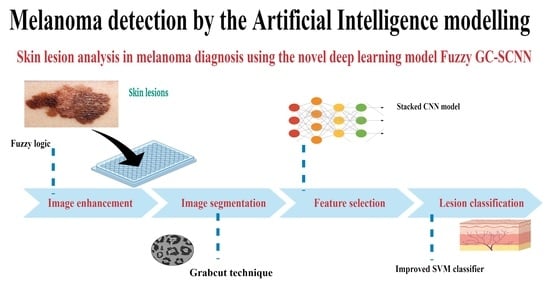
Skin Lesion Analysis for Melanoma Detection Using the Novel Deep Learning Model Fuzzy GC-SCNN
Abstract
:1. Introduction
2. Methods
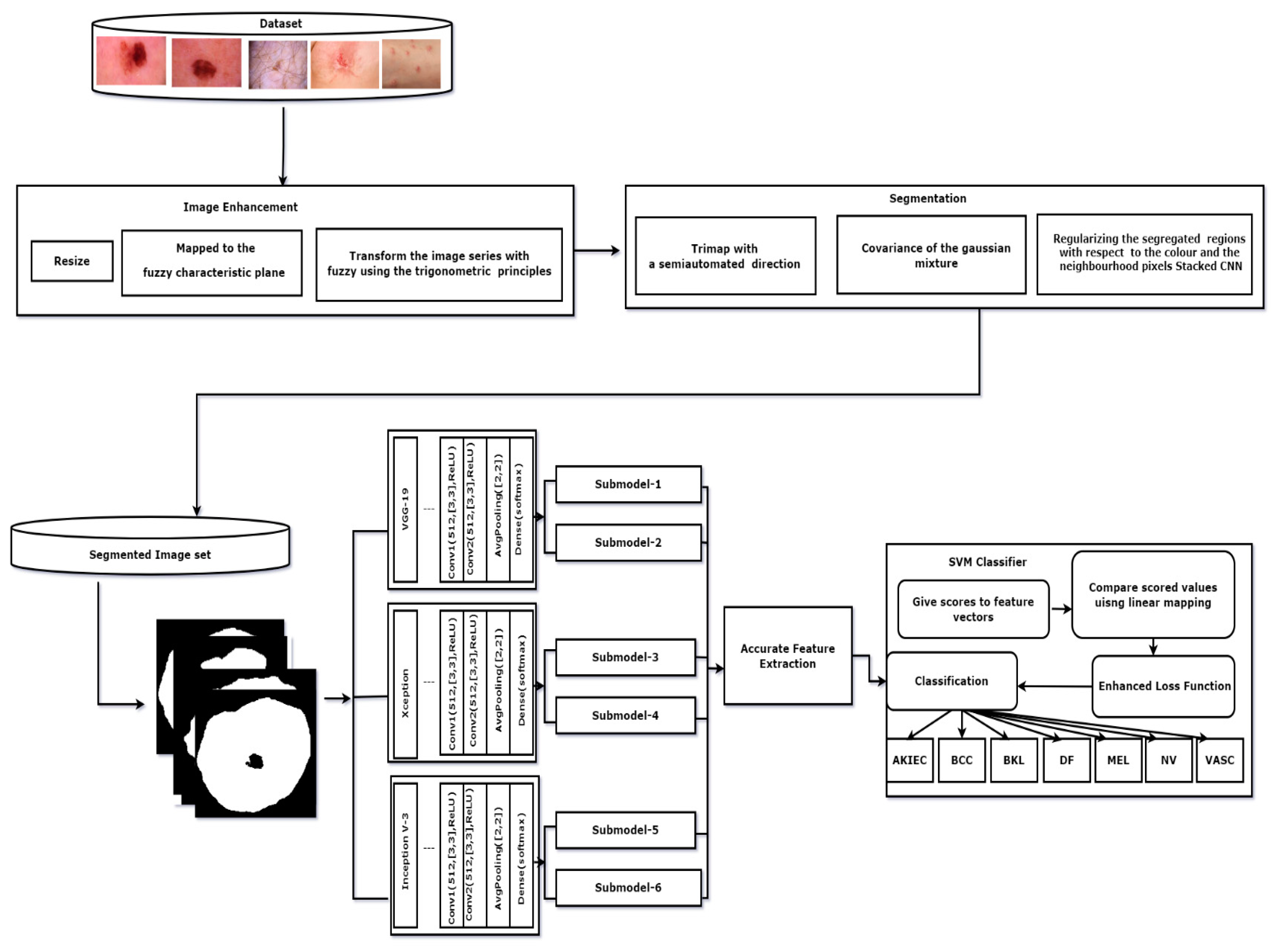
2.1. Dataset
2.2. Data Preprocessing
- ❖
- The pixels of the skin lesion domain are taken to a fuzzy domain. Let M be an image of p × q, and M (p,q) represent the intensity of the skin lesion image pixels that must be mapped to the fuzzy characteristic plane. It can be expressed as follows , p = 1, …, m and q = 1, …, n; where represent the pixels and µM (p, q) is the intensity level degree of the image ranging from zero to one.
