Molecular Cloning and Functional Characterization of CpMYC2 and CpBHLH13 Transcription Factors from Wintersweet (Chimonanthus praecox L.)
Abstract
1. Introduction
2. Results
2.1. Characterization of CpMYC2 and bHLH13
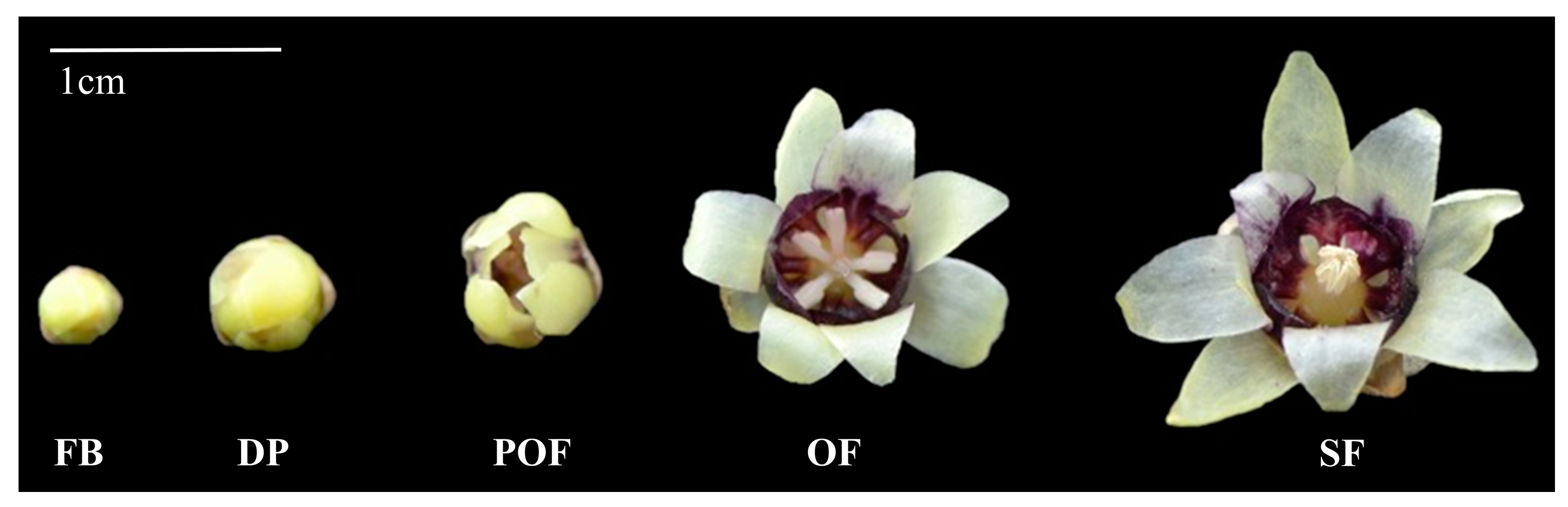
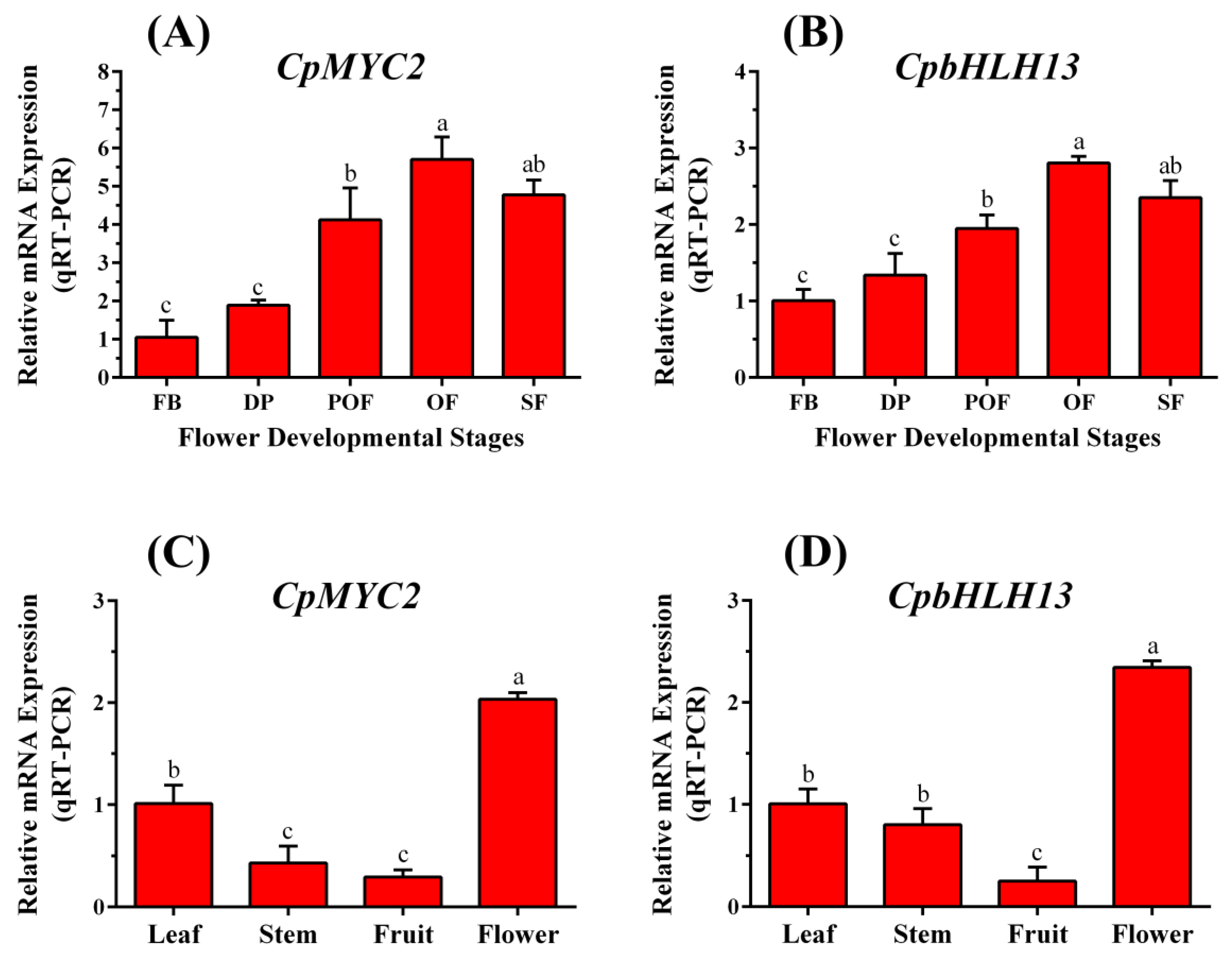
2.2. CpMYC2 and CpbHLH13 Expression in Wintersweet
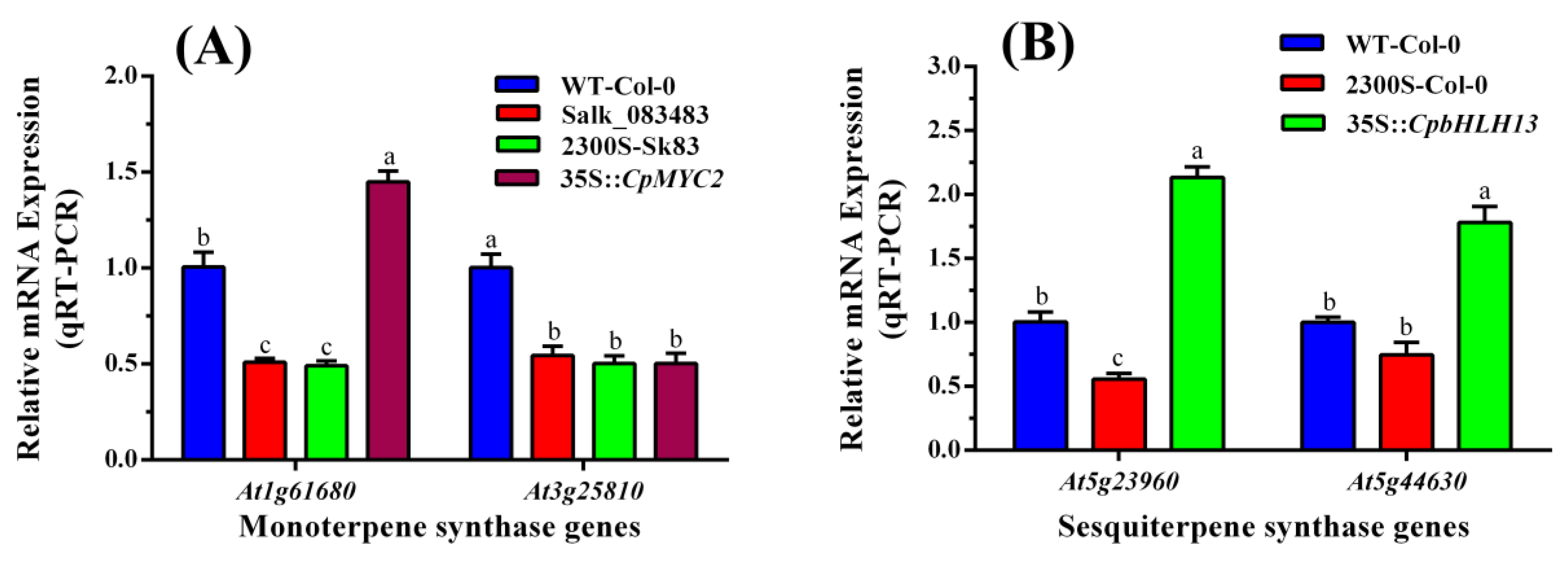
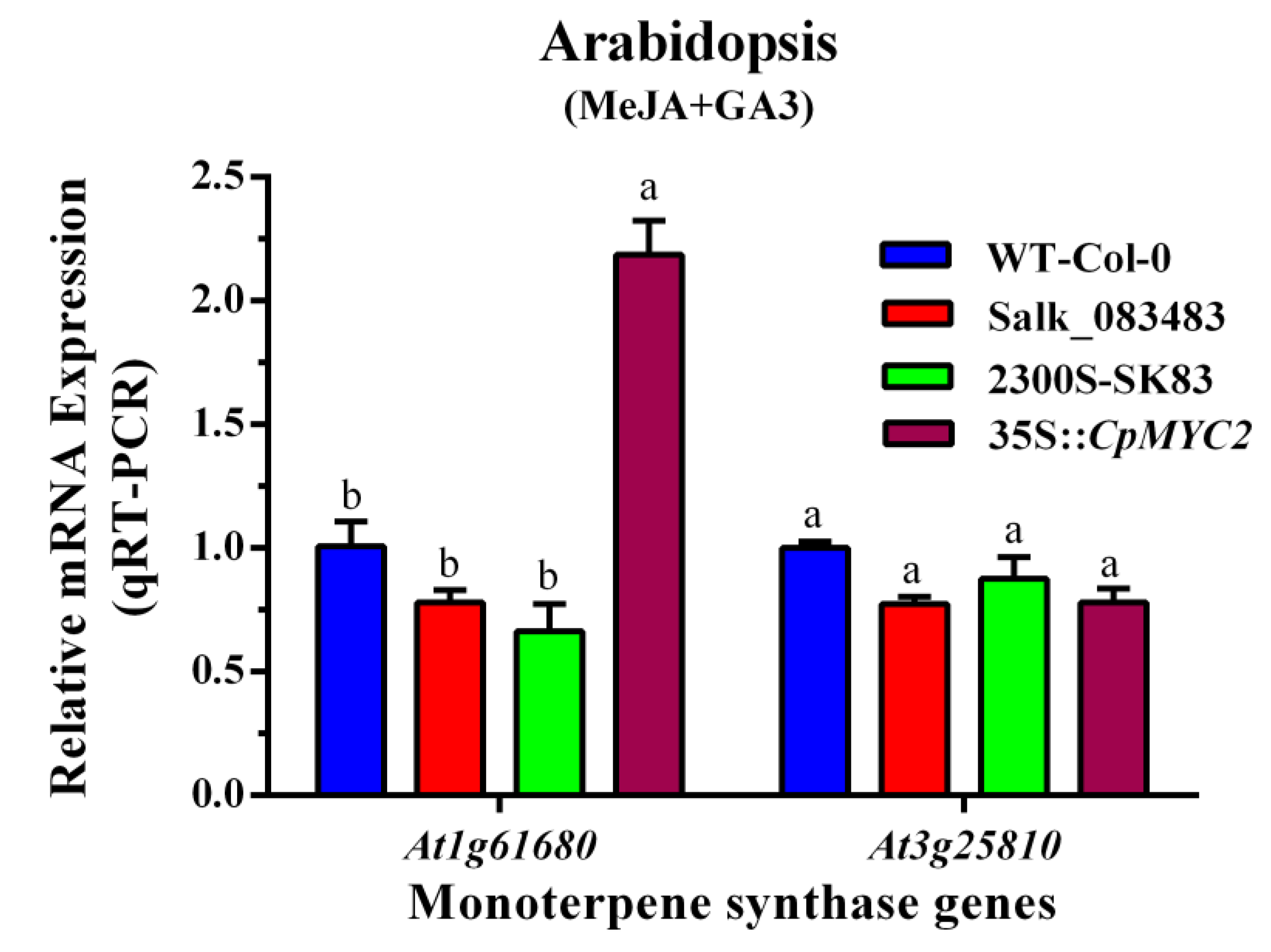
2.3. Expression Analysis of the Terpene Synthase Genes in Wild-Type and Transgenic Plants
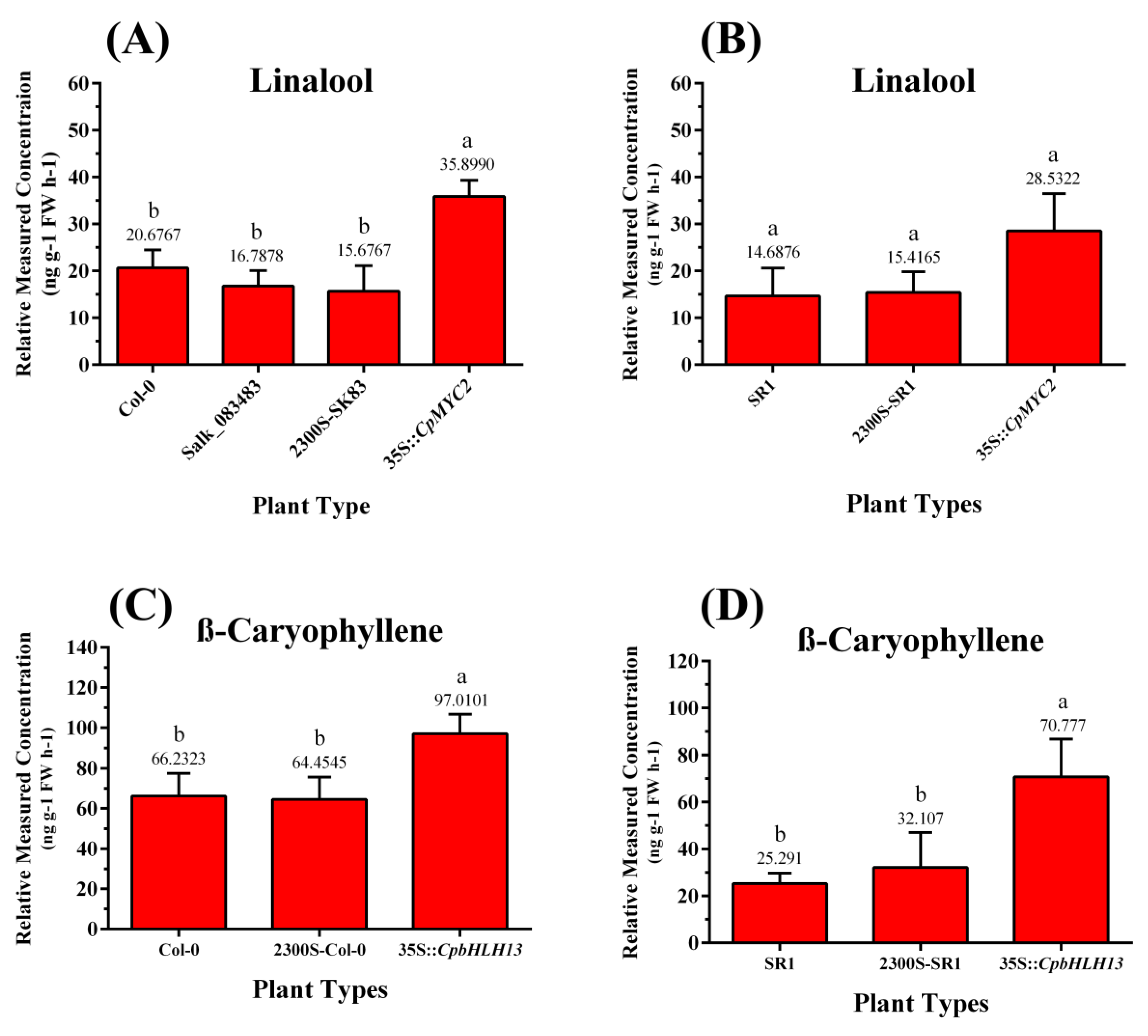
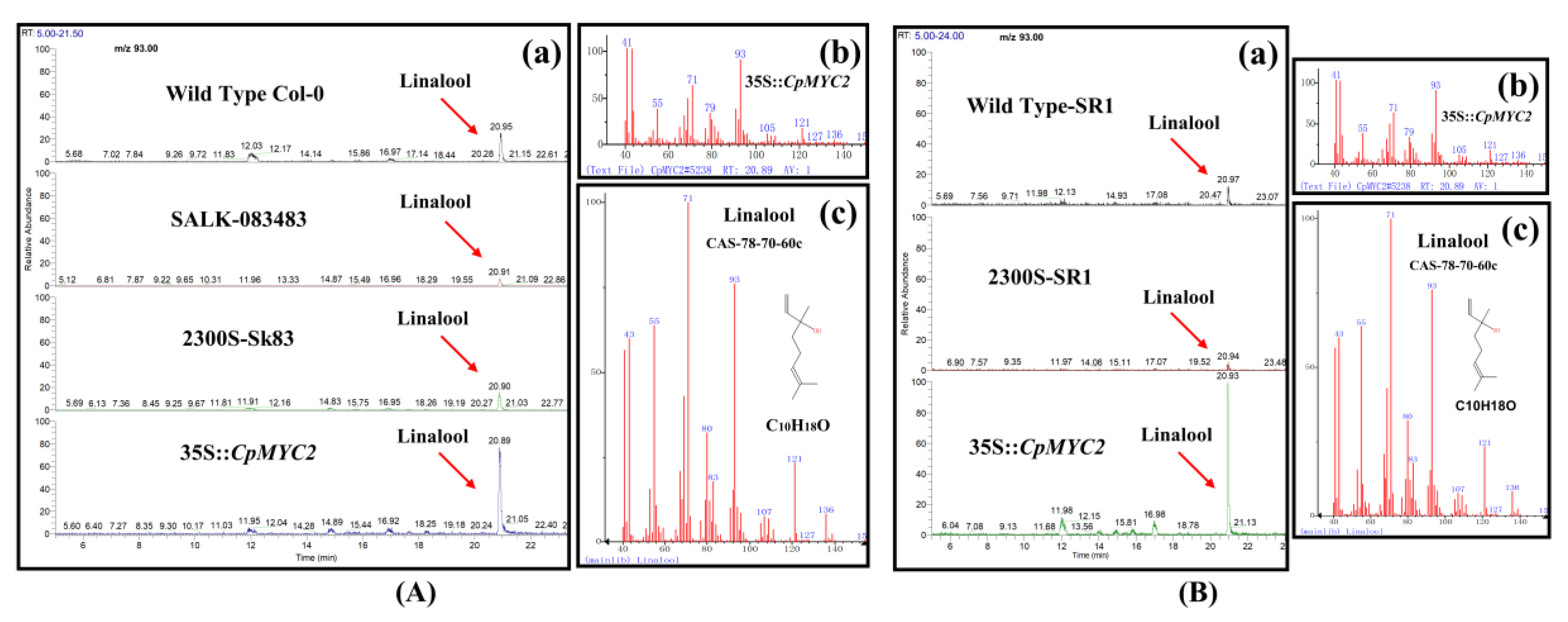
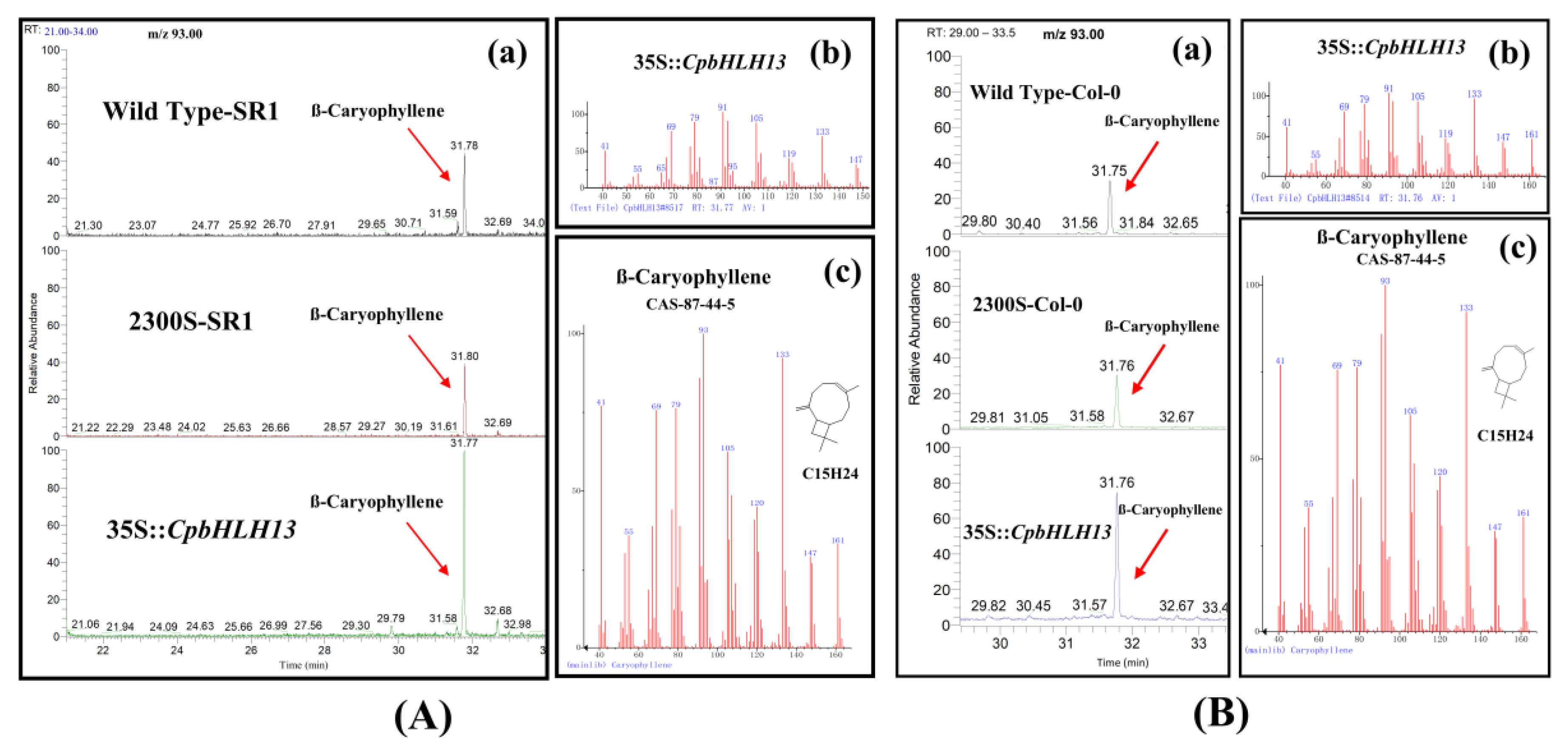
2.4. Overexpression of CpMYC2 and CpbHLH13 Induces VOC Emission
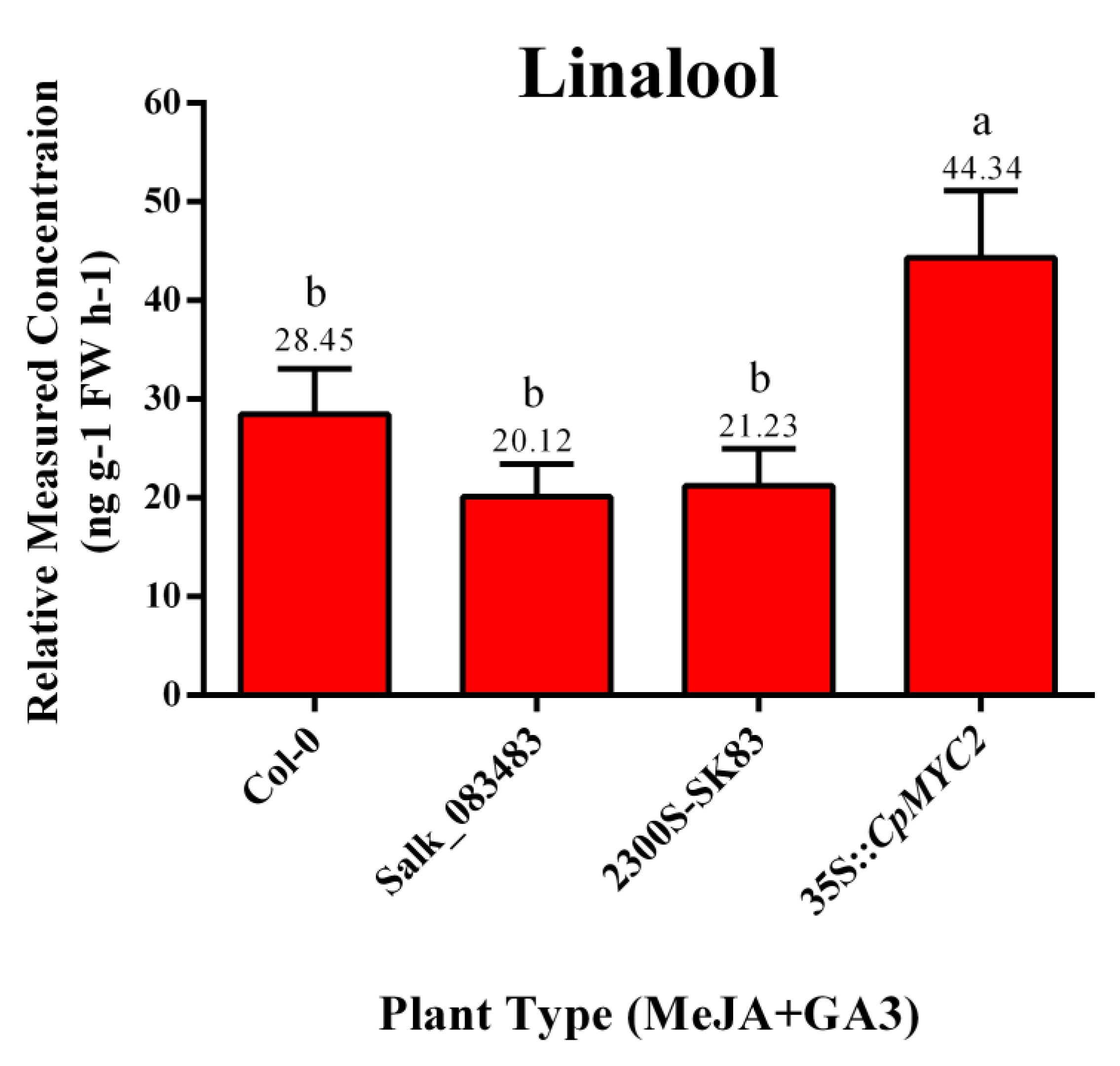
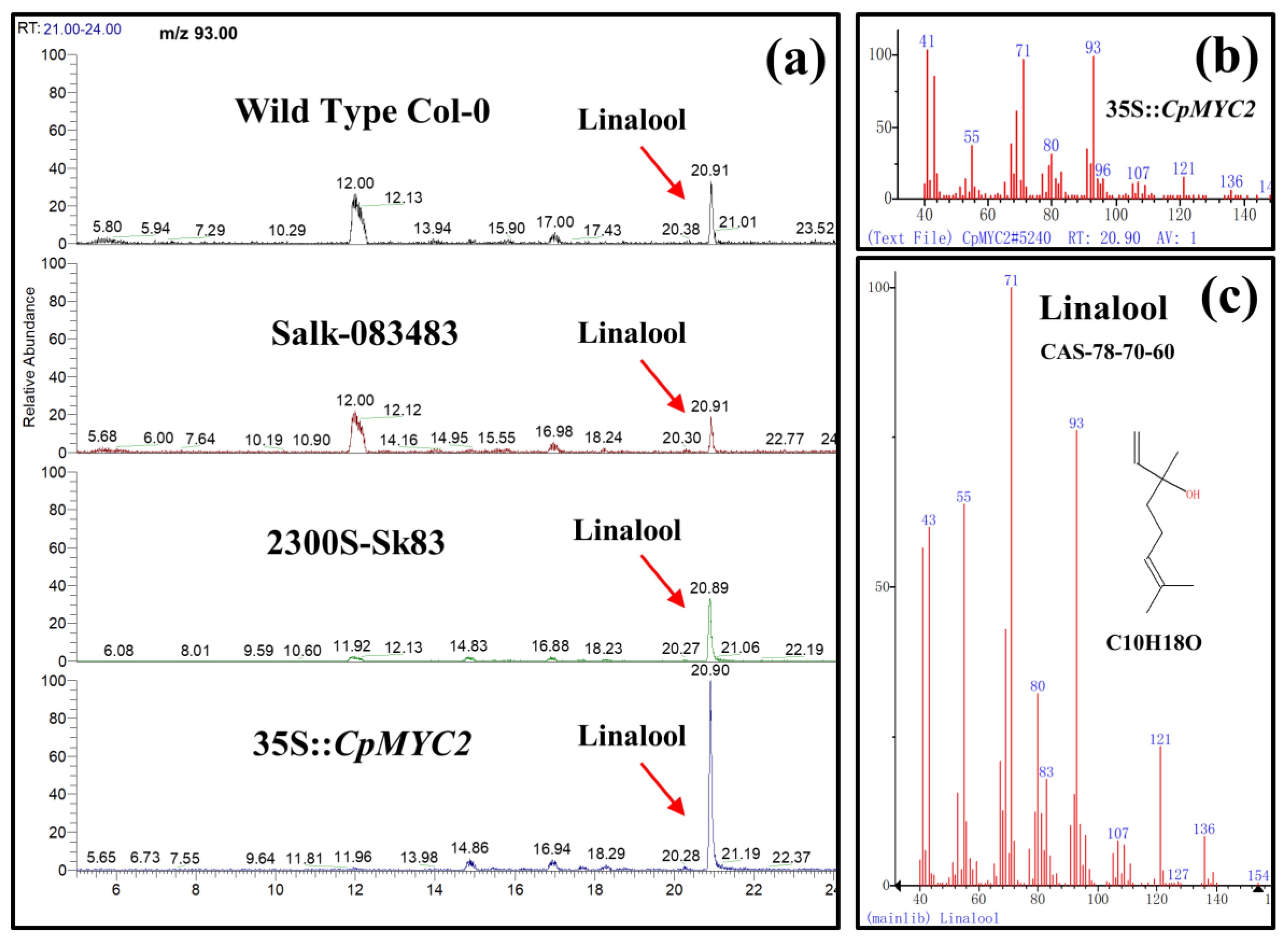
2.5. Application of MeJA and GA3 on Overexpressed CpMYC2 Plants
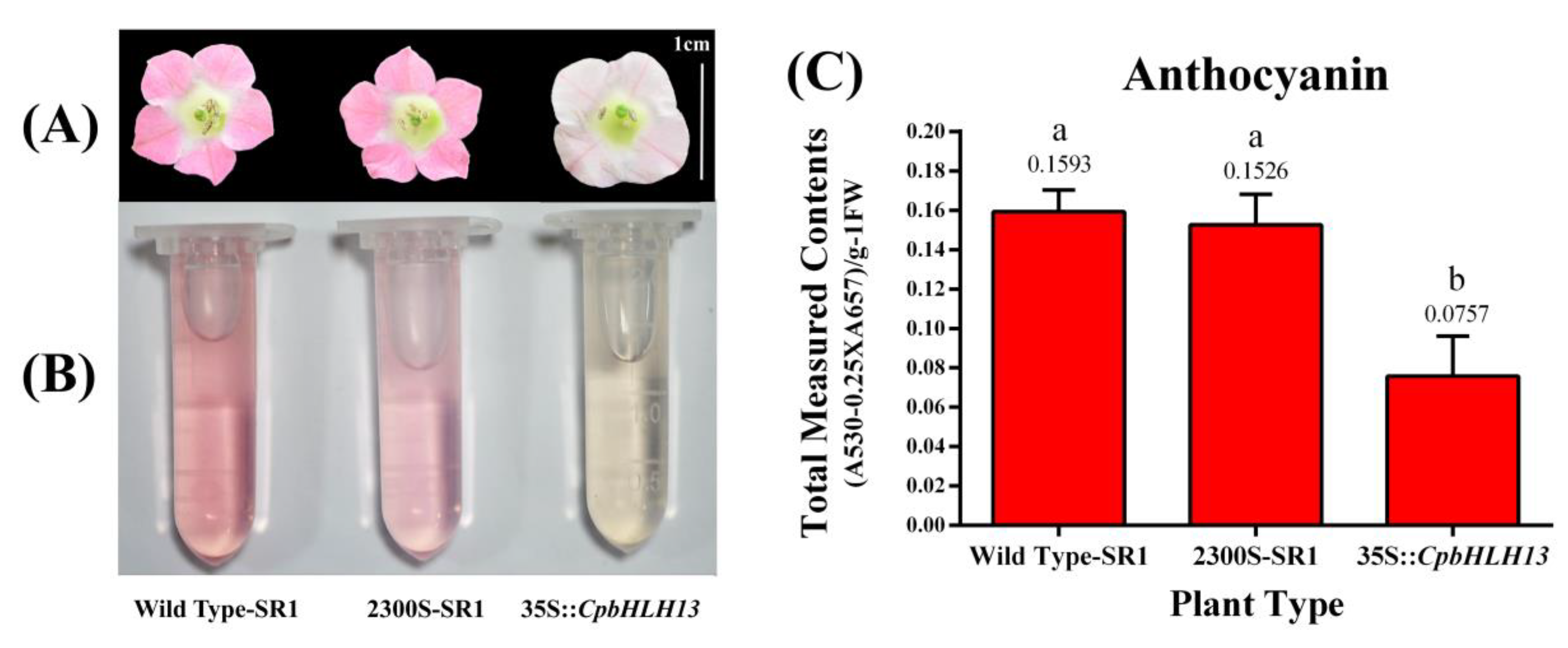
2.6. Overexpression of CpbHLH13 Reduces Flower Pigmentation in Tobacco Plants
3. Discussion
4. Materials and Methods
4.1. Selection and Analysis of Candidate Genes
4.2. Plant Materials
4.3. Total RNA Extraction, cDNA Synthesis, and Quantitative Real-Time PCR (qRT-PCR) Analysis
4.4. Isolation, Cloning, and Sequencing of Candidate Genes
4.5. Expression Vector Construction and Transformation
4.6. Volatile Organic Compounds Analysis
4.7. Application of MeJA and GA3 to Arabidopsis Plants
4.8. Anthocyanin Contents Measurements
4.9. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Howe, G.A.; Major, I.T.; Koo, A.J. Modularity in Jasmonate Signaling for Multistress Resilience. Annu. Rev. Plant Biol. 2018, 69, 387–415. [Google Scholar] [CrossRef]
- Sharifi, R.; Lee, S.-M.; Ryu, C.-M. Microbe-induced plant volatiles. New Phytol. 2017, 220, 684–691. [Google Scholar] [CrossRef]
- Dudareva, N.; Klempien, A.; Muhlemann, K.; Kaplan, I.; Muhlemann, J.K.; Kaplan, I. Biosynthesis, function and metabolic engineering of plant volatile organic compounds. New Phytol. 2013, 198, 16–32. [Google Scholar] [CrossRef]
- Muhlemann, J.K.; Klempien, A.; Dudareva, N. Floral volatiles: From biosynthesis to function. Plant Cell Environ. 2014, 37, 1936–1949. [Google Scholar] [CrossRef] [PubMed]
- Nagegowda, D.A.; Gutensohn, M.; Wilkerson, C.G.; Dudareva, N. Two nearly identical terpene synthases catalyze the formation of nerolidol and linalool in snapdragon flowers. Plant J. 2008, 55, 224–239. [Google Scholar] [CrossRef] [PubMed]
- Ginglinger, J.-F.; Boachon, B.; Höfer, R.; Paetz, C.; Köllner, T.G.; Miesch, L.; Lugan, R.; Baltenweck, R.; Mutterer, J.; Ullmann, P.; et al. Gene coexpression analysis reveals complex metabolism of the monoterpene alcohol linalool in Arabidopsis flowers. Plant Cell 2013, 25, 4640–4657. [Google Scholar] [CrossRef]
- Han, Y.; Wang, H.; Wang, X.; Li, K.; Dong, M.; Li, Y.; Zhu, Q.; Shang, F. Mechanism of floral scent production in Osmanthus fragrans and the production and regulation of its key floral constituents, β-ionone and linalool. Hortic. Res. 2019, 6. [Google Scholar] [CrossRef]
