The Genetic Diversity of Cranberry Crop Wild Relatives, Vaccinium macrocarpon Aiton and V. oxycoccos L., in the US, with Special Emphasis on National Forests
Abstract
1. Introduction
2. Results
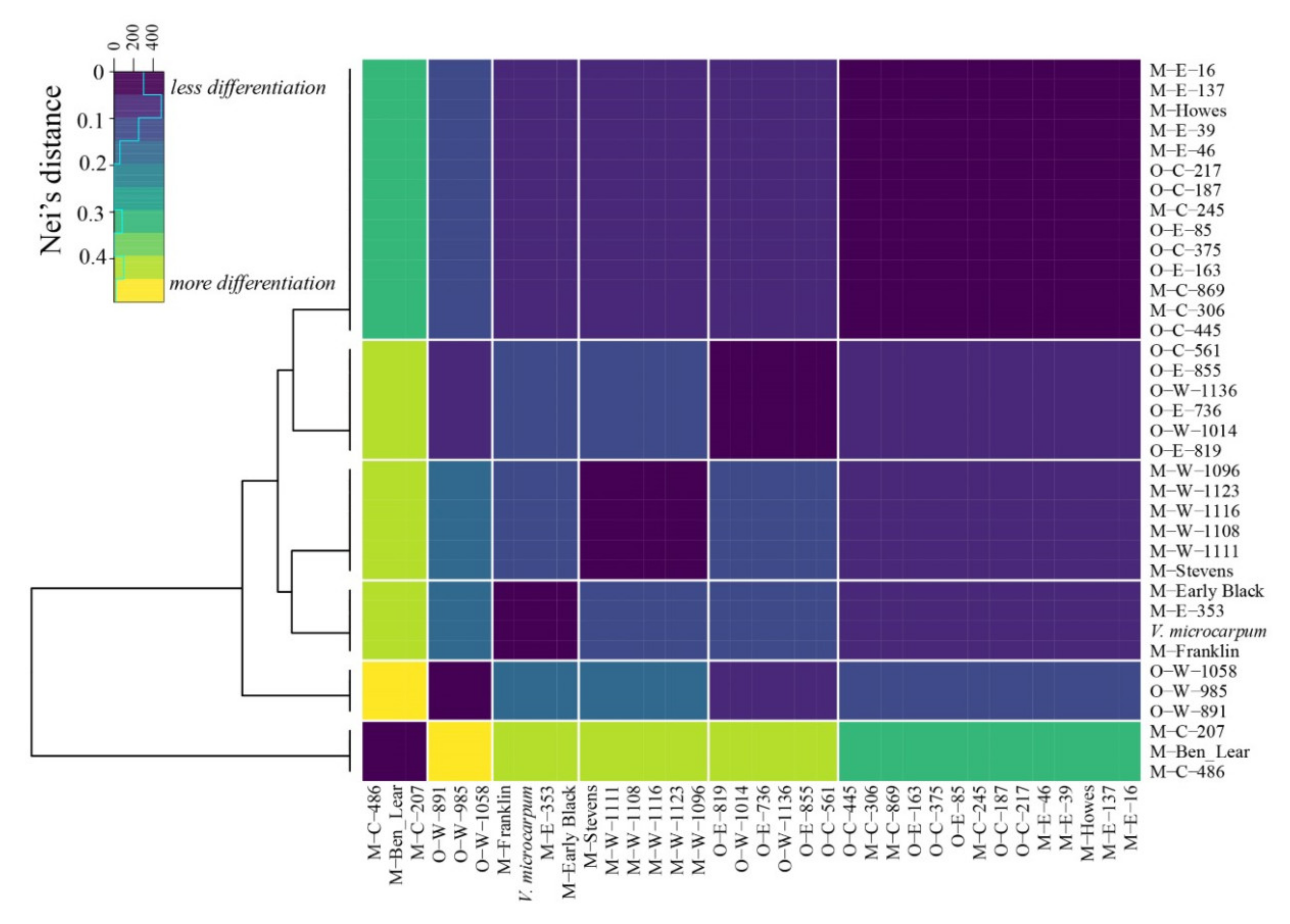
2.1. Exploring the Genetic Relationships of Wild Cranberries
2.2. Genetic Diversity Estimators and Population Structure of Vaccinium macrocarpon
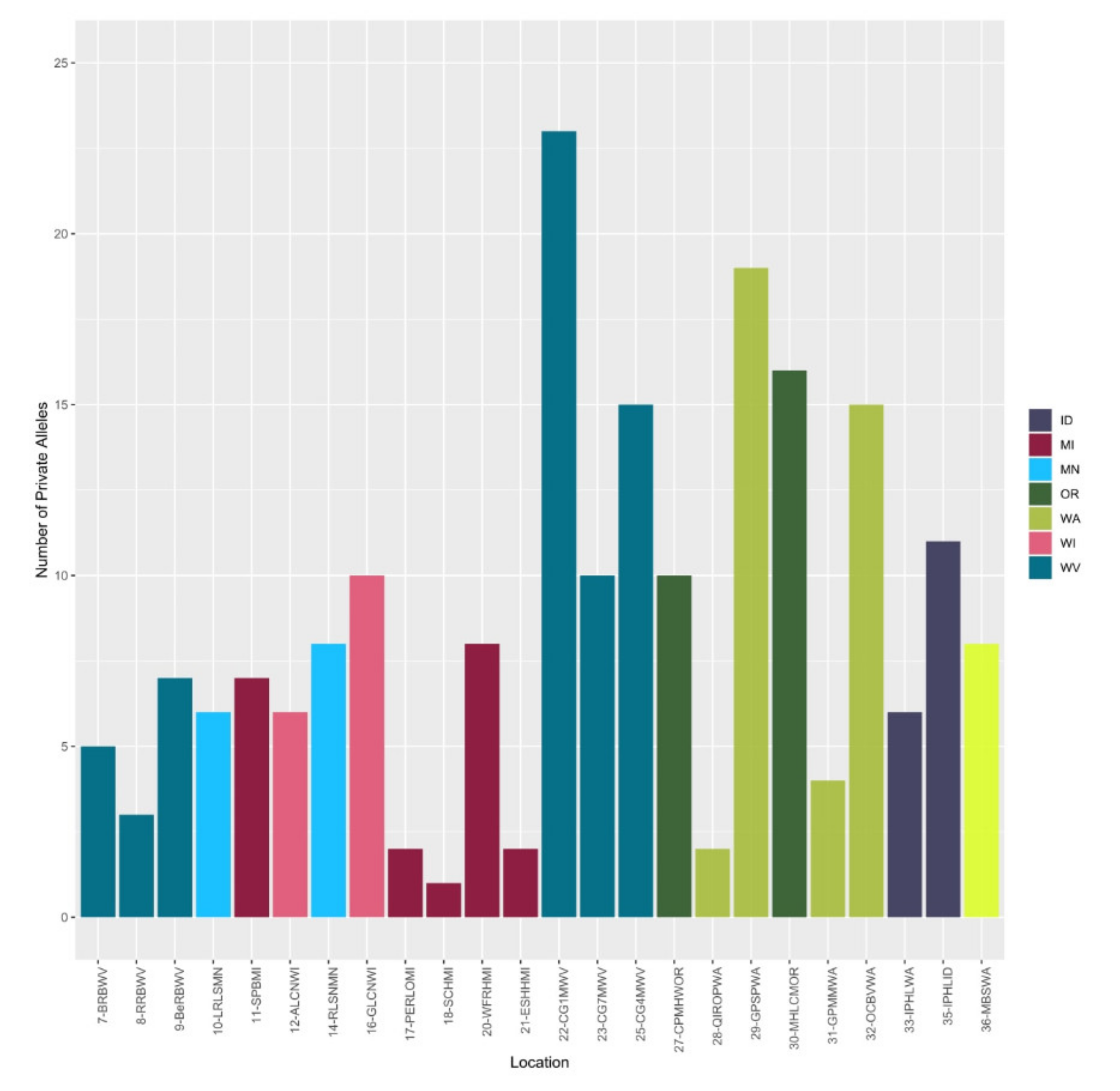
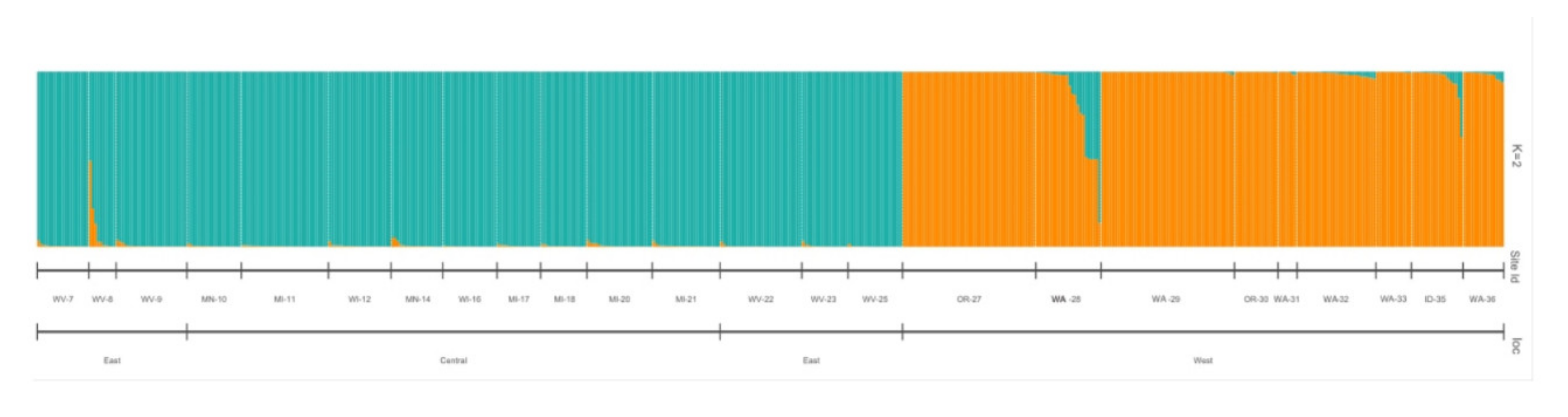
2.3. Genetic Diversity Estimators and Population Structure of Vaccinium oxycoccos
2.4. Analysis of Variable Regions in the Cranberry Organellar Genomes
3. Discussion
3.1. Genetic Analysis Shows High Differentiation between Morphologically Similar Species
3.2. Insight into the Genetic Diversity of Wild Populations of Vacciniium macrocarpon
3.3. Wild Vaccinium macrocarpon from Washington: An Escape or a Unique Population?
3.4. Wild Cranberry Populations from Washington Could Help Unravel a Complicated Evolutionary History
3.5. Insight into the Genetic Diversity of Wild Populations of Vaccinium oxycoccos
3.6. Understanding Genetic Variation in Cranberry and Its Implications for Conservation
4. Materials and Methods
4.1. Plant Materials
4.2. DNA Extraction
4.3. Simple Sequence Repeat (SSR) Markers, Polymerase Chain Reaction (PCR) Conditions and Analysis of Diversity
4.4. Analysis of Organelle Sequence Data and Genetic Relatedness Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Greene, S.L.; Khoury, C.K.; Williams, K.A. Wild Plant Genetic Resources in North America: An Overview. In North American Crop Wild Relatives, Volume 1: Conservation Strategies; Greene, S.L., Williams, K.A., Khoury, C.K., Eds.; Springer International Publishing: Cham, Switzerland, 2018; pp. 3–31. [Google Scholar]
- Khoury, C.K.; Greene, S.; Wiersema, J.; Maxted, N.; Jarvis, A.; Struik, P.C. An inventory of crop wild relatives of the United States. Crop Sci. 2013, 53, 1496–1508. [Google Scholar] [CrossRef]
- Jost, L.; Archer, F.; Flanagan, S.; Gaggiotti, O.; Hoban, S.; Latch, E. Differentiation measures for conservation genetics. Evol. Appl. 2018, 11, 1139–1148. [Google Scholar] [CrossRef] [PubMed]
- Vander Kloet, S.P. The Genus Vaccinium in North America; Agriculture Canada: Ottawa, ON, Canada, 1988. [Google Scholar]
- Song, G.-Q.; Hancock, J.F. Vaccinium. In Wild Crop Relatives: Genomic and Breeding Resources: Temperate Fruits; Kole, C., Ed.; Springer: Berlin/Heidelberg, Germany, 2011; pp. 197–221. [Google Scholar]
- Eck, P. The American Cranberry; Rutgers University Press: New Brunswick, NJ, USA, 1990. [Google Scholar]
- Schlautman, B.; Covarrubias-Pazaran, G.; Diaz-Garcia, L.; Iorizzo, M.; Polashock, J.; Grygleski, E.; Vorsa, N.; Zalapa, J. Construction of a High-Density American Cranberry (Vaccinium macrocarpon Ait.) Composite Map Using Genotyping-by-Sequencing for Multi-pedigree Linkage Mapping. G3 Genes Genomes Genet. 2017, 7, 1177–1189. [Google Scholar] [CrossRef] [PubMed]
