Semi-Automatic 3D Reconstruction of Atheroma Plaques from Intravascular Ultrasound Images Using an ad-hoc Algorithm
Abstract
1. Introduction
2. Methodology
2.1. Materials
2.2. Participants
2.3. Algorithms/Methods
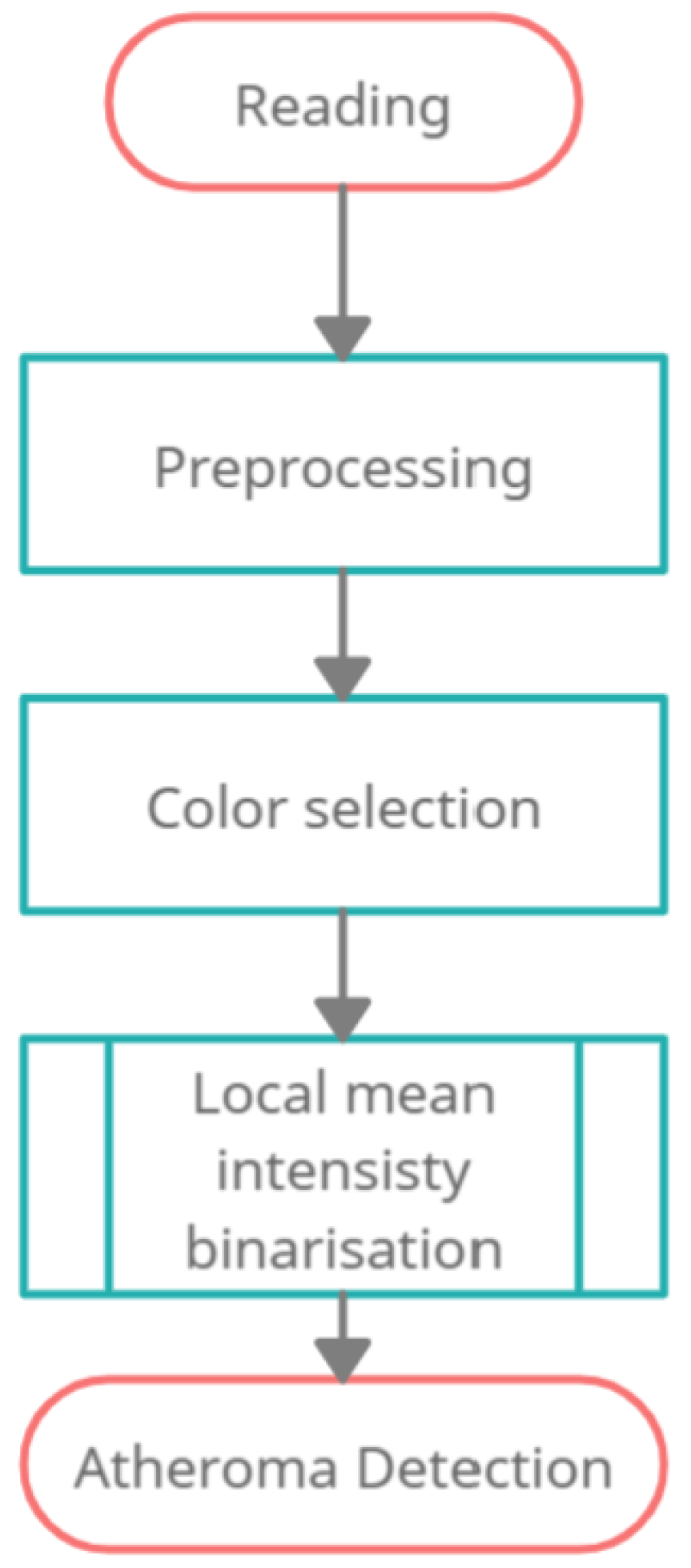
- Reading.
- Preprocessing.
- Extraction of the atheroma at each cross-sectional image.
- Extraction of the atheroma border.
- Postprocessing.
