Characteristics and Expression Pattern of MYC Genes in Triticum aestivum, Oryza sativa, and Brachypodium distachyon
Abstract
1. Introduction
2. Results
2.1. Wheat, Rice, and B. distachyon MYC TFs
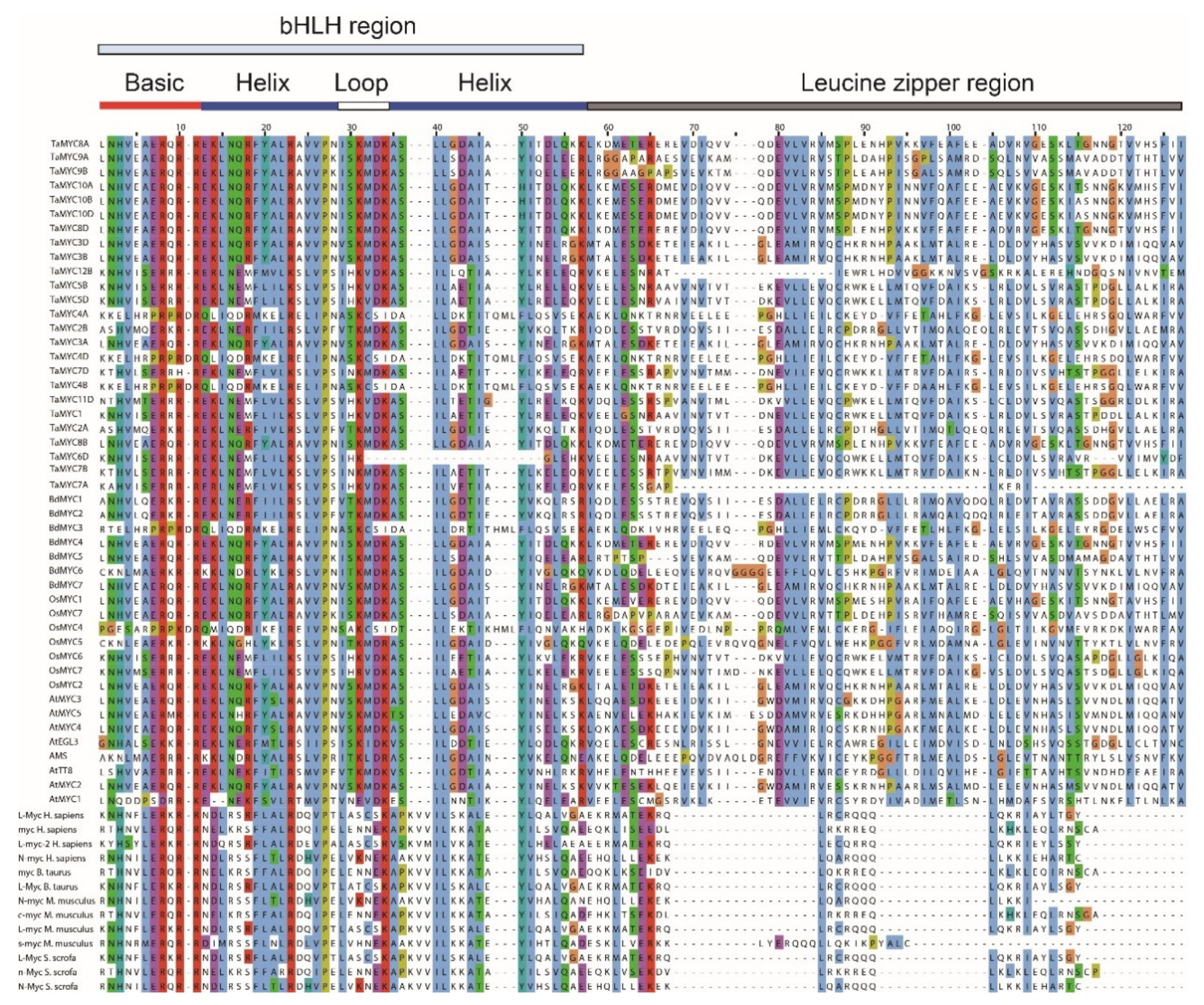
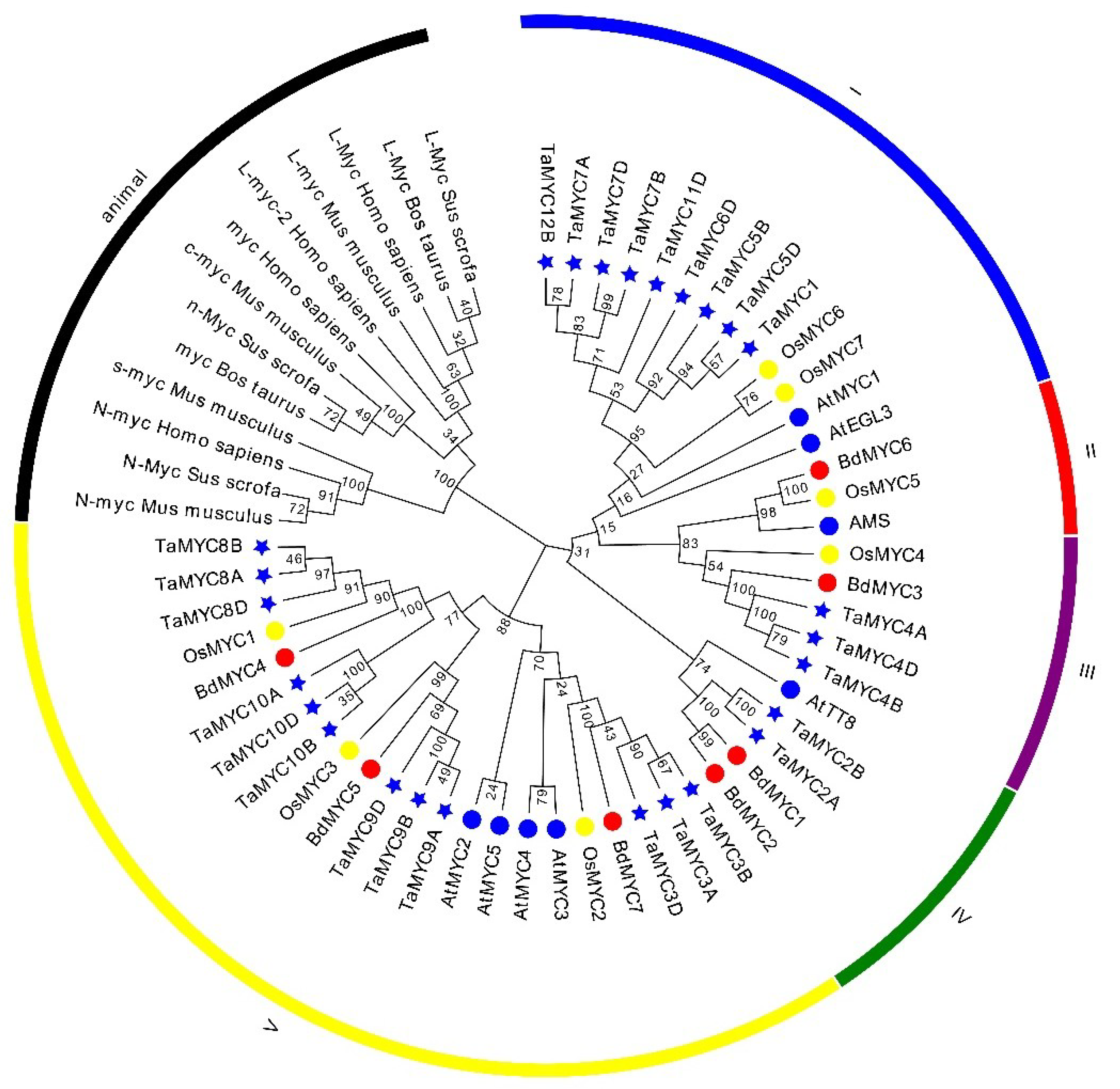
2.2. Sequence Alignment and Phylogenetic Tree of MYC TFs
2.3. Gene Structures and Conserved Motifs
2.4. Synteny and Homologous Gene Pairs
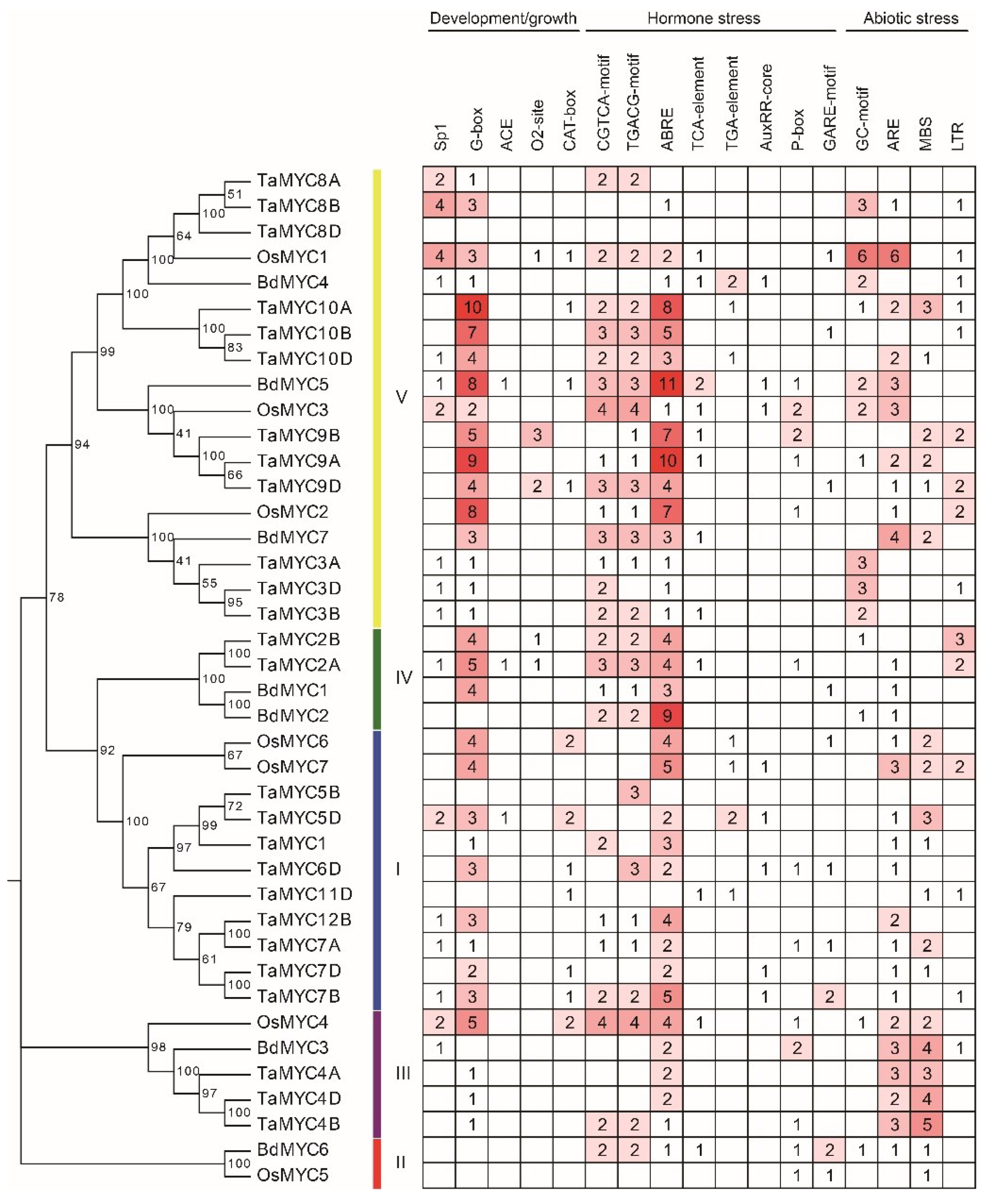
2.5. Identified Cis-Elements in MYC Gene Promoters
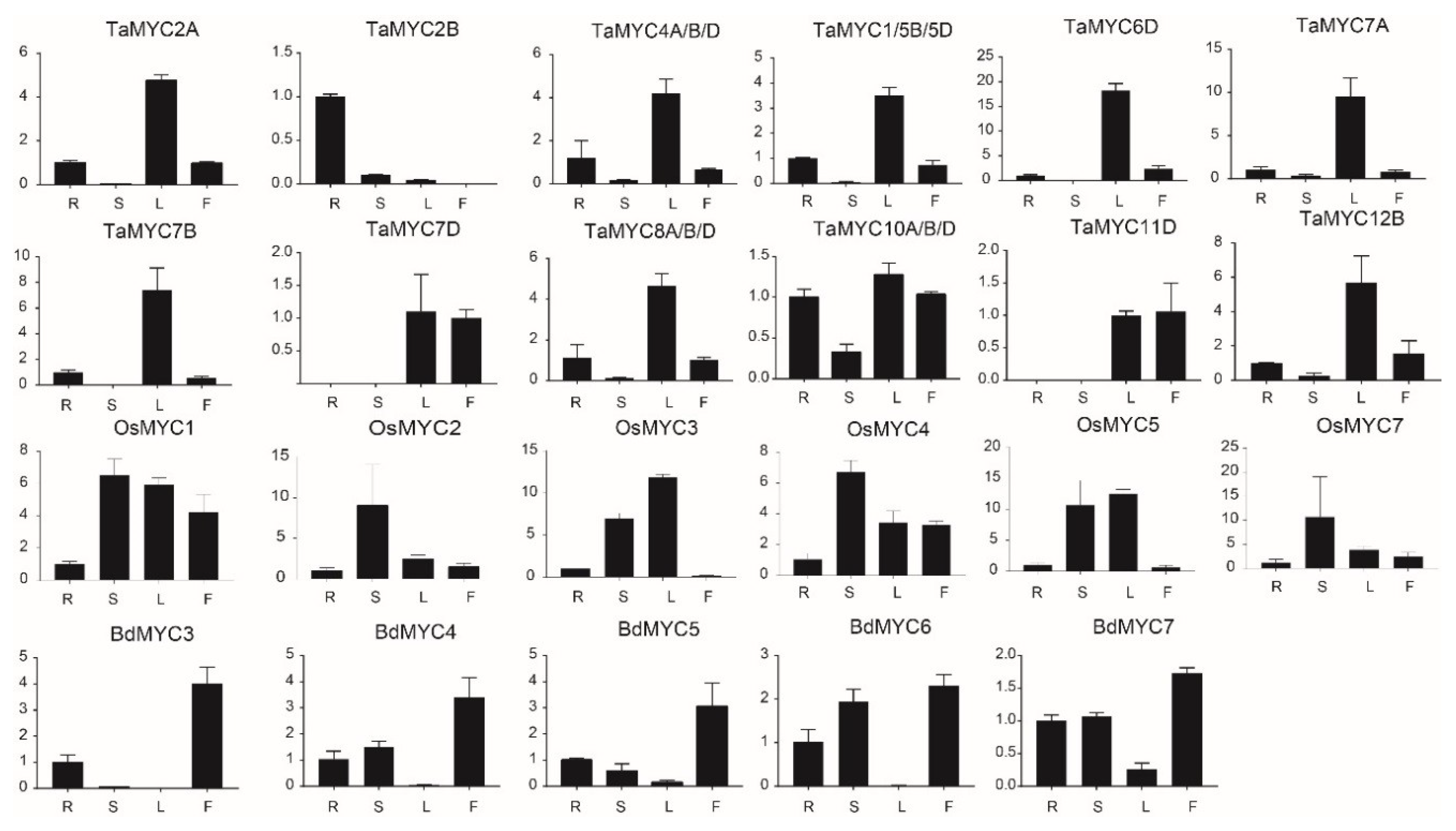
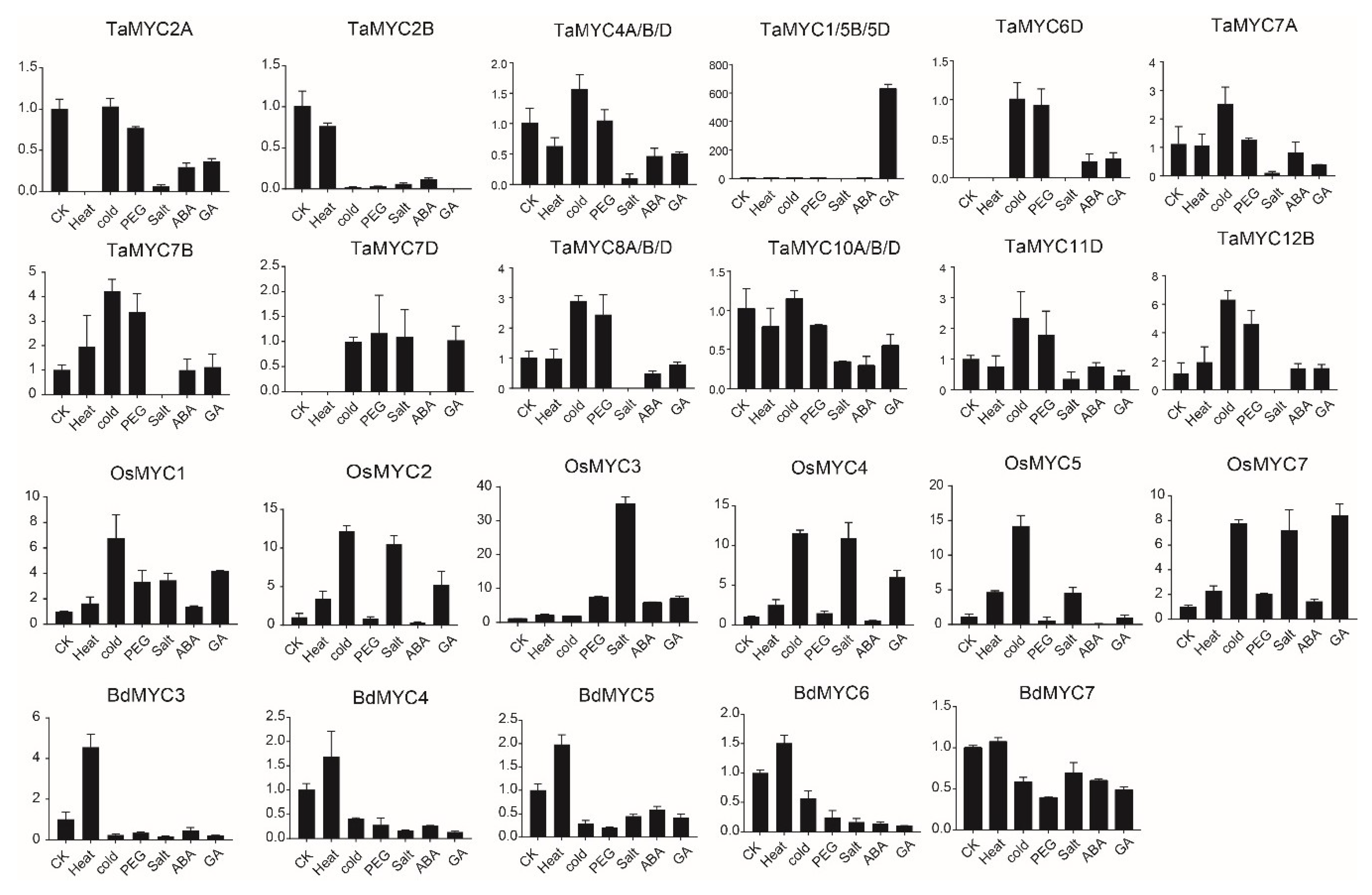
2.6. Expression Profiles of MYC Genes
3. Discussion
3.1. The Characteristics of MYCs
3.2. Functions of MYC TFs
4. Materials and Methods
4.1. Identification of MYC TFs in Wheat, Rice, and B. distachyon
4.2. Multiple Sequence Alignment and Phylogenetic Analysis of MYCs
4.3. Analysis of Gene Structures and Conserved Motifs
4.4. Analysis of Cis-Elements, Gene Duplication, and Synteny
4.5. Expression Profile Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Nuno, P.; Liam, D. Origin and diversification of basic-helix-loop-helix proteins in plants. Mol. Biol. Evol. 2010, 27, 862–874. [Google Scholar]
- Kazan, K.; Manners, J.M. MYC2: The Master in Action. Mol. Plant 2013, 6, 686–703. [Google Scholar] [CrossRef] [PubMed]
- Ledent, V.; Vervoort, M. The basic helix-loop-helix protein family: Comparative genomics and phylogenetic analysis. Genome Res. 2001, 11, 754–770. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.H.; Liao, Y.C.; Lv, F.F.; Zhang, Z.; Sun, P.W.; Gao, Z.H.; Hu, K.P.; Sui, C.; Jin, Y.; Wei, J.H. Transcription Factor AsMYC2 Controls the Jasmonate-responsive Expression of ASS1 Regulating Sesquiterpene Biosynthesis in Aquilaria sinensis (Lour.) Gilg. Plant Cell Physiol. 2017, 58, 1924–1933. [Google Scholar] [CrossRef] [PubMed]
- Oikawa, T.; Maeda, H.; Oguchi, T.; Yamaguchi, T.; Tanabe, N.; Ebana, K.; Yano, M.; Ebitani, T.; Izawa, T. The Birth of a Black Rice Gene and Its Local Spread by Introgression. Plant Cell 2015, 27, 2401–2414. [Google Scholar] [CrossRef] [PubMed]
- Tiancong, Q.; Jiaojiao, W.; Huang, H.; Bei, L.; Hua, G.; Yule, L.; Susheng, S.; Daoxin, X. Regulation of Jasmonate-Induced Leaf Senescence by Antagonism between bHLH Subgroup IIIe and IIId Factors in Arabidopsis. Plant Cell 2015, 27, 1634–1649. [Google Scholar]
- Gasperini, D.; Chételat, A.; Acosta, I.F.; Goossens, J.; Pauwels, L.; Goossens, A.; Dreos, R.; Alfonso, E.; Farmer, E.E. Multilayered Organization of Jasmonate Signalling in the Regulation of Root Growth. PLoS Genet. 2015, 11, e1005300. [Google Scholar] [CrossRef]
