Agrobacterium-Mediated Transformation of Chrysanthemum with Artemisinin Biosynthesis Pathway Genes
Abstract
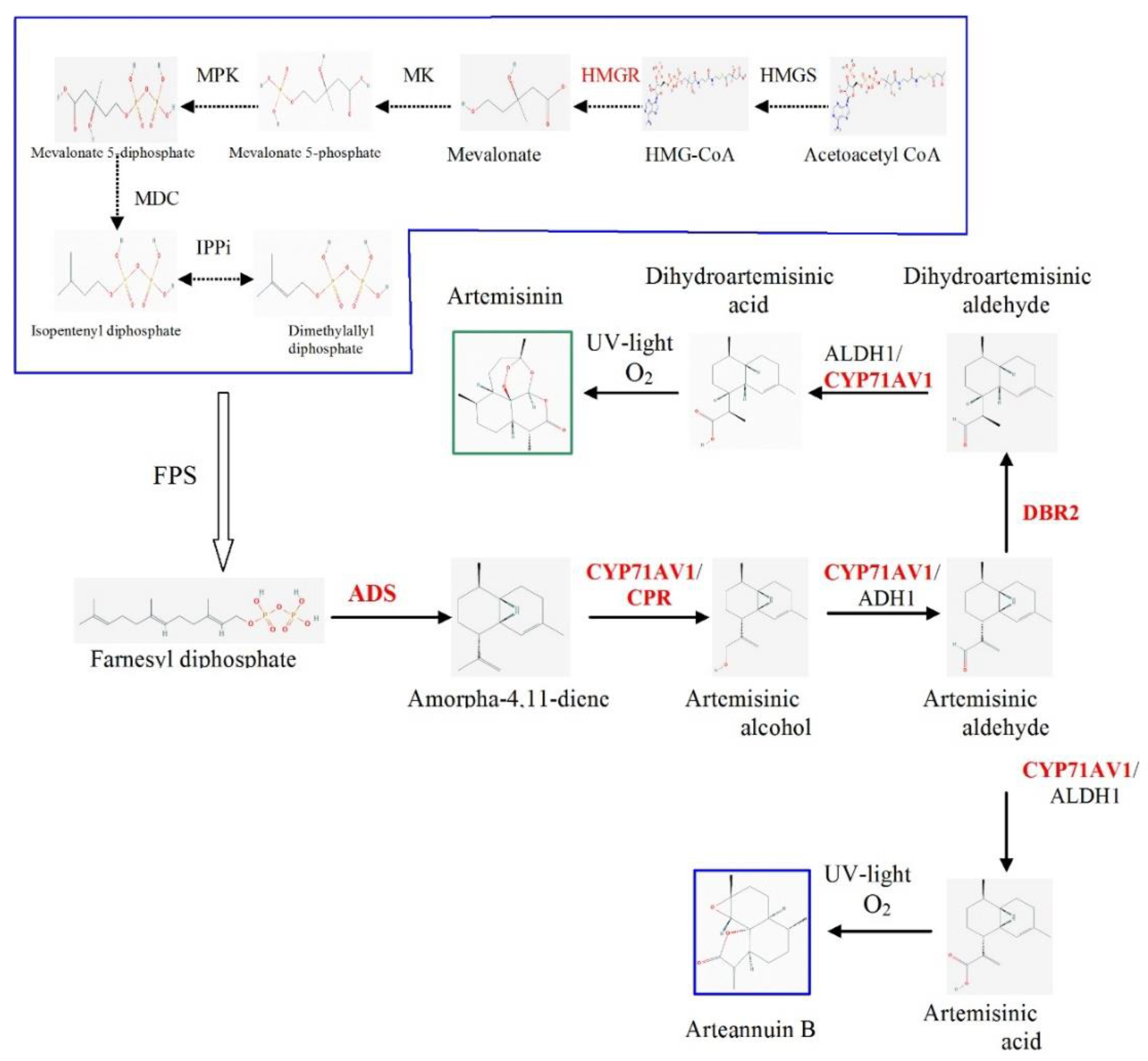
1. Introduction
2. Results and Discussion
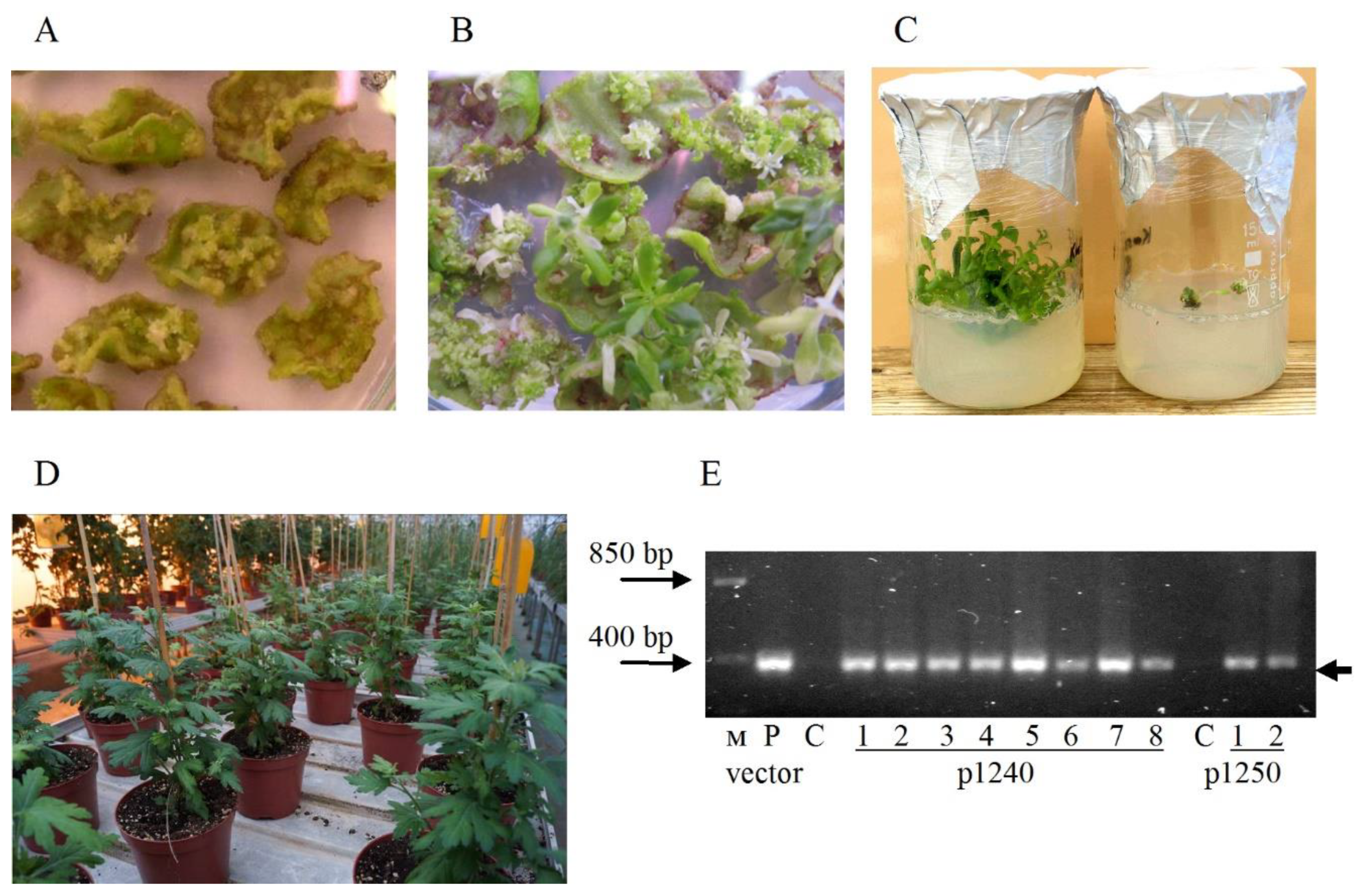
2.1. Agrobacterium-Mediated Transformation of Chrysanthemum Plants with p1240 and p1250 Vectors
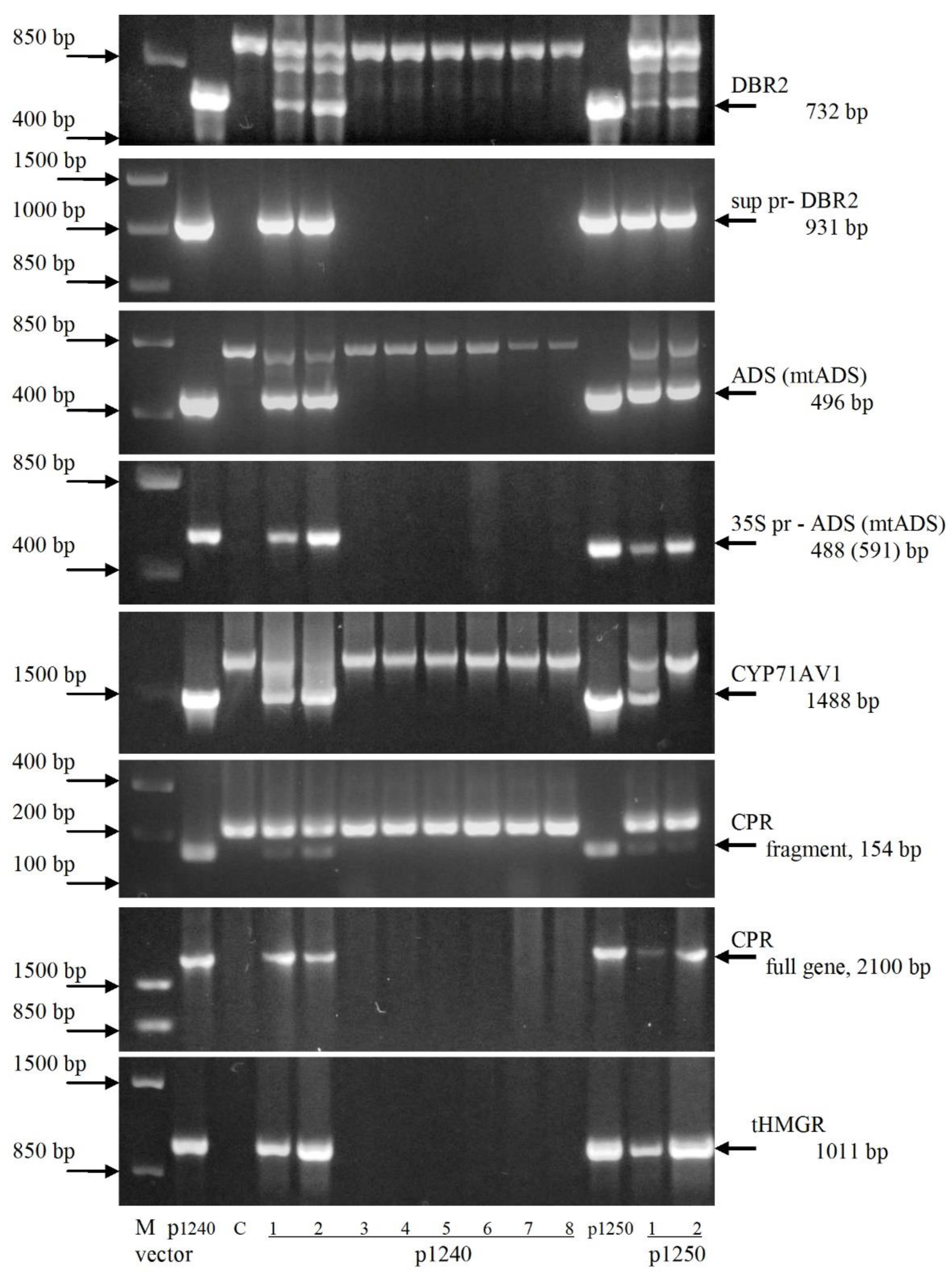
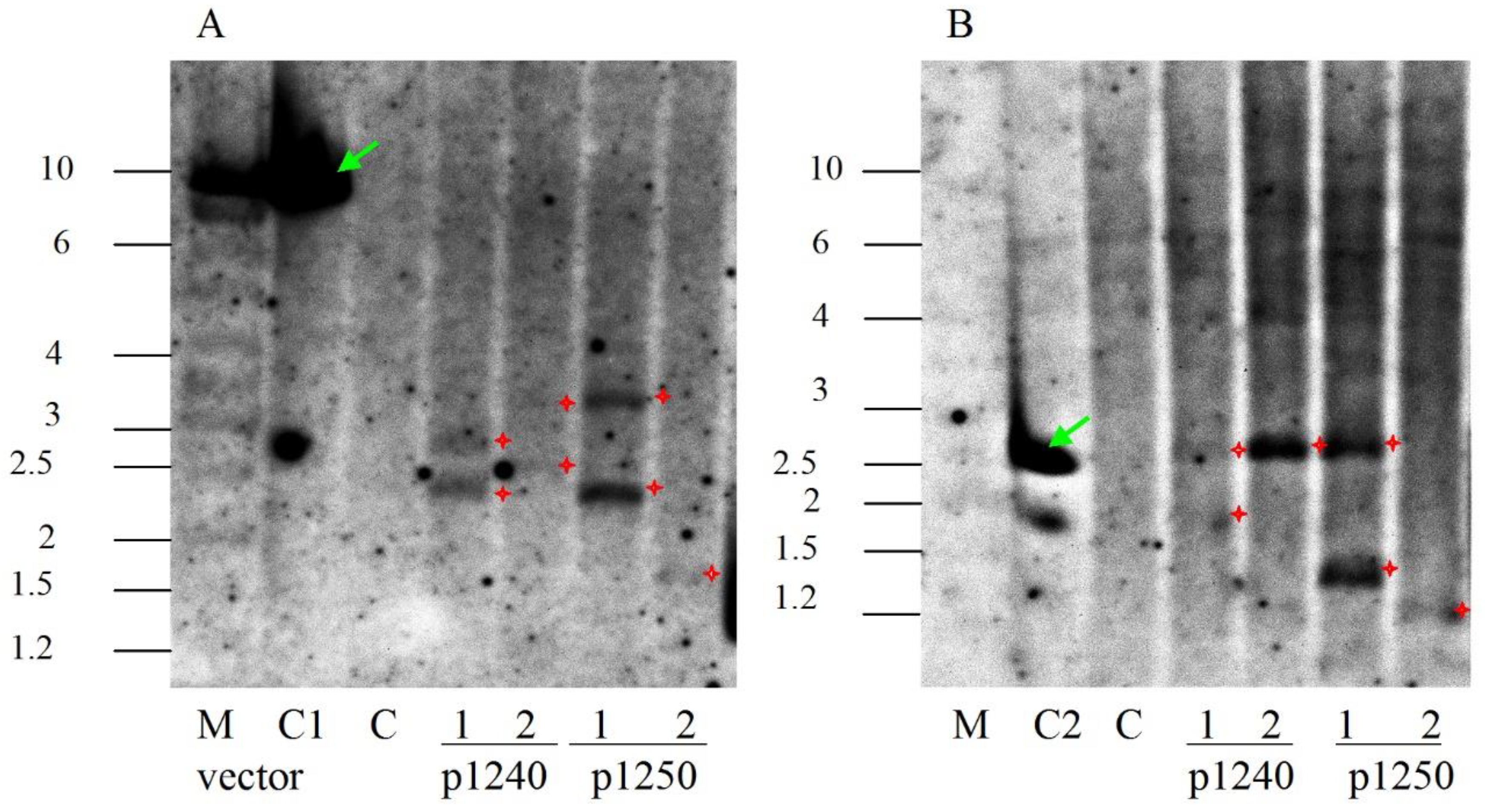
2.2. Analysis of the Integration of Artemisinin Biosynthetic Pathway Genes in Transformed Plants
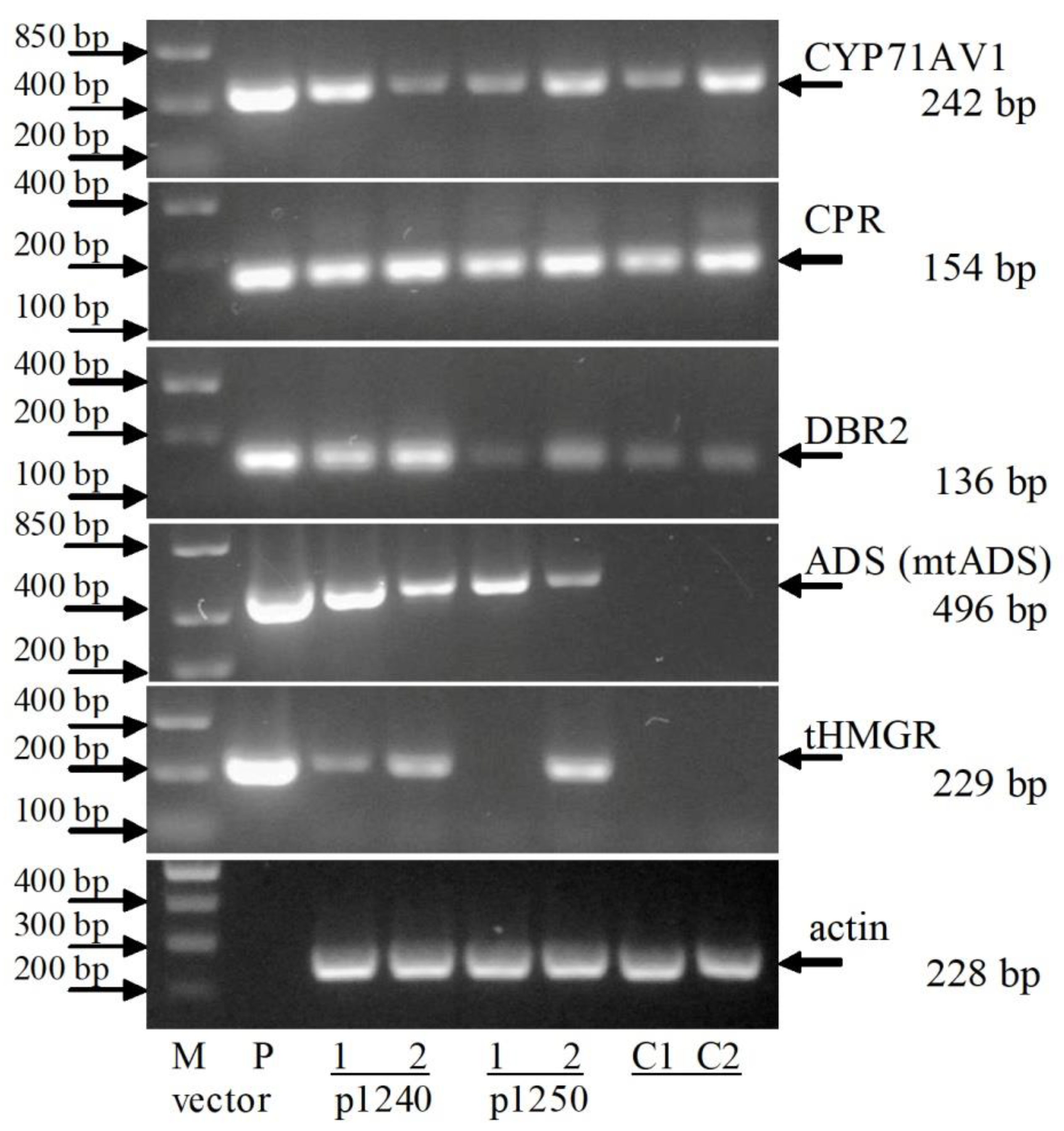
2.3. RT-PCR Analysis of Artemisinin Biosynthesis Genes Expression
2.4. Artemisinin Assay in the Transgenic Plants
3. Materials and Methods
3.1. The Plant Transformation Vectors
3.2. Agrobacterial Transformation of Chrysanthemum
3.3. PCR and Southern Blot Analysis
3.4. Reverse Transcription-Polymerase Chain Reaction Analysis
3.5. Artemisinin Assay in the Transgenic Plants
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ho, W.; Peh, H.; Chan, T.; Wong, W.S. Artemisinins: Pharmacological actions beyond anti-malarial. Pharm. Ther. 2014, 142, 126–139. [Google Scholar] [CrossRef] [PubMed]
- Johnston, G. Artemisia annua, Artemisinin, ACTs & Malaria Control in Africa: Tradition, Science and Public Policy by Dana G. Dalrymple. Q. Rev. Biol. 2013, 88, 257–258. [Google Scholar]
