Genetic Diversity and Population Structure of Maize Inbred Lines with Varying Levels of Resistance to Striga Hermonthica Using Agronomic Trait-Based and SNP Markers
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. Trait Measurements
2.3. Genotyping
3. Statistical Analysis
3.1. Phenotypic Analysis
3.2. Quality Control and Genotypic Analysis
3.3. Joint Analysis of Agronomic-Based and Molecular Data
4. Results
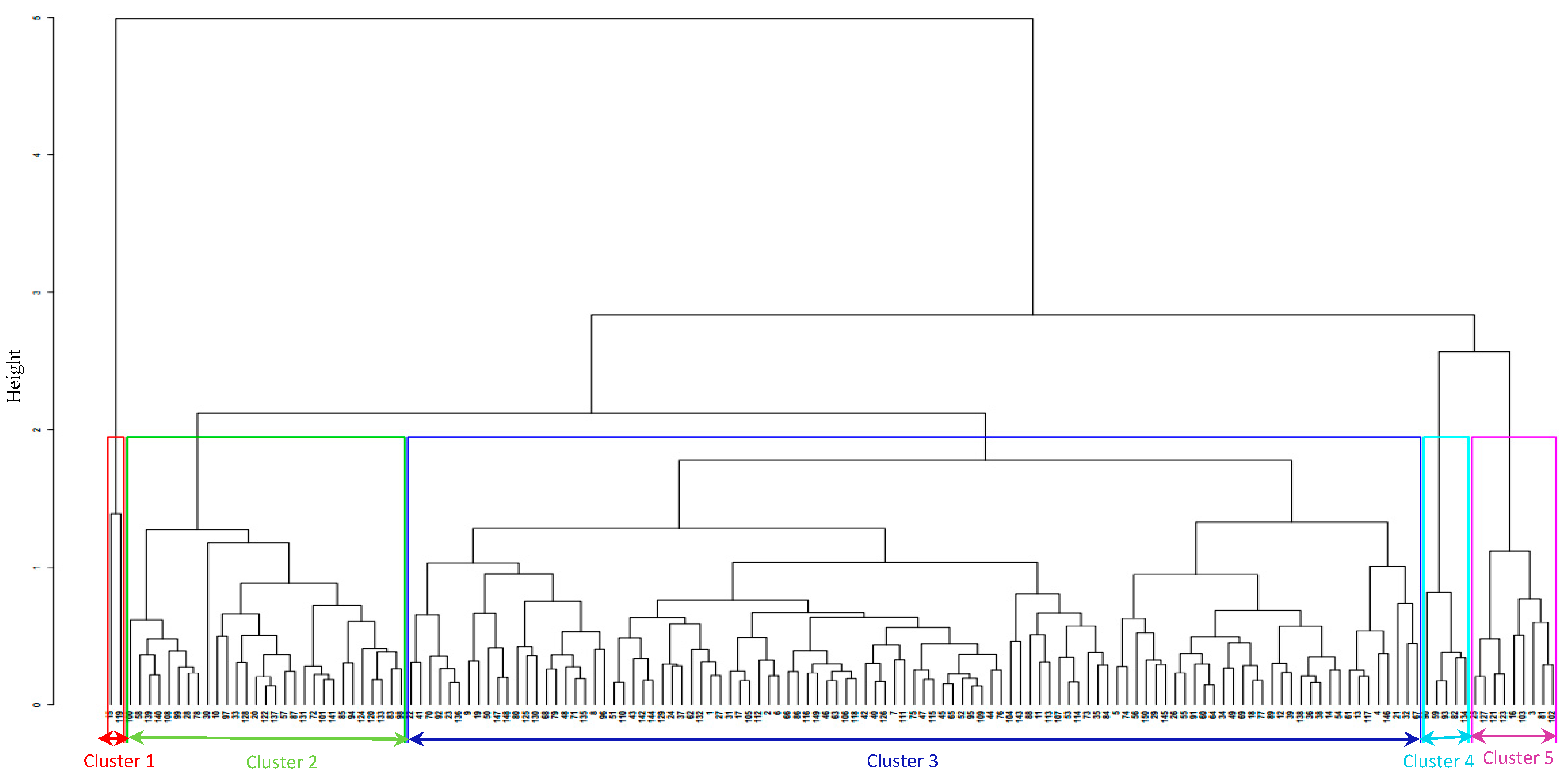
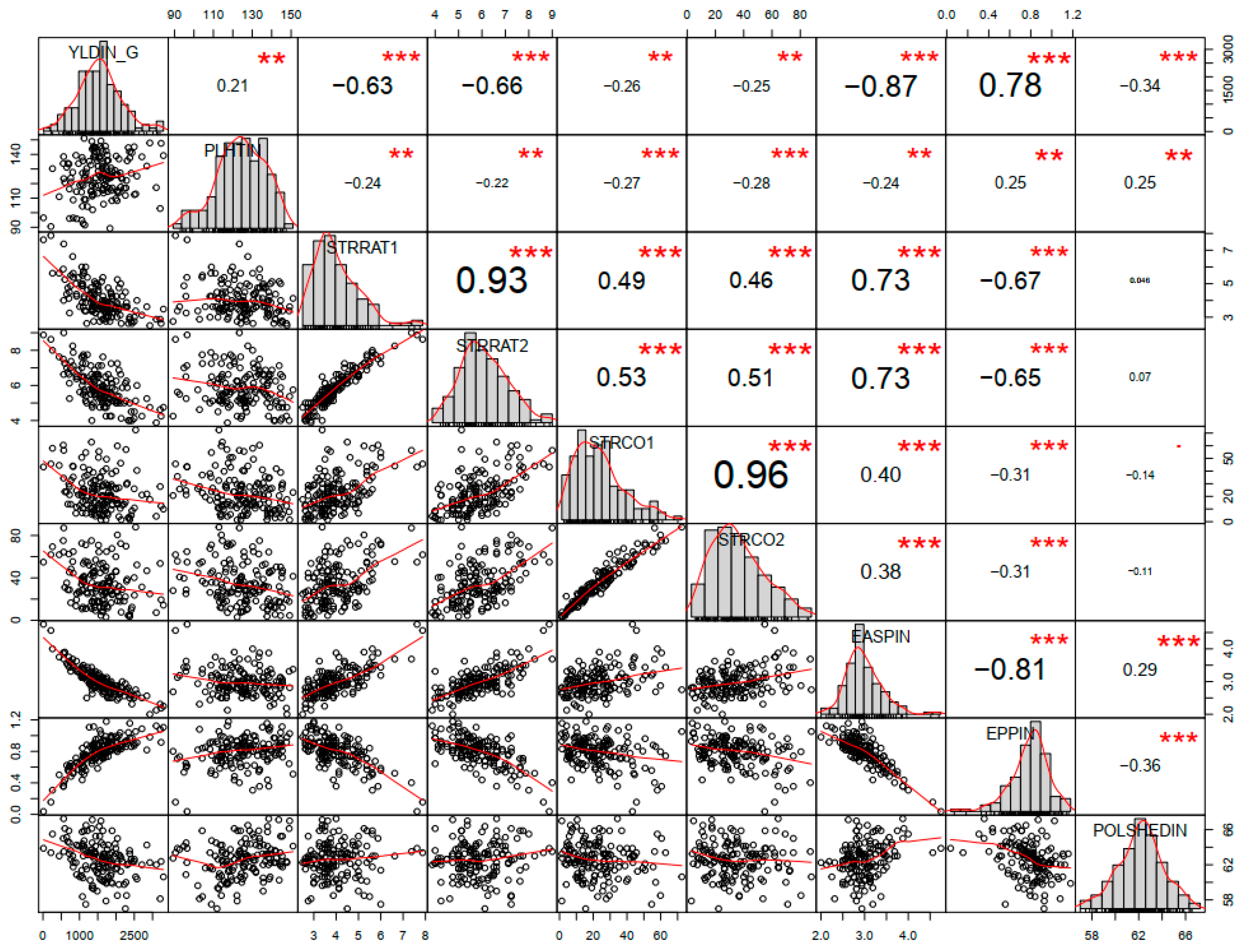
4.1. Diversity Based on Agronomic Traits
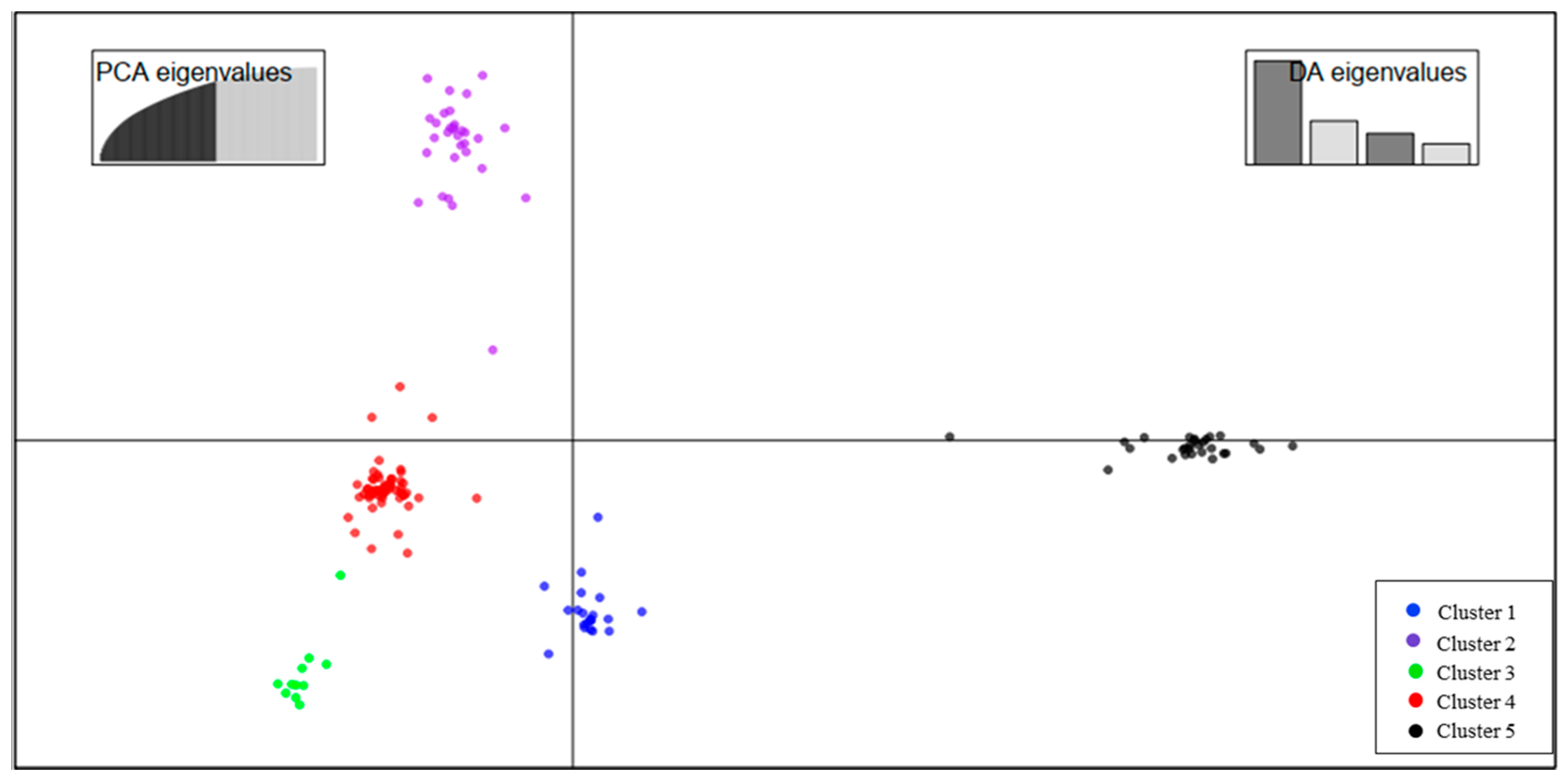
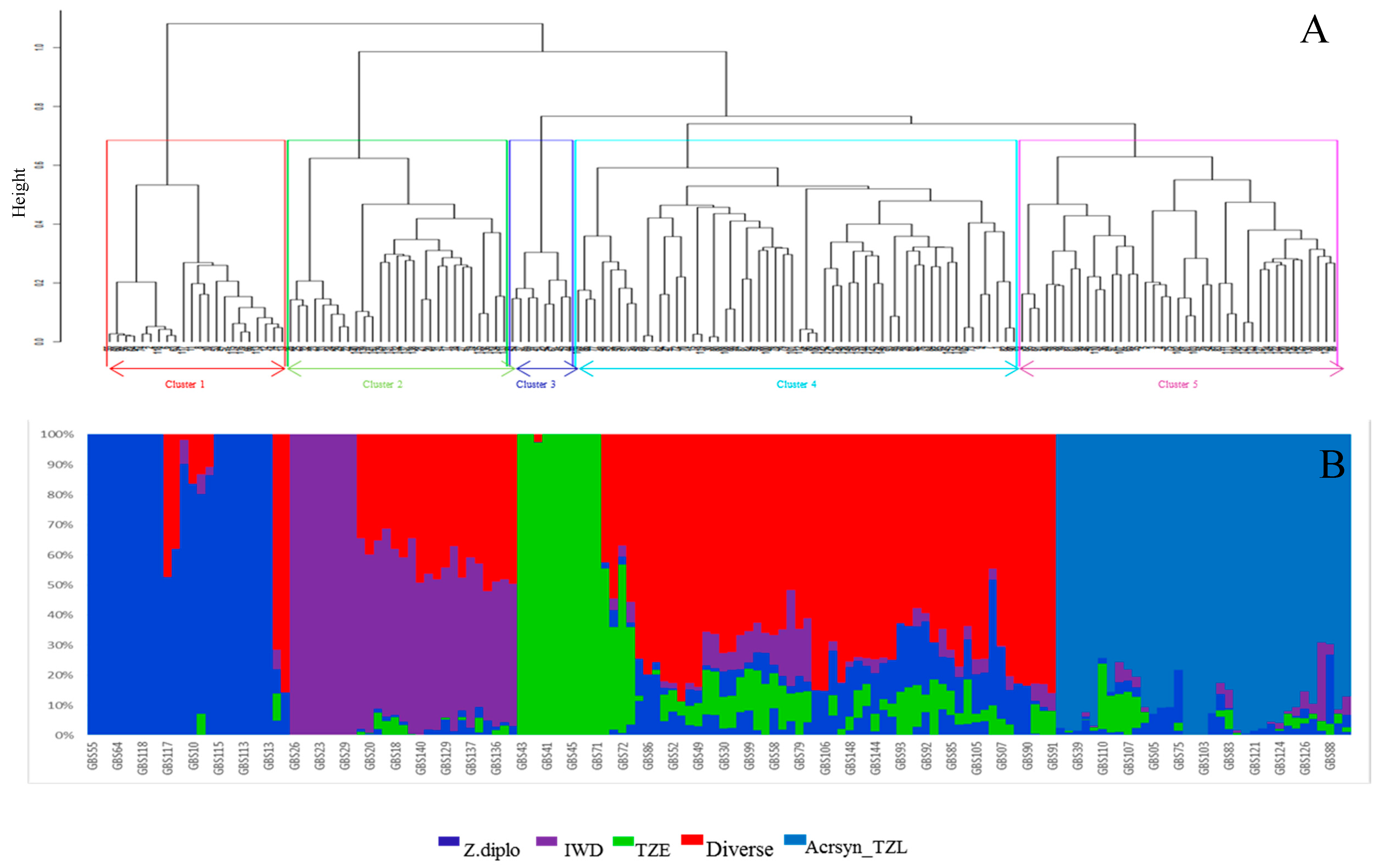
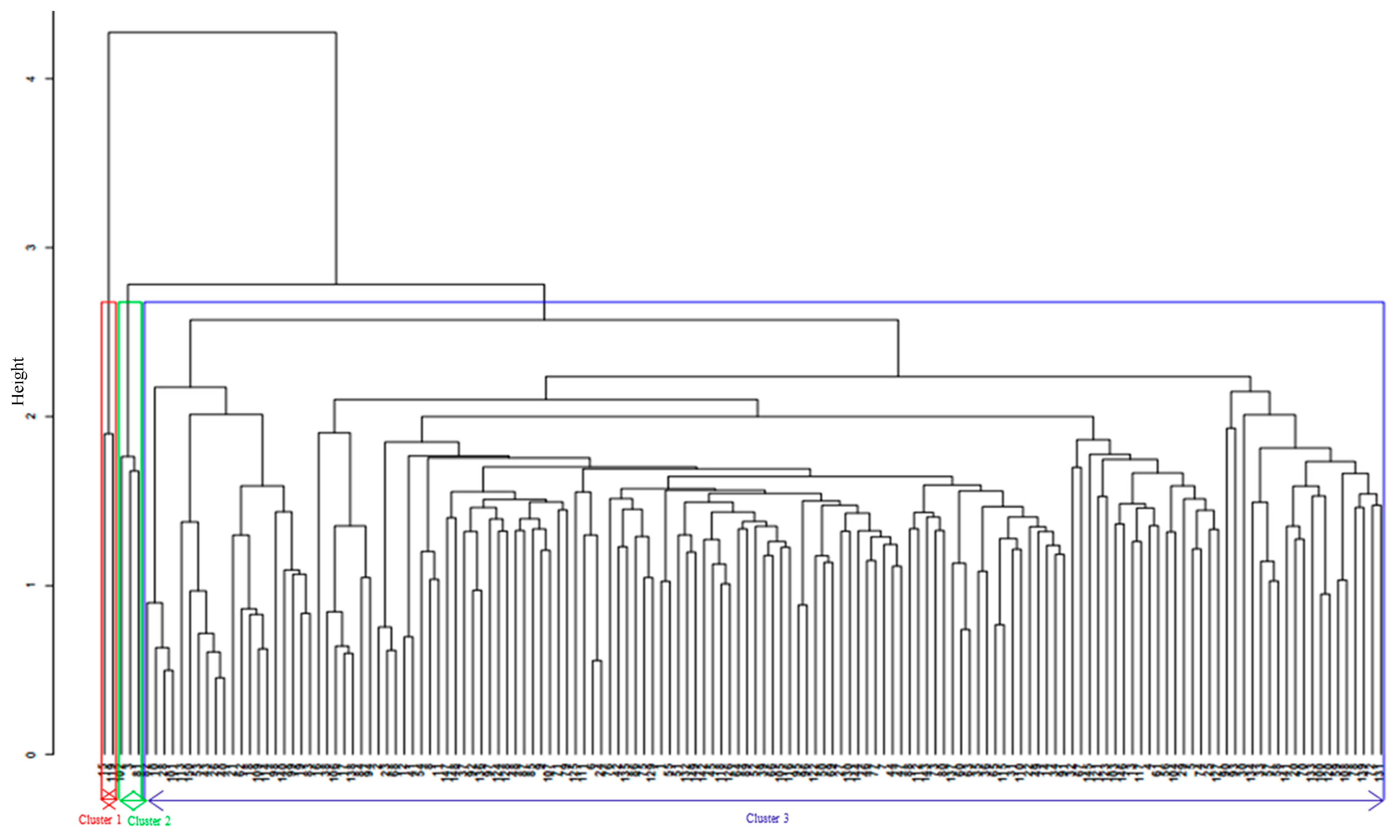
4.2. Genetic Diversity and Population Structure Based on SNP Markers
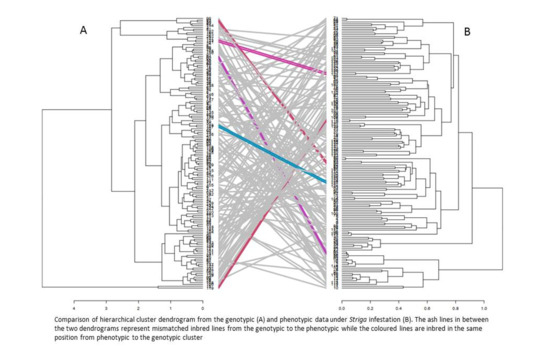
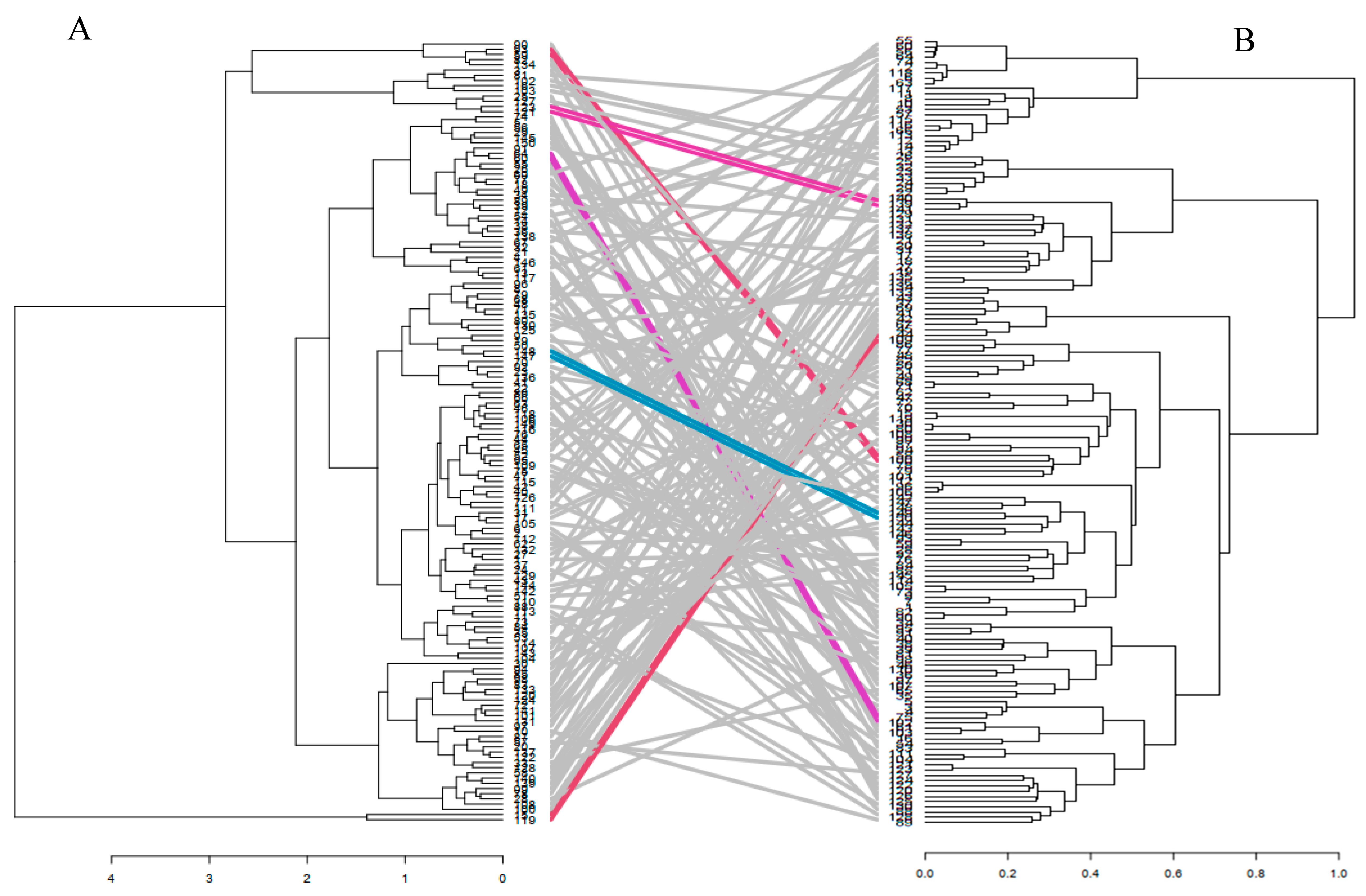
4.3. Combined Analysis of Phenotypic and Genotypic Data
5. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ANOVA | Analysis of variance |
| BIC | Bayesian information criterion |
| CTAB | Cetyltrimethylammonium bromide |
| DAPC | Discriminant analysis of principal components |
| DNA | Deoxyribonucleic acid |
| GBS | Genotyping-by-sequencing |
| IBS | Identity by State |
| LD | Linkage disequilibrium |
| SNP | Single nucleotide polymorphism |
| SSA | sub-Saharan Africa |
| PIC | Polymorphic information content |
| TASSEL | Trait Analysis by aSSociation, Evolution and Linkage |
References
- Shah, T.R.; Prasad, K.; Kumar, P. Maize: A potential source of human nutrition and health: A review. Cogent Food Agric. 2016, 2, 1166995. [Google Scholar]
- Shiferaw, B.; Prasanna, B.M.; Hellin, J.; Bänziger, M. Crops that feed the world 6. Past successes and future challenges to the role played by maize in global food security. Food Secur. 2011, 3, 307. [Google Scholar]
- FAOSTAT Food and Agriculture Organization of the United Nations Statistics Division; Economic and Social Development Department: Rome, Italy. Available online: http://faostat3.fao.org/home/E (accessed on 31 December 2016).
- Eyre-Walker, A.; Gaut, R.L.; Hilton, H.; Feldman, D.L.; Gaut, B.S. Investigation of the bottleneck leading to the domestication of maize. Proc. Natl. Acad. Sci. USA 1998, 95, 4441–4446. [Google Scholar]
