Exploring Microtubule-Dependent Cellulose-Synthase-Complex Movement with High Precision Particle Tracking
Abstract
1. Introduction
2. Results
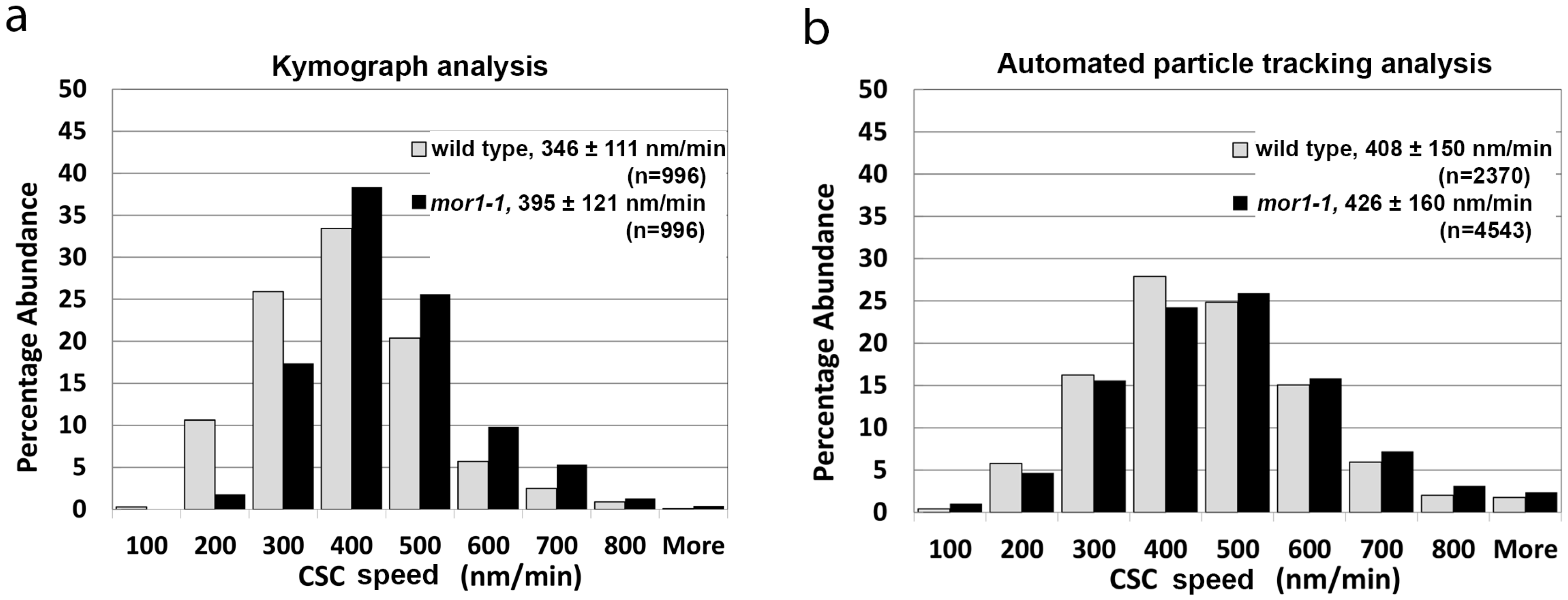
2.1. Automated Particle Tracking Quickly and Accurately Measures Fluorescent Protein Distribution and Dynamics
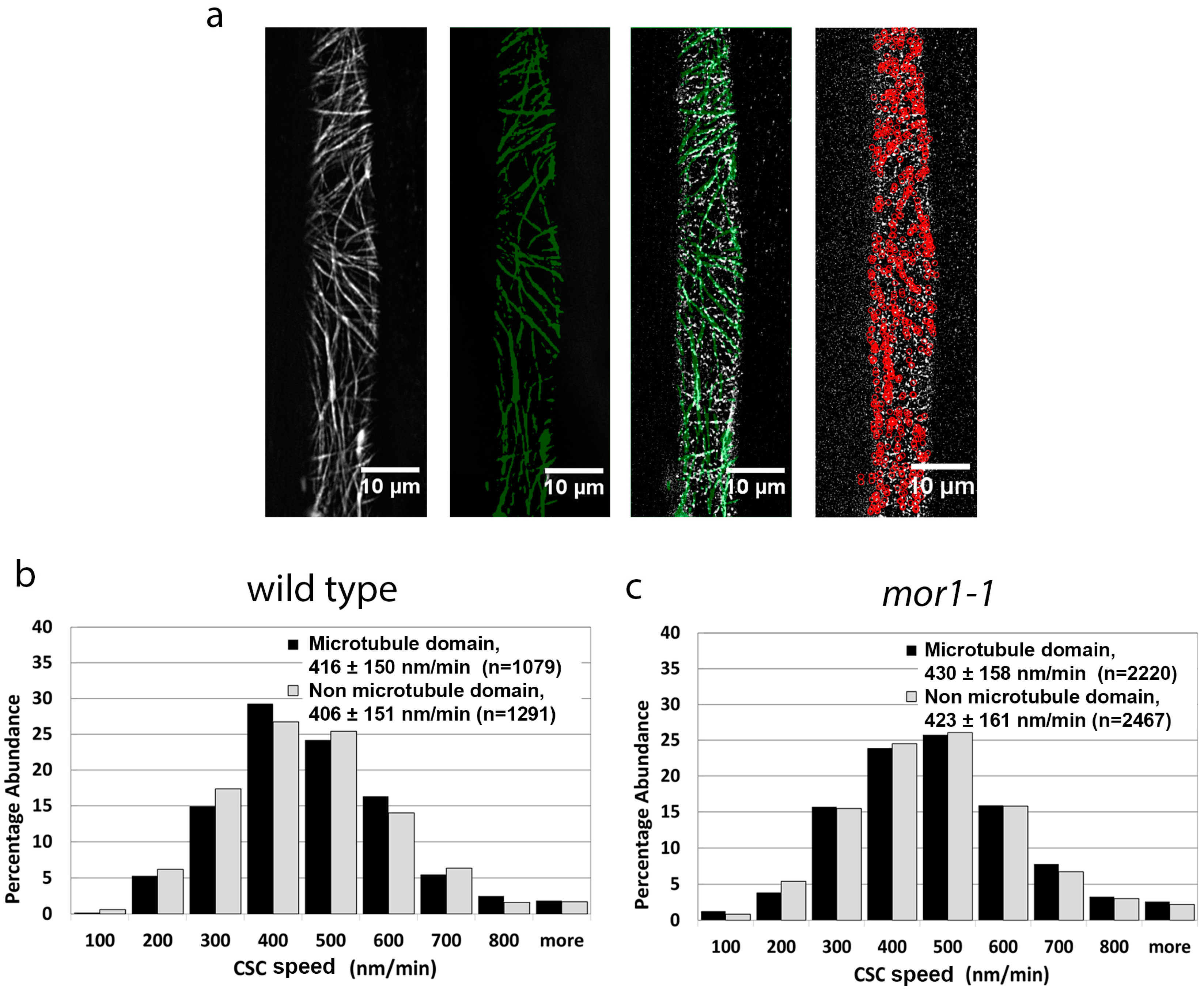
2.2. Cellulose Synthase Complex Speed Is Not Correlated with Proximity to Microtubules
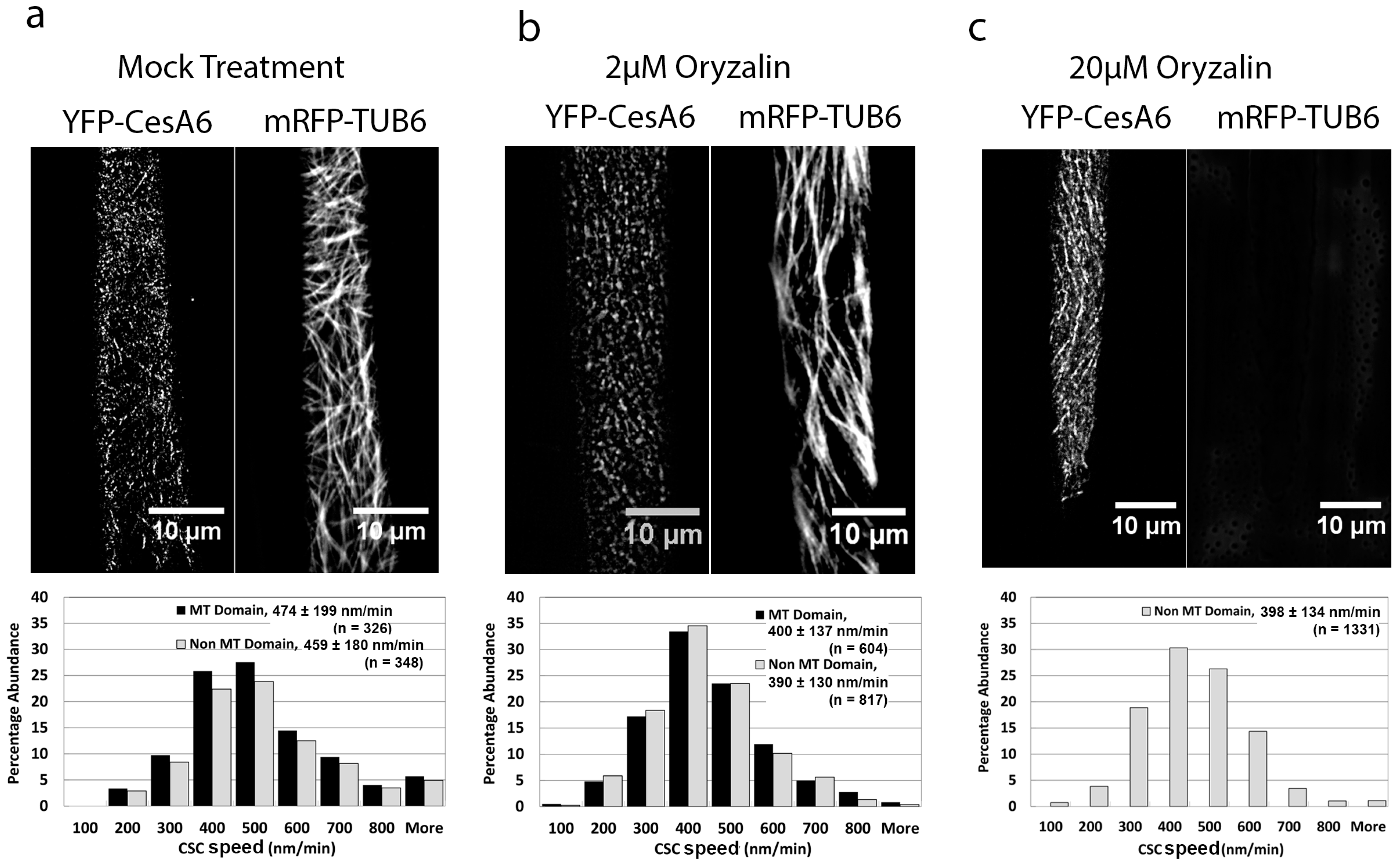
2.3. The Microtubule-Destabilizing Drug Oryzalin Reduces CSC Speed
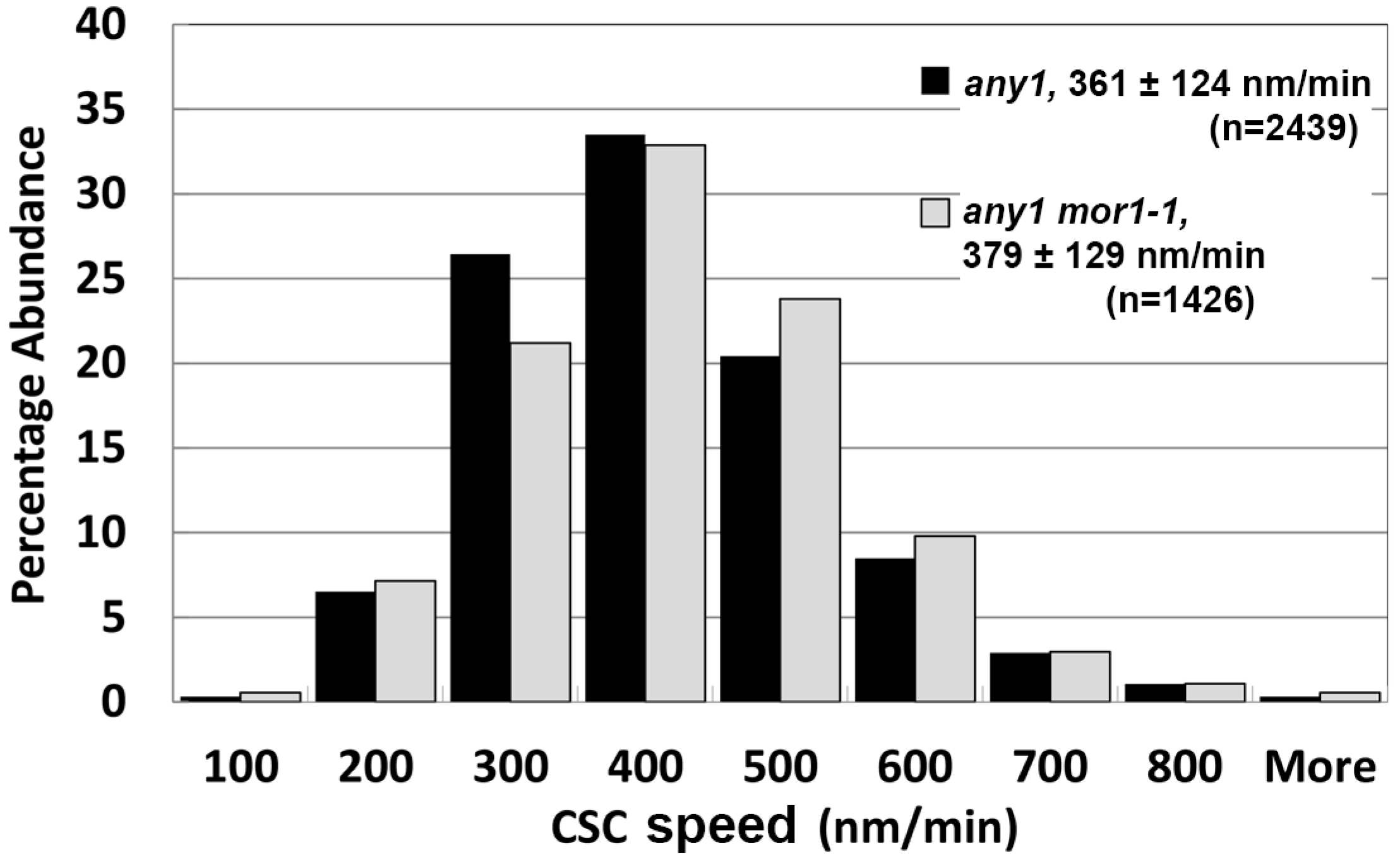
2.4. The mor1-1 Mutation Increases CSC Speed through a Mechanism That Is Independent of the CSC Catalytic Activity
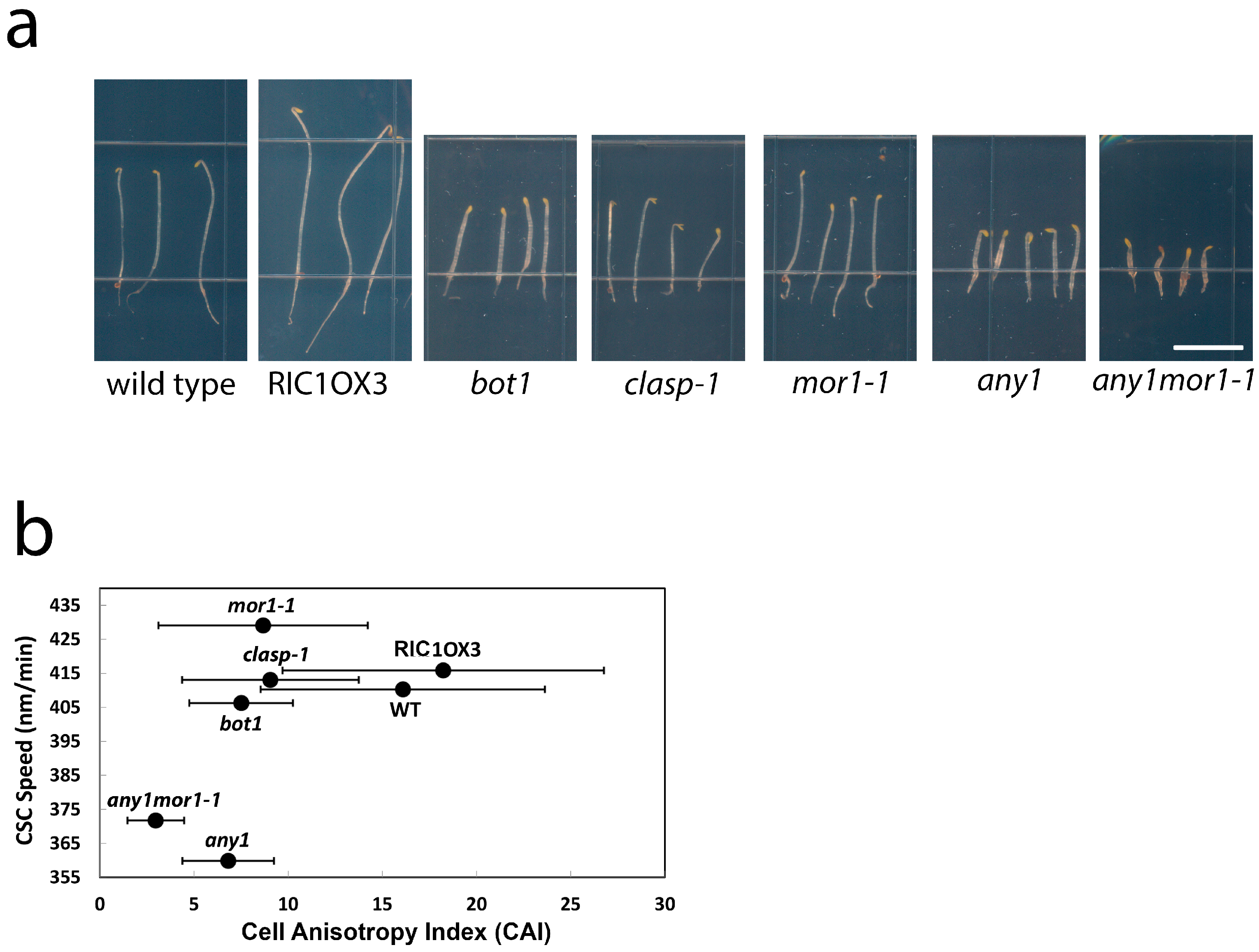
2.5. Moderate CSC Speed Is Required but Not Sufficient for Cell Growth Anisotropy
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Near-TIRF Imaging of CesAs
4.3. Image Processing
4.4. Particle Tracking
4.5. Cell Anisotropy Measurement
4.6. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Green, P.B. Mechanism for plant cellular morphogenesis. Science 1962, 138, 1404. [Google Scholar] [CrossRef] [PubMed]
