A Novel Computational Instrument Based on a Universal Mixture Density Network with a Gaussian Mixture Model as a Backbone for Predicting COVID-19 Variants’ Distributions
Abstract
1. Introduction
- (1)
- The number of compartments or stages (M);
- (2)
- The forward and backward inner flows between the compartments;
- (3)
- The weighting parameters;
- (4)
- The number of numerical computation libraries.
- (1)
- regional density distribution;
- (2)
- age categories;
- (3)
- social distancing and masking;
- (4)
- vaccination;
- (5)
- climate data;
- (6)
- asymptomatic infection;
- (7)
- healthcare resources.
2. COVID-19 Input Data Modes
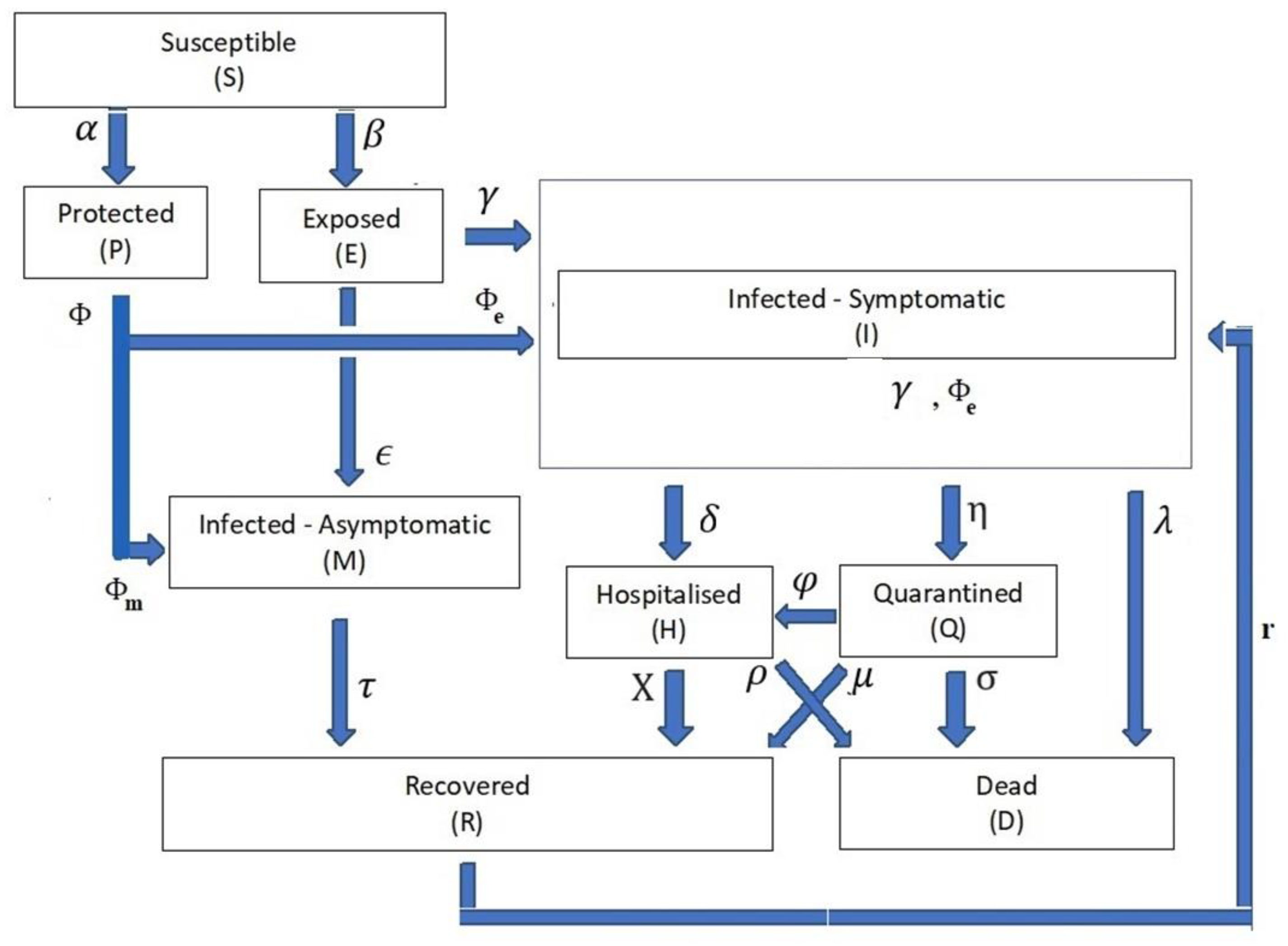
- One in which the time-dependent variable of the infection compartment’s population is Z(t), at which Z(t) {S(t), P(t), E(t), I(t), M(t), H(t), Q(t), D(t), R(t)}. Table 1 exhibits the definitions of the mentioned time-dependent variables. For the rest of this communication, the time-independent notation for the compartment rate variables is dropped to facilitate reading, when necessary.