- Matsuoka, Y.; Vigouroux, Y.; Goodman, M.M.; Sanchez, J.; Buckler, E.; Doebley, J. A single domestication for maize shown by multilocus microsatellite genotyping. Proc. Natl. Acad. Sci. USA 2002, 99, 6080–6084. [Google Scholar]
- Doebley, J. The genetics of maize evolution. Annu. Rev. Genet. 2004, 38, 37–59. [Google Scholar]
- Prasanna, B.M. Diversity in global maize germplasm: Characterization and utilization. J. Biosci. 2012, 37, 843–855. [Google Scholar]
- Buckler, E.S.; Gaut, B.S.; McMullen, M.D. Molecular and functional diversity of maize. Curr. Opin. Plant Biol. 2006, 9, 172–176. [Google Scholar]
- Whitt, S.R.; Wilson, L.M.; Tenaillon, M.I.; Gaut, B.S.; Buckler, E.S. Genetic diversity and selection in the maize starch pathway. Proc. Natl. Acad. Sci. USA 2002, 99, 12959–12962. [Google Scholar]
- M’mboyi, F.; Mugo, S.; Mwimali, M.; Ambani, L. Maize Production and Improvement in Sub-Saharan Africa; African Biotechnology Stakeholders Forum (ABSF): Nairobi, Kenya, 2010; Volume 2, pp. 14–17. [Google Scholar]
- Sauerborn, J. The economic importance of the phytoparasites Orobanche and Striga. In Proceedings of the 5th International Symposium of Parasitic Weeds, Nairobi, Kenya, 24–30 June 1991; pp. 137–143. [Google Scholar]
- Kanampiu, F.; Makumbi, D.; Mageto, E.; Omanya, G.; Waruingi, S.; Musyoka, P.; Ransom, J. Assessment of management options on Striga infestation and maize grain yield in Kenya. Weed Sci. 2018, 66, 516–524. [Google Scholar]
- Ransom, J.K. Long-term approaches for the control of Striga in cereals: Field management options. Crop Prot. 2000, 19, 759–763. [Google Scholar]
- Kim, S.K.; Winslow, M.D. Progress in breeding maize for tolerance/resistance at IITA. In Proceedings of the Fifth International Symposium on Parasitic Weeds, Nairobi, Kenya, 24–30 June 1991. [Google Scholar]
- Kling, J.G.; Fajemisin, J.M.; Badu-Apraku, B.; Diallo, A.; Menkir, A.; Melake-Berhan, A.; Haussmann, B.I. Striga resistance breeding in maize. In Breeding for Striga Resistance in Cereals; Margraf Verlag: Weikersheim, Germany, 2000; pp. 103–118. [Google Scholar]
- Mafakheri, K.; Bihamta, M.R.; Abbasi, A.R. Assessment of genetic diversity in cowpea (Vigna unguiculata L.) germplasm using morphological and molecular characterisation. Cogent Food Agric. 2017, 3, 1327092. [Google Scholar]