- ❖
- Assign the fuzzy plane pixels to the logarithmic function to map to the fuzzy domain f (M (p, q)) = ((1 + ); where Mmax and Mmin are the maximum and minimum intensity of the skin lesion image pixels.
- ❖
- To enhance the portions of the skin lesion images, transform the image using the trigonometric series with fuzzy principles as mentioned f (T (p, q)) = T (p, q) + f (M (p, q))2 where 0 ≤ f (M (p, q)) ≤ 0.5; where T (p, q) = and a = π (f (M (p, q) − 0.5) + 1.
- ❖
- The defuzzification can be expressed as D = Mmin + ((Mmax − Mmin) × 2 T(p, q) − 1)
- ❖
- Later, enhance the image quality by skin lesion image channel-wise.
2.3. Image Segmentation
2.4. Feature Extraction
| Algorithm 1: GC-SCNN. |
| Input: Segmented Images Output: Skin cancer Classification results for k = 1 to length (segmented images) do for j = 1 to 3 do sub-model j. predicts (segmented image) end for final = concatenation (P1, P2, P3) end for assess the SoftMax classifier on the feature vector final stacked CNN = Train (final, label) classification of skin cancer images prediction = classification (stacked CNN, testset) return prediction |
2.5. Lesion Classification
| Algorithm 2: Enhanced SVM algorithm. |
| Initialize the values in the training set Repeat for every i = 1 to N calculate the loss function for all values compare the extracted patches in the images end for Repeat for every score vector i- 1 to N Compute SVM with imputed labels argmax((w × xi) + b), i end for Evaluate for different weights and compute output. |
2.6. Experimental Framework
2.7. Performance Metrics
- ❖
- Accuracy measures the portion of the true results among the total number of the cases and is written as accuracy = ; Where FP-False positive, FN-False Negative, TP-True Positive, TN-True Negative
- ❖
- Sensitivity is the portion of the positive outcomes among the actual positive and it is defined as sensitivity =
- ❖
- Specificity is defined as the portion of the true negative outcomes among the negative outcomes and it is written as specificity =
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Rogers, H.W.; Weinstock, M.A.; Feldman, S.R.; Coldiron, B.M. Incidence estimate of non melanoma skin cancer in the U.S population. JAMA Dermatol. 2015, 155, 1081–1086. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.; Miah, S.; Haque, J.; Rahman, M.; Islam, K. An enhanced technique of skin cancer classification using deep convolutional neural network with transfer learning models. Mach. Learn. Appl. 2021, 5, 100036. [Google Scholar] [CrossRef]
- Nasir, M.; Attique Khan, M.; Sharif, M.; Lali, U.; Saba, T.; Iqbal, T. An improved strategy for skin lesion detection and classification using uniform segmentation and feature selection based approach. Microsc. Res. Tech. 2018, 81, 528–543. [Google Scholar] [CrossRef] [PubMed]
- Skin Cancer. Available online: https://www.skincancer.org/skin-cancer-information/skin-cancer-facts/ (accessed on 12 December 2021).