- Aros, D.; Gonzalez, V.; Allemann, R.K.; Müller, C.T.; Rosati, C.; Rogers, H.J. Volatile emissions of scented Alstroemeria genotypes are dominated by terpenes, and a myrcene synthase gene is highly expressed in scented Alstroemeria flowers. J. Exp. Bot. 2012, 63, 2739–2752. [Google Scholar] [CrossRef] [PubMed]
- Xiang, L.; Zhao, K.; Chen, L. Molecular cloning and expression of Chimonanthus praecox farnesyl pyrophosphate synthase gene and its possible involvement in the biosynthesis of floral volatile sesquiterpenoids. Plant Physiol. Biochem. 2010, 48, 845–850. [Google Scholar] [CrossRef]
- Roeder, S.; Hartmann, A.-M.; Effmert, U.; Piechulla, B. Regulation of simultaneous synthesis of floral scent terpenoids by the 1, 8-cineole synthase of Nicotiana suaveolens. Plant Mol. Biol. 2007, 65, 107–124. [Google Scholar] [CrossRef]
- Fan, Y.P.; Yu, R.C.; Huang, Y.; Chen, Y.F. Studies on the essential constituent of Hedychium flavum and H. coronarium. Acta Hortic. Sin. 2003, 30, 475. [Google Scholar]
- Fan, Y.; Wang, X.; Yu, R.; Yang, P. Analysis on the aroma components in several species of Hedychium. Acta Hortic. Sin. 2007, 34, 231. [Google Scholar]
- Shimada, T.; Endo, T.; Fujii, H.; Hara, M.; Omura, M. Isolation and characterization of (E)-beta-ocimene and 1, 8 cineole synthases in Citrus unshiu Marc. Plant Sci. 2005, 168, 987–995. [Google Scholar] [CrossRef]
- Tholl, D.; Chen, F.; Petri, J.; Gershenzon, J.; Pichersky, E. Two sesquiterpene synthases are responsible for the complex mixture of sesquiterpenes emitted from Arabidopsis flowers. Plant J. 2005, 42, 757–771. [Google Scholar] [CrossRef]
- Yue, Y.; Yu, R.; Fan, Y. Transcriptome profiling provides new insights into the formation of floral scent in Hedychium coronarium. BMC Genom. 2015, 16, 470. [Google Scholar] [CrossRef]
- Lucker, J.; Schwab, W.; Franssen, M.C.R.; Van Der Plas, L.H.W.; Bouwmeester, H.J.; Verhoeven, H.A.; Lücker, J.; Schwab, W.; Franssen, M.C.R.; Van Der Plas, L.H.W.; et al. Metabolic engineering of monoterpene biosynthesis: Two-step production of (+)-trans-isopiperitenol by tobacco. Plant J. 2004, 39, 135–145. [Google Scholar] [CrossRef]
- Green, S.A.; Chen, X.; Nieuwenhuizen, N.J.; Matich, A.J.; Wang, M.Y.; Bunn, B.J.; Yauk, Y.-K.K.; Atkinson, R.G. Identification, functional characterization, and regulation of the enzyme responsible for floral (E)-nerolidol biosynthesis in kiwifruit (Actinidia chinensis). J. Exp. Bot. 2012, 63, 1951–1967. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.; Kim, M.J.; Dhandapani, S.; Tjhang, J.G.; Yin, J.-L.; Wong, L.; Sarojam, R.; Chua, N.-H.; Jang, I.-C. The floral transcriptome of ylang ylang (Cananga odorata var. fruticosa) uncovers biosynthetic pathways for volatile organic compounds and a multifunctional and novel sesquiterpene synthase. J. Exp. Bot. 2015, 66, 3959–3975. [Google Scholar] [CrossRef] [PubMed]
- Jayaramaiah, R.H.; Anand, A.; Beedkar, S.D.; Dholakia, B.B.; Punekar, S.A.; Kalunke, R.M.; Gade, W.N.; Thulasiram, H.V.; Giri, A.P. Functional characterization and transient expression manipulation of a new sesquiterpene synthase involved in β-caryophyllene accumulation in Ocimum. Biochem. Biophys. Res. Commun. 2016, 473, 265–271. [Google Scholar] [CrossRef] [PubMed]