- Mahato, A.; Shahi, J.P.; Singh, P.K.; Kumar, M. Genetic diversity of sweet corn inbreds using agro-morphological traits and microsatellite markers. 3 Biotech 2018, 8, 332. [Google Scholar]
- Ali, M.L.; Rajewski, J.F.; Baenziger, P.S.; Gill, K.S.; Eskridge, K.M.; Dweikat, I. Assessment of genetic diversity and relationship among a collection of U.S. sweet sorghum germplasm by SSR markers. Mol. Breed. 2008, 21, 497–509. [Google Scholar]
- Menkir, A. Assessment of reactions of diverse maize inbred lines to Striga hermonthica (Del.) Benth. Plant Breed. 2006, 125, 131–139. [Google Scholar]
- Semagn, K.; Magorokosho, C.; Vivek, B.S.; Makumbi, D.; Beyene, Y.; Mugo, S.; Prasanna, B.M.; Warburton, M.L. Molecular characterization of diverse CIMMYT maize inbred lines from eastern and southern Africa using single nucleotide polymorphic markers. BMC Genom. 2012, 13, 1. [Google Scholar]
- Ertiro, B.T.; Semagn, K.; Das, B.; Olsen, M.; Labuschagne, M.; Worku, M.; Wegary, D.; Azmach, G.; Ogugo, V.; Keno, T.; et al. Genetic variation and population structure of maize inbred lines adapted to the mid-altitude sub-humid maize agro-ecology of Ethiopia using single nucleotide polymorphic (SNP) markers. BMC Genom. 2017, 18, 1. [Google Scholar]
- Hayward, M.D.; Breese, E.L. Population structure and variability. In Plant Breeding; Springer: Dordrecht, The Netherlands, 1993; pp. 16–29. [Google Scholar]
- Melchinger, A.E.; Gumber, R.K. Overview of heterosis and heterotic groups in agronomic crops. Concepts Breed. Heterosis Crop Plants 1998, 25, 29–44. [Google Scholar]
- Nadeem, M.A.; Nawaz, M.A.; Shahid, M.Q.; Doğan, Y.; Comertpay, G.; Yıldız, M.; Hatipoğlu, R.; Ahmad, F.; Alsaleh, A.; Labhane, N.; et al. DNA molecular markers in plant breeding: Current status and recent advancements in genomic selection and genome editing. Biotechnol. Biotechnol. Equip. 2018, 32, 261–285. [Google Scholar]
- Rahman, S.; Mia, M.M.; Quddus, T.; Hassan, L.; Haque, M.A. Assessing genetic diversity of maize (Zea mays L.) genotypes for agronomic traits. Res. Agric. Livest. Fish. 2015, 2, 53–61. [Google Scholar]
