1. Introduction
Severe acute respiratory syndrome coronavirus 2, also known as SARS-CoV-2, is reported as a virus strain causing the respiratory disease of COVID-19 [
1]. The World Health Organization (WHO) and the global nations confirmed the coronavirus disease to be extremely contagious [
2,
3]. The COVID-19 pandemic has been widely recognized as a public health emergency of international concern [
4]. To estimate the outbreak, identify the peak ahead of time, and also predict the mortality rate, epidemiological models have been widely used by officials and media. Outbreak prediction models have shown to be fundamental to provide insights into the damages caused by COVID-19. Furthermore, the prediction models are used as a reference to make new policies and to evaluate the conditions of curfew [
5]. The COVID-19 pandemic has been reported to be extremely aggressive to spread [
6]. Due to the uncertainty and complexity of the COVID-19 outbreak and its irregularity in different countries, the standard epidemiological models, i.e., susceptible-infected-resistant (SIR)-based models, have been challenged for delivering higher performance in individual nations. Furthermore, as the COVID-19 outbreak showed significant differences with the recent epidemics, e.g., swine fever, H1N1 influenza, Ebola, Cholera, dengue fever, and Zika, several advanced epidemiological models have emerged to provide higher accuracy [
7]. Nevertheless, due to the involvement of numerous unknown factors, the outbreaks, various curfew strategies in different countries, and existing double standards for evaluation metrics, the model uncertainty for the COVID-19 is generally reported extensive [
8,
9,
10]. As a result, SIR-based models are fundamentally challenged to provide reliable insight into the progress of COVID-19.
The general strategy behind the SIR-based models for predicting the COVID-19 outbreak, similar to other epidemics, is formed around the assumption of transmitting the contagious disease through social contacts. The SIR-based models assume that infection spreads through several groups, e.g., susceptible, exposed, infected, recovered, deceased, and immune [
11]. For instance, the standard SIR-based models assume that an epidemic includes susceptible to infection (class
S), infected (class
I), and the removed population (class
R). Such principal groups build the foundation of epidemiological modeling. Note that the definition of various classes of the outbreak may vary. For instance,
R is often referred to as those that have recovered, developed immunity, been isolated, or passed away. However, in various countries,
R may or may not be susceptible to infection again, and there exist uncertainties in allocating
R a value. Advancing SIR-based models require several assumptions. Thus, modeling with SIR-based models may include several contradicting assumptions.
Nevertheless, in most epidemiological models, it is generally agreed that class
I has a high probability of infecting class
S. The transmission probability is proportional to the total social contacts. The transmission can be estimated through the implementation of basic differential equations as follows [
12,
13,
14].
where
,
, and
represent the infected population, the susceptible population, and the daily reproduction rate, respectively. The time-series of
which is calculated by the differential equation, decreases gradually. However, it is observed that at the beginning of the pandemics, where the increment
is linear,
. Eventually, class
I is estimated as follows.
where
represents the parameter of the daily rate of spread. Furthermore, the susceptible individuals excluded from the SIR-based models are calculated. Furthermore, the class
R, representing individuals excluded from the spread of infection, is computed as follows:
Considering the above-mentioned assumptions, and under the unconstrained conditions of the excluded group, Equation (3), the outbreak modeling with SIR is finally stated as:
Furthermore, to evaluate the accuracy of the SIR-based models, the outbreak model’s median success function is used, which is represented as:
Several analytical solutions to the SIR models have been provided in the literature [
15,
16]. As the different nations take different actions toward slowing down the outbreak, the SIR-based model must be adapted according to the local assumptions [
17]. The inaccuracy of many SIR-based models in predicting the outbreak and mortality rate has been evidenced during the COVID-19 in many nations. The critical success of a SIR-based model relies on choosing the right model according to the context and the relevant assumptions. SIS (susceptible-infectious-susceptible), SIRD (susceptible-infected-recovered-deceased-model), MSIR (Maternally-derived-immunity-susceptible-infected-recovered), SEIR (Susceptible-exposed-infected-recovered), SEIS (Susceptible-exposed-infected- susceptible), MSEIR (Maternally-derived-immunity-susceptible-exposed-infected-recovered), and MSEIRS (Maternally-derived-immunity-susceptible- exposed-infected-recovered-susceptible) models are among the popular models used for predicting COVID-19 outbreaks worldwide. The more advanced variation of SIR-d models carefully considers the vital dynamics and constant population [
16]. For instance, at the presence of the long-lasting immunity assumption when the immunity is not realized upon recovery from infection, the susceptible-infectious-susceptible (SIS) model was suggested [
18]. In contrast, the susceptible-infected-recovered-deceased-model (SIRD) is used when immunity is assumed [
19]. In the case of COVID-19, different nations took different approaches in this regard.