- The compartment parameter variable ψ is defined as ψ {α, β, є, ϒ, δ, η, λ, μ, χ, σ, ρ, φ}. Table 2 exhibits the definitions of these parameters as the weights from the input compartment to the target compartment.
3. Mixture Density Network (MDN) with Multiple Outputs
3.1. Neural Networks (NNs): An Introductory Overview
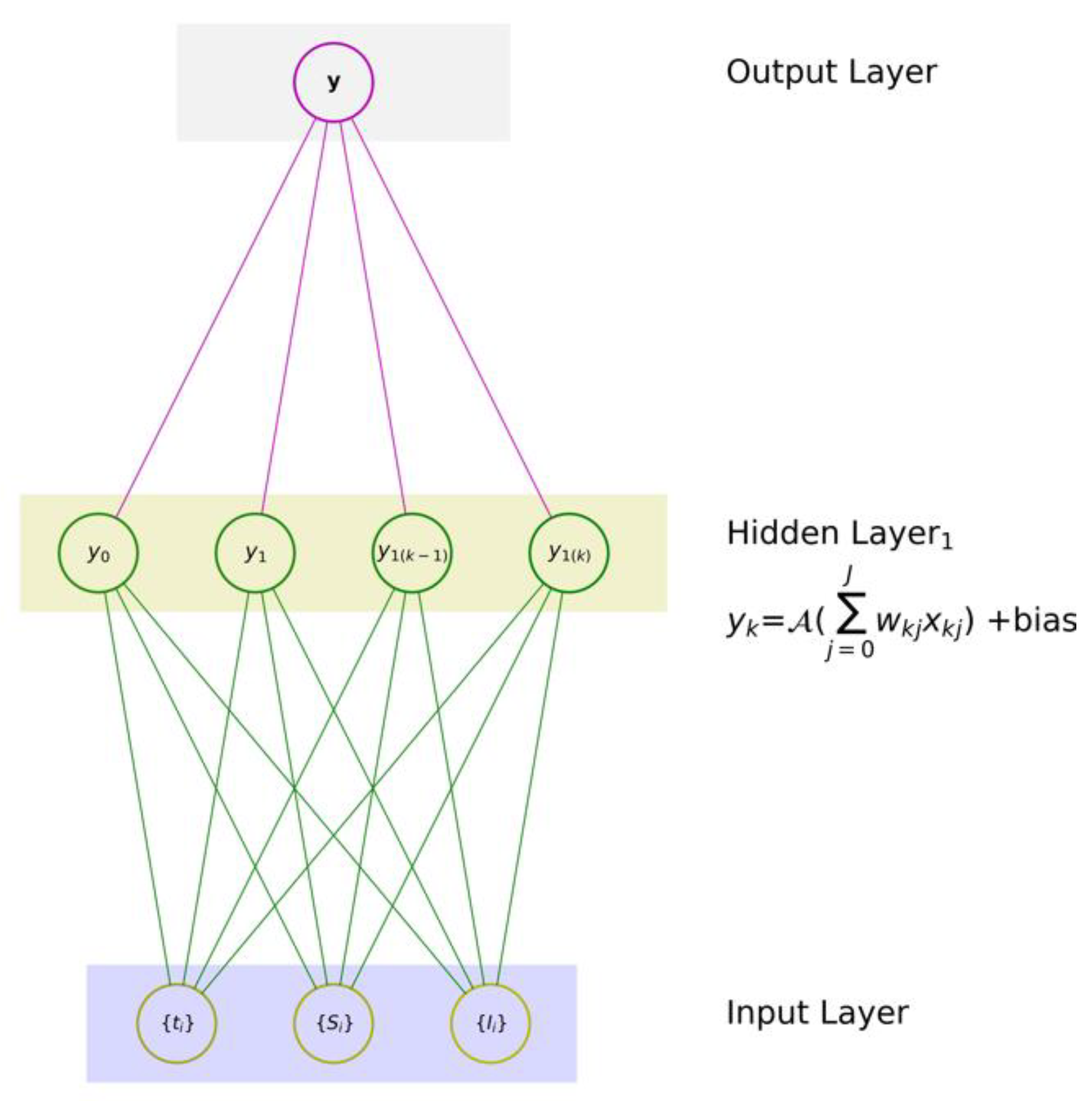
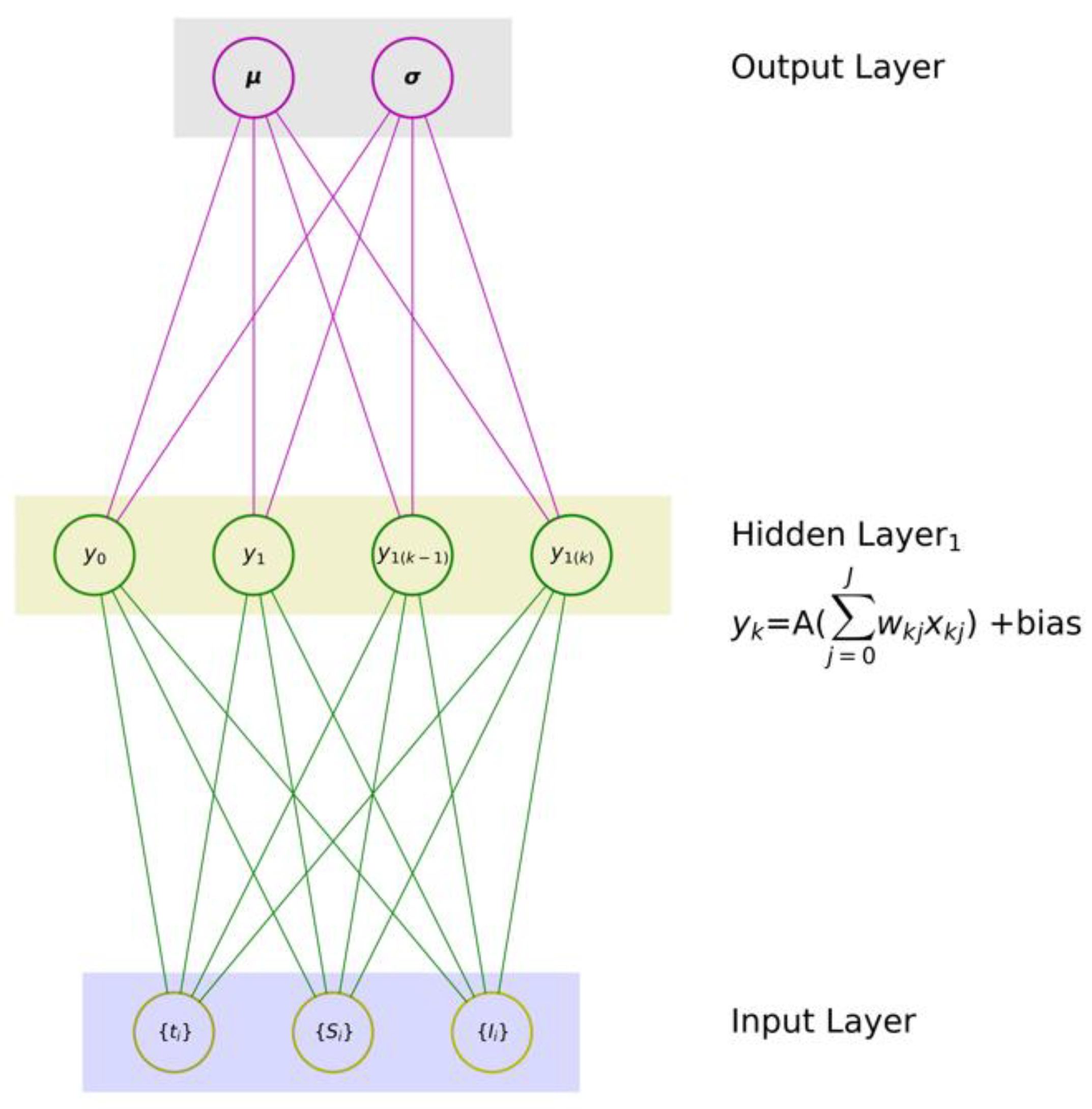
- (1)
- one input layer that digests a given set of input variables x ≡ {x1, x2, …, xJ}, where J is the number of input features,
- (2)
- one hidden layer with K neurons, and
- (3)
- one output layer with one neuron to produce an associated mapping y.
3.2. Mixture Density Network (MDN): The Theory
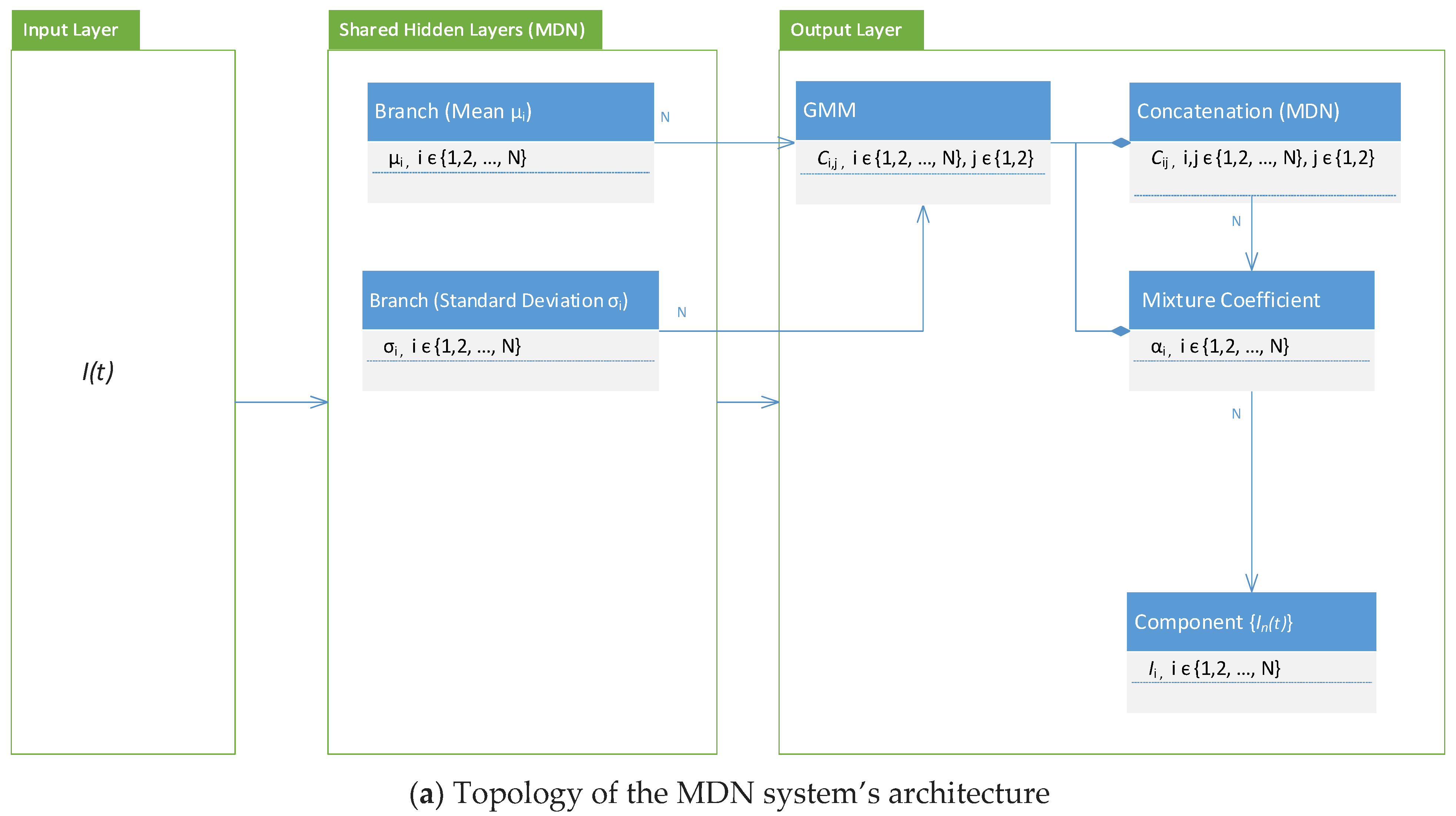
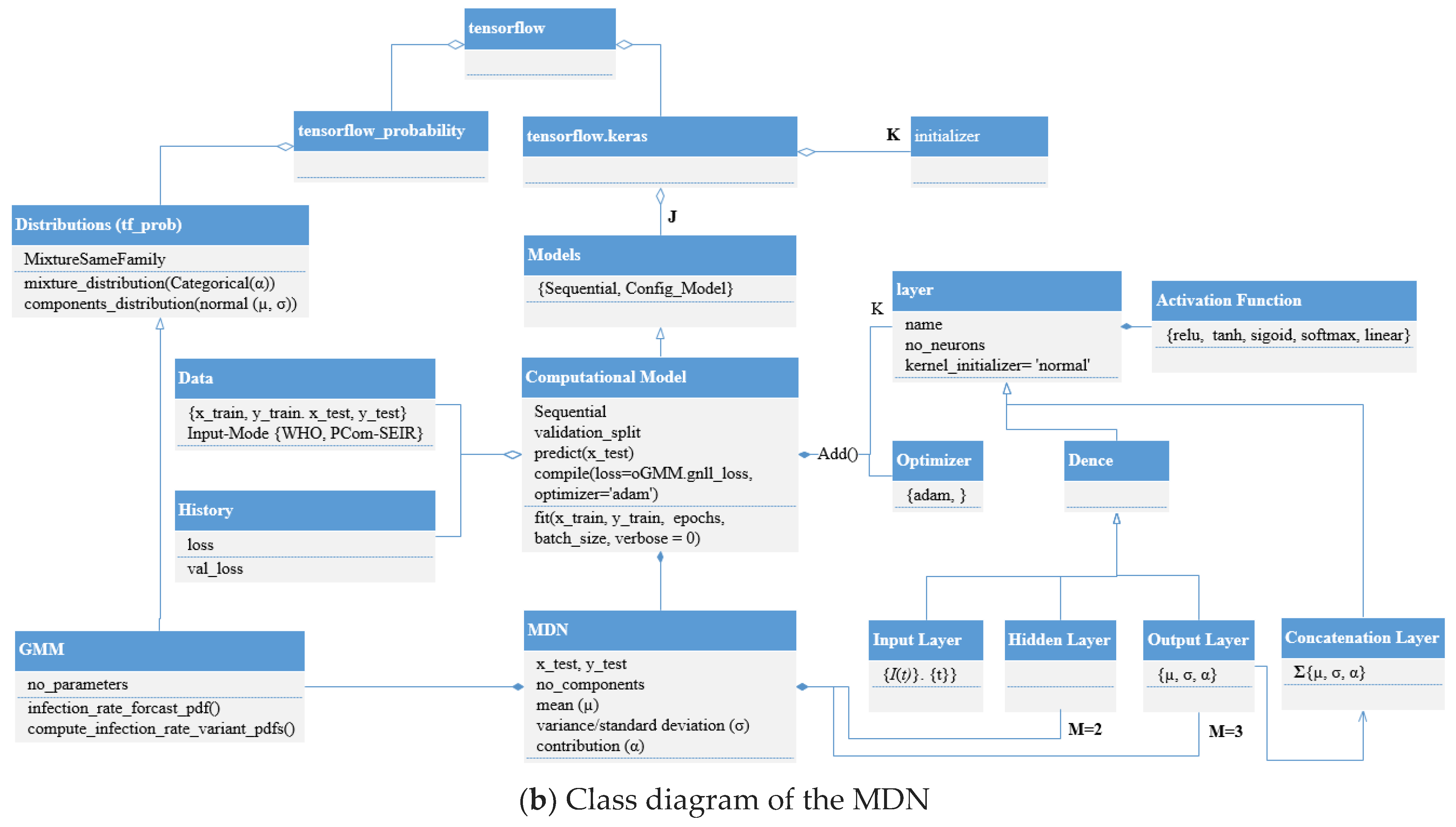
3.3. Architecture, Design, and Implementation Notes
4. Results and Discussion
4.1. Use Case Implementation Dictionary