- Govindaraj, M.; Vetriventhan, M.; Srinivasan, M. Importance of genetic diversity assessment in crop plants and its recent advances: An overview of its analytical perspectives. Genet. Res. Int. 2015, 2015, 431487. [Google Scholar]
- Giordani, W.; Scapim, C.A.; Ruas, P.M.; Ruas, C.D.; Contreras-Soto, R.; Coan, M.; Fonseca, I.C.; Gonçalves, L.S. Genetic diversity, population structure and AFLP markers associated with maize reaction to southern rust. Bragantia 2019, 78, 183–196. [Google Scholar]
- Sserumaga, J.P.; Makumbi, D.; Ji, H.; Njoroge, K.; Muthomi, J.W.; Chemining’wa, G.N.; Si-myung, L.; Asea, G.; Kim, H. Molecular characterization of tropical maize inbred lines using microsatellite DNA markers. Maydica 2014, 59, 267–274. [Google Scholar]
- Cömertpay, G.; Baloch, F.S.; Kilian, B.; Ülger, A.C.; Özkan, H. Diversity assessment of Turkish maize landraces based on fluorescent labelled SSR markers. Plant Mol. Biol. Rep. 2012, 30, 261–274. [Google Scholar]
- Deschamps, S.; Llaca, V.; May, G.D. Genotyping-by-sequencing in plants. Biology 2012, 1, 460–483. [Google Scholar]
- Mengesha, W.A.; Menkir, A.; Unakchukwu, N.; Meseka, S.; Farinola, A.; Girma, G.; Gedil, M. Genetic diversity of tropical maize inbred lines combining resistance to Striga hermonthica with drought tolerance using SNP markers. Plant Breed. 2017, 136, 338–343. [Google Scholar]
- Cooper, J.S.; Rice, B.R.; Shenstone, E.M.; Lipka, A.E.; Jamann, T.M. Genome-wide analysis and prediction of resistance to goss’s wilt in maize. Plant Genome 2019, 12, 1–10. [Google Scholar]
- Sartie, A.; Asiedu, R.; Franco, J. Genetic and phenotypic diversity in a germplasm working collection of cultivated tropical yams (Dioscorea spp.). Genet. Resour. Crop Evol. 2012, 59, 1753–1765. [Google Scholar]
- Andrade, E.K.; Júnior, A.V.C.; Laia, M.L.; Fernandes, J.S.; Oliveira, A.J.; Azevedo, A.M. Genetic dissimilarity among sweet potato genotypes using morphological and molecular descriptors. Acta Sci. Agron. 2017, 39, 447–455. [Google Scholar]
- Najaphy, A.; Parchin, R.A.; Farshadfar, E. Comparison of phenotypic and molecular characterizations of some important wheat cultivars and advanced breeding lines. Aust. J. Crop Sci. 2012, 6, 326. [Google Scholar]
