Phosphoregulation of the Plant Cellulose Synthase Complex and Cellulose Synthase-Like Proteins
Abstract
1. Introduction
2. The Cellulose Synthase Complex
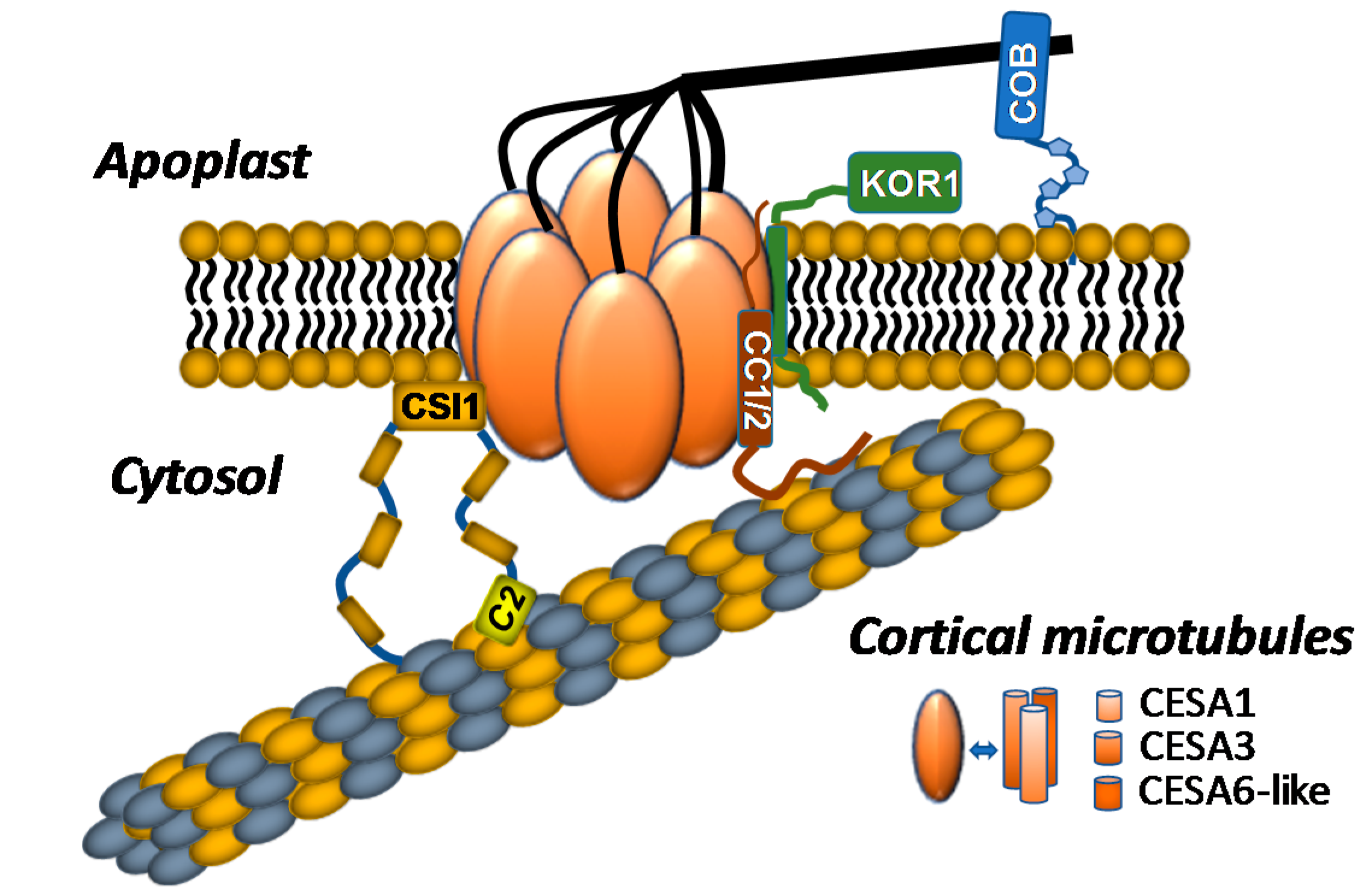
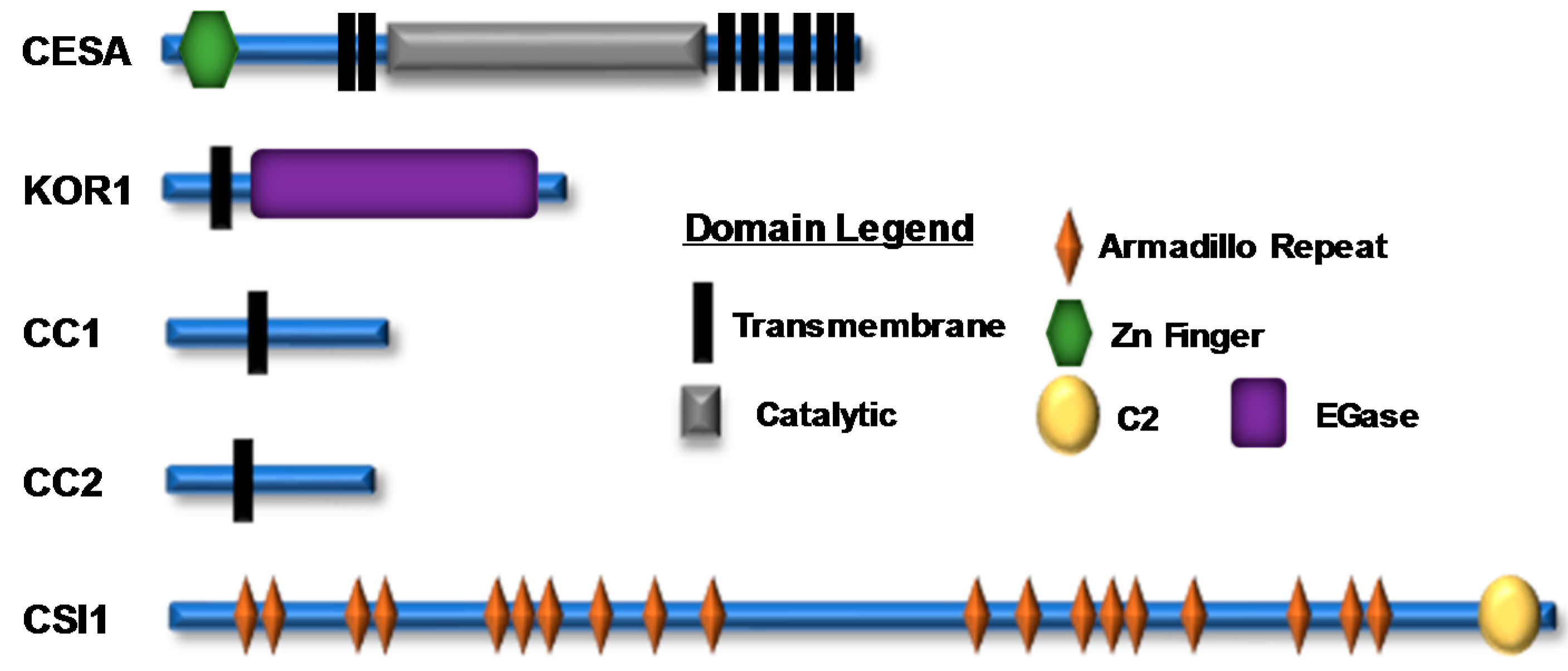
2.1. Composition of the Plant Cellulose Synthase Complex
2.2. Proteins Associated with the CSC
2.3. The In Vivo Dynamics of the Plant CSC
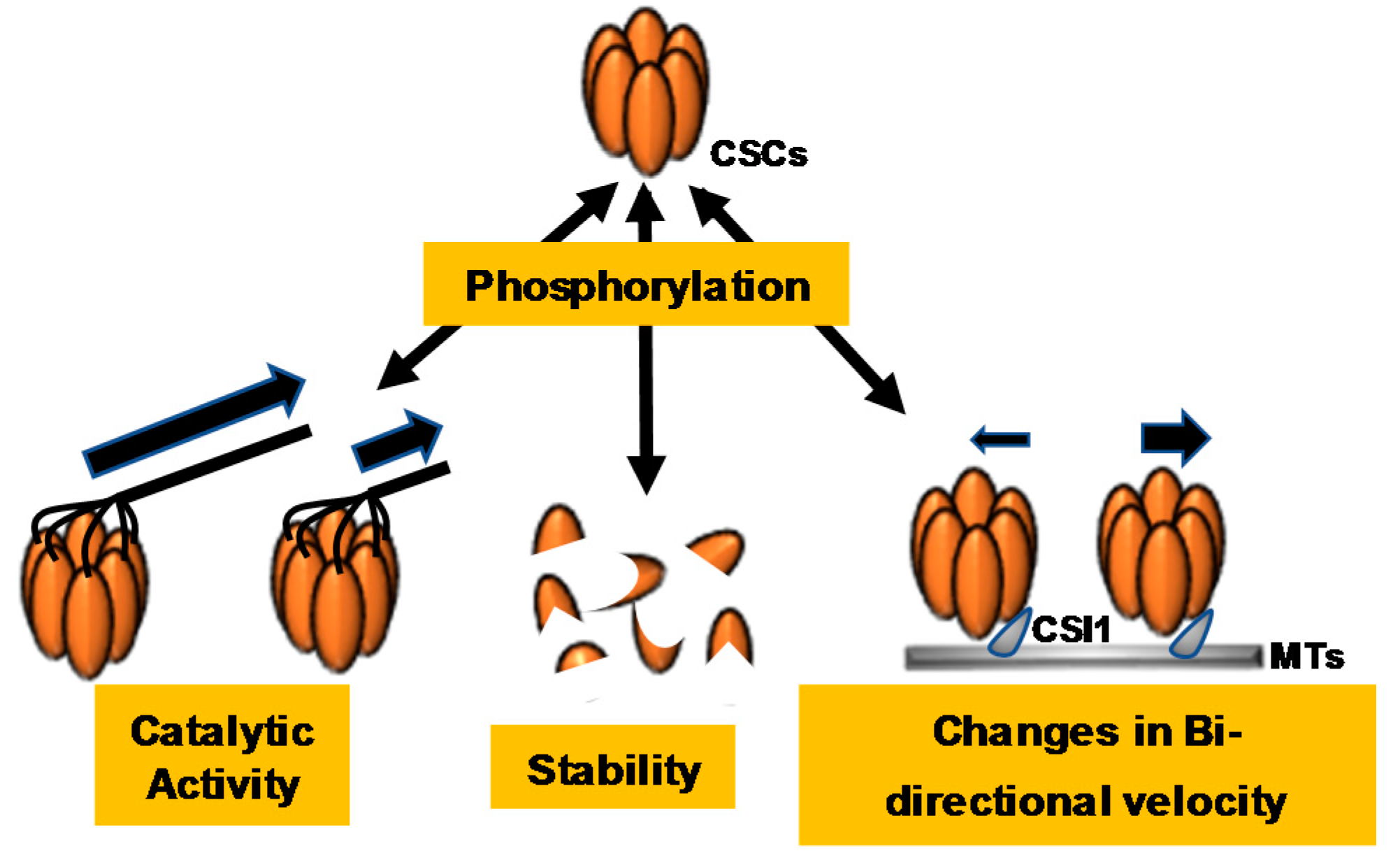
3. Phosphoregulation of CSC and CESA-Like Proteins
3.1. Identification and Characterization of CSC-Associated Phosphorylation Events
3.2. Brassinosteroid Regulation of the CSC and BIN2 Phoshorylation of CESA1
3.3. The Cellulose Synthase-Like Family of Glycosyltransferases
3.4. Evidence for CSL Phosphorylation
4. Receptor Kinases Implicated in Cellulose Biosynthesis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Somerville, C.; Bauer, S.; Brininstool, G.; Facette, M.; Hamann, T.; Milne, J.; Osborne, E.; Paredez, A.; Persson, S.; Raab, T.; et al. Toward a systems approach to understanding plant-cell walls. Science 2004, 306, 2206–2211. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, D.J. Growth of the plant cell wall. Nat. Rev. Mol. Cell Biol. 2005, 6, 850–861. [Google Scholar] [CrossRef] [PubMed]
- Burton, R.A.; Gidley, M.J.; Fincher, G.B. Heterogeneity in the chemistry, structure and function of plant cell walls. Nat. Chem. Biol. 2010, 6, 724–732. [Google Scholar] [CrossRef] [PubMed]
- Majda, M.; Grones, P.; Sintorn, I.M.; Vain, T.; Milani, P.; Krupinski, P.; Zagorska-Marek, B.; Viotti, C.; Jonsson, H.; Mellerowicz, E.J.; et al. Mechanochemical Polarization of Contiguous Cell Walls Shapes Plant Pavement Cells. Dev. Cell 2017, 43, 290–304. [Google Scholar] [CrossRef] [PubMed]
- Carter, R.; Woolfenden, H.; Baillie, A.; Amsbury, S.; Carroll, S.; Healicon, E.; Sovatzoglou, S.; Braybrook, S.; Gray, J.E.; Hobbs, J.; et al. Stomatal Opening Involves Polar, Not Radial, Stiffening of Guard Cells. Curr. Biol. 2017, 27, 2974–2983.e2. [Google Scholar] [CrossRef] [PubMed]
