Identification of Druggable Genes for Asthma by Integrated Genomic Network Analysis
Abstract
1. Introduction
2. Materials and Methods
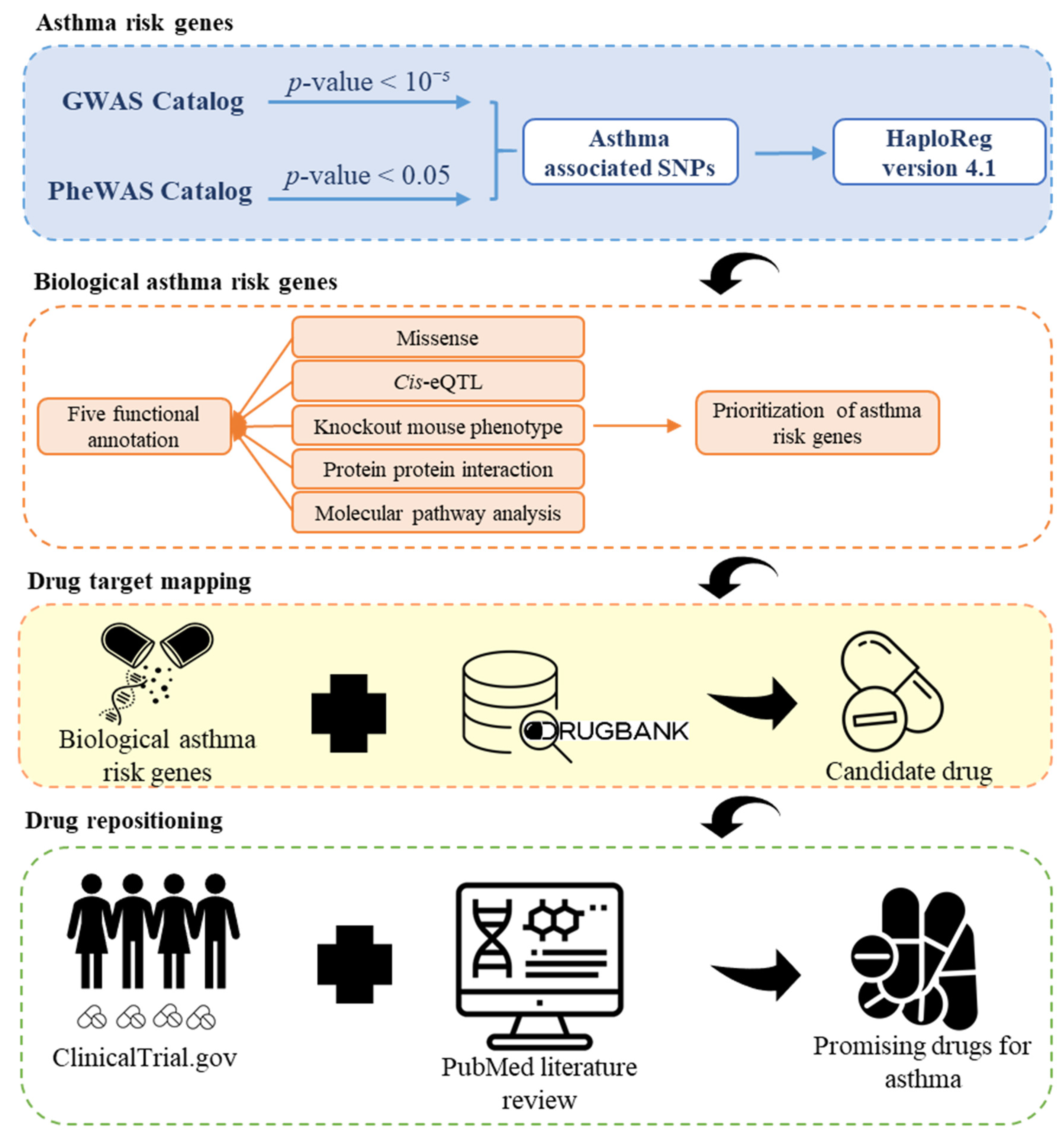
2.1. Study Design
2.2. Biological Asthma Risk Genes
2.3. Drug Mining and Prioritization
2.4. Statistical Analysis
3. Results
3.1. Identification of Asthma Associated SNP
3.2. Gene-Based Prioritization from Functional Annotation
3.3. Integrative Analysis for Drug Repositioning
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Holgate, S.T.; Wenzel, S.; Postma, D.S.; Weiss, S.T.; Renz, H.; Sly, P.D. Asthma. Nat. Rev. Dis. Prim. 2015, 1, 57–70. [Google Scholar] [CrossRef]
- Tarraf, H.; Aydin, O.; Mungan, D.; Albader, M.; Mahboub, B.; Doble, A.; Lahlou, A.; Tariq, L.; Aziz, F.; El Hasnaoui, A. Prevalence of asthma among the adult general population of five Middle Eastern countries: Results of the SNAPSHOT program. BMC Pulm. Med. 2018, 18, 68. [Google Scholar] [CrossRef] [PubMed]
- Global Initiative for Asthma. Global Strategy for Asthma Management and Prevention. 2021. Available online: https://ginasthma.org/ (accessed on 15 December 2021).
- Global Asthma Network. The Global Asthma Report 2018; Global Asthma Network: Auckland, New Zealand, 2018; ISBN 9780473465230. [Google Scholar]
- Huo, Y.; Zhang, H.Y. Genetic mechanisms of asthma and the implications for drug repositioning. Genes 2018, 9, 237. [Google Scholar] [CrossRef]
- Miller, S.M.; Ortega, V.E. Pharmacogenetics and the development of personalized approaches for combination therapy in asthma. Curr. Allergy Asthma Rep. 2013, 13, 443–452. [Google Scholar] [CrossRef]
- Slager, R.E.; Hawkins, G.A.; Li, X.; Postma, D.S.; Meyers, D.A.; Bleecker, E.R. Genetics of Asthma Susceptibility and Severity. Clin. Chest Med. 2012, 33, 431–443. [Google Scholar] [CrossRef]
