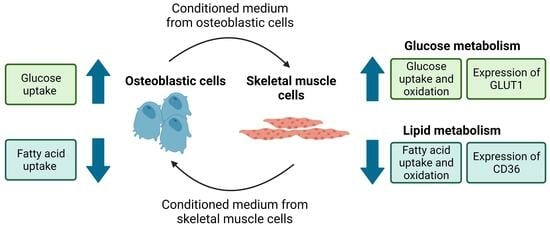
Interplay between Cultured Human Osteoblastic and Skeletal Muscle Cells: Effects of Conditioned Media on Glucose and Fatty Acid Metabolism
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Ethical Approvals
2.3. Cell Culturing and Harvesting of Conditioned Media
2.4. Total Protein Harvesting and Immunoblotting
2.5. RNA Isolation and qPCR
2.6. Acute Substrate Oxidation Assay
2.7. Lipid Distribution
2.8. Protein Synthesis by Leucine Incorporation
2.9. Proteomic Analysis
2.10. Statistics
3. Results
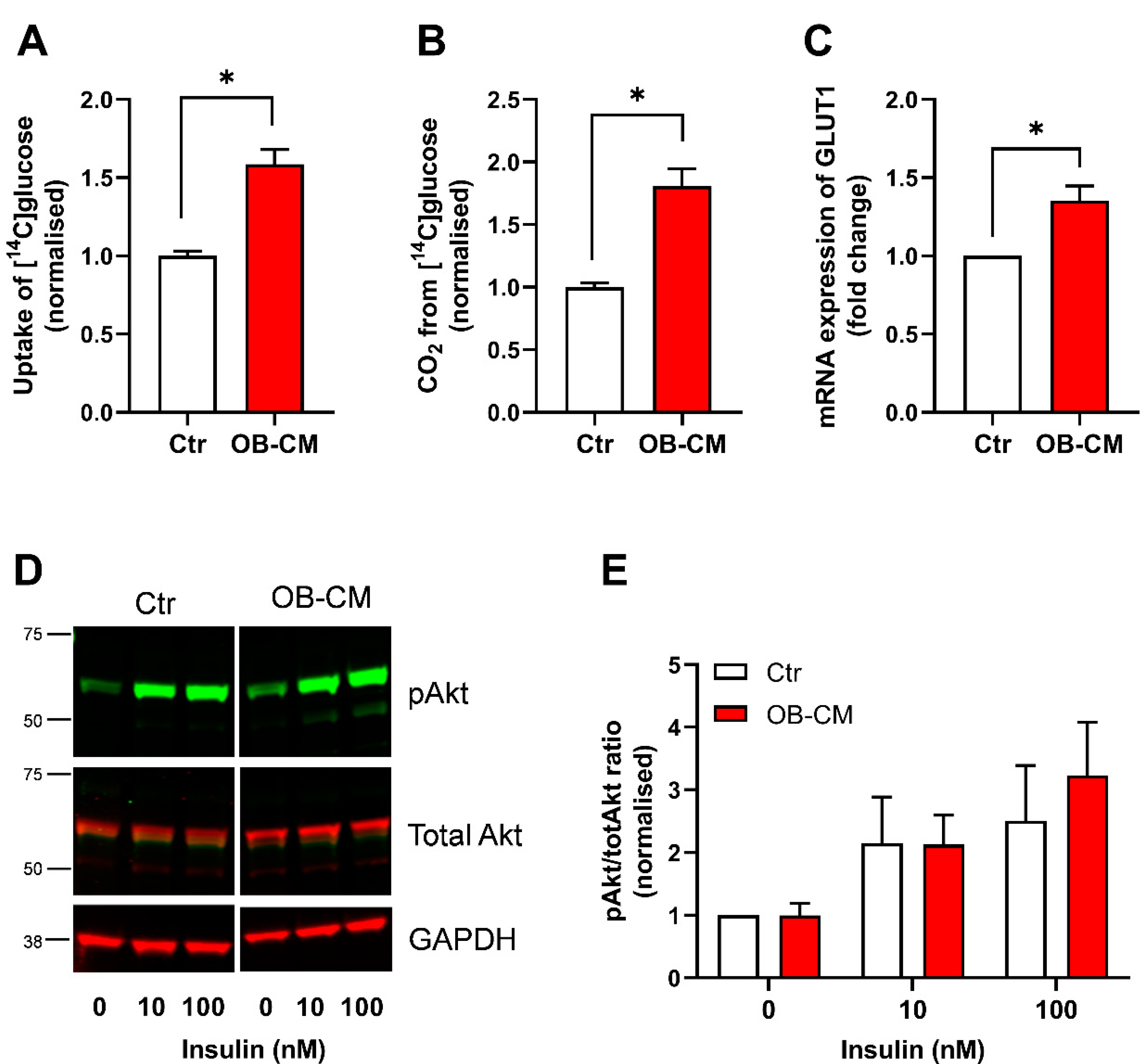
3.1. Conditioned Medium from Differentiated Osteoblastic Cells Increases Glucose Metabolism in Human Skeletal Muscle Cells
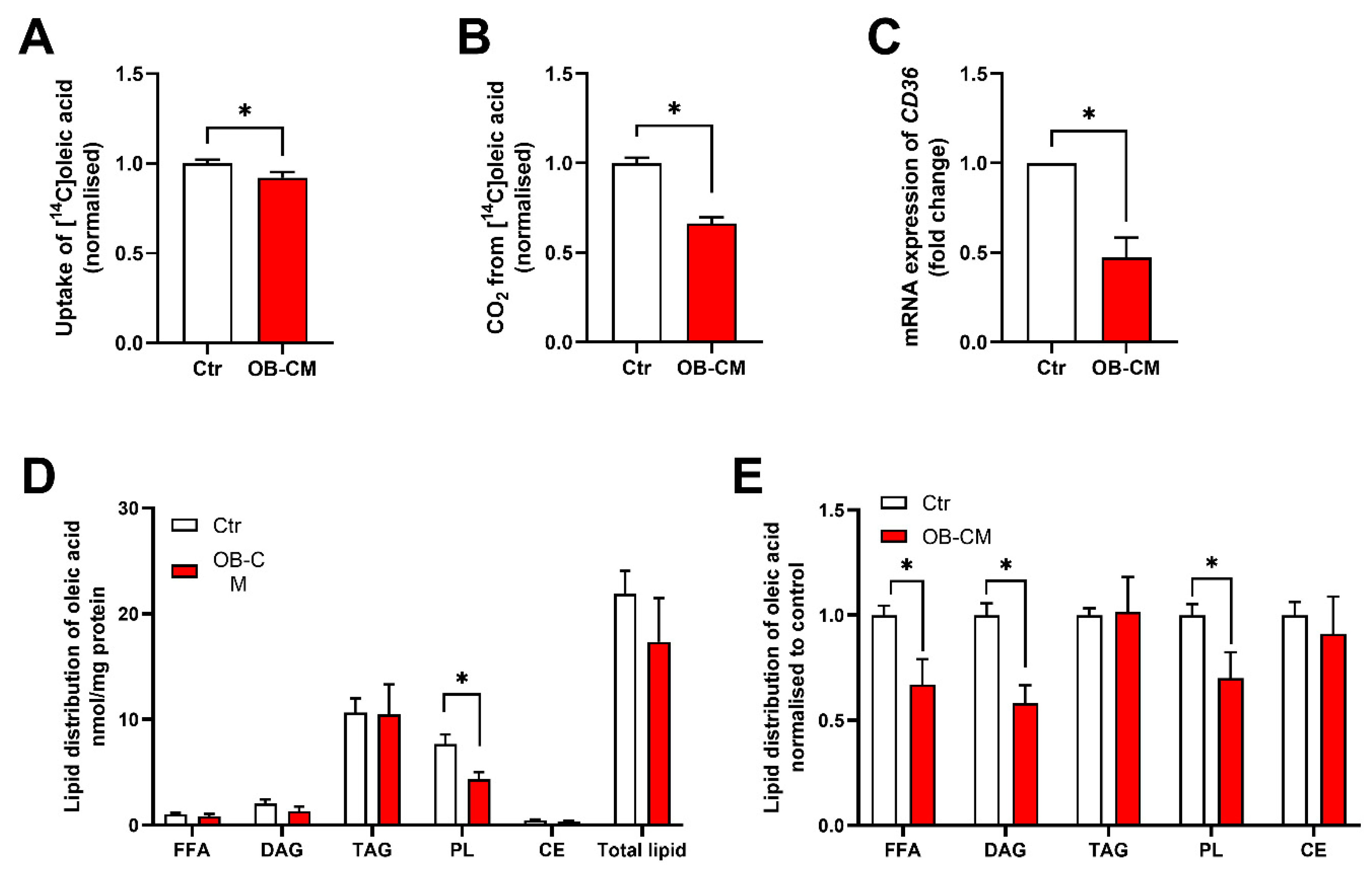
3.2. Conditioned Medium from Differentiated Osteoblastic Cells Reduces Oleic Acid Metabolism in Human Skeletal Muscle Cells
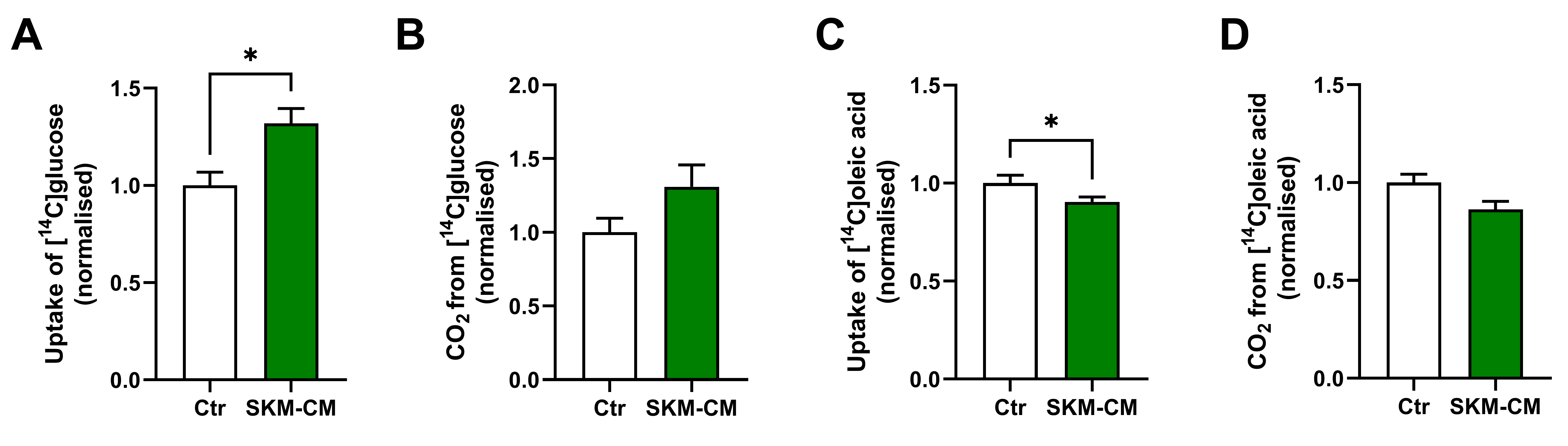
3.3. Conditioned Medium from Differentiated Myotubes Increases Glucose Uptake and Reduces Oleic Acid Uptake in Osteoblastic Cells
3.4. Secretome Analysis of Conditioned Medium from Differentiated Osteoblastic Cells and Myotubes
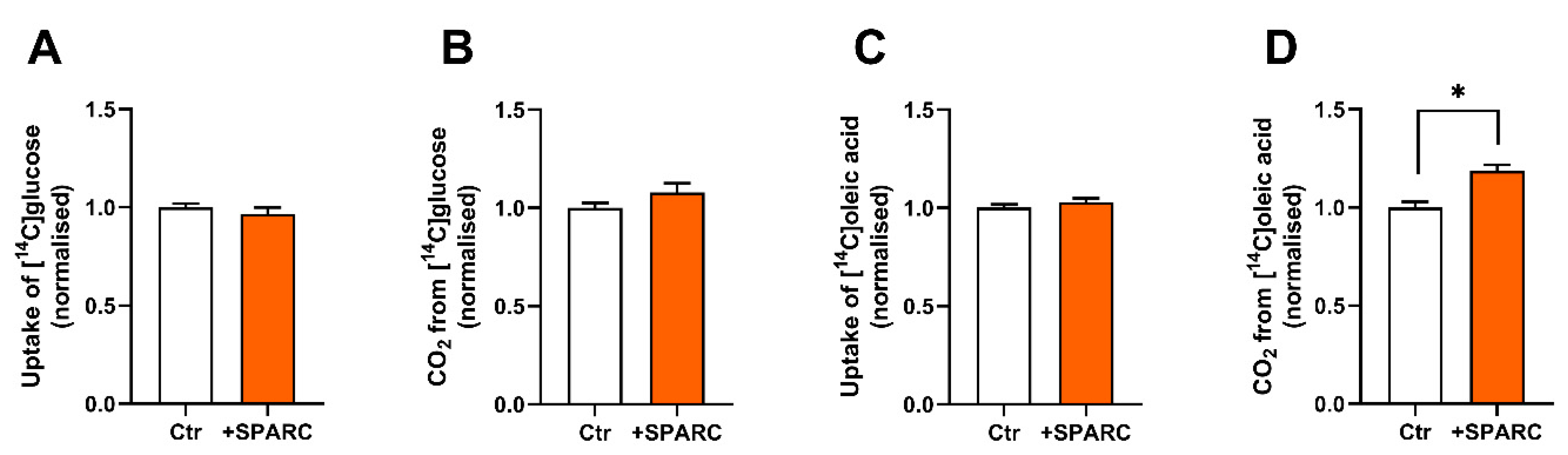
3.5. SPARC Increases Oleic Acid Oxidation in Human Skeletal Muscle Cells
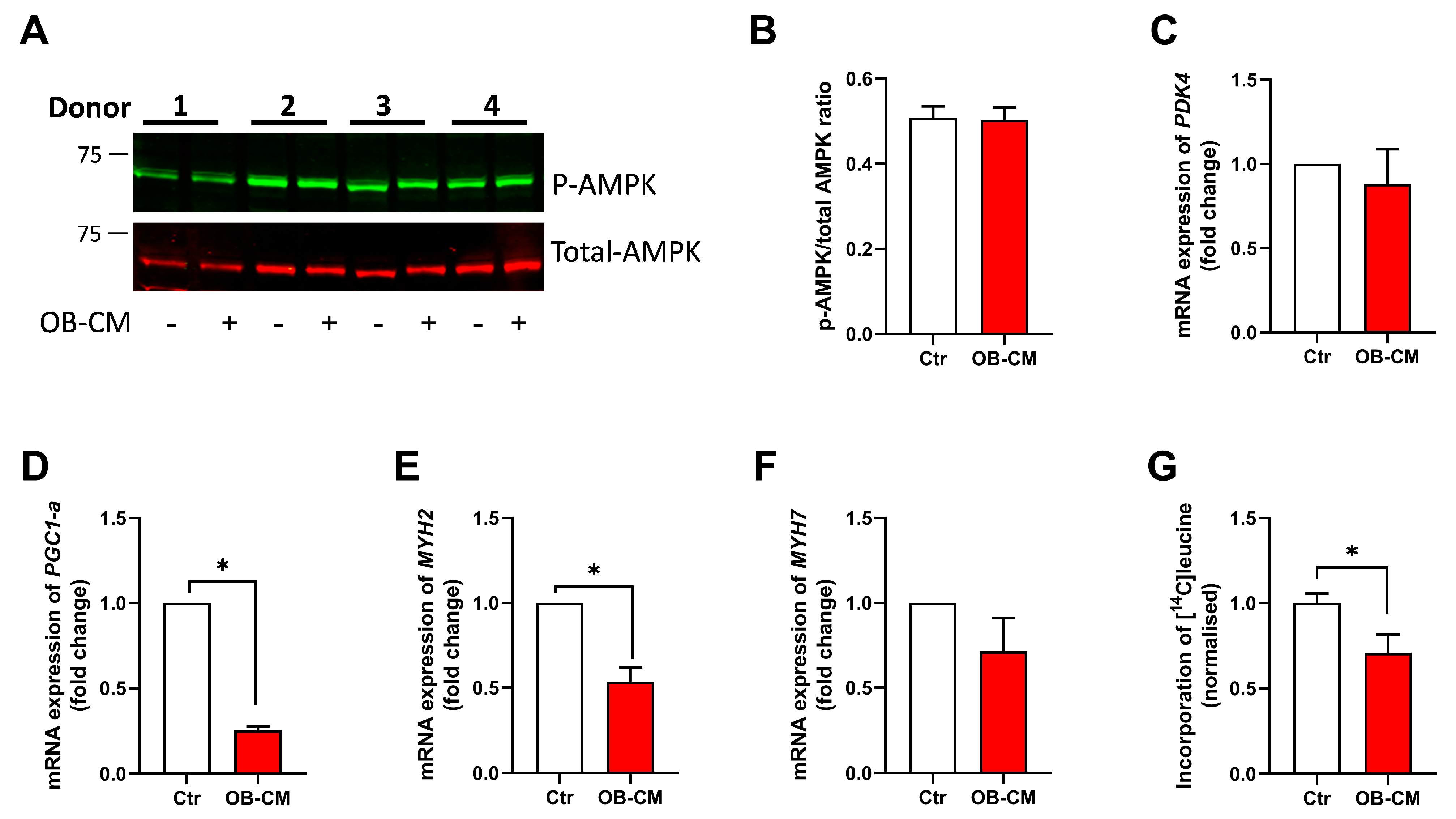
3.6. Conditioned Medium from Differentiated Osteoblastic Cells Inhibits PGC1α mRNA Expression and Protein Synthesis in Human Skeletal Muscle Cells
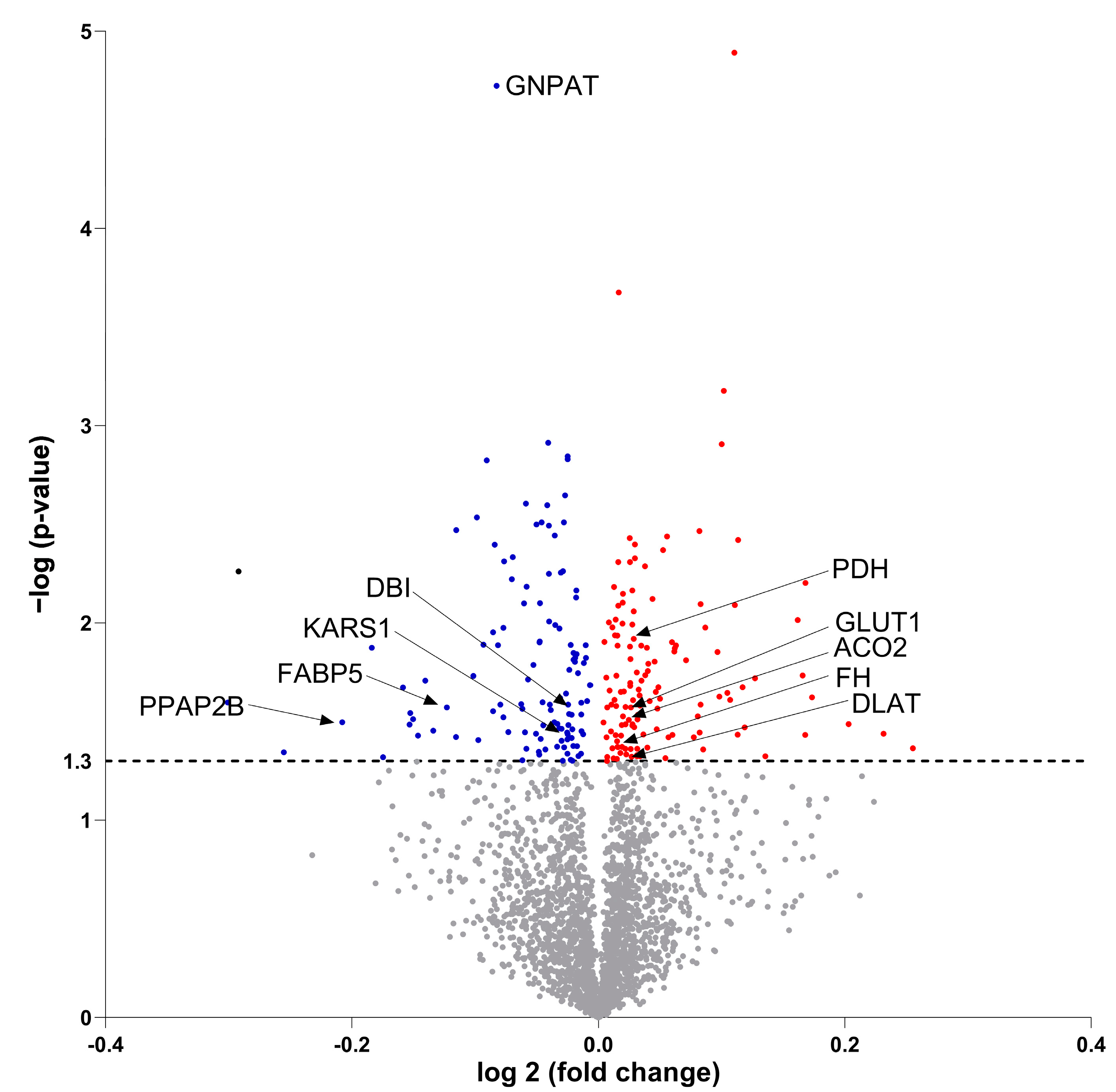
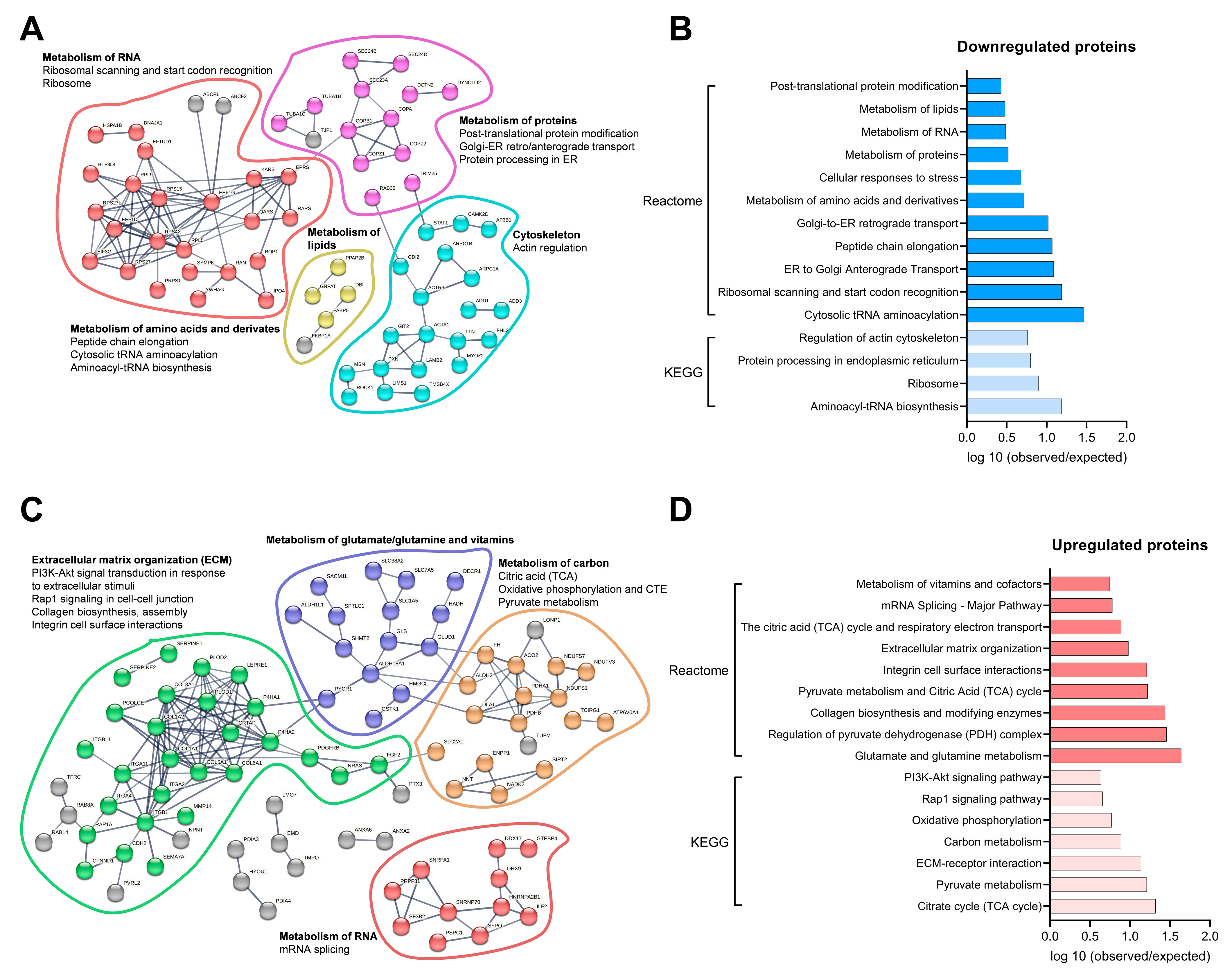
3.7. Molecular Proteomic Changes Correlate with Functional Effects of OB-CM on Myotubes Energy Metabolism
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kirk, B.; Feehan, J.; Lombardi, G.; Duque, G. Muscle, Bone, and Fat Crosstalk: The Biological Role of Myokines, Osteokines, and Adipokines. Curr. Osteoporos. Rep. 2020, 18, 388–400. [Google Scholar] [CrossRef]
- Li, G.; Zhang, L.; Wang, D.; AIQudsy, L.; Jiang, J.X.; Xu, H.; Shang, P. Muscle-bone crosstalk and potential therapies for sarco-osteoporosis. J. Cell. Biochem. 2019, 120, 14262–14273. [Google Scholar] [CrossRef] [PubMed]
- Buvinic, S.; Balanta-Melo, J.; Kupczik, K.; Vasquez, W.; Beato, C.; Toro-Ibacache, V. Muscle-Bone Crosstalk in the Masticatory System: From Biomechanical to Molecular Interactions. Front. Endocrinol. 2020, 11, 606947. [Google Scholar] [CrossRef]
