Contribution of Nanomaterials to the Development of Electrochemical Aptasensors for the Detection of Antimicrobial Residues in Food Products
Abstract
1. Introduction
2. Electrochemical Aptasensors for the Detection of Antimicrobial Residues
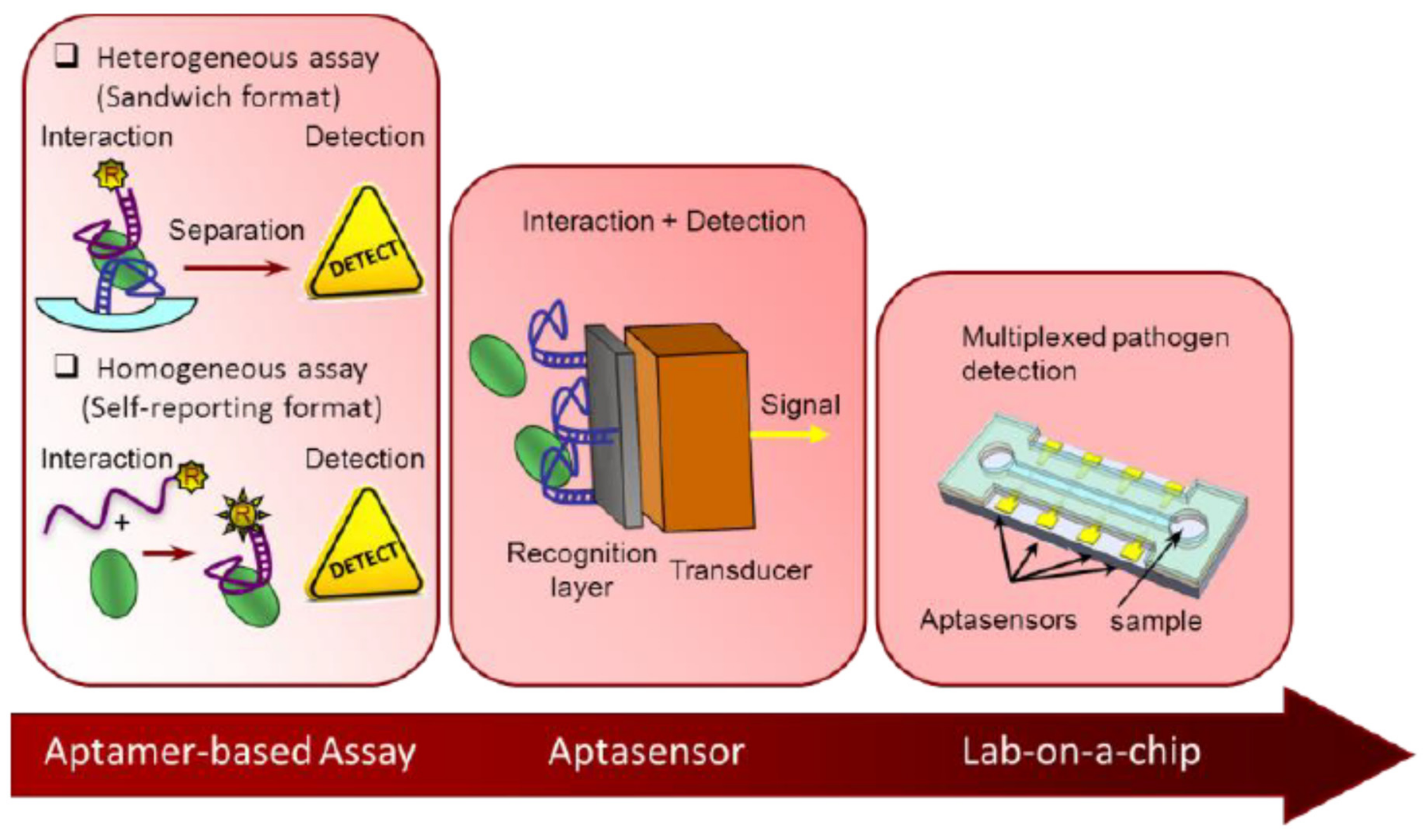
2.1. Different Strategies of Development of Aptasensors
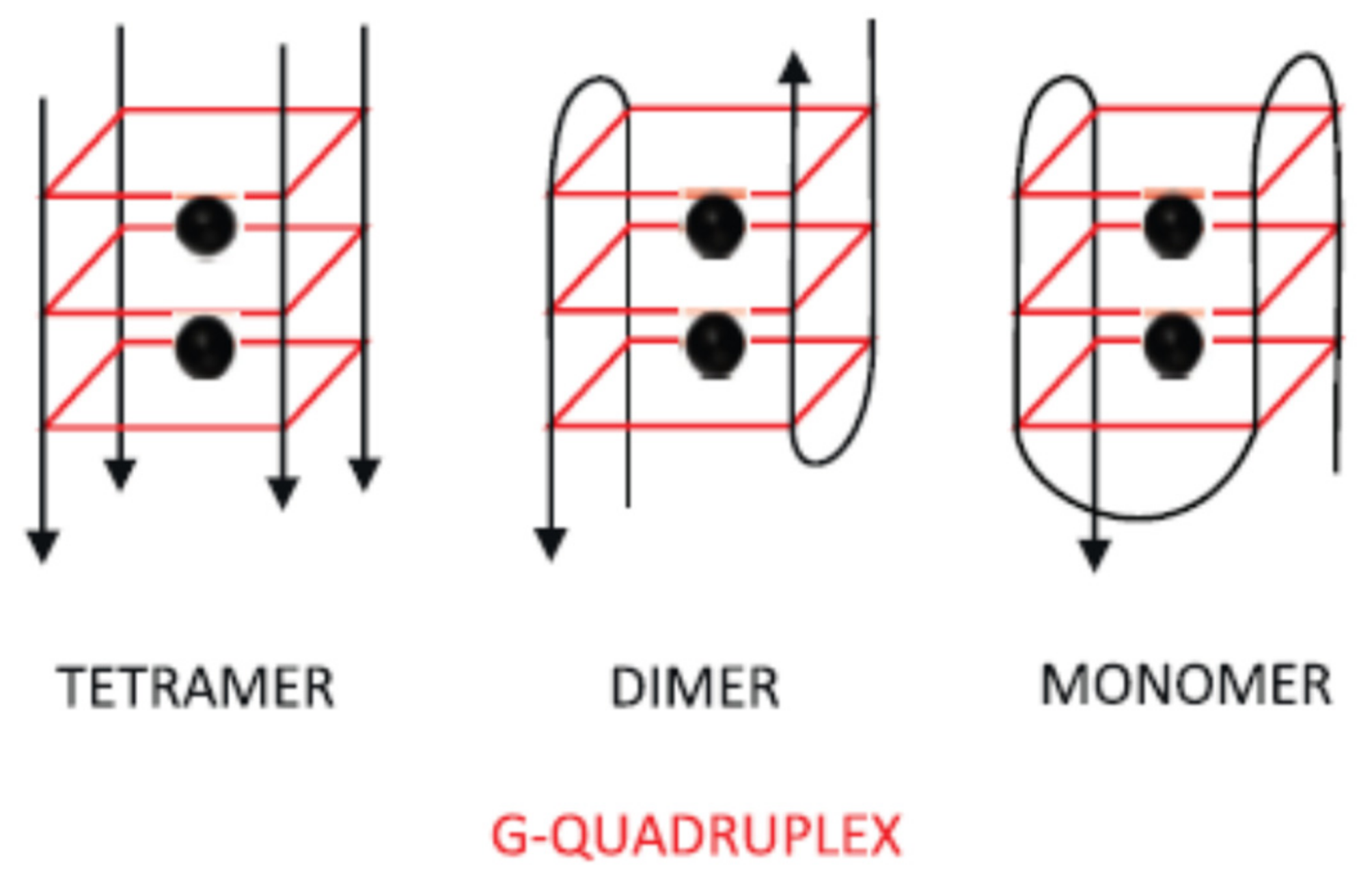
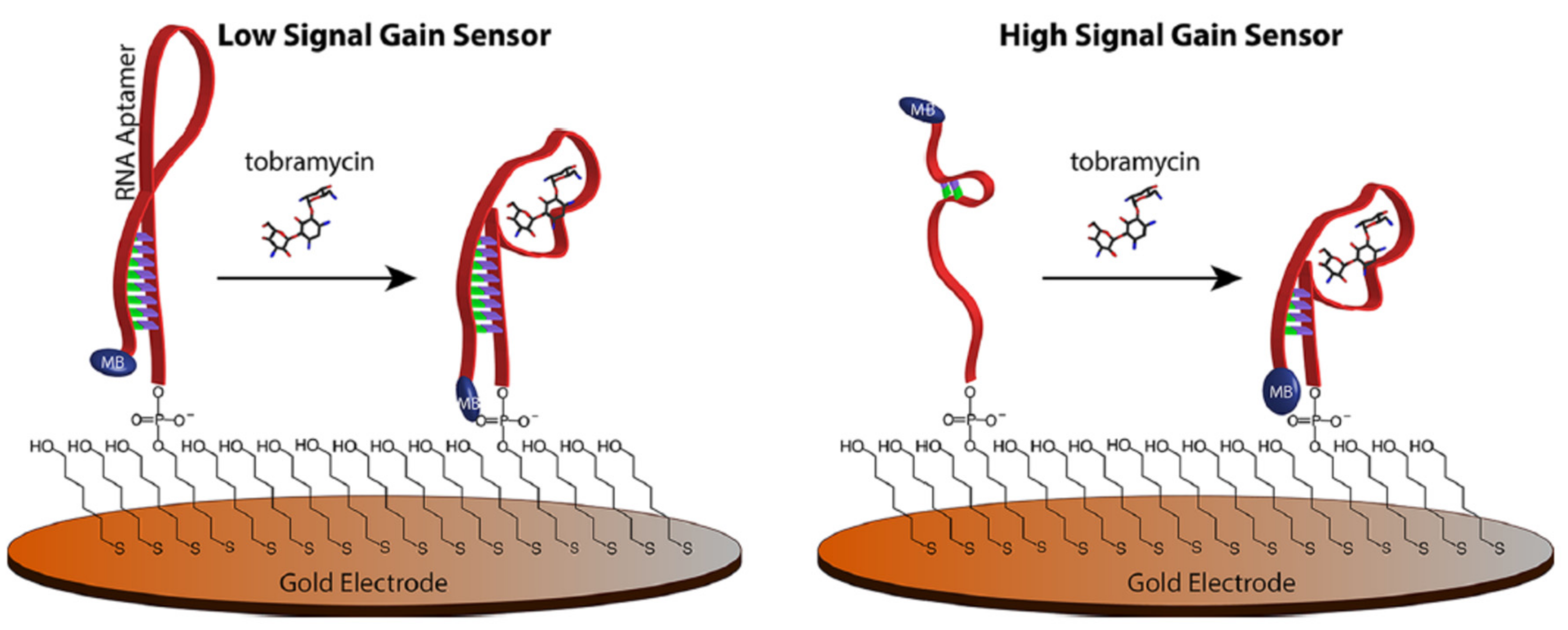
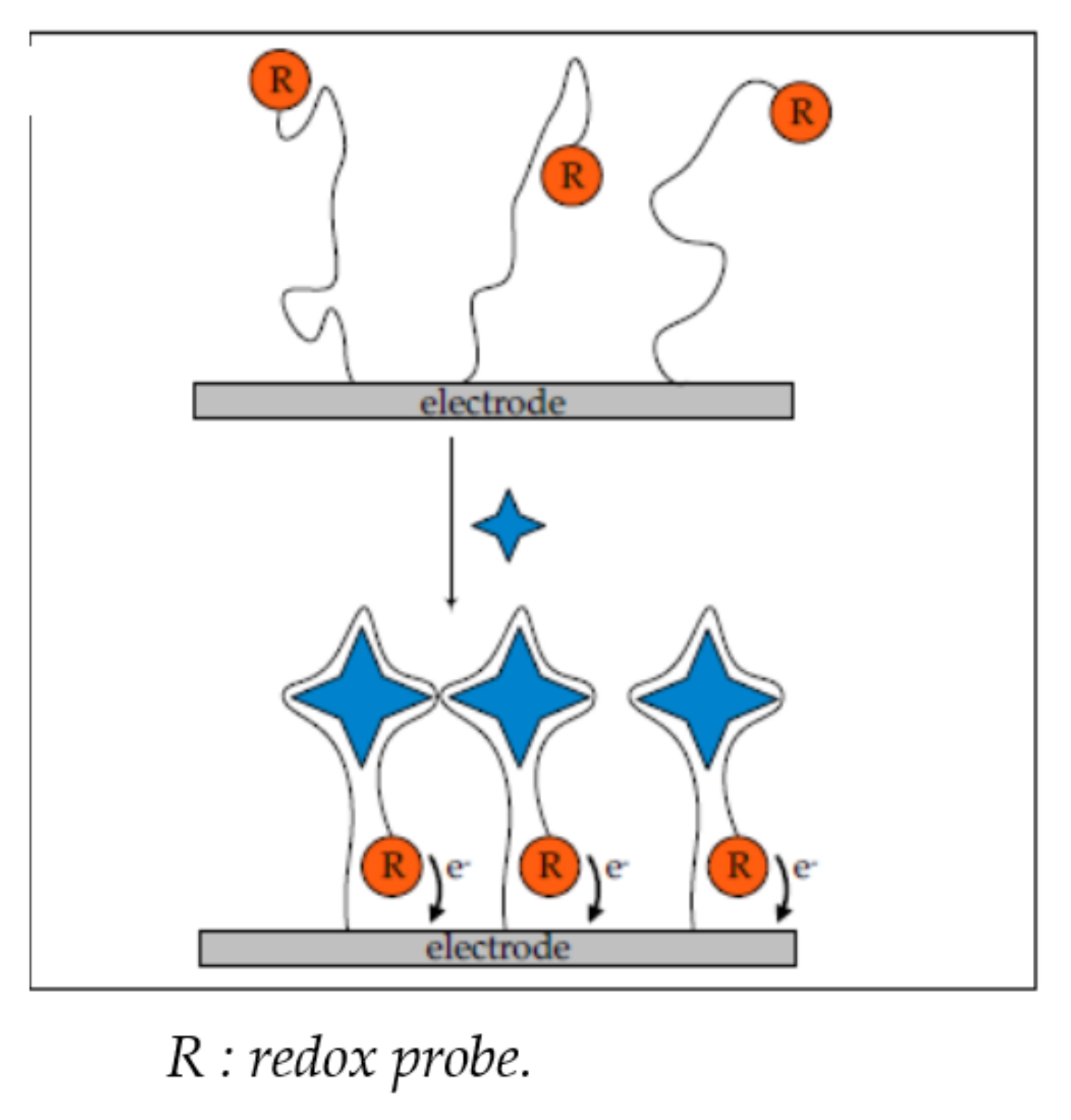
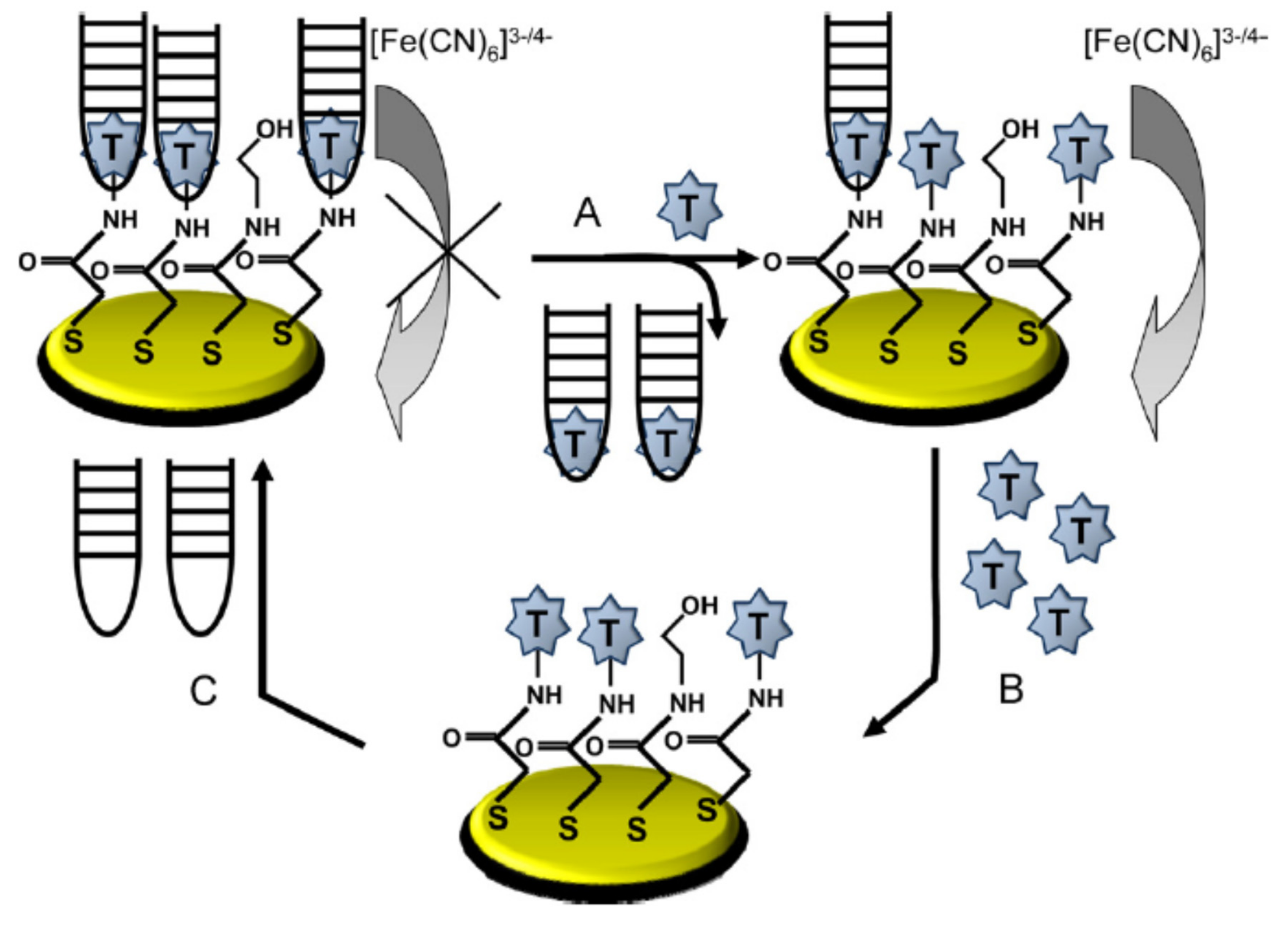
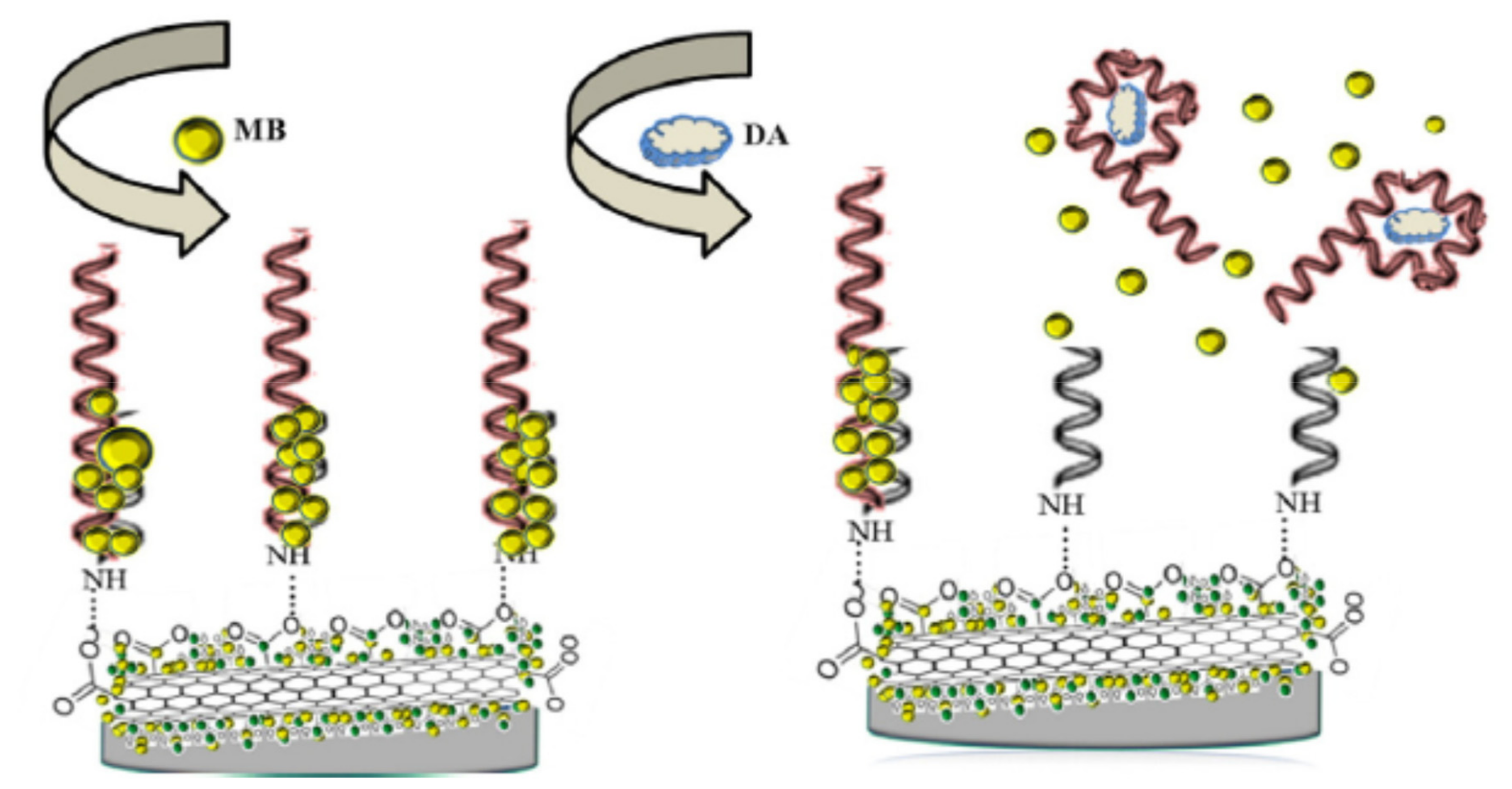
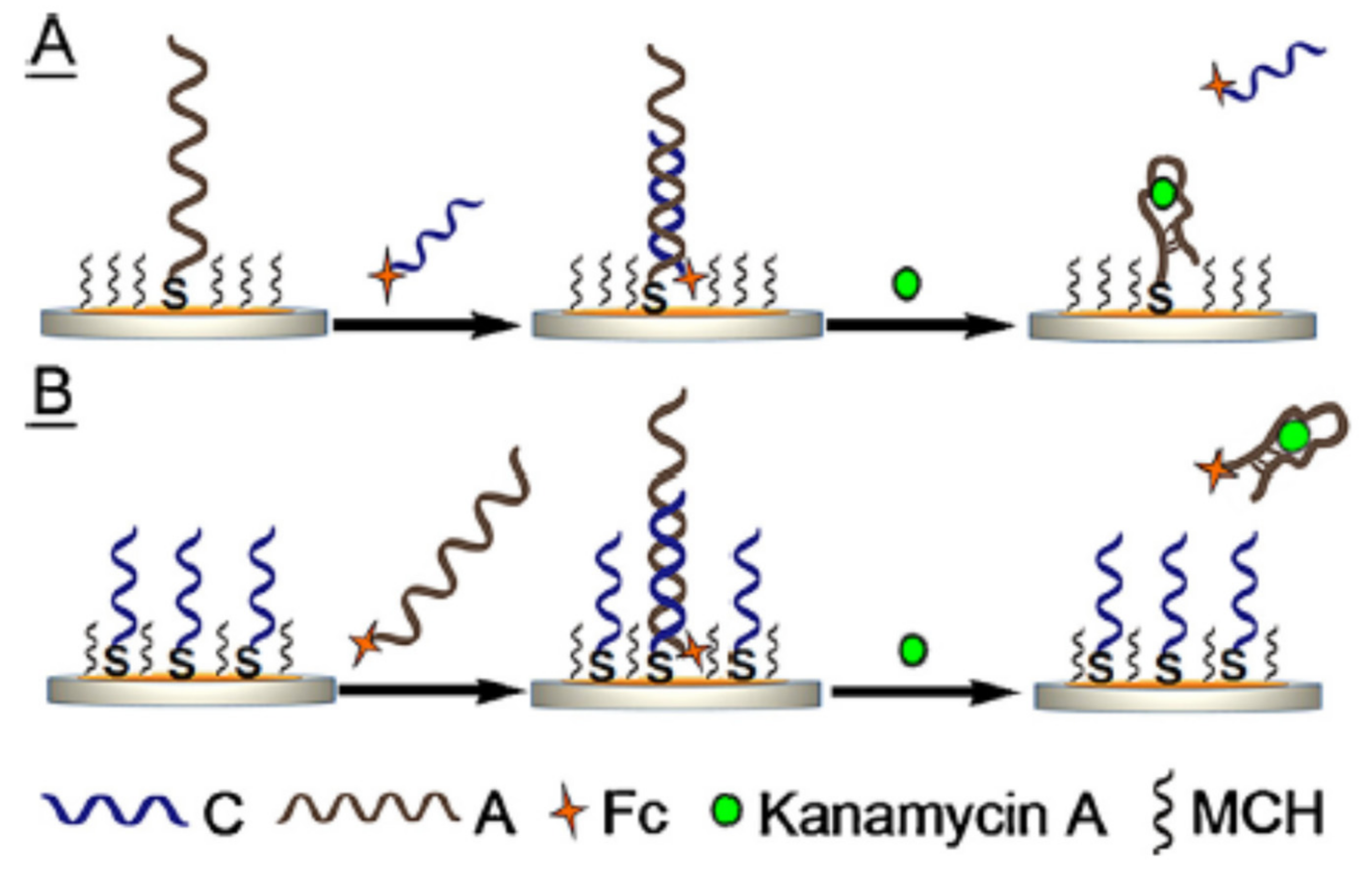
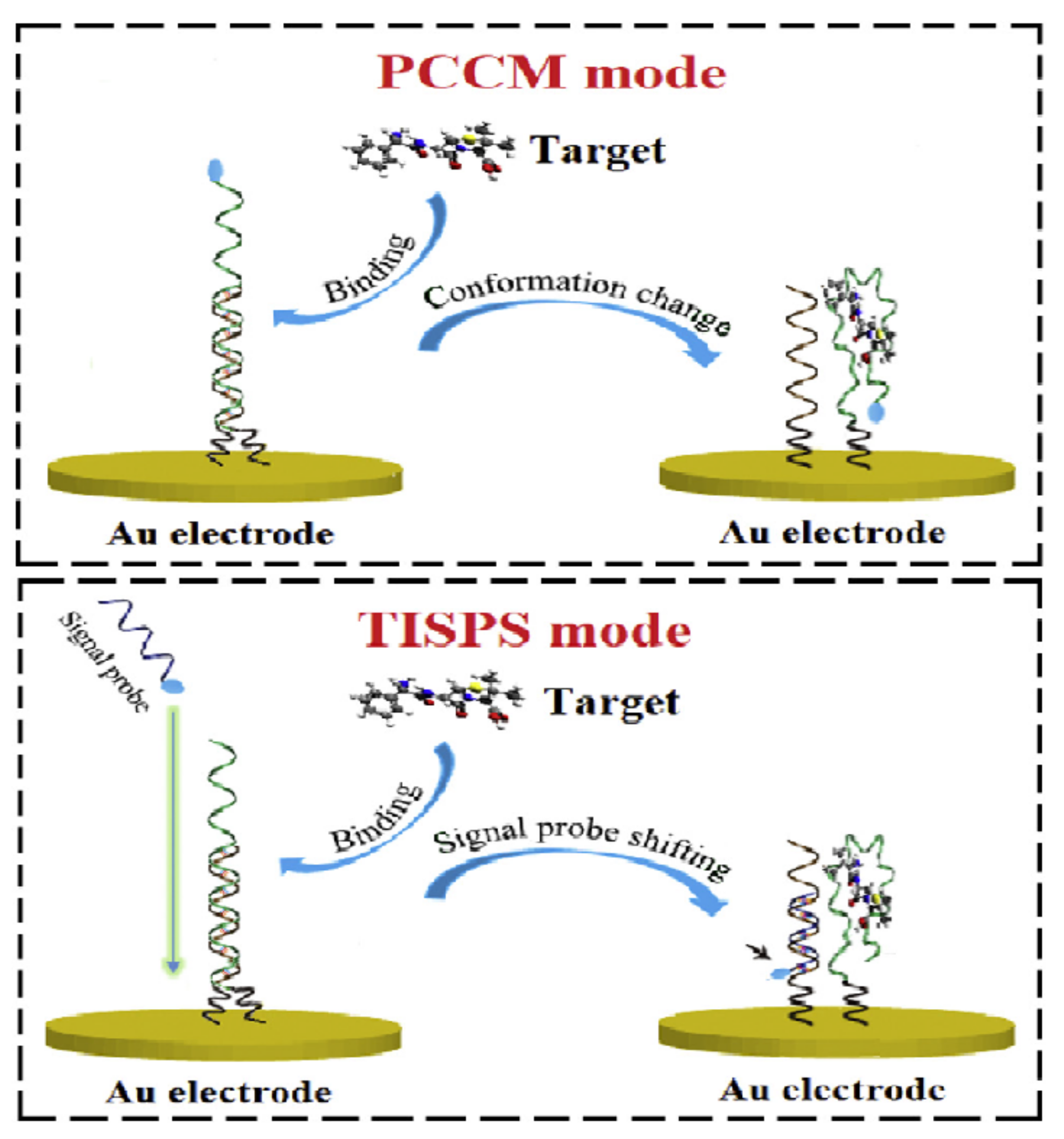
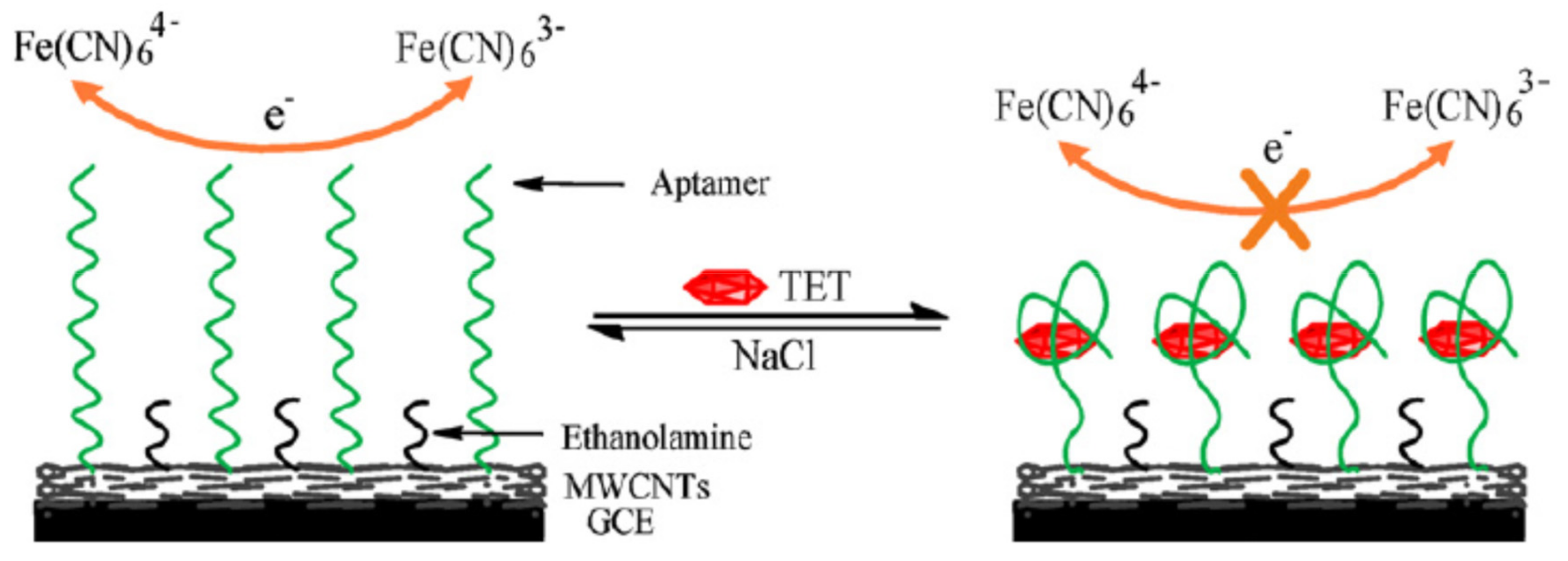
2.1.1. Change of Aptamer Conformation
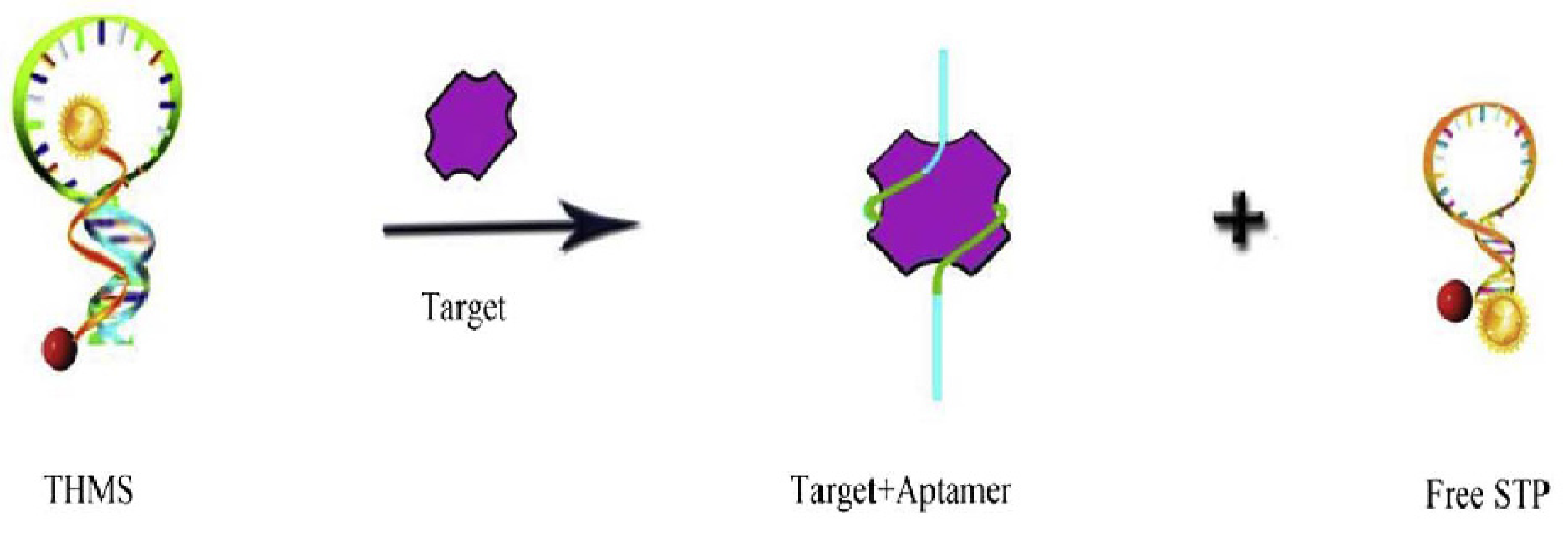
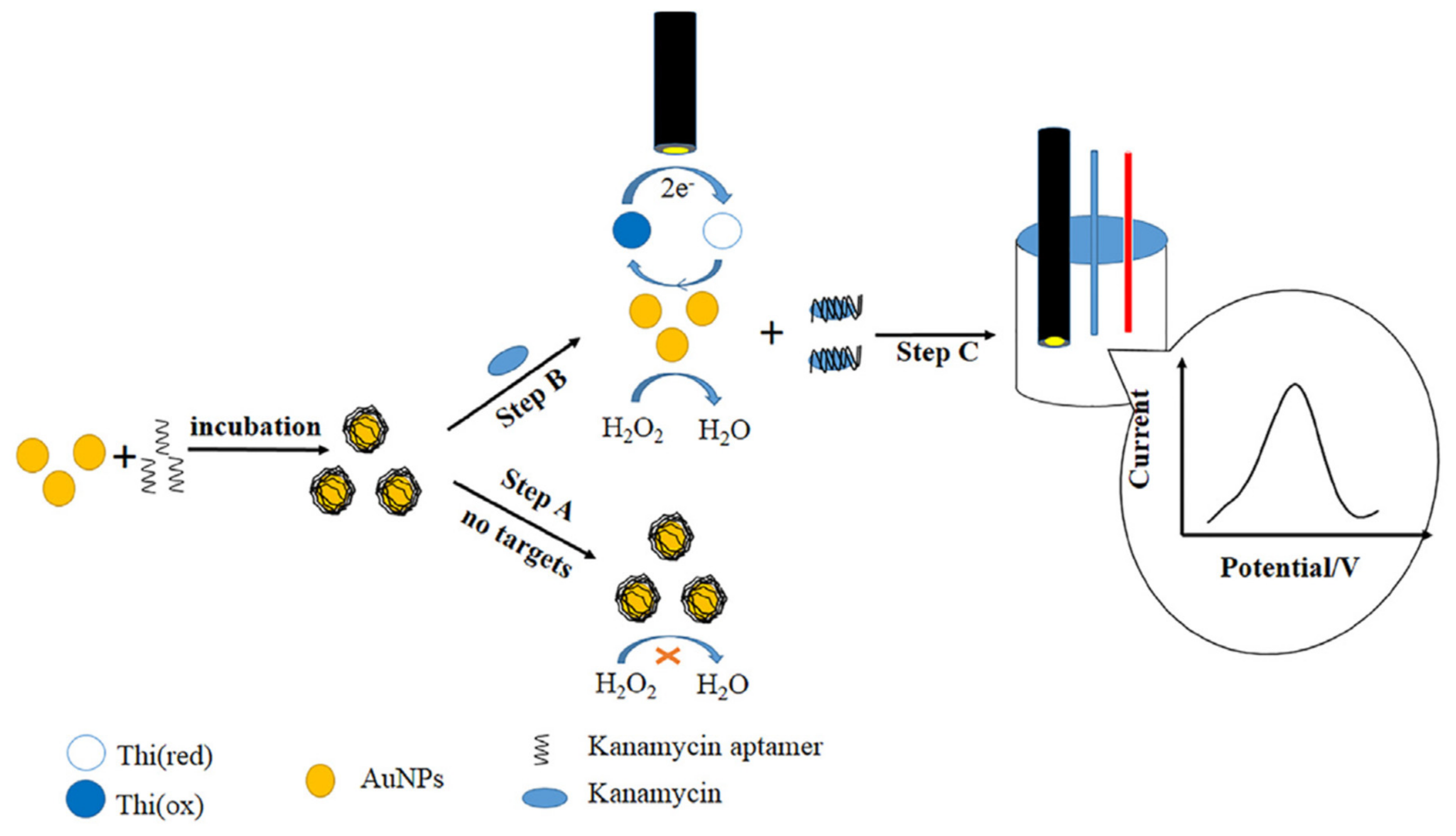
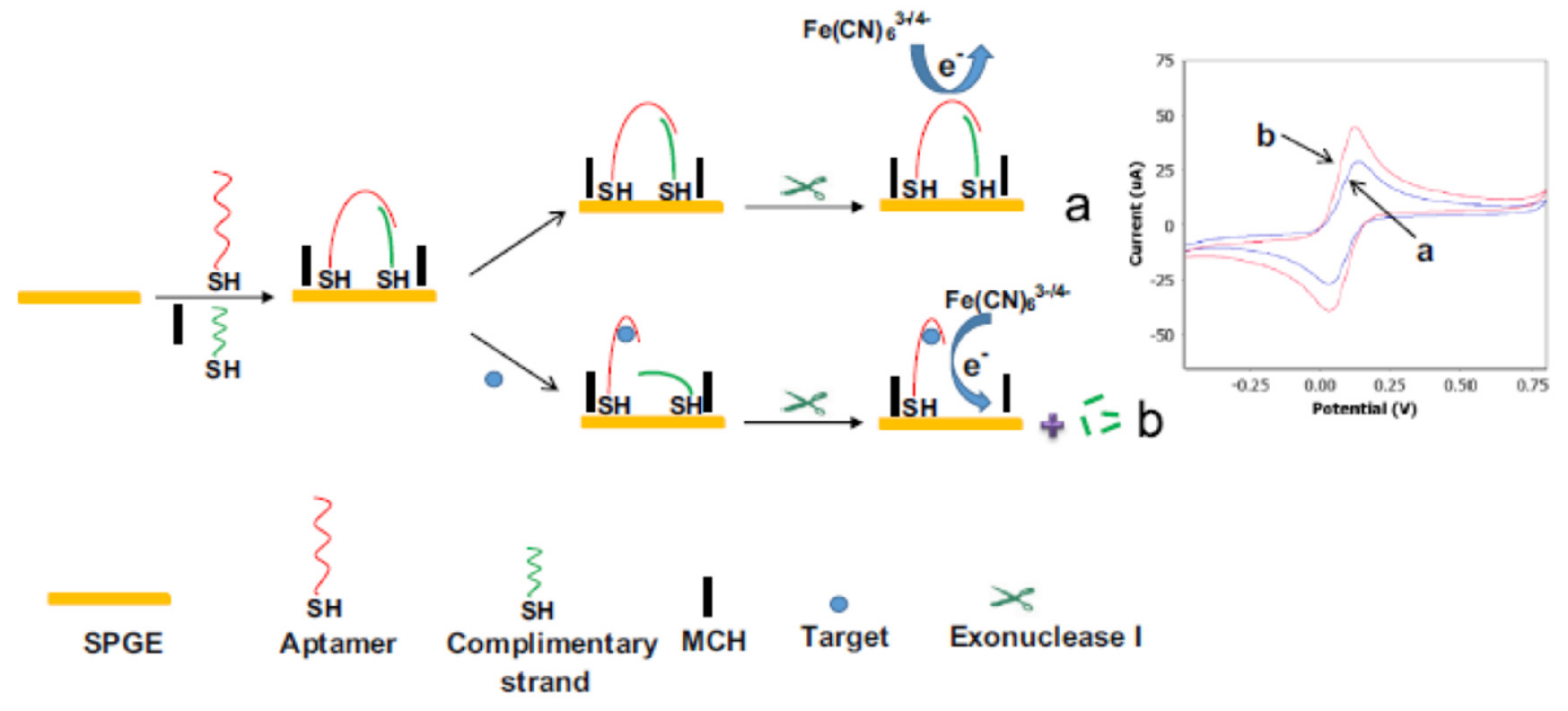
2.1.2. Target-Induced Displacement
2.2. Different Immobilisation Techniques of Aptamers
2.3. Nanomaterials
2.3.1. Carbon Nanomaterials (e.g., Carbon Nanotubes (CNT), Graphene, Graphene Oxide (GO))
2.3.2. Metallic Nanomaterials (e.g., Gold NPs, Silver NPs)
2.3.3. Metal Compound Nanomaterials (e.g., QDs, UCNPs, MOFs)
2.3.4. Conductive Polymers
2.3.5. Combination of Several Nanomaterials
2.4. Different Amplification Techniques
3. Conclusions and Perspectives
Funding
Acknowledgments
Conflicts of Interest
References
- van Duijkeren, E.; Schwarz, C.; Bouchard, D.; Catry, B.; Pomba, C.; Baptiste, K.E.; Moreno, M.A.; Rantala, M.; Ružauskas, M.; Sanders, P.; et al. The use of aminoglycosides in animals within the EU: Development of resistance in animals and possible impact on human and animal health: A review. J. Antimicrob. Chemother. 2019, 74, 2480–2496. [Google Scholar] [CrossRef]
- European Commission. Commission Regulation (EC) N° 470/2009 laying down Community procedures for the establishment of residue limits of pharmacologically active substances in foodstuffs of animal origin. Off. J. EU 2009, L152, 11–22.
- European Commission. Council Regulation (EU) (2010) N 37/2010 of 22 December 2009 on pharmacologically active substances and their classification regarding maximum residue limits in foodstuffs of animal origin. Off. J. EU 2009, L15, 1–72. [Google Scholar]
- European Commission. Commission Decision (EC) N° 2002/657 of 12 August 2002 implementing Council Directive 96/23/EC concerning the performance of analytical methods and interpretation of results. Off. J. EU 2002, L221, 8–36. [Google Scholar]
- European Commission. Commission Regulation (EU) 2019/1871 of 7 November 2019 on reference points for action for non-allowed pharmacologically active substances present in food of animal origin and repealing Decision 2005/34/EC (Text with EEA relevance). Off. J. EU 2019, L289, 41–46. [Google Scholar]
- European Commission. Regulation (EU) 2017/625 of the European Parliament and of the Council of 15 March 2017 on official controls and other official activities performed to ensure the application of food and feed law, rules on animal health and welfare, plant health and plant protection products. Off. J. EU 2017, L95, 1–142. [Google Scholar]
- Marco, M.-P.; Gee, S.; Hammock, B.D. Immunochemical techniques for environmental analysis I. Immunosensors. TrAC Trends Anal. Chem. 1995, 14, 341–350. [Google Scholar] [CrossRef]
- Gaudin, V. Advances in biosensor development for the screening of antibiotic residues in food products of animal origin—A comprehensive review. Biosens. Bioelectron. 2017, 90, 363–377. [Google Scholar] [CrossRef]
- Gaudin, V. The Growing Interest in Development of Innovative Optical Aptasensors for the Detection of Antimicrobial Residues in Food Products. Biosensors 2020, 10, 21. [Google Scholar] [CrossRef]
- Oberhaus, F.V.; Frense, D.; Beckmann, D. Immobilization Techniques for Aptamers on Gold Electrodes for the Electrochemical Detection of Proteins: A Review. Biosensors 2020, 10, 45. [Google Scholar] [CrossRef]
- Piro, B.; Shi, S.; Reisberg, S.; Noël, V.; Anquetin, G. Comparison of Electrochemical Immunosensors and Aptasensors for Detection of Small Organic Molecules in Environment, Food Safety, Clinical and Public Security. Biosensors 2016, 6, 7. [Google Scholar] [CrossRef] [PubMed]
- Amaya-González, S.; De-los-Santos-Álvarez, N.; Miranda-Ordieres, A.J.; Lobo-Castañón, M.J. Aptamer-Based Analysis: A Promising Alternative for Food Safety Control. Sensors 2013, 13, 16292–16311. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Killian, J.; Hamasaki, K.; Rando, R.R. RNA Molecules That Specifically and Stoichiometrically Bind Aminoglycoside Antibiotics with High Affinities. Biochemistry 1996, 35, 12338–12346. [Google Scholar] [CrossRef] [PubMed]
- Burke, D.H.; Hoffman, D.C.; Brown, A.; Hansen, M.; Pardi, A.; Gold, L. RNA aptamers to the peptidyl transferase inhibitor chloramphenicol. Chem. Biol. 1997, 4, 833–843. [Google Scholar] [CrossRef]
- Berens, C.; Thain, A.; Schroeder, R. A tetracycline-binding RNA aptamer. Bioorg. Med. Chem. 2001, 9, 2549–2556. [Google Scholar] [CrossRef]
- Mehlhorn, A.; Rahimi, P.; Joseph, Y. Aptamer-Based Biosensors for Antibiotic Detection: A Review. Biosensors 2018, 8, 54. [Google Scholar] [CrossRef]
- Kwon, Y.S.; Ahmad Raston, N.H.; Gu, M.B. An ultra-sensitive colorimetric detection of tetracyclines using the shortest aptamer with highly enhanced affinity. Chem. Commun. 2014, 50, 40–42. [Google Scholar] [CrossRef]
- Hianik, T.; Wang, J. Electrochemical Aptasensors—Recent Achievements and Perspectives. Electroanalysis 2009, 21, 1223–1235. [Google Scholar] [CrossRef]
- González-Fernández, E.; de-los-Santos-Álvarez, N.; Lobo-Castañón, M.J.; Miranda-Ordieres, A.J.; Tuñón-Blanco, P. Impedimetric aptasensor for tobramycin detection in human serum. Biosens. Bioelectron. 2011, 26, 2354–2360. [Google Scholar] [CrossRef]
- Kim, Y.-J.; Kim, Y.S.; Niazi, J.H.; Gu, M.B. Electrochemical aptasensor for tetracycline detection. Bioprocess Biosyst. Eng. 2010, 33, 31. [Google Scholar] [CrossRef]
