Enzymes as Tools in MIP-Sensors
Abstract
:1. Introduction
- Enzyme-initiated polymerization has been introduced as a green alternative to the well-established chemical polymerization and electrosynthesis.
- Removal of protein templates has been achieved under mild conditions by proteases, especially proteinase K.
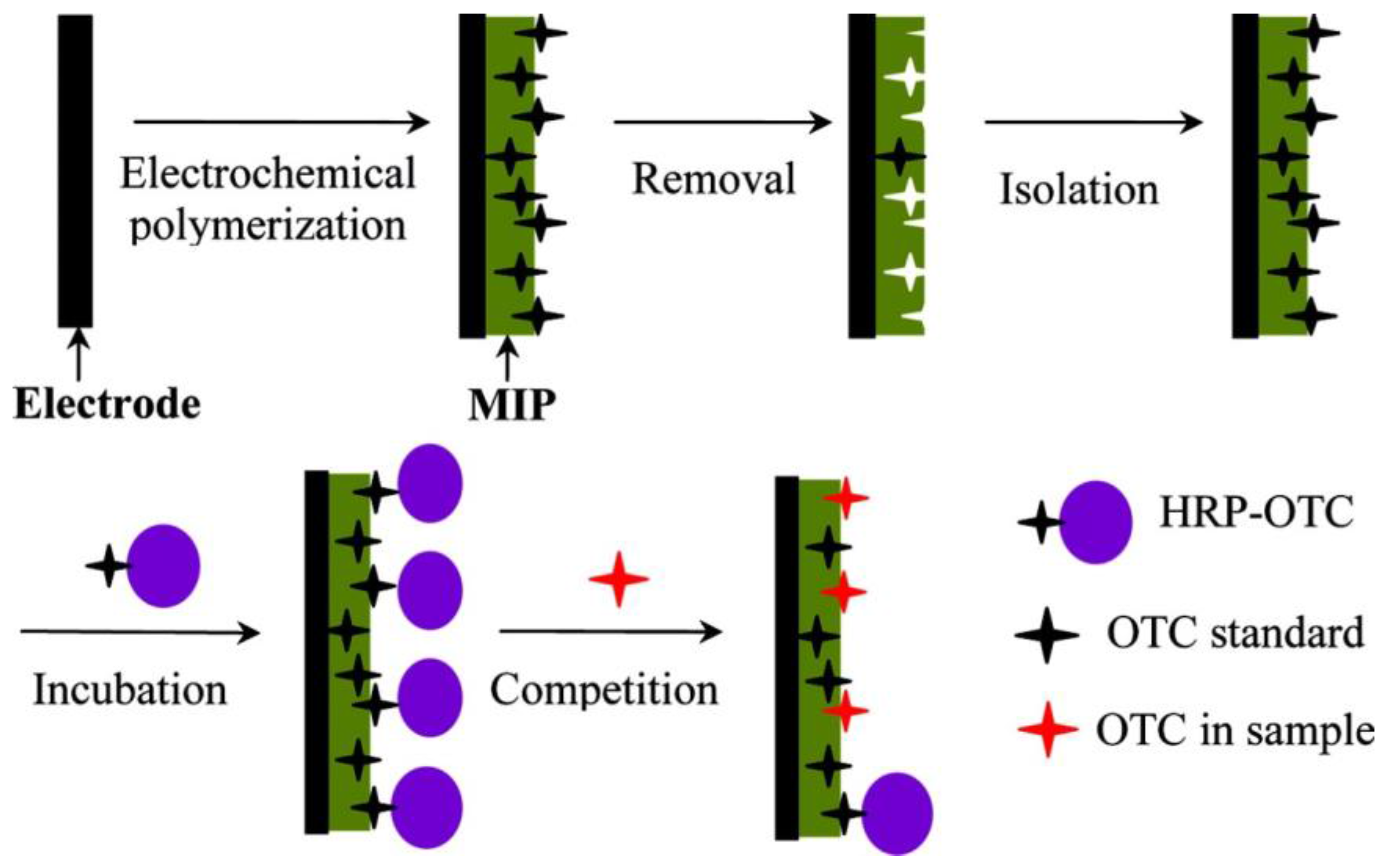
- Enzyme-labeled “tracers” have been used in analogy to competitive immunoassays in MIP sensors.
- The measuring signal of MIP-sensors has been amplified by electro-enzymatic recycling of the redox marker ferricyanide using horseradish peroxidase (HRP).
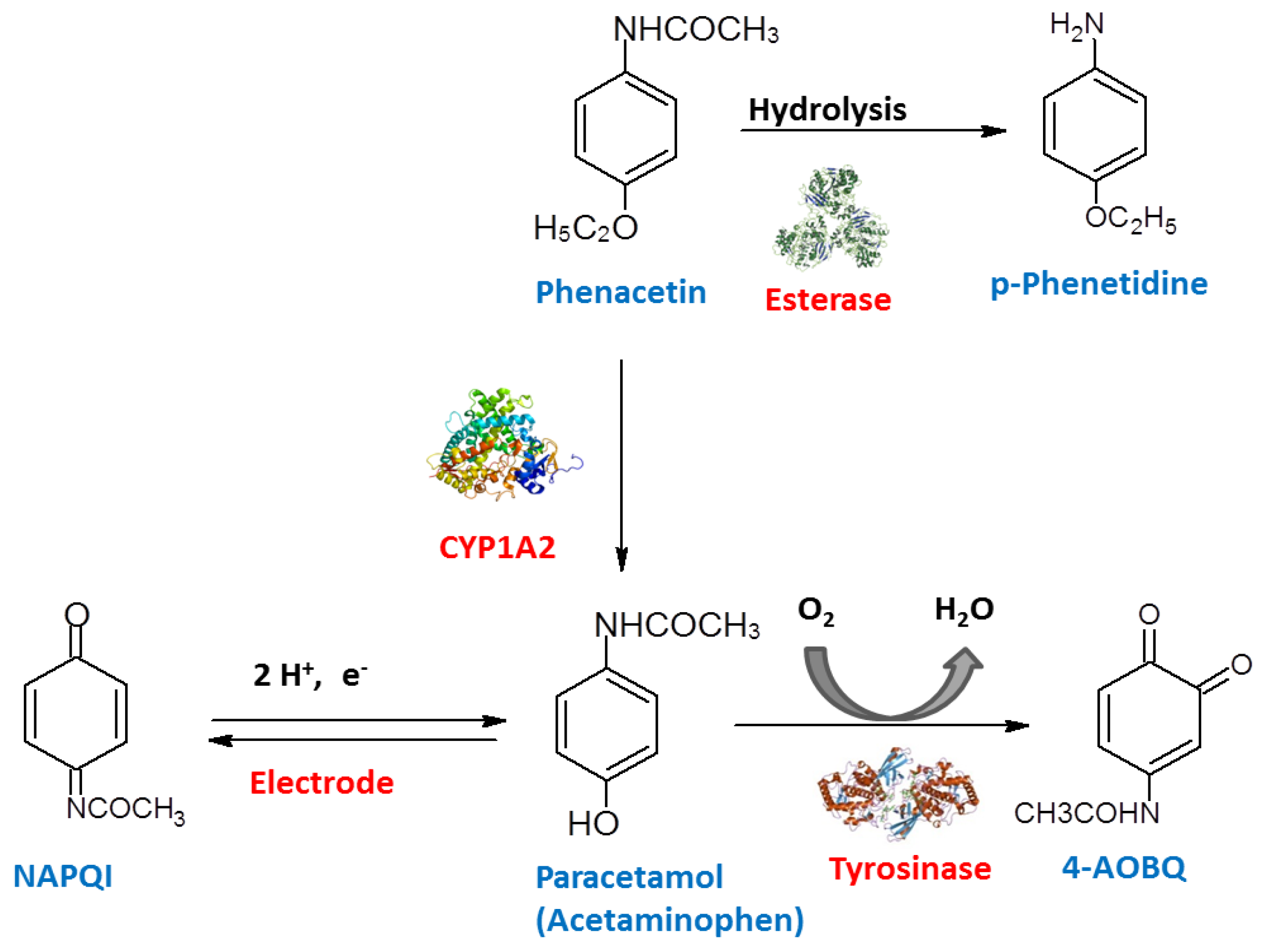
- The enzymatic pretreatment of the analyte allowed the interference-free electrochemical measurement or the conversion of a non-binding analyte into a target analog of the MIP.