- Fujita, M.; Himmelspach, R.; Hocart, C.H.; Williamson, R.E.; Mansfield, S.D.; Wasteneys, G.O. Cortical microtubules optimize cell-wall crystallinity to drive unidirectional growth in Arabidopsis. Plant J. 2011, 66, 915–928. [Google Scholar] [CrossRef] [PubMed]
- Fujita, M.; Himmelspach, R.; Ward, J.; Whittington, A.T.; Liu, C.; Truong, T.T.; Hasenbein, N.; Galway, M.E.; Mansfield, S.D.; Hocart, C.H.; Wasteneys, G.O. The any1 D604N mutation in the Arabidopsis thaliana cellulose synthase 1 catalytic domain reduces cell wall crystallinity and the velocity of cellulose synthase complexes. Plant Phys. 2013, 162, 74–85. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Rodriguez, C.; Bauer, S.; Hematy, K.; Saxe, F.; Ibanez, A.B.; Vodermaier, V.; Konlechner, C.; Sampathkumar, A.; Ruggeberg, M.; Aichinger, E.; et al. Chitinase-like1/pom-pom1 and its homolog ctl2 are glucan-interacting proteins important for cellulose biosynthesis in Arabidopsis. Plant Cell 2012, 24, 589–607. [Google Scholar] [CrossRef] [PubMed]