- Nguyen, T.; Hilton, J.P.; Lin, Q. Emerging applications of aptamers to micro- and nanoscale biosensing. Microfluid. Nanofluidics 2009, 6, 347. [Google Scholar] [CrossRef]
- Sharma, A.; Chandra Singh, A.; Bacher, G.; Bhand, S. Recent Advances in Aptamer- Based Biosensors for Detection of Antibiotic Residues. Aptamers Synth. Antibodies 2016, 2, 43–54. [Google Scholar]
- Viglasky, V.; Hianik, T. Potential uses of G-quadruplex-forming aptamers. Gen. Physiol. Biophys. 2013, 32, 149–172. [Google Scholar] [CrossRef] [PubMed]
- Chiorcea-Paquim, A.-M.; Oliveira-Brett, A.M. Redox behaviour of G-quadruplexes. Electrochim. Acta 2014, 126, 162–170. [Google Scholar] [CrossRef]
- Bagheri, E.; Abnous, K.; Alibolandi, M.; Ramezani, M.; Taghdisi, S.M. Triple-helix molecular switch-based aptasensors and DNA sensors. Biosens. Bioelectron. 2018, 111, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Jalalian, S.H.; Karimabadi, N.; Ramezani, M.; Abnous, K.; Taghdisi, S.M. Electrochemical and optical aptamer-based sensors for detection of tetracyclines. Trends Food Sci. Technol. 2018, 73, 45–57. [Google Scholar] [CrossRef]
- Schoukroun-Barnes, L.R.; Wagan, S.; White, R.J. Enhancing the Analytical Performance of Electrochemical RNA Aptamer-Based Sensors for Sensitive Detection of Aminoglycoside Antibiotics. Anal. Chem. 2014, 86, 1131–1137. [Google Scholar] [CrossRef]
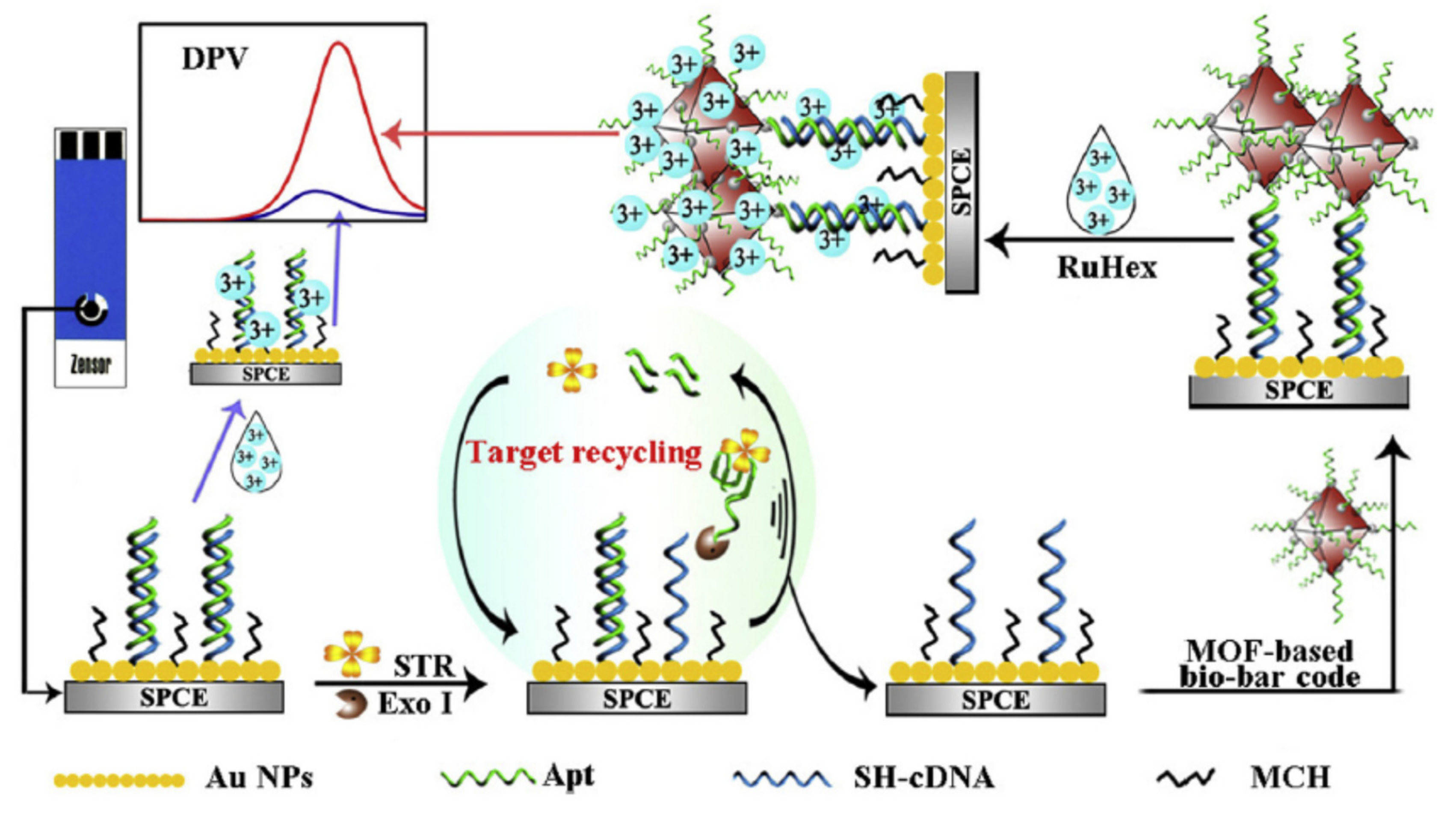
- Meng, X.; Gu, H.; Yi, H.; He, Y.; Chen, Y.; Sun, W. Sensitive detection of streptomycin in milk using a hybrid signal enhancement strategy of MOF-based bio-bar code and target recycling. Anal. Chim. Acta 2020, 1125, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Yu, Z.; Cui, P.; Xiang, Y.; Li, B.; Han, X.; Shi, W.; Yan, H.; Zhang, G. Developing a fast electrochemical aptasensor method for the quantitative detection of penicillin G residue in milk with high sensitivity and good anti-fouling ability. Microchem. J. 2020, 157, 105077. [Google Scholar] [CrossRef]
- Kim, Y.S.; Gu, M.B. Advances in Aptamer Screening and Small Molecule Aptasensors. In Biosensors Based on Aptamers and Enzymes; Gu, M.B., Kim, H.-S., Eds.; Springer: Berlin/Heidelberg, Germany, 2014; pp. 29–67. [Google Scholar] [CrossRef]
- Wang, Q.; Qin, X.; Geng, L.; Wang, Y. Label-Free Electrochemical Aptasensor for Sensitive Detection of Malachite Green Based on Au Nanoparticle/Graphene Quantum Dots/Tungsten Disulfide Nanocomposites. Nanomaterials 2019, 9, 229. [Google Scholar] [CrossRef]
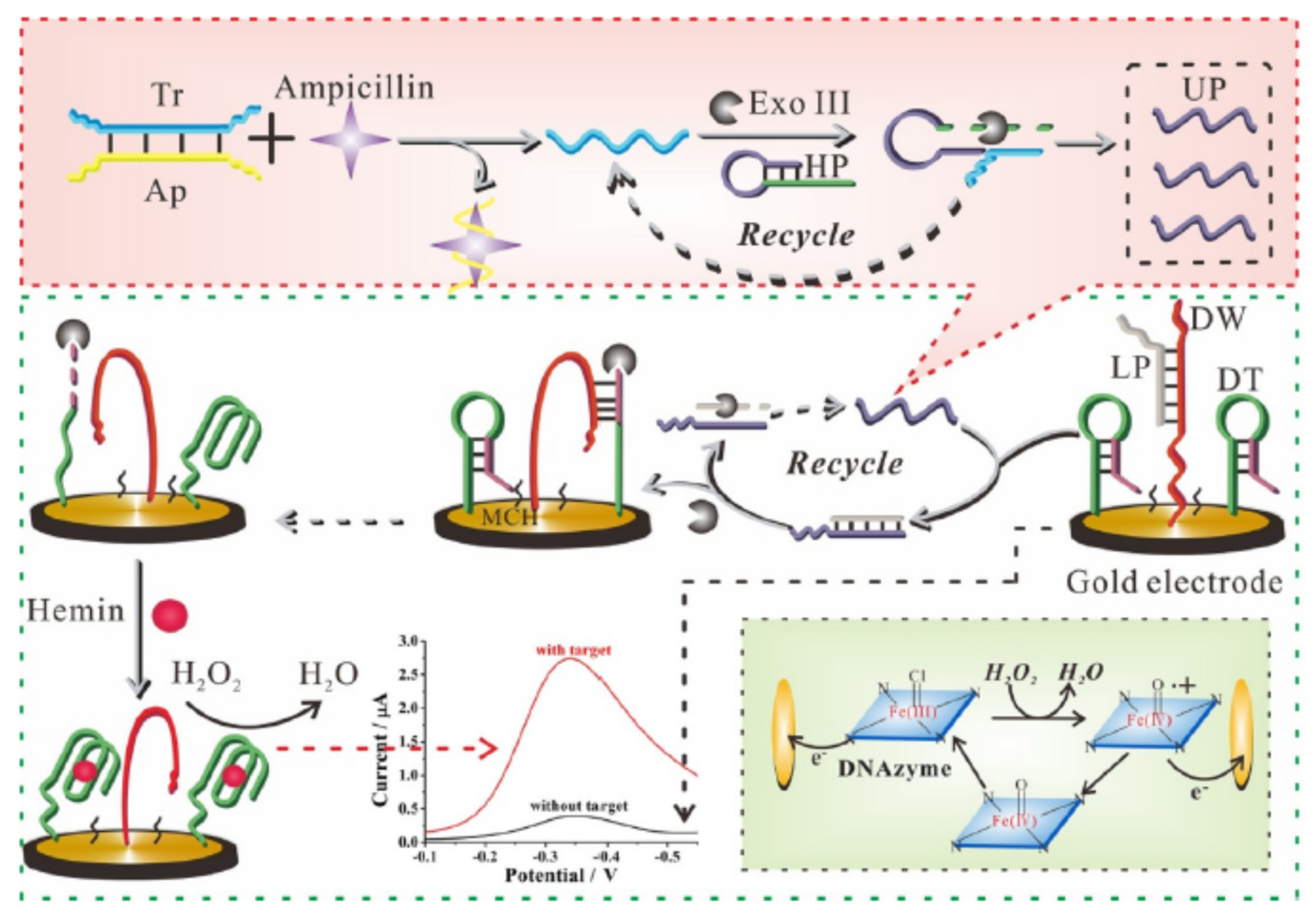
- Taghdisi, S.M.; Danesh, N.M.; Nameghi, M.A.; Ramezani, M.; Alibolandi, M.; Abnous, K. An electrochemical sensing platform based on ladder-shaped DNA structure and label-free aptamer for ultrasensitive detection of ampicillin. Biosens. Bioelectron. 2019. [Google Scholar] [CrossRef]
- Alizadeh, N.; Salimi, A.; Hallaj, R. Hemin/G-Quadruplex Horseradish Peroxidase-Mimicking DNAzyme: Principle and Biosensing Application. Adv. Biochem. Eng. Biotechnol. 2017, 170, 85–106. [Google Scholar] [CrossRef]
- Zhang, R.; Li, S.; Wang, J.; Qu, X.; Zhao, Y.; Liu, S.; Wang, Y.; Huang, J.; Yu, J. Entropy-driven spliced DNA walking machine for label-free electrochemical detection of antibiotics. Sens. Actuators B 2020, 320, 128385. [Google Scholar] [CrossRef]
- Zhang, R.; Wang, Y.; Qu, X.; Li, S.; Zhao, Y.; Liu, S.; Huang, J. Exonuclease III-powered DNA Walking Machine for Label-free and Ultrasensitive Electrochemical Sensing of Antibiotic. Sens. Actuators B 2019, 297, 126771. [Google Scholar] [CrossRef]
- Han, C.; Li, R.; Li, H.; Liu, S.; Xu, C.; Wang, J.; Wang, Y.; Huang, J. Ultrasensitive voltammetric determination of kanamycin using a target-triggered cascade enzymatic recycling couple along with DNAzyme amplification. Microchim. Acta 2017, 184, 2941–2948. [Google Scholar] [CrossRef]
- Huang, Y.; Zheng, J.; Wang, L.; Duan, X.; Wang, Y.; Xiang, Y.; Li, G. Sensitive Detection of Chloramphenicol Based on Ag-DNAzyme-Mediated Signal Amplification Modulated by DNA/Metal Ion Interaction. Biosens. Bioelectron. 2018. [Google Scholar] [CrossRef] [PubMed]
- Azadbakht, A.; Roushani, M.; Abbasi, A.R.; Menati, S.; Derikvand, Z. A label-free aptasensor based on polyethyleneimine wrapped carbon nanotubes in situ formed gold nanoparticles as signal probe for highly sensitive detection of dopamine. Mater. Sci. Eng. C 2016, 68, 585–593. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Yu, Z.; Han, X.; Shi, W.; Liu, Y.; Yan, H.; Zhang, G. A signal-on electrochemical aptasensor for highly sensitive and specific detection of kanamycin based on target-induced signaling probe shifting mechanism. Sens. Actuators B 2018, 273, 480–487. [Google Scholar] [CrossRef]
- Liu, R.; Yang, Z.; Guo, Q.; Zhao, J.; Ma, J.; Kang, Q.; Tang, Y.; Xue, Y.; Lou, X.; He, M. Signaling-Probe Displacement Electrochemical Aptamer-based Sensor (SD-EAB) for Detection of Nanomolar Kanamycin A. Electrochim. Acta 2015, 182, 516–523. [Google Scholar] [CrossRef]
- Han, X.; Yu, Z.; Li, F.; Shi, W.; Fu, C.; Yan, H.; Zhang, G. Two kanamycin electrochemical aptamer-based sensors using different signal transduction mechanisms: A comparison of electrochemical behavior and sensing performance. Bioelectrochemistry 2019, 129, 270–277. [Google Scholar] [CrossRef]
- Herne, T.M.; Tarlov, M.J. Characterization of DNA Probes Immobilized on Gold Surfaces. J. Am. Chem. Soc. 1997, 119, 8916–8920. [Google Scholar] [CrossRef]
- Nan, M.-N.; Bi, Y.; Xue, H.-L.; Long, H.-T.; Xue, S.-L.; Pu, L.-M.; Prusky, D. Modification performance and electrochemical characteristics of different groups of modified aptamers applied for label-free electrochemical impedimetric sensors. Food Chem. 2021, 337, 127761. [Google Scholar] [CrossRef]
- Li, F.; Wang, X.; Sun, X.; Guo, Y.; Zhao, W. A dual-signal amplification strategy for kanamycin based on ordered mesoporous carbon-chitosan/gold nanoparticles-streptavidin and ferrocene labelled DNA. Anal. Chim. Acta 2018, 1033, 185–192. [Google Scholar] [CrossRef]