- Pedersen, B.K. Muscles and their myokines. J. Exp. Biol. 2011, 214, 337–346. [Google Scholar] [CrossRef] [PubMed]
- Severinsen, M.C.K.; Pedersen, B.K. Muscle-Organ Crosstalk: The Emerging Roles of Myokines. Endocr. Rev. 2020, 41, 594–609. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, L.; Hojman, P. Muscle-to-organ cross talk mediated by myokines. Adipocyte 2012, 1, 164–167. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.Y.; Park, S.J.; Han, S.A.; Lee, S.H.; Koh, J.M.; Hamrick, M.W.; Kim, B.J. The effects of myokines on osteoclasts and osteoblasts. Biochem. Biophys. Res. Commun. 2019, 517, 749–754. [Google Scholar] [CrossRef]
- Mera, P.; Laue, K.; Wei, J.; Berger, J.M.; Karsenty, G. Osteocalcin is necessary and sufficient to maintain muscle mass in older mice. Mol. Metab. 2016, 5, 1042–1047. [Google Scholar] [CrossRef]
- Mera, P.; Laue, K.; Ferron, M.; Confavreux, C.; Wei, J.; Galan-Diez, M.; Lacampagne, A.; Mitchell, S.J.; Mattison, J.A.; Chen, Y.; et al. Osteocalcin Signaling in Myofibers Is Necessary and Sufficient for Optimum Adaptation to Exercise. Cell Metab. 2017, 25, 218. [Google Scholar] [CrossRef]
- Yoshida, Y.; Jain, S.S.; McFarlan, J.T.; Snook, L.A.; Chabowski, A.; Bonen, A. Exercise- and training-induced upregulation of skeletal muscle fatty acid oxidation are not solely dependent on mitochondrial machinery and biogenesis. J. Physiol. 2013, 591, 4415–4426. [Google Scholar] [CrossRef]
- Frontera, W.R.; Ochala, J. Skeletal muscle: A brief review of structure and function. Calcif. Tissue Int. 2015, 96, 183–195. [Google Scholar] [CrossRef] [PubMed]
- DeFronzo, R.A.; Tripathy, D. Skeletal muscle insulin resistance is the primary defect in type 2 diabetes. Diabetes Care 2009, 32 (Suppl. S2), S157–S163. [Google Scholar] [CrossRef] [PubMed]
- Civitelli, R.; Ziambaras, K. Calcium and phosphate homeostasis: Concerted interplay of new regulators. J. Endocrinol. Investig. 2011, 34, 3–7. [Google Scholar]
- DiGirolamo, D.J.; Kiel, D.P.; Esser, K.A. Bone and skeletal muscle: Neighbors with close ties. J. Bone Miner. Res. 2013, 28, 1509–1518. [Google Scholar] [CrossRef] [PubMed]