- Hong, G.-J.; Xue, X.-Y.; Mao, Y.-B.; Wang, L.-J.; Chen, X.-Y. Arabidopsis MYC2 interacts with DELLA proteins in regulating sesquiterpene synthase gene expression. Plant Cell 2012, 24, 2635–2648. [Google Scholar] [CrossRef] [PubMed]
- Yahyaa, M.; Tholl, D.; Cormier, G.; Jensen, R.; Simon, P.W.; Ibdah, M. Identification and characterization of terpene synthases potentially involved in the formation of volatile terpenes in carrot (Daucus carota L.) roots. J. Agric. Food Chem. 2015, 63, 4870–4878. [Google Scholar] [CrossRef] [PubMed]
- Wasternack, C.; Strnad, M. Jasmonates are signals in the biosynthesis of secondary metabolites—Pathways, transcription factors and applied aspects—A brief review. N. Biotechnol. 2017, 1–11. [Google Scholar] [CrossRef]
- Per, T.S.; Khan, M.I.R.; Anjum, N.A.; Masood, A.; Hussain, S.J.; Khan, N.A. Jasmonates in plants under abiotic stresses: Crosstalk with other phytohormones matters. Environ. Exp. Bot. 2018, 145, 104–120. [Google Scholar] [CrossRef]
- Wen, J.; Li, Y.; Qi, T.; Gao, H.; Liu, B.; Zhang, M.; Huang, H.; Song, S. The C-terminal domains of Arabidopsis GL3/EGL3/TT8 interact with JAZ proteins and mediate dimeric interactions. Plant Signal. Behav. 2018, 13, 1795–1814. [Google Scholar] [CrossRef] [PubMed]
- Miyamoto, K.; Kotake, T.; Boncela, A.J.; Saniewski, M.; Ueda, J. Hormonal regulation of gummosis and composition of gums from bulbs of hyacinth (Hyacinthus orientalis). J. Plant Physiol. 2015, 174, 1–4. [Google Scholar] [CrossRef]
- Kazan, K. Diverse roles of jasmonates and ethylene in abiotic stress tolerance. Trends Plant Sci. 2015, 20, 219–229. [Google Scholar] [CrossRef]
- Geetika, S.; Ruqia, M.; Poonam, S.; Harpreet, K.; Mudaser, A.M. Jasmonate Signaling and Stress Management in Plants. Mech. Plant Horm. Signal. Stress 2017, 143–171. [Google Scholar]
- Dar, T.A.; Uddin, M.; Khan, M.M.A.; Hakeem, K.R.; Jaleel, H. Jasmonates counter plant stress: A review. Environ. Exp. Bot. 2015, 115, 49–57. [Google Scholar] [CrossRef]
- Zhou, M.; Memelink, J. Jasmonate-responsive transcription factors regulating plant secondary metabolism. Biotechnol. Adv. 2016, 34, 441–449. [Google Scholar] [CrossRef]
- Zhang, X.; Luo, H.; Xu, Z.; Zhu, Y.; Ji, A.; Song, J.; Chen, S. Genome-wide characterisation and analysis of bHLH transcription factors related to tanshinone biosynthesis in Salvia miltiorrhiza. Sci. Rep. 2015, 5, 1–10. [Google Scholar] [CrossRef]
- Alquézar, B.; Volpe, H.X.L.; Magnani, R.F.; Miranda, M.P.; Santos, M.A.; Wulff, N.A.; Bento, J.M.S.; Parra, J.R.P.; Bouwmeester, H.; Peña, L. β-caryophyllene emitted from a transgenic Arabidopsis or chemical dispenser repels Diaphorina citri, vector of Candidatus liberibacters. Sci. Rep. 2017, 7, 5639. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Jia, M.; Huh, J.-H.; Muchlinski, A.; Peters, R.J.; Tholl, D. Identification of a dolabellane type diterpene synthase and other root-expressed diterpene synthases in Arabidopsis. Front. Plant Sci. 2016, 7, 1761. [Google Scholar] [CrossRef] [PubMed]
- Orlova, I.; Nagegowda, D.A.; Kish, C.M.; Gutensohn, M.; Maeda, H.; Varbanova, M.; Fridman, E.; Yamaguchi, S.; Hanada, A.; Kamiya, Y.; et al. The small subunit of snapdragon geranyl diphosphate synthase modifies the chain length specificity of tobacco geranylgeranyl diphosphate synthase in planta. Plant Cell 2009, 21, 4002–4017. [Google Scholar] [CrossRef] [PubMed]