- Khan, M.A.; Muhammad, K.; Sharif, M.; Akram, T.; de Albuquerque, V.H.C. Multi-Class Skin Lesion Detection and Classification via Teledermatology. IEEE J. Biomed. Health Inform. 2021, 25, 4267–4275. [Google Scholar] [CrossRef]
- Pacheco, A.G.; Krohling, R.A. The impact of patient clinical information on automated skin cancer detection. Comput. Biol. Med. 2020, 116, 103545. [Google Scholar] [CrossRef]
- Pacheco, A.G.; Sastry, C.S.; Trappenberg, T.; Oore, S.; Krohling, R.A. On out-of-Distribution Detection Algorithms with deep neural skin cancer classifiers. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) Workshops, Seattle, WA, USA, 14–19 June 2020; pp. 1–10. [Google Scholar]
- Abdelhalim, I.S.A.; Mohamed, M.F.; Mahdy, Y.B. Data augmentation for skin lesion self attention bassed progressive generative adversial network. Expert Syst. Appl. 2021, 165, 113922. [Google Scholar] [CrossRef]
- Liu, X.; Chen, C.H.; Karvela, M.; Toumazou, C. A DNA based intelligent expert system for personalised skin helath tecommentdations. IEEE J. Biomed. Health Inform. 2020, 24, 3276–3284. [Google Scholar] [CrossRef]
- Duggani, K.; Nath, M.K. A technical review report on deep learning approach for skin cancer detection and segmentation. Data Anal. Manag. 2021, 54, 87–99. [Google Scholar] [CrossRef]
- Khan, I.U.; Aslam, N.; Anwar, T.; Aljameel, S.S.; Ullah, M.; Khan, R.; Rehman, A.; Akhtar, N. Remote diagnosis and triaging model for skin cancer usign efficientnet and extreme gradient boosting. Complexity 2021, 2021, 5591614. [Google Scholar] [CrossRef]
- Alquran, H.; Qasmieh, A.; Alqudah, M.; Alhammouri, S.; Alawneh, E.; Abughazaleh, A.; Hasayen, F. The melanoma skin cancer detection and classification using support vector machine. In Proceedings of the International Conference on Applied Electrical Engineering and Computational Technology (AEECT), Aqaba, Jordan, 11–13 October 2017; pp. 1–5. [Google Scholar]
- Mahmuei, S.S.; Aldeen, M.; Stoecker, W.W.; Garnavi, R. Biologically inspired quadtree color detection in dermoscopy images of melanoma. IEEE J. Biomed. Health Inform. 2019, 23, 570–577. [Google Scholar] [CrossRef]
- Hameed, N.; Hameed, F.; Shabut, A.; Khan, S.; Cirstea, S.; Hossain, A. An intelligent computer aided scheme for classifying multiple skin lesions. Computers 2019, 8, 62. [Google Scholar] [CrossRef] [Green Version]
- Khan, M.Q.; Hussain, A.; Rehman, S.U.; Khan, U.; Maqsood, M.; Mehmood, K.; Khan, M.A. Classification of melanoma and nevus in digital images for diagnosis of skin cancer. IEEE Access 2019, 7, 90132–90144. [Google Scholar] [CrossRef]
- Celebi, M.E.; Codella, N.; Halpern, A. Dermoscopy Image Analysis: Overview and Future Directions. IEEE J. Biomed. Health Inform. 2019, 23, 474–478. [Google Scholar] [CrossRef]
- Guha, S.R.; Haque, S.R. Performance comparison of machine learning based classification of skin diseases from skin lesion images. In Proceedings of the International Conference of Communication Computational Electronics System, Coimbatore, India, 21–22 October 2020; pp. 15–25. [Google Scholar]
- Heckler, A.; Utikal, S.; Enk, H.A.; Hauschild, M.; Weichenthal, M.; Maron, R.C.; Berking, C.; Haferkamp, S.; Klode, J.; Schadendorf, D.; et al. Superior skin cancer classification by the combination of human and artificial intelligence. Eur. J. Cancer 2019, 120, 114–121. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Seeja, R.D.; Suresh, A. Deep learning based skin lesion segmentation and classification of melanoma using support vector machine (SVM). Asian Pac. J. Cancer Prev. 2019, 20, 1555–1561. [Google Scholar]
- Esteva, A.; Kuprel, B.; Novoa, A.; Ko, S.; Swetter, M.; Blau, H.M.; Thrun, S. Dermatologist level classification of skin cancer with dep neural networks. Nature 2017, 542, 115–118. [Google Scholar] [CrossRef]
- Brinker, T.J.; Hekler, A.; Utikal, J.S.; Grabe, N.; Schadendorf, D.; Klode, J.; Berking, C.; Steeb, T.; Enk, H.; Von Kalle, C. Skin cancer classification using convolutional neural networks: Systematic review. J. Med. Internet Residual 2018, 20, e11936. [Google Scholar] [CrossRef] [PubMed]
- Tschandl, P.; Rosendahl, C.; Akay, B.N.; Argenziano, G.; Blum, A.; Braun, R.P.; Cabo, H.; Gourhant, J.Y.; Kreusch, J.; Lallas, A.; et al. Expert level Diagnosis of Non Pigmented skin Cancer by Combined Convolution Neural Networks. JAMA Dermatol. 2019, 155, 58–65. [Google Scholar] [CrossRef]
- Saba, T.; Khan, M.A.; Rehman, A.; Sainte, S.L.M. Region Extraction and Classification of Skin cancer: A Heterogeneous Framework of Deep CNN features Fusion and Reduction. J. Med. Syst. 2019, 43, 289. [Google Scholar] [CrossRef]
- HAM1000dataset. Available online: https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/DBW86T (accessed on 18 March 2022).