- (1)
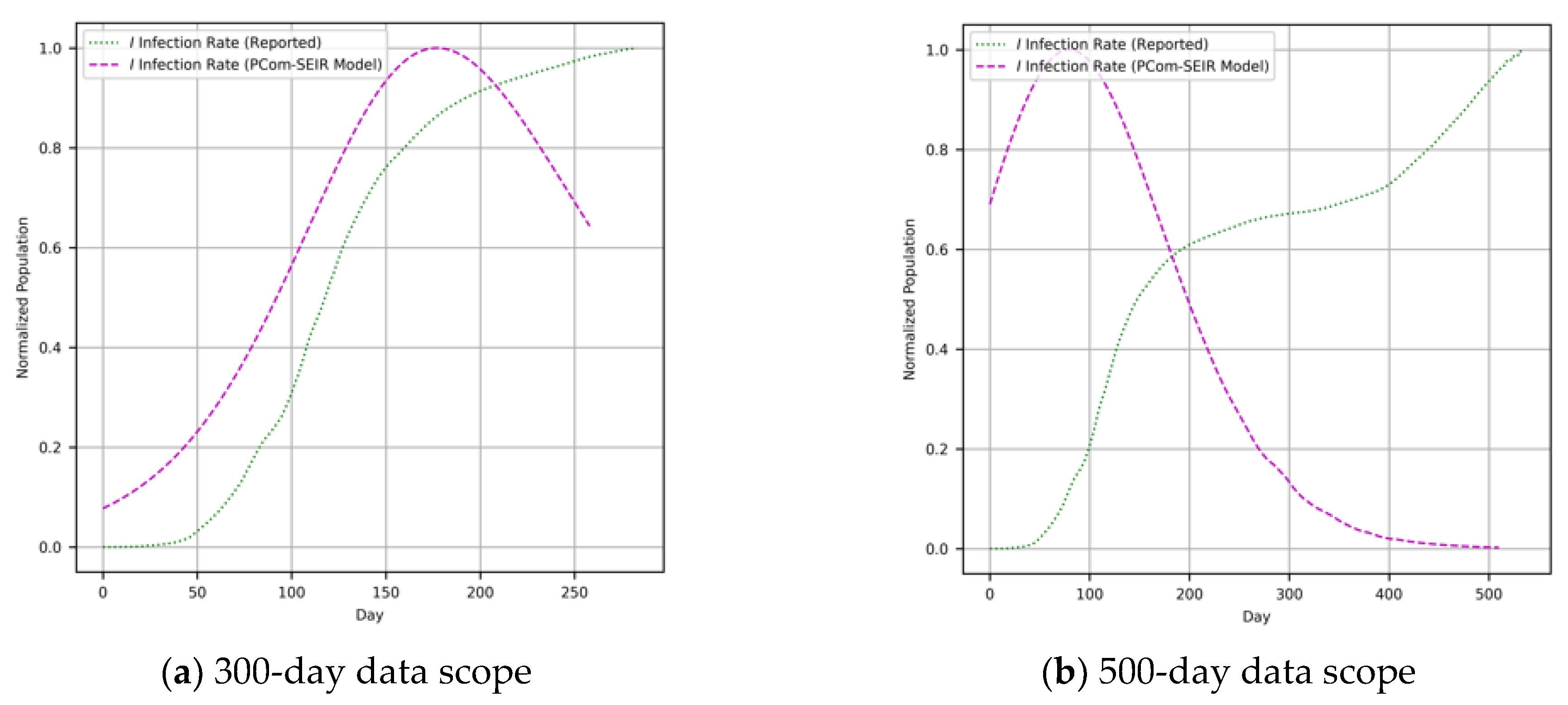
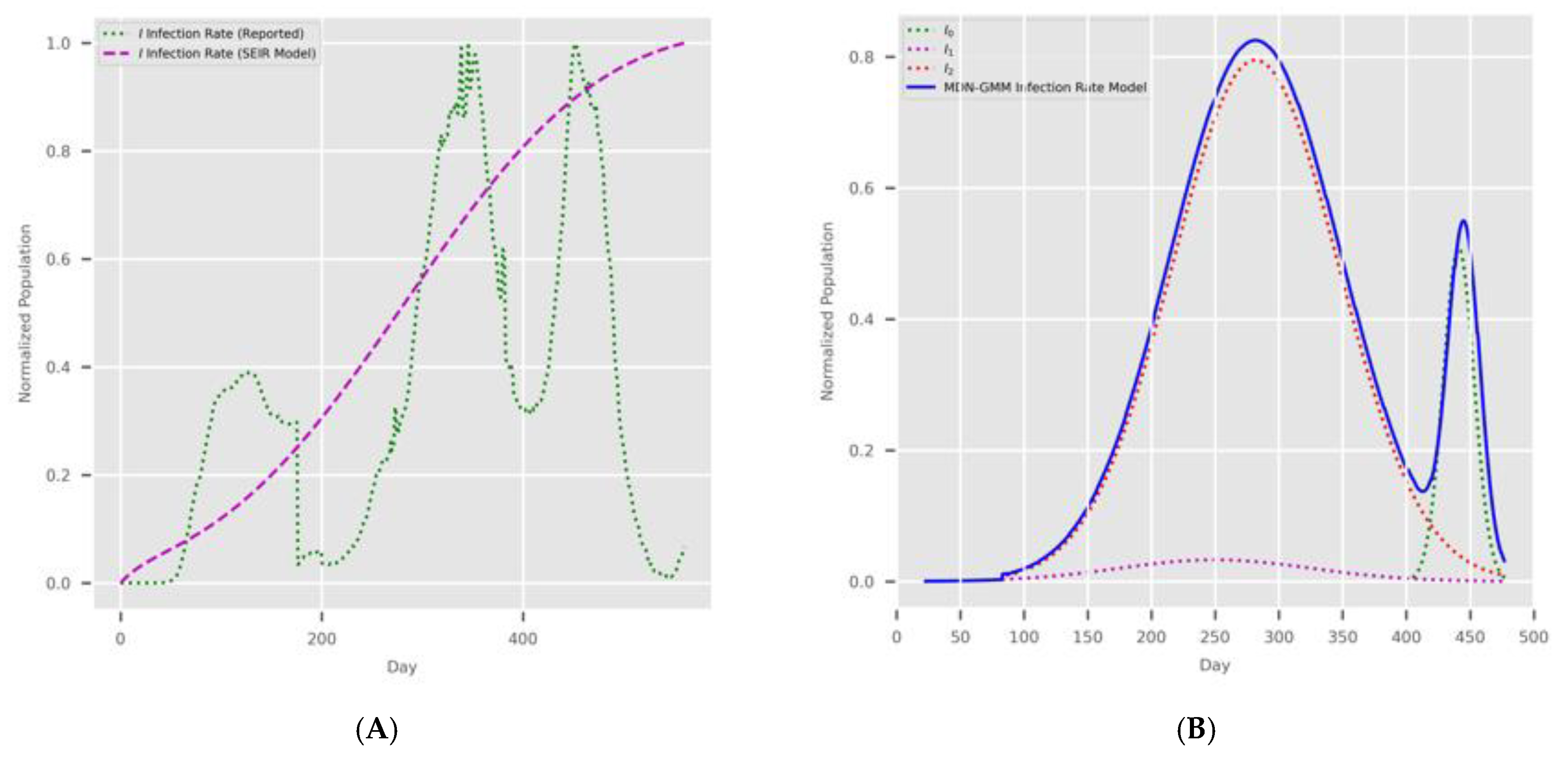
- Input Modes: In Figure 1, the diagram shows the deployment of the two implementations depending on the type of COVID-19 input data for Canada and Saudi Arabia. The first input feed was raw COVID-19 data from the WHO [37]; hence, we named this source “WHO COVID-19 data”. The second input feed was the optimal solution of the SODE in Equations (1)–(9), which was derived in Section 2. The results are depicted in Figure 4 (for Canada) and Figure 5 (for Saudi Arabia). In this study, we named this input feed “PCom-SEIR” because it relied on a modification of the PCom-SEIR model.
- (2)
- Data Ranges: The two input feeds covered two spans: 300 and 500 days.
- (3)
- The MDN implementation runs: There were three implementation runs; each run involved the utilization of one activation function in the hidden layers. The activation function set was {relu, tanh, sigmoid}.