- Tiancong, Q.; Huang, H.; Susheng, S.; Daoxin, X. Regulation of Jasmonate-Mediated Stamen Development and Seed Production by a bHLH-MYB Complex in Arabidopsis. Plant Cell 2015, 27, 1620–1633. [Google Scholar]
- Gao, C.; Qi, S.; Liu, K.; Li, D.; Jin, C.; Li, Z.; Huang, G.; Hai, J.; Zhang, M.; Chen, M. MYC2, MYC3, and MYC4 function redundantly in seed storage protein accumulation in Arabidopsis. Plant Physiol. Biochem. 2016, 108, 63–70. [Google Scholar] [CrossRef]
- Zhu, X.; Chen, J.; Xie, Z.; Gao, J.; Ren, G.; Gao, S.; Zhou, X.; Kuai, B. Jasmonic acid promotes degreening via MYC2/3/4- and ANAC019/055/072-mediated regulation of major chlorophyll catabolic genes. Plant J. 2015, 84, 597–610. [Google Scholar] [CrossRef]
- Rajani, S.; Sundaresan, V. The Arabidopsis myc/bHLH gene ALCATRAZ enables cell separation in fruit dehiscence. Curr. Biol. 2001, 11, 1914–1922. [Google Scholar] [CrossRef]
- Heisler, M.; Atkinson, A.; Bylstra, Y.; Walsh, R.; Smyth, D. SPATULA, a gene that controls development of carpel margin tissues in Arabidopsis, encodes a bHLH protein. Development 2001, 128, 1089–1098. [Google Scholar] [PubMed]
- Anna-Marie, S.; Sandra, K.B.; Unte, U.S.; Peter, H.; Koen, D.; Heinz, S. The Arabidopsis Aborted MicrosporeS (AMS) gene encodes a MYC class transcription factor. Plant J. Cell Mol. Biol. 2010, 33, 413–423. [Google Scholar]
- Na, L.; Da-Sheng, Z.; Hai-Sheng, L.; Chang-Song, Y.; Xiao-Xing, L.; Wan-Qi, L.; Zheng, Y.; Ben, X.; Huang-Wei, C.; Jia, W. The rice tapetum degeneration retardation gene is required for tapetum degradation and anther development. Plant Cell 2006, 18, 2999–3014. [Google Scholar]
- Cai, Q.; Yuan, Z.; Chen, M.; Yin, C.; Luo, Z.; Zhao, X.; Liang, W.; Hu, J.; Zhang, D. Jasmonic acid regulates spikelet development in rice. Nat. Commun. 2014, 5, 3476. [Google Scholar] [CrossRef] [PubMed]
- Uji, Y.; Taniguchi, S.; Tamaoki, D.; Shishido, H.; Akimitsu, K.; Gomi, K. Overexpression of OsMYC2 Results in the Up-Regulation of Early JA-Rresponsive Genes and Bacterial Blight Resistance in Rice. Plant Cell Physiol. 2016, 57, 1814–1827. [Google Scholar] [CrossRef] [PubMed]
- Niu, Y.; Figueroa, P.; Browse, J. Characterization of JAZ-interacting bHLH transcription factors that regulate jasmonate responses in Arabidopsis. J. Exp. Bot. 2011, 62, 2143–2154. [Google Scholar] [CrossRef]
- Fabian, S.; Patricia, F.C.; Mark, Z.; Monica, D.D.; Sandra, F.; Gaétan, G.; Lewsey, M.G.; Ecker, J.R.; Roberto, S.; Philippe, R. Arabidopsis basic helix-loop-helix transcription factors MYC2, MYC3, and MYC4 regulate glucosinolate biosynthesis, insect performance, and feeding behavior. Plant Cell 2013, 25, 3117–3132. [Google Scholar]
