Cost Matrix of Molecular Pathology in Glioma—Towards AI-Driven Rational Molecular Testing and Precision Care for the Future
Abstract
1. Introduction
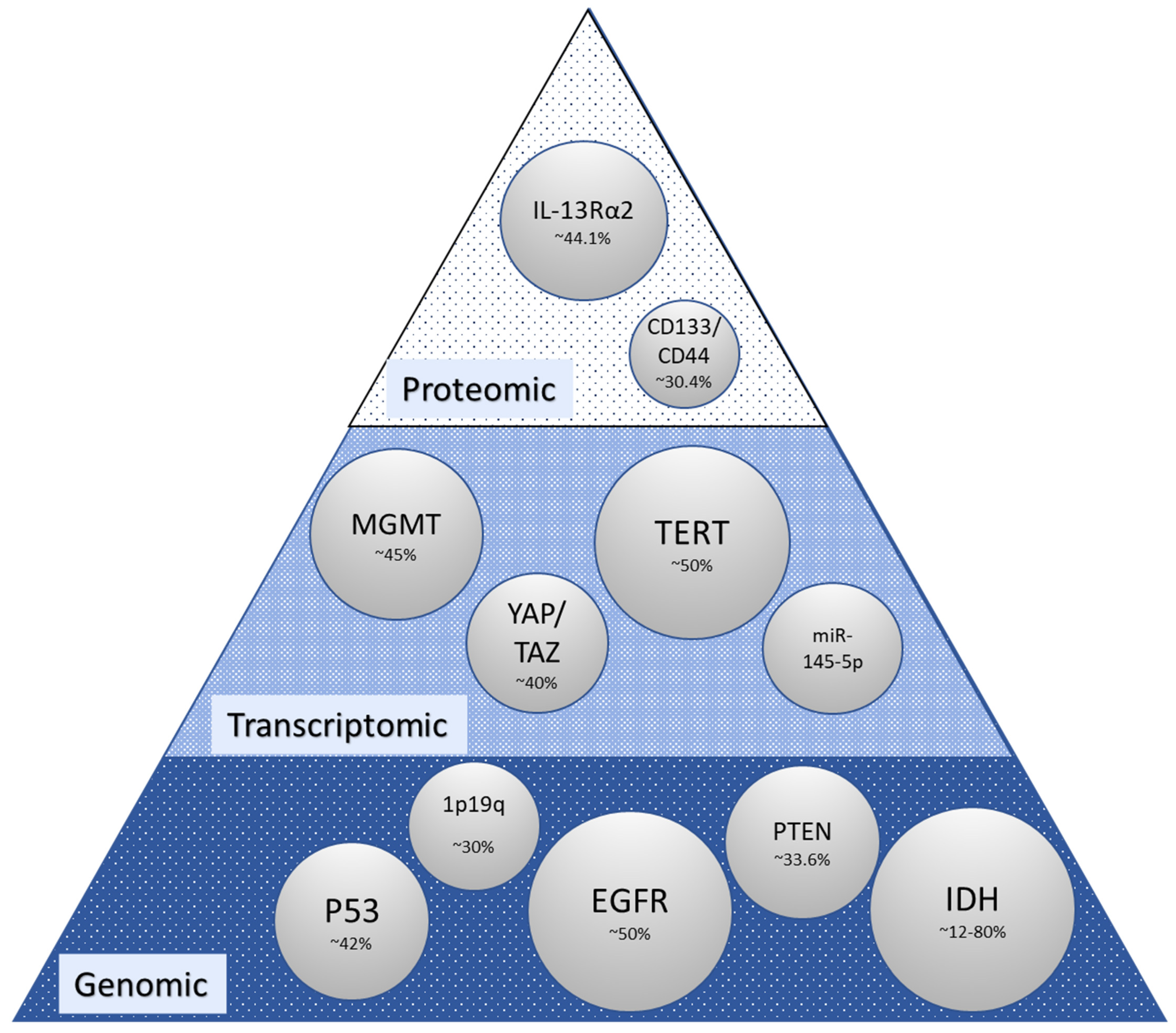
2. Molecular Alterations in Glioma
| Authors | Title | Finding |
|---|---|---|
| Kitange et al., 2009 [16] | Induction of MGMT expression is associated with temozolomide resistance in glioblastoma xenografts | MGMT expression is dynamically regulated in some MGMT nonmethylated tumors, and in these tumors protracted dosing regimens may not be effective. |
| Delfino et al., 2011 [45] | Therapy-, gender- and race-specific microRNA markers, target genes and networks related to glioblastoma recurrence and survival | Sensory perception and G-protein-coupled receptor processes were enriched among microRNA gene targets also associated with survival, and network visualization highlighted their relations. |
| Håvik et al., 2012 [46] | MGMT promoter methylation in gliomas-assessment by pyrosequencing and quantitative methylation-specific PCR | MGMT promoter methylation analysis gives sufficient prognostic information to merit its inclusion in the standard management of patients with high-grade gliomas, and in this study pyrosequencing seemed the better analytical method. |
| Aldape et al., 2015 [6] ** | Glioblastoma: pathology, molecular mechanisms and markers | IDH-mutant GBMs are clearly distinct from GBMs without IDH1/2 mutation with respect to molecular and clinical features, including prognosis. |
| Tanguturi et al., 2017 [47] | Characterization of MGMT and EGFR protein expression in glioblastoma and association with survival | There were several associations between GBM genomic subgroups and clinical or molecular prognostic covariates and validated known prognostic factors in all survival periods. |
| Asif et al., 2019 [27] | Comparative proteogenomic characterization of glioblastoma | Significantly mutated genes in GBM included TP53, EGFR, PIK3R1, PTEN, NF1, RET, and STAG2. MGMT methylation was present in two-thirds of cases. |
| Burgenske et al., 2019 [9] | Molecular profiling of long-term IDH-wildtype glioblastoma survivors | Unique attributes were observed in regard to altered gene expression and pathway enrichment. These attributes may be valuable prognostic markers and are worth further examination. |
| Gobin et al., 2019 [48] | A DNA Repair and Cell-Cycle Gene Expression Signature in Primary and Recurrent Glioblastoma: Prognostic Value and Clinical Implications | Classification of GBM tumors based on a DNA repair and cell cycle gene expression signature exposes vulnerabilities in standard-of-care therapies and offers the potential for personalized therapeutic strategies. |
| Neftel et al., 2019 [36] | An Integrative Model of Cellular States, Plasticity, and Genetics for Glioblastoma | Malignant cells in glioblastoma exist in four main cellular states that recapitulate distinct neural cell types, are influenced by the tumor microenvironment, and exhibit plasticity. The relative frequency of cells in each state varies between glioblastoma samples and is influenced by copy number amplifications of the CDK4, EGFR, and PDGFRA loci and by mutations in the NF1 locus, which each favor a defined state. |
| Oh et al., 2020 [49] | Integrated pharmaco-proteogenomics defines two subgroups in isocitrate dehydrogenase wild-type glioblastoma with prognostic and therapeutic opportunities | Two distinct binary classifications of IDH wild-type GBM tumors. GBM proteomic cluster 1 (GPC1) tumors exhibit Warburg-like features, neural stem cell markers, immune checkpoint ligands, and a poor prognostic biomarker, FKBP prolyl isomerase 9 (FKBP9). Meanwhile, GPC2 tumors show elevated oxidative phosphorylation-related proteins, differentiated oligodendrocyte and astrocyte markers, and a favorable prognostic biomarker, phosphoglycerate dehydrogenase (PHGDH). |
| Mata et al., 2020 [50] | Genetic and epigenetic landscape of IDH-wildtype glioblastomas with FGFR3-TACC3 fusions | Despite being older at diagnosis and having similar frequencies of MGMT promoter hypermethylation, patients with F3T3-positive GBMs lived about 8 months longer than those with F3T3 wild-type tumors. Consistent with IDH wild-type GBMs, F3T3-positive GBMs exhibited distinct biological features. |
| Egaña et al., 2020 [15] | Methylation of MGMT promoter does not predict response to temozolomide in patients with glioblastoma in Donostia Hospital | No association was detected between methylation of MGMT promoter and molecular markers such as ATRX, IDH, p53, and Ki67. |