- Geitmann, A. Experimental approaches used to quantify physical parameters at cellular and subcellular levels. Am. J. Bot. 2006, 93, 1380–1390. [Google Scholar] [CrossRef] [PubMed]
- Geldner, N. The Endodermis. In Annual Review of Plant Biology; Merchant, S.S., Ed.; Annual Reviews: Palo Alto, CA, USA, 2013; Volume 64, pp. 531–558. ISBN 978-0-8243-0664-9. [Google Scholar]
- Li, Z.; Fernie, A.R.; Persson, S. Transition of primary to secondary cell wall synthesis. Sci. Bull. 2016, 61, 838–846. [Google Scholar] [CrossRef]
- Keegstra, K. Plant Cell Walls. Plant Physiol. 2010, 154, 483–486. [Google Scholar] [CrossRef] [PubMed]
- Turner, S.R.; Somerville, C.R. Collapsed xylem phenotype of Arabidopsis identifies mutants deficient in cellulose deposition in the secondary cell wall. Plant Cell 1997, 9, 689–701. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Omranian, N.; Neumetzler, L.; Wang, T.; Herter, T.; Usadel, B.; Demura, T.; Giavalisco, P.; Nikoloski, Z.; Persson, S. A Transcriptional and Metabolic Framework for Secondary Wall Formation in Arabidopsis. Plant Physiol. 2016, 172, 1334–1351. [Google Scholar] [CrossRef] [PubMed]
- Hill, J.L.; Hammudi, M.B.; Tien, M. The Arabidopsis Cellulose Synthase Complex: A Proposed Hexamer of CESA Trimers in an Equimolar Stoichiometry. Plant Cell 2014, 26, 4834–4842. [Google Scholar] [CrossRef] [PubMed]
- McFarlane, H.E.; Doring, A.; Persson, S. The Cell Biology of Cellulose Synthesis. Annu. Rev. Plant Biol. 2014, 65, 69–94. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, D.J. Wall extensibility: Its nature, measurement and relationship to plant cell growth. New Phytol. 1993, 124, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Scheller, H. V.; Ulvskov, P. Hemicelluloses. Annu. Rev. Plant Biol. 2010, 61, 263–289. [Google Scholar] [CrossRef] [PubMed]