- Zong, Y.; Xi, X.; Li, S.; Chen, W.; Zhang, B.; Liu, D.; Liu, B.; Wang, D.; Zhang, H. Allelic Variation and Transcriptional Isoforms of WheatTaMYC1Gene Regulating Anthocyanin Synthesis in Pericarp. Front. Plant Sci. 2017, 8, 1645. [Google Scholar] [CrossRef]
- Shoeva, O.Y.; Mock, H.P.; Kukoeva, T.V.; Börner, A.; Khlestkina, E.K. Regulation of the Flavonoid Biosynthesis Pathway Genes in Purple and Black Grains of Hordeum vulgare. PLoS ONE 2016, 11, e0163782. [Google Scholar] [CrossRef]
- Todd, A.T.; Enwu, L.; Polvi, S.L.; Pammett, R.T.; Page, J.E. A functional genomics screen identifies diverse transcription factors that regulate alkaloid biosynthesis in Nicotiana benthamiana. Plant J. 2010, 62, 589–600. [Google Scholar] [CrossRef] [PubMed]
- Yang, N.; Zhou, W.; Su, J.; Wang, X.; Li, L.; Wang, L.; Cao, X.; Wang, Z. Overexpression of SmMYC2 Increases the Production of Phenolic Acids in Salvia miltiorrhiza. Front. Plant Sci. 2017, 8, 1804. [Google Scholar] [CrossRef] [PubMed]
- Lenka, S.K.; Nims, N.E.; Vongpaseuth, K.; Boshar, R.A.; Roberts, S.C.; Walker, E.L. Jasmonate-responsive expression of paclitaxel biosynthesis genes in Taxus cuspidata cultured cells is negatively regulated by the bHLH transcription factors TcJAMYC1, TcJAMYC2, and TcJAMYC4. Front. Plant Sci. 2015, 6, 115. [Google Scholar] [CrossRef] [PubMed]
- Abe, H.; Yamaguchi-Shinozaki, K.; Urao, T.; Iwasaki, T.; Hosokawa, D.; Shinozaki, K. Role of arabidopsis MYC and MYB homologs in drought- and abscisic acid-regulated gene expression. Plant Cell 1997, 9, 1859–1868. [Google Scholar] [PubMed]
- Shinozaki, K.; Yamaguchi-Shinozaki, K. Gene networks involved in drought stress response and tolerance. J. Exp.Bot. 2007, 58, 221. [Google Scholar] [CrossRef]
- Ohta, M.; Sato, A.; Na, R. MYC-type transcription factors, MYC67 and MYC70, interact with ICE1 and negatively regulate cold tolerance in Arabidopsis. Sci. Rep. 2018, 8, 11622. [Google Scholar] [CrossRef]
- Lorenzo, O.; Chico JMSanchez-Serrano, J.J.; Solano, R. Jasmonate-insensitive1 encodes a MYC transcription factor essential to discriminate between different jasmonate-regulated defense responses in Arabidopsis. Plant Cell 2004, 16, 1938–1950. [Google Scholar] [CrossRef]
- Chini, A.; Fonseca, S.; Fernández, G.; Adie, B.; Chico, J.M.; Lorenzo, O.; García-Casado, G.; López-Vidriero, I.; Lozano, F.M.; Ponce, M.R. The JAZ family of repressors is the missing link in jasmonate signalling. Nature 2007, 448, 666–671. [Google Scholar] [CrossRef]