2.3.1. Preprocessing
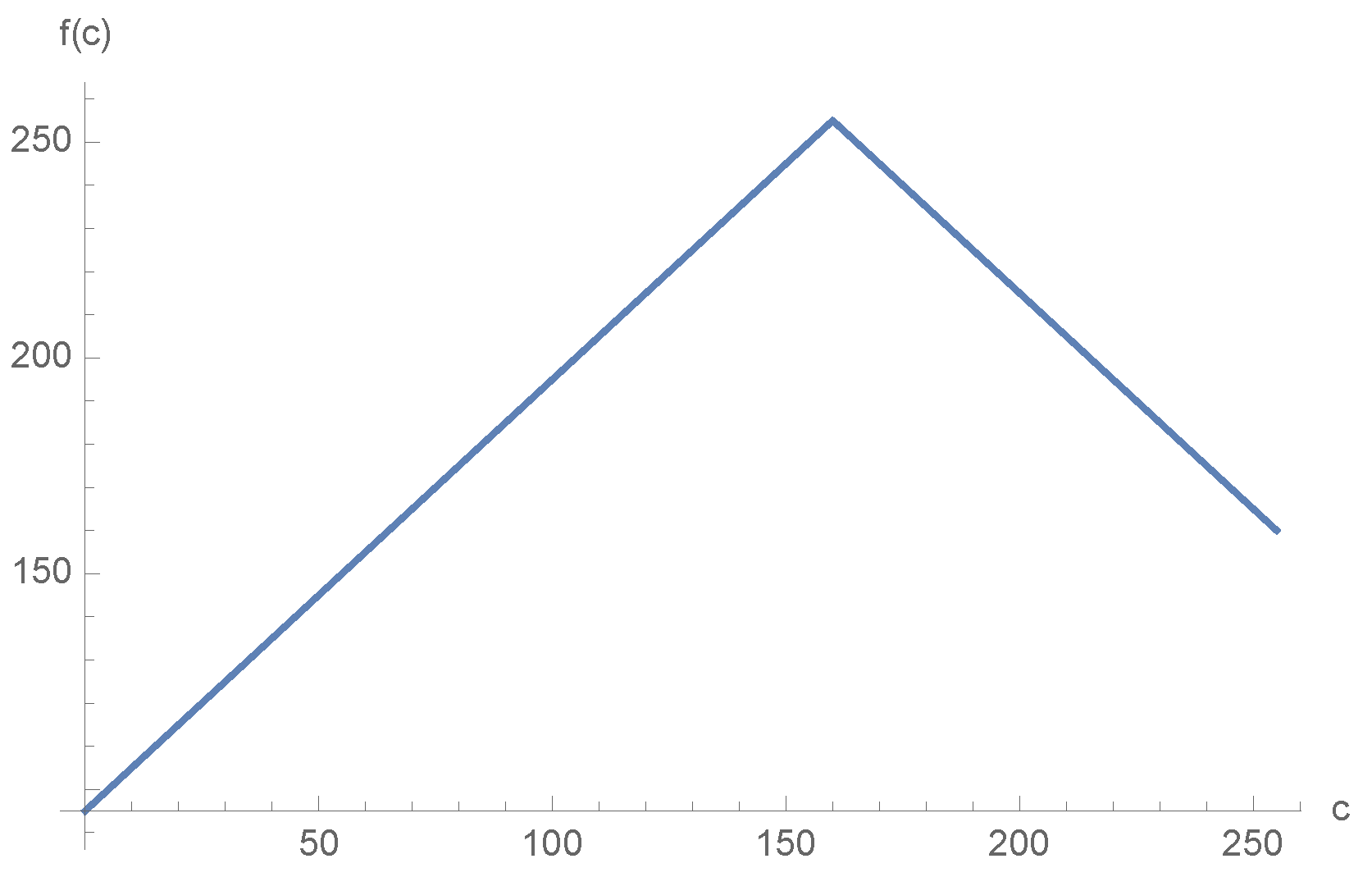
- Before starting the algorithm, the specialist is requested to provide the intensity of the atheroma . If the atheroma is not the brightest region of the image (i.e., ), then each pixel of the image is transformed following a tent-like map given by the function:In such way, it is ensured that the brightest region is the region that needs to be selected. Figure 2 gives an example of such map with a value . Once the transformation is used, the algorithm extracts the whitest regions in the next step.
- Binarisation of the image. The image is binarised by following an adaptive threshold based on the local mean intensity. In such way, the resulting image is smoother and less dependent on difference of light due to the noise that can be introduced by the capture device. Other methods have been implemented and tested, such as the Otsu filter or a fixed threshold value in order to decide whether the pixel is white or black. However, the extraction obtained is not as precise as with the adaptative threshold when comparing against the extraction made by a specialist. Thanks to the tent-like transformation, the intensity corresponding to the atheroma is always captured and is never lost due to the effects of the local mean intensity.