- Wang, C.; Chen, D.; Hao, L.; Liu, X.; Zeng, Y.; Chen, J.; Zhang, G. Pulmonary image classification based on Inception v3 transfer learing model. IEEE Access 2019, 7, 146533–146541. [Google Scholar] [CrossRef]
- Chollet, F. Xception: Dep Learning with depth wise separable convolutions. In Proceedings of the IEEE Conference on Computer Viion and Patter Recognition (CVPR-2017), Honolulu, HI, USA, 21–26 July 2017; pp. 1800–1807. [Google Scholar]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. Imagenet classification with deep convolution neural networks. Adv. Neural Inf. Process. Syst. 2012, 25, 1097–1105. [Google Scholar]
- Usharani, B. ILF-LSTM: Enhanced loss function in LSTM to predict the sea surface temperature. Soft Comput. 2022, 1433–7479. [Google Scholar] [CrossRef]
- Kaseem, M.A.; Hosny, K.M.; Fouad, M.M. Skin lesion classification into eight classes for ISIC2019 using deep convolution neural networks and transfer learning. IEEE Access 2020, 8, 114822–114832. [Google Scholar] [CrossRef]
- Chaturvedi, S.S.; Gupta, K.; Prasad, P.S. Skin Lesion analyser: An efficient seven way multi class skin cancer classification using mobile net. In Proceedings of the International Conference on Advanced Machine Learning Technologies and Applications, Cairo, Egypt, 20–22 March 2020; Volume 12, pp. 165–276. [Google Scholar]
- Khan, M.A.; Sharif, M.; Akram, T.; Damasevicius, R.; Maskeliunas, R. Skin Lesion Segmentation and multiclass classification using deep learning features and improved moth flame optimization. Diagnostics 2021, 11, 811. [Google Scholar] [CrossRef] [PubMed]
- Rahul, A.R.; Mozaffari, M.H.; Lee, W.S.; Pari, B.E. Skin lesions classification using deep learning based on dilated convolution. bioRxiv 2019, 860700. [Google Scholar] [CrossRef] [Green Version]
- Gessert, N.; Nielsen, M.; Shaikh, M.; Werner, R.; Schlaefer, A. Skin lesion classification using ensembles of multi-resolution EfficientNets with meta data. MethodsX 2020, 7, 100864. [Google Scholar] [CrossRef]
- Ailin, S.; Chao, H.; Minjie, C.; Hui, X.; Yali, Y. Skin lesion classification using additional patient information. Biomed Res. Int. 2021, 2021, 6673852. [Google Scholar] [CrossRef]
- Khan, M.A.; Javed, M.Y.; Sharif, M.; Saba, T.; Rehman, A. Multimodal deep neural network based features extraction and optimal selection approach for skin lesion classification. In Proceedings of the International Conference on Computer and Inflrmation Science (ICCIS2019), Sakaka, Saudi Arabia, 3–4 April 2019; pp. 1–7. [Google Scholar]
- Mohamed, E.H.; Behaidy, E.W.H. Enhanced skin lesions classification using deep convolutional networks. In Proceedings of the International Conference on Intelligent computing and information systems (ICICIS2019), Cairo, Egypt, 8–10 December 2019; pp. 180–188. [Google Scholar]
- Huang, H.W.; Hsu, B.W.Y.; Lee, C.H.; Tseng, V.S. Development of a light weight deep learning model for cloud applications and remote diagnosis of skin cancers. J. Dermatol. 2021, 48, 310–316. [Google Scholar] [CrossRef]