2. Enzymes in the Workflow of the Preparation of Surface-Imprinted MIPs
2.1. Preparation of Surface Imprinted MIPs
2.1.1. Photo- and Chemically Initiated Polymerization
2.1.2. Electropolymerization
2.1.3. Self-Polymerization
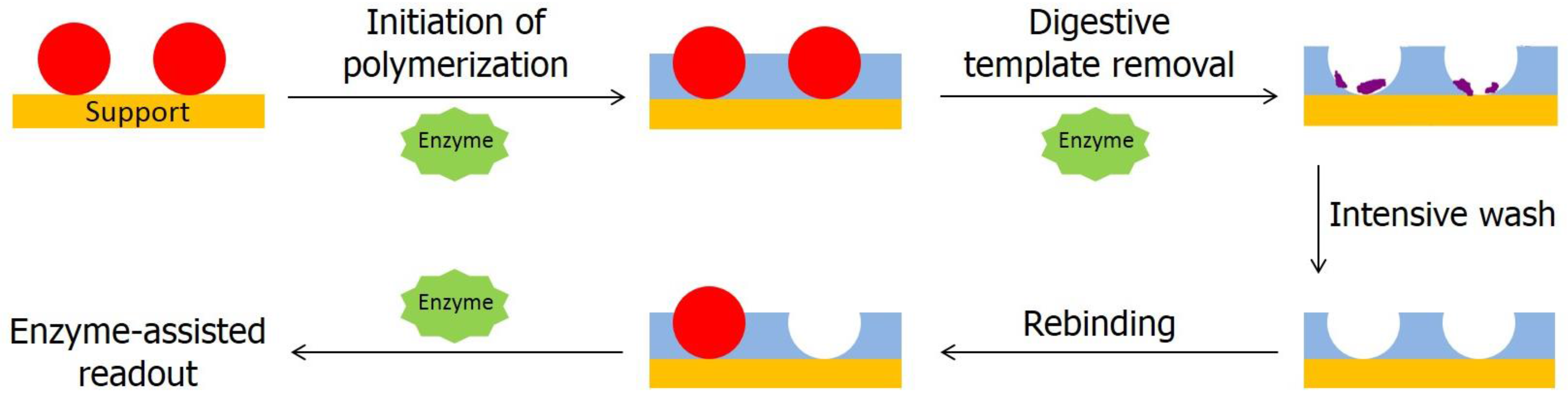
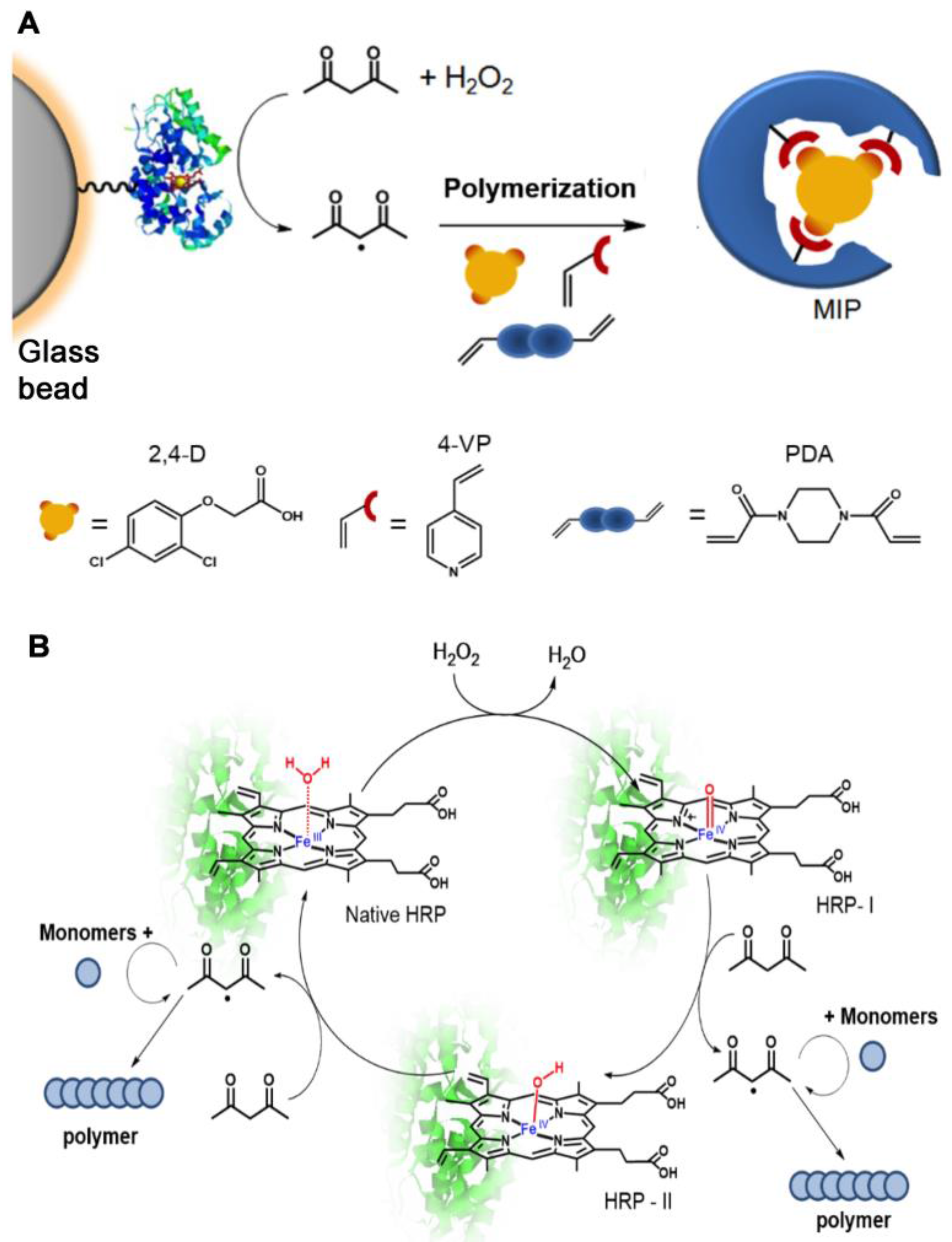
2.1.4. Enzyme-Initiated MIP Synthesis
2.2. Template Removal by Enzymes
3. Enzymes for the Enhancement of the Analytical Performance of MIP Sensors
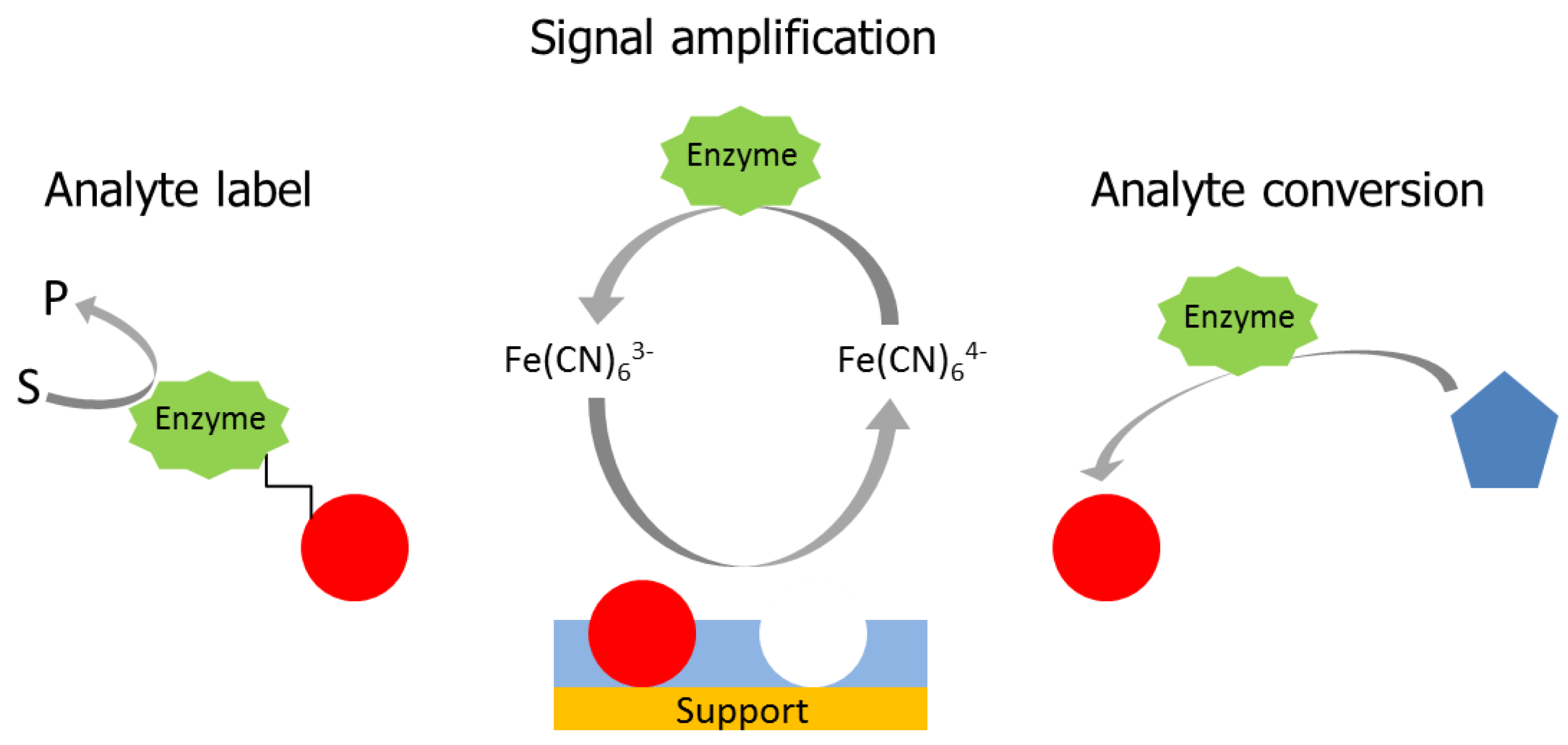
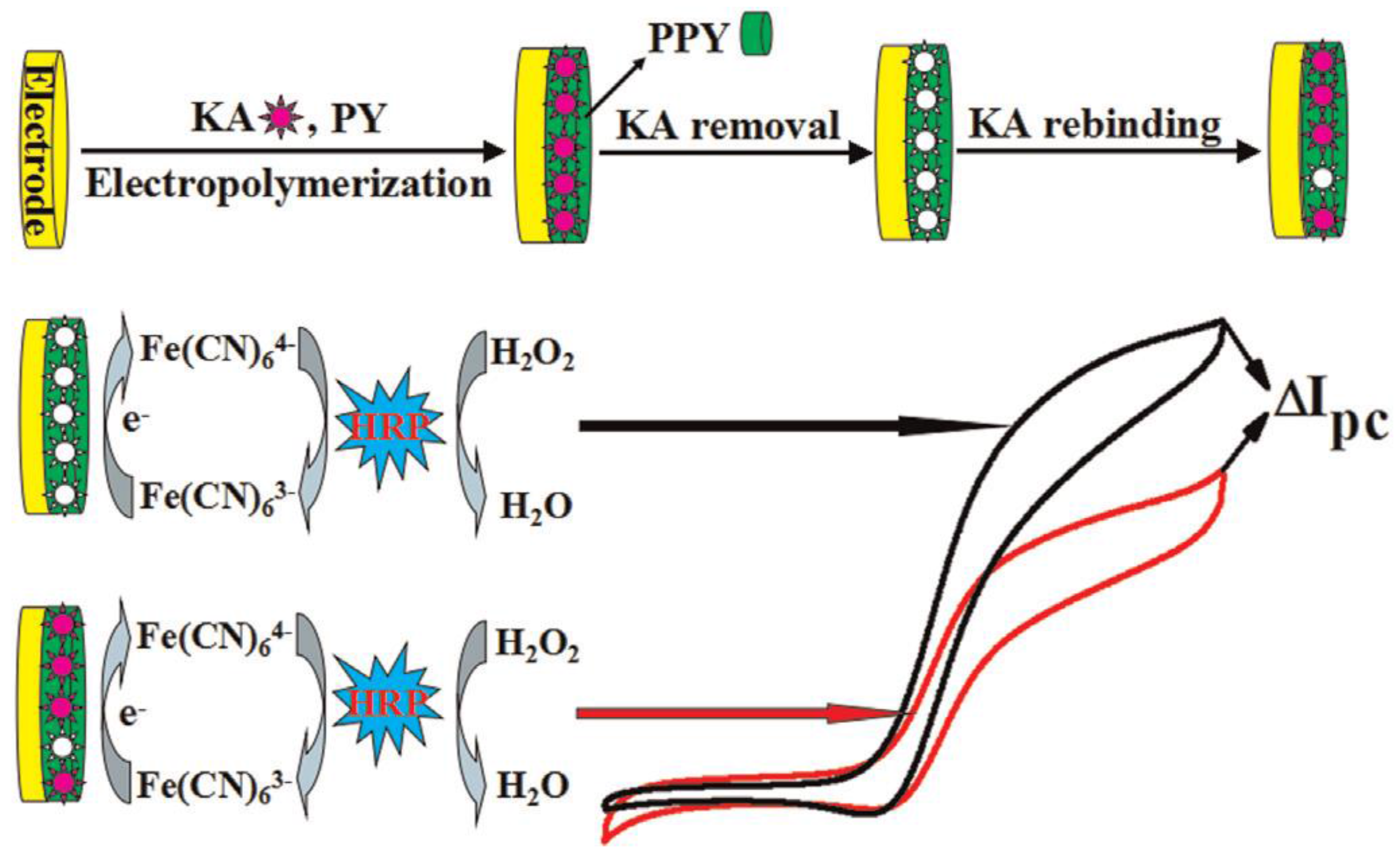
3.1. Signal Amplification in Electrochemical MIP Sensors
- (i)
- Electroactive targets, such as morphine, paracetamol, tamoxifen, and diclofenac can permeate through the cavities of the MIP to the electrode and an electrochemical signal can be generated by the conversion of the target using different electrochemical methods. This principle has been frequently used for drugs which contain phenolic structures but also for a few proteins which show direct electron transfer, e.g., cytochrome c, hemoglobin, and hexameric tyrosine-coordinated heme protein (HTHP) [21,70,71].
