang, L.; Zhang, R.; Guo, G.; Wang, G.; Wen, Y.; Lin, Y.; Zhang, X.; Yu, X.; Huang, Z.; Zhao, D.; et al. Development and validation of a prediction model for lung adenocarcinoma based on RNA-binding protein. Ann. Transl. Med. 2021, 9, 474. [Google Scholar] [CrossRef]
- Kim, B.E.; Nevitt, T.; Thiele, D.J. Mechanisms for copper acquisition, distribution and regulation. Nat. Chem. Biol. 2008, 4, 176–185. [Google Scholar] [CrossRef] [PubMed]
- Lutsenko, S. Human copper homeostasis: A network of interconnected pathways. Curr. Opin. Chem. Biol. 2010, 14, 211–217. [Google Scholar] [CrossRef]
- Tsvetkov, P.; Coy, S.; Petrova, B.; Dreishpoon, M.; Verma, A.; Abdusamad, M.; Rossen, J.; Joesch-Cohen, L.; Humeidi, R.; Spangler, R.D.; et al. Copper induces cell death by targeting lipoylated TCA cycle proteins. Science 2022, 375, 1254–1261. [Google Scholar] [CrossRef]
- Ishida, S.; Andreux, P.; Poitry-Yamate, C.; Auwerx, J.; Hanahan, D. Bioavailable copper modulates oxidative phosphorylation and growth of tumors. Proc. Natl. Acad. Sci. USA 2013, 110, 19507–19512. [Google Scholar] [CrossRef] [PubMed]
- Director’s Challenge Consortium for the Molecular Classification of Lung Adenocarcinoma. Gene expression–based survival prediction in lung adenocarcinoma: A multi-site, blinded validation study. Nat. Med. 2008, 14, 822–827. [Google Scholar] [CrossRef] [PubMed]
- Rl, S.; KD, M.; Jemal, A. Cancer statistics, 2020. CA Cancer J. Clin. 2020, 70, 7–30. [Google Scholar]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef]
- Löfling, L.; Bahmanyar, S.; Kieler, H.; Lambe, M.; Wagenius, G. Temporal trends in lung cancer survival: A population-based study. Acta Oncol. 2022, 61, 625–631. [Google Scholar] [CrossRef]
- Yun, Y.; Wang, Y.; Yang, E.; Jing, X. Cuproptosis-Related Gene–SLC31A1, FDX1 and ATP7B–Polymorphisms are Associated with Risk of Lung Cancer. Pharmacogenom. Pers. Med. 2022, 15, 733–742. [Google Scholar] [CrossRef] [PubMed]
- Inoue, Y.; Matsumoto, H.; Yamada, S.; Kawai, K.; Suemizu, H.; Gika, M.; Takanami, I.; Nakamura, M.; Iwazaki, M. ATP7B expression is associated with in vitro sensitivity to cisplatin in non-small cell lung cancer. Oncol. Lett. 2010, 1, 279–282. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.Q.; Chen, J.; Yin, J.Y.; Liu, Z.Q.; Li, X.P. Gene expression and single nucleotide polymorphism of ATP7B are associated with platinum-based chemotherapy response in non-small cell lung cancer patients. J. Cancer 2018, 9, 3532. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, T.; Inoue, Y.; Kodama, H.; Yamazaki, H.; Kawai, K.; Suemizu, H.; Masuda, R.; Iwazaki, M.; Yamada, S.; Ueyama, Y.; et al. Expression of copper-transporting P-type adenosine triphosphatase (ATP7B) correlates with cisplatin resistance in human non-small cell lung cancer xenografts. Oncol. Rep. 2008, 20, 265–270. [Google Scholar]
- Sun, J.; Li, J.; Guo, Z.; Sun, L.; Juan, C.; Zhou, Y.; Gu, H.; Yu, Y.; Hu, Q.; Yu, Z.; et al. Overexpression of pyruvate dehydrogenase E1α subunit inhibits Warburg effect and induces cell apoptosis through mitochondria-mediated pathway in hepatocellular carcinoma. Oncol. Res. 2019, 27, 407. [Google Scholar] [CrossRef]
- Liu, L.; Cao, J.; Zhao, J.; Li, X.; Suo, Z.; Li, H. PDHA1 gene knockout in human esophageal squamous cancer cells resulted in greater Warburg effect and aggressive features in vitro and in vivo. OncoTargets Ther. 2019, 12, 9899. [Google Scholar] [CrossRef]