SEIR models have been reported among the most popular tools to predict the notable outbreaks. To advance an SEIR model, often, the incubation period of an infected person is carefully estimated to achieve more accurate predictions. In the case of Varicella and Zika outbreaks, the SEIR models showed increased model accuracy [
20,
21]. To do so, SEIR models might assign the incubation period to a random variable. Furthermore, similar to the standard SIR-based models, SEIR models work on the ideology of the disease-free-equilibrium [
22,
23]. However, it should be noted that SEIR models can not fit well where the contact network is non-stationary through time [
24]. Social mixing as a critical factor of non-stationarity determines the reproductive number
which is the number of susceptible individuals for infection. The value of
for COVID-19 was estimated to be 4, which greatly trigged the pandemic [
1]. The lockdown measures aimed at reducing the
value down to 1. Nevertheless, the SEIR models are reported to be difficult to fit in the case of COVID-19 due to the non-stationarity of mixing caused by nudging intervention measures. Therefore, to develop more accurate SIR-based models, in-depth information about the social movement and the quality of lockdown measures would be essential. Another drawback of SIR-based models is the short lead-time. For long-term prediction, the accuracy of the most SIR-based models declines. For the COVID-19 outbreak case of Italy, for instance, the accuracy of the model drops significantly. The performance of
for the lead-time of 120 h reduces to
for 144 h lead time [
17]. Overall, the SIR-based models would be accurate if firstly the status of social interactions is stable. Secondly, class
R can be computed precisely. To better estimate class
R, several data sources can be integrated with SIR-based models, e.g., CCTVs, social media, mobile apps, and call data records. Nevertheless, using such systems are reported to still associate with significant complexity and uncertainties [
25,
26,
27,
28,
29,
30,
31,
32]. Due to the high level of uncertainties involved in the advancement of SIR-based models, the generalization ability is yet to be improved to achieve a scalable model with high performance [
33].
Due to the presence of uncertainties and a high degree of complexity in advancing epidemiological models of the outbreak, machine learning has increasingly been seen as a potential technology. ML has already shown promising results in a contribution to developing better SIR-based models with higher performance with generalization ability and robustness [
34,
35,
36,
37,
38]. Machine learning has already been recognized as a computing technique with great potential in outbreak prediction. The notable machine learning algorithms include, e.g., random forest for swine fever [
39,
40], neural networks for H1N1 flu, dengue fever, and Oyster norovirus [
11,
41,
42], genetic programming for Oyster norovirus [
43], classification and regression tree (CART) for Dengue [
44], Bayesian Network for Dengue and Aedes [
45], LogitBoost for Dengue [
46], multi-regression and Naïve Bayes for Dengue outbreak prediction [
47]. Machine learning has often been used as a complementary computation tool to enhance SIR-based models [
11,
39,
40,
41,
42,
43,
44,
45,
46,
47,
48,
49]. Nevertheless, there is a gap in using machine learning in the case of COVID-19. However, several recent research works point out the enormous potential of machine learning for the fight against COVID-19 [
49,
50,
51,
52]. Machine learning delivered promising results in several aspects for mitigation and prevention and have been endorsed in the scientific community for, e.g., case identifications [
53], classification of novel pathogens [
54], modification of SIR-based models [
55], diagnosis [
56,
57], survival prediction [
58], and ICU demand prediction [
59]. Furthermore, the non-peer reviewed sources suggest novel applications for fighting COVID-19 [
60]. Among the applications of machine learning improvement of the existing models of prediction, identifying vulnerable groups, early diagnosis, the advancement of drugs delivery, evaluation of the probability of next pandemic, the advancement of the integrated systems for Spatio-temporal prediction, evaluating the risk of infection, advancing reliable biomedical knowledge graphs, and data mining the social networks are being noted.