- Vorsa, N.; Zalapa, J. Domestication, Genetics, and Genomics of the American Cranberry. Plant Breed. Rev. 2019, 43, 279–310. [Google Scholar]
- Yan, X.; Murphy, B.T.; Hammond, G.B.; Vinson, J.A.; Neto, C.C. Antioxidant activities and antitumor screening of extracts from cranberry fruit (Vaccinium macrocarpon). J. Agric. Food Chem. 2002, 50, 5844–5849. [Google Scholar] [CrossRef]
- McKay, D.L.; Blumberg, J.B. Cranberries (Vaccinium macrocarpon) and cardiovascular disease risk factors. Nutr. Rev. 2007, 65, 490–502. [Google Scholar] [CrossRef] [PubMed]
- Brown, P.N.; Turi, C.E.; Shipley, P.R.; Murch, S.J. Comparisons of large (Vaccinium macrocarpon Ait.) and small (Vaccinium oxycoccos L.; Vaccinium vitis-idaea L.) cranberry in British Columbia by phytochemical determination, antioxidant potential, and metabolomic profiling with chemometric analysis. Planta Med. 2012, 78, 630–640. [Google Scholar] [CrossRef]
- Roper, T.R.; Vorsa, N. Cranberry: Botany and Horticulture. In Horticultural Reviews; Janick, J., Ed.; John Wiley & Sons, Inc.: Oxford, UK, 1997; pp. 215–249. [Google Scholar]
- Vorsa, N.; Johnson-Cicalese, J. American Cranberry. In Fruit Breeding; Badenes, M.L., Byrne, D.H., Eds.; Springer US: Boston, MA, USA, 2012; pp. 191–223. [Google Scholar]
- Vander Kloet, S.P. The Taxonomy of Vaccinium § Oxycoccus. Rhodora 1983, 85, 1–43. [Google Scholar]
- Jacquemart, A.-L. Vaccinium oxycoccos L. (Oxycoccus Palustris Pers.) and Vaccinium Microcarpum (Turcz. ex Rupr.) Schmalh. (Oxycoccus Microcarpus Turcz. ex Rupr.). J. Ecol. 1997, 85, 381–396. [Google Scholar] [CrossRef]
- Areškevičiūtė, J.; Paulauskas, A.; Česonienė, L.; Daubaras, R. Genetic characterisation of wild cranberry (Vaccinium oxycoccos) from Čepkeliai reserve by the RAPD method. Biologija 2006, 1, 5–7. [Google Scholar]
- Mao, D.; Yu, L.; Chen, D.; Chen, D.; Li, L.; Zhu, Y.; Xiao, Y.; Zhang, D.; Chen, C. Multiple cold resistance loci confer the high cold tolerance adaptation of Dongxiang wild rice (Oryza rufipogon) to its high-latitude habitat. Theor. Appl. Genet. 2015, 128, 1359–1371. [Google Scholar] [CrossRef] [PubMed]
- Hokanson, S.C.; Lamboy, W.F.; Szewc-McFadden, A.K.; McFerson, J.R. Microsatellite (SSR) variation in a collection of Malus (apple) species and hybrids. Euphytica 2001, 118, 281–294. [Google Scholar] [CrossRef]
- Salehi, M.; Arzani, A.; Talebi, M.; Rokhzadi, A. Genetic diversity of wheat wild relatives using SSR markers. Genetika 2018, 50, 131–141. [Google Scholar] [CrossRef]
- Ali, A.; Pan, Y.; Wang, Q.; Wang, J.-D.; Chen, J.-L.; Gao, S.-J. Genetic diversity and population structure analysis of Saccharum and Erianthus genera using microsatellite (SSR) markers. Sci. Rep. 2019, 9, 395. [Google Scholar] [CrossRef]
- Rodriguez-Bonilla, L.; Cuevas, H.E.; Montero-Rojas, M.; Bird-Pico, F.; Luciano-Rosario, D.; Siritunga, D. Assessment of genetic diversity of sweet potato in Puerto Rico. PLoS ONE 2014, 9, e116184. [Google Scholar] [CrossRef]