- Scanlon, M.; Timmermans, M. Growth and development: From genes to networks and a mechanistic understanding of plant development. Curr. Opin. Plant Biol. 2013, 16, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Miart, F.; Desprez, T.; Biot, E.; Morin, H.; Belcram, K.; Hofte, H.; Gonneau, M.; Vernhettes, S. Spatio-temporal analysis of cellulose synthesis during cell plate formation in Arabidopsis. Plant J. 2014, 77, 71–84. [Google Scholar] [CrossRef] [PubMed]
- Drakakaki, G. Polysaccharide deposition during cytokinesis: Challenges and future perspectives. Plant Sci. 2015, 236, 177–184. [Google Scholar] [CrossRef] [PubMed]
- Fujimoto, M.; Suda, Y.; Vernhettes, S.; Nakano, A.; Ueda, T. Phosphatidylinositol 3-Kinase and 4-Kinase Have Distinct Roles in Intracellular Trafficking of Cellulose Synthase Complexes in Arabidopsis thaliana. Plant Cell Physiol. 2015, 56, 287–298. [Google Scholar] [CrossRef] [PubMed]
- Lane, D.R.; Wiedemeier, A.; Peng, L.C.; Hofte, H.; Vernhettes, S.; Desprez, T.; Hocart, C.H.; Birch, R.J.; Baskin, T.I.; Burn, J.E.; et al. Temperature-sensitive alleles of RSW2 link the KORRIGAN endo-1,4-beta-glucanase to cellulose synthesis and cytokinesis in arabidopsis. Plant Physiol. 2001, 126, 278–288. [Google Scholar] [CrossRef] [PubMed]
- Beeckman, T.; Przemeck, G.K.H.; Stamatiou, G.; Lau, R.; Terryn, N.; De Rycke, R.; Inze, D.; Berleth, T. Genetic complexity of cellulose synthase A gene function in Arabidopsis embryogenesis. Plant Physiol. 2002, 130, 1883–1893. [Google Scholar] [CrossRef] [PubMed]
- Bringmann, M.; Li, E.Y.; Sampathkumar, A.; Kocabek, T.; Hauser, M.T.; Persson, S. POM-POM2/CELLULOSE SYNTHASE INTERACTING1 Is Essential for the Functional Association of Cellulose Synthase and Microtubules in Arabidopsis. Plant Cell 2012, 24, 163–177. [Google Scholar] [CrossRef] [PubMed]
- Paredez, A.R.; Somerville, C.R.; Ehrhardt, D.W. Visualization of cellulose synthase demonstrates functional association with microtubules. Science 2006, 312, 1491–1495. [Google Scholar] [CrossRef] [PubMed]