- Paredez, A.R.; Somerville, C.R.; Ehrhardt, D.W. Visualization of cellulose synthase demonstrates functional association with microtubules. Science 2006, 312, 1491–1495. [Google Scholar] [CrossRef] [PubMed]
- Desprez, T.; Juraniec, M.; Crowell, E.F.; Jouy, H.; Pochylova, Z.; Parcy, F.; Hofte, H.; Gonneau, M.; Vernhettes, S. Organization of cellulose synthase complexes involved in primary cell wall synthesis in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2007, 104, 15572–15577. [Google Scholar] [CrossRef] [PubMed]
- DeBolt, S.; Gutierrez, R.; Ehrhardt, D.W.; Melo, C.V.; Ross, L.; Cutler, S.R.; Somerville, C.; Bonetta, D. Morlin, an inhibitor of cortical microtubule dynamics and cellulose synthase movement. Proc. Natl. Acad. Sci. USA 2007, 104, 5854–5859. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Kaplinsky, N.; Bringmann, M.; Cobb, A.; Carroll, A.; Sampathkumar, A.; Baskin, T.I.; Persson, S.; Somerville, C.R. Identification of a cellulose synthase-associated protein required for cellulose biosynthesis. Proc. Natl. Acad. Sci. USA 2010, 107, 12866–12871. [Google Scholar] [CrossRef] [PubMed]