- (ii)
- Binding of the target modulates the diffusive permeation of redox markers in a concentration-dependent manner. This effect has been frequently applied to characterize each step of MIP preparation for electro-inactive targets, such as melamine, methyl parathion, phenobarbital, caffeine, 17β-estradiol, acetylsalicylic acid, and warfarin [72,73,74,75,76]. In addition, this method is frequently applied to quantify the binding of the target analyte. However, it supplies an indirect signal which integrates all changes of the MIP-layer. Using this approach, several papers claim measuring ranges over more than four decades of target concentration and lower limits of detection (LOD) in the sub-nanomolar range for both low- [77,78] and high-molecular weight targets [79,80,81]. Rebinding of the target in the pores of the MIP could be strong for small molecules. On the other hand, the film thickness for surface imprinted layers is lower than the dimension of macromolecular targets. Thus, affinity constants for non-covalent MIPs could hardly reach the sub-nanomolar region. From the practical point of view, it seems questionable to evaluate the tiny current decreases per concentration decade of the cyclic and differential pulse voltammograms.
3.2. Enzyme-Labels in MIP-Based Affinity Sensors
3.3. Combinations of MIPs with the Enzymatic Conversion of the Analyte
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Wulff, G. Molecular Imprinting in Cross-Linked Materials with the Aid of Molecular Templates—A Way towards Artificial Antibodies. Angew. Chem. Int. Ed. 1995, 34, 1812–1832. [Google Scholar] [CrossRef]
- Haupt, K.; Mosbach, K. Molecularly imprinted polymers and their use in biomimetic sensors. Chem. Rev. 2000, 100, 2495–2504. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Xu, S.; Li, J. Recent advances in molecular imprinting technology: Current status, challenges and highlighted applications. Chem. Soc. Rev. 2011, 40, 2922. [Google Scholar] [CrossRef] [PubMed]
- Hayden, O.; Lieberzeit, P.A.; Blaas, D.; Dickert, F.L. Artificial antibodies for bioanalyte detection - Sensing viruses and proteins. Adv. Funct. Mater. 2006, 16, 1269–1278. [Google Scholar] [CrossRef]
- Alexander, C.; Andersson, H.S.; Andersson, L.I.; Ansell, R.J.; Kirsch, N.; Nicholls, I.A.; O’Mahony, J.; Whitcombe, M.J. Molecular imprinting science and technology: A survey of the literature for the years up to and including 2003. J. Mol. Recognit. 2006, 19, 106–180. [Google Scholar] [CrossRef] [PubMed]
- Whitcombe, M.J.; Kirsch, N.; Nicholls, I.A. Molecular imprinting science and technology: A survey of the literature for the years 2004–2011. J. Mol. Recognit. 2014, 27, 297–401. [Google Scholar] [PubMed]
- Erdőssy, J.; Horváth, V.; Yarman, A.; Scheller, F.W.; Gyurcsányi, R.E. Electrosynthesized molecularly imprinted polymers for protein recognition. TrAC Trends Anal. Chem. 2016, 79, 179–190. [Google Scholar] [CrossRef]
- Menger, M.; Yarman, A.; Erdőssy, J.; Yildiz, H.; Gyurcsányi, R.; Scheller, F. MIPs and Aptamers for Recognition of Proteins in Biomimetic Sensing. Biosensors 2016, 6, 35. [Google Scholar] [CrossRef] [PubMed]
- Uzun, L.; Turner, A.P.F. Molecularly-imprinted polymer sensors: Realising their potential. Biosens. Bioelectron. 2016, 76, 131–144. [Google Scholar] [CrossRef] [PubMed]
- Daoud Attieh, M.; Zhao, Y.; Elkak, A.; Falcimaigne-Cordin, A.; Haupt, K. Enzyme-initiated Free-Radical Polymerization of Molecularly Imprinted Polymer Nanogels on a Solid Phase with Immobilized Radical Source. Angew. Chem. Int. Ed. 2017, 56, 3339–3343. [Google Scholar] [CrossRef] [PubMed]
- Ambrosini, S.; Beyazit, S.; Haupt, K.; Tse Sum Bui, B. Solid-phase synthesis of molecularly imprinted nanoparticles for protein recognition. Chem. Commun. 2013, 49, 6746. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Ambrosini, S.; Tamahkar, E.; Rossi, C.; Haupt, K.; Tse Sum Bui, B. Toward a universal method for preparing molecularly imprinted polymer nanoparticles with antibody-like affinity for proteins. Biomacromolecules 2016, 17, 345–353. [Google Scholar] [CrossRef] [PubMed]
- Cutivet, A.; Schembri, C.; Kovensky, J.; Haupt, K. Molecularly imprinted microgels as enzyme inhibitors. J. Am. Chem. Soc. 2009, 131, 14699–14702. [Google Scholar] [CrossRef] [PubMed]
- Matsunaga, T.; Hishiya, T.; Takeuchi, T. Surface plasmon resonance sensor for lysozyme based on molecularly imprinted thin films. Anal. Chim. Acta 2007, 591, 63–67. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Zheng, J.; Fang, G.; Xie, W. Improvement of the homogeneity of protein-imprinted polymer films by orientated immobilization of the template. Anal. Chim. Acta 2012, 726, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Zheng, C.; Zhang, X.-L.; Liu, W.; Liu, B.; Yang, H.-H.; Lin, Z.-A.; Chen, G.-N. A Selective Artificial Enzyme Inhibitor Based on Nanoparticle-Enzyme Interactions and Molecular Imprinting. Adv. Mater. 2013, 25, 5922–5927. [Google Scholar] [CrossRef] [PubMed]
- Guerreiro, A.; Poma, A.; Karim, K.; Moczko, E.; Takarada, J.; de Vargas-Sansalvador, I.P.; Turner, N.; Piletska, E.; de Magalhães, C.S.; Glazova, N.; et al. Influence of Surface-Imprinted Nanoparticles on Trypsin Activity. Adv. Healthc. Mater. 2014, 3, 1426–1429. [Google Scholar] [CrossRef] [PubMed]
- Jetzschmann, K.J.; Jágerszki, G.; Dechtrirat, D.; Yarman, A.; Gajovic-Eichelmann, N.; Gilsing, H.-D.; Schulz, B.; Gyurcsányi, R.E.; Scheller, F.W. Vectorially Imprinted Hybrid Nanofilm for Acetylcholinesterase Recognition. Adv. Funct. Mater. 2015, 25, 5178–5183. [Google Scholar] [CrossRef]