- Fäldt, J.; Arimura, G.I.; Gershenzon, J.; Takabayashi, J.; Bohlmann, J. Functional identification of AtTPS03 as (E)-beta-ocimene synthase: A monoterpene synthase catalyzing jasmonate- and wound-induced volatile formation in Arabidopsis thaliana. Planta 2003, 216, 745–751. [Google Scholar] [CrossRef]
- Cheng, Z.; Sun, L.; Qi, T.; Zhang, B.; Peng, W.; Liu, Y.; Xie, D. The bHLH transcription factor MYC3 interacts with the jasmonate ZIM-domain proteins to mediate jasmonate response in arabidopsis. Mol. Plant 2011, 4, 279–288. [Google Scholar] [CrossRef]
- Yang, J.; Dai, P.; Zhou, T.; Huang, Z.; Feng, L.; Su, H.; Liu, Z.; Zhao, G. Genetic diversity and structure of wintersweet (Chimonanthus praecox) revealed by EST-SSR markers. Sci. Hortic. (Amsterdam) 2013, 150, 1–10. [Google Scholar] [CrossRef]
- Liu, D.; Sui, S.; Ma, J.; Li, Z.; Guo, Y.; Luo, D.; Yang, J.; Li, M. Transcriptomic analysis of flower development in Wintersweet (Chimonanthus praecox). PLoS ONE 2014, 9. [Google Scholar] [CrossRef]
- Zhang, S.; Zhang, H.; Chen, L.; Xia, W.; Zhang, W. Phytochemical profiles and antioxidant activities of different varieties of Chimonanthus Praecox. Ind. Crops Prod. 2016, 85, 11–21. [Google Scholar] [CrossRef]
- Zhang, S.H.; Wei, Y.; Liu, J.L.; Yu, H.M.; Yin, J.H.; Pan, H.Y.; Baldwin, T.C. An apoplastic chitinase CpCHT1 isolated from the corolla of wintersweet exhibits both antifreeze and antifungal activities. Biol. Plant. 2011, 55, 141–148. [Google Scholar] [CrossRef]
- Dai, P.F.; Yang, J.; Zhou, T.H.; Huang, Z.H.; Feng, L.; Su, H.L.; Liu, Z.L.; Zhao, G.F. Genetic diversity and differentiation in Chimonanthus praecox and Ch. salicifolius (Calycanthaceae) as revealed by inter-simple sequence repeat (ISSR) markers. Biochem. Syst. Ecol. 2012, 44, 149–156. [Google Scholar] [CrossRef]
- Sui, S.; Luo, J.; Ma, J.; Zhu, Q.; Lei, X.; Li, M. Generation and analysis of expressed sequence tags from Chimonanthus praecox (Wintersweet) flowers for discovering stress-responsive and floral development-related genes. Comp. Funct. Genom. 2012, 2012, 13. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Chen, Z. Polymer Monolith Microextraction Coupled with HPLC for Determination of Jasmonates in Wintersweet Flowers. Anal. Lett. 2012, 46, 74–86. [Google Scholar] [CrossRef]
- Zeng, X.; Liu, C.; Zheng, R.; Cai, X.; Luo, J.; Zou, J.; Wang, C. Emission and Accumulation of Monoterpene and the Key Terpene Synthase (TPS) Associated with Monoterpene Biosynthesis in Osmanthus fragrans Lour. Front. Plant Sci. 2016, 6, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Park, J.; Sun, B.; Ahn, H.; Wang, G. Wintersweet-flower-like CoFe2O4/MWCNTs hybrid material for high-capacity reversible lithium storage. Chem. An Asian J. 2012, 7, 1940–1946. [Google Scholar] [CrossRef]
- Tian, J.-P.; Ma, Z.-Y.; Zhao, K.-G.; Zhang, J.; Xiang, L.; Chen, L.-Q. Transcriptomic and proteomic approaches to explore the differences in monoterpene and benzenoid biosynthesis between scented and unscented genotypes of wintersweet. Physiol. Plant. 2018. [Google Scholar] [CrossRef]