- Figueroa, P.; Browse, J.; Figueroa, P.; Browse, J. Male sterility in Arabidopsis induced by overexpression of a MYC5-SRDX chimeric repressor. Plant J. 2015, 81, 849–860. [Google Scholar] [CrossRef]
- Fernandez-Calvo, P.; Chini, A.; Fernandez-Barbero, G.; Chico, J.M.; Gimenez-Ibanez, S.; Geerinck, J.; Eeckhout, D.; Schweizer, F.; Godoy, M.; Franco-Zorrilla, J.M.; et al. The Arabidopsis bHLH transcription factors MYC3 and MYC4 are targets of JAZ repressors and act additively with MYC2 in the activation of jasmonate responses. Plant Cell 2011, 23, 701–715. [Google Scholar] [CrossRef]
- Ogawa, S.; Kawahara-Miki, R.; Miyamoto, K.; Yamane, H.; Nojiri, H.; Tsujii, Y.; Okada, K. OsMYC2 mediates numerous defence-related transcriptional changes via jasmonic acid signalling in rice. Biochem. Biophys. Res. Commun. 2017, 486, 796–803. [Google Scholar] [CrossRef] [PubMed]
- Wusirika, R.; Jorge, D.; Yong-Jin, P.; Carlos, B.; John, E.; Phillip, S.M.; Bennetzen, J.L. Different types and rates of genome evolution detected by comparative sequence analysis of orthologous segments from four cereal genomes. Genetics 2002, 162, 1389. [Google Scholar]
- Yu, C.S.; Chen, Y.C.; Hwang, J.K. Prediction of protein subcellular localization. Proteins-Struct. Funct. Bioinform. 2010, 64, 643–651. [Google Scholar] [CrossRef] [PubMed]
- Simionato, E.; Ledent, V.; Richards, G.; Thomas-Chollier, M.; Kerner, P.; Coornaert, D.; Degnan, B.M.; Vervoort, M. Origin and diversification of the basic helix-loop-helix gene family in metazoans: Insights from comparative genomics. BMC Evol. Biol. 2007, 7, 33. [Google Scholar] [CrossRef] [PubMed]
- Ledent, V.; Paquet, O.; Vervoort, M. Phylogenetic analysis of the human basic helix-loop-helix proteins. Genome Biol. 2002, 3, 1–18. [Google Scholar] [CrossRef]
- Zhang, D.; Li, G.; Wang, Y. A genome-wide identification and analysis of basic helix-loop-helix transcription factors in cattle. Gene 2017, 626, 241. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Song, S.; Huang, H.; Wang, J.; Qi, T.; Xie, D. MYC5 is Involved in Jasmonate-Regulated Plant Growth, Leaf Senescence and Defense Responses. Plant Cell Physiol. 2017, 58, 1752–1763. [Google Scholar]
- Urao, T.; Yamaguchishinozaki, K.; Mitsukawa, N.; Shibata, D.; Shinozaki, K. Molecular cloning and characterization of a gene that encodes a MYC-related protein in Arabidopsis. Plant Mol. Biol. 1996, 32, 571–576. [Google Scholar] [CrossRef]
- Benson, A.; GenBank, D. GenBank: update. Nucleic Acids Res. 2004, 32, D23–D26. [Google Scholar] [CrossRef]