- Lee, N.K.; Sowa, H.; Hinoi, E.; Ferron, M.; Ahn, J.D.; Confavreux, C.; Dacquin, R.; Mee, P.J.; McKee, M.D.; Jung, D.Y.; et al. Endocrine regulation of energy metabolism by the skeleton. Cell 2007, 130, 456–469. [Google Scholar] [CrossRef] [PubMed]
- Mengeste, A.M.; Nikolic, N.; Dalmao Fernandez, A.; Feng, Y.Z.; Nyman, T.A.; Kersten, S.; Haugen, F.; Kase, E.T.; Aas, V.; Rustan, A.C.; et al. Insight into the Metabolic Adaptations of Electrically Pulse-Stimulated Human Myotubes Using Global Analysis of the Transcriptome and Proteome. Front. Physiol. 2022, 13, 928195. [Google Scholar] [CrossRef] [PubMed]
- Lund, J.; Rustan, A.C.; Løvsletten, N.G.; Mudry, J.M.; Langleite, T.M.; Feng, Y.Z.; Stensrud, C.; Brubak, M.G.; Drevon, C.A.; Birkeland, K.I.; et al. Exercise in vivo marks human myotubes in vitro: Training-induced increase in lipid metabolism. PLoS ONE 2017, 12, e0175441. [Google Scholar] [CrossRef]
- Jafari, A.; Qanie, D.; Andersen, T.L.; Zhang, Y.; Chen, L.; Postert, B.; Parsons, S.; Ditzel, N.; Khosla, S.; Johansen, H.T.; et al. Legumain Regulates Differentiation Fate of Human Bone Marrow Stromal Cells and Is Altered in Postmenopausal Osteoporosis. Stem Cell Rep. 2017, 8, 373–386. [Google Scholar] [CrossRef]
- Wensaas, A.J.; Rustan, A.C.; Lovstedt, K.; Kull, B.; Wikstrom, S.; Drevon, C.A.; Hallen, S. Cell-based multiwell assays for the detection of substrate accumulation and oxidation. J. Lipids Res. 2007, 48, 961–967. [Google Scholar] [CrossRef]
- Folch, J.; Lees, M.; Sloane Stanley, G.H. A simple method for the isolation and purification of total lipides from animal tissues. J. Biol. Chem. 1957, 226, 497–509. [Google Scholar] [CrossRef]
- Batth, T.S.; Tollenaere, M.X.; Rüther, P.; Gonzalez-Franquesa, A.; Prabhakar, B.S.; Bekker-Jensen, S.; Deshmukh, A.S.; Olsen, J.V. Protein Aggregation Capture on Microparticles Enables Multipurpose Proteomics Sample Preparation. Mol. Cell. Proteom. 2019, 18, 1027–1035. [Google Scholar] [CrossRef] [PubMed]
- Rath, S.; Sharma, R.; Gupta, R.; Ast, T.; Chan, C.; Durham, T.J.; Goodman, R.P.; Grabarek, Z.; Haas, M.E.; Hung, W.H.W.; et al. MitoCarta3.0: An updated mitochondrial proteome now with sub-organelle localization and pathway annotations. Nucleic Acids Res. 2021, 49, D1541–D1547. [Google Scholar] [CrossRef] [PubMed]