- Tholl, D.; Boland, W.; Hansel, A.; Loreto, F.; Röse, U.S.R.; Schnitzler, J.P. Practical approaches to plant volatile analysis. Plant J. 2006, 45, 540–560. [Google Scholar] [CrossRef] [PubMed]
- Zhao, K.G.; Zhou, M.Q.; Chen, L.Q.; Zhang, D.; Robert, G.W. Genetic diversity and discrimination of Chimonanthus praecox (L.) link germplasm using ISSR and RAPD markers. HortScience 2007, 42, 1144–1148. [Google Scholar] [CrossRef]
- Zhang, F.; Yao, J.; Ke, J.; Zhang, L.; Lam, V.Q.; Xin, X.F.; Zhou, X.E.; Chen, J.; Brunzelle, J.; Griffin, P.R.; et al. Structural basis of JAZ repression of MYC transcription factors in jasmonate signalling. Nature 2015, 525, 269–273. [Google Scholar] [CrossRef]
- Chen, F.; Tholl, D.; D’Auria, J.C.; Farooq, A.; Pichersky, E.; Gershenzon, J. Biosynthesis and Emission of Terpenoid Volatiles from Arabidopsis Flowers. Plant Cell 2003, 15, 481–494. [Google Scholar] [CrossRef]
- Nakata, M.; Mitsuda, N.; Herde, M.; Koo, A.J.K.K.; Moreno, J.E.; Suzuki, K.; Howe, G.A.; Ohme-takagi, M. A bHLH-type transcription factor, ABA-INDUCIBLE BHLH-TYPE TRANSCRIPTION FACTOR/JA-ASSOCIATED MYC2-LIKE1, acts as a repressor to negatively regulate jasmonate signaling in arabidopsis. Plant Cell 2013, 25, 1641–1656. [Google Scholar] [CrossRef]
- Sasaki-Sekimoto, Y.; Jikumaru, Y.; Obayashi, T.; Saito, H.; Masuda, S.; Kamiya, Y.; Ohta, H.; Shirasu, K. Basic helix-loop-helix transcription factors JASMONATE-ASSOCIATED MYC2-LIKE1 (JAM1), JAM2, and JAM3 are negative regulators of jasmonate responses in Arabidopsis. Plant Physiol. 2013, 163, 291–304. [Google Scholar] [CrossRef] [PubMed]
- Song, S.; Huang, H.; Gao, H.; Wang, J.; Wu, D.; Liu, X.; Yang, S.; Zhai, Q.; Li, C.; Qi, T.; et al. Interaction between MYC2 and ETHYLENE INSENSITIVE3 modulates antagonism between jasmonate and ethylene signaling in Arabidopsis. Plant Cell 2014, 26, 263–279. [Google Scholar] [CrossRef] [PubMed]
- Yue, Y.; Yin, C.; Guo, R.; Peng, H.; Yang, Z. An anther-specific gene PhGRP is regulated by PhMYC2 and causes male sterility when overexpressed in petunia anthers. Plant Cell Rep. 2017. [Google Scholar] [CrossRef] [PubMed]
- Schweizer, F.; Colinas, M.; Pollier, J.; Van Moerkercke, A.; Bossche, R.V.; de Clercq, R.; Goossens, A. An engineered combinatorial module of transcription factors boosts production of monoterpenoid indole alkaloids in Catharanthus roseus. Metab. Eng. 2018. [Google Scholar] [CrossRef] [PubMed]
- Zvi, M.M.B.; Shklarman, E.; Masci, T.; Kalev, H.; Debener, T.; Shafir, S.; Ovadis, M.; Vainstein, A. PAP1 transcription factor enhances production of phenylpropanoid and terpenoid scent compounds in rose flowers. New Phytol. 2012, 195, 335–345. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.-H.; Wang, J.-Y.J.-W.; Wang, S.; Wang, J.-Y.J.-W.; Chen, X.-Y. Characterization of GaWRKY1, a cotton transcription factor that regulates the sesquiterpene synthase gene (+)-δ-cadinene synthase-A. Plant Physiol. 2004, 135, 507–515. [Google Scholar] [CrossRef]