- Glushakova, L.G.; Lisankie, M.J.; Eruslanov, E.B.; Ojano-Dirain, C.; Zolotukhin, I.; Liu, C.; Srivastava, A.; Stacpoole, P.W. AAV3-mediated transfer and expression of the pyruvate dehydrogenase E1 alpha subunit gene causes metabolic remodeling and apoptosis of human liver cancer cells. Mol. Genet. Metab. 2009, 98, 289–299. [Google Scholar] [CrossRef]
- Sabater-Lleal, M.; Huffman, J.E.; De Vries, P.S.; Marten, J.; Mastrangelo, M.A.; Song, C.; Pankratz, N.; Ward-Caviness, C.K.; Yanek, L.R.; Trompet, S.; et al. Genome-wide association transethnic meta-analyses identifies novel associations regulating coagulation factor VIII and von Willebrand factor plasma levels. Circulation 2019, 139, 620–635. [Google Scholar] [CrossRef]
- Zhu, Y.; Wu, G.; Yan, W.; Zhan, H.; Sun, P. miR-146b-5p regulates cell growth, invasion, and metabolism by targeting PDHB in colorectal cancer. Am. J. Cancer Res. 2017, 7, 1136. [Google Scholar]
- Mayr, J.A.; Feichtinger, R.G.; Tort, F.; Ribes, A.; Sperl, W. Lipoic acid biosynthesis defects. J. Inherit. Metab. Dis. 2014, 37, 553–563. [Google Scholar] [CrossRef] [PubMed]
- Soreze, Y.; Boutron, A.; Habarou, F.; Barnerias, C.; Nonnenmacher, L.; Delpech, H.; Mamoune, A.; Chrétien, D.; Hubert, L.; Bole-Feysot, C.; et al. Mutations in human lipoyltransferase gene LIPT1cause a Leigh disease with secondary deficiency for pyruvate and alpha-ketoglutarate dehydrogenase. Orphanet J. Rare Dis. 2013, 8, 192. [Google Scholar] [CrossRef] [PubMed]
- Tort, F.; Ferrer-Cortès, X.; Thió, M.; Navarro-Sastre, A.; Matalonga, L.; Quintana, E.; Bujan, N.; Arias, A.; García-Villoria, J.; Acquaviva, C.; et al. Mutations in the lipoyltransferase LIPT1 gene cause a fatal disease associated with a specific lipoylation defect of the 2-ketoacid dehydrogenase complexes. Hum. Mol. Genet. 2014, 23, 1907–1915. [Google Scholar] [CrossRef]
- Chen, Y.; Xu, T.; Xie, F.; Wang, L.; Liang, Z.; Li, D.; Liang, Y.; Zhao, K.; Qi, X.; Yang, X.; et al. Evaluating the biological functions of the prognostic genes identified by the Pathology Atlas in bladder cancer. Oncol. Rep. 2021, 45, 191–201. [Google Scholar] [CrossRef] [PubMed]
- Lv, H.; Liu, X.; Zeng, X.; Liu, Y.; Zhang, C.; Zhang, Q.; Xu, J. Comprehensive analysis of cuproptosis-related genes in immune infiltration and prognosis in melanoma. Front. Pharmacol. 2022, 13, 930041. [Google Scholar] [CrossRef] [PubMed]
- Yan, C.; Niu, Y.; Ma, L.; Tian, L.; Ma, J. System analysis based on the cuproptosis-related genes identifies LIPT1 as a novel therapy target for liver hepatocellular carcinoma. J. Transl. Med. 2022, 20, 452. [Google Scholar] [CrossRef]
- Zhang, W.T.; Gong, Y.M.; Zhang, C.Y.; Pan, J.S.; Huang, T.; Li, Y.X. A novel cuprotosis-related gene Fdx1 signature for overall survival prediction in clear cell renal cell carcinoma patients. BioMed Res. Int. 2022, 2022, 9196540. [Google Scholar] [CrossRef]
- Zhang, C.; Zeng, Y.; Guo, X.; Zhang, J.; Wang, K.; Ji, M.; Huang, S. Pan-cancer analyses confirmed the cuproptosis-related gene FDX1 as an immunotherapy predictor and prognostic biomarker. Front. Genet. 2022, 13, 923737. [Google Scholar] [CrossRef]
- Huang, Z.; Xiao, C.; Yang, L.; Jin, L.; Lin, W.; Zhang, F.; Huang, S. Prognostic and immunological role of cuproptosis-related protein FDX1 in pan-cancer. Front. Genet. 2022, 13, 962028. [Google Scholar]