- Kesten, C.; Menna, A.; Sanchez-Rodriguez, C. Regulation of cellulose synthesis in response to stress. Curr. Opin. Plant Biol. 2017, 40, 106–113. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; McFarlane, H.E.; Persson, S. The impact of abiotic factors on cellulose synthesis. J. Exp. Bot. 2016, 67, 543–552. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.S.; Frank, J.; Kang, C.H.; Kajiura, H.; Vikram, M.; Ueda, A.; Kim, S.; Bahk, J.D.; Triplett, B.; Fujiyama, K.; et al. Salt tolerance of Arabidopsis thaliana requires maturation of N-glycosylated proteins in the Golgi apparatus. Proc. Natl. Acad. Sci. USA 2008, 105, 5933–5938. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Rodriguez, C.; Bauer, S.; Hematy, K.; Saxe, F.; Ibanez, A.B.; Vodermaier, V.; Konlechner, C.; Sampathkumar, A.; Ruggeberg, M.; Aichinger, E.; et al. CHITINASE-LIKE1/POM-POM1 and Its Homolog CTL2 Are Glucan-Interacting Proteins Important for Cellulose Biosynthesis in Arabidopsis. Plant Cell 2012, 24, 589–607. [Google Scholar] [CrossRef] [PubMed]
- Endler, A.; Kesten, C.; Schneider, R.; Zhang, Y.; Ivakov, A.; Froehlich, A.; Funke, N.; Persson, S. A Mechanism for Sustained Cellulose Synthesis during Salt Stress. Cell 2015, 162, 1353–1364. [Google Scholar] [CrossRef] [PubMed]
- Barnes, W.J.; Anderson, C.T. Release, Recycle, Rebuild: Cell-Wall Remodeling, Autodegradation, and Sugar Salvage for New Wall Biosynthesis during Plant Development. Mol. Plant 2018, 11, 31–46. [Google Scholar] [CrossRef] [PubMed]
- Verbancic, J.; Lunn, J.E.; Stitt, M.; Persson, S. Carbon Supply and the Regulation of Cell Wall Synthesis. Mol. Plant 2018, 11, 75–94. [Google Scholar] [CrossRef] [PubMed]
- McNamara, J.T.; Morgan, J.L.W.; Zimmer, J. A Molecular Description of Cellulose Biosynthesis. Annu. Rev. Biochem. 2015, 84, 895–921. [Google Scholar] [CrossRef] [PubMed]