- Bouchard, C.; Staller, P.; Eilers, M. Control of Cell Proliferation by Myc. Trends Cell Biol. 1998, 8, 202–206. [Google Scholar] [CrossRef]
- Xiaoxing, L.; Xuepeng, D.; Haixiong, J.; Yujin, S.; Yuanping, T.; Zheng, Y.; Jingkang, G.; Wanqi, L.; Liang, C.; Jingyuan, Y. Genome-wide analysis of basic/helix-loop-helix transcription factor family in rice and Arabidopsis. Plant Physiol. 2006, 141, 1167. [Google Scholar]
- Blackwood, E.; Eisenman, R. Max: A helix-loop-helix zipper protein that forms a sequence-specific DNA-binding complex with Myc. Science 1991, 251, 1211. [Google Scholar] [CrossRef] [PubMed]
- Niu, X.; Guan, Y.; Chen, S.; Li, H. Genome-wide analysis of basic helix-loop-helix (bHLH) transcription factors in Brachypodium distachyon. BMC Genom. 2017, 18, 619. [Google Scholar] [CrossRef] [PubMed]
- Yupeng, W.; Haibao, T.; Debarry, J.D.; Xu, T.; Jingping, L.; Xiyin, W.; Tae-Ho, L.; Huizhe, J.; Barry, M.; Hui, G. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar]
- Magali, L.; Patrice, D.; Gert, T.; Kathleen, M.; Yves, M.; Yves, V.D.P.; Pierre, R.; Stephane, R. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar]
- Giuliano, G.; Pichersky, E.; Malik, V.S.; Timko, M.P.; Scolnik, P.A.; Cashmore, A.R. An evolutionarily conserved protein binding sequence upstream of a plant light-regulated gene. Proc. Natl. Acad. Sci. USA 1988, 85, 7089–7093. [Google Scholar] [CrossRef] [PubMed]
- Haidar, M.A.; Henning, D.; Busch, H. Sp1 is essential and its position is important for p120 gene transcription: A 35 bp juxtaposed positive regulatory element enhances transcription 2.5 fold. Nucleic Acids Res. 1991, 19, 6559–6563. [Google Scholar] [CrossRef][Green Version]
- Carlini, L.E.; Ketudat, M.; Parsons, R.L.; Prabhakar, S.; Schmidt, R.J.; Guiltinan, M.J. The maize EmBP-1 orthologue differentially regulates Opaque2-dependent gene expression in yeast and cultured maize endosperm cells. Plant Mol. Biol. 1999, 41, 339–349. [Google Scholar] [CrossRef]
- Zhang, L.F.; Li, W.F.; Han, S.Y.; Yang, W.H.; Qi, L.W. cDNA cloning, genomic organization and expression analysis during somatic embryogenesis of the translationally controlled tumor protein (TCTP) gene from Japanese larch (Larix leptolepis). Gene 2013, 529, 150–158. [Google Scholar] [CrossRef]
- He, Y.; Gan, S. Identical promoter elements are involved in regulation of the OPR1 gene by senescence and jasmonic acid in Arabidopsis. Plant Mol. Biol. 2001, 47, 595–605. [Google Scholar] [CrossRef]