- Zhang, Z.; Ma, Y.; Guo, X.; Du, Y.; Zhu, Q.; Wang, X.; Duan, C. FDX1 can Impact the Prognosis and Mediate the Metabolism of Lung Adenocarcinoma. Front. Pharmacol. 2021, 12, 749134. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Li, L.; Sun, Q. High expression of cuproptosis-related SLC31A1 gene in relation to unfavorable outcome and deregulated immune cell infiltration in breast cancer: An analysis based on public databases. BMC Bioinform. 2022, 23, 350. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhao, X.; Li, J.; Wang, X.; Hu, W.; Zhang, X. Four m6A RNA methylation gene signatures and their prognostic values in lung adenocarcinoma. Technol. Cancer Res. Treat. 2022, 21, 15330338221085373. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Wang, H.; Huang, H.; Zhang, L.; Wang, D.; Wan, Y. m6A RNA methylation regulators participate in the malignant progression and have clinical prognostic value in lung adenocarcinoma. Front. Genet. 2020, 11, 994. [Google Scholar] [CrossRef] [PubMed]
- Tian, J.; Fu, C.; Peng, Q.; Yang, J.; Fan, X.; Zeng, X.; Qing, W.; Wu, Y. Construction of immune cell infiltration score model to assess prognostic ability of tumor immune environment in lung adenocarcinoma. Am. J. Transl. Res. 2023, 15, 1730. [Google Scholar]
- Liu, S.Y.; Huang, D.J.; Tang, E.Y.; Zhang, R.X.; Zhang, Z.M.; Gao, T.; Xu, G.Q. Construction of a non-negative matrix factorization model of immunogenic cell death-related genes in lung adenocarcinoma and analysis of survival prognosis. Heliyon 2023, 9, e14820. [Google Scholar] [CrossRef]
- Xiaona, X.; Liu, Q.; Zhou, X.; Liang, R.; Yang, S.; Xu, M.; Zhao, H.; Li, C.; Chen, Y.; Xueding, C. Comprehensive analysis of cuproptosis-related genes in immune infiltration and prognosis in lung adenocarcinoma. Comput. Biol. Med. 2023, 158, 106831. [Google Scholar] [CrossRef]
- Jiang, G.; Song, C.; Wang, X.; Xu, Y.; Li, H.; He, Z.; Cai, Y.; Zheng, M.; Mao, W. The multi-omics analysis identifies a novel cuproptosis-anoikis-related gene signature in prognosis and immune infiltration characterization of lung adenocarcinoma. Heliyon 2023, 9, e14091. [Google Scholar] [CrossRef]





| Model | Year 1 AUC | Year 3 AUC | Year 5 AUC |
|---|---|---|---|
| DNN | 0.606 | 0.621 | 0.603 |
| Cox | 0.601 | 0.586 | 0.584 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liang, P.; Chen, J.; Yao, L.; Hao, Z.; Chang, Q. A Deep Learning Approach for Prognostic Evaluation of Lung Adenocarcinoma Based on Cuproptosis-Related Genes. Biomedicines 2023, 11, 1479. https://doi.org/10.3390/biomedicines11051479
Liang P, Chen J, Yao L, Hao Z, Chang Q. A Deep Learning Approach for Prognostic Evaluation of Lung Adenocarcinoma Based on Cuproptosis-Related Genes. Biomedicines. 2023; 11(5):1479. https://doi.org/10.3390/biomedicines11051479
Chicago/Turabian StyleLiang, Pengchen, Jianguo Chen, Lei Yao, Zezhou Hao, and Qing Chang. 2023. "A Deep Learning Approach for Prognostic Evaluation of Lung Adenocarcinoma Based on Cuproptosis-Related Genes" Biomedicines 11, no. 5: 1479. https://doi.org/10.3390/biomedicines11051479
APA StyleLiang, P., Chen, J., Yao, L., Hao, Z., & Chang, Q. (2023). A Deep Learning Approach for Prognostic Evaluation of Lung Adenocarcinoma Based on Cuproptosis-Related Genes. Biomedicines, 11(5), 1479. https://doi.org/10.3390/biomedicines11051479