- Zalapa, J.E.; Bougie, T.C.; Bougie, T.A.; Schlautman, B.J.; Wiesman, E.; Guzman, A.; Fajardo, D.A.; Steffan, S.; Smith, T. Clonal diversity and genetic differentiation revealed by SSR markers in wild Vaccinium macrocarpon and Vaccinium oxycoccos. Ann. Appl. Biol. 2015, 166, 196–207. [Google Scholar] [CrossRef]
- Rodríguez-Bonilla, L.; Bonilla, F.R.; Matusinec, D.; Wiesman, E.; Schoville, S.D.; Atucha, A.; Zalapa, J. Exploring the Genetic Diversity of Wild Cranberry Populations in the Upper Midwestern United States. Crop Sci. 2019, 59, 2413–2428. [Google Scholar] [CrossRef]
- Rodriguez-Bonilla, L.; Rohde, J.; Matusinec, D.; Zalapa, J. Cross-transferability analysis of SSR markers developed from the American Cranberry (Vaccinium macrocarpon Ait.) to other Vaccinium species of agricultural importance. Genet. Resour. Crop Evol. 2019, 66, 1713–1725. [Google Scholar] [CrossRef]
- Segarra-Moragues, J.G.; Palop-Esteban, M.; González-Candelas, F.; Catalán, P. On the verge of extinction: Genetics of the critically endangered Iberian plant species, Borderea chouardii (Dioscoreaceae) and implications for conservation management. Mol. Ecol. 2005, 14, 969–982. [Google Scholar] [CrossRef]
- Marfil, C.F.; Hidalgo, V.; Masuelli, R.W. In situ conservation of wild potato germplasm in Argentina: Example and possibilities. Glob. Ecol. Conserv. 2015, 3, 461–476. [Google Scholar] [CrossRef]
- Stewart, C.N.; Neal Stewart, C.; Nilsen, E.T. Phenotypic Plasticity and Genetic Variation of Vaccinium macrocarpon, the American Cranberry. I. Reaction Norms of Clones from Central and Marginal Populations in a Common Garden. Int. J. Plant Sci. 1995, 156, 687–697. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [PubMed]
- Earl, D.A.; von Holdt, B.M. Structure Harvester: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Fajardo, D.; Schlautman, B.; Steffan, S.; Polashock, J.; Vorsa, N.; Zalapa, J. The American cranberry mitochondrial genome reveals the presence of selenocysteine (tRNA-Sec and SECIS) insertion machinery in land plants. Gene 2014, 536, 336–343. [Google Scholar] [CrossRef]
- Diaz-Garcia, L.; Rodriguez-Bonilla, L.; Rohde, J.; Smith, T.; Zalapa, J. Pacbio Sequencing Reveals Identical Organelle Genomes between American Cranberry (Vaccinium macrocarpon Ait.) and a Wild Relative. Genes 2019, 10, 291. [Google Scholar] [CrossRef]
- Dempewolf, H.; Baute, G.; Anderson, J.; Kilian, B.; Smith, C.; Guarino, L. Past and future use of wild relatives in crop breeding. Crop Sci. 2017, 57, 1070–1082. [Google Scholar] [CrossRef]
- Gallardo, R.K.; Klingthong, P.; Zhang, Q.; Polashock, J.; Atucha, A.; Zalapa, J.; Rodriguez-Soana, C.; Vorsa, N.; Iorizzo, M. Breeding Trait Priorities of the Cranberry Industry in the United States and Canada. HortScience 2018, 53, 1467–1474. [Google Scholar] [CrossRef]
- Smith, T.W.; Walinga, C.; Wang, S.; Kron, P.; Suda, J.; Zalapa, J. Evaluating the relationship between diploid and tetraploid Vaccinium oxycoccos (Ericaceae) in eastern Canada. Botany 2015, 93, 623–636. [Google Scholar] [CrossRef]