- Perez-Riverol, Y.; Bai, J.; Bandla, C.; Hewapathirana, S.; García-Seisdedos, D.; Kamatchinathan, S.; Kundu, D.; Prakash, A.; Frericks-Zipper, A.; Eisenacher, M.; et al. The PRIDE database resources in 2022: A Hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 2022, 50, D543–D552. [Google Scholar] [CrossRef] [PubMed]
- Melouane, A.; Carbonell, A.; Yoshioka, M.; Puymirat, J.; St-Amand, J. Implication of SPARC in the modulation of the extracellular matrix and mitochondrial function in muscle cells. PLoS ONE 2018, 13, e0192714. [Google Scholar] [CrossRef] [PubMed]
- Melouane, A.; Yoshioka, M.; Kanzaki, M.; St-Amand, J. Sparc, an EPS-induced gene, modulates the extracellular matrix and mitochondrial function via ILK/AMPK pathways in C2C12 cells. Life Sci. 2019, 229, 277–287. [Google Scholar] [CrossRef] [PubMed]
- Mathes, S.; Fahrner, A.; Luca, E.; Krutzfeldt, J. Growth hormone/IGF-I-dependent signaling restores decreased expression of the myokine SPARC in aged skeletal muscle. J. Mol. Med. 2022, 100, 1647–1658. [Google Scholar] [CrossRef]
- Hardie, D.G. Energy sensing by the AMP-activated protein kinase and its effects on muscle metabolism. Proc. Nutr. Soc. 2011, 70, 92–99. [Google Scholar] [CrossRef]
- Battafarano, G.; Rossi, M.; Marampon, F.; Minisola, S.; Del Fattore, A. Bone Control of Muscle Function. Int. J. Mol. Sci. 2020, 21, 1178. [Google Scholar] [CrossRef]
- Colaianni, G.; Storlino, G.; Sanesi, L.; Colucci, S.; Grano, M. Myokines and Osteokines in the Pathogenesis of Muscle and Bone Diseases. Curr. Osteoporos. Rep. 2020, 18, 401–407. [Google Scholar] [CrossRef]
- Laurens, C.; Parmar, A.; Murphy, E.; Carper, D.; Lair, B.; Maes, P.; Vion, J.; Boulet, N.; Fontaine, C.; Marquès, M.; et al. Growth and differentiation factor 15 is secreted by skeletal muscle during exercise and promotes lipolysis in humans. JCI Insight 2020, 5, e131870. [Google Scholar] [CrossRef] [PubMed]
- Koshal, P.; Matera, I.; Abruzzese, V.; Ostuni, A.; Bisaccia, F. The Crosstalk between HepG2 and HMC-III Cells: In Vitro Modulation of Gene Expression with Conditioned Media. Int. J. Mol. Sci. 2022, 23, 14443. [Google Scholar] [CrossRef] [PubMed]
- Vis, M.A.M.; Ito, K.; Hofmann, S. Impact of Culture Medium on Cellular Interactions in in vitro Co-culture Systems. Front. Bioeng. Biotechnol. 2020, 8, 911. [Google Scholar] [CrossRef]