- Agre, P.; Asibe, F.; Darkwa, K.; Edemodu, A.; Bauchet, G.; Asiedu, R.; Adebola, P.; Asfaw, A. Phenotypic and molecular assessment of genetic structure and diversity in a panel of winged yam (Dioscorea alata) clones and cultivars. Sci. Rep. 2019, 9, 18221. [Google Scholar]
- Kwabena, D.; Paterne, A.; Bunmi, O.; Kohtaro, I.; Ryo, M.; Powell, A.; Guillaume, B.; Satoru, M.; Adebola, P.; Asiedu, R.; et al. Comparative assessment of genetic diversity matrices and clustering methods in white Guinea yam (Dioscorea rotundata) based on morphological and molecular markers. Sci. Rep. 2020, 10, 13191. [Google Scholar]
- Belalia, N.; Lupini, A.; Djemel, A.; Morsli, A.; Mauceri, A.; Lotti, C.; Khelifi-Slaoui, M.; Khelifi, L.; Sunseri, F. Analysis of genetic diversity and population structure in Saharan maize (Zea mays L.) populations using phenotypic traits and SSR markers. Genet. Resour. Crop Evol. 2019, 66, 243–257. [Google Scholar]
- Kim, S.K.; Akintunde, A.Y.; Walker, P. Responses of maize, sorghum and millet host plants to infestation by Striga hermonthica. Crop Prot. 1994, 13, 582–590. [Google Scholar]
- Saghai-Maroof, M.A.; Soliman, K.M.; Jorgensen, R.A.; Allard, R.W. Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. Proc. Natl. Acad. Sci. USA 1984, 81, 8014–8018. [Google Scholar]
- Elshire, R.J.; Glaubitz, J.C.; Sun, Q.; Poland, J.A.; Kawamoto, K.; Buckler, E.S.; Mitchell, S.E. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 2011, 6, e19379. [Google Scholar]
- Glaubitz, J.C.; Casstevens, T.M.; Lu, F.; Harriman, J.; Elshire, R.J.; Sun, Q.; Buckler, E.S. TASSEL-GBS: A high capacity genotyping by sequencing analysis pipeline. PLoS ONE 2014, 9, e90346. [Google Scholar]
- Vargas, M.; Combs, E.; Alvarado, G.; Atlin, G.; Mathews, K.; Crossa, J. META: A suite of SAS programs to analyze multienvironment breeding trials. Agron. J. 2013, 105, 9–11. [Google Scholar]
- Lê, S.; Josse, J.; Husson, F. FactoMineR: An R package for multivariate analysis. J. Stat. Softw. 2008, 25, 1–8. [Google Scholar]
- Peres-Neto, P.R.; Jackson, D.A.; Somers, K.M. Giving meaningful interpretation to ordination axes: Assessing loading significance in principal component analysis. Ecology 2003, 84, 2347–2363. [Google Scholar]