- Bruederle, L.P.; Hugan, M.S.; Dignan, J.M.; Vorsa, N. Genetic variation in natural populations of the large cranberry, Vaccinium macrocarpon Ait.(Ericaceae). Bull. Torrey Bot. Club 1996, 123, 41–47. [Google Scholar] [CrossRef]
- Llácer, G.; Badenes, M.L.; Calvo, J.M. Plant material of loquat in Mediterranean countries. In: Proc. First Int. Loquat Symp. Options Méditerranéennes: Série A. Séminaires Méditerranéens 2003, 58, 45–52. [Google Scholar]
- Montero-Rojas, M.; Correa, A.M.; Siritunga, D. Molecular differentiation and diversity of cassava (Manihot esculenta) taken from 162 locations across Puerto Rico and assessed with microsatellite markers. AoB Plants 2011, 2011, plr010. [Google Scholar] [CrossRef]
- Stewart, C.N.; Excoffier, L. Assessing population genetic structure and variability with RAPD data: Application to Vaccinium macrocarpon (American Cranberry). J. Evol. Biol. 1996, 9, 153–171. [Google Scholar] [CrossRef]
- Novy, R.G.; Vorsa, N. Identification of intracultivar genetic heterogeneity in cranberry using silver-stained RAPDs. HortScience 1995, 30, 600–604. [Google Scholar] [CrossRef]
- Mahy, G.; Bruederle, L.P.; Connors, B.; Van Hofwegen, M.; Vorsa, N. Allozyme evidence for genetic autopolyploidy and high genetic diversity in tetraploid cranberry, Vaccinium oxycoccos (Ericaceae). Am. J. Bot. 2000, 87, 1882–1889. [Google Scholar] [CrossRef] [PubMed]
- Kalyan Chakravarthi, B.; Naravaneni, R. SSR marker based DNA fingerprinting and diversity study in rice (Oryza sativa. L). Afr. J. Biotechnol. 2006, 5, 684–688. [Google Scholar]
- Montero-Rojas, M.; Ortiz, M.; Beaver, J.S.; Siritunga, D. Genetic, morphological and cyanogen content evaluation of a new collection of Caribbean Lima bean (Phaseolus lunatus L.) landraces. Genet. Resour. Crop Evol. 2013, 60, 2241–2252. [Google Scholar] [CrossRef]
- Schlautman, B.; Covarrubias-Pazaran, G.; Rodriguez-Bonilla, L.; Hummer, K.; Bassil, N.; Smith, T.; Zalapa, J. Genetic diversity and cultivar variants in the National Clonal Germplasm Repository cranberry (Vaccinium macrocarpon Aiton) collection. J. Genet. 2018, 97, 1339–1351. [Google Scholar] [CrossRef]
- Raymond, O.; Gouzy, J.; Just, J.; Badouin, H.; Verdenaud, M.; Lemainque, A.; Vergne, P.; Moja, S.; Choisne, N.; Pont, C.; et al. The Rosa genome provides new insights into the domestication of modern roses. Nat. Genet. 2018, 50, 772–777. [Google Scholar] [CrossRef]
- Van Strien, M.J.; Holderegger, R.; Van Heck, H.J. Isolation-by-distance in landscapes: Considerations for landscape genetics. Heredity 2015, 114, 27–37. [Google Scholar] [CrossRef]
- Nei, M.; Maruyama, T.; Chakraborty, R. The Bottleneck Effect and Genetic Variability in Populations. Evolution 1975, 29, 1–10. [Google Scholar] [CrossRef]
- Heintzman, P.D.; Froese, D.; Ives, J.W.; Soares, A.E.; Zazula, G.D.; Letts, B.; Andrews, T.D.; Driver, J.C.; Hall, E.; Hare, P.G.; et al. Bison phylogeography constrains dispersal and viability of the Ice Free Corridor in western Canada. Proc. Natl. Acad. Sci. USA 2016, 113, 8057–8063. [Google Scholar] [CrossRef]