- Zheng, Z.; Xiaoming, Y.; Yaping, F.; Longfei, Z.; Hantian, W.; Xinchun, L. Overexpression of PvPin1, a Bamboo Homolog of PIN1-Type Parvulin 1, Delays Flowering Time in Transgenic Arabidopsis and Rice. Front. Plant Sci. 2017, 8, 1526. [Google Scholar] [CrossRef] [PubMed]
- Ezcurra, I.; Ellerström, M.; Wycliffe, P.; Stålberg, K.; Rask, L. Interaction between composite elements in the napA promoter: Both the B-box ABA-responsive complex and the RY/G complex are necessary for seed-specific expression. Plant Mol. Biol. 1999, 40, 699–709. [Google Scholar] [CrossRef] [PubMed]
- Sutoh, K.; Yamauchi, D. Two cis-acting elements necessary and sufficient for gibberellin-upregulated proteinase expression in rice seeds. Plant J. 2010, 34, 635–645. [Google Scholar] [CrossRef]
- Liu, Z.-B. Soybean GH3 promoter contains multiple auxin-inducible elements. Plant Cell 1994, 6, 645–657. [Google Scholar] [PubMed]
- Ballas, N.; Wong, L.-M.; Theologis, A. Identification of the Auxin-responsive Element, AuxRE, in the Primary indoleacetic Acid-inducible Gene, PS-IAA4/5, of Pea (Pisum sativum). J. Mol. Biol. 1993, 233, 580–596. [Google Scholar] [CrossRef] [PubMed]
- Goldsbrough, A.P.; Albrecht, H.; Stratford, R. Salicylic acid-inducible binding of a tobacco nuclear protein to a 10 bp sequence which is highly conserved amongst stress-inducible genes. Plant J. Cell Mol. Biol. 2010, 3, 563–571. [Google Scholar] [CrossRef]
- Zhang, Z.Y.; Zhao, J.; Hu, Y.; Zhang, T.Z. Isolation of GhMYB9 gene promoter and characterization of its activity in transgenic cotton. Biol. Plant. 2015, 59, 629–636. [Google Scholar] [CrossRef]
- Dunn, M.A.; White, A.J.; Vural, S.; Hughes, M.A. Identification of promoter elements in a low-temperature-responsive gene (blt4.9) from barley (Hordeum vulgare L.). Plant Mol. Biol. 1998, 38, 551–564. [Google Scholar] [CrossRef]
- Geffers, R.; Sell, S.; Cerff, R.; Hehl, R. The TATA box and a Myb binding site are essential for anaerobic expression of a maize GapC4 minimal promoter in tobacco. Biochim. Biophys. Acta 2001, 1521, 120–125. [Google Scholar] [CrossRef]
- Estes, K.S.; Anderson, D.G.; Stoler, D. Anoxic induction of a sarcoma virus-related VL30 retrotransposon is mediated by a cis-acting element which binds hypoxia-inducible factor 1 and an anoxia-inducible factor. J. Virol. 1995, 69, 6335–6341. [Google Scholar]
- Feller, A.; Machemer, K.; Braun, E.L.; Grotewold, E. Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. Plant J. Cell Mol. Biol. 2011, 66, 94–116. [Google Scholar] [CrossRef] [PubMed]