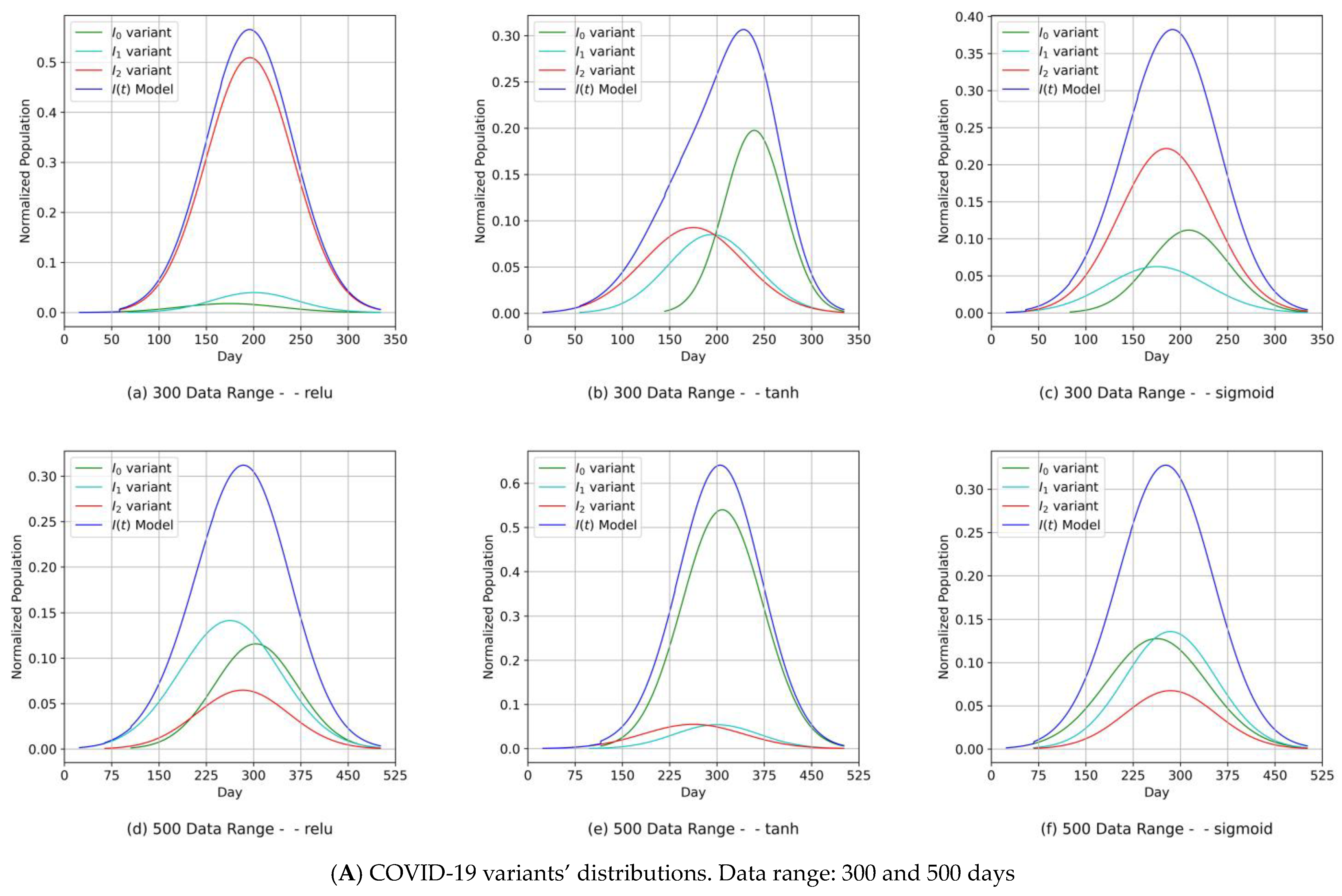
- (1)
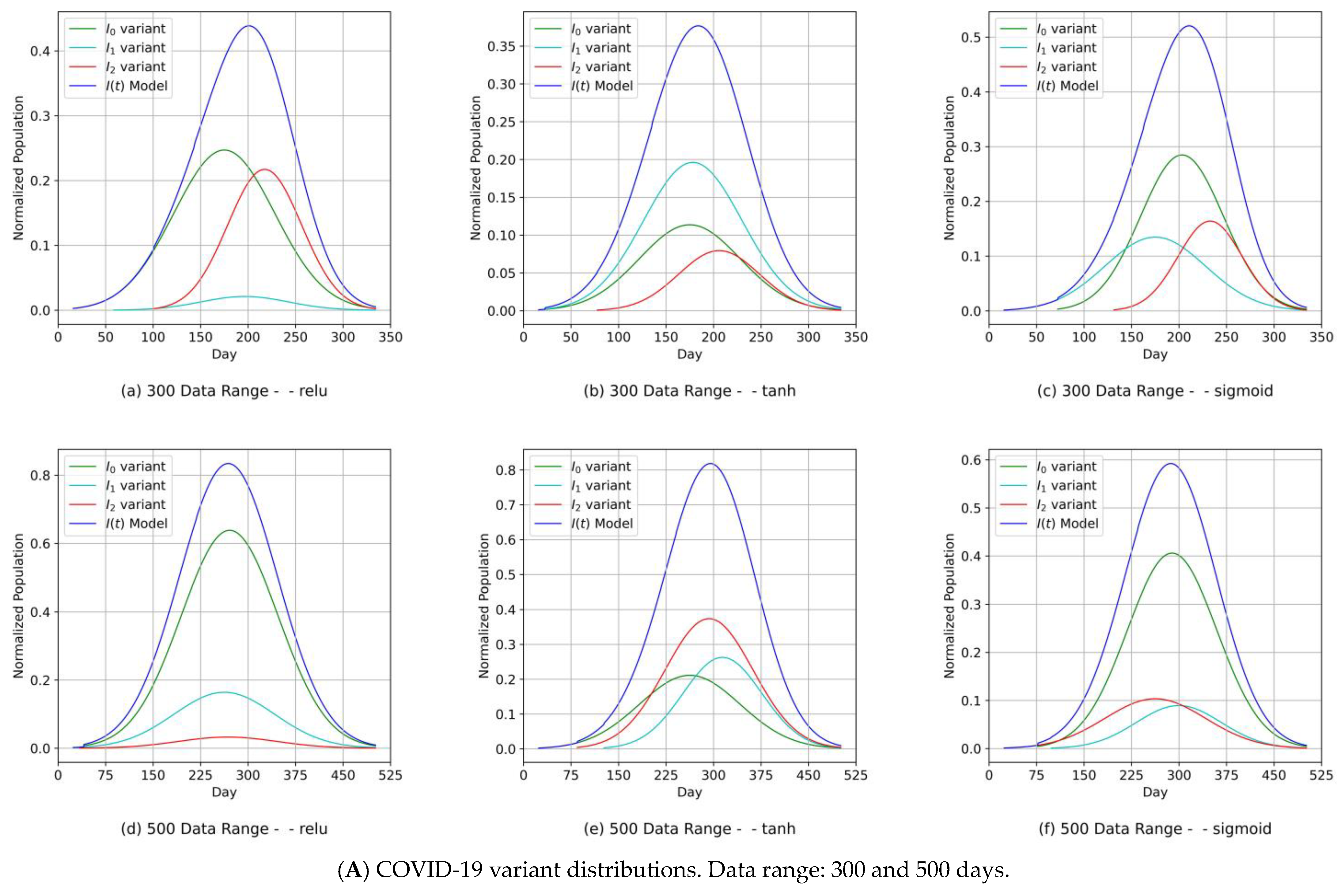
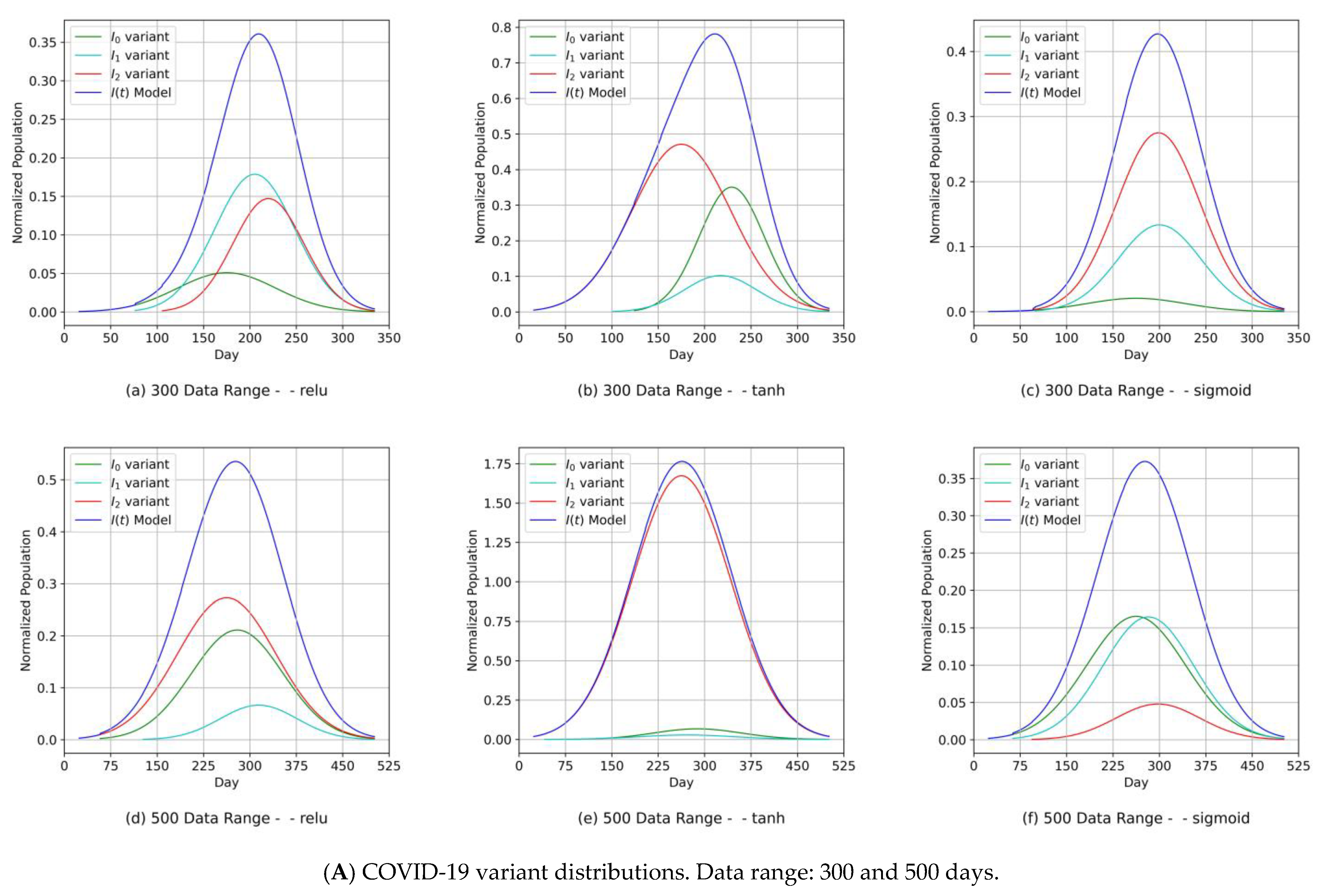
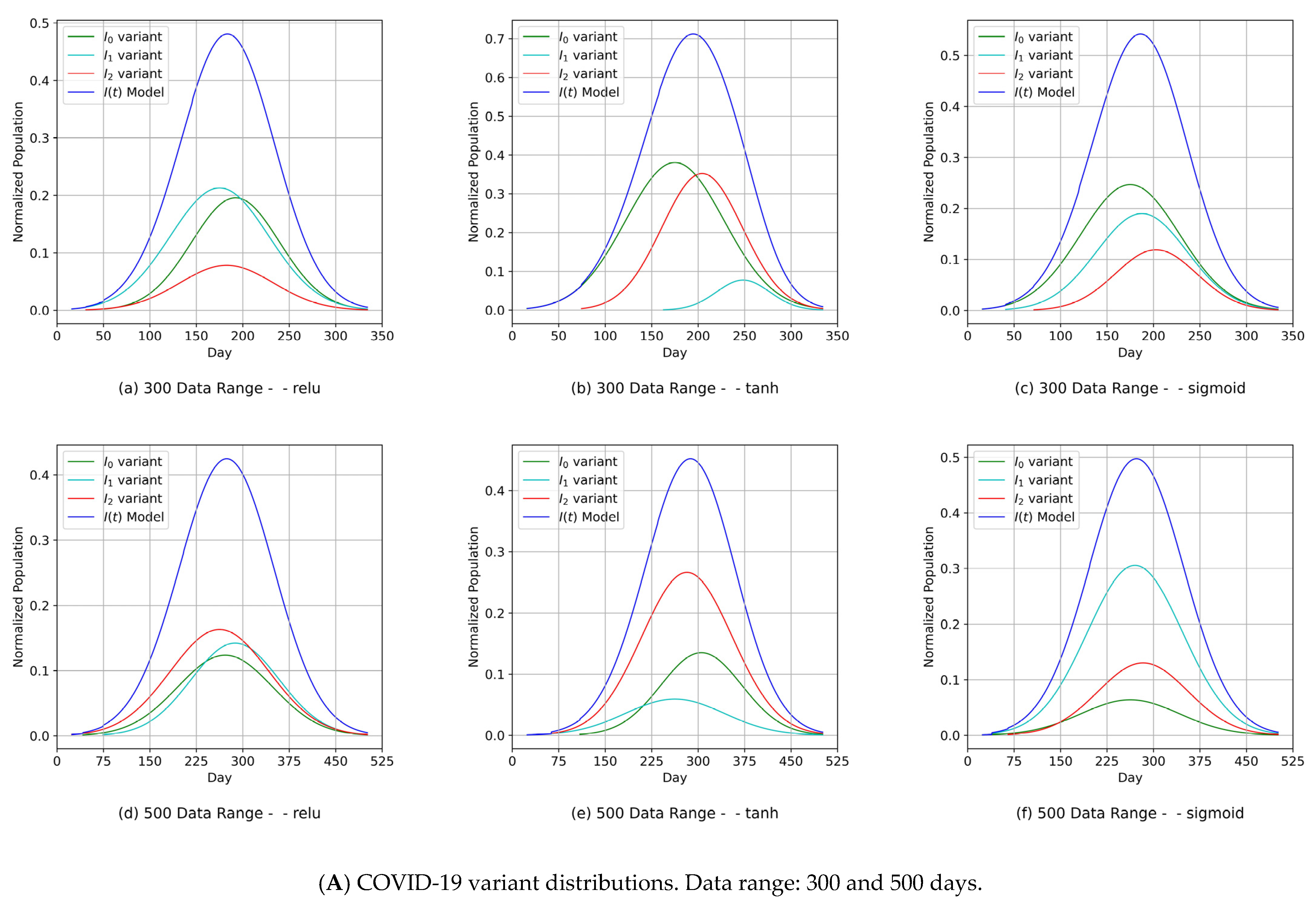
- Diagrams of the COVID-19 variants’ distributions {In(t)} and their predicted I(t) values as a synthesized Gaussian distribution;
- (2)
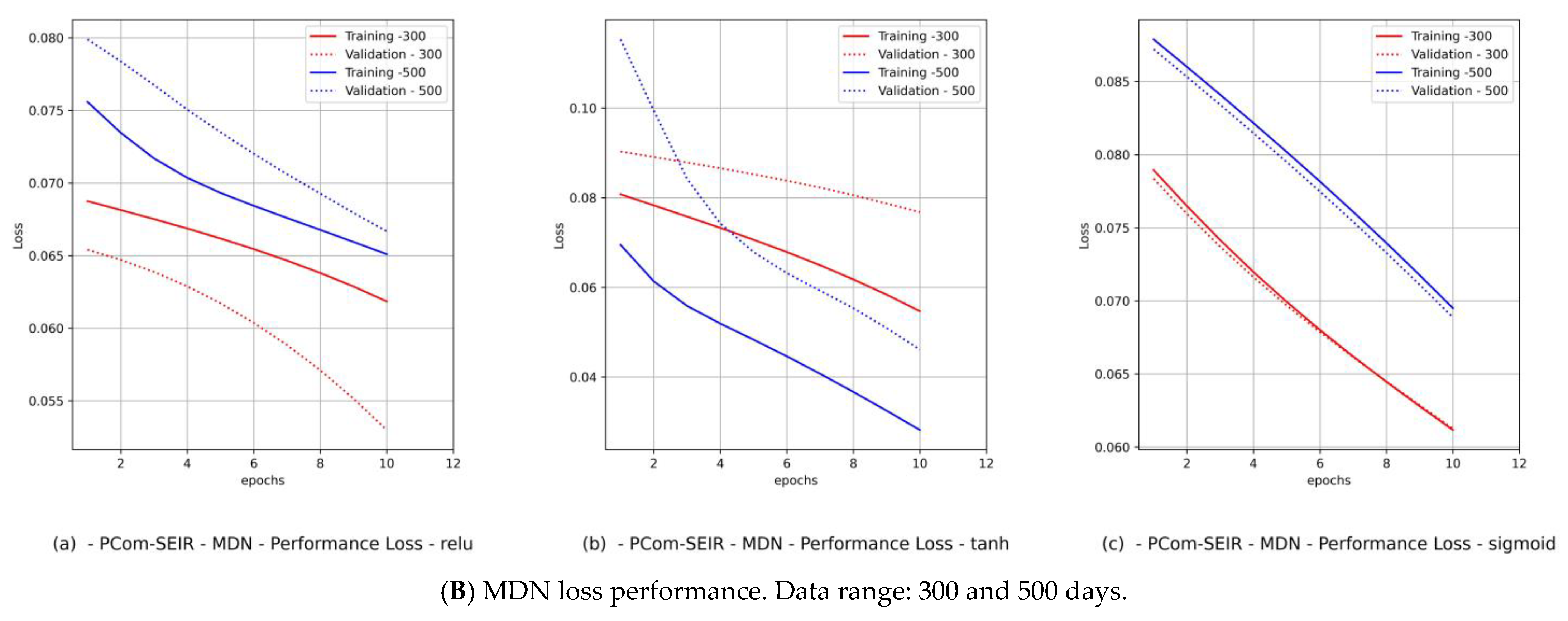
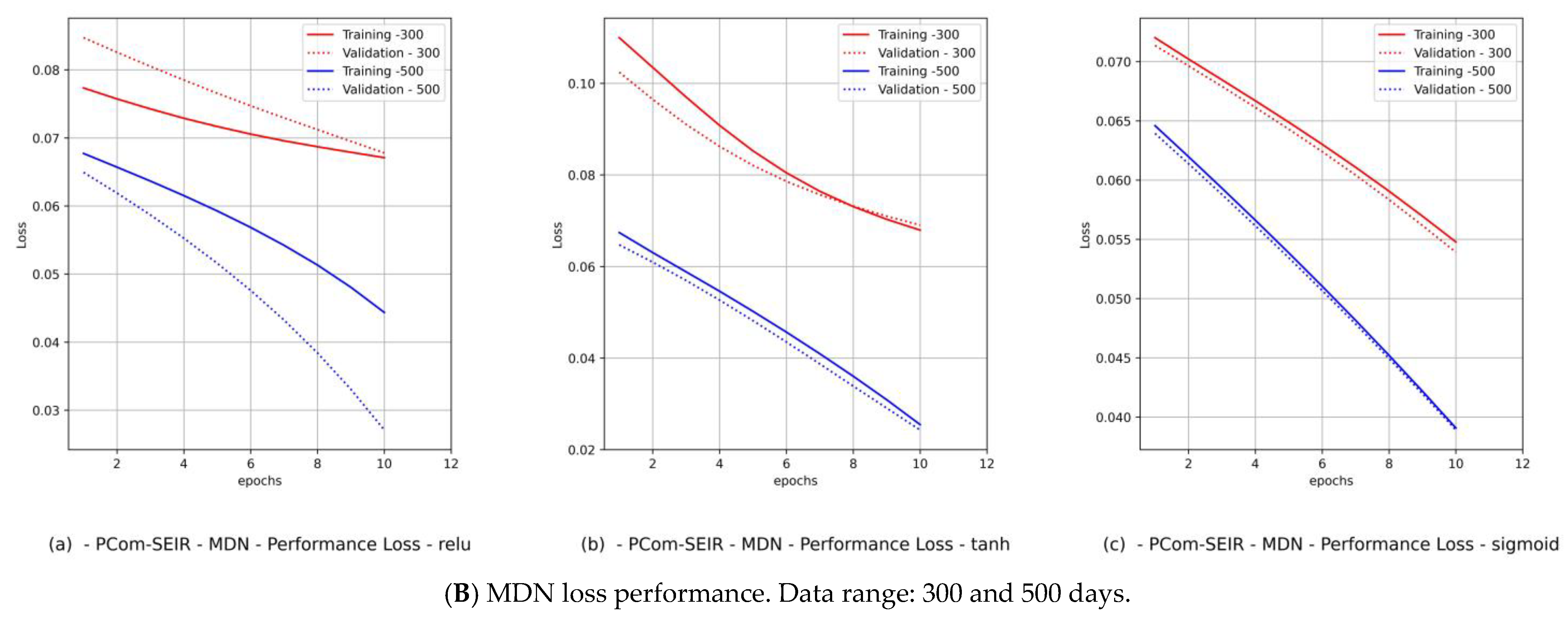