As stated in our recent paper, machine learning can be used for data preprocessing. Improving the quality of data can particularly improve the quality of the SIR-based model. For instance, the number of cases reported by Worldometer is not precisely the number of infected cases (
E in the SEIR model), or calculating the number of infectious people (
I in SEIR) cannot be easily determined, as many people who might be infectious may not turn up for testing although the number of people who are admitted to hospital and deceased will not support
R as most COVID-19 positive cases recover without entering the hospital. Considering this data problem, it is challenging to fit SEIR models satisfactorily. Considering such challenges, for future research, the ability of machine learning for estimation of the missing information on the number of exposed
E or infected individuals can be evaluated. Along with the prediction of the outbreak, prediction of the total mortality rate (
n(deaths)/
n(infecteds)) is also essential to accurately estimate the number of potential patients in the critical situation and the required beds in intensive care units. Although the research is in the very early stage, the trend in outbreak prediction with machine learning can be classified in two directions. Firstly, improvement of the SIR-based models, e.g., [
55,
61], and secondly time-series prediction [
62,
63]. Consequently, the state-of-the-art machine learning methods for outbreak modeling suggest two major research gaps for machine learning to address. Firstly, the improvement of SIR-based models and secondly advancement in outbreak time series. Considering the drawbacks of the SIR-based models, machine learning should be able to contribute. This paper contributes to the advancement of time-series modeling and prediction of COVID-19. Although ML has been currently established in predicting several scientific phenomena [
64,
65,
66,
67,
68], its advancement for pandemic modeling remains at the early stages [
69]. More sophisticated ML methods are yet to be explored. A recent paper by Ardabili et al., [
51] explored the potential of MLP and ANFIS in time series prediction of COVID-19 in several countries. The contribution of the present paper is to improve the quality of prediction by proposing a hybrid machine learning and compare the results with ANFIS. In the present paper, the time series of the total mortality is also included. This article continues as follows.
Section 2 describes the methods and materials. The results are given in
Section 3.
Section 4 presents the conclusions.
5. Conclusions
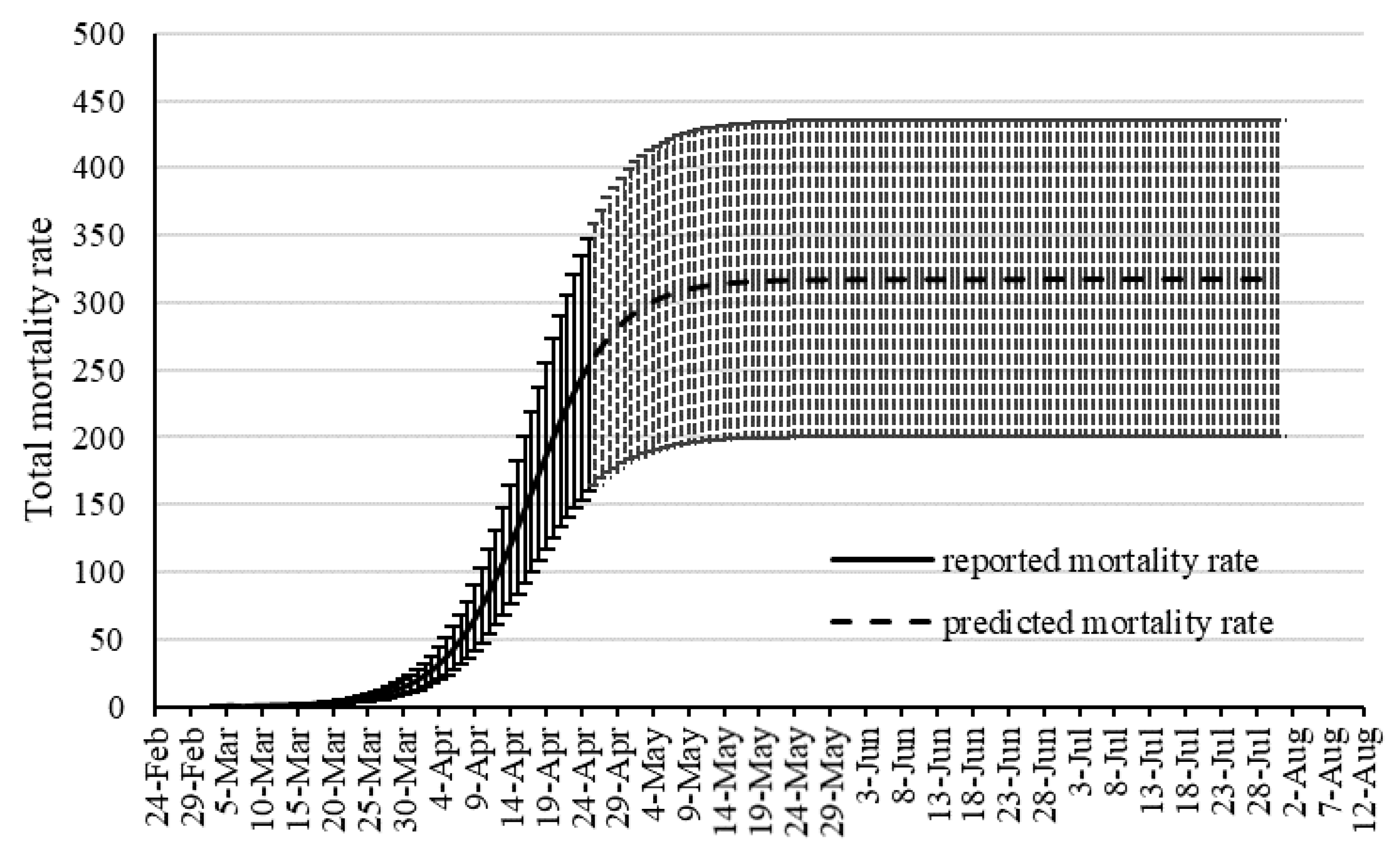
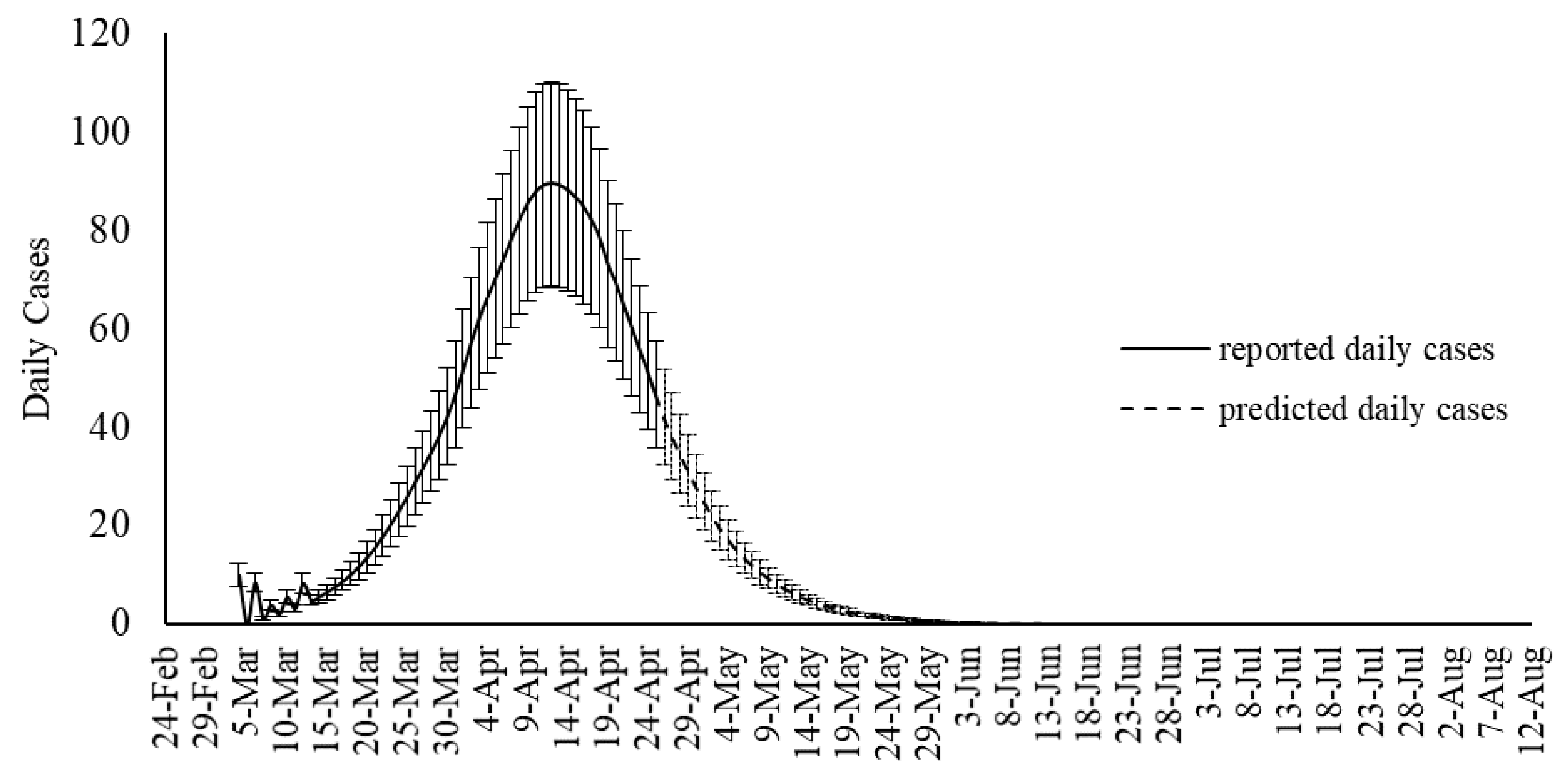
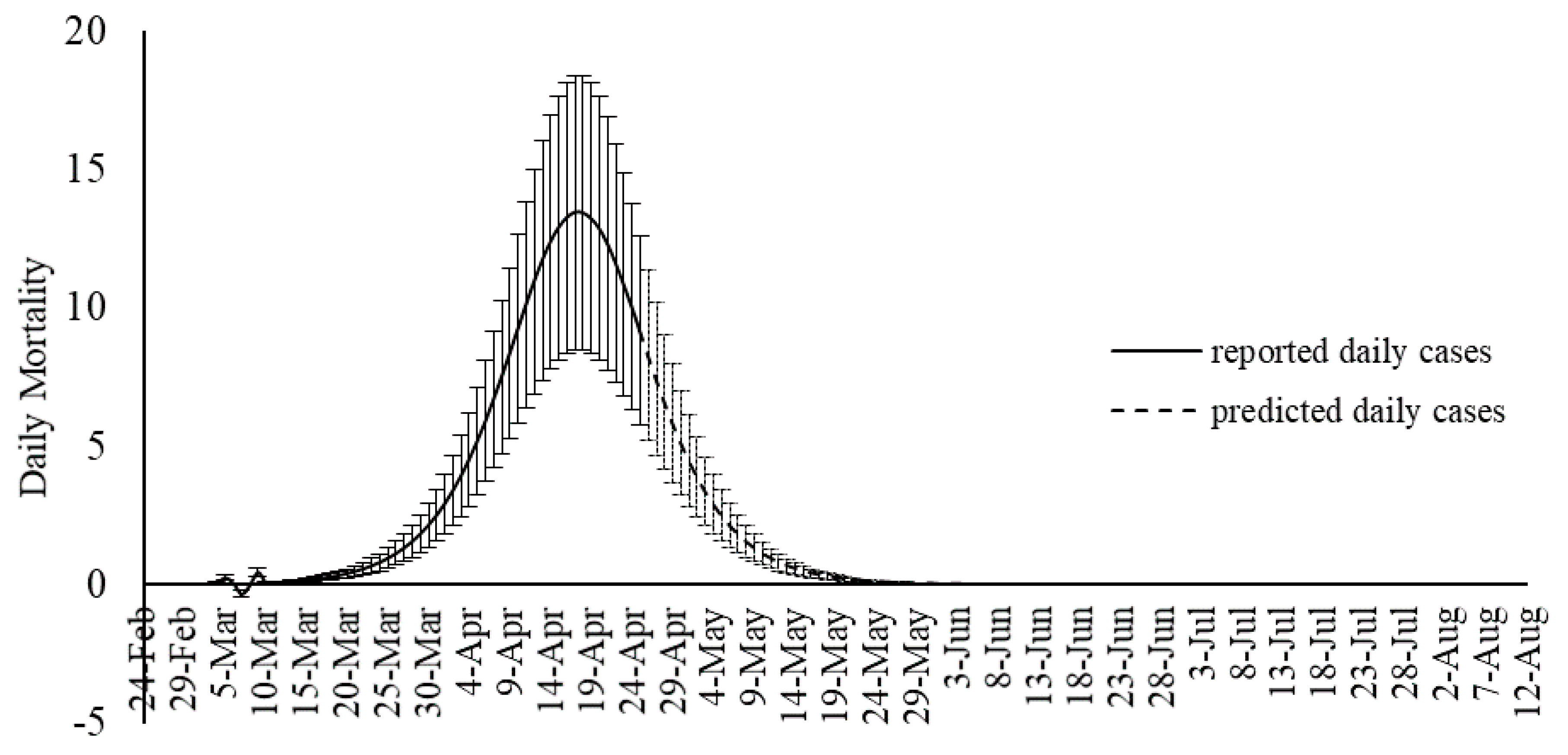
Although SIR-based models been widely used for modeling the COVID-19 outbreak, they include some degree of uncertainties. Several advancements are emerging to improve the quality of SIR-based models suitable to the COVID-19 outbreak. As an alternative to the SIR-based models, this study proposed