- Amoutzias, G.D.; Robertson, D.L.; Peer, Y.V.; Oliver, S.G. Choose your partners: Dimerization in eukaryotic transcription factors. Trends Biochem. Sci. 2008, 33, 220–229. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Jiang, H.; Li, L.; Zhai, Q.; Qi, L.; Zhou, W.; Liu, X.; Li, H.; Zheng, W.; Sun, J.; et al. The Arabidopsis Mediator Subunit MED25 Differentially Regulates Jasmonate and Abscisic Acid Signaling through Interacting with the MYC2 and ABI5 Transcription Factors. Plant Cell. 2012, 24, 2898–2916. [Google Scholar] [CrossRef] [PubMed]
- Baxevanis, A.D.; Vinson, C.R. Interactions of coiled coils in transcription factors: Where is the specificity? Curr. Opin. Genet. Dev. 1993, 3, 278–285. [Google Scholar] [CrossRef]
- Wei, K.; Chen, H. Comparative functional genomics analysis of bHLH gene family in rice, maize and wheat. BMC Plant Biol. 2018, 18, 309. [Google Scholar] [CrossRef] [PubMed]
- Bruno, D.; Ping, X.G.; Sprague, S.J.; Kirkegaard, J.A.; Ross, J.J.; Reid, J.B.; Fitt, G.P.; Nasser, S.; Schenk, P.M.; Manners, J.M. MYC2 differentially modulates diverse jasmonate-dependent functions in Arabidopsis. Plant Cell 2007, 19, 2225–2245. [Google Scholar]
- Hongtao, Z.; Sabah, H.; Grégory, M.; Yanxia, Z.; Guillaume, C.; Martial, P.; Pascal, G.; Johan, M. The basic helix-loop-helix transcription factor CrMYC2 controls the jasmonate-responsive expression of the ORCA genes that regulate alkaloid biosynthesis in Catharanthus roseus. Plant J. 2011, 67, 61–71. [Google Scholar]
- Dan, M.B.; Staines, D.M.; Perry, E.; Kersey, P.J. Ensembl Plants: Integrating Tools for Visualizing, Mining, and Analyzing Plant Genomic Data. Methods Mol. Biol. 2017, 1533, 1. [Google Scholar]
- Finn, R.D.; John, T.; Jaina, M.; Coggill, P.C.; Stephen John, S.; Hans-Rudolf, H.; Goran, C.; Kristoffer, F.; Eddy, S.R.; Sonnhammer, E.L.L. The Pfam protein families database. Nucleic Acids Res. 2008, 36, 281–288. [Google Scholar] [CrossRef]
- Panu, A.; Manohar, J.; Konstantin, A.; Delphine, B.; Gabor, C.; Edouard, D.C.; Séverine, D.; Volker, F.; Arnaud, F.; Elisabeth, G. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 2012, 40, W597. [Google Scholar]
- Johnson, M.; Zaretskaya, I.; Raytselis, Y.; Merezhuk, Y.; McGinnis, S.; Madden, T.L. NCBI BLAST: A better web interface. Nucleic Acids Res. 2008, 36, W5–W9. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.; Blackshields, G.; Brown, N.; Chenna, R.; Mcgettigan, P.; Mcwilliam, H.; Valentin, F.; Wallace, I.; Wilm, A.; Lopez, R.; et al. Clustal W and clustal X version 2.0. Bioinformatics 2007, 3, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, A.; Procter, J.; Martin, D.A.; Barton, G.J. Jalview: Visualization and Analysis of Molecular Sequences, Alignments, and Structures. BMC Bioinform. 2005, 6, 1. [Google Scholar] [CrossRef]