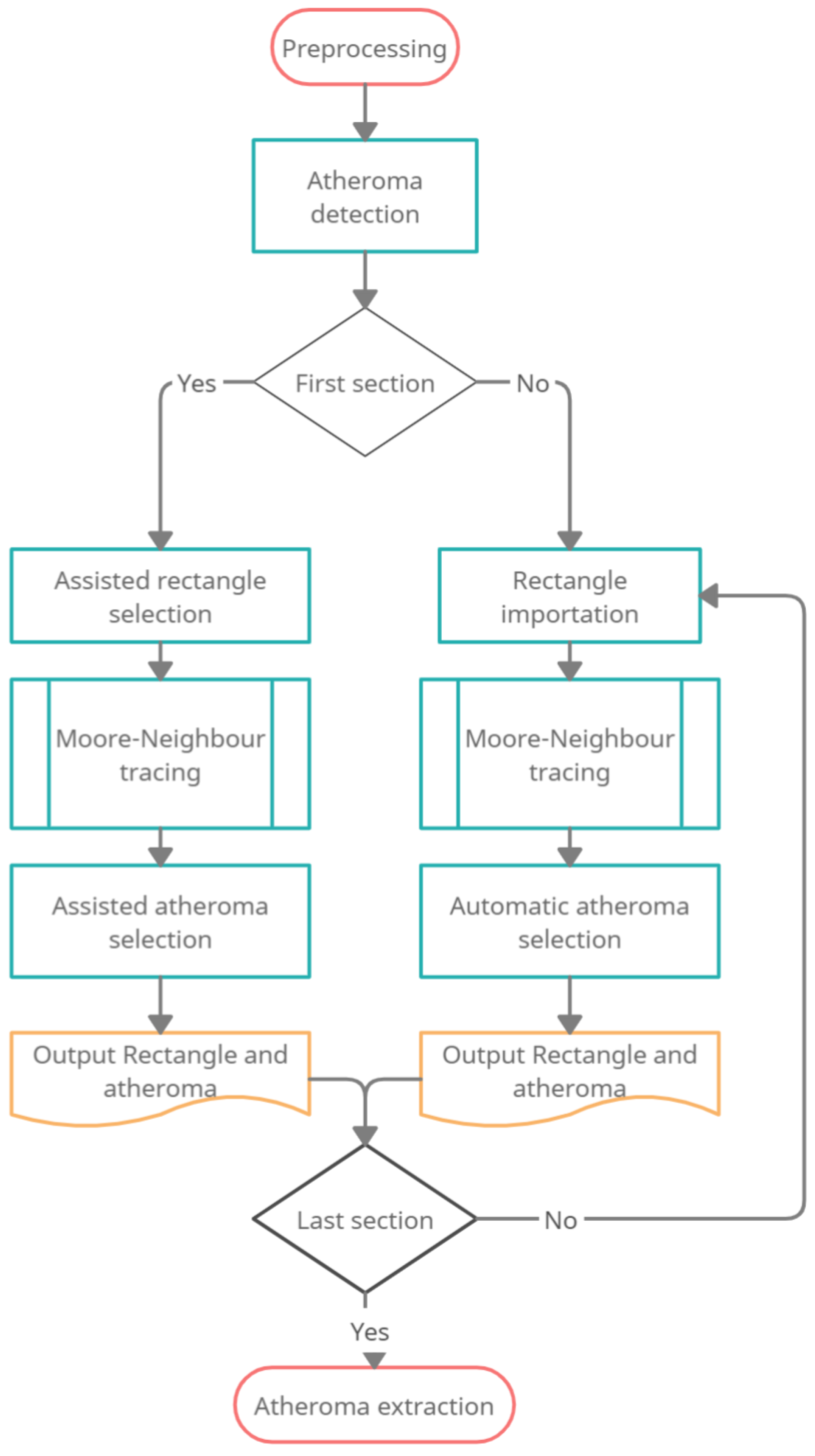
2.3.2. Detection of the Atheroma at Each Cross-Sectional Image
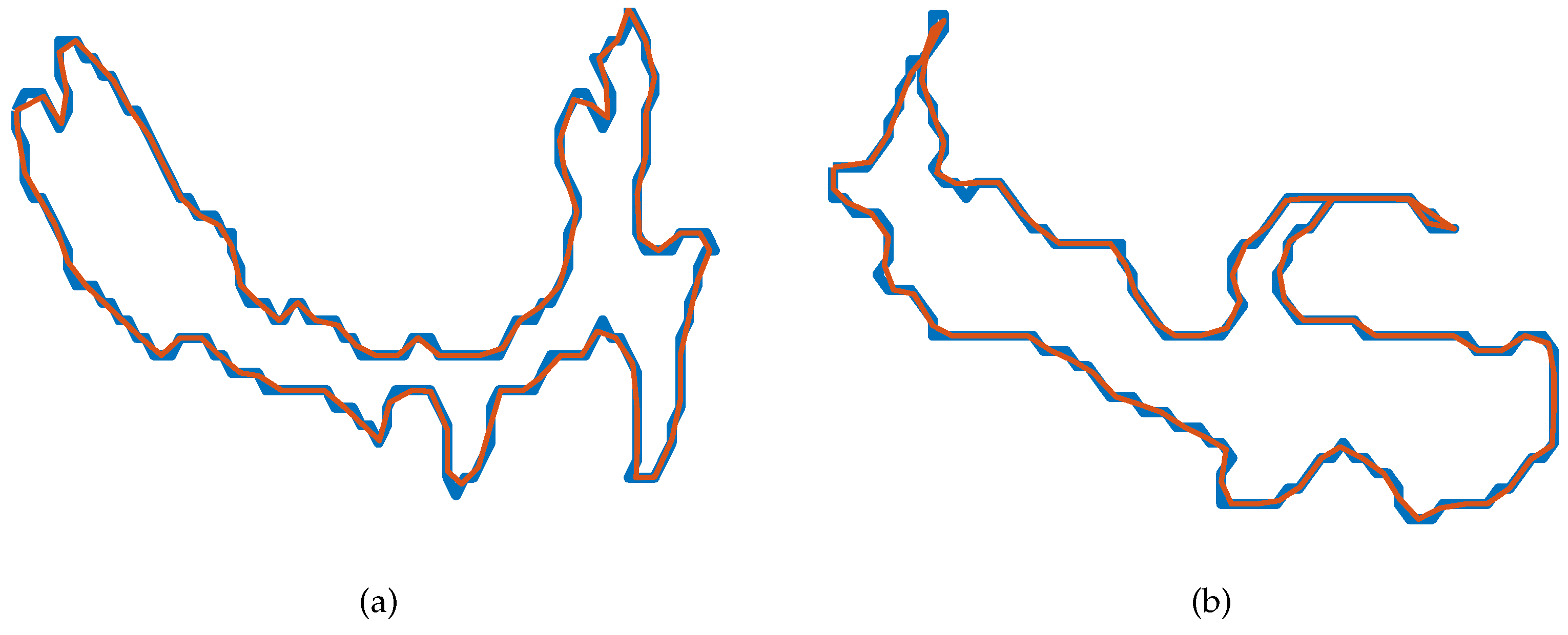
2.3.3. Extraction of the Atheroma Edge
- Compute , where we set to close the boundary. l gives the total length of the boundary following a piecewise connection.