- Brisibe, E.A.; Chukwurah, P.N. Production of artemisinin in planta and in microbial systems need not be mutually exclusive. In Artemisia annua-Pharmacology and Biotechnology; Aftab, T., Ferreira, J.F.S., Eds.; Springer: Berlin/Heidelberg, Germany, 2014; pp. 269–292. [Google Scholar]
- Kumar, S.; Gupta, S.; Singh, P.; Bajpai, P.; Gupta, M.; Singh, D.; Gupta, A.; Ram, G.; Shasany, A.; Sharma, S. High yields of artemisinin by multi-harvest of Artemisia annua crops. Ind. Crops Prod. 2004, 19, 77–90. [Google Scholar] [CrossRef]
- Tu, Y. Metabolic Engineering of Artemisia annua L. In From Artemisia annua L. to Artemisinins. The Discovery and Development of Artemisinins and Antimalarial Agents; Chemical Industry Press: Beijing, China, 2017; Chapter 8; pp. 163–208. [Google Scholar]
- Brown, G.D. The biosynthesis of artemisinin (Qinghaosu) and the phytochemistry of Artemisia annua L. (Qinghao). Molecules 2010, 15, 7603–7698. [Google Scholar] [CrossRef] [PubMed]
- Ikram, N.K.B.; Beyraghdar Kashkooli, A.; Peramuna, A.V.; van der Krol, A.R.; Bouwmeester, H.; Simonsen, H.T. Stable production of the antimalarial drug artemisinin in the moss Physcomitrella patens. Front. Bioeng. Biotechnol. 2017, 5, 47. [Google Scholar] [CrossRef] [PubMed]