- Sharma, A.; Istamboulie, G.; Hayat, A.; Catanante, G.; Bhand, S.; Marty, J.L. Disposable and portable aptamer functionalized impedimetric sensor for detection of kanamycin residue in milk sample. Sens. Actuators B 2017, 245, 507–515. [Google Scholar] [CrossRef]
- Tanimu, A.; Lawal, M.A. electrochemical-sensors-using-nanomaterials—A-mini-review. Res. Rev. J. Chem. 2017, 6, 38–48. [Google Scholar]
- Malhotra, B.D.; Ali, M.A. Chapter 1—Nanomaterials in Biosensors: Fundamentals and Applications. In Nanomaterials for Biosensors; Malhotra, B.D., Ali, M.A., Eds.; William Andrew Publishing: Amsterdam, The Netherlands, 2018; pp. 1–74. [Google Scholar] [CrossRef]
- Yang, T.; Huang, H.; Zhu, F.; Lin, Q.; Zhang, L.; Liu, J. Recent Progresses in Nanobiosensing for Food Safety Analysis. Sensors 2016, 16, 1118. [Google Scholar] [CrossRef]
- Abi, A.; Mohammadpour, Z.; Zuo, X.; Safavi, A. Nucleic acid-based electrochemical nanobiosensors. Biosens. Bioelectron. 2018, 102, 479–489. [Google Scholar] [CrossRef] [PubMed]
- Lim, M.C.; Kim, Y.R. Analytical Applications of Nanomaterials in Monitoring Biological and Chemical Contaminants in Food. J Microbiol. Biotechnol. 2016, 26, 1505–1516. [Google Scholar] [CrossRef]
- Sharma, R.; Ragavan, K.V.; Thakur, M.S.; Raghavarao, K.S.M.S. Recent advances in nanoparticle based aptasensors for food contaminants. Biosens. Bioelectron. 2015, 74, 612–627. [Google Scholar] [CrossRef]
- Hu, M.; Wang, Y.; Yang, J.; Sun, Y.; Xing, G.; Deng, R.; Hu, X.; Zhang, G. Competitive electrochemical immunosensor for maduramicin detection by multiple signal amplification strategy via hemin@Fe-MIL-88NH2/AuPt. Biosens. Bioelectron. 2019, 111554. [Google Scholar] [CrossRef]
- Liu, S.; Lai, G.; Zhang, H.; Yu, A. Amperometric aptasensing of chloramphenicol at a glassy carbon electrode modified with a nanocomposite consisting of graphene and silver nanoparticles. Microchim. Acta 2017, 184, 1445–1451. [Google Scholar] [CrossRef]
- Huang, S.; Gan, N.; Zhang, X.; Wu, Y.; Shao, Y.; Jiang, Z.; Wang, Q. Portable fluoride-selective electrode as signal transducer for sensitive and selective detection of trace antibiotics in complex samples. Biosens. Bioelectron. 2019, 128, 113–121. [Google Scholar] [CrossRef]
- Yao, Y.; Jiang, C.; Ping, J. Flexible Freestanding Graphene Paper-based Potentiometric Enzymatic Aptasensor for Ultrasensitive Wireless Detection of Kanamycin. Biosens. Bioelectron. 2018. [Google Scholar] [CrossRef]
- Ebrahimi Vafaye, S.; Rahman, A.; Safaeian, S.; Adabi, M. An electrochemical aptasensor based on electrospun carbon nanofiber mat and gold nanoparticles for the sensitive detection of Penicillin in milk. J. Food Meas. Charact. 2021, 15, 876–882. [Google Scholar] [CrossRef]
- Alawad, A.; Istamboulié, G.; Calas-Blanchard, C.; Noguer, T. A reagentless aptasensor based on intrinsic aptamer redox activity for the detection of tetracycline in water. Sens. Actuators B 2019, 288, 141–146. [Google Scholar] [CrossRef]
- Shi, Z.; Hou, W.; Jiao, Y.; Guo, Y.; Sun, X.; Zhao, J.; Wang, X. Ultra-sensitive aptasensor based on IL and Fe3O4 nanoparticles for tetracycline detection. Int. J. Electrochem. Sci. 2017, 12, 7426–7434. [Google Scholar] [CrossRef]
- Guo, Y.; Zhang, Q.; Xu, Q.; Ma, N.; Sun, X.; Wang, X. Fabrication of aptasensors modified by MWCNTs-CS/Fe3O4-CS based on SPEs. Int. J. Electrochem. Sci. 2016, 11, 1691–1698. [Google Scholar]
- Taghdisi Heidarian, S.M.; Tavanaee Sani, A.; Danesh, N.M.; Ramezani, M.; Alibolandi, M.; Gerayelou, G.; Abnous, K.; Taghdisi, S.M. A novel electrochemical approach for the ultrasensitive detection of fluoroquinolones based on a double-labelled aptamer to surpass complementary strands of aptamer lying flat. Sens. Actuators B 2021, 334, 129632. [Google Scholar] [CrossRef]
- Huang, W.; Zhou, Y.; Zhan, D.; Lai, G. Homogeneous biorecognition reaction-induced assembly of DNA nanostructures for ultrasensitive electrochemical detection of kanamycin antibiotic. Anal. Chim. Acta 2021, 1154, 338317. [Google Scholar] [CrossRef]
- Lu, M.; Cao, C.; Wang, F.; Liu, G. A polyethyleneimine reduced graphene oxide/gold nanocubes based electrochemical aptasensor for chloramphenicol detection using single-stranded DNA-binding protein. Mater. Des. 2021, 199, 109409. [Google Scholar] [CrossRef]
- Roushani, M.; Rahmati, Z.; Farokhi, S.; Hoseini, S.J.; Fath, R.H. The development of an electrochemical nanoaptasensor to sensing chloramphenicol using a nanocomposite consisting of graphene oxide functionalized with (3-Aminopropyl) triethoxysilane and silver nanoparticles. Mater. Sci. Eng. C 2020, 108, 110388. [Google Scholar] [CrossRef]
- Bi, H.; Wu, Y.; Wang, Y.; Liu, G.; Ning, G.; Xu, Z. A molecularly imprinted polymer combined with dual functional Au@Fe3O4 nanocomposites for sensitive detection of kanamycin. J. Electroanal. Chem. 2020, 870, 114216. [Google Scholar] [CrossRef]
- Guo, W.; Umar, A.; Alsaiari, M.A.; Wang, L.; Pei, M. Ultrasensitive and selective label-free aptasensor for the detection of penicillin based on nanoporous PtTi/graphene oxide-Fe3O4/MWCNT-Fe3O4 nanocomposite. Microchem. J. 2020, 158, 105270. [Google Scholar] [CrossRef]
- Li, F. Magnetic bead-based electrochemical aptasensor doped with multi-wall carbon nanotubes for the detection of ampicillin in milk. Int. J. Electrochem. Sci. 2020, 7520–7530. [Google Scholar] [CrossRef]
- Sebastian, N.; Yu, W.-C.; Balram, D. Electrochemical detection of an antibiotic drug chloramphenicol based on a graphene oxide/hierarchical zinc oxide nanocomposite. Inorg. Chem. Front. 2019, 6, 82–93. [Google Scholar] [CrossRef]
- Zhang, R.; Wang, Y.; Qu, X.; Li, S.; Zhao, Y.; Liu, S.; Zhang, F.; Huang, J.; Yu, J. A Label-free Electrochemical Platform for Antibiotics Detection Based on Cascade Enzymatic Amplification Coupled with Split G-quadruplex DNAzyme. Analyst 2019, 144. [Google Scholar] [CrossRef]