- Kamon, Y.; Matsuura, R.; Kitayama, Y.; Ooya, T.; Takeuchi, T. Precisely controlled molecular imprinting of glutathione-s-transferase by orientated template immobilization using specific interaction with an anchored ligand on a gold substrate. Polym. Chem. 2014, 5, 4764. [Google Scholar] [CrossRef]
- Wang, S.; Ye, J.; Bie, Z.; Liu, Z. Affinity-tunable specific recognition of glycoproteins via boronate affinity-based controllable oriented surface imprinting. Chem. Sci. 2014, 5, 1135–1140. [Google Scholar] [CrossRef]
- Peng, L.; Yarman, A.; Jetzschmann, K.J.; Jeoung, J.-H.; Schad, D.; Dobbek, H.; Wollenberger, U.; Scheller, F.W. Molecularly imprinted electropolymer for a hexameric heme protein with direct electron transfer and peroxide electrocatalysis. Sensors 2016, 16, 272. [Google Scholar] [CrossRef] [PubMed]
- Lettau, K.; Warsinke, A.; Katterle, M.; Danielsson, B.; Scheller, F.W. A Bifunctional Molecularly Imprinted Polymer (MIP): Analysis of Binding and Catalysis by a Thermistor. Angew. Chem. Int. Ed. 2006, 45, 6986–6990. [Google Scholar] [CrossRef] [PubMed]
- Wulff, G.; Liu, J. Design of biomimetic catalysts by molecular imprinting in synthetic polymers: The role of transition state stabilization. Acc. Chem. Res. 2012, 45, 239–247. [Google Scholar] [CrossRef] [PubMed]
- Wulff, G.; Vesper, W.; Grobe-Einsler, R.; Sarhan, A. Enzyme-analogue built polymers, 4. On the synthesis of polymers containing chiral cavities and their use for the resolution of racemates. Die Makromol. Chem. 1977, 178, 2799–2816. [Google Scholar] [CrossRef]
- De Jesus Rodrigues Santos, W.; Lima, P.R.; Tarley, C.R.T.; Kubota, L.T. A catalytically active molecularly imprinted polymer that mimics peroxidase based on hemin: Application to the determination of p-aminophenol. Anal. Bioanal. Chem. 2007, 389, 1919–1929. [Google Scholar] [CrossRef] [PubMed]
- Lakshmi, D.; Bossi, A.; Whitcombe, M.J.; Chianella, I.; Fowler, S.A.; Subrahmanyam, S.; Piletska, E.V.; Piletsky, S.A. Electrochemical Sensor for Catechol and Dopamine Based on a Catalytic Molecularly Imprinted Polymer-Conducting Polymer Hybrid Recognition Element. Anal. Chem. 2009, 81, 3576–3584. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Yin, Y.; Liu, Y.; Bai, X.; Zhang, Z.; Xu, J.; Shen, J.; Liu, J. Incorporation of glutathione peroxidase active site into polymer based on imprinting strategy. Biosens. Bioelectron. 2009, 25, 657–660. [Google Scholar] [CrossRef] [PubMed]
- Sode, K.; Ohta, S.; Yanai, Y.; Yamazaki, T. Construction of a molecular imprinting catalyst using target analogue template and its application for an amperometric fructosylamine sensor. Biosens. Bioelectron. 2003, 18, 1485–1490. [Google Scholar] [CrossRef]
- Lohmann, W.; Karst, U. Biomimetic modeling of oxidative drug metabolism. Anal. Bioanal. Chem. 2008, 391, 79–96. [Google Scholar] [CrossRef] [PubMed]
- Resmini, M. Molecularly imprinted polymers as biomimetic catalysts. Anal. Bioanal. Chem. 2012, 402, 3021–3026. [Google Scholar] [CrossRef] [PubMed]
- Dechtrirat, D.; Jetzschmann, K.J.; Stöcklein, W.F.M.; Scheller, F.W.; Gajovic-Eichelmann, N. Protein rebinding to a surface-confined imprint. Adv. Funct. Mater. 2012, 22, 5231–5237. [Google Scholar] [CrossRef]
- Poma, A.; Turner, A.P.F.; Piletsky, S.A. Advances in the manufacture of MIP nanoparticles. Trends Biotechnol. 2010, 28, 629–637. [Google Scholar] [CrossRef] [PubMed]
- Pérez, N.; Whitcombe, M.J.; Vulfson, E.N. Surface imprinting of cholesterol on submicrometer core-shell emulsion particles. Macromolecules 2001, 34, 830–836. [Google Scholar] [CrossRef]
- Li, L.; He, X.; Chen, L.; Zhang, Y. Preparation of novel bovine hemoglobin surface-imprinted polystyrene nanoparticles with magnetic susceptibility. Sci. China Ser. B Chem. 2009, 52, 1402–1411. [Google Scholar] [CrossRef]
- Poma, A.; Guerreiro, A.; Whitcombe, M.J.; Piletska, E.V.; Turner, A.P.F.; Piletsky, S.A. Solid-Phase Synthesis of Molecularly Imprinted Polymer Nanoparticles with a Reusable Template—“Plastic Antibodies”. Adv. Funct. Mater. 2013, 23, 2821–2827. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, Y.; Hong, M.; Bin, Q.; Lin, Z.; Lin, Z.; Cai, Z.; Chen, G. Highly sensitive protein molecularly imprinted electro-chemical sensor based on gold microdendrites electrode and prussian blue mediated amplification. Biosens. Bioelectron. 2013, 42, 612–617. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Shi, X.-W.; Kim, E.; Robinson, L.M.; Nye, C.K.; Ghodssi, R.; Rubloff, G.W.; Bentley, W.E.; Payne, G.F. Chitosan to electroaddress biological components in lab-on-a-chip devices. Carbohydr. Polym. 2011, 84, 704–708. [Google Scholar] [CrossRef]
- Malitesta, C.; Losito, I.; Zambonin, P.G. Molecularly imprinted electrosynthesized polymers: New materials for biomimetic sensors. Anal. Chem. 1999, 71, 1366–1370. [Google Scholar] [CrossRef] [PubMed]
- Panasyuk, T.L.; Mirsky, V.M.; Piletsky, S.A.; Wolfbeis, O.S. Electropolymerized molecularly imprinted polymers as receptor layers in capacitive chemical sensors. Anal. Chem. 1999, 71, 4609–4613. [Google Scholar] [CrossRef]
- Ramanavicius, A.; Oztekin, Y.; Ramanaviciene, A. Electrochemical formation of polypyrrole-based layer for immunosensor design. Sens. Actuators B Chem. 2014, 197, 237–243. [Google Scholar] [CrossRef]
- Menaker, A.; Syritski, V.; Reut, J.; Öpik, A.; Horváth, V.; Gyurcsányi, R.E. Electrosynthesized Surface-Imprinted Conducting Polymer Microrods for Selective Protein Recognition. Adv. Mater. 2009, 21, 2271–2275. [Google Scholar] [CrossRef]
- Chen, H.-J.; Zhang, Z.-H.; Xie, D.; Cai, R.; Chen, X.; Liu, Y.-N.; Yao, S.-Z. Surface-Imprinting Sensor Based on Carbon Nanotubes/Graphene Composite for Determination of Bovine Serum Albumin. Electroanalysis 2012, 24, 2109–2116. [Google Scholar] [CrossRef]