- Farhi, M.; Marhevka, E.; Ben-Ari, J.; Algamas-Dimantov, A.; Liang, Z.; Zeevi, V.; Edelbaum, O.; Spitzer-Rimon, B.; Abeliovich, H.; Schwartz, B.; et al. Generation of the potent anti-malarial drug artemisinin in tobacco. Nat Biotechnol. 2011, 29, 1072–1074. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Kashkooli, A.B.; Sallets, A.; Ting, H.-M.; De Ruijter, N.C.; Olofsson, L.; Brodelius, P.; Pottier, M.; Boutry, M.; Bouwmeester, H.; et al. Transient production of artemisinin in Nicotiana benthamiana is boosted by a specific lipid transfer protein from A. annua. Metab. Eng. 2016, 38, 159–169. [Google Scholar] [CrossRef] [PubMed]
- Malhotra, K.; Subramaniyan, M.; Rawat, K.; Kalamuddin, M.; Qureshi, M.I.; Malhotra, P.; Mohmmed, A.; Cornish, K.; Daniell, H.; Kumar, S. Compartmentalized metabolic engineering for artemisinin biosynthesis and effective malaria treatment by oral delivery of plant. Cells. Mol. Plant 2016, 9, 1464–1477. [Google Scholar] [CrossRef]
- Wen, W.; Yu, R. Artemisinin biosynthesis and its regulatory enzymes: Progress and perspective. Pharmacogn. Rev. 2011, 5, 189–194. [Google Scholar]
- Bruno, M.; Bancheva, S.; Rosselli., S.; Maggio, A. Sesquiterpenoids in subtribe Centaureinae (Cass.) Dumort (tribe Cardueae, Asteraceae): Distribution, (13)C NMR spectral data and biological properties. Phytochemistry 2013, 95, 19–93. [Google Scholar]
- Wang, Y.; Li, X.; Jiang, Q.; Sun, H.; Jiang, J.; Chen, S.; Guan, Z.; Fang, W.; Chen, F. GC-MS Analysis of the Volatile Constituents in the Leaves of 14 Compositae Plants. Molecules 2018, 23, 166. [Google Scholar] [CrossRef]