- Maechler, M. Finding Groups in Data: Cluster Analysis Extended Rousseeuw et al. R Package Version 2.0. Available online: ftp://128.61.111.11/pub/CRAN/web/packages/cluster/cluster.pdf (accessed on 7 June 2019).
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.; Daly, M.J.; et al. PlinK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar]
- Alexander, D.H.; Novembre, J.; Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 2009, 19, 1655–1664. [Google Scholar]
- Alexander, D.H.; Lange, K. Enhancements to the ADMIXTURE algorithm for individual ancestry estimation. BMC Bioinform. 2011, 12, 246. [Google Scholar]
- Jombart, T.; Devillard, S.; Balloux, F. Discriminant analysis of principal components: A new method for the analysis of genetically structured populations. BMC Genet. 2010, 11, 94. [Google Scholar]
- R Core Team. A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018. [Google Scholar]
- Paradis, E.; Claude, J.; Strimmer, K. APE: Analyses of phylogenetics and evolution in R language. Bioinformatics 2004, 20, 289–290. [Google Scholar]
- Galili, T. Dendextend: An R package for visualizing, adjusting and comparing trees of hierarchical clustering. Bioinformatics 2015, 31, 3718–3720. [Google Scholar]
- Menkir, A.; Makumbi, D.; Franco, J. Assessment of reaction patterns of hybrids to Striga hermonthica (Del.) Benth. under artificial infestation in Kenya and Nigeria. Crop Sci. 2012, 52, 2528–2537. [Google Scholar]
- Menkir, A.; Meseka, S. Genetic improvement in resistance to Striga in tropical maize hybrids. Crop Sci. 2019, 59, 2484–2497. [Google Scholar]
- Al-Naggar, A.M.; Shafik, M.M.; Musa, R.Y. Genetic diversity based on morphological traits of 19 maize genotypes using principal component analysis and G.T. biplot. Annu. Res. Rev. Biol. 2020, 35, 68–85. [Google Scholar]
- Wang, C.H.; Zheng, X.M.; Xu, Q.; Yuan, X.P.; Huang, L.; Zhou, H.F.; Wei, X.H.; Ge, S. Genetic diversity and classification of Oryza sativa with emphasis on Chinese rice germplasm. Heredity 2014, 112, 489–496. [Google Scholar]
- Fatokun, C.; Girma, G.; Abberton, M.; Gedil, M.; Unachukwu, N.; Oyatomi, O.; Yusuf, M.; Rabbi, I.; Boukar, O. Genetic diversity and population structure of a mini-core subset from the world cowpea (Vigna unguiculata (L.) Walp.) germplasm collection. Sci. Rep. 2018, 8, 16035. [Google Scholar]