- Karimian, N.; Vagin, M.; Zavar, M.H.A.; Chamsaz, M.; Turner, A.P.F.; Tiwari, A. An ultrasensitive molecularly-imprinted human cardiac troponin sensor. Biosens. Bioelectron. 2013, 50, 492–498. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Dong, J.; Ming, H.; Ai, S. Sensing of glycoprotein via a biomimetic sensor based on molecularly imprinted polymers and graphene–Au nanoparticles. Analyst 2013, 138, 1219. [Google Scholar] [CrossRef] [PubMed]
- Moreira, F.T.C.; Sharma, S.; Dutra, R.A.F.; Noronha, J.P.C.; Cass, A.E.G.; Sales, M.G.F. Protein-responsive polymers for point-of-care detection of cardiac biomarker. Sens. Actuators B Chem. 2014, 196, 123–132. [Google Scholar] [CrossRef]
- Bognár, J.; Szucs, J.; Dorkõ, Z.; Horváth, V.; Gyurcsányi, R.E. Nanosphere lithography as a versatile method to generate surface-imprinted polymer films for selective protein recognition. Adv. Funct. Mater. 2013, 23, 4703–4709. [Google Scholar] [CrossRef]
- Ceolin, G.; Orbán, Á.; Kocsis, V.; Gyurcsányi, R.E.; Kézsmárki, I.; Horváth, V. Electrochemical template synthesis of protein-imprinted magnetic polymer microrods. J. Mater. Sci. 2013, 48, 5209–5218. [Google Scholar] [CrossRef]
- Lee, H.; Dellatore, S.M.; Miller, W.M.; Messersmith, P.B. Mussel-inspired surface chemistry for multifunctional coatings. Science 2007, 318, 426–430. [Google Scholar] [CrossRef] [PubMed]
- Waite, J.H.; Qin, X. Polyphosphoprotein from the adhesive pads of Mytilus edulis. Biochemistry 2001, 40, 2887–2893. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.-H.; Lu, C.-H.; Guo, X.-C.; Chen, F.-R.; Yang, H.-H.; Wang, X.-R. Mussel-inspired molecularly imprinted polymer coating superparamagnetic nanoparticles for protein recognition. J. Mater. Chem. 2010, 20, 880. [Google Scholar] [CrossRef]
- Zhang, M.; Zhang, X.; He, X.; Chen, L.; Zhang, Y. A self-assembled polydopamine film on the surface of magnetic nanoparticles for specific capture of protein. Nanoscale 2012, 4, 3141. [Google Scholar] [CrossRef] [PubMed]
- Yao, G.-H.; Liang, R.-P.; Huang, C.-F.; Wang, Y.; Qiu, J.-D. Surface plasmon resonance sensor based on magnetic molecularly imprinted polymers amplification for pesticide recognition. Anal. Chem. 2013, 85, 11944–11951. [Google Scholar] [CrossRef] [PubMed]
- Lv, Y.; Qin, Y.; Svec, F.; Tan, T. Molecularly imprinted plasmonic nanosensor for selective SERS detection of protein biomarkers. Biosens. Bioelectron. 2016, 80, 433–441. [Google Scholar] [CrossRef] [PubMed]
- Nematollahzadeh, A.; Shojaei, A.; Abdekhodaie, M.J.; Sellergren, B. Molecularly imprinted polydopamine nano-layer on the pore surface of porous particles for protein capture in HPLC column. J. Colloid Interface Sci. 2013, 404, 117–126. [Google Scholar] [CrossRef] [PubMed]
- Xia, Z.; Lin, Z.; Xiao, Y.; Wang, L.; Zheng, J.; Yang, H.; Chen, G. Facile synthesis of polydopamine-coated molecularly imprinted silica nanoparticles for protein recognition and separation. Biosens. Bioelectron. 2013, 47, 120–126. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Shao, M.; Xu, H.; Zhuo, S.; Liu, S.; Lee, S.-T. Molecularly imprinted polymer-coated silicon nanowires for protein specific recognition and fast separation. J. Mater. Chem. 2012, 22, 3990–3996. [Google Scholar] [CrossRef]
- Lin, Z.; Wang, J.; Tan, X.; Sun, L.; Yu, R.; Yang, H.; Chen, G. Preparation of boronate-functionalized molecularly imprinted monolithic column with polydopamine coating for glycoprotein recognition and enrichment. J. Chromatogr. A 2013, 1319, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Tretjakov, A.; Syritski, V.; Reut, J.; Boroznjak, R.; Volobujeva, O.; Öpik, A. Surface molecularly imprinted polydopamine films for recognition of immunoglobulin G. Microchim. Acta 2013, 180, 1433–1442. [Google Scholar] [CrossRef]
- Li, L.; Fan, L.; Dai, Y.; Kan, X. Recognition and determination of bovine hemoglobin using a gold electrode modified with gold nanoparticles and molecularly imprinted self-polymerized dopamine. Microchim. Acta 2015, 182, 2477–2483. [Google Scholar] [CrossRef]
- Liu, R.; Sha, M.; Jiang, S.; Luo, J.; Liu, X. A facile approach for imprinting protein on the surface of multi-walled carbon nanotubes. Talanta 2014, 120, 76–83. [Google Scholar] [CrossRef] [PubMed]
- Yu, B.; Liu, J.; Liu, S.; Zhou, F. Pdop layer exhibiting zwitterionicity: A simple electrochemical interface for governing ion permeability. Chem. Commun. 2010, 46, 5900. [Google Scholar] [CrossRef] [PubMed]
- Hollmann, F.; Arends, I.W.C.E. Enzyme initiated radical polymerizations. Polymers 2012, 4, 759–793. [Google Scholar] [CrossRef]
- Kobayashi, S.; Makino, A. Enzymatic polymer synthesis: An opportunity for green polymer chemistry. Chem. Rev. 2009, 109, 5288–5353. [Google Scholar] [CrossRef] [PubMed]
- Lopes, G.R.; Pinto, D.C.G.A.; Silva, A.M.S. Horseradish peroxidase (HRP) as a tool in green chemistry. RSC Adv. 2014, 4, 37244. [Google Scholar] [CrossRef]
- Lizardi, P.M.; Engelberg, A. Rapid isolation of RNA using proteinase K and sodium perchlorate. Anal. Biochem. 1979, 98, 116–122. [Google Scholar] [CrossRef]
- Rao, M.B.; Tanksale, A.M.; Ghatge, M.S.; Deshpande, V. Molecular and biotechnological aspects of microbial proteases. Microbiol. Mol. Biol. Rev. 1998, 62, 597–635. [Google Scholar] [PubMed]
- Hawkins, D.M.; Stevenson, D.; Reddy, S.M. Investigation of protein imprinting in hydrogel-based molecularly imprinted polymers (HydroMIPs). Anal. Chim. Acta 2005, 542, 61–65. [Google Scholar] [CrossRef]
- Moreira, F.T.C.; Ferreira, M.J.M.S.; Puga, J.R.T.; Sales, M.G.F. Screen-printed electrode produced by printed-circuit board technology. Application to cancer biomarker detection by means of plastic antibody as sensing material. Sens. Actuators B Chem. 2016, 223, 927–935. [Google Scholar] [CrossRef]