- Bringmann, M.; Li, E.; Sampathkumar, A.; Kocabek, T.; Hauser, M.T.; Persson, S. Pom-pom2/cellulose synthase interacting1 is essential for the functional association of cellulose synthase and microtubules in Arabidopsis. Plant Cell 2012, 24, 163–177. [Google Scholar] [CrossRef] [PubMed]
- Holman, L.; Head, M.L.; Lanfear, R.; Jennions, M.D. Evidence of experimental bias in the life sciences: Why we need blind data recording. PLoS Biol. 2015, 13, e1002190. [Google Scholar] [CrossRef] [PubMed]
- Mangeol, P.; Prevo, B.; Peterman, E.J.G. Kymographclear and kymographdirect: Two tools for the automated quantitative analysis of molecular and cellular dynamics using kymographs. Mol. Biol. Cell 2016, 27, 1948–1957. [Google Scholar] [CrossRef] [PubMed]
- De Chaumont, F.; Dallongeville, S.; Chenouard, N.; Herve, N.; Pop, S.; Provoost, T.; Meas-Yedid, V.; Pankajakshan, P.; Lecomte, T.; Le Montagner, Y.; et al. Icy: An open bioimage informatics platform for extended reproducible research. Nat. Methods 2012, 9, 690–696. [Google Scholar] [CrossRef] [PubMed]
- Kawamura, E.; Wasteneys, G.O. MOR1, the Arabidopsis thaliana homologue of Xenopus MAP215, promotes rapid growth and shrinkage, and suppresses the pausing of microtubules in vivo. J. Cell Sci. 2008, 121, 4114–4123. [Google Scholar] [CrossRef] [PubMed]