- Singh, S.P.; Nodari, R.; Gepts, P. Genetic diversity in cultivated common bean: I. Allozymes. Crop Sci. 1991, 31, 19–23. [Google Scholar]
- Alves, A.A.; Bhering, L.L.; Rosado, T.B.; Laviola, B.G.; Formighieri, E.F.; Cruz, C.D. Joint analysis of phenotypic and molecular diversity provides new insights on the genetic variability of the Brazilian physic nut germplasm bank. Genet. Mol. Biol. 2013, 36, 371–381. [Google Scholar]
- Collard, B.C.; Jahufer, M.Z.; Brouwer, J.B.; Pang, E.C. An introduction to markers, quantitative trait loci (QTL) mapping and marker-assisted selection for crop improvement: The basic concepts. Euphytica 2005, 142, 169–196. [Google Scholar]
- Sunil, N.; Sujatha, M.; Kumar, V.; Vanaja, M.; Basha, S.D.; Varaprasad, K.S. Correlating the phenotypic and molecular diversity in Jatropha curcas L. Biomass Bioenergy 2011, 35, 1085–1096. [Google Scholar]
- Ristić, D.; Babić, V.; Anđelković, V.; Vančetović, J.; Mladenović-Drinić, S.; Ignjatović-Micić, D. Genetic diversity in maize dent landraces assessed by morphological and molecular markers. Genetika 2013, 45, 811–824. [Google Scholar]
- Reis, R.V.; Viana, A.P.; Oliveira, E.J.; Silva, M.G. Phenotypic and molecular selection of yellow passion fruit progenies in the second cycle of recurrent selection. Crop Breed. Appl. Biotechnol. 2012, 12, 17–24. [Google Scholar]
| Source Population | Genetic Background of Source Population | Lines Evaluated |
|---|---|---|
| ZDIP | Lines derived from a backcross (BC4) containing a Z. diploperennis accession as a donor parent plus lines derived from bi-parental crosses involving one parent derived from the same Z. diploperennis backcross | 39 |
| TZLC | A late-maturing composite formed by crossing TZB-SR with seven inbred lines developed for field resistance to S. hermonthica plus lines derived from bi-parental crosses involving one parent derived from the same source populations | 44 |
| TZEC | An early maturing composite formed by crossing TZESR-W C3 with eight inbred lines developed for field resistance to S. hermonthica plus lines derived from bi-parental crosses involving one parent derived from the same composite | 13 |