- Van Der Fits, L.; Memelink, J. ORCA3, a jasmonate-responsive transcriptional regulator of plant primary and secondary metabolism. Science 2000, 289, 295–297. [Google Scholar] [CrossRef]
- Yu, Z.X.; Li, J.X.; Yang, C.Q.; Hu, W.L.; Wang, L.J.; Chen, X.Y. The jasmonate-responsive AP2/ERF transcription factors AaERF1 and AaERF2 positively regulate artemisinin biosynthesis in Artemisia annua L. Mol. Plant 2012, 5, 353–365. [Google Scholar] [CrossRef]
- Song, S.; Qi, T.; Wasternack, C.; Xie, D. Jasmonate signaling and crosstalk with gibberellin and ethylene. Curr. Opin. Plant Biol. 2014, 21, 112–119. [Google Scholar] [CrossRef]
- Schuman, M.C.; Palmer-Young, E.C.; Schmidt, A.; Gershenzon, J.; Baldwin, I.T. Ectopic Terpene Synthase Expression Enhances Sesquiterpene Emission in Nicotiana attenuata without Altering Defense or Development of Transgenic Plants or Neighbors. Plant Physiol. 2014, 166, 779–797. [Google Scholar] [CrossRef] [PubMed]
- Song, S.; Qi, T.; Fan, M.; Zhang, X.; Gao, H.; Huang, H.; Wu, D.; Guo, H.; Xie, D. The bHLH subgroup IIId factors negatively regulate jasmonate-mediated plant defense and development. PLoS Genet. 2013, 9, e1003653. [Google Scholar] [CrossRef] [PubMed]
- Nakata, M.; Ohme-Takagi, M. Two bHLH-type transcription factors, JA-ASSOCIATED MYC2-LIKE2 and JAM3, are transcriptional repressors and affect male fertility. Plant Signal. Behav. 2013, 8, e26473. [Google Scholar] [CrossRef] [PubMed]
- Sandra, F.; Patricia, F.-C.; Guillermo, M.F.; Monica, D.-D.; Selena, G.-I.; Irene, L.-V.; Marta, G.; Gemma, F.-B.; Jelle, V.L.; Geert, D.J.; et al. bHLH003, bHLH013 and bHLH017 are new targets of JAZ repressors negatively regulating JA responses. PLoS One 2014, 9, e86182. [Google Scholar] [CrossRef]
- Huang, H.; Gao, H.; Liu, B.; Fan, M.; Wang, J.; Wang, C.; Tian, H.; Wang, L.; Xie, C.; Wu, D.; et al. bHLH13 Regulates Jasmonate-Mediated Defense Responses and Growth. Evol. Bioinform. 2018, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Hall, T.; Biosciences, I.; Carlsbad, C. BioEdit: An important software for molecular biology. GERF Bull. Biosci. 2011, 2, 60–61. [Google Scholar]
- Li, Y.X.; Chau, K.T.; Wei, K.T.; Kang, Q. Bin A comparative study on two XML editors (oxygon and ultraedit). In Proceedings of the 3rd International Conference on Cryptography, Security and Privacy, Kuala Lumpur, Malaysia, January 2019; pp. 284–288. [Google Scholar]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; Mcgettigan, P.A.; Mcwilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Alonso, J.M.; Stepanova, A.N.; Leisse, T.J.; Kim, C.J.; Chen, H.; Shinn, P.; Stevenson, D.K.; Zimmerman, J.; Barajas, P.; Cheuk, R.; et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 2003, 301, 653–657. [Google Scholar] [CrossRef]
- Maliga, P.; BREZNOVITS, Á.S.; Marton, L. Streptomycin-resistant plants from callus culture of haploid tobacco. Nat. New Biol. 1973, 244, 29. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Yang, N.; Zhao, K.; Li, X.; Zhao, R.; z Aslam, M.; Yu, L.; Chen, L. Comprehensive analysis of wintersweet flower reveals key structural genes involved in flavonoid biosynthetic pathway. Gene 2018, 676, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Li, Z.; Wang, B.; Sui, S.; Li, M. Cloning of an expansin gene from Chimonanthus praecox flowers and its expression in flowers treated with ethephon or 1-methylcyclopropene. HortScience 2012, 47, 1472–1477. [Google Scholar] [CrossRef]