- Česonienė, L.; Daubaras, R.; Paulauskas, A.; Zukauskiene, J.; Zych, M. Morphological and genetic diversity of European cranberry (Vaccinium oxycoccos L.; Ericaceae) clones in Lithuanian reserves. Acta Soc. Bot. Pol. 2013, 82, 211–217. [Google Scholar] [CrossRef]
- Zhang, Y.; Zalapa, J.; Jakubowski, A.R.; Price, D.L.; Acharya, A.; Wei, Y.; Brummer, C.; Kaeppler, S.M.; Casler, M.D. Natural hybrids and gene flow between upland and lowland switchgrass. Crop Sci. 2011, 51, 2626–2641. [Google Scholar] [CrossRef]
- Szpiech, Z.A.; Rosenberg, N.A. On the size distribution of private microsatellite alleles. Theor. Popul. Biol. 2011, 80, 100–113. [Google Scholar] [CrossRef] [PubMed]
- Doyle, J.J.; Doyle, J.L. CTAB DNA extraction in plants. Phytochem. Bull. 1987, 19, 11–15. [Google Scholar]
- Fajardo, D.; Morales, J.; Zhu, H.; Steffan, S.; Harbu, R.; Bassil, N.; Hummer, K.; Polashock, J.; Vorsa, N.; Zalapa, J. Discrimination of American cranberry cultivars and assessment of clonal heterogeneity using microsatellite markers. Plant Mol. Biol. 2013, 31, 264–271. [Google Scholar] [CrossRef]
- Schlautman, B.; Fajardo, D.; Bougie, T.; Wiesman, E.; Polashock, J.; Vorsa, N.; Steffan, S.; Zalapa, J. Development and validation of 697 novel polymorphic genomic and EST-SSR markers in the American cranberry (Vaccinium macrocarpon Ait.). Molecules 2015, 20, 2001–2013. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research--an update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018. [Google Scholar]
- Kamvar, Z.N.; Tabima, J.F.; Grünwald, N.J. Poppr: An R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2014, 2, e281. [Google Scholar] [CrossRef]
- Meirmans, P.G.; Van Tienderen, P.H. Genotype and Genodive: Two programs for the analysis of genetic diversity of asexual organisms. Mol. Ecol. Notes 2004, 4, 792–794. [Google Scholar] [CrossRef]
- Jombart, T. Adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 2008, 24, 1403–1405. [Google Scholar] [CrossRef] [PubMed]
- Wickham, H. ggplot2. WIREs Comp. Stat. 2011, 3, 180–185. [Google Scholar] [CrossRef]
- Kassambara, A.; Mundt, F. Package Factoextra: Extract and Visualize the Results of Multivariate Data Analyses; Comprehensive R Arch. Network: Vienna, Austria, 2016. [Google Scholar]
- Galili, T. Dendextend: An R package for visualizing, adjusting and comparing trees of hierarchical clustering. Bioinformatics 2015, 31, 3718–3720. [Google Scholar] [CrossRef]
- Gu, Z.; Gu, L.; Eils, R.; Schlesner, M.; Brors, B. circlize implements and enhances circular visualization in R. Bioinformatics 2014, 30, 2811–2812. [Google Scholar] [CrossRef] [PubMed]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar]
- Francis, R.M. Pophelper: An R package and web app to analyse and visualize population structure. Mol. Ecol. Resour. 2017, 17, 27–32. [Google Scholar] [CrossRef] [PubMed]
- Becker, R.A.; Wilks, A.R.; Brownrigg, R.; Minka, T.P.; Deckmyn, A. Maps: Draw Geographical Maps; R Package Version 2. 2013. Available online: https://cran.r-project.org/web/packages/maps/index.html (accessed on 21 October 2020).