- Alekos, N.S.; Moorer, M.C.; Riddle, R.C. Dual Effects of Lipid Metabolism on Osteoblast Function. Front. Endocrinol. 2020, 11, 578194. [Google Scholar] [CrossRef]
- Goodpaster, B.H.; Sparks, L.M. Metabolic Flexibility in Health and Disease. Cell Metab. 2017, 25, 1027–1036. [Google Scholar] [CrossRef]
- Donat, A.; Knapstein, P.R.; Jiang, S.; Baranowsky, A.; Ballhause, T.M.; Frosch, K.H.; Keller, J. Glucose Metabolism in Osteoblasts in Healthy and Pathophysiological Conditions. Int. J. Mol. Sci. 2021, 22, 4120. [Google Scholar] [CrossRef]
- Shen, L.; Hu, G.; Karner, C.M. Bioenergetic Metabolism in Osteoblast Differentiation. Curr. Osteoporos. Rep. 2022, 20, 53–64. [Google Scholar] [CrossRef]
- Alghannam, A.F.; Ghaith, M.M.; Alhussain, M.H. Regulation of Energy Substrate Metabolism in Endurance Exercise. Int. J. Environ. Res. Public Health 2021, 18, 4963. [Google Scholar] [CrossRef]
- Son, J.S.; Chae, S.A.; Testroet, E.D.; Du, M.; Jun, H.P. Exercise-induced myokines: A brief review of controversial issues of this decade. Expert Rev. Endocrinol. Metab. 2018, 13, 51–58. [Google Scholar] [CrossRef]
- Norheim, F.; Raastad, T.; Thiede, B.; Rustan, A.C.; Drevon, C.A.; Haugen, F. Proteomic identification of secreted proteins from human skeletal muscle cells and expression in response to strength training. Am. J. Physiol. Endocrinol. Metab. 2011, 301, E1013–E1021. [Google Scholar] [CrossRef]
- Alquier, T.; Christian-Hinman, C.A.; Alfonso, J.; Faergeman, N.J. From benzodiazepines to fatty acids and beyond: Revisiting the role of ACBP/DBI. Trends Endocrinol. Metab. 2021, 32, 890–903. [Google Scholar] [CrossRef]
- Busnelli, M.; Manzini, S.; Parolini, C.; Escalante-Alcalde, D.; Chiesa, G. Lipid phosphate phosphatase 3 in vascular pathophysiology. Atherosclerosis 2018, 271, 156–165. [Google Scholar] [CrossRef] [PubMed]
- George Warren, W.; Osborn, M.; Yates, A.; Wright, K.; O’Sullivan, S.E. The emerging role of fatty acid binding protein 5 (FABP5) in cancers. Drug Discov. Today 2023, 28, 103628. [Google Scholar] [CrossRef] [PubMed]
- Dias, J.; Octobre, G.; Kobbi, L.; Comisso, M.; Flisiak, S.; Mirande, M. Activation of human mitochondrial lysyl-tRNA synthetase upon maturation of its premitochondrial precursor. Biochemistry 2012, 51, 909–916. [Google Scholar] [CrossRef] [PubMed]