- Compute the step .
- Fix , and .
- For ,
- (a)
- Compute the accumulated distance .
- (b)
- While a new point should be added. Use the following steps:
- Use linear interpolation to add the next boundary point to the uniformised boundary:Where .
- Increase .
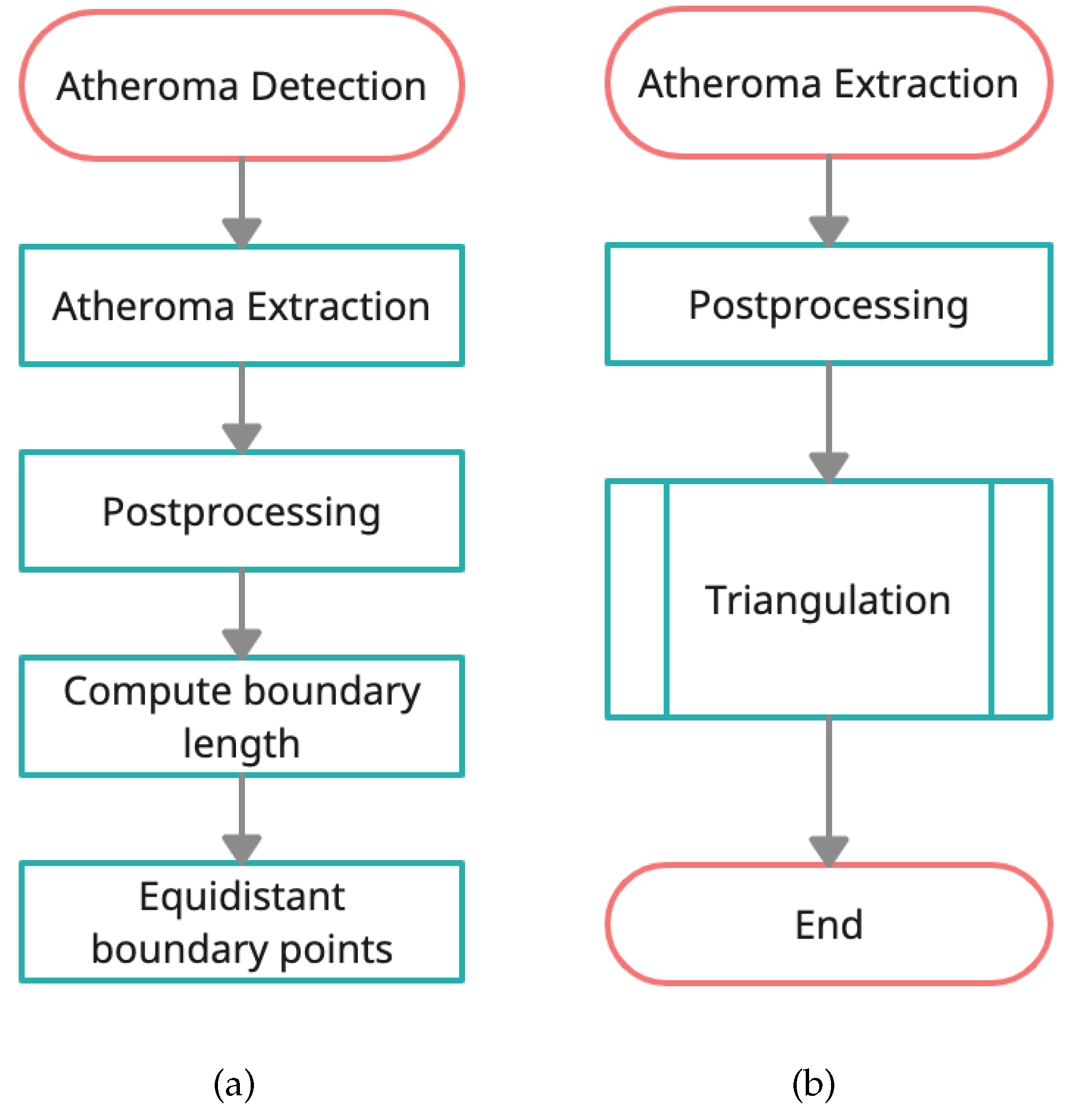
2.3.4. Postprocessing
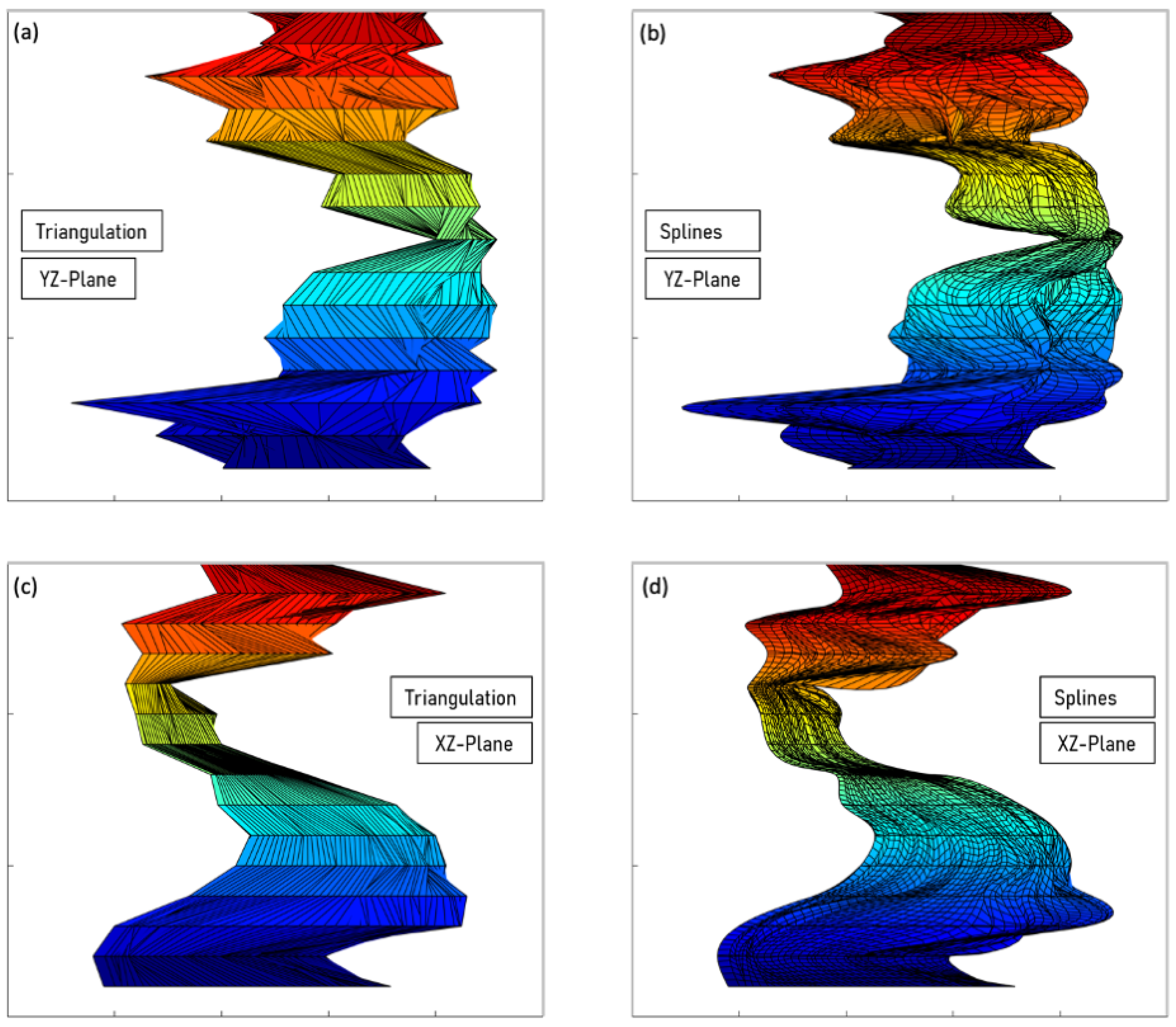
- Triangulation.After the last step for each image , the points at the boundary of the atheroma have been selected. We follow the triangulation (see flowchart in Figure 4b), as follows:
- Connect each layer with the layer by adding triangles using the vertices:
- –
- and .
- –
- and .
- Once layer i is connected, proceed with the next layer, .
In such way, each layer is connected to the previous one by triangulation. The boundary of the atheroma is a 2-dimensional manifold; therefore, it can be covered by a mesh of triangles to join triplets of points. We note to the reader that here triangulation does not refer to the location of a point in the 3D space given two or more images but to cover a shape by a mesh of triangles. For each set of four points (, , and ). The triangles and are stored in order to reconstruct the 3D figure. With this, the atheroma is reconstructed. - Cubic spline surface reconstruction.Following the previous method, an additional attempt was made to reconstruct the surface of the atheroma from a set of splines that approximated its contour. Thus, using the initial vertical discretization (i.e., the 15 contours with 100 points per level), clusters of 15 points in the z-direction were used to fit a set of 100 cubic splines that had individually described functions that fit the lateral surface and allowed an improved discretization. Final atheroma’s reconstruction is achieved by joining adjacent points, which are the result of a denser sampling based on the previously obtained functions, with straight lines.