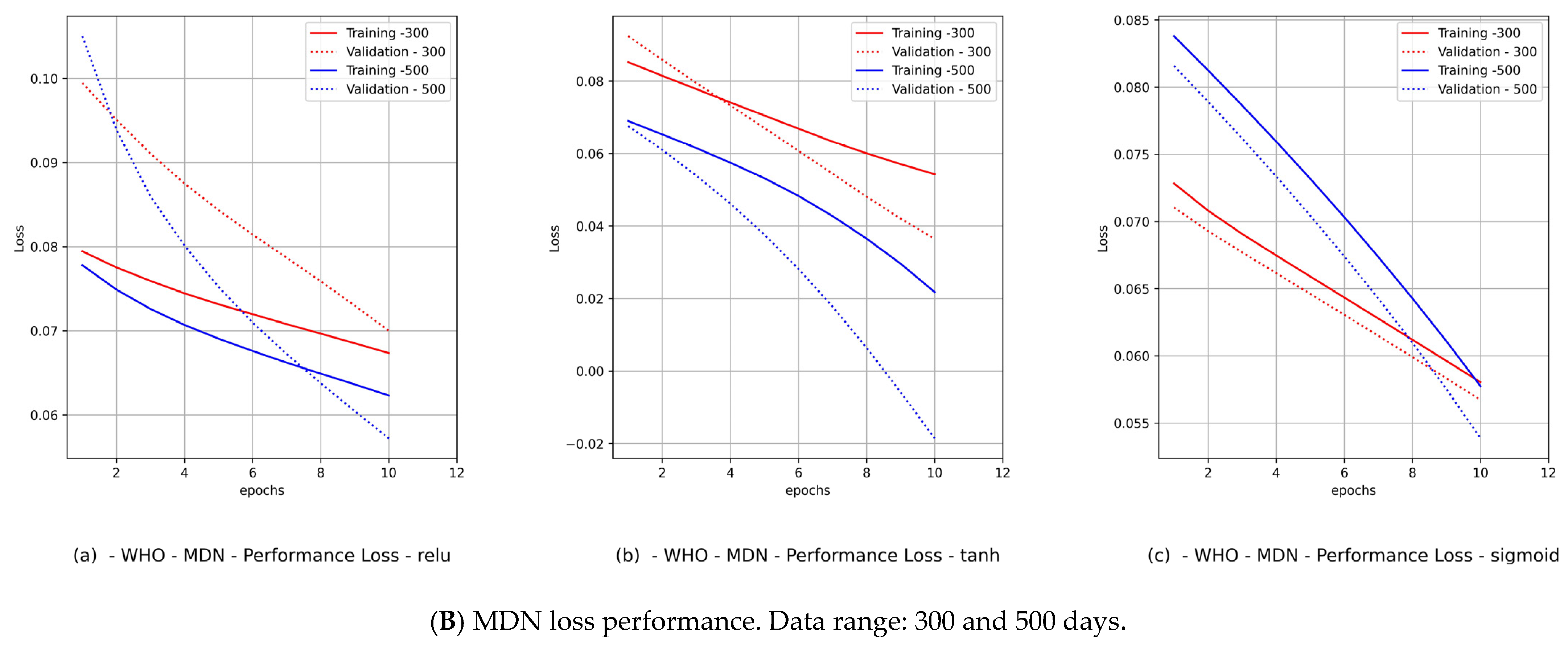
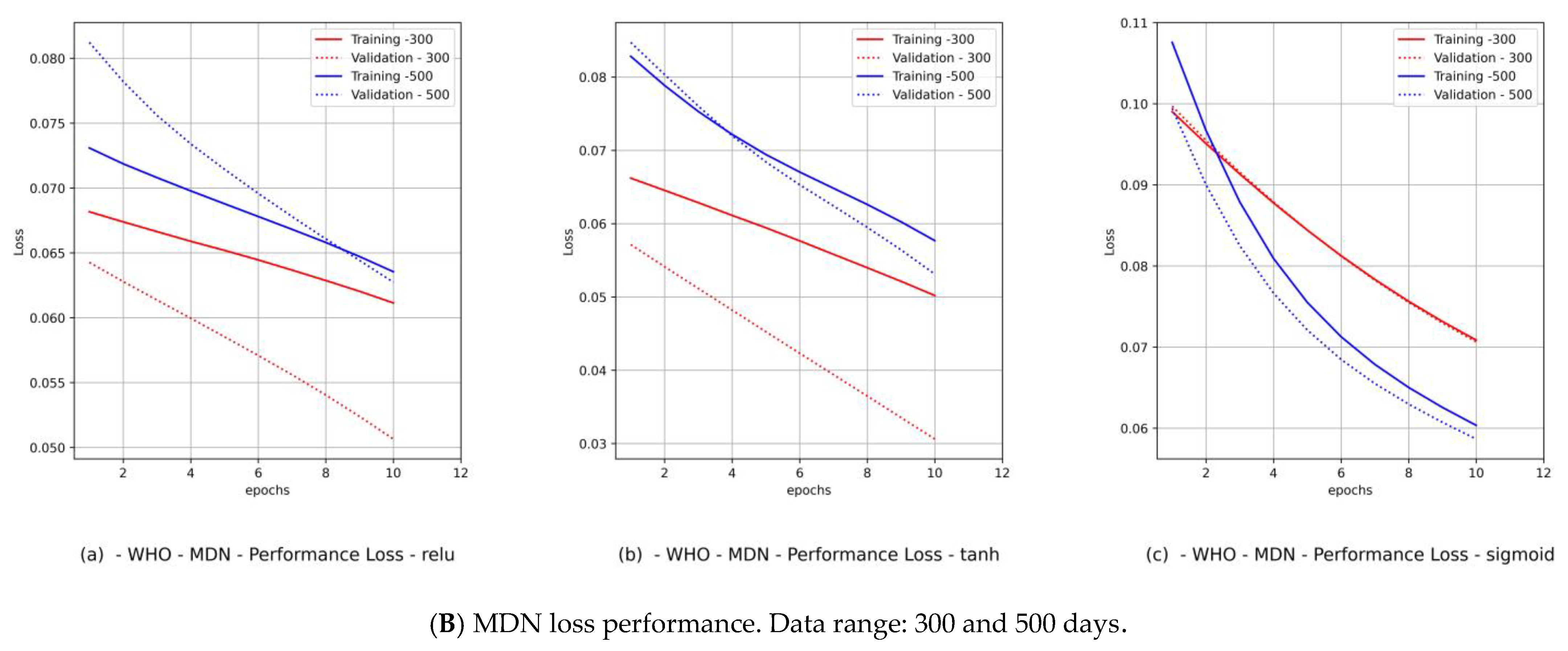
- Diagrams of the MDN’s loss performance, which includes
- (a)
- loss vs. epochs;
- (b)
- val_loss vs. epochs.
4.2. Implementation Configurations and Environments
4.3. The MDN’s Predictions of COVID-19 Variants
4.4. The Impacts of Epistemic Uncertainty and Aleatoric Uncertainty on Component Predictions
- (1)
- The fitting level of the PCom-SEIR data feed for both data ranges and both countries was 100% optimal, regardless of the activation function.
- (2)
- The fitting level of the WHO data for both data ranges and both countries was between 33% and 66%. The 33% optimal fitting occurred when we used the sigmoid activation function.