- Wolf, S.; Hematy, K.; Hofte, H. Growth Control and Cell Wall Signaling in Plants. Annu. Rev. Plant Biol. 2012, 63, 381–407. [Google Scholar] [CrossRef] [PubMed]
- Mueller, S.C.; Brown, R.M. Evidence for an Intramembrane Component Associated with a Cellulose Microfibril-Synthesizing Complex in Higher-Plants. J. Cell Biol. 1980, 84, 315–326. [Google Scholar] [CrossRef] [PubMed]
- Kimura, S.; Laosinchai, W.; Itoh, T.; Cui, X.J.; Linder, C.R.; Brown, R.M. Immunogold labeling of rosette terminal cellulose-synthesizing complexes in the vascular plant Vigna angularis. Plant Cell 1999, 11, 2075–2085. [Google Scholar] [CrossRef] [PubMed]
- Purushotham, P.; Cho, S.H.; Diaz-Moreno, S.M.; Kumar, M.; Nixon, B.T.; Bulone, V.; Zimmer, J. A single heterologously expressed plant cellulose synthase isoform is sufficient for cellulose microfibril formation in vitro. Proc. Natl. Acad. Sci. USA 2016, 113, 11360–11365. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.H.; Purushotham, P.; Fang, C.; Maranas, C.; Diaz-Moreno, S.M.; Bulone, V.; Zimmer, J.; Kumar, M.; Nixon, B.T. Synthesis and Self-Assembly of Cellulose Microfibrils from Reconstituted Cellulose Synthase. Plant Physiol. 2017, 175, 146–156. [Google Scholar] [CrossRef] [PubMed]
- Persson, S.; Paredez, A.; Carroll, A.; Palsdottir, H.; Doblin, M.; Poindexter, P.; Khitrov, N.; Auer, M.; Somerville, C.R. Genetic evidence for three unique components in primary cell-wall cellulose synthase complexes in Arabidopsis. Proc. Natl. Acad. Sci. USA 2007, 104, 15566–15571. [Google Scholar] [CrossRef] [PubMed]
- Desprez, T.; Juraniec, M.; Crowell, E.F.; Jouy, H.; Pochylova, Z.; Parcy, F.; Hofte, H.; Gonneau, M.; Vernhettes, S. Organization of cellulose synthase complexes involved in primary cell wall synthesis in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2007, 104, 15572–15577. [Google Scholar] [CrossRef] [PubMed]