| Schaff et al., 2020 [44] | Characterization of MGMT and EGFR protein expression in glioblastoma and association with survival | A weak association was seen between MGMT protein expression and promoter methylation. Quantification of MGMT protein expression was inferior to MGMT methylation for prognostication in GBM. |
| Cong et al., 2021 [51] | Identification of the Role and Clinical Prognostic Value of Target Genes of m6A RNA Methylation Regulators in Glioma | The study established and validated a seven-gene signature comprising METTL3, COL18A1, NASP, PHLPP2, TIMP1, U2AF2, and VEGFA, with a good capability for predicting glioma survival. These genes were identified to influence 81 anticancer drug responses, which further contributes to the early-phase clinical trials of drug development. |
| Digregorio et al., 2021 [52] | The expression of B7-H3 isoforms in newly diagnosed glioblastoma and recurrence and their functional role | B7-H3 was a marker for SVZ-GBM cells. B7-H3 inhibition in GBM cells reduced their tumorigenicity. Out of the two B7-H3 isoforms, only 2IgB7-H3 was detected in non-cancerous brain tissue, whereas 4IgB7-H3 was specific to GBM. 2IgB7-H3 expression was higher in GBM recurrences and increased resistance to temozolomide-mediated apoptosis. |
| Syafruddin et al., 2021 [37] | Integration of RNA-Seq and proteomics data identifies glioblastoma multiforme surfaceome signature | The results identify six high-confidence GBM genes, HLA-DRA, CD44, SLC1A5, EGFR, ITGB2, PTPRJ, which were significantly upregulated in GBM. High expression of CD44, PTPRJ, and HLA-DRA was significantly associated with poor disease-free survival. |
| Wang et al., 2022 [53] | Identification of Prognostic Biomarkers for Glioblastoma Based on Transcriptome and Proteome Association Analysis | Fibronectin 1(FN1) was a prognostic risk factor and significantly upregulated in GBM samples. FN1 may play a role in GBM progression through ECM-receptor interaction and PI3K-Akt signaling pathways. |
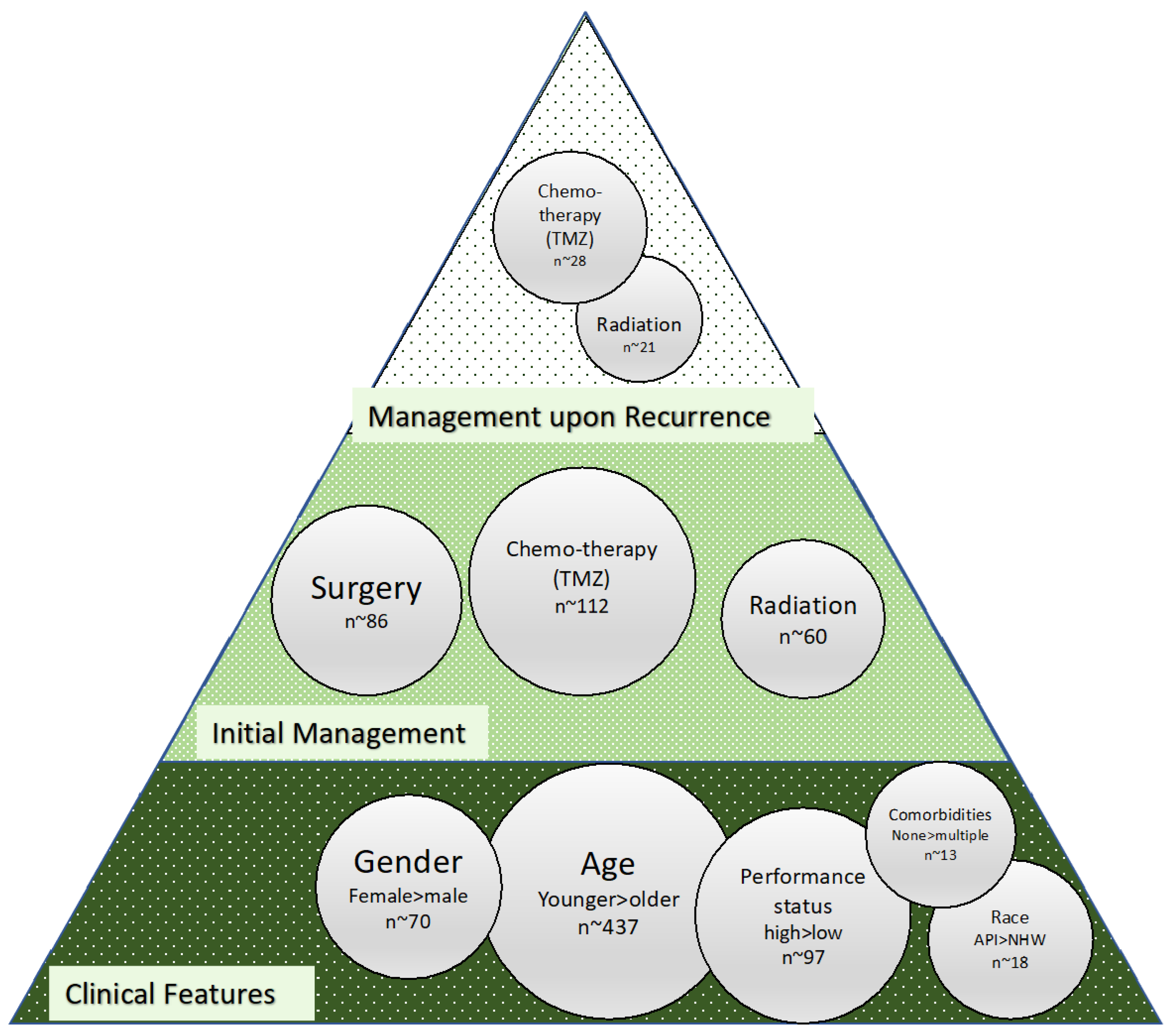
3. Clinical and Management Features of Significance in Glioma
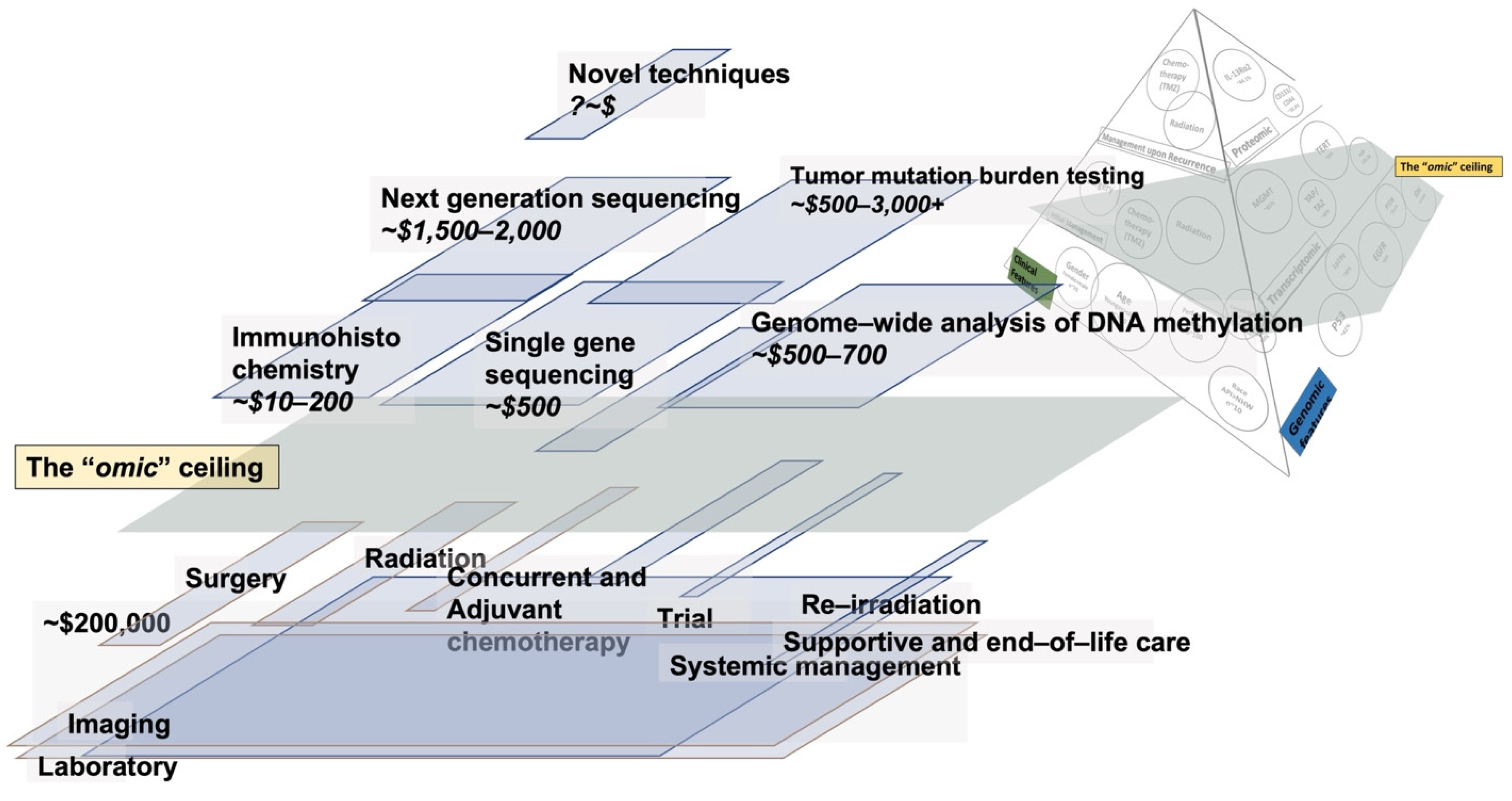
4. The Cost of Care and Omics in Glioma
5. The Cost of Neuropathology and Omics in Glioma
6. Optimizing Cost-Benefit in Glioma to Advance Patient Outcomes
7. Future Directions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| CD44 | Cluster of Differentiation 44 |
| CD133 | Cluster of Differentiation 133 |
| CGGA | Chinese Glioma Genome Atlas |
| CISH | Chromogenic In Situ Hybridization |
| CNS | Central Nervous System |
| EGFR | Epidermal Growth Factor Receptor |
| FISH | Fluorescence In Situ Hybridization |
| GBM | Glioblastoma |
| GSC | Glioma Stem Cells |
| HDMT | Histone Deacetylase |
| ICER | Incremental Cost-Effectiveness Ratio |
| IDH | Isocitrate Dehydrogenase |
| KDMT | Histone Demethylates |
| LGG | Low-Grade Glioma |
| LYG | Life Years Gained |
| MAPK | Mitogen Activated Protein Kinases |
| MGMT | O6-methylguanine-DNA Methyltransferase |
| MLPA | Multiplex Ligation-Dependent Probe Amplification |
| MRI | Magnetic Resonance Imaging |
| NF1 | Neurofibromatosis 1 |
| NGS | Next-Generation Sequencing |
| NHS | National Health Service |
| PDFRA | Platelet-Derived Growth Factors |
| PTEN | Phosphatase and Tensin Homolog |
| P13K-AKT | Phosphoinositide 3-kinasem-Protein Kinase B |
| QALY | Quality-Adjusted Life Years 18-FET-PET |
| RPA | Recursive Partitioning Analysis |
| RT | Radiation therapy |
| SEER | Surveillance, Epidemiology, and End Results |
| TAZ | WW Domain-Containing Transcriptional Regulator 1 |
| TCGA | The Cancer Genome Atlas |
| TERT | Telomerase Reverse Transcriptase |
| TMB | Tumor Mutation Burden Testing |
| TMZ | Temozolomide |
| TP53 | Tumor Protein p53 |
| YAP | Yes-Associated Protein |
| VEGF | Vascular Endothelial Growth Factor |
| 18F-FET PET | O-(2-[18F]fluoroethyl)-L-tyrosine Positron Emission Tomography |
References
- Stupp, R.; Mason, W.P.; van den Bent, M.J.; Weller, M.; Fisher, B.; Taphoorn, M.J.B.; Belanger, K.; Brandes, A.A.; Marosi, C.; Bogdahn, U.; et al. Radiotherapy plus Concomitant and Adjuvant Temozolomide for Glioblastoma. N. Engl. J. Med. 2005, 352, 987–996. [Google Scholar] [CrossRef] [PubMed]