- Dechtrirat, D.; Gajovic-Eichelmann, N.; Bier, F.F.; Scheller, F.W. Hybrid Material for Protein Sensing Based on Electrosynthesized MIP on a Mannose Terminated Self-Assembled Monolayer. Adv. Funct. Mater. 2014, 24, 2233–2239. [Google Scholar] [CrossRef]
- Bosserdt, M.; Gajovic-Eichelman, N.; Scheller, F.W. Modulation of direct electron transfer of cytochrome c by use of a molecularly imprinted thin film. Anal. Bioanal. Chem. 2013, 405, 6437–6444. [Google Scholar] [CrossRef] [PubMed]
- Reddy, S.M.; Sette, G.; Phan, Q. Electrochemical probing of selective haemoglobin binding in hydrogel-based molecularly imprinted polymers. Electrochim. Acta 2011, 56, 9203–9208. [Google Scholar] [CrossRef]
- Liu, Y.T.; Deng, J.; Xiao, X.L.; Ding, L.; Yuan, Y.L.; Li, H.; Li, X.T.; Yan, X.N.; Wang, L.L. Electrochemical sensor based on a poly(para-aminobenzoic acid) film modified glassy carbon electrode for the determination of melamine in milk. Electrochim. Acta 2011, 56, 4595–4602. [Google Scholar] [CrossRef]
- Xue, X.; Wei, Q.; Wu, D.; Li, H.; Zhang, Y.; Feng, R.; Du, B. Determination of methyl parathion by a molecularly imprinted sensor based on nitrogen doped graphene sheets. Electrochim. Acta 2014, 116, 366–371. [Google Scholar] [CrossRef]
- Yu, H.C.; Huang, X.Y.; Lei, F.H.; Tan, X.C.; Wei, Y.C.; Li, H. Molecularly imprinted electrochemical sensor based on nickel nanoparticle-modified electrodes for phenobarbital determination. Electrochim. Acta 2014, 141, 45–50. [Google Scholar] [CrossRef]
- Rezaei, B.; Khalili Boroujeni, M.; Ensafi, A.A. Caffeine electrochemical sensor using imprinted film as recognition element based on polypyrrole, sol-gel, and gold nanoparticles hybrid nanocomposite modified pencil graphite electrode. Biosens. Bioelectron. 2014, 60, 77–83. [Google Scholar] [CrossRef] [PubMed]
- Rezaei, B.; Rahmanian, O.; Ensafi, A.A. An electrochemical sensor based on multiwall carbon nanotubes and molecular imprinting strategy for warfarin recognition and determination. Sens. Actuators B Chem. 2014, 196, 539–545. [Google Scholar] [CrossRef]
- Pasha, S.K.; Kaushik, A.; Vasudev, A.; Snipes, S.A.; Bhansali, S. Electrochemical Immunosensing of Saliva Cortisol. J. Electrochem. Soc. 2013, 161, B3077–B3082. [Google Scholar] [CrossRef]
- Florea, A.; Guo, Z.; Cristea, C.; Bessueille, F.; Vocanson, F.; Goutaland, F.; Dzyadevych, S.; Săndulescu, R.; Jaffrezic-Renault, N. Anticancer drug detection using a highly sensitive molecularly imprinted electrochemical sensor based on an electropolymerized microporous metal organic framework. Talanta 2015, 138, 71–76. [Google Scholar] [CrossRef] [PubMed]
- Prasad, B.B.; Prasad, A.; Tiwari, M.P. Multiwalled carbon nanotubes-ceramic electrode modified with substrate-selective imprinted polymer for ultra-trace detection of bovine serum albumin. Biosens. Bioelectron. 2013, 39, 236–243. [Google Scholar] [CrossRef] [PubMed]
- Cieplak, M.; Szwabinska, K.; Sosnowska, M.; Chandra, B.K.C.; Borowicz, P.; Noworyta, K.; D’Souza, F.; Kutner, W. Selective electrochemical sensing of human serum albumin by semi-covalent molecular imprinting. Biosens. Bioelectron. 2015, 74, 960–966. [Google Scholar] [CrossRef] [PubMed]
- Cai, D.; Ren, L.; Zhao, H.; Xu, C.; Zhang, L.; Yu, Y.; Wang, H.; Lan, Y.; Roberts, M.F.; Chuang, J.H.; et al. A molecular-imprint nanosensor for ultrasensitive detection of proteins. Nat. Nanotechnol. 2010, 5, 597–601. [Google Scholar] [CrossRef] [PubMed]
- Lian, W.; Liu, S.; Wang, L.; Liu, H. A novel strategy to improve the sensitivity of antibiotics determination based on bioelectrocatalysis at molecularly imprinted polymer film electrodes. Biosens. Bioelectron. 2015, 73, 214–220. [Google Scholar] [CrossRef] [PubMed]
- Vlatakis, G.; Andersson, L.I.; Müller, R.; Mosbach, K. Drug assay using antibody mimics made by molecular imprinting. Nature 1993, 361, 645–647. [Google Scholar] [CrossRef] [PubMed]
- Baggiani, C.; Anfossi, L.; Giovannoli, C. MIP-based immunoassays: State of the Art, limitations and Perspectives. Mol. Impr. 2013, 1, 41–54. [Google Scholar] [CrossRef]
- Surugiu, I.; Ye, L.; Yilmaz, E.; Dzgoev, A.; Danielsson, B.; Mosbach, K.; Haupt, K. An enzyme-linked molecularly imprinted sorbent assay. Analyst 2000, 125, 13–16. [Google Scholar] [CrossRef]
- Surugiu, I.; Danielsson, B.; Ye, L.; Mosbach, K.; Haupt, K. Chemiluminescence imaging ELISA using an imprinted polymer as the recognition element instead of an antibody. Anal. Chem. 2001, 73, 487–491. [Google Scholar] [CrossRef] [PubMed]
- Surugiu, I.; Svitel, J.; Ye, L.; Haupt, K.; Danielsson, B. Development of a flow injection capillary chemiluminescent ELISA using an imprinted polymer instead of the antibody. Anal. Chem. 2001, 73, 4388–4392. [Google Scholar] [CrossRef] [PubMed]
- Piletsky, S.A.; Piletska, E.V.; Bossi, A.; Karim, K.; Lowe, P.; Turner, A.P. Substitution of antibodies and receptors with molecularly imprinted polymers in enzyme-linked and fluorescent assays. Biosens. Bioelectron. 2001, 16, 701–707. [Google Scholar] [CrossRef]
- Wang, S.; Xu, Z.; Fang, G.; Zhang, Y.; Liu, B.; Zhu, H. Development of a Biomimetic Enzyme-Linked Immunosorbent Assay Method for the Determination of Estrone in Environmental Water using Novel Molecularly Imprinted Films of Controlled Thickness as Artificial Antibodies. J. Agric. Food Chem. 2009, 57, 4528–4534. [Google Scholar] [CrossRef] [PubMed]