2.4. Reconstruction of the Artery
3. Results
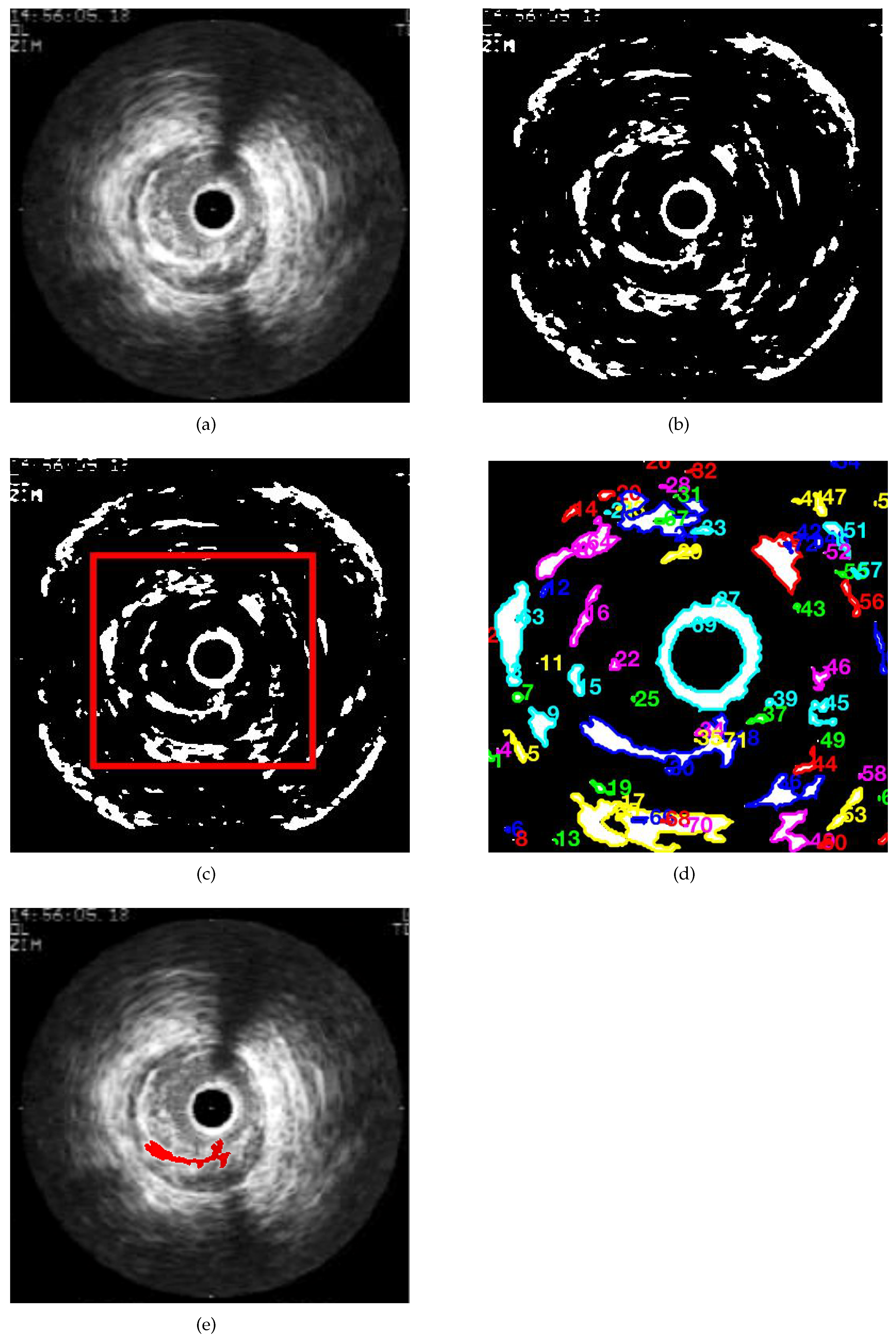
- First, the initial figure is obtained by the specialist.
- Then, the binarised image is shown.
- The user select the red square in the third.
- Finally, the boundaries of the possible regions are highlighted by the method. The user must select the one of interest to start the process.