- Moore, W.C.; Meyers, D.A.; Wenzel, S.E.; Teague, W.G.; Li, H.; Li, X.; D’Agostino, R., Jr.; Castro, M.; Curran-Everett, D.; Fitzpatrick, A.M.; et al. Identification of asthma phenotypes using cluster analysis in the severe asthma research program. Am. J. Respir. Crit. Care Med. 2010, 181, 315–323. [Google Scholar] [CrossRef]
- Barnes, P.J. Drugs for asthma. Br. J. Pharmacol. 2006, 147, 297–303. [Google Scholar] [CrossRef]
- Green, R. New therapies and management strategies in the treatment of asthma: Patient-focused developments. J. Asthma Allergy 2010, 4, 1. [Google Scholar] [CrossRef][Green Version]
- Ashburn, T.T.; Thor, K.B. Drug repositioning: Identifying and developing new uses for existing drugs. Nat. Rev. Drug Discov. 2004, 3, 673–683. [Google Scholar] [CrossRef] [PubMed]
- Jin, G.; Wong, S.T.C. Toward better drug repositioning: Prioritizing and integrating existing methods into efficient pipelines. Drug Discov. Today 2014, 19, 637–644. [Google Scholar] [CrossRef] [PubMed]
- Mullard, A. Drug repurposing programmes get lift off. Nat. Rev. Drug Discov. 2012, 11, 505–506. [Google Scholar] [CrossRef]
- Shoaib, M.; Kamal, M.A.; Danish Rizvi, S.M. Repurposed Drugs as Potential Therapeutic Candidates for the Management of Alzheimer’s Disease. Curr. Drug Metab. 2017, 18, 842–852. [Google Scholar] [CrossRef]
- Sertkaya, A.; Birkenbach, A.; Berlind, A.; Euraud, J. Examination of clinical trials costs and barriers for drug development. In US Department of Health and Human Services, Office of the Assistant Secretary for Planning and Evaluation Report; U.S. Department of Health and Human Services: Washington, DC, USA, 2014; pp. 1–92. [Google Scholar]
- Turanli, B.; Grøtli, M.; Boren, J.; Nielsen, J.; Uhlen, M.; Arga, K.Y.; Mardinoglu, A. Drug repositioning for effective prostate cancer treatment. Front. Physiol. 2018, 9, 500. [Google Scholar] [CrossRef] [PubMed]
- Buniello, A.; Macarthur, J.A.L.; Cerezo, M.; Harris, L.W.; Hayhurst, J.; Malangone, C.; McMahon, A.; Morales, J.; Mountjoy, E.; Sollis, E.; et al. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 2019, 47, D1005–D1012. [Google Scholar] [CrossRef] [PubMed]