- Hegi, M.E.; Diserens, A.C.; Gorlia, T.; Hamou, M.F.; De Tribolet, N.; Weller, M.; Kros, J.M.; Hainfellner, J.A.; Mason, W.; Mariani, L.; et al. MGMT gene silencing and benefit from temozolomide in glioblastoma. N. Engl. J. Med. 2005, 352, 997–1003. [Google Scholar] [CrossRef] [PubMed]
- Benitez, J.A.; Finlay, D.; Castanza, A.; Parisian, A.D.; Ma, J.; Longobardi, C.; Campos, A.; Vadla, R.; Izurieta, A.; Scerra, G.; et al. PTEN deficiency leads to proteasome addiction: A novel vulnerability in glioblastoma. Neuro-Oncology 2021, 23, 1072–1086. [Google Scholar] [CrossRef]
- Mahlokozera, T.; Patel, B.; Chen, H.; Desouza, P.; Qu, X.; Mao, D.D.; Hafez, D.; Yang, W.; Taiwo, R.; Paturu, M.; et al. Competitive binding of E3 ligases TRIM26 and WWP2 controls SOX2 in glioblastoma. Nat. Commun. 2021, 12, 6321. [Google Scholar] [CrossRef] [PubMed]
- Verhaak, R.G.W.; Hoadley, K.A.; Purdom, E.; Wang, V.; Wilkerson, M.D.; Miller, C.R.; Ding, L.; Golub, T.; Jill, P.; Alexe, G.; et al. Integrated Genomic Analysis Identifies Clinically Relevant Subtypes of Glioblastoma Characterized by Abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 2010, 17, 98–110. [Google Scholar] [CrossRef]
- Aldape, K.; Zadeh, G.; Mansouri, S.; Reifenberger, G.; von Deimling, A. Glioblastoma: Pathology, molecular mechanisms and markers. Acta Neuropathol. 2015, 129, 829–848. [Google Scholar] [CrossRef]
- Akbari, H.; Rathore, S.; Bakas, S.; Nasrallah, M.P.; Shukla, G.; Mamourian, E.; Rozycki, M.; Bagley, S.J.; Rudie, J.D.; Flanders, A.E.; et al. Histopathology-validated machine learning radiographic biomarker for noninvasive discrimination between true progression and pseudo-progression in glioblastoma. Cancer 2020, 126, 2625–2636. [Google Scholar] [CrossRef] [PubMed]
- Booth, T.; Williams, M.; Luis, A.; Cardoso, J.; Ashkan, K.; Shuaib, H. Machine learning and glioma imaging biomarkers. Clin. Radiol. 2020, 75, 20–32. [Google Scholar] [CrossRef]
- Burgenske, D.M.; Yang, J.; A Decker, P.; Kollmeyer, T.M.; Kosel, M.L.; Mladek, A.C.; A Caron, A.; A Vaubel, R.; Gupta, S.K.; Kitange, G.J.; et al. Molecular profiling of long-term IDH-wildtype glioblastoma survivors. Neuro-Oncology 2019, 21, 1458–1469. [Google Scholar] [CrossRef]
- Byrne, N.; Tambe, P.; Coulter, J. Radiation Response in the Tumour Microenvironment: Predictive Biomarkers and Future Perspectives. J. Pers. Med. 2021, 11, 53. [Google Scholar] [CrossRef]
- Davatzikos, C.; Barnholtz-Sloan, J.S.; Bakas, S.; Colen, R.; Mahajan, A.; Quintero, C.B.; Font, J.C.; Puig, J.; Jain, R.; E Sloan, A.; et al. AI-based prognostic imaging biomarkers for precision neuro-oncology: The ReSPOND consortium. Neuro-Oncology 2020, 22, 886–888. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.; Yong, R.L.; Paddison, P.; Zhu, J. Comparison of glioblastoma (GBM) molecular classification methods. In Seminars in Cancer Biology; Academic Press: New York, NY, USA, 2018. [Google Scholar]
- Phillips, H.S.; Kharbanda, S.; Chen, R.; Forrest, W.F.; Soriano, R.H.; Wu, T.D.; Misra, A.; Nigro, J.M.; Colman, H.; Soroceanu, L.; et al. Molecular subclasses of high-grade glioma predict prognosis, delineate a pattern of disease progression, and resemble stages in neurogenesis. Cancer Cell 2006, 9, 157–173. [Google Scholar] [CrossRef] [PubMed]
- Wang, A.; Zhang, G. Differential gene expression analysis in glioblastoma cells and normal human brain cells based on GEO database. Oncol. Lett. 2017, 14, 6040–6044. [Google Scholar] [CrossRef] [PubMed]
- Egaña, L.; Auzmendi-Iriarte, J.; Andermatten, J.; Villanua, J.; Ruiz, I.; Elua-Pinin, A.; Aldaz, P.; Querejeta, A.; Sarasqueta, C.; Zubia, F.; et al. Methylation of MGMT promoter does not predict response to temozolomide in patients with glioblastoma in Donostia Hospital. Sci. Rep. 2020, 10, 18445. [Google Scholar] [CrossRef]
- Kitange, G.J.; Carlson, B.L.; Schroeder, M.A.; Grogan, P.T.; Lamont, J.D.; Decker, P.A.; Wu, W.; James, C.D.; Sarkaria, J.N. Induction of MGMT expression is associated with temozolomide resistance in glioblastoma xenografts. Neuro-Oncology 2009, 11, 281–291. [Google Scholar] [CrossRef]
- Brennan, C.W.; Verhaak, R.G.W.; McKenna, A.; Campos, B.; Noushmehr, H.; Salama, S.R.; Zheng, S.; Chakravarty, D.; Sanborn, J.Z.; Berman, S.H.; et al. The Somatic Genomic Landscape of Glioblastoma. Cell 2013, 155, 462–477. [Google Scholar] [CrossRef]
- Brown, C.E.; Warden, C.D.; Starr, R.; Deng, X.; Badie, B.; Yuan, Y.-C.; Forman, S.J.; Barish, M.E. Glioma IL13Rα2 Is Associated with Mesenchymal Signature Gene Expression and Poor Patient Prognosis. PLoS ONE 2013, 8, e77769. [Google Scholar] [CrossRef]