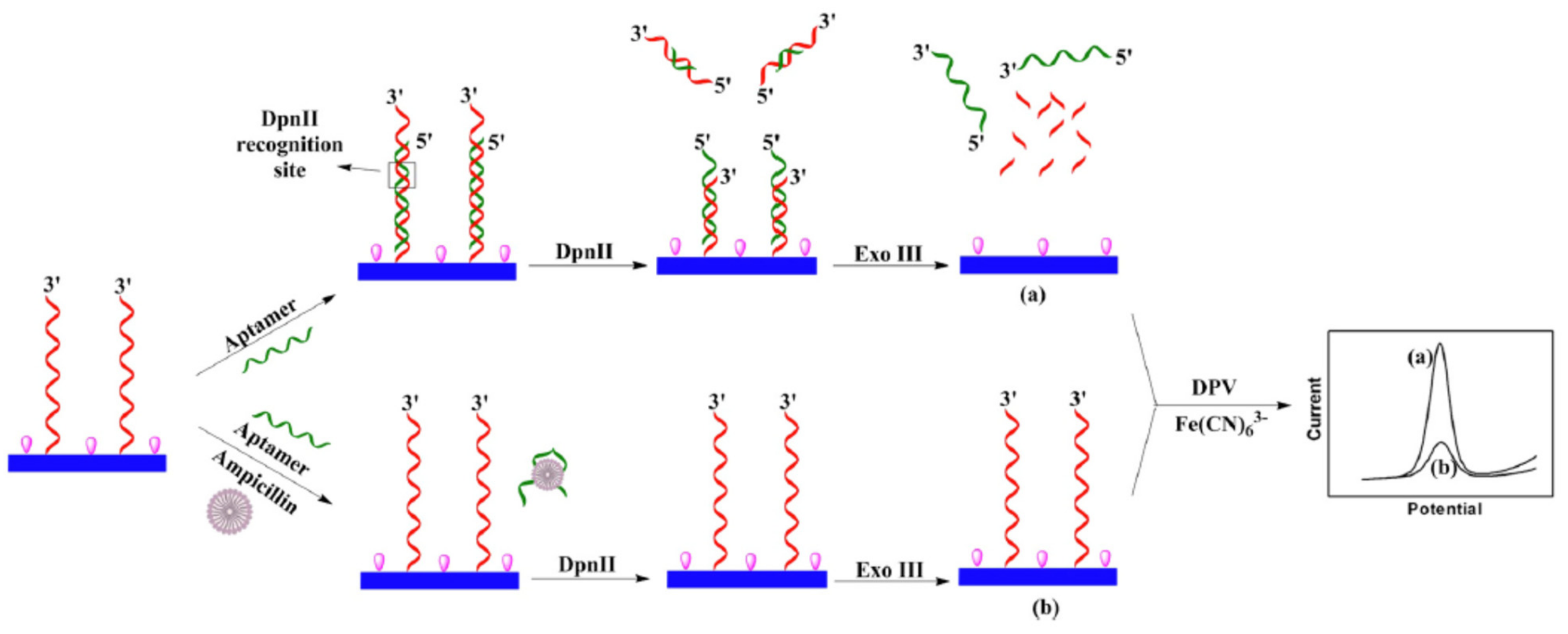
- Wang, T.; Yin, H.; Zhang, Y.; Wang, L.; Du, Y.; Zhuge, Y.; Ai, S. Electrochemical aptasensor for ampicillin detection based on the protective effect of aptamer-antibiotic conjugate towards DpnII and Exo III digestion. Talanta 2019, 197, 42–48. [Google Scholar] [CrossRef] [PubMed]
- Bai, Z.; Chen, Y.; Li, F.; Zhou, Y.; Yin, H.; Ai, S. Electrochemical aptasensor for sulfadimethoxine detection based on the triggered cleavage activity of nuclease P1 by aptamer-target complex. Talanta 2019, 204, 409–414. [Google Scholar] [CrossRef]
- Li, L.; Liu, X.; Yang, L.; Zhang, S.; Zheng, H.; Tang, Y.; Wong, D.K.Y. Amplified oxygen reduction signal at a Pt-Sn-modified TiO2 nanocomposite on an electrochemical aptasensor. Biosens. Bioelectron. 2019, 142, 111525. [Google Scholar] [CrossRef]
- He, B.; Wang, L.; Dong, X.; Yan, X.; Li, M.; Yan, S.; Yan, D. Aptamer-based thin film gold electrode modified with gold nanoparticles and carboxylated multi-walled carbon nanotubes for detecting oxytetracycline in chicken samples. Food Chem. 2019, 300, 125179. [Google Scholar] [CrossRef]
- You, H.; Bai, L.; Yuan, Y.; Zhou, J.; Bai, Y.; Mu, Z. An amperometric aptasensor for ultrasensitive detection of sulfadimethoxine based on exonuclease-assisted target recycling and new signal tracer for amplification. Biosens. Bioelectron. 2018, 117, 706–712. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Guo, Y.; Wang, X.; Sun, X. Multiplexed aptasensor based on metal ions labels for simultaneous detection of multiple antibiotic residues in milk. Biosens. Bioelectron. 2018, 115, 7–13. [Google Scholar] [CrossRef]
- Chen, Z.; Lai, G.; Liu, S.; Yu, A. Ultrasensitive electrochemical aptasensing of kanamycin antibiotic by enzymatic signal amplification with a horseradish peroxidase-functionalized gold nanoprobe. Sens. Actuators B 2018, 273, 1762–1767. [Google Scholar] [CrossRef]
- Hong, F.; Chen, X.; Cao, Y.; Dong, Y.; Wu, D.; Hu, F.; Gan, N. Enzyme- and label-free electrochemical aptasensor for kanamycin detection based on double stir bar-assisted toehold-mediated strand displacement reaction for dual-signal amplification. Biosens. Bioelectron. 2018, 112, 202–208. [Google Scholar] [CrossRef] [PubMed]
- Yin, J.; Guo, W.; Qin, X.; Zhao, J.; Pei, M.; Ding, F. A sensitive electrochemical aptasensor for highly specific detection of streptomycin based on the porous carbon nanorods and multifunctional graphene nanocomposites for signal amplification. Sens. Actuators B 2017, 241, 151–159. [Google Scholar] [CrossRef]
- Taghdisi, S.M.; Danesh, N.M.; Ramezani, M.; Abnous, K. A novel M-shape electrochemical aptasensor for ultrasensitive detection of tetracyclines. Biosens. Bioelectron. 2016, 85, 509–514. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Guo, W.; Pei, M.; Ding, F. GR–Fe3O4NPs and PEDOT–AuNPs composite based electrochemical aptasensor for the sensitive detection of penicillin. Anal. Methods 2016, 8, 4391–4397. [Google Scholar] [CrossRef]
- Mohammad Danesh, N.; Ramezani, M.; Sarreshtehdar Emrani, A.; Abnous, K.; Taghdisi, S.M. A novel electrochemical aptasensor based on arch-shape structure of aptamer-complimentary strand conjugate and exonuclease I for sensitive detection of streptomycin. Biosens. Bioelectron. 2016, 75, 123–128. [Google Scholar] [CrossRef] [PubMed]
- Yin, J.; Guo, W.; Qin, X.; Pei, M.; Wang, L.; Ding, F. A regular “signal attenuation” electrochemical aptasensor for highly sensitive detection of streptomycin. New J. Chem. 2016, 40, 9711–9718. [Google Scholar] [CrossRef]
- Hamidi-Asl, E.; Dardenne, F.; Blust, R.; De Wael, K. An Improved Electrochemical Aptasensor for Chloramphenicol Detection Based on Aptamer Incorporated Gelatine. Sensors 2015, 15, 7605–7618. [Google Scholar] [CrossRef]
- Yan, L.; Luo, C.; Cheng, W.; Mao, W.; Zhang, D.; Ding, S. A simple and sensitive electrochemical aptasensor for determination of Chloramphenicol in honey based on target-induced strand release. J. Electroanal. Chem. 2012, 687, 89–94. [Google Scholar] [CrossRef]
- Shen, Z.; He, L.; Cao, Y.; Hong, F.; Zhang, K.; Hu, F.; Lin, J.; Wu, D.; Gan, N. Multiplexed electrochemical aptasensor for antibiotics detection using metallic-encoded apoferritin probes and double stirring bars-assisted target recycling for signal amplification. Talanta 2019, 197, 491–499. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Gan, N.; Li, T.; Zhou, Y.; Cao, Y.; Dong, Y. Electrochemical aptasensor for multi-antibiotics detection based on endonuclease and exonuclease assisted dual recycling amplification strategy. Talanta 2018, 179, 28–36. [Google Scholar] [CrossRef]
- Chen, M.; Gan, N.; Zhou, Y.; Li, T.; Xu, Q.; Cao, Y.; Chen, Y. A novel aptamer- metal ions- nanoscale MOF based electrochemical biocodes for multiple antibiotics detection and signal amplification. Sens. Actuators B 2017, 242, 1201–1209. [Google Scholar] [CrossRef]
- Chen, M.; Gan, N.; Li, T.; Wang, Y.; Xu, Q.; Chen, Y. An electrochemical aptasensor for multiplex antibiotics detection using Y-shaped DNA-based metal ions encoded probes with NMOF substrate and CSRP target-triggered amplification strategy. Anal. Chim. Acta 2017, 968, 30–39. [Google Scholar] [CrossRef]
- Yan, Z.; Gan, N.; Li, T.; Cao, Y.; Chen, Y. A sensitive electrochemical aptasensor for multiplex antibiotics detection based on high-capacity magnetic hollow porous nanotracers coupling exonuclease-assisted cascade target recycling. Biosens. Bioelectron. 2016, 78, 51–57. [Google Scholar] [CrossRef]
- Pilehvar, S.; Mehta, J.; Dardenne, F.; Robbens, J.; Blust, R.; De Wael, K. Aptasensing of Chloramphenicol in the Presence of Its Analogues: Reaching the Maximum Residue Limit. Anal. Chem. 2012, 84, 6753–6758. [Google Scholar] [CrossRef]
- Xue, J.; Liu, J.; Wang, C.; Tian, Y.; Zhou, N. Simultaneous electrochemical detection of multiple antibiotic residues in milk based on aptamers and quantum dots. Anal. Methods 2016, 8, 1981–1988. [Google Scholar] [CrossRef]