- Russell, A.P. PGC-1alpha and exercise: Important partners in combating insulin resistance. Curr. Diabetes Rev. 2005, 1, 175–181. [Google Scholar] [CrossRef]
- Schiaffino, S.; Reggiano, C. Fiber types in mammalian skeletal muscles. Physiol. Rev. 2011, 91, 1447–1531. [Google Scholar] [CrossRef]
| Gene | Acc.no. | Forward Sequence | Reverse Sequence |
|---|---|---|---|
| CD36 | L06850 | AGTCACTGCGACATGATTAATGGT | CTGCAATACCTGGCTTTTCTCAA |
| GLUT1/SLC2A1 | K03195 | CAGCAGCCCTAAGGATCTCTCA | CCGGCTCGGCTGACATC |
| MYH2 | C5814 | AAGGTCGGCAATGAGTATGTCA | CAACCATCCACAGGAACATCTTC |
| MYH7 | NM_000257.2 | CTCTGCACAGGGAAAATCTGAA | CCCCTGGACTTTGTCTCATT |
| PDK4 | BC040239 | TTTCCAGAACCAACCAATTCACA | TGC CCG CAT TGC ATT CTT A |
| PGC1α | NM013261.3 | AAAGGATGCGCTCTCGTTCA | TCTACTGCCTGGAGACCTTGATC |
| GAPDH | NM002046 | TGCACCACCACCTGCTTAGC | GGCATGGACTGTGGTCAT GAG |
| RPLP0 | M17885 | CCATTCTATCATCAACGGGTACAA | AGCAAGTGGGAAGGTGTAATCC |
| Uniprot ID | Protein Symbol | Protein Name | Biological Process (Gene Ontology) | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | P06733 | ENO1 | Alpha-enolase | ||||||||
| 2 | P62937 | PPIA | Peptidyl-prolyl cis-trans isomerase A | ||||||||
| 3 | P63261 | ACTG1 | Actin, cytoplasmic 2 | ||||||||
| 4 | P08670 | VIM | Vimentin | ||||||||
| 5 | P09486 | SPARC | Secreted protein acidic and cysteine rich | ||||||||
| 6 | P02765 | AHSG | Alpha-2-HS-glycoprotein | ||||||||
| 7 | P23528 | CFL1 | Cofilin-1 | ||||||||
| 8 | P09382 | LGALS1 | Galectin-1 | ||||||||
| 9 | P14618 | PKM | Pyruvate kinase | ||||||||
| 10 | P37802 | TAGLN2 | Transgelin-2 | ||||||||
| 11 | P68363 | TUBA1B | Tubulin alpha-1B chain | ||||||||
| 12 | O00299 | CLIC1 | Chloride intracellular channel protein 1 | ||||||||
| 13 | P26022 | PTX3 | Pentraxin-related protein | ||||||||
| 14 | P04792 | HSPB1 | Heat shock protein beta-1 | ||||||||
| 15 | P60709 | ACTB | Actin, cytoplasmic 1 | ||||||||
| 16 | P00338 | LDHA | L-lactate dehydrogenase A chain | ||||||||
| 17 | P09211 | GSTP1 | Glutathione S-transferase P | ||||||||
| 18 | P07437 | TUBB | Tubulin beta chain | ||||||||
| 19 | P07737 | PFN1 | Profilin-1 | ||||||||
| 20 | P07355 | ANXA2 | Annexin 2 | ||||||||
| Uniprot ID | Protein Symbol | Protein Name | Biological Process (Gene Ontology) | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | P63313 | TMSB10 | Thymosin beta 10 | ||||||||