- Denny, J.C.; Bastarache, L.; Ritchie, M.D.; Carroll, R.J.; Zink, R.; Mosley, J.D.; Field, J.R.; Pulley, J.M.; Ramirez, A.H.; Bowton, E.; et al. Systematic comparison of phenome-wide association study of electronic medical record data and genome-wide association study data. Nat. Biotechnol. 2013, 31, 1102–1110. [Google Scholar] [CrossRef] [PubMed]
- Lau, A.; So, H.C. Turning genome-wide association study findings into opportunities for drug repositioning. Comput. Struct. Biotechnol. J. 2020, 18, 1639–1650. [Google Scholar] [CrossRef] [PubMed]
- Ward, L.D.; Kellis, M. HaploReg v4: Systematic mining of putative causal variants, cell types, regulators and target genes for human complex traits and disease. Nucleic Acids Res. 2016, 44, D877–D881. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Feunang, Y.D.; Guo, A.C.; Lo, E.J.; Marcu, A.; Grant, J.R.; Sajed, T.; Johnson, D.; Li, C.; Sayeeda, Z.; et al. DrugBank 5.0: A major update to the DrugBank database for 2018. Nucleic Acids Res. 2018, 46, D1074–D1082. [Google Scholar] [CrossRef] [PubMed]
- Okada, Y.; Wu, D.; Trynka, G.; Raj, T.; Terao, C.; Ikari, K.; Kochi, Y.; Ohmura, K.; Suzuki, A.; Yoshida, S.; et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 2013, 506, 376–381. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Wang, J.; Jaehnig, E.J.; Shi, Z.; Zhang, B. WebGestalt 2019: Gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res. 2019, 47, W199–W205. [Google Scholar] [CrossRef] [PubMed]
- Zhbannikov, I.Y.; Arbeev, K.; Yashin, A.I. Package ‘haploR’. Available online: https://cran.r-project.org/web/packages/haploR/index.html (accessed on 10 September 2021).
- Wang, J.; Liao, Y.; Jaehnig, E.; Shi, Z.; Sheng, Q. Package ‘WebGestaltR’. Available online: https://cran.r-project.org/web/packages/WebGestaltR/index.html (accessed on 10 September 2021).