- Chianella, I.; Guerreiro, A.; Moczko, E.; Caygill, J.S.; Piletska, E.V.; De Vargas Sansalvador, I.M.P.; Whitcombe, M.J.; Piletsky, S.A. Direct Replacement of Antibodies with Molecularly Imprinted Polymer Nanoparticles in ELISA—Development of a Novel Assay for Vancomycin. Anal. Chem. 2013, 85, 8462–8468. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Tao, H.; Li, J. Molecularly Imprinted Electrochemical Luminescence Sensor Based on Enzymatic Amplification for Ultratrace Isoproturon Determination. Electroanalysis 2012, 24, 1664–1670. [Google Scholar] [CrossRef]
- Li, J.; Jiang, F.; Wei, X. Molecularly Imprinted Sensor Based on an Enzyme Amplifier for Ultratrace Oxytetracycline Determination. Anal. Chem. 2010, 82, 6074–6078. [Google Scholar] [CrossRef]
- Yarman, A.; Scheller, F. The First Electrochemical MIP Sensor for Tamoxifen. Sensors 2014, 14, 7647–7654. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Caballero, A.; Goicolea, M.A.; Barrio, R.J. Paracetamol voltammetric microsensors based on electrocopolymerized–molecularly imprinted film modified carbon fiber microelectrodes. Analyst 2005, 130, 1012. [Google Scholar] [CrossRef] [PubMed]
- Özcan, L.; Şahin, Y. Determination of paracetamol based on electropolymerized-molecularly imprinted polypyrrole modified pencil graphite electrode. Sens. Actuators B Chem. 2007, 127, 362–369. [Google Scholar] [CrossRef]
- Wang, X.; Luo, J.; Yi, C.; Liu, X. Paracetamol Sensor Based on Molecular Imprinting by Photosensitive Polymers. Electroanalysis 2013, 25, 1907–1916. [Google Scholar] [CrossRef]
- Luo, J.; Fan, C.; Wang, X.; Liu, R.; Liu, X. A novel electrochemical sensor for paracetamol based on molecularly imprinted polymeric micelles. Sens. Actuators B Chem. 2013, 188, 909–916. [Google Scholar] [CrossRef]
- Tan, Y.; Zhou, Z.; Wang, P.; Nie, L.; Yao, S. A study of a bio-mimetic recognition material for the BAW sensor by molecular imprinting and its application for the determination of paracetamol in the human serum and urine. Talanta 2001, 55, 337–347. [Google Scholar] [CrossRef]
- Pernites, R.; Ponnapati, R.; Felipe, M.J.; Advincula, R. Electropolymerization molecularly imprinted polymer (E-MIP) SPR sensing of drug molecules: Pre-polymerization complexed terthiophene and carbazole electroactive monomers. Biosens. Bioelectron. 2011, 26, 2766–2771. [Google Scholar] [CrossRef] [PubMed]
- Carralero Sanz, V.; Mena, M.L.; González-Cortés, A.; Yáñez-Sedeño, P.; Pingarrón, J.M. Development of a tyrosinase biosensor based on gold nanoparticles-modified glassy carbon electrodes: Application to the measurement of a bioelectrochemical polyphenols index in wines. Anal. Chim. Acta 2005, 528, 1–8. [Google Scholar] [CrossRef]
- Boujakhrout, A.; Sánchez, A.; Díez, P.; Jiménez-Falcao, S.; Martínez-Ruiz, P.; Peña-Álvarez, M.; Pingarrón, J.M.; Villalonga, R. Decorating graphene oxide/nanogold with dextran-based polymer brushes for the construction of ultrasensitive electrochemical enzyme biosensors. J. Mater. Chem. B 2015, 3, 3518–3524. [Google Scholar] [CrossRef]
- Vidal, J.C.; Esteban, S.; Gil, J.; Castillo, J.R. A comparative study of immobilization methods of a tyrosinase enzyme on electrodes and their application to the detection of dichlorvos organophosphorus insecticide. Talanta 2006, 68, 791–799. [Google Scholar] [CrossRef] [PubMed]
- Yildiz, H.B.; Castillo, J.; Guschin, D.A.; Toppare, L.; Schuhmann, W. Phenol biosensor based on electrochemically controlled integration of tyrosinase in a redox polymer. Microchim. Acta 2007, 159, 27–34. [Google Scholar] [CrossRef]
- Kurbanoglu, S.; Mayorga-Martinez, C.C.; Medina-Sánchez, M.; Rivas, L.; Ozkan, S.A.; Merkoçi, A. Antithyroid drug detection using an enzyme cascade blocking in a nanoparticle-based lab-on-a-chip system. Biosens. Bioelectron. 2015, 67, 670–676. [Google Scholar] [CrossRef] [PubMed]
- Valero, E.; Varón, R.; García-Carmona, F. Tyrosinase-Mediated Oxidation of Acetaminophen to 4-Acetamido-o- Benzoquinone. Biol. Chem. 2002, 383, 1931–1939. [Google Scholar] [CrossRef] [PubMed]
- Calas-Blanchard, C.; Istamboulié, G.; Bontoux, M.; Plantard, G.; Goetz, V.; Noguer, T. Biosensor-based real-time monitoring of paracetamol photocatalytic degradation. Chemosphere 2015, 131, 124–129. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Naser, N.; Wollenberger, U. Use of tyrosinase for enzymatic elimination of acetaminophen interference in amperometric sensing. Anal. Chim. Acta 1993, 281, 19–24. [Google Scholar] [CrossRef]
- Yarman, A.; Scheller, F.W. MIP-esterase/Tyrosinase Combinations for Paracetamol and Phenacetin. Electroanalysis 2016, 28, 2222–2227. [Google Scholar] [CrossRef]
- Bacon, J.; Adams, R.N. Anodic oxidations of aromatic amines. III. Substituted anilines in aqueous media. J. Am. Chem. Soc. 1968, 90, 6596–6599. [Google Scholar] [CrossRef]
- Yarman, A.; Scheller, F.W. Coupling biocatalysis with molecular imprinting in a biomimetic sensor. Angew. Chem. Int. Ed. 2013, 52, 11521–11525. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yarman, A.; Jetzschmann, K.J.; Neumann, B.; Zhang, X.; Wollenberger, U.; Cordin, A.; Haupt, K.; Scheller, F.W. Enzymes as Tools in MIP-Sensors. Chemosensors 2017, 5, 11. https://doi.org/10.3390/chemosensors5020011
Yarman A, Jetzschmann KJ, Neumann B, Zhang X, Wollenberger U, Cordin A, Haupt K, Scheller FW. Enzymes as Tools in MIP-Sensors. Chemosensors. 2017; 5(2):11. https://doi.org/10.3390/chemosensors5020011
Chicago/Turabian StyleYarman, Aysu, Katharina J. Jetzschmann, Bettina Neumann, Xiaorong Zhang, Ulla Wollenberger, Aude Cordin, Karsten Haupt, and Frieder W. Scheller. 2017. "Enzymes as Tools in MIP-Sensors" Chemosensors 5, no. 2: 11. https://doi.org/10.3390/chemosensors5020011
APA StyleYarman, A., Jetzschmann, K. J., Neumann, B., Zhang, X., Wollenberger, U., Cordin, A., Haupt, K., & Scheller, F. W. (2017). Enzymes as Tools in MIP-Sensors. Chemosensors, 5(2), 11. https://doi.org/10.3390/chemosensors5020011