- Xue, G.-M.; Li, X.-Q.; Chen, C.; Chen, K.; Wang, X.-B.; Gu, Y.-C.; Luo, J.-G.; Kong, L.-Y. Highly Oxidized Guaianolide Sesquiterpenoids with Potential Anti-inflammatory Activity from Chrysanthemum indicum. J. Nat. Prod. 2018, 81, 378–386. [Google Scholar] [CrossRef] [PubMed]
- Chang, K.M.; Kim, G.H. Volatile aroma constituents of gukhwa (Chrysanthemum morifolium R.). Food Sci Biotechnol. 2013, 22, 659–663. [Google Scholar] [CrossRef]
- Yang, L.; Nuerbiye, A.; Cheng, P.; Wang, J.H.; Li, H. Analysis of Floral Volatile Components and Antioxidant Activity of Different Varieties of Chrysanthemum morifolium. Molecules 2017, 22, E1790. [Google Scholar] [CrossRef] [PubMed]
- Spaargaren, J.; van Geest, G. Chrysanthemum. In Ornamental Crops; Van Huylenbroeck, J., Ed.; Springer International Publishing: Cham, Switzerland, 2018; pp. 319–348. [Google Scholar]
- Won, S.Y.; Kwon, S.-J.; Lee, T.-H.; Jung, J.-A.; Kim, J.S.; Kang, S.-H.; Sohn, S.-H. Comparative transcriptome analysis reveals whole-genome duplications and gene selection patterns in cultivated and wild Chrysanthemum species. Plant Mol. Biol. 2017, 95, 451–461. [Google Scholar] [CrossRef]
- Hirakawa, H.; Sumitomo, K.; Hisamatsu, T.; Nagano, S.; Shirasawa, K.; Higuchi, Y.; Kusaba, M.; Koshioka, M.; Nakano, Y.; Yagi, M.; et al. De novo whole-genome assembly in Chrysanthemum seticuspe, a model species of Chrysanthemums, and its application to genetic and gene discovery analysis. Dna Res. 2019, 26, 195–203. [Google Scholar] [CrossRef]