- Guo, Y.; Li, R.; Asgher, Z.; Shamsi, I.H.; Yu, S.; Zhao, J.; Ren, X. Genome-Wide Selection and Testing of Superior Reference Genes for Quantitative Gene Expression Normalization in Tobacco. Res. Rev. J. Microbiol. Biotechnol. 2016, 5, 68–75. [Google Scholar]
- Wang, H.; Wang, J.; Jiang, J.; Chen, S.; Guan, Z.; Liao, Y.; Chen, F. Reference genes for normalizing transcription in diploid and tetraploid Arabidopsis. Sci. Rep. 2014, 4, 6781. [Google Scholar] [CrossRef] [PubMed]
- Zmienko, A.; Samelak-Czajka, A.; Goralski, M.; Sobieszczuk-Nowicka, E.; Kozlowski, P.; Figlerowicz, M. Selection of Reference Genes for qPCR- And ddPCR-Based Analyses of Gene Expression in Senescing Barley Leaves. PLoS ONE 2015, 10, 1–16. [Google Scholar] [CrossRef]
- Kozera, B.; Rapacz, M. Reference genes in real-time PCR. J. Appl. Genet. 2013, 54, 391–406. [Google Scholar] [CrossRef]
- HOrsch, R.B.; Fry, J.E.; Hoffmann, N.L.; Eichholtz, D.; Rogers, S.G.; Fraley, R.T. A Simple and General Method for hybridization revealed the expected. Science 1985, 227, 1229–1231. [Google Scholar] [CrossRef]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method forAgrobacterium-mediated transformation ofArabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef]
- Donath, J.; Boland, W. Biosynthesis of acyclic homoterpenes: Enzyme selectivity and absolute configuration of the nerolidol precursor. Phytochemistry 1995, 39, 785–790. [Google Scholar] [CrossRef]
- Ai, T.N.; Naing, A.H.; Arun, M.; Lim, S.H.; Kim, C.K. Sucrose-induced anthocyanin accumulation in vegetative tissue of Petunia plants requires anthocyanin regulatory transcription factors. Plant Sci. 2016, 252, 144–150. [Google Scholar] [CrossRef] [PubMed]
- Chu, H.; Jeong, J.C.; Kim, W.J.; Chung, D.M.; Jeon, H.K.; Ahn, Y.O.; Kim, S.H.; Lee, H.S.; Kwak, S.S.; Kim, C.Y. Expression of the sweetpotato R2R3-type IbMYB1a gene induces anthocyanin accumulation in Arabidopsis. Physiol. Plant. 2013, 148, 189–199. [Google Scholar] [CrossRef] [PubMed]



| Plant Name | Gene | Identity % | Gene Function |
|---|---|---|---|
| Arabidopsis | AtMYC2 (At1g32640) | 54.01 | Activate terpene synthesase |
| Tobacco | Transcription factor MYC2-like | 57.62 | Activation of JA responses |
| Tomato | SlMYC2 | 57.56 | Methyl jasmonate inducer |
| Grape vine | Predicted: transcription factor MYC2 | 62.01 | Unknown (Genome data) * |
| Lindera | Transcription factor MYC | 62.78 | Accumulation mechanism of β-ocimene |
| Hayata | Transcription factor MYC2 | 65.13 | Unknown (Genome data) |
| Lotus | Transcription factor MYC2-like | 62.01 | Unknown (Genome data) |
| Plant Name | Gene | Identity % | Gene Function |
|---|---|---|---|
| Arabidopsis | Transcription factor bHLH13/JAM2 (At1g01260) | 42.06 | Interacts with JAZ proteins to negatively regulate jasmonate responses |
| Arabidopsis | Transcription factor bHLH17/JAM1 (At2g46510) | 44.78 | Interacts with JAZ proteins to negatively regulate jasmonate responses |