- Yuan, R.; Yan, Z.; Shaga, A.; He, H. Design and fabrication of an electrochemical sensing platform based on a porous organic polymer for ultrasensitive ampicillin detection. Sens. Actuators B 2021, 327, 128949. [Google Scholar] [CrossRef]
- He, X.; Han, H.; Shi, W.; Dong, J.; Lu, X.; Yang, W.; Lu, X. A label-free electrochemical DNA biosensor for kanamycin detection based on diblock DNA with poly-cytosine as a high affinity anchor on graphene oxide. Anal. Methods 2020, 12, 3462–3469. [Google Scholar] [CrossRef]
- Mohammad-Razdari, A.; Ghasemi-Varnamkhasti, M.; Rostami, S.; Izadi, Z.; Ensafi, A.A.; Siadat, M. Development of an electrochemical biosensor for impedimetric detection of tetracycline in milk. J. Food Sci. Technol. 2020. [Google Scholar] [CrossRef]
- Kulikova, T.; Gorbatchuk, V.; Stoikov, I.; Rogov, A.; Evtugyn, G.; Hianik, T. Impedimetric Determination of Kanamycin in Milk with Aptasensor Based on Carbon Black-Oligolactide Composite. Sensors 2020, 20, 4738. [Google Scholar] [CrossRef]
- He, H.; Wang, S.-Q.; Han, Z.-Y.; Tian, X.-H.; Zhang, W.-W.; Li, C.-P.; Du, M. Construction of electrochemical aptasensors with Ag(I) metal−organic frameworks toward high-efficient detection of ultra-trace penicillin. Appl. Surf. Sci. 2020, 531, 147342. [Google Scholar] [CrossRef]
- Zhou, N.; Ma, Y.; Hu, B.; He, L.; Wang, S.; Zhang, Z.; Lu, S. Construction of Ce-MOF@COF hybrid nanostructure: Label-free aptasensor for the ultrasensitive detection of oxytetracycline residues in aqueous solution environments. Biosens. Bioelectron. 2019, 127, 92–100. [Google Scholar] [CrossRef]
- Yildirim-Tirgil, N.; Lee, J.; Cho, H.; Lee, H.; Somu, S.; Busnaina, A.; Gu, A.Z. A SWCNT based aptasensor system for antibiotic oxytetracycline detection in water samples. Anal. Methods 2019, 11, 2692–2699. [Google Scholar] [CrossRef]
- Mohammad-Razdari, A.; Ghasemi-Varnamkhasti, M.; Izadi, Z.; Ensafi, A.A.; Rostami, S.; Siadat, M. An impedimetric aptasensor for ultrasensitive detection of Penicillin G based on the use of reduced graphene oxide and gold nanoparticles. Microchim. Acta 2019, 186, 372. [Google Scholar] [CrossRef]
- Mohammad-Razdari, A.; Ghasemi-Varnamkhasti, M.; Izadi, Z.; Rostami, S.; Ensafi, A.A.; Siadat, M.; Losson, E. Detection of sulfadimethoxine in meat samples using a novel electrochemical biosensor as a rapid analysis method. J. Food Compos. Anal. 2019, 82, 103252. [Google Scholar] [CrossRef]
- Hu, X.; Goud, K.Y.; Kumar, V.S.; Catanante, G.; Li, Z.; Zhu, Z.; Marty, J.L. Disposable electrochemical aptasensor based on carbon nanotubes- V2O5-chitosan nanocomposite for detection of ciprofloxacin. Sens. Actuators B 2018, 268, 278–286. [Google Scholar] [CrossRef]
- Liu, X.; Hu, M.; Wang, M.; Song, Y.; Zhou, N.; He, L.; Zhang, Z. Novel Nanoarchitecture of Co-MOF-on-TPN-COF Hybrid: Ultralowly Sensitive Bioplatform of Electrochemical Aptasensor toward Ampicillin. Biosens. Bioelectron. 2018. [Google Scholar] [CrossRef] [PubMed]
- Paniel, N.; Istamboulié, G.; Triki, A.; Lozano, C.; Barthelmebs, L.; Noguer, T. Selection of DNA aptamers against penicillin G using Capture-SELEX for the development of an impedimetric sensor. Talanta 2017, 162, 232–240. [Google Scholar] [CrossRef]
- Xu, Q.-C.; Zhang, Q.-Q.; Sun, X.; Guo, Y.-M.; Wang, X.-Y. Aptasensors modified by antimony tin oxide nanoparticle-chitosan based on interdigitated array microelectrodes for tetracycline detection. RSC Adv. 2016, 6, 17328–17335. [Google Scholar] [CrossRef]
- Pilehvar, S.; Dierckx, T.; Blust, R.; Breugelmans, T.; De Wael, K. An Electrochemical Impedimetric Aptasensing Platform for Sensitive and Selective Detection of Small Molecules Such as Chloramphenicol. Sensors 2014, 14, 12059–12069. [Google Scholar] [CrossRef]
- Peng, J.; Yang, J.; Chen, B.; Zeng, S.; Zheng, D.; Chen, Y.; Gao, W. Design of ultrathin nanosheet subunits ZnIn2S4 hollow nanocages with enhanced photoelectric conversion for ultrasensitive photoelectrochemical sensing. Biosens. Bioelectron. 2021, 175, 112873. [Google Scholar] [CrossRef]
- Liu, D.; Meng, S.; Shen, X.; Li, Y.; Yan, X.; You, T. Dual-ratiometric aptasensor for streptomycin detection based on the in-situ coupling of photoelectrochemical and electrochemical assay with a bifunctional probe of methylene blue. Sens. Actuators B 2021, 332, 129529. [Google Scholar] [CrossRef]
- Liu, X.; Jiang, Y.; Luo, J.; Guo, X.; Ying, Y.; Wen, Y.; Yang, H.; Wu, Y. A SnO2/Bi2S3-based photoelectrochemical aptasensor for sensitive detection of tobramycin in milk. Food Chem. 2021, 344, 128716. [Google Scholar] [CrossRef] [PubMed]
- Yan, T.; Zhang, X.; Ren, X.; Lu, Y.; Li, J.; Sun, M.; Yan, L.; Wei, Q.; Ju, H. Fabrication of N-GQDs and AgBiS2 dual-sensitized ZIFs-derived hollow ZnxCo3-xO4 dodecahedron for sensitive photoelectrochemical aptasensing of ampicillin. Sens. Actuators B 2020, 320, 128387. [Google Scholar] [CrossRef]
- Chen, Y.; Wang, Y.; Yan, P.; Ouyang, Q.; Dong, J.; Qian, J.; Chen, J.; Xu, L.; Li, H. Co3O4 nanoparticles/graphitic carbon nitride heterojunction for photoelectrochemical aptasensor of oxytetracycline. Anal. Chim. Acta 2020, 1125, 299–307. [Google Scholar] [CrossRef]
- Dong, J.; Li, H.; Yan, P.; Xu, L.; Zhang, J.; Qian, J.; Chen, J.; Li, H. A label-free photoelectrochemical aptasensor for tetracycline based on Au/BiOI composites. Inorg. Chem. Commun. 2019, 109, 107557. [Google Scholar] [CrossRef]
- Zhu, Y.; Yan, K.; Xu, Z.; Wu, J.; Zhang, J. Cathodic “signal-on” photoelectrochemical aptasensor for chloramphenicol detection using hierarchical porous flower-like Bi-BiOI@C composite. Biosens. Bioelectron. 2019. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Gao, L.; Tang, L.; Peng, B.; Huang, H.; Wang, J.; Yu, J.; Ouyang, X.; Tan, J. Ultrathin PtNi nanozyme based self-powered photoelectrochemical aptasensor for ultrasensitive chloramphenicol detection. Biosens. Bioelectron. 2019, 111756. [Google Scholar] [CrossRef]