- Brown, D.V.; Daniel, P.M.; D’Abaco, G.M.; Gogos, A.; Ng, W.; Morokoff, A.P.; Mantamadiotis, T. Coexpression analysis of CD133 and CD44 identifies Proneural and Mesenchymal subtypes of glioblastoma multiforme. Oncotarget 2015, 6, 6267–6280. [Google Scholar] [CrossRef]
- Olympios, N.; Gilard, V.; Marguet, F.; Clatot, F.; Di Fiore, F.; Fontanilles, M. TERT Promoter Alterations in Glioblastoma: A Systematic Review. Cancers 2021, 13, 1147. [Google Scholar] [CrossRef]
- Zhang, Y.; Ta, W.-W.; Sun, P.-F.; Meng, Y.-F.; Zhao, C.-Z. Diagnostic and prognostic significance of serum miR-145-5p expression in glioblastoma. Int. J. Clin. Exp. Pathol. 2019, 12, 2536–2543. [Google Scholar]
- Yao, J.; Hagiwara, A.; Raymond, C.; Shabani, S.; Pope, W.B.; Salamon, N.; Lai, A.; Ji, M.; Nghiemphu, P.L.; Liau, L.M.; et al. Human IDH mutant 1p/19q co-deleted gliomas have low tumor acidity as evidenced by molecular MRI and PET: A retrospective study. Sci. Rep. 2020, 10, 11922. [Google Scholar] [CrossRef] [PubMed]
- The Cancer Genome Atlas Research Network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 2008, 455, 1061–1068. [Google Scholar]
- Cohen, A.L.; Holmen, S.L.; Colman, H. IDH1 and IDH2 Mutations in Gliomas. Curr. Neurol. Neurosci. Rep. 2013, 13, 345. [Google Scholar] [CrossRef] [PubMed]
- Listernick, R.; Charrow, J.; Gutmann, D.H. Intracranial gliomas in neurofibromatosis type 1. Am. J. Med Genet. 1999, 89, 38–44. [Google Scholar] [CrossRef]
- Yan, H.; Parsons, D.W.; Jin, G.; McLendon, R.; Rasheed, B.A.; Yuan, W.; Kos, I.; Batinic-Haberle, I.; Jones, S.; Riggins, G.J.; et al. IDH1 and IDH2 mutations in gliomas. N. Engl. J. Med. 2009, 360, 765–773. [Google Scholar] [CrossRef] [PubMed]
- Asif, S.; Fatima, R.; Krc, R.; Bennett, J.; Raza, S. Comparative proteogenomic characterization of glioblastoma. CNS Oncol. 2019, 8, CNS37. [Google Scholar] [CrossRef] [PubMed]
- Mandel, J.J.; Cachia, D.; Liu, D.; Wilson, C.; Aldape, K.; Fuller, G.; De Groot, J.F. Impact of IDH1 mutation status on outcome in clinical trials for recurrent glioblastoma. J. Neuro-Oncol. 2016, 129, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Stancheva, G.; Goranova, T.; Laleva, M.; Kamenova, M.; Mitkova, A.; Velinov, N.; Poptodorov, G.; Mitev, V.; Kaneva, R.; Gabrovsky, N. IDH1/IDH2 but not TP53 mutations predict prognosis in Bulgarian glioblastoma patients. BioMed Res. Int. 2014, 2014, 654727. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Berezov, A.; Wang, Q.; Zhang, G.; Drebin, J.; Murali, R.; Greene, M.I. ErbB receptors: From oncogenes to targeted cancer therapies. J. Clin. Investig. 2007, 117, 2051–2058. [Google Scholar] [CrossRef] [PubMed]
- Vigneswaran, K.; Boyd, N.H.; Oh, S.-Y.; Lallani, S.; Boucher, A.; Neill, S.G.; Olson, J.J.; Read, R.D. YAP/TAZ Transcriptional Coactivators Create Therapeutic Vulnerability to Verteporfin in EGFR-mutant Glioblastoma. Clin. Cancer Res. 2021, 27, 1553–1569. [Google Scholar] [CrossRef]
- Zhang, Y.; Dube, C.; Gibert, M.; Cruickshanks, N.; Wang, B.; Coughlan, M.; Yang, Y.; Setiady, I.; Deveau, C.; Saoud, K.; et al. The p53 Pathway in Glioblastoma. Cancers 2018, 10, 297. [Google Scholar] [CrossRef]
- Qian, X.; Li, X.; Shi, Z.; Xia, Y.; Cai, Q.; Xu, D.; Tan, L.; Du, L.; Zheng, Y.; Zhao, D.; et al. PTEN Suppresses Glycolysis by Dephosphorylating and Inhibiting Autophosphorylated PGK1. Mol. Cell 2019, 76, 516–527.e7. [Google Scholar] [CrossRef] [PubMed]
- Simon, M.; Hosen, I.; Gousias, K.; Rachakonda, S.; Heidenreich, B.; Gessi, M.; Schramm, J.; Hemminki, K.; Waha, A.; Kumar, R. TERT promoter mutations: A novel independent prognostic factor in primary glioblastomas. Neuro-Oncology 2015, 17, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Labussière, M.; Boisselier, B.; Mokhtari, K.; Di Stefano, A.L.; Rahimian, A.; Rossetto, M.; Ciccarino, P.; Saulnier, O.; Paterra, R.; Marie, Y.; et al. Combined analysis of TERT, EGFR, and IDH status defines distinct prognostic glioblastoma classes. Neurology 2014, 83, 1200–1206. [Google Scholar] [CrossRef] [PubMed]
- Neftel, C.; Laffy, J.; Filbin, M.G.; Hara, T.; Shore, M.E.; Rahme, G.J.; Richman, A.R.; Silverbush, D.; Shaw, M.L.; Hebert, C.M.; et al. An Integrative Model of Cellular States, Plasticity, and Genetics for Glioblastoma. Cell 2019, 178, 835–849.e21. [Google Scholar] [CrossRef]
- Syafruddin, S.E.; Nazarie, W.F.; Moidu, N.A.; Soon, B.H.; Mohtar, M.A. Integration of RNA-Seq and proteomics data identifies glioblastoma multiforme surfaceome signature. BMC Cancer 2021, 21, 850. [Google Scholar] [CrossRef]