- Jevnikar, Z.; Östling, J.; Ax, E.; Calvén, J.; Thörn, K.; Israelsson, E.; Öberg, L.; Singhania, A.; Lau, L.C.K.; Wilson, S.J.; et al. Epithelial IL-6 trans-signaling defines a new asthma phenotype with increased airway inflammation. J. Allergy Clin. Immunol. 2010, 143, 577–590. [Google Scholar] [CrossRef]
- Akira, S. IL-6-regulated transcription factors. Int. J. Biochem. Cell Biol. 1997, 29, 1401–1418. [Google Scholar] [CrossRef]
- Akira, S.; Nishio, Y.; Inoue, M.; Wang, X.J.; We, S.; Matsusaka, T.; Yoshida, K.; Sudo, T.; Naruto, M.; Kishimoto, T. Molecular cloning of APRF, a novel IFN-stimulated gene factor 3 p91-related transcription factor involved in the gp130-mediated signaling pathway. Cell 1994, 77, 63–71. [Google Scholar] [CrossRef]
- Akira, S.; Kishimoto, T. IL-6 and NF-IL6 in Acute-Phase Response and Viral Infection. Immunol. Rev. 1992, 127, 25–50. [Google Scholar] [CrossRef]
- Pelaia, G.; Gallelli, L.; D’Agostino, B.; Vatrella, A.; Cuda, G.; Fratto, D.; Renda, T.; Galderisi, U.; Piegari, E.; Crimi, N.; et al. Effects of TGF-β and Glucocorticoids on Map Kinase Phosphorylation, IL-6/IL-11 Secretion and Cell Proliferation in Primary Cultures of Human Lung Fibroblasts. J. Cell. Physiol. 2006, 211, 736–747. [Google Scholar] [CrossRef]
- Rincon, M.; Irvin, C.G. Role of IL-6 in asthma and other inflammatory pulmonary diseases. Int. J. Biol. Sci. 2012, 8, 1281–1290. [Google Scholar] [CrossRef] [PubMed]
- Ullah, M.A.; Revez, J.A.; Loh, Z.; Simpson, J.; Zhang, V.; Bain, L.; Varelias, A.; Rose-John, S.; Blumenthal, A.; Smyth, M.J.; et al. Allergen-induced IL-6 trans-signaling activates γδ T cells to promote type 2 and type 17 airway inflammation. J. Allergy Clin. Immunol. 2015, 136, 1065–1073. [Google Scholar] [CrossRef]
- Revez, J.A.; Bain, L.M.; Watson, R.M.; Towers, M.; Collins, T.; Killian, K.J.; O’Byrne, P.M.; Gauvreau, G.M.; Upham, J.W.; Ferreira, M.A.R. Effects of interleukin-6 receptor blockade on allergen-induced airway responses in mild asthmatics. Clin. Transl. Immunol. 2019, 8, e1044. [Google Scholar] [CrossRef] [PubMed]
- Falcone, D.; Gallelli, L.; Di Virgilio, A.; Tucci, L.; Scaramuzzino, M.; Terracciano, R.; Pelaia, G.; Savino, R. Effects of simvastatin and rosuvastatin on RAS protein, matrix metalloproteinases and NF-κB in lung cancer and in normal pulmonary tissues. Cell Prolif. 2013, 46, 172–182. [Google Scholar] [CrossRef]
- Zeki, A.A.; Elbadawi-Sidhu, M. Innovations in asthma therapy: Is there a role for inhaled statins? Expert Rev. Respir. Med. 2018, 12, 461–473. [Google Scholar] [CrossRef] [PubMed]
- Zeki, A.A.; Franzi, L.; Last, J.; Kenyon, N.J. Simvastatin inhibits airway hyperreactivity: Implications for the mevalonate pathway and beyond. Am. J. Respir. Crit. Care Med. 2009, 180, 731–740. [Google Scholar] [CrossRef]
- Alexeeff, S.E.; Litonjua, A.A.; Sparrow, D.; Vokonas, P.S.; Schwartz, J. Statin use reduces decline in lung function: VA Normative Aging Study. Am. J. Respir. Crit. Care Med. 2007, 176, 742–747. [Google Scholar] [CrossRef]