- Whittington, A.T.; Vugrek, O.; Wei, K.J.; Hasenbein, N.G.; Sugimoto, K.; Rashbrooke, M.C.; Wasteneys, G.O. MOR1 is essential for organizing cortical microtubules in plants. Nature 2001, 411, 610–613. [Google Scholar] [CrossRef] [PubMed]
- Fujita, M.; Lechner, B.; Barton, D.A.; Overall, R.L.; Wasteneys, G.O. The missing link: Do cortical microtubules define plasma membrane nanodomains that modulate cellulose biosynthesis? Protoplasma 2012, 249, S59–S67. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Gu, Y.; Zheng, Z.; Wasteneys, G.; Yang, Z. Arabidopsis interdigitating cell growth requires two antagonistic pathways with opposing action on cell morphogenesis. Cell 2005, 120, 687–700. [Google Scholar] [CrossRef] [PubMed]
- Bichet, A.; Desnos, T.; Turner, S.; Grandjean, O.; Hofte, H. Botero1 is required for normal orientation of cortical microtubules and anisotropic cell expansion in Arabidopsis. Plant J. 2001, 25, 137–148. [Google Scholar] [CrossRef] [PubMed]
- Ambrose, J.C.; Shoji, T.; Kotzer, A.M.; Pighin, J.A.; Wasteneys, G.O. The Arabidopsis CLASP gene encodes a microtubule-associated protein involved in cell expansion and division. Plant Cell 2007, 19, 2763–2775. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Lei, L.; Somerville, C.R.; Gu, Y. Cellulose synthase interactive protein 1 (CSI1) links microtubules and cellulose synthase complexes. Proc. Natl. Acad. Sci. USA 2012, 109, 185–190. [Google Scholar] [CrossRef] [PubMed]
- Rui, Y.; Anderson, C.T. Functional analysis of cellulose and xyloglucan in the walls of stomatal guard cells of Arabidopsis. Plant Physiol. 2016, 170, 1398–1419. [Google Scholar] [CrossRef] [PubMed]
- Ambrose, C.; Allard, J.F.; Cytrynbaum, E.N.; Wasteneys, G.O. A clasp-modulated cell edge barrier mechanism drives cell-wide cortical microtubule organization in Arabidopsis. Nat. Commun. 2011, 2, 430. [Google Scholar] [CrossRef] [PubMed]
- Vineyard, L.; Elliott, A.; Dhingra, S.; Lucas, J.R.; Shaw, S.L. Progressive transverse microtubule array organization in hormone-induced Arabidopsis hypocotyl cells. Plant Cell 2013, 25, 662–676. [Google Scholar] [CrossRef] [PubMed]