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A

References
- Handels, H.; Ross, T.; Kreusch, J.; Wolff, H.H.; Pöppl, S.J. Feature selection for optimized skin tumor recognition using genetic algorithms. Artif. Intell. Med. 1999, 16, 183–197. [Google Scholar] [CrossRef] [PubMed]
- Gniadecka, M.; Philipsen, P.A.; Sigurdsson, S.; Wessel, S.; Nielsen, O.F.; Christensen, D.H.; Hercogova, J.; Rossen, K.; Thomsen, H.K.; Gniadecki, R.; et al. Melanoma diagnosis by Raman spectroscopy and neural networks: Structure alterations in proteins and lipids in intact cancer tissue. J. Investig. Dermatol. 2004, 122, 443–449. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.M.; Kecman, V. Gene extraction for cancer diagnosis by support vector machines—An improvement. Artif. Intell. Med. 2005, 35, 185–194. [Google Scholar] [CrossRef]
- Tan, B.K.J.; Teo, C.B.; Tadeo, X.; Peng, S.; Soh, H.P.L.; Du, S.D.X.; Luo, V.W.Y.; Bandla, A.; Sundar, R.; Ho, D.; et al. Personalised, Rational, Efficacy-Driven Cancer Drug Dosing via an Artificial Intelligence SystEm (PRECISE): A Protocol for the PRECISE CURATE.AI Pilot Clinical Trial. Front. Digit. Health 2021, 3, 635524. [Google Scholar] [CrossRef] [PubMed]
- Pawłowski, J.; Majchrowska, S.; Golan, T. Generation of microbial colonies dataset with deep learning style transfer. Sci. Rep. 2022, 12, 5212. [Google Scholar] [CrossRef]
- Itchhaporia, D.; Almassy, R.; Kaufman, L.; Snow, P.; Oetgen, W. Artificial Neural Networks Can Predict Significant Coronary Disease. J. Am. Coll. Cardiol. 1995, 25, 328A. [Google Scholar] [CrossRef]
- Poli, R.; Cagnoni, A.; Livi, R.; Coppini, G.; Valli, G. A neural network expert system for diagnosing and treating hypertension. Computer 1991, 24, 64–71. [Google Scholar] [CrossRef]
- Cilla, M.; Pérez-Rey, I.; Martínez, M.A.; Peña, E.; Martínez, J. On the use of Machine Learning Techniques for the Mechanical Characterization of Soft Biological Tissues. Int. J. Numer. Methods Biomed. Eng. 2018, e3121, 1–12. [Google Scholar] [CrossRef]
- Martínez, J.; Pérez-Rey, I.; Cilla, M. What are the possibilities of machine learning techniques on the mechanical characterization of biological tissues? Int. J. Biosens. Bioelectron. 2020, 6, 1–2. [Google Scholar]
- Jambukia, S.H.; Dabhi, V.K.; Prajapati, H.B. Classification of ECG signals using machine learning techniques: A survey. In Proceedings of the 2015 International Conference on Advances in Computer Engineering and Applications, Ghaziabad, India, 19–20 March 2015; pp. 714–721. [Google Scholar]
- Stary, H.C.; Chandler, A.B.; Dinsmore, R.E.; Fuster, V.; Glagov, S.; Insull, W.J.; Rosenfeld, M.E.; Schwartz, C.J.; Wagner, W.D.; Wissler, R.W. A definition of advanced types of atherosclerotic lesions and a histological classification of atherosclerosis. A report from the Committee on Vascular Lesions of the Council on Arteriosclerosis, American Heart Association. Circulation 1995, 92, 1355–1374. [Google Scholar] [CrossRef]
- Grandón-Pastén, N.; Aracena-Pizarro, D.; Tozzi, C.L. Reconstrucción de objeto 3D a partir de imágenes calibradas. Ingeniare Rev. Chil. Ing. 2007, 15, 158–168. [Google Scholar] [CrossRef]
- Piquer, A.; Company, P. Situación Actual de la Reconstrucción 3D de Dibujos Lineales. Inf. Tecnol. 2004, 15, 71–80. [Google Scholar] [CrossRef]
- Liritzis, I.; Volonakis, P.; Vosinakis, S. 3D Reconstruction of Cultural Heritage Sites as an Educational Approach. The Sanctuary of Delphi. Appl. Sci. 2021, 11, 3635. [Google Scholar] [CrossRef]
- Zingoni, A.; Diani, M.; Cosrini, G.; Masini, A. Real-time 3D reconstruction from images taken from an UAV. Int. Arch. Photogramm. Remote. Sens. Spat. Inf. Sci. 2015, XL-3/W3, 313–319. [Google Scholar] [CrossRef]
- Maurer, M.; Rumpler, M.; Wendel, A.; Hoppe, C.; Irschara, A.; Bischof, H. Geo-referenced 3D reconstruction: Fusing public geographic data and aerial imagery. In Proceedings of the 2012 IEEE International Conference on Robotics and Automation, Saint Paul, MN, USA, 14–18 May 2012; pp. 3557–3558. [Google Scholar]
- Tian, Y.; Long, Y.; Xia, D.; Yao, H.; Zhang, J. Handling occlusions in augmented reality based on 3D reconstruction method. Neurocomputing 2015, 156, 96–104. [Google Scholar] [CrossRef]
- Tsang, W.; Piazzese, C.; Kronzon, I.; Sotaquira, M.; Weinert, L.; Lang, R.M.; Caiani, E. Semi-automated detection and quantification of aortic atheromas from 3d transesophageal echocardiography. J. Am. Coll. Cardiol. 2013, 61, E974. [Google Scholar] [CrossRef][Green Version]