- (3)
- By observing the performance in Figure 9B(c) and Figure 11B(c) for Canada and in Figure 10B(c) and Figure 12B(c) for Saudi Arabia, one can notice that the sigmoid activation function produced 100% optimal fitting for both input data feeds. Relu and Tanh had a 100% optimal fit for the PCom-SEIR input data feed and nearly zero for the WHO input data feed. This meant that the algorithm, alongside the statistical profile of the input data, played a part in governing the model fitting for the same set of batch sizes and epochs.
4.5. The MDN Model’s Accuracy
4.6. General Remarks
5. Conclusions
- (1)
- The new MDN-based computational instrument was proven to be valid and credible through the examination of twelve use cases covering COVID-19 data sources, date ranges (i.e., data scope), and several MDN implementation configurations. The MDN’s outputs for these use cases clearly indicated that it produced an interpretable model for the growth of the COVID-19 infection rate [45].
- (2)
- Our approach can provide vital information to health authorities when planning health control measures and preparing for a severe spread of infections. This can be argued because our results indicated that the MDN produced COVID-19 variants’ In(t) values as Gaussian distributions with different sets of distribution parameters {µ, σ, α}. Different values of µ and σ imply that each variant candidate in the {In(t)} set has a different extent of time (length of days), different start/end days, and different variant peaks mostly taking place on different days (which is the unit of time of observation). Accordingly, the information above is vital to health authorities when planning health control measures and preparing for the impact of the severe spread of infections. On the other hand, in [1], the COVID-19 variant profiles had the same extent of time and practically the same peaks, as evidenced when comparing the prediction results for the COVID-19 variants’ In(t) values in this study.
- (3)
- Another indicator of the universality and practicality of our proposed MDN as a tool for predicting COVID-19 variants was demonstrated. It was shown that the relu, tanh, and sigmoid activation functions essentially had comparable sets of COVID-19 variant distributions, each with different parameter sets {µ, σ, α} that were compatible with the reality of the emergence and spread of COVID-19 variants.
- (4)
- (5)
- The MDN with the sigmoid activation function produced a multipeaked I(t) value by producing one of the components as a late-peaking variant component (Figure 13).
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Al-Hadeethi, Y.; El Ramley, I.F.; Mohammed, H.; Barasheed, A.Z. A New Polymorphic Comprehensive Model for COVID-19 Transition Cycle Dynamics with Extended Feed Streams to Symptomatic and Asymptomatic Infections. Mathematics 2023, 11, 1119. [Google Scholar] [CrossRef]
- Al-Hadeethi, Y.; Ramley, I.F.E.; Sayyed, M.I. Convolution model for COVID-19 rate predictions and health effort levels computation for Saudi Arabia, France, and Canada. Sci. Rep. 2021, 11, 22664. [Google Scholar] [CrossRef] [PubMed]
- Anastassopoulou, C.; Russo, L.; Tsakris, A.; Siettos, C. Data-based analysis, modelling and forecasting of the COVID-19 outbreak. PLoS ONE 2020, 15, e0230405. [Google Scholar] [CrossRef] [PubMed]
- Bakhta, A.; Boiveau, T.; Maday, Y.; Mula, O. Epidemiological forecasting with model reduction of compartmental models. Application to the COVID-19 pandemic. Biology 2020, 10, 22. [Google Scholar] [CrossRef]
- Chang, Y.-C.; Liu, C.-T. A Stochastic Multi-Strain SIR Model with Two-Dose Vaccination Rate. Mathematics 2022, 10, 1804. [Google Scholar] [CrossRef]
- Liu, X.; Ding, Y. Stability and numerical simulations of a new SVIR model with two delays on COVID-19 booster vaccination. Mathematics 2022, 10, 1772. [Google Scholar] [CrossRef]
- Putra, S.; Mutamar, Z.K. Estimation of parameters in the SIR epidemic model using particle swarm optimisation. Am. J. Math. Comput. Model. 2019, 4, 83–93. [Google Scholar] [CrossRef]
- Margenov, S.; Popivanov, N.; Ugrinova, I.; Hristov, T. Mathematical Modeling and Short-Term Forecasting of the COVID-19 Epidemic in Bulgaria: SEIRS Model with Vaccination. Mathematics 2022, 10, 2570. [Google Scholar] [CrossRef]
- Mamis, K.; Farazmand, M. Stochastic compartmental models of the COVID-19 pandemic must have temporally correlated uncertainties. Proc. R. Soc. A 2023, 479, 20220568. [Google Scholar] [CrossRef]
- Mbuvha, R.; Marwala, T. On data-driven management of the COVID-19 outbreak in South Africa. medRxiv 2020. [Google Scholar] [CrossRef]
- Gatto, A.; Accarino, G.; Aloisi, V.; Immorlano, F.; Donato, F.; Aloisio, G. Limits of Compartmental Models and New Opportunities for Machine Learning: A Case Study to Forecast the Second Wave of COVID-19 Hospitalizations in Lombardy, Italy. Informatics 2021, 8, 57. [Google Scholar] [CrossRef]
- Wondyfraw, T.A.; Sama, S.T. Stochastic model of the transmission dynamics of COVID-19 pandemic. Adv. Differ. Equ. 2021, 2021, 457. [Google Scholar]
- Hoertel, N.; Blachier, M.; Blanco, C.; Olfson, M.; Massetti, M.; Rico, M.S.; Limosin, F.; Leleu, H. A stochastic agent-based model of the SARS-CoV-2 epidemic in France. Nat. Med. 2020, 26, 1417–1421. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Zhang, H.T.; Xiao, Y.; Wang, M.; Guo, Y.; Sun, C.; Tang, X.; Jing, L.; Li, S.; Zhang, M.; et al. Prediction of criticality in patients with severe COVID-19 infection using three clinical features: A machine learning-based prognostic model with clinical data in Wuhan. medRxiv 2020. [CrossRef]
- Frausto-Solís, J.; Hernández-González, L.J.; González-Barbosa, J.J.; Sánchez-Hernández, J.P.; Román-Rangel, E. Convolutional Neural Network–Component Transformation (CNN–CT) for Confirmed COVID-19 Cases. Math. Comput. Appl. 2021, 26, 29. [Google Scholar] [CrossRef]
- Alanazi, S.A.; Kamruzzaman, M.M.; Alruwaili, M.; Alshammari, N.; Alqahtani, S.A.; Karime, A. Measuring and Preventing COVID-19 Using the SIR Model and Machine Learning in Smart Health Care. J. Healthc. Eng. 2020, 2020, 8857346. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, Z.; Almaspoor, Z.; Khan, F.; El-Morshedy, M. On predictive modeling using a new flexible Weibull distribution and machine learning approach: Analysing the COVID-19 data. Mathematics 2022, 10, 1792. [Google Scholar] [CrossRef]
- Yadav, S.K.; Akhter, Y. Statistical Modeling for the Prediction of Infectious Disease Dissemination with Special Reference to COVID-19 Spread. Front. Public Health 2021. [Google Scholar] [CrossRef]
- Zain, Z.M.; Alturki, N.M. COVID-19 pandemic forecasting using CNN-LSTM: A hybrid approach. J. Control Sci. Eng. 2021, 2021, 8785636. [Google Scholar] [CrossRef]
- Wang, L.; Lin, Z.Q.; Wong, A. Covid-net: A tailored deep convolutional neural network design for detection of COVID-19 cases from chest x-ray images. Sci. Rep. 2020, 10, 19549. [Google Scholar] [CrossRef]
- Zisad, S.N.; Hossain, M.S.; Hossain, M.S.; Andersson, K. An Integrated Neural Network and SEIR Model to Predict COVID-19. Algorithms 2021, 14, 94. [Google Scholar] [CrossRef]
- Wieczorek, M.; Siłka, J.; Wo’zniak, M. Neural network powered COVID-19 spread Forecasting model. Chaos Solitons Fractals 2020, 140, 110203. [Google Scholar] [CrossRef]
- Schiassi, E.; de Florio, M.; D’Ambrosio, A.; Mortari, D.; Furfaro, R. Physics-informed neural networks and functional interpolation for data-driven parameters discovery of epidemiological compartmental models. Mathematics 2021, 9, 2069. [Google Scholar] [CrossRef]
- Hussein, H.I.; Mohammed, A.O.; Hassan, M.M.; Mstafa, R.J. Lightweight deep CNN-based models for early detection of COVID-19 patients from chest X-ray images. Expert Syst. Appl. 2023, 223, 119900. [Google Scholar] [CrossRef] [PubMed]
- Tamang, S.K.; Singh, P.D.; Datta, B. Forecasting of COVID-19 cases based on prediction using artificial neural network curve fitting technique. Glob. J. Environ. Sci. Manag. 2020, 6, 53–64. [Google Scholar]
- Huang, C.J.; Chen, Y.H.; Ma, Y.; Kuo, P.H. Multiple-input deep convolutional neural network model for covid-19 forecasting in china. medRxiv 2020. [Google Scholar] [CrossRef]
- Gomez-Cravioto, D.A.; Diaz-Ramos, R.E.; Cantu-Ortiz, F.J.; Ceballos, H.G. Data Analysis and Forecasting of the COVID-19 Spread: A Comparison of Recurrent Neural Networks and Time Series Models. Cogn. Comput. 2021. [Google Scholar] [CrossRef]
- Feng, C.; Wang, L.; Chen, X.; Zhai, Y.; Zhu, F.; Chen, H.; Wang, Y.; Su, X.; Huang, S.; Tian, L.; et al. A Novel triage tool of artificial intelligence-assisted diagnosis aid system for suspected COVID-19 pneumonia in fever clinics. Ann Transl Med. 2021, 9, 201. [Google Scholar] [CrossRef]
- Jin, C.; Chen, W.; Cao, Y.; Xu, Z.; Tan, Z.; Zhang, X.; Deng, L.; Zheng, C.; Zhou, J.; Shi, H.; et al. Development and evaluation of an artificial intelligence system for COVID-19 diagnosis. Nat. Commun. 2020, 11, 5088. [Google Scholar] [CrossRef]
- Xie, J.; Hungerford, D.; Chen, H.; Abrams, S.T.; Li, S.; Wang, G.; Wang, Y.; Kang, H.; Bonnett, L.; Zheng, R.; et al. Development and external validation of a prognostic multivariable model on admission for hospitalised patients with COVID-19. medRxiv 2020. [CrossRef]
- Wynants, L.; van Calster, B.; Collins, G.S.; Riley, R.D.; Heinze, G.; Schuit, E.; Bonten, M.M.J.; Dahly, D.L.; Damen, J.A.; Debray, T.P.A.; et al. Prediction models for diagnosis and prognosis of COVID-19: Systematic review and critical appraisal. BMJ 2020, 369. [Google Scholar] [CrossRef]
- Rahimi, I.; Chen, F.; Gandomi, A.H. A review on COVID-19 forecasting models. Neural Comput. Appl. 2023, 35, 23671–23681. [Google Scholar] [CrossRef] [PubMed]
- Naudé, W. Artificial intelligence vs COVID-19: Limitations, constraints and pitfalls. AI Soc. 2020, 35, 761–765. [Google Scholar] [CrossRef] [PubMed]
- Britton, T. Stochastic epidemic models: A survey. Math. Biosci. 2010, 225, 24–35. [Google Scholar] [CrossRef] [PubMed]
- Storn, R.; Price, K. Differential evolution–a simple and efficient heuristic for global optimisation over continuous spaces. J. Glob. Optim. 1997, 11, 341–359. [Google Scholar] [CrossRef]
- Technical Report. Available online: https://publications.aston.ac.uk/ (accessed on 14 April 2024).