- Pagès, H.; Aboyoun, P.; Gentleman, R.; DebRoy, S. Biostrings: Efficient Manipulation of Biological Strings; R Package Version 2.54.0. 2019. Available online: https://bioconductor.riken.jp/packages/3.10/bioc/html/Biostrings.html (accessed on 21 October 2020).
- Nei, M. Genetic Distance between Populations. Am. Nat. 1972, 106, 283–292. [Google Scholar] [CrossRef]
- Pembleton, L.W.; Cogan, N.O.I.; Forster, J.W. StAMPP: An R package for calculation of genetic differentiation and structure of mixed-ploidy level populations. Mol. Ecol. Resour. 2013, 13, 946–952. [Google Scholar] [CrossRef]
- Kahle, D.; Wickham, H. Ggmap: Spatial Visualization with ggplot2. R J. 2013, 5, 144–161. [Google Scholar] [CrossRef]


| Site ID | Species | Sample IDs | State | National Forest/ Administrative Area | Location | Latitude | Longitude | Regional Group |
|---|---|---|---|---|---|---|---|---|
| 2-GPVA (VA-2) * | V. macrocarpon | VAC2013 (4–23) | Virginia | George Washington National Forest | Green Pond | 37.94087 | −79.0526 | East |
| 3-JBCTN (TN-3) * | V. macrocarpon | VAC2013 (25–40) | Tennessee | Cherokee National Forest | John’s Bog | 36.52908 | −81.964 | East |
| 4-OBCTN (TN-4) * | V. macrocarpon | VAC2013 (41–47) | Tennessee | Cherokee National Forest | Osborne Bog | 36.48826 | −81.9652 | East |
| 5-BBKPNC (NC-5) * | V. macrocarpon | VAC2013 (48–53) | North Carolina | Pisgah National Forest | Black Balsam Knob | 35.32943 | −82.8782 | East |
| 6-IGSPNC (NC-6) * | V. macrocarpon | VAC2013 (54–69) | North Carolina | Pisgah National Forest | Investor Gap Seep | 35.34345 | −82.8697 | East |
| 7-BRBWV (WV-7) * | V. macrocarpon V. oxycoccos | WV2014-2 (1–18) WV2014-1 (1–19) | West Virginia | Monongahela National Forest | Big Run Bog | 39.11805 | −79.5845 | East |
| 8-RRBWV (WV-8) * | V. macrocarpon V. oxycoccos | WV2014-4 (1–24) WV2014-3 (1–10) | West Virginia | Monongahela National Forest | Red Run Bog | 39.07227 | −79.4784 | East |
| 9-BeRBWV (WV-9) * | V. macrocarpon V. oxycoccos | WV2014-6 (1–2) WV2014-5 (1–26) | West Virginia | Monongahela National Forest | Bear Rocks Bog | 39.06535 | −79.3051 | East |
| 10-LRLSMN (MN-10) * | V. macrocarpon V. oxycoccos | VAC2015-M (165–184) VAC2015-O (145–164) | Minnesota | Superior National Forest | Little Rice Lake | 47.70847 | −92.4411 | Central |
| 11-SPBMI (MI-11) * | V. macrocarpon V. oxycoccos | VAC2015-M (220–254) VAC2015-O (185–219) | Michigan | Keweenaw Bay Indian Community | Sand Point Bog | 46.79143 | −88.4669 | Central |
| 12-ALCNWI (WI-12) * | V. macrocarpon V. oxycoccos | VAC2015-M (278–298) VAC2015-O (255–277) | Wisconsin | Chequamegon-Nicolet National Forest | Atkins Lake | 45.64951 | −89.0397 | Central |
| 13-ANFPA (PA-13) * | V. macrocarpon | VAC2015-M (299–336) | Pennsylvania | Allegheny National Forest | Along both sides of FR320, approximately 0.2 miles south of junction with FR455 | 41.81581 | −78.7341 | East |
| 14-RLSNMN (MN-14) * | V. oxycoccos | VAC2015-O (337–356) | Minnesota | Superior National Forest | Rice Lake | 47.56709 | −92.3689 | Central |
| 15-CLSNMN (MN-15) * | V. macrocarpon | VAC2015-M (357–381) | Minnesota | Superior National Forest | Cranberry Lake | 47.51161 | −92.0166 | Central |
| 16-GLCNWI (WI-16) * | V. macrocarpon V. oxycoccos | VAC2015-M (382–402) VAC2015-O (403–422) | Wisconsin | Chequamegon-Nicolet National Forest | Glocke Lake | 45.33362 | −88.5702 | Central |
| 17-PERLOMI (MI-17) * | V. macrocarpon V. oxycoccos | USFS-ONF-2015-1-1, USFS-ONF-2015- 2 (1–16) USFS-ONF-2015-2 (1o–160) | Michigan | Ottawa National Forest | Pond east of Raven Lake | 46.26263 | −89.21718 | Central |