- Van Herpen, T.W.J.M.; Cankar, K.; Nogueira, M.; Bosch, D.; Bouwmeester, H.J.; Beekwilder, J. Nicotiana benthamiana as a Production Platform for Artemisinin Precursors. PLoS ONE 2010, 5, e14222. [Google Scholar] [CrossRef]
- Farhi, M.; Marhevka, E.; Masci, T.; Marcos, E.; Eyal, Y.; Ovadis, M.; Abeliovich, H.; Vainstein, A. Harnessing yeast subcellular compartments for the production of plant terpenoids. Metab. Eng. 2011, 13, 474–481. [Google Scholar] [CrossRef]
- Farhi, M.; Kozin, M.; Duchin, S.; Vainstein, A. Metabolic engineering of plants for artemisinin synthesis. Biotechnol. Genet. Eng. Rev. 2013, 29, 135–148. [Google Scholar] [CrossRef]
- Shinoyama, H.; Mitsuhara, I.; Ichikawa, H.; Kato, K.; Mochizuki, A. Transgenic chrysanthemums (Chrysanthemum morifolium Ramat.) carrying both insect and disease resistance. Acta Hortic. 2015, 1087, 485–497. [Google Scholar] [CrossRef]
- Naing, A.H.; Ai, T.N.; Jeon, S.M.; Lim, S.H.; Kim, C.K. An efficient protocol for Agrobacterium-mediated genetic transformation of recalcitrant chrysanthemum cultivar Shinma. Acta Physiol. Plant. 2016, 38, 38. [Google Scholar] [CrossRef]
- Mitiouchkina, T.; Firsov, A.; Titova, S.; Shulga, O.; Dolgov, S. Efficiency assessment of genetic designs with coat protein in transgene-mediated resistance against Chrysanthemum virus B. Acta Hortic. 2018, 1193, 89–94. [Google Scholar] [CrossRef]