- Senbanjo, L.T.; Chellaiah, M.A. CD44: A Multifunctional Cell Surface Adhesion Receptor Is a Regulator of Progression and Metastasis of Cancer Cells. Front. Cell Dev. Biol. 2017, 5, 18. [Google Scholar] [CrossRef]
- Brown, C.E.; Aguilar, B.; Starr, R.; Yang, X.; Chang, W.-C.; Weng, L.; Chang, B.; Sarkissian, A.; Brito, A.; Sanchez, J.F.; et al. Optimization of IL13Rα2-Targeted Chimeric Antigen Receptor T Cells for Improved Anti-tumor Efficacy against Glioblastoma. Mol. Ther. 2018, 26, 31–44. [Google Scholar] [CrossRef]
- Kunadis, E.; Lakiotaki, E.; Korkolopoulou, P.; Piperi, C. Targeting post-translational histone modifying enzymes in glioblastoma. Pharmacol. Ther. 2021, 220, 107721. [Google Scholar] [CrossRef]
- Tasci, E.; Zhuge, Y.; Kaur, H.; Camphausen, K.; Krauze, A.V. Hierarchical Voting-based Feature Selection and Ensemble Learning Model Scheme for Glioma Grading with Clinical and Molecular Characteristics. Submitt. Int. J. Mol. Sci. 2022, 23, 14155. [Google Scholar] [CrossRef]
- Bakas, S.; Sako, C.; Akbari, H.; Bilello, M.; Sotiras, A.; Shukla, G.; Rudie, J.D.; Santamaría, N.F.; Kazerooni, A.F.; Pati, S.; et al. The University of Pennsylvania glioblastoma (UPenn-GBM) cohort: Advanced MRI, clinical, genomics, & radiomics. Sci. Data 2022, 9, 453. [Google Scholar] [PubMed]
- Chen, S.; Xu, Y.; Ye, M.; Li, Y.; Sun, Y.; Liang, J.; Lu, J.; Wang, Z.; Zhu, Z.; Zhang, X.; et al. Predicting MGMT Promoter Methylation in Diffuse Gliomas Using Deep Learning with Radiomics. J. Clin. Med. 2022, 11, 3445. [Google Scholar] [CrossRef] [PubMed]
- Schaff, L.R.; Yan, D.; Thyparambil, S.; Tian, Y.; Cecchi, F.; Rosenblum, M.; Reiner, A.S.; Panageas, K.S.; Hembrough, T.; Lin, A.L. Characterization of MGMT and EGFR protein expression in glioblastoma and association with survival. J. Neuro-Oncol. 2020, 146, 163–170. [Google Scholar] [CrossRef] [PubMed]
- Delfino, K.R.; Serão, N.V.L.; Southey, B.R.; Rodriguez-Zas, S.L. Therapy-, gender- and race-specific microRNA markers, target genes and networks related to glioblastoma recurrence and survival. Cancer Genom.-Proteom. 2011, 8, 173–183. [Google Scholar]
- Håvik, A.B.; Brandal, P.; Honne, H.; Dahlback, H.-S.S.; Scheie, D.; Hektoen, M.; Meling, T.R.; Helseth, E.; Heim, S.; Lothe, R.A.; et al. MGMT promoter methylation in gliomas-assessment by pyrosequencing and quantitative methylation-specific PCR. J. Transl. Med. 2012, 10, 36. [Google Scholar] [CrossRef] [PubMed]
- Tanguturi, S.K.; Trippa, L.; Ramkissoon, S.H.; Pelton, K.; Knoff, D.; Sandak, D.; Lindeman, N.I.; Ligon, A.H.; Beroukhim, R.; Parmigiani, G.; et al. Leveraging molecular datasets for biomarker-based clinical trial design in glioblastoma. Neuro-Oncology 2017, 19, 908–917. [Google Scholar] [CrossRef]
- Gobin, M.; Nazarov, P.V.; Warta, R.; Timmer, M.; Reifenberger, G.; Felsberg, J.; Vallar, L.; Chalmers, A.J.; Herold-Mende, C.C.; Goldbrunner, R.; et al. A DNA Repair and Cell-Cycle Gene Expression Signature in Primary and Recurrent Glioblastoma: Prognostic Value and Clinical Implications. Cancer Res. 2019, 79, 1226–1238. [Google Scholar] [CrossRef]
- Oh, S.; Yeom, J.; Cho, H.J.; Kim, J.-H.; Yoon, S.-J.; Kim, H.; Sa, J.K.; Ju, S.; Lee, H.; Oh, M.J.; et al. Integrated pharmaco-proteogenomics defines two subgroups in isocitrate dehydrogenase wild-type glioblastoma with prognostic and therapeutic opportunities. Nat. Commun. 2020, 11, 3288. [Google Scholar] [CrossRef]
- Mata, D.A.; Benhamida, J.K.; Lin, A.L.; Vanderbilt, C.M.; Yang, S.-R.; Villafania, L.B.; Ferguson, D.C.; Jonsson, P.; Miller, A.M.; Tabar, V.; et al. Genetic and epigenetic landscape of IDH-wildtype glioblastomas with FGFR3-TACC3 fusions. Acta Neuropathol. Commun. 2020, 8, 186. [Google Scholar] [CrossRef]
- Cong, P.; Wu, T.; Huang, X.; Liang, H.; Gao, X.; Tian, L.; Li, W.; Chen, A.; Wan, H.; He, M.; et al. Identification of the Role and Clinical Prognostic Value of Target Genes of m6A RNA Methylation Regulators in Glioma. Front. Cell Dev. Biol. 2021, 9, 709022. [Google Scholar] [CrossRef]
- Digregorio, M.; Coppieters, N.; Lombard, A.; Lumapat, P.N.; Scholtes, F.; Rogister, B. The expression of B7-H3 isoforms in newly diagnosed glioblastoma and recurrence and their functional role. Acta Neuropathol. Commun. 2021, 9, 59. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Yan, S.; Chen, X.; Wang, A.; Han, Z.; Liu, B.; Shen, H. Identification of Prognostic Biomarkers for Glioblastoma Based on Transcriptome and Proteome Association Analysis. Technol. Cancer Res. Treat. 2022, 21, 15330338211035270. [Google Scholar] [CrossRef] [PubMed]