- Liu, Q.; Yu, L.; Luo, L.; Dou, Q.; Heng, P.A. Semi supervised medical image classification with relation driven self ensembling model. IEEE Trans. Med. Imaging 2020, 39, 3429–3440. [Google Scholar] [CrossRef]
- Gu, Y.; Ge, Z.; Bonnington, C.P.; Zhou, J. Progressive transfer learning and adversarial domain adaption for ross domain skin disease classification. IEEE J. Biomed. Health Inf. 2019, 24, 1379–1393. [Google Scholar] [CrossRef]
- Zhou, L.; Luo, Y. Deep features fusion with mutual attention transformr for skin lesion diagnosis. In Proceedings of the IEEE International conference on Imge processing (ICIP2021), Anchorage, AK, USA, 19–22 September 2021; pp. 3797–3801. [Google Scholar]
- Cai, G.; Zhu, Y.; Wu, Y.; Jiang, X.; Ye, J.; Yang, D. A multimodal transformer to fue images and metadata for skin disease classification. Vis. Comput. 2022, 1–13. [Google Scholar] [CrossRef]
- Ahmed, S.A.A.; Yanikouglu, B.; Goksu, O.; Aptoula, E. Skin lesion classification with deep CNN ensembles. In Proceedings of the Proceedings SIU, Gaziantep, Turkey, 5–7 October 2020; pp. 1–4. [Google Scholar]
- Pacheco, A.G.C.; Ali, A.R.; Trappenberg, T. Skin cancer detection based on deep learning and entropy to detect outlier samples. arXiv 2019, arXiv:1909.04525. [Google Scholar]
- Molina, E.O.; Solorza, S.; Alvarez, J. Classification of dermoscopy skin lesion color images using fractal deep learning features. Appl. Sci. 2020, 10, 5954. [Google Scholar] [CrossRef]
- Sun, Q.; Huang, C.; Chen, M.; Xu, H.; Yang, Y. Skin Lesion Classification Using Additional Patient Information. BioMed Res. Int. 2021, 2021, 6673852. [Google Scholar] [CrossRef]
- Iqbla, I.; Younus, M.; Walayat, K.; Kakar, M.U.; Ma, J. Automated multi class classification of skin lesions through deep convolutional neural network with dermoscopic images. Comput. Med. Imaging Graph. 2021, 88, 101843. [Google Scholar] [CrossRef] [PubMed]
- Villa-Pulgarin, J.P.; Ruales-Torres, A.A.; Arias-Garzon, D.; Bravo-Ortiz, M.A.; Arteaga-Arteaga, H.B.; Mora-Rubio, A.; Alzate-Grisales, J.A.; Mercado-Ruiz, E.; Hassaballah, M.; Orozco-Arias, S.; et al. Optimized convolutional neural network models for skin lesion classification. Comput. Mater. Contin. 2022, 70, 2131–2148. [Google Scholar] [CrossRef]
- Afza, F.; Khan, M.A.; Sharif, M.; Rehman, A. Microscopic Skin Laceration segmentation and classification: A Framework of statistical normal distribution and optimal feature selection. Microsc. Res. Tech. 2019, 82, 1471–1488. [Google Scholar] [CrossRef] [PubMed]
- Sadria, R.; Azarianpour, S.; Zekri, M.; Celebi, M.E. WN based approach to melanoma diagnosis from dermoscopy images. IET Image Process. 2017, 11, 475–482. [Google Scholar] [CrossRef]
- Schaefer, G.; Krawczyk, B.; Celebi, M.E.; Iyatomi, H. An ensemble classification approach for melanoma diagnosis. Memetic Comput. 2014, 6, 233–240. [Google Scholar] [CrossRef]