| Tobacco | Predicted: transcription factor bHLH13-like | 60.26 | Unknown (Genome data) * |
| Tomato | Transcription factor bHLH13-like | 59.51 | Unknown (Genome data) |
| Grape vine | Transcription factor bHLH13 | 62.90 | Unknown (Genome data) |
| Hayata | Transcription factor bHLH13 | 69.77 | Unknown (Genome data) |
| Lotus | Predicted: transcription factor bHLH13-like | 62.46 | Unknown (Genome data) |
| Primer Name | Primer Sequence (5′–3′) | TM C⁰ |
|---|---|---|
| Salk_083483 LP | TGGTTTTTCTTGGTTTCGATG | 60 |
| Salk_083483 RP | CTCTAATCATTGCGTCCCAAC | |
| LBb1.3 | ATTTTGCCGATTTCGGAAC | 60 |
| Primer Name | Primer Sequence (5′–3′) | TM C⁰ |
|---|---|---|
| CpMYC2q-F | TCCAGTCCAACAAGAAGAACCACCC | 63.5 |
| CpMYC2q-R | CTATCTGTTGCCAATTTTCTCCGCC | |
| CpbHLH13q-F | CCTTTGGACACCCACCCG | 60.5 |
| CpbHLH13q-R | CTATAATGAGTTTGATTCACGACTAAATGC | |
| RPL8-F | ACATTGCCGACCATGAGATTG | 59 |
| RPL8-R | CACTTGCCCGAGTTACCTTT | |
| CpTublin-F | GTGCATCTCTATCCACATCG | 60 |
| CpTublin-R | CAAGCTTCCTTATGCGATCC | |
| AtCACS-F | TCGATTGCTTGGTTTGGAAGAT | 60 |
| AtCACS-R | GCACTTAGCGTGGACTCTGTTTGATC | |
| AtTubulin-F | TCAAGAGGTTCTCAGCAGTA | 59 |
| AtTubulin-R | TCACCTTCTTCATCCGCAGTT | |
| NtNADH-F | TTGGTGGATCTGACCTAGTG | 60 |
| NtNADH-R | ATGGTGTGAAAGAGCGTTCG | |
| NtCPN60-2-F | ATGGCACTCTTGATGGGTTC | 60 |
| NtCPN60-2-R | AGCACTAGGCATTGCCATTG | |
| 35S-F | ACGCACAATCCCACTATCCT | 59 |
| 35S-R | TGCTCAACACATGAGCGAAAC | |
| At1g61680-F | ATGATCGATGTCATTCAAAGT | 52 |
| At1g61680-R | TTAAATGTTTGAGACATTTCTC | |
| At3g25810-F | TATATTTGATGTAATCATCG | 53 |
| At3g25810-R | TTGAACCATAGCAGTGAAGAG | |
| At5g23960-F | GGAACTGAGACGTTCAAAGAG | 57 |
| At5g23960-R | CGCTGTGAATAAGATTAGTGC | |
| At5g44630-F | TGGAGGAAAATATAGTGATAT | 56 |
| At5g44630-R | CGGTGCTGAGGTATGTGAAGA |
| Primer Name | Primer Sequence (5′–3′) | TM C⁰ |
|---|---|---|
| CpMYC2-F | CGGGGTACCATGAATCTCTGGTCCGACGA | 63.5 |
| CpMYC2-R | CTAGTCTAGACTATCTGTTGCCAATTTTCTCC | |
| CpbHLH13-F | CGGGGTACCATGAAATCGGAGATTGGAA | 60.5 |
| CpbHLH13-R | CTAGTCTAGACTATAATGAGTTTGATTCACGA |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aslam, M.Z.; Lin, X.; Li, X.; Yang, N.; Chen, L. Molecular Cloning and Functional Characterization of CpMYC2 and CpBHLH13 Transcription Factors from Wintersweet (Chimonanthus praecox L.). Plants 2020, 9, 785. https://doi.org/10.3390/plants9060785
Aslam MZ, Lin X, Li X, Yang N, Chen L. Molecular Cloning and Functional Characterization of CpMYC2 and CpBHLH13 Transcription Factors from Wintersweet (Chimonanthus praecox L.). Plants. 2020; 9(6):785. https://doi.org/10.3390/plants9060785
Chicago/Turabian StyleAslam, Muhammad Zeshan, Xiang Lin, Xiang Li, Nan Yang, and Longqing Chen. 2020. "Molecular Cloning and Functional Characterization of CpMYC2 and CpBHLH13 Transcription Factors from Wintersweet (Chimonanthus praecox L.)" Plants 9, no. 6: 785. https://doi.org/10.3390/plants9060785
APA StyleAslam, M. Z., Lin, X., Li, X., Yang, N., & Chen, L. (2020). Molecular Cloning and Functional Characterization of CpMYC2 and CpBHLH13 Transcription Factors from Wintersweet (Chimonanthus praecox L.). Plants, 9(6), 785. https://doi.org/10.3390/plants9060785