| 18-SCHMI (MI-18) * | V. macrocarpon V. oxycoccos | USFS-HNF-2015-1 (1–8, 12–17, 29–35, 38–47) USFS-HNF-2015-1 (9–11, 18–28, 36–37, 48) | Michigan | Hiawatha National Forest | South side FR2268 to east of H94, 0.8 miles south of Stutts Creek crossing | 46.29175 | −86.455 | Central |
| 19-NHHMI (MI-19) * | V. macrocarpon | USFS-HNF-2015-2 (1– 16) | Michigan | Hiawatha National Forest | North of Haywire Trail, unmarked two track West of Highway 94 | 46.29177 | −86.454 | Central |
| 20-WFRHMI (MI-20) * | V. oxycoccos | USFS-HNF-2015-3 (1–24) | Michigan | Hiawatha National Forest | West Side of FR13, north of FR2447 | 46.1807 | −86.4242 | Central |
| 21-ESHHMI (MI-21) * | V. oxycoccos | USFS-HNF-2015-4 (1–24) | Michigan | Hiawatha National Forest | East Side of Highway 13, north of FR2020 | 46.06647 | −86.6438 | Central |
| 22-CG1MWV (WV-22) * | V. macrocarpon V. oxycoccos | USFS-MNF-2015-1-(11–12), USFS-MNF-2015-1-13 (k-p) USFS-MNF-2015-1- (1–10), USFS-MNF-2015-1 13 (a-j), (USFS-MNF-2015-1 (14–22) | West Virginia | Monongahela National Forest | Cranberry Glades 1 | 38.19939 | −80.272 | East |
| 23-CG7MWV (WV-23) * | V. oxycoccos | USFS-MNF-2015-7 (1–16) | West Virginia | Monongahela National Forest | Cranberry Glades 7 | 38.20603 | −80.2773 | East |
| 24-CG5MWV (WV-24) * | V. macrocarpon | USFS-MNF-2015-5 (1–17) | West Virginia | Monongahela National Forest | Cranberry Glades 5 | 38.19943 | −80.2654 | East |
| 25-CG4MWV (WV-25) * | V. oxycoccos | USFS-MNF-2015-4 (1–20) | West Virginia | Monongahela National Forest | Cranberry Glades 4 | 38.20012 | −80.2651 | East |
| 26-UILCNWI (WV-26) * | V. macrocarpon | USFS-CNNF-2015-4 (1–17) | Wisconsin | Chequamegon-Nicolet National Forest | Upper Island Lake | 45.25023 | −88.5576 | Central |
| 27-CPMHWOR (OR-27) * | V. oxycoccos | USFS-MHNF-2017-1 (1–49) | Oregon | Mount Hood National Forest | Camas Prairie | 45.1382 | −121.566 | West |
| 28-QIROPWA (WA-28) * | V. oxycoccos | QIR-2017-1 (1–24) | Washington | Quinault Indian Reservation | Otook Prairie | 47.41137 | −124.155 | West |
| 29-GPSPWA (WA-29) * | V. oxycoccos | USFS-GPNF-2017-2 (1–49) | Washington | Gifford Pinchot National Forest | South Prairie | 45.90969 | −121.699 | West |
| 30-MHLCMOR (OR-30) * | V. oxycoccos | USFS-MHNF-2017-2 (1–16) | Oregon | Mount Hood National Forest | Little Crater Meadow | 45.14545 | −121.741 | West |
| 31-GPMMWA (WA-31) * | V. oxycoccos | USFS-GPNF-2017-1 (1–7) | Washington | Gifford Pinchot National Forest | McClellan Meadows | 45.99633 | −121.89 | West |
| 32-OCBVWA (WA-32) * | V. oxycoccos | USFS-OLNF-2017-1 (1–29) | Washington | Olympic National Forest | Cranberry Bog Botanical Area | 47.98635 | −123.114 | West |
| 33-IPHLWA (WA-33) * | V. oxycoccos | USFS-IPMF-2018-1 (1–13) | Washington | Idaho Panhandle National Forest | Huff Lake | 48.74059 | −117.063 | West |
| 34-OWFLBWA (WA-34) * | V. macrocarpon | USFS-OWNF-2018-1 (1–36) | Washington | Okanogan-Wenatchee | Fish Lake Bog | 47.8253 | −120.723 | West |
| 35-IPHLID (ID-35) * | V. oxycoccos | USFS-IPNF-2018-2 (1–19) | Idaho | Idaho Panhandle National Forest | Hager Lake | 48.59713 | −116.97 | West |
| 36-MBSWA (WA-36) * | V. oxycoccos | USFSMBSNF-2018-1 (1–15) | Washington | Mt. Baker-Snoqualmie | Morovitz Wetland Complex | 48.74092 | −121.674 | West |
| V. macrocarpon | V. oxycoccos | |
|---|---|---|
| Total alleles | 613 | 881 |
| N 1 | 388 | 539 |
| Num 2 | 19.15 | 27.51 |
| Ho 3 | 0.99 | 0.71 |
| Hs 4 | 0.51 | 0.72 |
| Ht 5 | 0.75 | 0.80 |
| Gis 6 | −0.95 | 0.02 |
| G’st(Nei) 7 | 0.33 | 0.09 |
| Site ID | N 1 | Num 2 | Ho 3 | Hs 4 | Ht 5 | Gis 6 |
|---|---|---|---|---|---|---|
| 2-GPVA | 11 | 3.25 | 1.00 | 0.43 | 0.43 | −1.28 |
| 3-JBCTN | 16 | 2.31 | 1.00 | 0.23 | 0.23 | −3.20 |
| 4-OBCTN | 7 | 2.40 | 1.00 | 0.33 | 0.33 | −2.01 |
| 5-BBKPNC | 6 | 2.00 | 1.00 | 0.21 | 0.21 | −3.58 |