- Jia, Z.; Li, X.; Yan, Y.; Shen, X.; Wang, J.; Yang, H.; Liu, S.; Han, C.; Hu, Y. Exploring the relationship between age and prognosis in glioma: Rethinking current age stratification. BMC Neurol. 2022, 22, 350. [Google Scholar] [CrossRef]
- Lin, Z.; Yang, R.; Li, K.; Yi, G.; Li, Z.; Guo, J.; Zhang, Z.; Junxiang, P.; Liu, Y.; Qi, S.; et al. Establishment of age group classification for risk stratification in glioma patients. BMC Neurol. 2020, 20, 310. [Google Scholar] [CrossRef]
- Curran, W.J., Jr.; Scott, C.B.; Horton, J.R.; Nelson, J.S.; Weinstein, A.S.; Fischbach, A.J.; Chang, C.H.; Rotman, M.; Asbell, S.O.; Krisch, R.E.; et al. Recursive Partitioning Analysis of Prognostic Factors in Three Radiation Therapy Oncology Group Malignant Glioma Trials. J. Natl. Cancer Inst. 1993, 85, 704–710. [Google Scholar] [CrossRef] [PubMed]
- Tavelin, B.; Malmström, A. Sex Differences in Glioblastoma—Findings from the Swedish National Quality Registry for Primary Brain Tumors between 1999–2018. J. Clin. Med. 2022, 11, 486. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.T.; Prajapati, B.; Lakhina, S.; Sharma, M.; Prajapati, S.; Chosdol, K.; Sinha, S. Identification of Gender-Specific Molecular Differences in Glioblastoma (GBM) and Low-Grade Glioma (LGG) by the Analysis of Large Transcriptomic and Epigenomic Datasets. Front. Oncol. 2021, 11, 699594. [Google Scholar] [CrossRef]
- Hodges, T.R.; Labak, C.M.; Mahajan, U.V.; Wright, C.H.; Wright, J.; Cioffi, G.; Gittleman, H.; Herring, E.Z.; Zhou, X.; Duncan, K.; et al. Impact of race on care, readmissions, and survival for patients with glioblastoma: An analysis of the National Cancer Database. Neuro-Oncol. Adv. 2021, 3, vdab040. [Google Scholar] [CrossRef] [PubMed]
- Bohn, A.; Braley, A.; De La Vega, P.R.; Zevallos, J.C.; Barengo, N.C. The association between race and survival in glioblastoma patients in the US: A retrospective cohort study. PLoS ONE 2018, 13, e0198581. [Google Scholar] [CrossRef]
- Ostrom, Q.T.; Cote, D.J.; Ascha, M.; Kruchko, C.; Barnholtz-Sloan, J. Adult Glioma Incidence and Survival by Race or Ethnicity in the United States From 2000 to 2014. JAMA Oncol. 2018, 4, 1254–1262. [Google Scholar] [CrossRef]
- Chen, P.; Aldape, K.; Wiencke, J.K.; Kelsey, K.T.; Miike, R.; Davis, R.L.; Liu, J.; Kesler-Diaz, A.; Takahashi, M.; Wrensch, M. Ethnicity delineates different genetic pathways in malignant glioma. Cancer Res. 2001, 61, 3949–3954. [Google Scholar]
- Wiencke, J.K.; Aldape, K.; McMillan, A.; Wiemels, J.; Moghadassi, M.; Miike, R.; Kelsey, K.T.; Patoka, J.; Long, J.; Wrensch, M. Molecular Features of Adult Glioma Associated with Patient Race/Ethnicity, Age, and a Polymorphism in O6-Methylguanine-DNA-Methyltransferase. Cancer Epidemiol. Biomark. Prev. 2005, 14, 1774–1783. [Google Scholar] [CrossRef] [PubMed]
- Al Feghali, K.A.; Buszek, S.M.; Elhalawani, H.; Chevli, N.; Allen, P.K.; Chung, C. Real-world evaluation of the impact of radiotherapy and chemotherapy in elderly patients with glioblastoma based on age and performance status. Neuro-Oncol. Pr. 2020, 8, 199–208. [Google Scholar] [CrossRef]
- Bell, E.H.; Pugh, S.L.; McElroy, J.P.; Gilbert, M.R.; Mehta, M.; Klimowicz, A.C.; Magliocco, A.; Bredel, M.; Robe, P.; Grosu, A.L.; et al. Molecular-Based Recursive Partitioning Analysis Model for Glioblastoma in the Temozolomide Era: A Correlative Analysis Based on NRG Oncology RTOG 0525. JAMA Oncol. 2017, 3, 784–792. [Google Scholar] [CrossRef] [PubMed]
- Caloglu, M.; Yurut-Caloglu, V.; Karagol, H.; Bayir-Angin, G.; Turan, F.N.; Uzal, C. Prognostic factors other than the performance status and age for glioblastoma multiforme: A single-institution experience. J BUON 2009, 14, 211–218. [Google Scholar] [PubMed]
- Li, L.; Wang, Y.; Li, Y.; Fang, S.; Jiang, T. Role of molecular biomarkers in glioma resection: A systematic review. Chin. Neurosurg. J. 2020, 6, 18. [Google Scholar] [CrossRef] [PubMed]
- Jaroch, K.; Modrakowska, P.; Bojko, B. Glioblastoma Metabolomics—In Vitro Studies. Metabolites 2021, 11, 315. [Google Scholar] [CrossRef]
- Cheng, J.; Liu, J.; Kuang, H.; Wang, J. A Fully Automated Multimodal MRI-Based Multi-Task Learning for Glioma Segmentation and IDH Genotyping. IEEE Trans. Med Imaging 2022, 41, 1520–1532. [Google Scholar] [CrossRef]