- Zhang, X.; Yan, T.; Wu, T.; Feng, Y.; Sun, M.; Yan, L.; Du, B.; Wei, Q. Fabrication of hierarchical MIL-68(In)-NH2/MWCNT/CdS composites for constructing label-free photoelectrochemical tetracycline aptasensor platform. Biosens. Bioelectron. 2019. [Google Scholar] [CrossRef]
- You, F.; Zhu, M.; Ding, L.; Xu, Y.; Wang, K. Design and construction of Z-scheme Bi2S3/nitrogen-doped graphene quantum dots: Boosted photoelectric conversion efficiency for high-performance photoelectrochemical aptasensing of sulfadimethoxine. Biosens. Bioelectron. 2019, 130, 230–235. [Google Scholar] [CrossRef] [PubMed]
- Ge, L.; Xu, Y.; Ding, L.; You, F.; Liu, Q.; Wang, K. Perovskite-type BiFeO3/ultrathin graphite-like carbon nitride nanosheets p-n heterojunction: Boosted visible-light-driven photoelectrochemical activity for fabricating ampicillin aptasensor. Biosens. Bioelectron. 2019, 124–125, 33–39. [Google Scholar] [CrossRef]
- Ge, L.; Liu, Q.; Jiang, D.; Ding, L.; Wen, Z.; Guo, Y.; Ding, C.; Wang, K. Oxygen vacancy enhanced photoelectrochemical performance of Bi2MoO6/B, N co-doped graphene for fabricating lincomycin aptasensor. Biosens. Bioelectron. 2019, 135, 145–152. [Google Scholar] [CrossRef]
- Qin, X.; Wang, Q.; Geng, L.; Shu, X.; Wang, Y. A “signal-on” photoelectrochemical aptasensor based on graphene quantum dots-sensitized TiO2 nanotube arrays for sensitive detection of chloramphenicol. Talanta 2019, 197, 28–35. [Google Scholar] [CrossRef] [PubMed]
- Qin, X.; Geng, L.; Wang, Q.; Wang, Y. Photoelectrochemical aptasensing of ofloxacin based on the use of a TiO2 nanotube array co-sensitized with a nanocomposite prepared from polydopamine and Ag2S nanoparticles. Microchim. Acta 2019, 186, 430. [Google Scholar] [CrossRef]
- Wang, Y.; Bian, F.; Qin, X.; Wang, Q. Visible light photoelectrochemical aptasensor for chloramphenicol by using a TiO2 nanorod array sensitized with Eu(III)-doped CdS quantum dots. Microchim. Acta 2018, 185, 161. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Sui, C.; Yin, H.; Wang, Y.; Wang, M.; Ai, S. Tungsten disulfide (WS2) nanosheet-based photoelectrochemical aptasensing of chloramphenicol. Microchim. Acta 2018, 185, 453. [Google Scholar] [CrossRef] [PubMed]
- Ge, L.; Li, H.; Du, X.; Zhu, M.; Chen, W.; Shi, T.; Hao, N.; Liu, Q.; Wang, K. Facile one-pot synthesis of visible light-responsive BiPO4/nitrogen doped graphene hydrogel for fabricating label-free photoelectrochemical tetracycline aptasensor. Biosens. Bioelectron. 2018, 111, 131–137. [Google Scholar] [CrossRef]
- Liu, X.; Liu, P.; Tang, Y.; Yang, L.; Li, L.; Qi, Z.; Li, D.; Wong, D.K.Y. A photoelectrochemical aptasensor based on a 3D flower-like TiO2-MoS2-gold nanoparticle heterostructure for detection of kanamycin. Biosens. Bioelectron. 2018, 112, 193–201. [Google Scholar] [CrossRef]
- Xu, X.; Liu, D.; Luo, L.; Li, L.; Wang, K.; You, T. Photoelectrochemical aptasensor based on CdTe quantum dots-single walled carbon nanohorns for the sensitive detection of streptomycin. Sens. Actuators B 2017, 251, 564–571. [Google Scholar] [CrossRef]
- Li, Y.; Tian, J.; Yuan, T.; Wang, P.; Lu, J. A sensitive photoelectrochemical aptasensor for oxytetracycline based on a signal “switch off-on” strategy. Sens. Actuators B 2017, 240, 785–792. [Google Scholar] [CrossRef]
- Okoth, O.K.; Yan, K.; Liu, Y.; Zhang, J. Graphene-doped Bi2S3 nanorods as visible-light photoelectrochemical aptasensing platform for sulfadimethoxine detection. Biosens. Bioelectron. 2016, 86, 636–642. [Google Scholar] [CrossRef]
- Cheng, S.; Liu, H.; Zhang, H.; Chu, G.; Guo, Y.; Sun, X. Ultrasensitive electrochemiluminescence aptasensor for kanamycin detection based on silver nanoparticle-catalyzed chemiluminescent reaction between luminol and hydrogen peroxide. Sens. Actuators B 2020, 304, 127367. [Google Scholar] [CrossRef]
- Li, S.; Liu, C.; Yin, G.; Zhang, Q.; Luo, J.; Wu, N. Aptamer-molecularly imprinted sensor base on electrogenerated chemiluminescence energy transfer for detection of lincomycin. Biosens. Bioelectron. 2017, 91, 687–691. [Google Scholar] [CrossRef] [PubMed]
- Miao, Y.-B.; Ren, H.-X.; Gan, N.; Zhou, Y.; Cao, Y.; Li, T.; Chen, Y. A triple-amplification SPR electrochemiluminescence assay for chloramphenicol based on polymer enzyme-linked nanotracers and exonuclease-assisted target recycling. Biosens. Bioelectron. 2016, 86, 477–483. [Google Scholar] [CrossRef]
- Feng, X.; Gan, N.; Zhang, H.; Yan, Q.; Li, T.; Cao, Y.; Hu, F.; Yu, H.; Jiang, Q. A novel “dual-potential” electrochemiluminescence aptasensor array using CdS quantum dots and luminol-gold nanoparticles as labels for simultaneous detection of malachite green and chloramphenicol. Biosens. Bioelectron. 2015, 74, 587–593. [Google Scholar] [CrossRef]
- Feng, X.; Gan, N.; Lin, S.; Li, T.; Cao, Y.; Hu, F.; Jiang, Q.; Chen, Y. Ratiometric electrochemiluminescent aptasensor array for antibiotic based on internal standard method and spatial-resolved technique. Sens. Actuators B 2016, 226, 305–311. [Google Scholar] [CrossRef]
- Gai, P.; Gu, C.; Hou, T.; Li, F. Ultrasensitive Self-Powered Aptasensor Based on Enzyme Biofuel Cell and DNA Bioconjugate: A Facile and Powerful Tool for Antibiotic Residue Detection. Anal. Chem. 2017, 89, 2163–2169. [Google Scholar] [CrossRef]
- Vasilescu, A.; Hayat, A.; Gáspár, S.; Marty, J.-L. Advantages of Carbon Nanomaterials in Electrochemical Aptasensors for Food Analysis. Electroanalysis 2018, 30, 2–19. [Google Scholar] [CrossRef]
- Zhou, L.; Li, D.-J.; Gai, L.; Wang, J.-P.; Li, Y.-B. Electrochemical aptasensor for the detection of tetracycline with multi-walled carbon nanotubes amplification. Sens. Actuators B 2012, 162, 201–208. [Google Scholar] [CrossRef]
- Golkarieh, A.-M.; Nasirizadeh, N.; Jahanmardi, R. Fabrication of an electrochemical sensor with Au nanorods-graphene oxide hybrid nanocomposites for in situ measurement of cloxacillin. Mater. Sci. Eng. C 2021, 118, 111317. [Google Scholar] [CrossRef]