| 6-IGSPNC | 15 | 2.20 | 1.00 | 0.33 | 0.33 | −2.03 |
| 7-BRBWV | 18 | 4.10 | 0.99 | 0.55 | 0.55 | −0.79 |
| 8-RRBWV | 24 | 5.73 | 0.99 | 0.66 | 0.66 | −0.48 |
| 10-LRLSMN | 20 | 4.46 | 1.00 | 0.55 | 0.55 | −0.81 |
| 11-SPBMI | 31 | 7.15 | 1.00 | 0.68 | 0.68 | −0.46 |
| 12-ALCNWI | 21 | 5.31 | 1.00 | 0.62 | 0.62 | −0.60 |
| 13-ANFPA | 38 | 3.45 | 1.00 | 0.52 | 0.52 | −0.89 |
| 15-CLSNMN | 25 | 4.64 | 1.00 | 0.60 | 0.60 | −0.65 |
| 16-GLCNWI | 21 | 5.37 | 1.00 | 0.60 | 0.60 | −0.66 |
| 17-PERLOMI | 17 | 4.75 | 1.00 | 0.51 | 0.51 | −0.93 |
| 18-SCHMI | 31 | 6.40 | 1.00 | 0.63 | 0.63 | −0.56 |
| 19-NHHMI | 15 | 3.96 | 1.00 | 0.51 | 0.51 | −0.94 |
| 24-CG5MWV | 16 | 5.34 | 0.99 | 0.62 | 0.62 | −0.58 |
| 26-UILCNWI | 14 | 6.06 | 1.00 | 0.66 | 0.66 | −0.49 |
| 34-OWFLBWA | 35 | 3.12 | 0.97 | 0.48 | 0.48 | −1.00 |
| Site ID | N 1 | Num 2 | Ho 3 | Hs 4 | Ht 5 | Gis 6 |
|---|---|---|---|---|---|---|
| 7-BRBWV | 19 | 6.43 | 0.75 | 0.73 | 0.73 | −0.02 |
| 8-RRBWV | 10 | 4.75 | 0.72 | 0.83 | 0.83 | 0.12 |
| 9-BeRBWV | 26 | 8.12 | 0.64 | 0.74 | 0.74 | 0.12 |
| 10-LRLSMN | 20 | 8.81 | 0.69 | 0.74 | 0.74 | 0.06 |
| 11-SPBMI | 32 | 11.3 | 0.69 | 0.77 | 0.77 | 0.10 |
| 12-ALCNWI | 23 | 11.0 | 0.70 | 0.77 | 0.77 | 0.09 |
| 14-RLSNMN | 19 | 9.16 | 0.70 | 0.78 | 0.78 | 0.10 |
| 16-GLCNWI | 20 | 9.81 | 0.79 | 0.79 | 0.79 | −0.00 |
| 17-PERLOMI | 16 | 9.06 | 0.80 | 0.77 | 0.77 | −0.03 |
| 18-SCHMI | 16 | 6.50 | 0.73 | 0.70 | 0.70 | −0.03 |
| 20-WFRHMI | 24 | 9.65 | 0.74 | 0.77 | 0.77 | 0.03 |
| 21-ESHHMI | 25 | 9.65 | 0.75 | 0.76 | 0.76 | 0.01 |
| 22-CG1MWV | 30 | 10.3 | 0.74 | 0.76 | 0.76 | 0.01 |
| 23-CG7MWV | 16 | 8.78 | 0.77 | 0.76 | 0.76 | −0.01 |
| 25-CG4MWV | 20 | 9.93 | 0.75 | 0.77 | 0.77 | 0.02 |
| 27-CPMHWOR | 49 | 7.25 | 0.68 | 0.65 | 0.65 | −0.05 |
| 28-QIROPWA | 24 | 4.66 | 0.45 | 0.59 | 0.59 | 0.23 |
| 29-GPSPWA | 49 | 9.45 | 0.75 | 0.69 | 0.69 | −0.08 |
| 30-MHLCMOR | 16 | 6.25 | 0.70 | 0.66 | 0.66 | −0.07 |
| 31-GPMMWA | 7 | 5.06 | 0.76 | 0.69 | 0.69 | −0.11 |
| 32-OCBVWA | 29 | 9.54 | 0.71 | 0.68 | 0.68 | −0.03 |
| 33-IPHLWA | 13 | 2.71 | 0.67 | 0.51 | 0.51 | −0.31 |
| 35-IPHLID | 19 | 4.78 | 0.72 | 0.64 | 0.64 | −0.11 |
| 36-MBSWA | 15 | 4.96 | 0.71 | 0.65 | 0.65 | −0.10 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rodriguez-Bonilla, L.; Williams, K.A.; Rodríguez Bonilla, F.; Matusinec, D.; Maule, A.; Coe, K.; Wiesman, E.; Diaz-Garcia, L.; Zalapa, J. The Genetic Diversity of Cranberry Crop Wild Relatives, Vaccinium macrocarpon Aiton and V. oxycoccos L., in the US, with Special Emphasis on National Forests. Plants 2020, 9, 1446. https://doi.org/10.3390/plants9111446
Rodriguez-Bonilla L, Williams KA, Rodríguez Bonilla F, Matusinec D, Maule A, Coe K, Wiesman E, Diaz-Garcia L, Zalapa J. The Genetic Diversity of Cranberry Crop Wild Relatives, Vaccinium macrocarpon Aiton and V. oxycoccos L., in the US, with Special Emphasis on National Forests. Plants. 2020; 9(11):1446. https://doi.org/10.3390/plants9111446
Chicago/Turabian StyleRodriguez-Bonilla, Lorraine, Karen A. Williams, Fabian Rodríguez Bonilla, Daniel Matusinec, Andrew Maule, Kevin Coe, Eric Wiesman, Luis Diaz-Garcia, and Juan Zalapa. 2020. "The Genetic Diversity of Cranberry Crop Wild Relatives, Vaccinium macrocarpon Aiton and V. oxycoccos L., in the US, with Special Emphasis on National Forests" Plants 9, no. 11: 1446. https://doi.org/10.3390/plants9111446
APA StyleRodriguez-Bonilla, L., Williams, K. A., Rodríguez Bonilla, F., Matusinec, D., Maule, A., Coe, K., Wiesman, E., Diaz-Garcia, L., & Zalapa, J. (2020). The Genetic Diversity of Cranberry Crop Wild Relatives, Vaccinium macrocarpon Aiton and V. oxycoccos L., in the US, with Special Emphasis on National Forests. Plants, 9(11), 1446. https://doi.org/10.3390/plants9111446