| 2 | P62328 | TMSB4X | Thymosin beta 4 X-linked | ||||||||
| 3 | P63261 | ACTG1 | Actin, cytoplasmic 2 | ||||||||
| 4 | P07737 | PFN1 | Profilin 1 | ||||||||
| 5 | P02795 | MT2A | Metallothionein 2A | ||||||||
| 6 | P08670 | VIM | Vimentin | ||||||||
| 7 | P09486 | SPARC | Secreted protein acidic and cysteine rich | ||||||||
| 8 | P63104 | YWHAZ | Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation zeta | ||||||||
| 9 | P08123 | COL1A2 | Collagen type I alpha 2 chain | ||||||||
| 10 | P09382 | LGALS1 | Galectin 1 | ||||||||
| 11 | P02452 | COL1A1 | Collagen type I alpha 1 chain | ||||||||
| 12 | P0DP25 | CALM3 | Calmodulin 3 | ||||||||
| 13 | P07951 | TPM2 | Tropomyosin 2 | ||||||||
| 14 | P62258 | YWHAE | Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation epsilon | ||||||||
| 15 | Q16270 | IGFBP7 | Insulin-like growth factor binding protein 7 | ||||||||
| 16 | P17661 | DES | Desmin | ||||||||
| 17 | P41222 | PTGDS | Prostaglandin D2 synthase | ||||||||
| 18 | P10599 | TXN | Thioredoxin | ||||||||
| 19 | P01033 | TIMP1 | TIMP metallopeptidase inhibitor 1 | ||||||||
| 20 | P06454 | PTMA | Prothymosin alpha | ||||||||
| Uniprot ID | Protein Symbol | Protein Name | Log2 (Fold Change) | p-Value |
|---|---|---|---|---|
| Q16698 | DECR1 | 2,4-dienoyl-CoA reductase 1 | 0.025562 | 0.01318 |
| P35914 | HMGCL | 3-hydroxy-3-methylglutaryl-CoA lyase | 0.0276 | 0.010166 |
| Q99798 | ACO2 | Aconitase 2 | 0.010616 | 0.025911 |
| O75891 | ALDH1L1 | Aldehyde dehydrogenase 1 family member L1 | 0.029607 | 0.004703 |
| P54886 | ALDH18A | Aldehyde dehydrogenase 18 family member A1 | 0.014905 | 0.048835 |
| P05091 | ALDH2 | Aldehyde dehydrogenase 2 family member | 0.027491 | 0.006844 |
| Q9Y2R0 | COA3 | Cytochrome C oxidase assembly factor 3 | 0.203074 | 0.032505 |
| P10515 | DLAT | Dihydrolipoamide S-acetyltransferase | 0.022512 | 0.046349 |
| P07954 | FH | Fumarate hydratase | 0.007098 | 0.047717 |
| P00367 | GLUD1 | Glutamate dehydrogenase 1 | 0.014062 | 0.037031 |
| O94925 | GLS | Glutaminase | 0.048643 | 0.021184 |
| Q9Y2Q3 | GSTK1 | Glutathione S-transferase kappa 1 | 0.004066 | 0.031943 |
| Q16836 | HADH | Hydroxyacyl-CoA dehydrogenase | 0.02538 | 0.003717 |
| P36776 | LONP1 | Lon peptidase 1, mitochondrial | 0.026353 | 0.026783 |