- Krauze, A.V.; Zhuge, Y.; Zhao, R.; Tasci, E.; Camphausen, K. AI-Driven Image Analysis in Central Nervous System Tumors-Traditional Machine Learning, Deep Learning and Hybrid Models. J. Biotechnol. Biomed. 2022, 5, 1–19. [Google Scholar]
- Pasquini, L.; Napolitano, A.; Lucignani, M.; Tagliente, E.; Dellepiane, F.; Rossi-Espagnet, M.C.; Ritrovato, M.; Vidiri, A.; Villani, V.; Ranazzi, G.; et al. AI and High-Grade Glioma for Diagnosis and Outcome Prediction: Do All Machine Learning Models Perform Equally Well? Front. Oncol. 2021, 11, 601425. [Google Scholar] [CrossRef]
- Wang, Z.; Wang, Y.; Yang, T.; Xing, H.; Wang, Y.; Gao, L.; Guo, X.; Xing, B.; Wang, Y.; Ma, W. Machine learning revealed stemness features and a novel stemness-based classification with appealing implications in discriminating the prognosis, immunotherapy and temozolomide responses of 906 glioblastoma patients. Briefings Bioinform. 2021, 22, bbab032. [Google Scholar] [CrossRef] [PubMed]
- Jagasia, S. PubMed Literature Search. Available online: https://pubmed.ncbi.nlm.nih.gov/ (accessed on 9 November 2022).
- Latif, A.Z.; Signorini, D.; Gregor, A.; Whittle, I.R. The costs of managing patients with malignant glioma at a neuro-oncology clinic. Br. J. Neurosurg. 1998, 12, 118–122. [Google Scholar] [CrossRef] [PubMed]
- Butenschoen, V.M.; Kelm, A.; Meyer, B.; Krieg, S.M. Quality-adjusted life years in glioma patients: A systematic review on currently available data and the lack of evidence-based utilities. J. Neuro-Oncol. 2019, 144, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Qian, Y.; Maruyama, S.; Kim, H.; Pollom, E.L.; A Kumar, K.; Chin, A.L.; Harris, J.P.; Chang, D.T.; Pitt, A.; Bendavid, E.; et al. Cost-effectiveness of radiation and chemotherapy for high-risk low-grade glioma. Neuro-Oncology 2017, 19, 1651–1660. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Rosen, J.; Ceccon, G.; Bauer, E.K.; Werner, J.M.; Tscherpel, C.; Dunkl, V.; Rapp, M.; Sabel, M.; Herrlinger, U.; Heinzel, A.; et al. Cost-effectiveness of 18F-FET PET for early treatment response assessment in glioma patients following adjuvant temozolomide chemotherapy. J. Nucl. Med. 2022, 63, 1677–1682. [Google Scholar] [CrossRef]
- Jiang, S.; Hill, K.; Patel, D.; Waldeck, A.R.; Botteman, M.; Aly, A.; Norden, A.D. Direct medical costs of treatment in newly-diagnosed high-grade glioma among commercially insured US patients. J. Med Econ. 2017, 20, 1237–1243. [Google Scholar] [CrossRef]
- Louis, D.N.; Perry, A.; Reifenberger, G.; Von Deimling, A.; Figarella-Branger, D.; Cavenee, W.K.; Ohgaki, H.; Wiestler, O.D.; Kleihues, P.; Ellison, D.W. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A summary. Acta Neuropathol. 2016, 131, 803–820. [Google Scholar] [CrossRef]
- Soffietti, R.; Bettegowda, C.; Mellinghoff, I.K.; E Warren, K.; Ahluwalia, M.S.; De Groot, J.F.; Galanis, E.; Gilbert, M.R.; A Jaeckle, K.; Le Rhun, E.; et al. Liquid biopsy in gliomas: A RANO review and proposals for clinical applications. Neuro-Oncology 2022, 24, 855–871. [Google Scholar] [CrossRef]
- Goel, N.J.; Bird, C.E.; Hicks, W.H.; Abdullah, K.G. Economic implications of the modern treatment paradigm of glioblastoma: An analysis of global cost estimates and their utility for cost assessment. J. Med. Econ. 2021, 24, 1018–1024. [Google Scholar] [CrossRef]
- Norden, A.D.; Korytowsky, B.; You, M.; Le, T.K.; Dastani, H.; Bobiak, S.; Singh, P. A Real-World Claims Analysis of Costs and Patterns of Care in Treated Patients with Glioblastoma Multiforme in the United States. J. Manag. Care Spéc. Pharm. 2019, 25, 428–436. [Google Scholar] [CrossRef]
- Mansouri, A.; Hachem, L.D.; Mansouri, S.; Nassiri, F.; Laperriere, N.J.; Xia, D.; I Lindeman, N.; Wen, P.Y.; Chakravarti, A.; Mehta, M.P.; et al. MGMT promoter methylation status testing to guide therapy for glioblastoma: Refining the approach based on emerging evidence and current challenges. Neuro-Oncology 2019, 21, 167–178. [Google Scholar] [CrossRef] [PubMed]
- McAleenan, A.; E Jones, H.; Kernohan, A.; Faulkner, C.L.; Palmer, A.; Dawson, S.; Wragg, C.; Jefferies, S.; Brandner, S.; Vale, L.; et al. Diagnostic test accuracy and cost-effectiveness of tests for codeletion of chromosomal arms 1p and 19q in people with glioma. Cochrane Database Syst. Rev. 2022, 3, CD013387. [Google Scholar] [CrossRef] [PubMed]