- Waheed, Z.; Zafar, M.; Waheed, A.; Riaz, F. An efficient machine learning approach for the detection f melanoma using dermoscopic images. In Proceedings of the International Conference on Communication Computing and Digital Systems (C-CODE), Islamabad, Pakistan, 8–9 March 2017; pp. 316–319. [Google Scholar]
- Sivaraj, S.; Malmathanraj, R.; Palanisamy, P. Detecting anomalous growth of skin lesion uisng threshold based segmentation algorithm and fuzzy k nearest neighbor classifier. J. Cnacer Res. Ther. 2020, 16, 40–52. [Google Scholar] [CrossRef]
- Bi, L.; Kim, J.; Ahn, E.; Feng, D.; Fulham, M. Automatic melanoma detection via multi scale lesion biased representation and joint reverse classification. In Proceedings of the International Symposium on Biomedical Imaging(ISBI), Prague, Czech Republic, 13–16 April 2016; pp. 1055–1058. [Google Scholar]
- Abbes, W.; Sellami, D. Deep Neural network for fuzzy automatic melanoma diagnosis. In Proceedings of the International Joint Conference on Computer Vision, Imaging and Computer Graphics Theory and Applications (VISIGRAPP2019), Prague, Czech Republic, 25–27 February 2019; pp. 47–56. [Google Scholar]
- Kawahara, J.; Hamarneh, G. Multi resolution tract CNN with hybrid pretrained and skin lesion trained layers. In Machine Learning in Medical Imaging. MLMI 2016; Lecture Notes in Computer Science; Wang, L., Adeli, E., Wang, Q., Shi, Y., Suk, H.I., Eds.; Springer: Cham, Switzerland, 2016; Volume 10019, pp. 164–171. [Google Scholar]
- Kawahara, J.; Benraieb, A.; Hamarneh, G. Deep features to classify skin lesions. In Proceedings of the IEEE International Symposium on Biomedical Imaging (ISBI2016), Prague, Czech Republic, 13–16 April 2016; pp. 1397–1400. [Google Scholar]
- Nyiri, T.; Kiss, A. Novel ensembling methods for dermatological image classification. In Theory and Practice of Natural Computing. TPNC 2018; Lecture Notes in Computer Science; Fagan, D., Martín-Vide, C., O’Neill, M., Vega-Rodríguez, M.A., Eds.; Springer: Cham, Switzerland, 2018; Volume 11324, pp. 438–448. [Google Scholar]
- Shahin, A.H.; Kamal, A.; Elattat, M.A. Deep ensemble learning for skin lesion classification from dermoscopic images. In Proceedings of the IEEE International Biomedical Engineering Conferences (CIBEC2018), Cairo, Egypt, 20–22 December 2018; pp. 150–153. [Google Scholar]
- Pennisi, A.; Bloisi, D.D.; Nardi, D.; Giampetruzzi, A.R.; Mondino, C.; Facchiano, A. Skin lesions image segmentation using Delaunay Triangulation for melanoma detection. Comput. Med. Imaging Graph. 2016, 52, 89–103. [Google Scholar] [CrossRef] [Green Version]
- Warsi, F.; Khanam, R.; Kamya, S. An efficient 3D color texture feature and neural network technique for melanoma detection. Inform. Med. Unlocked 2019, 1, 100176. [Google Scholar] [CrossRef]
- Khatib, E.; Popescu, D.; Ichim, L. Deep learning based methods for automatic diagnosis of skin lesions. Sensors 2020, 20, 1753. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jain, S.; Singhania, U.; Tripathy, B.; Nasr, E.A.; Aboudaif, M.K.; Kamrani, A.K. Deep Learning-Based Transfer Learning for Classifi-cation of Skin Cancer. Sensors 2021, 21, 8142. [Google Scholar] [CrossRef]