| Q8IXM3 | MRPL41 | Mitochondrial ribosomal protein L41 | 0.117037 | 0.021162 |
| Q4G0N4 | NADK2 | NAD kinase 2, mitochondrial | 0.08224 | 0.0359 |
| P28331 | NDUFS1 | NADH:Ubiquinone oxidoreductase core subunit S1 | 0.015448 | 0.04241 |
| O75251 | NDUFS7 | NADH:Ubiquinone oxidoreductase core subunit S7 | 0.033855 | 0.047395 |
| Q13423 | NNT | Nicotinamide nucleotide transhydrogenase | 0.012271 | 0.048518 |
| Q10713 | PMPCA | Peptidase, mitochondrial processing subunit alpha | 0.029524 | 0.004 |
| Q9Y3E5 | PTRH2 | Peptidyl-TRNA hydrolase 2 | 0.019982 | 0.029719 |
| P32322 | PYCR1 | Pyrroline-5-carboxylate reductase 1 | 0.034958 | 0.019611 |
| P08559 | PDHA1 | Pyruvate dehydrogenase E1 subunit alpha 1 | 0.028673 | 0.012049 |
| P11177 | PDHB | Pyruvate dehydrogenase E1 subunit beta | 0.028679 | 0.008729 |
| P34897 | SHMT2 | Serine hydroxymethyltransferase 2 | 0.02749 | 0.033103 |
| P49411 | TUFM | Tu translation elongation factor, mitochondrial | 0.010111 | 0.035421 |
| P07108 | DBI | Acyl-CoA binding protein | −0.04511 | 0.02523 |
| Q15046 | KARS1 | Lysyl-TRNA synthetase 1 | −0.01335 | 0.035324 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lunde, N.N.; Osoble, N.M.M.; Fernandez, A.D.; Antobreh, A.S.; Jafari, A.; Singh, S.; Nyman, T.A.; Rustan, A.C.; Solberg, R.; Thoresen, G.H. Interplay between Cultured Human Osteoblastic and Skeletal Muscle Cells: Effects of Conditioned Media on Glucose and Fatty Acid Metabolism. Biomedicines 2023, 11, 2908. https://doi.org/10.3390/biomedicines11112908
Lunde NN, Osoble NMM, Fernandez AD, Antobreh AS, Jafari A, Singh S, Nyman TA, Rustan AC, Solberg R, Thoresen GH. Interplay between Cultured Human Osteoblastic and Skeletal Muscle Cells: Effects of Conditioned Media on Glucose and Fatty Acid Metabolism. Biomedicines. 2023; 11(11):2908. https://doi.org/10.3390/biomedicines11112908
Chicago/Turabian StyleLunde, Ngoc Nguyen, Nimo Mukhtar Mohamud Osoble, Andrea Dalmao Fernandez, Alfreda S. Antobreh, Abbas Jafari, Sachin Singh, Tuula A. Nyman, Arild C. Rustan, Rigmor Solberg, and G. Hege Thoresen. 2023. "Interplay between Cultured Human Osteoblastic and Skeletal Muscle Cells: Effects of Conditioned Media on Glucose and Fatty Acid Metabolism" Biomedicines 11, no. 11: 2908. https://doi.org/10.3390/biomedicines11112908