- Gao, S.; Wang, C.; Li, W.; Shu, S.; Zhou, J.; Yuan, Z.; Wang, L. Allergic asthma aggravated atherosclerosis increases cholesterol biosynthesis and foam cell formation in apolipoprotein E-deficient mice. Biochem. Biophys. Res. Commun. 2019, 519, 861–867. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Yang, R.; Wang, G. Anti-asthmatic effect of pitavastatin through aerosol inhalation is associated with CD4+ CD25+ Foxp3+ T cells in an asthma mouse model. Sci. Rep. 2017, 7, 6084. [Google Scholar] [CrossRef] [PubMed]
- Imamura, M.; Okunishi, K.; Ohtsu, H.; Nakagome, K.; Harada, H.; Tanaka, R.; Yamamoto, K.; Dohi, M. Pravastatin attenuates allergic airway inflammation by suppressing antigen sensitisation, interleukin 17 production and antigen presentation in the lung. Thorax 2009, 64, 44–49. [Google Scholar] [CrossRef] [PubMed]
- Liou, C.J.; Cheng, P.Y.; Huang, W.C.; Chan, C.C.; Chen, M.C.; Kuo, M.L.; Shen, J.J. Oral lovastatin attenuates airway inflammation and mucus secretion in ovalbumin-induced murine model of asthma. Allergy Asthma Immunol. Res. 2014, 6, 548–557. [Google Scholar] [CrossRef] [PubMed]
- Samson, K.T.R.; Minoguchi, K.; Tanaka, A.; Oda, N.; Yokoe, T.; Yamamoto, Y.; Yamamoto, M.; Ohta, S.; Adachi, M. Inhibitory effects of fluvastatin on cytokine and chemokine production by peripheral blood mononuclear cells in patients with allergic asthma. Clin. Exp. Allergy 2006, 36, 475–482. [Google Scholar] [CrossRef]
- Wang, W.; Le, W.; Ahuja, R.; Cho, D.; Hwang, P.H.; Upadhyay, D. Inhibition of Inflammatory Mediators: Role of Statins in Airway Inflammation. Otolaryngol. Head Neck Surg. 2011, 144, 982–987. [Google Scholar] [CrossRef]
- Liu, J.; Suh, D.; Yang, E.; Lee, S.; Park, H.; Shin, Y.S. Attenuation of airway inflammation by simvastatin and the implications for asthma treatment: Is the jury still out? Exp. Mol. Med. 2014, 46, e113. [Google Scholar] [CrossRef]
- Polosa, R.; Holgate, S.T. Adenosine Receptors as Promising Therapeutic Targets for Drug Development in Chronic Airway Inflammation. Curr. Drug Targets 2006, 7, 699–706. [Google Scholar] [CrossRef]
- Brown, R.A.; Spina, D.; Page, C.P. Adenosine receptors and asthma. Br. J. Pharmacol. 2008, 153, 446–456. [Google Scholar] [CrossRef] [PubMed]
- Keane-Myers, A.M.; Gause, W.C.; Finkelman, F.D.; Xhou, X.D.; Wills-Karp, M. Development of murine allergic asthma is dependent upon B7-2 costimulation. J. Immunol. 1998, 160, 1036–1043. [Google Scholar] [PubMed]
- Haczku, A.; Takeda, K.; Redai, I.; Hamelmann, E.; Cieslewicz, G.; Joetham, A.; Loader, J.; Lee, J.J.; Irvin, C.; Gelfand, E.W.; et al. Airway Hyperresponsiveness in Mice. Am. J. Respir. Crit. Care Med. 1999, 86, 1638–1643. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.-N.; Tseng, H.-I.; Kao, C.-C.; Chu, Y.-T.; Chen, W.-Y.; Wu, P.-F.; Lee, C.-H.; Ko, Y.-C. The effects of NOS1 gene on asthma and total IgE levels in Taiwanese children, and the interactions with environmental factors. Pediatr. Allergy Immunol. 2010, 21, 1064–1071. [Google Scholar] [CrossRef]
- Schmalbach, T.; Fuhr, R.; Albayaty, M.; Allen, K.; Douglas, M.; Dunbar, J.; McLaughlin, J.; Alexander, L.; McKee, C. Duvelisib, a PI3K-δ,γ inhibitor, in subjects with mild asthma. Eur. Respir. J. 2015, 46, PA2122. [Google Scholar]