- WHO Data. Available online: https://covid19.who.int/WHO-COVID-19-global-data.csv (accessed on 14 April 2024).
- Python Optimization (scipy. Optimise). Available online: https://docs.scipy.org/doc/scipy/tutorial/optimize.html (accessed on 10 October 2022).
- Lerch, F.; Ultsch, A.; Lötsch, J. Distribution Optimization: An evolutionary algorithm to separate Gaussian mixtures. Sci. Rep. 2020, 10, 648. [Google Scholar] [CrossRef] [PubMed]
- Covões, T.F.; Hruschka, E.R.; Ghosh, J. Evolving gaussian mixture models with splitting and merging mutation operators. Evol. Comput. 2016, 24, 293–317. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Dushoff, J.; Bolker, B.M. Fitting mechanistic epidemic models to data: A comparison of simple Markov chain Monte Carlo approaches. Stat. Methods Med. Res. 2018, 27, 1956–1967. [Google Scholar] [CrossRef] [PubMed]
- Rafique, D.; Velasco, L. Machine learning for network automation: Overview, architecture, and applications [Invited Tutorial]. J. Opt. Commun. Netw. 2018, 10, D126–D143. [Google Scholar] [CrossRef]
- Saleh, B. Photoelectron Statistics: With Applications to Spectroscopy and Optical Communication; Springer: Cham, Switzerland, 2013; Volume 6. [Google Scholar]
- Hüllermeier, E.; Waegeman, W. Aleatoric and epistemic uncertainty in machine learning: An introduction to concepts and methods. Mach. Learn. 2021, 110, 457–506. [Google Scholar] [CrossRef]
- Montesinos López, O.A.; Montesinos López, A.; Crossa, J. Multivariate Statistical Machine Learning Methods for Genomic Prediction; Springer Nature: Berlin, Germany, 2022; Volume 691. [Google Scholar]
- Rahmani, R.; Yusof, R. A new simple, fast and efficient algorithm for global optimization over continuous search-space problems: Radial movement optimization. Appl. Math. Comput. 2014, 248, 287–300. [Google Scholar] [CrossRef]



| Variable (Z(t)) | Description | |
|---|---|---|
| 1 | S(t) | The population of the susceptible compartment. |
| 2 | P(t) | The population of the protected compartment. |
| 3 | E(t) | The population of the exposed compartment. |
| 4 | I(t) | The infection population of the symptomatic infection compartment. |
| 5 | M(t) | The population with asymptomatic infection. |
| 6 | Q(t) | The population of the quarantined compartment. |
| 7 | H(t) | The population of the hospitalised compartment. |
| 8 | D(t) | The population of the dead compartment. |
| Parameter (ψ) | Input Parameter | ||
|---|---|---|---|
| From Compartment | To Compartment | ||
| 1 | α | Susceptible (S) | Protected (P) |
| 2 | Β | Exposed (E) | |
| 3 | Φ | Protected (P) | |
| 4 | Φe | Protected Pe = P * Φe component | Symptomatic infection (I) |
| 5 | Φm | Protected Pm = P * Φm component | Asymptomatic infection (M) |
| 6 | ϒ | Exposed (E) | Symptomatic infection (I) |
| 7 | є | Asymptomatic infection (M) | |
| 8 | I | Symptomatic infection (I) | Symptomatic infection |
| 9 | ϒ | Exposed component | Symptomatic infection (I) |
| 10 | η | Quarantine (Q) | |
| 11 | λ | Death (D) | |
| 12 | Τ | Asymptomatic (M) | Recovered (R) |
| 13 | Χ | Hospitalisation (H) | Recovered (R) |
| 14 | Ρ | Death (D) | |
| 15 | Μ | Quarantine (Q) | Recovered (R) |
| 16 | Σ | Death (D) | |
| 17 | Φ | Hospitalisation (H) | |
| 18 | r | Recovered component | Symptomatic infection |
| MDN–GMM Network Data | ||||
|---|---|---|---|---|
| 300 Data Range | 500 Data Range | 300 Data Range | 500 Data Range | |
| Input Data Size | 319 | 571 | 283 | 535 |
| Training Data Size | 212 | 380 | 188 | 356 |
| Testing Data Size | 107 | 191 | 95 | 179 |
| Output Data Size | 107 | 191 | 95 | 179 |
| Neurons | 10 | 10 | 10 | 10 |
| Batch Size | 64 | 64 | 64 | 64 |
| (A) Canada | (B) Saudi Arabia | |||
| Country | Canada | Saudi Arabia | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Data Range | 300–500 Days | 300–500 Days | ||||||||||
| Data Input Mode | PCom-SEIR | WHO Data | PCom-SEIR | WHO Data | ||||||||
| Activation Function | relu | tanh | sigmoid | relu | tanh | sigmoid | relu | tanh | sigmoid | relu | tanh | sigmoid |
| Variant’s Distribution | Figure 9A | Figure 9A | Figure 9A | Figure 11A | Figure 11A | Figure 11A | Figure 10A | Figure 10A | Figure 10A | Figure 12A | Figure 12A | Figure 12A |
| Loss Performance | Figure 9B(a) | Figure 9B(b) | Figure 9B(c) | Figure 11B(a) | Figure 11B(b) | Figure 11B(c) | Figure 10B(a) | Figure 10B(b) | Figure 10B(c) | Figure 12B(a) | Figure 12B(b) | Figure 12B(c) |
| Network Parameters | Table 3A | Table 3B | ||||||||||
| Country | Canada | Saudi Arabia | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Input Mode | PCom-SEIR | WHO | PCom-SEIR | WHO | |||||
| Data Range | 300 | 500 | 300 | 500 | 300 | 500 | 300 | 500 | |
| relu | loss |  |  |  |  |  |  |  |  |
| Accuracy | 97.49 | 98.25 | 97.49 | 76.88 | 96.47 | 98.50 | 96.11 | 99.07 | |
| tanh | loss |  |  |  |  |  |  |  |  |
| Accuracy | 97.49 | 99.30 | 97.81 | 77.93 | 97.88 | 98.88 | 98.23 | 98.88 | |
| sigmoid | loss |  |  |  |  |  |  |  |  |
| Accuracy | 97.18 | 99.12 | 96.87 | 77.76 | 96.11 | 98.13 | 97.17 | 99.44 | |
| Fitting Level % | 100% | 100% | 33% | 33% | 100% | 100% | 33% | 66% | |
| Loss Legend | Optimal fit  | Underfitting  | Overfitting  | ||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Al-Hadeethi, Y.; El Ramley, I.F.; Mohammed, H.; Bedaiwi, N.M.; Barasheed, A.Z. A Novel Computational Instrument Based on a Universal Mixture Density Network with a Gaussian Mixture Model as a Backbone for Predicting COVID-19 Variants’ Distributions. Mathematics 2024, 12, 1254. https://doi.org/10.3390/math12081254
Al-Hadeethi Y, El Ramley IF, Mohammed H, Bedaiwi NM, Barasheed AZ. A Novel Computational Instrument Based on a Universal Mixture Density Network with a Gaussian Mixture Model as a Backbone for Predicting COVID-19 Variants’ Distributions. Mathematics. 2024; 12(8):1254. https://doi.org/10.3390/math12081254
Chicago/Turabian StyleAl-Hadeethi, Yas, Intesar F. El Ramley, Hiba Mohammed, Nada M. Bedaiwi, and Abeer Z. Barasheed. 2024. "A Novel Computational Instrument Based on a Universal Mixture Density Network with a Gaussian Mixture Model as a Backbone for Predicting COVID-19 Variants’ Distributions" Mathematics 12, no. 8: 1254. https://doi.org/10.3390/math12081254
APA StyleAl-Hadeethi, Y., El Ramley, I. F., Mohammed, H., Bedaiwi, N. M., & Barasheed, A. Z. (2024). A Novel Computational Instrument Based on a Universal Mixture Density Network with a Gaussian Mixture Model as a Backbone for Predicting COVID-19 Variants’ Distributions. Mathematics, 12(8), 1254. https://doi.org/10.3390/math12081254