- Thevenaz, P.; Ruttimann, U.E.; Unser, M. A pyramid approach to subpixel registration based on intensity. IEEE Trans. Image Process 1998, 7, 27–41. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Woodley, M.; Mulvihill, A.; Fujita, M.; Wasteneys, G.O. Exploring Microtubule-Dependent Cellulose-Synthase-Complex Movement with High Precision Particle Tracking. Plants 2018, 7, 53. https://doi.org/10.3390/plants7030053
Woodley M, Mulvihill A, Fujita M, Wasteneys GO. Exploring Microtubule-Dependent Cellulose-Synthase-Complex Movement with High Precision Particle Tracking. Plants. 2018; 7(3):53. https://doi.org/10.3390/plants7030053
Chicago/Turabian StyleWoodley, Marcus, Adam Mulvihill, Miki Fujita, and Geoffrey O. Wasteneys. 2018. "Exploring Microtubule-Dependent Cellulose-Synthase-Complex Movement with High Precision Particle Tracking" Plants 7, no. 3: 53. https://doi.org/10.3390/plants7030053
APA StyleWoodley, M., Mulvihill, A., Fujita, M., & Wasteneys, G. O. (2018). Exploring Microtubule-Dependent Cellulose-Synthase-Complex Movement with High Precision Particle Tracking. Plants, 7(3), 53. https://doi.org/10.3390/plants7030053