- Wu, D.; Luo, X.; Shang, M.; He, Y.; Wang, G.; Wu, X. A data characteristic aware latent factor model for web services QOS prediction. IEEE Trans. Knowl. Data Eng. 2022, 34, 2525–2538. [Google Scholar] [CrossRef]
- Wu, D.; He, Q.; Luo, X.; Shang, M.; He, Y.; Wang, G. A posterior neighborhood regularized latent factor model for highly accurate web service QOS prediction. IEEE Trans. Serv. Comput. 2022, 15, 793–805. [Google Scholar] [CrossRef]




| Batch Size | Optimizer | Dense | Learning Rate | Weight Decay Values | Epoch | Loss | Processing Time (ms) |
|---|---|---|---|---|---|---|---|
| 32 | RMSProp | 4 | 0.0001 | 0.0001 | 50 | 6.50 | 4 |
| RMSProp | 4 | 0.0001 | 0.001 | 100 | 6.56 | 5 | |
| RMSProp | 4 | 0.001 | 0.0001 | 50 | 6.25 | 4 | |
| RMSProp | 4 | 0.001 | 0.001 | 100 | 6.57 | 5 | |
| 64 | RMSProp | 4 | 0.0001 | 0.0001 | 50 | 7.27 | 5 |
| RMSProp | 4 | 0.0001 | 0.001 | 100 | 7.37 | 6 | |
| RMSProp | 4 | 0.001 | 0.0001 | 50 | 7.52 | 5 | |
| RMSProp | 4 | 0.001 | 0.001 | 100 | 7.79 | 7 | |
| 32 | RMSProp | 5 | 0.0001 | 0.0001 | 50 | 8.46 | 6 |
| RMSProp | 5 | 0.0001 | 0.001 | 100 | 8.63 | 7 | |
| RMSProp | 5 | 0.001 | 0.0001 | 50 | 8.21 | 6 | |
| RMSProp | 5 | 0.001 | 0.001 | 100 | 8.35 | 7 | |
| 64 | RMSProp | 5 | 0.0001 | 0.0001 | 50 | 8.32 | 8 |
| RMSProp | 5 | 0.0001 | 0.001 | 100 | 8.34 | 7 | |
| RMSProp | 5 | 0.001 | 0.0001 | 50 | 8.25 | 7 | |
| RMSProp | 5 | 0.001 | 0.001 | 100 | 8.31 | 7 | |
| 32 | ADAM | 4 | 0.0001 | 0.0001 | 50 | 6.26 | 3 |
| ADAM | 4 | 0.0001 | 0.001 | 100 | 6.28 | 4 | |
| ADAM | 4 | 0.001 | 0.0001 | 50 | 6.27 | 4 | |
| ADAM | 4 | 0.001 | 0.001 | 100 | 6.55 | 5 | |
| 64 | ADAM | 4 | 0.0001 | 0.0001 | 50 | 7.04 | 4 |
| ADAM | 4 | 0.0001 | 0.001 | 100 | 7.06 | 5 | |
| ADAM | 4 | 0.001 | 0.0001 | 50 | 7.26 | 4 | |
| ADAM | 4 | 0.001 | 0.001 | 100 | 7.27 | 6 | |
| 32 | ADAM | 5 | 0.0001 | 0.0001 | 50 | 7.67 | 4 |
| ADAM | 5 | 0.0001 | 0.001 | 100 | 7.63 | 5 | |
| ADAM | 5 | 0.001 | 0.0001 | 50 | 7.21 | 4 | |
| ADAM | 5 | 0.001 | 0.001 | 100 | 7.35 | 6 | |
| 64 | ADAM | 5 | 0.0001 | 0.0001 | 50 | 8.02 | 4 |
| ADAM | 5 | 0.0001 | 0.001 | 100 | 8.14 | 5 | |
| ADAM | 5 | 0.001 | 0.0001 | 50 | 8.05 | 5 | |
| ADAM | 5 | 0.001 | 0.001 | 100 | 8.10 | 6 | |
| 32 | AdaGrad | 4 | 0.0001 | 0.0001 | 50 | 6.47 | 4 |
| AdaGrad | 4 | 0.0001 | 0.001 | 100 | 6.74 | 4 | |
| AdaGrad | 4 | 0.001 | 0.0001 | 50 | 6.25 | 5 | |
| AdaGrad | 4 | 0.001 | 0.001 | 100 | 6.55 | 5 | |
| 64 | AdaGrad | 4 | 0.0001 | 0.0001 | 50 | 7.14 | 5 |
| AdaGrad | 4 | 0.0001 | 0.001 | 100 | 7.06 | 6 | |
| AdaGrad | 4 | 0.001 | 0.0001 | 50 | 7.16 | 6 | |
| AdaGrad | 4 | 0.001 | 0.001 | 100 | 7.29 | 7 | |
| 32 | AdaGrad | 5 | 0.0001 | 0.0001 | 50 | 7.77 | 5 |
| AdaGrad | 5 | 0.0001 | 0.001 | 100 | 7.61 | 6 | |
| AdaGrad | 5 | 0.001 | 0.0001 | 50 | 7.23 | 6 | |
| AdaGrad | 5 | 0.001 | 0.001 | 100 | 7.32 | 7 | |
| 64 | AdaGrad | 5 | 0.0001 | 0.0001 | 50 | 8.06 | 5 |
| AdaGrad | 5 | 0.0001 | 0.001 | 100 | 8.18 | 5 | |
| AdaGrad | 5 | 0.001 | 0.0001 | 50 | 8.09 | 6 | |
| AdaGrad | 5 | 0.001 | 0.001 | 100 | 8.11 | 7 | |