- Collier, R.; Thomson, J.G.; Thilmony, R. A versatile and robust Agrobacterium-based gene stacking system generates high-quality transgenic Arabidopsis plants. Plant J. 2018, 95, 573–583. [Google Scholar] [CrossRef] [PubMed]
- Untergasser, A.; Bijl, G.J.; Liu, W.; Bisseling, T.; Schaart, J.G.; Geurts, R. One-step Agrobacterium mediated transformation of eight genes essential for rhizobium symbiotic signaling using the novel binary vector system pHUGE. PLoS ONE 2012, 7, e47885. [Google Scholar] [CrossRef]
- Fujisawa, M.; Takita, E.; Harada, H.; Sakurai, N.; Suzuki, H.; Ohyama, K.; Shibata, D.; Misawa, N. Pathway engineering of Brassica napus seeds using multiple key enzyme genes involved in ketocarotenoid formation. J. Exp. Bot. 2009, 60, 1319–1332. [Google Scholar] [CrossRef] [PubMed]
- Ward, J.; Xu, J.; Lamb, J.; Gilbertson, L. The use of Agrobacterium tumefaciens to deliver ten genes on a single vector to corn. In Proceedings of the 9th International Plant Molecular Biology (IPMB) Congress 2009, Saint Louis, MO, USA, 25–30 October 2009. [Google Scholar]
- Chung, S.M.; Frankman, E.L.; Tzfira, T. A versatile vector system for multiple gene expression in plants. Trends Plant Sci. 2005, 10, 357–361. [Google Scholar] [CrossRef] [PubMed]