machine learning as a new trend in advancing outbreak models. The machine learning approach makes no assumption on the pandemic and spread of the infection. Instead, it predicts the time series of the infected cases as well as total mortality cases. In this study, the hybrid machine learning model of MLP-ICA and ANFIS is used to predict the COVID-19 outbreak in Hungary. The models predict that by late May, the outbreak and the total morality will drop substantially. Based on the promising results reported in this study, and due to the complex phenomenon of COVID-19 outbreak, this study, as an alternative modeling strategy, suggests machine learning as a potential technology to be considered to model the outbreak. However, further research would be essential to validate the results and improve the quality of prediction.
In this study, two scenarios were proposed for sampling the data. Scenario 1 considered sampling the odd days, and Scenario 2 used even days for training the data. Training the two machine learning models of ANFIS and MLP-ICA, were considered through using two scenarios. It is concluded that using different scenarios for data sampling has a minimum effect on the model performance. A detailed investigation was carried out to explore the most suitable number of neurons. Furthermore, the performance of the proposed algorithm is evaluated using both training and validation data. The training data are used to train the algorithm and define the best set of parameters to be used in ANFIS and MLP-ICA. After that, the best setup for each algorithm is used to predict outbreaks on the validation samples. The validation is performed for 9 days with promising results, which confirms the model accuracy. In this study, due to the lack of adequate sample data to avoid overfitting, the training is used to choose and evaluate the model with higher performance. In future research, as the COVID-19 progresses in time and with the availability of more sample data, further testing and validation can be used to better evaluate the models.
Both models showed promising results in terms of predicting the time series without the assumptions that epidemiological models require. Both machine learning models, as an alternative to epidemiological models, showed potential in predicting COVID-19 outbreak as well as estimating total mortality. Yet, MLP-ICA outperformed ANFIS with delivering accurate results on validation samples. Considering the availability of a small amount of training data, further investigation would be essential to explore the true capability of the proposed hybrid model. It is expected that the model maintains its accuracy as long as no major interruption occurs. For instance, if other outbreaks would initiate in the other cities, or the prevention regime changes, naturally the model will not maintain its accuracy. For future studies, advancing deep learning and deep reinforcement learning models is strongly encouraged for comparative studies on various ML models for individual countries.