- Bender, K.; Matoq, A.; Nelson, J.S. Transesophageal echocardiography in congenital heart surgery: Perioperative considerations. Prog. Pediatr. Cardiol. 2020, 58, 101263. [Google Scholar] [CrossRef]
- Kigka, V.B.; Rigas, G.; Sakellarios, A.; Siogkas, P.; Andrikos, I.O.; Exarchos, T.P.; Loggitsi, D.; Anagnostopoulos, C.D.; Michalis, L.K.; Negliaf, D.; et al. 3D reconstruction of coronary arteries and atherosclerotic plaques based on computed tomography angiography images. Biomed. Signal Process. Control 2018, 40, 286–294. [Google Scholar] [CrossRef]
- Vásquez, S.H.; Vesga, B.E.; Hernández, H.J. Imagenología coronaria: Ultrasonido intravascular (IVUS). Rev. Colomb. Cardiol. 2017, 24, 101–106. [Google Scholar] [CrossRef]
- Salido, L.; Mestre, J.L.; Jiménez-Mena, M.; Hernández, R.; Zamorano, J.L. Disección coronaria espontánea multivaso posparto. Rev. Argent. Cardiol. 2015, 83, 152–154. [Google Scholar] [CrossRef]
- Mariani, J., Jr.; Guedes, C.; Soares, P.; Zalc, S.; Campos, C.M.; Lopes, A.C.; Spadaro, A.G.; Perin, M.A.; Filho, A.E.; Takimura, C.K.; et al. Intravascular ultrasound guidance to minimize the use of iodine contrast in percutaneous coronary intervention: The MOZART (Minimizing cOntrast utiliZation with IVUS Guidance in coRonary angioplasTy) randomized controlled trial. J. Am. Coll. Cardiol. Cardiovasc. Interv. 2014, 7, 1287–1293. [Google Scholar] [CrossRef] [PubMed]
- Sanz, R.; Bodí, V.; Sanchís, J.; Moratal, D.; Núñez, J.; Palau, P.; García, D.; Rieta, J.J.; Sanchís, J.M.; Chorro, F.J.; et al. Desarrollo de software para la reconstrucción tridimensional y cuantificación automática de secuencias de ultrasonido intravascular. Experiencia inicial. Rev. Esp. Cardiol. 2006, 59, 879–888. [Google Scholar] [CrossRef] [PubMed]
- Lo Vercio, L.; Arguiñarena, E.; Romero, L.; Jonas, I.; del Fresno, M.; Vénere, M. Reconstrucción Arterial a partir de Ultrasonido Intravascular Sincronizado con Electrocardiograma. Mec. Comput. 2012, XXXI, 3053–3060. [Google Scholar]
- Long, Q.; Ariff, B.; Zhao, S.Z.; Thom, S.A.; Hughes, A.D.; Xu, X.Y. Reproducibility study of 3D geometrical reconstruction of the human carotid bifurcation from magnetic resonance images. Magn. Reson. Med. 2003, 49, 665–674. [Google Scholar] [CrossRef] [PubMed]
- Delaunay, B. Sur la sphère vide. A la mémoire de Georges Voronoï. Bulletin de l’Académie des Sciences de l’URSS. VII Série. 1934, pp. 793–800. Available online: http://mi.mathnet.ru/im4937 (accessed on 18 October 2022).
- Cazals, F.; Giesen, J. Delaunay Triangulation Based Surface Reconstruction: Ideas and Algorithms; No. 5393; INRIA: Rocquencourt, France, 2004.
- Burger, W.; Burge, M.J. Principles of Digital Image Processing Core Algorithms; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2010. [Google Scholar]
- Gonzalez, R.C.; Woods, R.E.; Eddins, S.L. Digital Image Processing Using MATLAB; Gatesmark Publishing: Knoxville, TN, USA, 2009. [Google Scholar]
- Fu, F.; Wei, J.; Zhang, M.; Yu, F.; Xiao, Y.; Rong, D.; Shan, Y.; Li, Y.; Zhao, C.; Liao, F.; et al. Rapid vessel segmentation and reconstruction of head and neck angiograms using 3D convolutional neural network. Nat. Commun. 2020, 11, 4829. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Martínez, J.; Pérez-Palau, D.; Cilla, M.; Garrido, N.; Larrañaga, A.; Pérez-Rey, I. Semi-Automatic 3D Reconstruction of Atheroma Plaques from Intravascular Ultrasound Images Using an ad-hoc Algorithm. Mathematics 2023, 11, 537. https://doi.org/10.3390/math11030537
Martínez J, Pérez-Palau D, Cilla M, Garrido N, Larrañaga A, Pérez-Rey I. Semi-Automatic 3D Reconstruction of Atheroma Plaques from Intravascular Ultrasound Images Using an ad-hoc Algorithm. Mathematics. 2023; 11(3):537. https://doi.org/10.3390/math11030537
Chicago/Turabian StyleMartínez, Javier, Daniel Pérez-Palau, Myriam Cilla, Neus Garrido, Ana Larrañaga, and Ignacio Pérez-Rey. 2023. "Semi-Automatic 3D Reconstruction of Atheroma Plaques from Intravascular Ultrasound Images Using an ad-hoc Algorithm" Mathematics 11, no. 3: 537. https://doi.org/10.3390/math11030537
APA StyleMartínez, J., Pérez-Palau, D., Cilla, M., Garrido, N., Larrañaga, A., & Pérez-Rey, I. (2023). Semi-Automatic 3D Reconstruction of Atheroma Plaques from Intravascular Ultrasound Images Using an ad-hoc Algorithm. Mathematics, 11(3), 537. https://doi.org/10.3390/math11030537