- He, Z.; Zhang, H.; Gao, S.; Lercher, M.J.; Chen, W.H.; Hu, S. Evolview v2: An online visualization and management tool for customized and annotated phylogenetic trees. Nucleic Acids Res. 2016, 44, W236–W241. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2014, 31, 1296. [Google Scholar] [CrossRef]
- Bailey, T.L.; Nadya, W.; Chris, M.; Li, W.W. MEME: Discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res. 2006, 34, 369–373. [Google Scholar] [CrossRef]
- Liu, C.; Xie, T.; Chen, C.; Luan, A.; Long, J.; Li, C.; Ding, Y.; He, Y. Genome-wide organization and expression profiling of the R2R3-MYB transcription factor family in pineapple (Ananas comosus). BMC Genom. 2017, 18, 503. [Google Scholar] [CrossRef]
- Hong, S.Y.; Seo, P.J.; Yang, M.-S.; Xiang, F.; Park, C.-M. Exploring valid reference genes for gene expression studies inBrachypodium distachyonby real-time PCR. BMC Plant Biol. 2008, 8, 112. [Google Scholar] [CrossRef]
- Ramesh, S.A.; Kamran, M.; Sullivan, W.; Chirkova, L.; Okamoto, M.; Degryse, F.; McLauchlin, M.; Gilliham, M.; Tyerman, S.D. Aluminium-Activated Malate Transporters Can Facilitate GABA Transport. Plant Cell 2018, 30, 1147–1164. [Google Scholar] [CrossRef]
- Kim, B.R.; Nam, H.-Y.; Kim, S.-U.; Kim, S.-I.; Chang, Y.-J. Normalization of reverse transcription quantitative-PCR with housekeeping genes in rice. Biotechnol. Lett. 2003, 25, 1869–1872. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔ C T Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]


© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, S.; Zhao, H.; Luo, T.; Liu, Y.; Nie, X.; Li, H. Characteristics and Expression Pattern of MYC Genes in Triticum aestivum, Oryza sativa, and Brachypodium distachyon. Plants 2019, 8, 274. https://doi.org/10.3390/plants8080274
Chen S, Zhao H, Luo T, Liu Y, Nie X, Li H. Characteristics and Expression Pattern of MYC Genes in Triticum aestivum, Oryza sativa, and Brachypodium distachyon. Plants. 2019; 8(8):274. https://doi.org/10.3390/plants8080274
Chicago/Turabian StyleChen, Shoukun, Hongyan Zhao, Tengli Luo, Yue Liu, Xiaojun Nie, and Haifeng Li. 2019. "Characteristics and Expression Pattern of MYC Genes in Triticum aestivum, Oryza sativa, and Brachypodium distachyon" Plants 8, no. 8: 274. https://doi.org/10.3390/plants8080274
APA StyleChen, S., Zhao, H., Luo, T., Liu, Y., Nie, X., & Li, H. (2019). Characteristics and Expression Pattern of MYC Genes in Triticum aestivum, Oryza sativa, and Brachypodium distachyon. Plants, 8(8), 274. https://doi.org/10.3390/plants8080274