- Shulga, O.A.; Mitiouchkina, T.Y.; Shchennikova, A.V.; Skryabin, K.G.; Dolgov, S.V. Overexpression of AP1-like genes from Asteraceae induces early-flowering in transgenic Chrysanthemum plants. In Vitro Cell. Dev. Biol. Plant. 2011, 47, 553–560. [Google Scholar] [CrossRef]
- Mitiouchkina, T.Y.; Firsov, A.P.; Titova, S.M.; Pushin, A.S.; Shulga, O.A.; Dolgov, S.V. Different approaches to produce transgenic virus B Resistant Chrysanthemum. Agronomy 2018, 8, 28. [Google Scholar] [CrossRef]
- Dellaporta, S.L.; Wood, J.; Hicks, J.B. A plant DNA minipreparation version II. Plant Mol. Biol. Report. 1983, 1, 19–21. [Google Scholar] [CrossRef]
- Bhandari, P.; Gupta, A.P.; Singh, B.K.; Vijay, K. Simultaneous densitometric determination of artemisinin, artemisinic acid and arteannuin B in Artemisia annua using reversed-phase thin layer chromatography. J. Sep. Sci. 2005, 28, 2288–2292. [Google Scholar] [CrossRef]
- Khan, S.; Ali, A.; Ahmad, S.; Abdin, M.Z. Affordable and rapid HPTLC method for the simultaneous analysis of artemisinin and its metabolite artemisinic acid in Artemisia annua L. Biomed. Chromatogr. 2015, 29, 1594–1603. [Google Scholar] [CrossRef]



| Stages of chrysanthemum transformation and analysis | Vector | |
| 1240 | 1250 | |
| Transformed explants | 600 | 600 |
| The explants with regenerants | 20 | 18 |
| Adventive shoots per one regenerating explant | 1.2 | 1.3 |