| 32 | Adadelta | 4 | 0.0001 | 0.0001 | 50 | 6.69 | 4 |
| Adadelta | 4 | 0.0001 | 0.001 | 100 | 6.56 | 5 | |
| Adadelta | 4 | 0.001 | 0.0001 | 50 | 6.28 | 4 | |
| Adadelta | 4 | 0.001 | 0.001 | 100 | 6.47 | 4 | |
| 64 | Adadelta | 4 | 0.0001 | 0.0001 | 50 | 6.85 | 4 |
| Adadelta | 4 | 0.0001 | 0.001 | 100 | 7.44 | 4 | |
| Adadelta | 4 | 0.001 | 0.0001 | 50 | 7.16 | 5 | |
| Adadelta | 4 | 0.001 | 0.001 | 100 | 7.26 | 5 | |
| 32 | Adadelta | 5 | 0.0001 | 0.0001 | 50 | 7.67 | 6 |
| Adadelta | 5 | 0.0001 | 0.001 | 100 | 7.77 | 6 | |
| Adadelta | 5 | 0.001 | 0.0001 | 50 | 7.73 | 5 | |
| Adadelta | 5 | 0.001 | 0.001 | 100 | 7.31 | 7 | |
| 64 | Adadelta | 5 | 0.0001 | 0.0001 | 50 | 7.55 | 6 |
| Adadelta | 5 | 0.0001 | 0.001 | 100 | 8.08 | 7 | |
| Adadelta | 5 | 0.001 | 0.0001 | 50 | 8.19 | 5 | |
| Adadelta | 5 | 0.001 | 0.001 | 100 | 8.12 | 6 |
| Classifier | Accuracy (%) | Sensitivity (%) | Specificity (%) |
|---|---|---|---|
| DCN transfer learning [29] | 94.92 | 80.36 | 79.8 |
| Mobile Net [30] | 83.1 | 89 | 83 |
| Kernel extreme learning machine [31] | 90.67 | 90.20 | 89.43 |
| DilatInceptV3 [32] | 90.10 | 87 | 87 |
| Proposed | 99.75 | 100 | 100 |
| Project | Accuracy (%) | Sensitivity (%) | Specificity (%) |
|---|---|---|---|
| Gessert et al. [33] | 98.70 | 80.9 | 98.4 |
| Ailin et al. [34] | 98.20 | 89.5 | 98.1 |
| Khan et al. [35] | 89.80 | 89.7 | 94.5 |
| Mohamed et al. [36] | 92.70 | 72.42 | 97.14 |
| Huang et al. [37] | 85.80 | 69.04 | 95.92 |
| Liu et al. [38] | 92.54 | 71.47 | 92.72 |
| Gu et al. [39] | 91.4 | 83.74 | 93.24 |
| Zhou et al. [40] | 92.55 | 84.67 | 93.63 |
| Gan et al. [41] | 93.81 | 90.14 | 98.36 |
| Proposed | 99.78 | 100 | 100 |
| Project | Accuracy (%) | Sensitivity (%) | Specificity (%) |
|---|---|---|---|
| Gessert et al. [33] | 92.3 | 80.9 | 98.4 |
| Ailin et al. [34] | 91.5 | 89.5 | 98.1 |
| Ahmed et al. [42] | 94 | 89.7 | 94.5 |
| Pacheco et al. [43] | 92 | 72.42 | 97.14 |
| Molina et al. [44] | 97 | 69.04 | 95.92 |
| Kaseem et al. [45] | 94 | 71.47 | 92.72 |
| Iqbla et al. [46] | 90 | 83.74 | 93.24 |
| Pulgarin et al. [47] | 92 | 89.53 | 93.57 |
| Proposed | 99.51 | 100 | 100 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bhimavarapu, U.; Battineni, G. Skin Lesion Analysis for Melanoma Detection Using the Novel Deep Learning Model Fuzzy GC-SCNN. Healthcare 2022, 10, 962. https://doi.org/10.3390/healthcare10050962
Bhimavarapu U, Battineni G. Skin Lesion Analysis for Melanoma Detection Using the Novel Deep Learning Model Fuzzy GC-SCNN. Healthcare. 2022; 10(5):962. https://doi.org/10.3390/healthcare10050962
Chicago/Turabian StyleBhimavarapu, Usharani, and Gopi Battineni. 2022. "Skin Lesion Analysis for Melanoma Detection Using the Novel Deep Learning Model Fuzzy GC-SCNN" Healthcare 10, no. 5: 962. https://doi.org/10.3390/healthcare10050962
APA StyleBhimavarapu, U., & Battineni, G. (2022). Skin Lesion Analysis for Melanoma Detection Using the Novel Deep Learning Model Fuzzy GC-SCNN. Healthcare, 10(5), 962. https://doi.org/10.3390/healthcare10050962