- Mishra, G. Serious adverse effects of anticancer drugs—A review. Ideal Res. 2018, 3. [Google Scholar]
- Fetro, C.; Scherman, D. Drug repurposing in rare diseases: Myths and reality. Therapies 2020, 75, 157–160. [Google Scholar] [CrossRef]
- Brightling, C.E.; Nair, P.; Cousins, D.J.; Louis, R.; Singh, D. Risankizumab in Severe Asthma—A Phase 2a, Placebo-Controlled Trial. N. Engl. J. Med. 2021, 385, 1669–1679. [Google Scholar] [CrossRef]



| Drug Candidate | Gene Target | Drug Action | Current Drug Indication | Phase of Development | N.C.T. Number/PubMed ID |
|---|---|---|---|---|---|
| Gabapentin | ADORA1 | Agonist | Postherpetic neuralgia | Phase IV | NCT00153283 |
| Lamotrigine | ADORA1 | Inhibitor | Epilepsy | Phase IV | NCT00153244 |
| Simvastatin | HMGCR | Inhibitor | Hypercholesterolemia | Phase III | NCT01266434 |
| Ketamine | NOS1 | Inhibitor | General anaesthesia | Phase III | NCT03338205 |
| Atorvastatin | HMGCR | Inhibitor | Hypercholesterolemia | Phases II/III | NCT00126048 |
| Imatinib | BCR | Inhibitor | Chronic myelogenous leukaemia | Phase II | NCT01097694 |
| Abatacept | CD86 | Antagonist | Rheumatoid arthritis | Phase II | NCT00784459 |
| Duvelisib | PIK3CB | Inhibitor | Small lymphocytic lymphoma | Phase II | NCT01653756 |
| Adenosine | ADORA1 | Agonist | Tachycardia | Phase II | NCT01006655 |
| Rosuvastatin | HMGCR | Inhibitor | Hypercholesterolemia | Phase I | NCT01411111 |
| Tocilizumab | IL6R | Inhibitor | Rheumatoid arthritis | Phases I/II | ACTRN12614000123640, 25930193, 30885880 |
| Caffein | ADORA1 | Inhibitor | Apnea of prematurity | NA | NCT01057875 |
| Pentoxifylline * | ADORA1 | Intermittent claudication | - | 19905913 | |
| Pitavastatin * | HMGCR | Inhibitor | Hypercholesterolemia | - | 28729731 |
| Pravastatin * | HMGCR | Inhibitor | Hypercholesterolemia | - | 18835962 |
| Lovastatin * | HMGCR | Inhibitor | Hypercholesterolemia | - | 25374755 |
| Fluvastatin * | HMGCR | Inhibitor | Hypercholesterolemia | - | 16630152 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Adikusuma, W.; Chou, W.-H.; Lin, M.-R.; Ting, J.; Irham, L.M.; Perwitasari, D.A.; Chang, W.-P.; Chang, W.-C. Identification of Druggable Genes for Asthma by Integrated Genomic Network Analysis. Biomedicines 2022, 10, 113. https://doi.org/10.3390/biomedicines10010113
Adikusuma W, Chou W-H, Lin M-R, Ting J, Irham LM, Perwitasari DA, Chang W-P, Chang W-C. Identification of Druggable Genes for Asthma by Integrated Genomic Network Analysis. Biomedicines. 2022; 10(1):113. https://doi.org/10.3390/biomedicines10010113
Chicago/Turabian StyleAdikusuma, Wirawan, Wan-Hsuan Chou, Min-Rou Lin, Jafit Ting, Lalu Muhammad Irham, Dyah Aryani Perwitasari, Wei-Pin Chang, and Wei-Chiao Chang. 2022. "Identification of Druggable Genes for Asthma by Integrated Genomic Network Analysis" Biomedicines 10, no. 1: 113. https://doi.org/10.3390/biomedicines10010113
APA StyleAdikusuma, W., Chou, W.-H., Lin, M.-R., Ting, J., Irham, L. M., Perwitasari, D. A., Chang, W.-P., & Chang, W.-C. (2022). Identification of Druggable Genes for Asthma by Integrated Genomic Network Analysis. Biomedicines, 10(1), 113. https://doi.org/10.3390/biomedicines10010113