| Planted regenerants on the proliferation medium with kanamycin | 24 | 23 |
| The regenerants died on the proliferation medium with kanamycin | 16 | 21 |
| Kanamycin-resistant lines | 8 | 2 |
| NPT II -positive lines | 8 | 2 |
| Lines with all target genes | 2 | 1 |
| Transcription of all target genes | 2 | 0 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Firsov, A.; Mitiouchkina, T.; Shaloiko, L.; Pushin, A.; Vainstein, A.; Dolgov, S. Agrobacterium-Mediated Transformation of Chrysanthemum with Artemisinin Biosynthesis Pathway Genes. Plants 2020, 9, 537. https://doi.org/10.3390/plants9040537
Firsov A, Mitiouchkina T, Shaloiko L, Pushin A, Vainstein A, Dolgov S. Agrobacterium-Mediated Transformation of Chrysanthemum with Artemisinin Biosynthesis Pathway Genes. Plants. 2020; 9(4):537. https://doi.org/10.3390/plants9040537
Chicago/Turabian StyleFirsov, Aleksey, Tatiana Mitiouchkina, Lyubov Shaloiko, Alexander Pushin, Alexander Vainstein, and Sergey Dolgov. 2020. "Agrobacterium-Mediated Transformation of Chrysanthemum with Artemisinin Biosynthesis Pathway Genes" Plants 9, no. 4: 537. https://doi.org/10.3390/plants9040537
APA StyleFirsov, A., Mitiouchkina, T., Shaloiko, L., Pushin, A., Vainstein, A., & Dolgov, S. (2020). Agrobacterium-Mediated Transformation of Chrysanthemum with Artemisinin Biosynthesis Pathway Genes. Plants, 9(4), 537. https://doi.org/10.3390/plants9040537