| IWDS | A synthetic formed from medium maturing white inbred lines and improved for resistance to Striga and drought plus lines derived from bi-parental crosses involving one parent derived from these synthetic | 30 |
| MIXED | Lines derived from diverse source populations plus the tolerant lines expensively used as donors of field resistance to form populations | 24 |
| Variables | PC1 | PC2 | PC3 |
|---|---|---|---|
| Days to silking | −0.441 | 0.855 | −0.068 |
| Ear aspect | 0.884 | 0.070 | −0.303 |
| Ear per plant | −0.824 | −0.064 | 0.324 |
| Plant height | −0.388 | 0.212 | −0.221 |
| Striga count at 8 WAP | 0.651 | 0.148 | 0.712 |
| Striga count at 10 WAP | 0.633 | 0.182 | 0.719 |
| Striga damage at 8 WAP | 0.878 | 0.107 | −0.092 |
| Striga damage at 10 WAP | 0.881 | 0.201 | −0.085 |
| Yield | −0.786 | −0.173 | 0.434 |
| Anthesis silking interval | −0.302 | 0.908 | 0.014 |
| Eigenvalue | 4.886 | 1.745 | 1.478 |
| Variance (%) | 48.863 | 17.459 | 14.784 |
| Cumulative variance (%) | 48.863 | 66.321 | 81.105 |
| MAF | Ho | He | PIC | |
|---|---|---|---|---|
| Minimum | 0.05 | 0.00 | 0.10 | 0.1 |
| Maximum | 0.50 | 0.98 | 0.50 | 0.47 |
| Mean | 0.24 | 0.10 | 0.32 | 0.26 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stanley, A.; Menkir, A.; Paterne, A.; Ifie, B.; Tongoona, P.; Unachukwu, N.; Meseka, S.; Mengesha, W.; Gedil, M. Genetic Diversity and Population Structure of Maize Inbred Lines with Varying Levels of Resistance to Striga Hermonthica Using Agronomic Trait-Based and SNP Markers. Plants 2020, 9, 1223. https://doi.org/10.3390/plants9091223
Stanley A, Menkir A, Paterne A, Ifie B, Tongoona P, Unachukwu N, Meseka S, Mengesha W, Gedil M. Genetic Diversity and Population Structure of Maize Inbred Lines with Varying Levels of Resistance to Striga Hermonthica Using Agronomic Trait-Based and SNP Markers. Plants. 2020; 9(9):1223. https://doi.org/10.3390/plants9091223
Chicago/Turabian StyleStanley, Adekemi, Abebe Menkir, Agre Paterne, Beatrice Ifie, Pangirayi Tongoona, Nnanna Unachukwu, Silvestro Meseka, Wende Mengesha, and Melaku Gedil. 2020. "Genetic Diversity and Population Structure of Maize Inbred Lines with Varying Levels of Resistance to Striga Hermonthica Using Agronomic Trait-Based and SNP Markers" Plants 9, no. 9: 1223. https://doi.org/10.3390/plants9091223