- Cao, L.; Zhang, Q.; Dai, H.; Fu, Y.; Li, Y. Separation/Concentration-signal-amplification in-One Method Based on Electrochemical Conversion of Magnetic Nanoparticles for Electrochemical Biosensing. Electroanalysis 2018, 30, 517–524. [Google Scholar] [CrossRef]
- Wang, C.; Liu, C.; Luo, J.; Tian, Y.; Zhou, N. Direct electrochemical detection of kanamycin based on peroxidase-like activity of gold nanoparticles. Anal. Chim. Acta 2016, 936, 75–82. [Google Scholar] [CrossRef] [PubMed]
- Daprà, J.; Lauridsen, L.H.; Nielsen, A.T.; Rozlosnik, N. Comparative study on aptamers as recognition elements for antibiotics in a label-free all-polymer biosensor. Biosens. Bioelectron. 2013, 43, 315–320. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Wang, X.; Sun, X.; Guo, Y. Multiplex electrochemical aptasensor for detecting multiple antibiotics residues based on carbon fiber and mesoporous carbon-gold nanoparticles. Sens. Actuators B 2018, 265, 217–226. [Google Scholar] [CrossRef]
- Qian, M.; Hu, W.; Wang, L.; Wang, Y.; Dong, Y. A Non-Enzyme and Non-Label Sensitive Fluorescent Aptasensor Based on Simulation-Assisted and Target-Triggered Hairpin Probe Self-Assembly for Ochratoxin a Detection. Toxins 2020, 12, 376. [Google Scholar] [CrossRef]
- Wang, Y.; Yao, L.; Ning, G.; Wu, Y.; Wu, S.; Mao, S.; Liu, G.-Q. An electrochemical strategy for tetracycline detection coupled triple helix aptamer probe with catalyzed hairpin assembly signal amplification. Biosens. Bioelectron. 2019, 143, 111613. [Google Scholar] [CrossRef]
- Leonardo, S.; Toldrà, A.; Campàs, M. Biosensors Based on Isothermal DNA Amplification for Bacterial Detection in Food Safety and Environmental Monitoring. Sensors 2021, 21, 602. [Google Scholar] [CrossRef] [PubMed]


| Nanomaterials (NMs) | |||||
|---|---|---|---|---|---|
| Main Function | Metallic NM | Metal Compound NM | Non-Metallic NM | Carbon NMs | Nanostructures |
| Carrier | AuNPs | QDs | SiO2 NPs | Graphene, CNTs | DNA nanostructures |
| Enhancer | AuNPs, AgNPs | CuO NPs | Polyaniline NPs | CNTs | |
| Reporter | AuNPs, AgNPs | QDs, UCNPs | CDs | ||
| Quencher | AuNPs | Graphene, CNTs | |||
| Catalyst | PtNPs | CuO NPs | |||
| Reporter | Metal nanoclusters | ||||
| Separator | Fe3O4 NPs | ||||
| Electrode@NMs | Analyte | Food Product | Detection Technique | LOD (ng/mL) | References |
|---|---|---|---|---|---|
| GCE@AuNPs/MD-BSA/Ab1 + hemin@MOFs/AuPt-Ab2-HRP + HRP | Maduramicin | Eggs | Amperometric | 0.045 ng/mL | [52] |
| GCE@rGO/AgNPs | CAP | Milk | Amperometric | 0.65 ng/mL | [53] |
| Au bar a@AuNPs/cDNA (S2) + Apt-MOFs (F-) + Au bar b-cDNA (S3) + FSE | Kanamycin, CAP | Milk, fish | Potentiometric | 0.17 ng/mL, 0.15 ng/mL | [54] |
| GNPE@Apt + DNase I | Kanamycin | Milk | Potentiometric | 0.00003 ng/mL | [55] |
| ECNF@AuNPs-Apt | Penicillin | Milk | CV | 0.6 ng/mL | [56] |
| SPCE@4-CP-Apt | Tetracycline | Water | CV | 0.035 ng/mL | [57] |
| SPE@IL/Fe3O4/Apt | Tetracycline | Milk | CV | 0.44 ng/mL | [58] |
| SPCE@CS-MWCNT/CS-Fe3O4/Apt | Tetracycline | Milk | CV | 0.44 ng/mL | [59] |
| SPGE@cDNA1/cDNA2 + MB-Apt-MB + [Fe(CN)6]3−/4− | Ciprofloxacin | Milk | DPV | 0.033 ng/mL | [60] |
| AuE@S3/S4/Apt/S1/S2/MB/AuNP/HRP | Kanamycin | Milk, honey | DPV | 0.0000091 ng/mL | [61] |
| AuE@PEI-rGO/AuNCs-Apt | CAP | Chicken muscle | DPV | 0.67 ng/mL | [62] |
| GCE@GO-f-/[Si-NH2]/AgNPs-Apt + [Fe(CN)6]3−/4− | CAP | Honey, milk | DPV | 0.001 ng/mL | [63] |
| GCE@CS-rGO-AuNPs-MIP + KAN + Au@Fe3O4/SH-β-CD-Fc-Apt | Kanamycin | Milk | DPV | 0.91 ng/mL | [64] |
| AuE@AD/TS + Apt-Primer + DW | Ampicillin | Water | DPV | 0.0003 ng/mL | [34] |
| GCE@ GO-Fe3O4/MWCNT-Fe3O4/PtTi/Apt | Penicillin | Milk | DPV | 0.0253 ng/mL | [65] |
| SPCE@Au-APt | Streptomycin | Milk | DPV | 0.0026 ng/mL | [28] |
| MBs-Ampi + GCE + biotinylated-Apt + SA-HRP + MWCNT | Ampicillin | Milk | DPV | 0.00003 ng/mL | [66] |
| GCE@GO/ZnO | CAP | Milk, honey | DPV | 3.23 ng/mL | [67] |
| AuE@captDNA + Apt-Primer + ExoIII + Hemin + H2O2 | Kanamycin | Milk | DPV | 0.00004 ng/mL | [68] |
| AuE@captDNA + Apt-Primer + ExoIII + Hemin + H2O2 | Ampicillin | Milk | DPV | 0.027 ng/mL | [35] |
| GCE@cDNA/Apt + DpnII + Exo III + [Fe(CN)6]3−/4− | Ampicillin | Milk | DPV | 0.011 ng/mL | [69] |
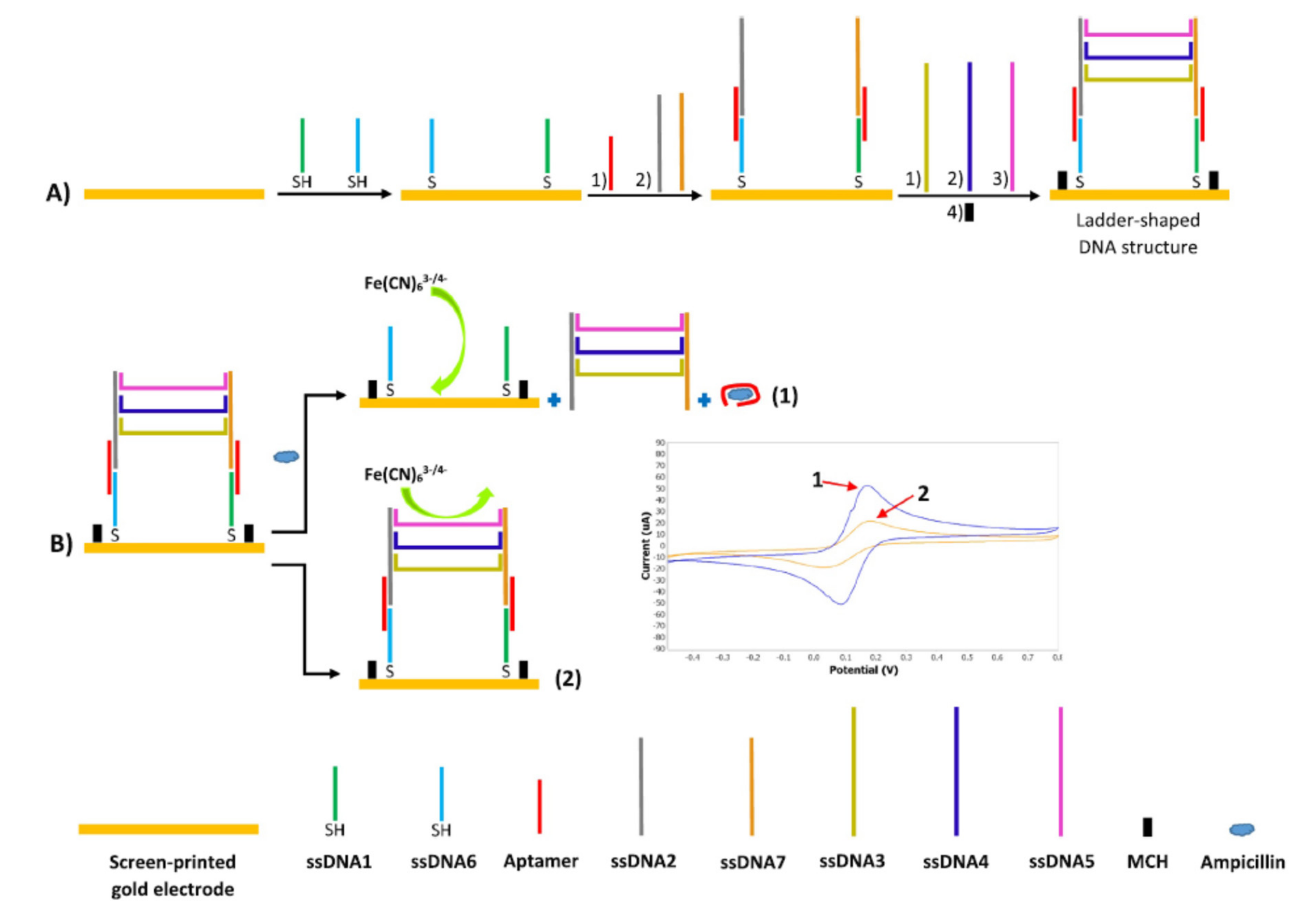
| SPGE@ssDNA1/ssDNA6/Apt/ssDNA2/ssDNA7/ssDNA3/ssDNA4/ssDNA5 + [Fe(CN)6]3−/4− | Ampicillin | Milk | DPV | 0.00035 ng/mL | [32] |
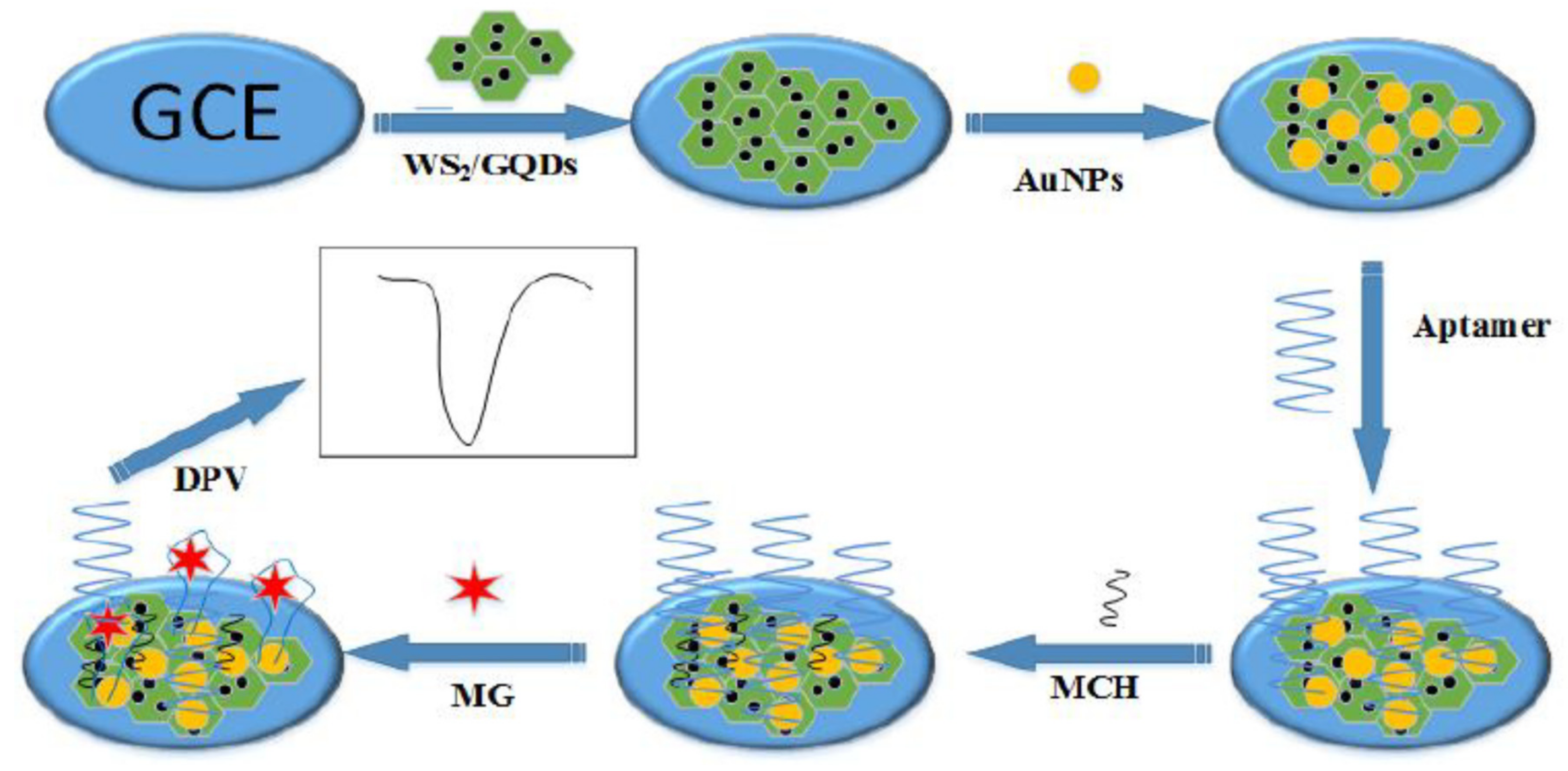
| GCE@MCH/Apt/AuNPs/GQDs-WS2/ | Malachite green | Fish | DPV | 1.11 ng/mL | [31] |
| AuE@cDNA/Apt + Nuclease P1 + Ab anti-dsDNA + [Fe(CN)6]3−/4− | SDMX | Milk | DPV | 0.01 ng/mL | [70] |
| GCE@Pt–Sn@TiO2/cDNA/Apt + RecJf exonuclease | Streptomycin | Milk | DPV | 0.01 ± 0.003 ng/mL | [71] |
| TFGE@Apt + cDNA/MWCNT/AuNPs + thionine | OTC | Chicken meat | DPV | 3.1 × 10−5 ng/mL | [72] |
| GCE/OMC-CS/AuNPs-SA/Biot-cDNA1/Fc-DNA2 (Apt) | Kanamycin | Milk | DPV | 0.017 ng/mL | [44] |
| AuE@Apt-cDNA + C60-rGO/Tb/AuNPs + ExoNase | SDMX | Milk | DPV | 10 × 10−6 ng/mL | [73] |
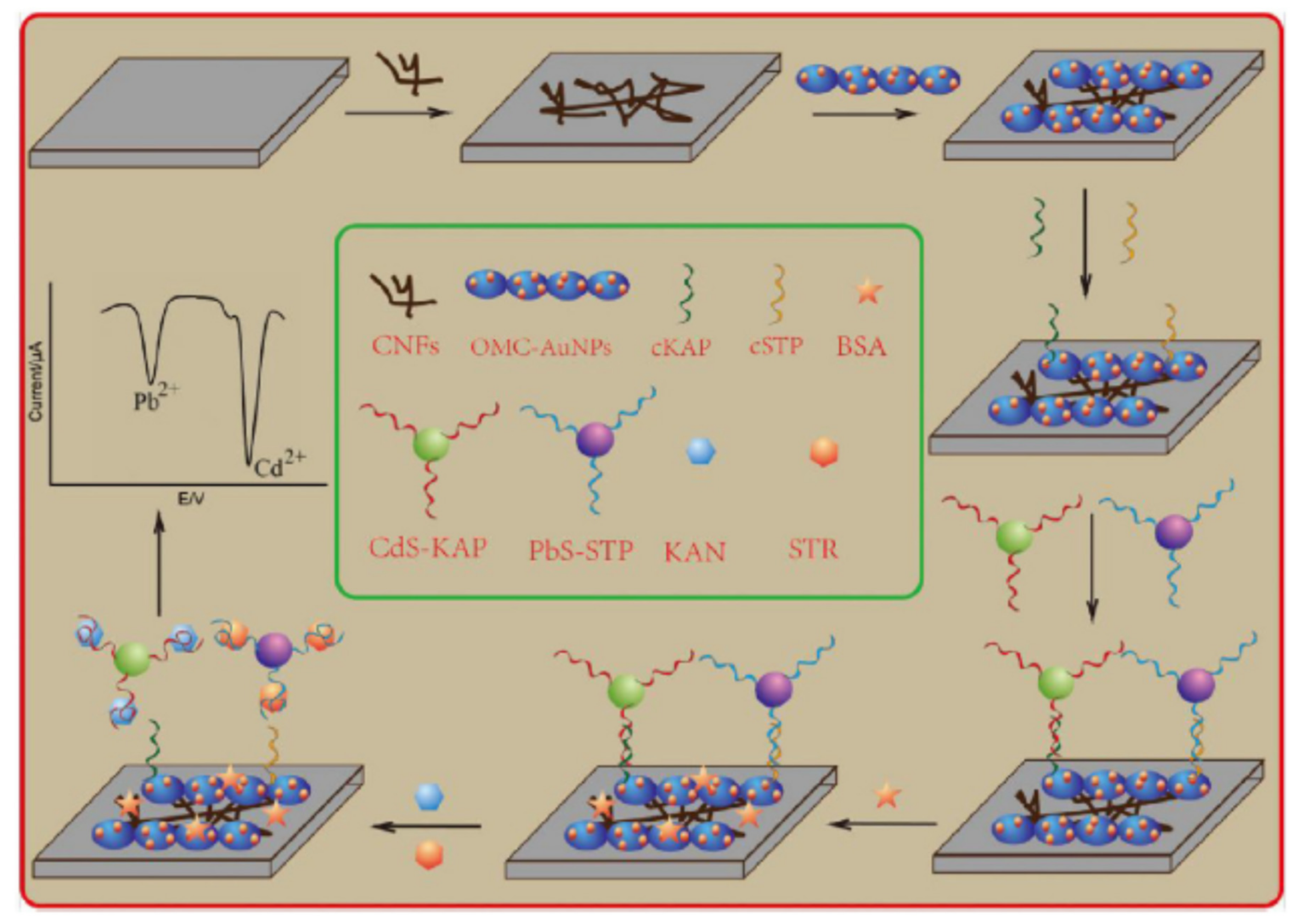
| E@AuNPsCNFs/MWCNTs/cDNA-KAN/polyA/cDNA-STR + CdS-AptKAN + PbS/AptSTR | Kanamycin, streptomycin | Milk | DPV | 0.036 ng/mL 0.021 ng/mL | [74] |
| AuE@cDNA + biotinylated Apt + MB + AuNPs/SA/HRP + H2O2 | Kanamycin | Milk | DPV | 0.00088 ng/mL | [75] |
| AuE1 and AuE2/S1-S2(Apt)/S3/MB | Kanamycin | Milk | DPV | 7.8 × 10−6 ng/mL | [76] |
| AuE@HP2 + Apt-primer cDNA + Exo I + HP1 + nb.BbvCI + Hemin/K++ H2O2 | Kanamycin | Milk | DPV | 0.00024 ng/mL | [36] |
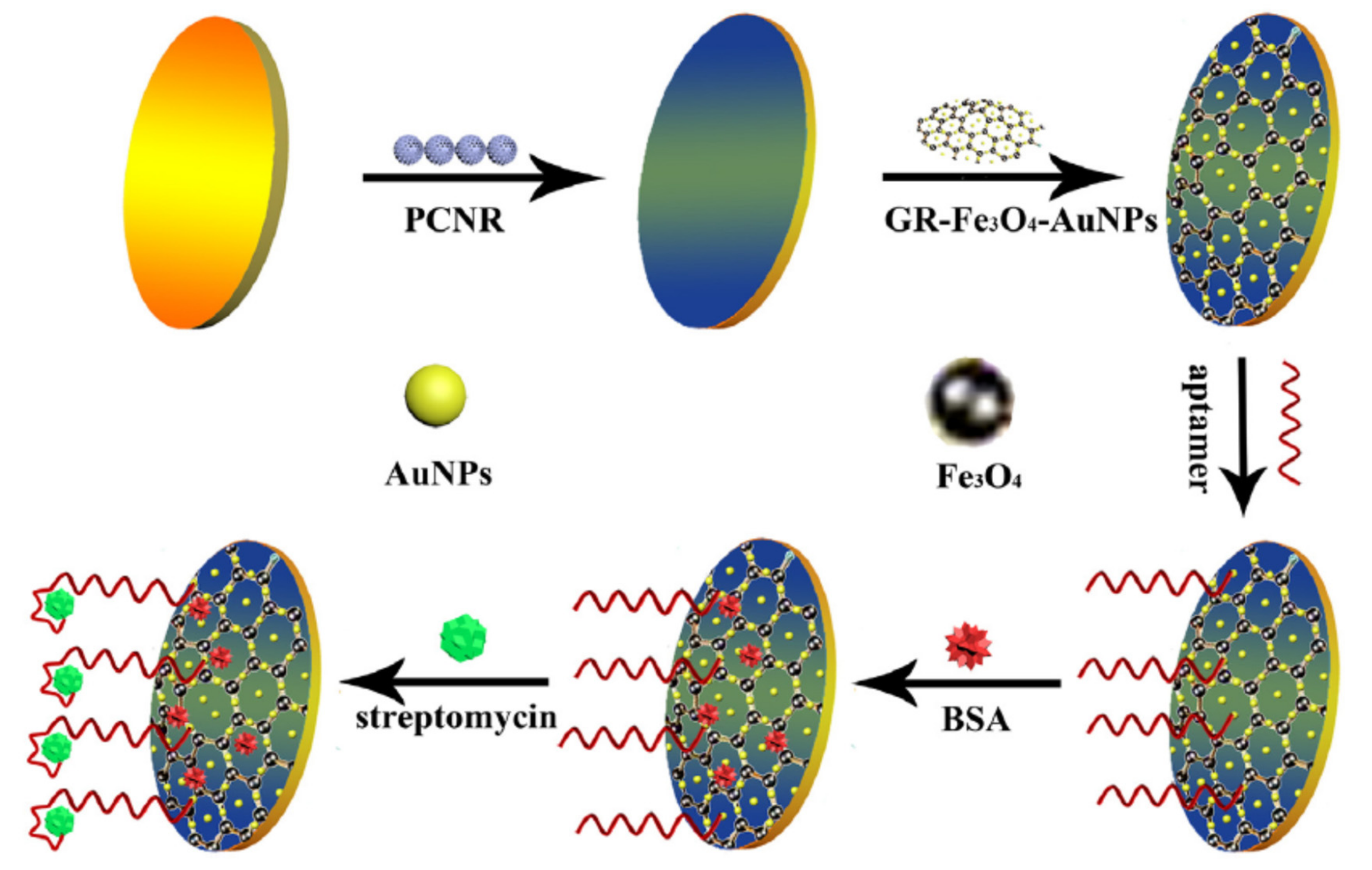
| GCE@PCNR/GR–Fe3O4–AuNPs/Apt | Streptomycin | milk | DPV | 0.028 ng/mL | [77] |
| AuE@cDNA1/cDNA2/cDNA3/Apt +Exo I + [Fe(CN)6]3−/4− | Tetracycline, doxycyline | Milk | DPV | 0.2 ng/mL | [78] |
| GCE@PEDOT–AuNPs/GR–Fe3O4NPs/Apt | Penicillin | Milk | DPV | 0.057 ng/mL | [79] |
| AuE@cDNA/Apt + Exo I + [Fe(CN)6]3−/4− | Streptomycin | Milk | DPV | 8.2 ng/mL | [80] |