- DeWitt, J.C.; Jordan, J.T.; Frosch, M.P.; Samore, W.R.; Iafrate, A.J.; Louis, D.N.; Lennerz, J.K. Cost-effectiveness of IDH testing in diffuse gliomas according to the 2016 WHO classification of tumors of the central nervous system recommendations. Neuro-Oncology 2017, 19, 1640–1650. [Google Scholar] [CrossRef]
- Desai, K.; Hooker, G.; Gilbert, K.; Cropper, C.; Metcalf, R.; Kachroo, S. Real-world trends in costs of next generation sequencing (NGS) testing in U.S. setting. J. Clin. Oncol. 2021, 39 (Suppl. 15), e18824. [Google Scholar] [CrossRef]
- Hsiao, S.J.; Sireci, A.N.; Pendrick, D.; Freeman, C.; Fernandes, H.; Schwartz, G.K.; Henick, B.S.; Mansukhani, M.M.; Roth, K.A.; Carvajal, R.D.; et al. Clinical Utilization, Utility, and Reimbursement for Expanded Genomic Panel Testing in Adult Oncology. JCO Precis. Oncol. 2020, 4, 1038–1048. [Google Scholar] [CrossRef]
- Zhang, Y.; Lucas, C.H.; Young, J.S.; Morshed, R.A.; McCoy, L.; Oberheim Bush, N.A.; Taylor, J.W.; Daras, M.; Butowski, N.A.; Villanueva-Meyer, J.E.; et al. Prospective genomically guided identification of “early/evolving” and “undersampled” IDH-wildtype glioblastoma leads to improved clinical outcomes. Neuro-Oncology 2022, 24, 1749–1762. [Google Scholar] [CrossRef]
- Dinnes, J.; Cave, C.; Huang, S.; Major, K.; Milne, R. The effectiveness and cost-effectiveness of temozolomide for the treatment of recurrent malignant glioma: A rapid and systematic review. Heal. Technol. Assess. 2001, 5, 1–73. [Google Scholar] [CrossRef]
- Haider, S.A.; Asmaro, K.; Kalkanis, S.N.; Lee, I.Y.; Bazydlo, M.; Nerenz, D.R.; Salloum, R.G.; Snyder, J.; Walbert, T. The economic impact of glioma survivorship: The cost of care from a patient perspective. Neurology 2020, 95, e1575–e1581. [Google Scholar] [CrossRef]
- Konski, A.; Bracy, P.; Weiss, S.; Grigsby, P. Cost-utility analysis of a malignant glioma protocol. Int. J. Radiat. Oncol. 1997, 39, 575–578. [Google Scholar] [CrossRef]
- Pendharkar, A.V.; Rezaii, P.G.; Ho, A.L.; Sussman, E.S.; Li, G.; Desai, A.M. Functional Mapping for Glioma Surgery: A Propensity-Matched Analysis of Outcomes and Cost. World Neurosurg. 2020, 137, e328–e335. [Google Scholar] [CrossRef]
- Jiang, W.; Jones, J.C.; Shankavaram, U.; Sproull, M.; Camphausen, K.; Krauze, A.V. Analytical Considerations of Large-Scale Aptamer-Based Datasets for Translational Applications. Cancers 2022, 14, 2227. [Google Scholar] [CrossRef] [PubMed]
- Tasci, E.; Zhuge, Y.; Camphausen, K.; Krauze, A.V. Bias and Class Imbalance in Oncologic Data—Towards Inclusive and Transferrable AI in Large Scale Oncology Data Sets. Cancers 2022, 14, 2897. [Google Scholar] [CrossRef] [PubMed]
- Macyszyn, L.; Akbari, H.; Pisapia, J.M.; Da, X.; Attiah, M.; Pigrish, V.; Bi, Y.; Pal, S.; Davuluri, R.V.; Roccograndi, L.; et al. Imaging patterns predict patient survival and molecular subtype in glioblastoma via machine learning techniques. Neuro-Oncology 2016, 18, 417–425. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.; Mu, W.; Wang, X.-C.; Yang, G.-Q.; Gillies, R.J.; Zhang, H. Improving survival prediction of high-grade glioma via machine learning techniques based on MRI radiomic, genetic and clinical risk factors. Eur. J. Radiol. 2019, 120, 108609. [Google Scholar] [CrossRef]
- Zanella, L.; Facco, P.; Bezzo, F.; Cimetta, E. Feature Selection and Molecular Classification of Cancer Phenotypes: A Comparative Study. Int. J. Mol. Sci. 2022, 23, 9087. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jagasia, S.; Tasci, E.; Zhuge, Y.; Camphausen, K.; Krauze, A.V. Cost Matrix of Molecular Pathology in Glioma—Towards AI-Driven Rational Molecular Testing and Precision Care for the Future. Biomedicines 2022, 10, 3029. https://doi.org/10.3390/biomedicines10123029
Jagasia S, Tasci E, Zhuge Y, Camphausen K, Krauze AV. Cost Matrix of Molecular Pathology in Glioma—Towards AI-Driven Rational Molecular Testing and Precision Care for the Future. Biomedicines. 2022; 10(12):3029. https://doi.org/10.3390/biomedicines10123029
Chicago/Turabian StyleJagasia, Sarisha, Erdal Tasci, Ying Zhuge, Kevin Camphausen, and Andra Valentina Krauze. 2022. "Cost Matrix of Molecular Pathology in Glioma—Towards AI-Driven Rational Molecular Testing and Precision Care for the Future" Biomedicines 10, no. 12: 3029. https://doi.org/10.3390/biomedicines10123029
APA StyleJagasia, S., Tasci, E., Zhuge, Y., Camphausen, K., & Krauze, A. V. (2022). Cost Matrix of Molecular Pathology in Glioma—Towards AI-Driven Rational Molecular Testing and Precision Care for the Future. Biomedicines, 10(12), 3029. https://doi.org/10.3390/biomedicines10123029