| GCE@PCNR/MWCNT-CuO-AuNPs/Apt | Streptomycin | Milk, honey | DPV | 0.036 ng/mL | [81] |
| SPGE@GelB-Apt | CAP | Milk | DPV | 0.059 ng/mL | [82] |
| AuE@Apt + Biotinylated detection DNA + ST-AP | CAP | Honey | DPV | 0.29 ng/mL | [83] |
| GCE + Au bar a@AuNPs/S1/S2 5AptKAN)-S’1/S’2 (AptAMP) + cDNAKAN (S4)/Apo (Pb) + cDNAAMP (S’4)/Apo (Cd) + Au bar b@AuNPs/S3 | Kanamycin, ampicillin | Milk, fish | SWV | 8.7 × 10−6 ng/mL, 0.00001 ng/mL | [84] |
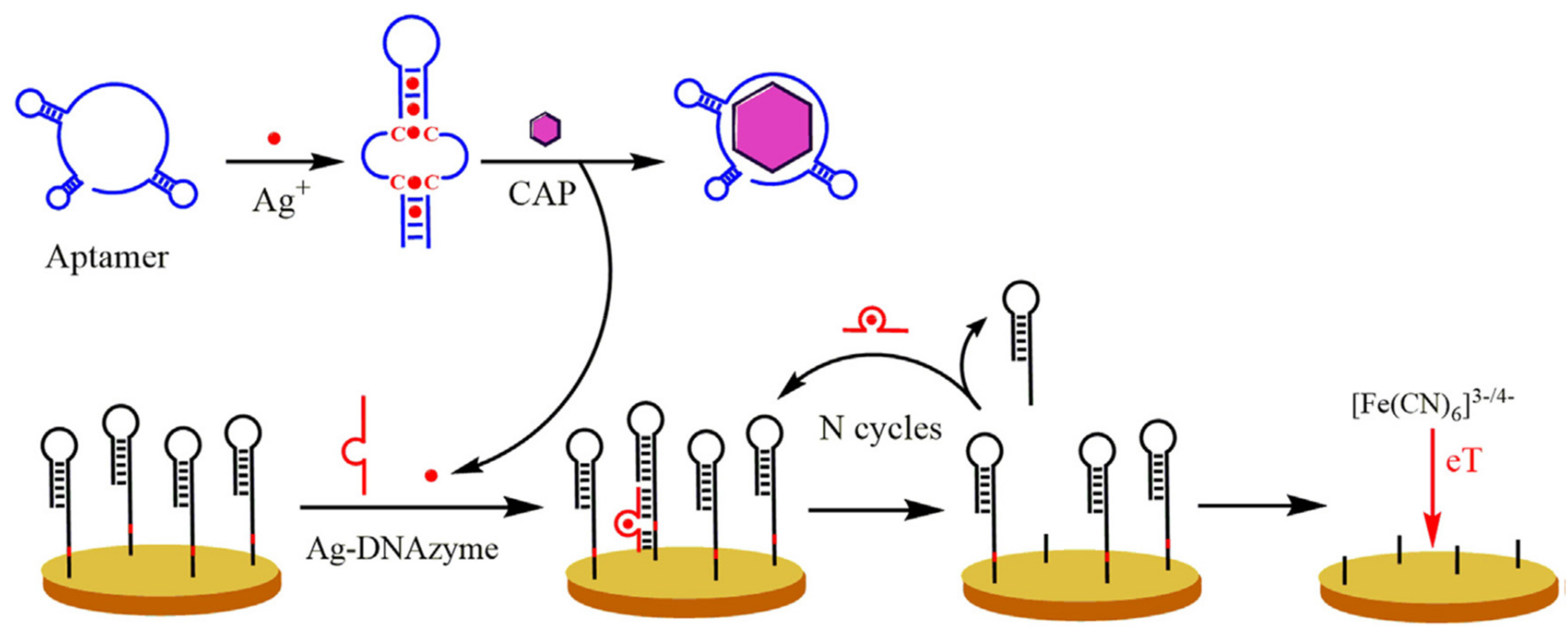
| AuE@HP (DNAzyme) + Apt/Ag + [Fe(CN)6]3−/4− | CAP | Milk | SWV | 0.0004 ng/mL | [37] |
| Au bar a@Apt1/cDNA1/AuNPs-EcoR1-Apt2/cDNA2/AuNPs-BamH1 + Exo I + Au bar b-dsDNA1&2/MOFs (MB, Fc) | Kanamycin, CAP | Milk | SWV | 0.0000170 ng/mL, 0.0000068 ng/mL | [85] |
| GCE@MB-Apt Probe cDNA, M-NMOF (Pb2+, Cd2+) | Kanamycin, CAP | Milk | SWV | 0.0000775 ng/mL, 0.0000061 ng/mL | [86] |
| GCE + MGNPs@aDNA/CaptDNA1&2 (=Apt + primer) + cDNA-MOFs (sDNA1 Pb, sDNA2 Cd) | CAP, OTC | Milk | SWV | 0.0000107 ng/mL, 0.000022 ng/mL | [87] |
| GCE@S3/S4 + Apt-CAP + Apt-OTC + S1-MOF (Cd) + S2-MOF (Pb) + Exo I | CAP, OTC | Milk | SWV | 0.15 and 0.10 ng/mL | [88] |
| AuE@Apt | CAP | Milk | SWV | 0.52 ng/mL | [89] |
| AuE@CaptDNA + 3 Apt (STR, CAP, TTC) + 3 cDNA1s + 3 QD–cDNA2 (PbS, CdS, ZnS NCs) + HNO3 | Streptomycin, CAP, TTC | Milk | SWASV | 5.82 ng/mL, 1.62 ng/mL, 8.89 ng/mL | [90] |
| AuE@POP-Apt | Ampicillin | Milk | EIS | 0.00000133 ng/mL | [91] |
| GCE@MWCNT/GO/Apt | Tetracycline | Milk | Impedimetric | 0.0000212 ng/mL | [92] |
| PGE@AuNPs/RGO/Apt | Tetracycline | Milk | EIS | 0.000000017 ng/mL | [93] |
| GCE@CB/CS-Apt | Kanamycin | Milk, yoghurt | EIS | 0.15 ng/mL | [94] |
| AuE@Ag(I) MOFs/Apt + [Fe(CN)6]3−/4− | Penicillin | Milk | EIS | 0.000849 ng/mL | [95] |
| AuE@Ce-MOF/MCA/Apt | OTC | Milk | EIS | 17.4 × 10−6 ng/mL | [96] |
| AuE@SWCNT-cDNA + Apt | OTC | Water | Impedimetric | 1.125 ng/mL | [97] |
| AuE@Ce-MOF/MCA-Apt | OTC | Milk | EIS | 0.00000037 ng/mL | [96] |
| PGE@rGO/AuNPs-Apt | Penicillin G | Milk | EIS | 0.00000027 ng/mL | [98] |
| PGE@rGO/AuNPs-Apt | SDMX | Fish, chicken, beef | EIS | 0.000000115 ng/mL | [99] |
| SPCE@MWCNT-V2O5-CS-Apt + [Fe(CN)6]4−/3− | Ciprofloxacin | Milk | EIS | 0.5 ng/mL | [100] |
| AuE@TPN-COF@Co-MOF-Apt | Ampicillin | Milk | EIS | 0.217 × 10−6 ng/mL | [101] |
| SPCE@4-CP-Apt + [Fe(CN)6]4−/3− | Kanamycin | Milk | EIS | 0.11 ng/mL | [45] |
| SPCE@4-NB-Apt + [Fe(CN)6]4−/3− | Penicillin G | Milk | EIS | 0.17 ng/mL | [102] |
| IDAM@CS/TiO2 NPs | Tetracycline | Milk | EIS | 3 ng/mL | [103] |
| AuE@Apt + [Fe(CN)6]4−/3− | CAP | / | EIS | 0.57 ng/mL | [104] |
| Electrode@NMs | Analyte | Food Product | Detection Technique | LOD (ng/g or nM) | References |
|---|---|---|---|---|---|
| ITO@ZIS-HNCs/CS-Apt | Lincomycin | Milk | PEC | 0.000034 ng/mL | [105] |
| ITO@AuNPs/sDNA/Fc-rp/Fc-Apt + HP1-HP2 + MB | Streptomycin | Water | PEC&EC dual-mode | 10.06 ng/mL | [106] |
| ITO@SnO2/Bi2S3-Apt | Tobramycin | Milk | PEC | 2 ng/mL | [107] |
| ITO@ZnxCo3-xO4/N-GQDs/AgBiS2-Apt | Ampicillin | Water | PEC | 0.00009 ng/mL | [108] |
| ITO@g-CN/Co3O4-Apt | OTC | Water | PEC | 0.0016 ng/mL | [109] |
| ITO@BiOI/Au-Apt | Tetracycline | PEC | 0.0002 ng/mL | [110] | |
| GCE@Bi-BiOI@C/Apt | CAP | / | PEC | 0.26 ng/mL | [111] |
| FTO@BR-CN/cDNA/biotinylatedApt + SAPtNi + H2O2 | CAP | Milk | PEC | 0.0000084 ng/mL | [112] |
| ITO@MIL−68(In)-NH2/MWCNT/CdS/Apt | TTC | Water | PEC | 0.0067 ng/mL | [113] |
| ITO@Bi2S3/NGQDs/CS/GA/Apt | SDMX | Milk | PEC | 0.009 ng/mL0 | [114] |
| ITO@BiFeO3/utg-C3N4-Apt | Ampicillin | Milk | PEC | 0.0000115 ng/mL | [115] |
| ITO@Bi2MoO6/BNG-Apt | Lincomycin | Milk | PEC | 0.0015 ng/mL | [116] |
| TiE@ TiO2 NTs/GQDs | CAP | Honey | PEC | 0.019 ng/mL | [117] |
| TiE@ TiO2 NTs/Ag2S/PDA-Apt | Ofloxacin | Milk | PEC | 0.00027 ng/mL | [118] |
| FTO@TiO2NRA/Eu3+ QDs (CdS)-Apt | CAP | Milk | PEC | 0.00012 ng/mL | [119] |
| ITO@WS2-Apt + DNase | CAP | Milk powder | PEC | 0.0012 ng/mL | [120] |
| ITO@BiPO4/3DNGH-Apt | Tetracycline | Milk | PEC | 0.0147 ng/mL | [121] |
| ITO@TiO2-MoS2-AuNPs-Apt | Kanamycin | Milk | PEC | 0.024 ng/mL | [122] |
| ITO@CdTe QDs/SWCNHs-Apt | Streptomycin | Honey | PEC | 0.02 ng/mL | [123] |
| ITO@TiO2/H-DNA@CdTe QDs-Apt | OTC | Chicken, raw milk | PEC | 0.09 ng/mL | [124] |
| FTO@G-Bi2S3-Apt | SDMX | Milk | PEC | 0.17 ng/mL | [125] |
| PE@Luminol/H2O2/AgNPs-Apt | Kanamycin | Milk | ECL | 0.06 ng/mL | [126] |
| GCE@Au-GO + CDots-Apt + MIP | Lincomycin | Meat | ECL | 0.000065 ng/mL | [127] |
| GCE@CdS NCs/Apt-ssDNA/EV (HRP)-Au-SSB + ExoI | CAP | Fish | SPR-ECL | 0.000011 ng/mL | [128] |
| WE1: SPCE@CdS QDs-MG Apt-Cy5WE2: SPCE@ L-AuNPs-CAP Apt-CA | MG, CAP | Fish | ECL | 0.023 ng/mL MG, 0.01 ng/mL CAP | [129] |
| WE1: SPCE@L-AuNPs/cDNA/CA/AptWE2: SPCE@CdS QDs | CAP | Fish | R-ECL | 0.01 ng/mL | [130] |
| CPE@AuNPs/GOx-Apt−csDNA/AuNPs/SiO2 + CPE@AuNPs/PDA/Laccase + glucose | Ampicillin | Milk | EBFCs | 0.7 ng/mL | [131] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gaudin, V. Contribution of Nanomaterials to the Development of Electrochemical Aptasensors for the Detection of Antimicrobial Residues in Food Products. Chemosensors 2021, 9, 69. https://doi.org/10.3390/chemosensors9040069
Gaudin V. Contribution of Nanomaterials to the Development of Electrochemical Aptasensors for the Detection of Antimicrobial Residues in Food Products. Chemosensors. 2021; 9(4):69. https://doi.org/10.3390/chemosensors9040069
Chicago/Turabian StyleGaudin, Valérie. 2021. "Contribution of Nanomaterials to the Development of Electrochemical Aptasensors for the Detection of Antimicrobial Residues in Food Products" Chemosensors 9, no. 4: 69. https://doi.org/10.3390/chemosensors9040069
APA StyleGaudin, V. (2021). Contribution of Nanomaterials to the Development of Electrochemical Aptasensors for the Detection of Antimicrobial Residues in Food Products. Chemosensors, 9(4), 69. https://doi.org/10.3390/chemosensors9040069