- Taylor, N.G.; Howells, R.M.; Huttly, A.K.; Vickers, K.; Turner, S.R. Interactions among three distinct CesA proteins essential for cellulose synthesis. Proc. Natl. Acad. Sci. USA 2003, 100, 1450–1455. [Google Scholar] [CrossRef] [PubMed]
- Gonneau, M.; Desprez, T.; Guillot, A.; Vernhettes, S.; Hofte, H. Catalytic Subunit Stoichiometry within the Cellulose Synthase Complex. Plant Physiol. 2014, 166, 1709–1712. [Google Scholar] [CrossRef] [PubMed]
- Vandavasi, V.G.; Putnam, D.K.; Zhang, Q.; Petridis, L.; Heller, W.T.; Nixon, B.T.; Haigler, C.H.; Kalluri, U.; Coates, L.; Langan, P.; et al. A Structural Study of CESA1 Catalytic Domain of Arabidopsis Cellulose Synthesis Complex: Evidence for CESA Trimers. Plant Physiol. 2016, 170, 123–135. [Google Scholar] [CrossRef] [PubMed]
- Nixon, B.T.; Mansouri, K.; Singh, A.; Du, J.; Davis, J.K.; Lee, J.G.; Slabaugh, E.; Vandavasi, V.G.; O’Neill, H.; Roberts, E.M.; et al. Comparative Structural and Computational Analysis Supports Eighteen Cellulose Synthases in the Plant Cellulose Synthesis Complex. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Crowell, E.F.; Bischoff, V.; Desprez, T.; Rolland, A.; Stierhof, Y.D.; Schumacher, K.; Gonneau, M.; Hofte, H.; Vernhettes, S. Pausing of Golgi Bodies on Microtubules Regulates Secretion of Cellulose Synthase Complexes in Arabidopsis. Plant Cell 2009, 21, 1141–1154. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, R.; Lindeboom, J.J.; Paredez, A.R.; Emons, A.M.C.; Ehrhardt, D.W. Arabidopsis cortical microtubules position cellulose synthase delivery to the plasma membrane and interact with cellulose synthase trafficking compartments. Nat. Cell Biol. 2009, 11, 797–806. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Kaplinsky, N.; Bringmann, M.; Cobb, A.; Carroll, A.; Sampathkumar, A.; Baskin, T.I.; Persson, S.; Somerville, C.R. Identification of a cellulose synthase-associated protein required for cellulose biosynthesis. Proc. Natl. Acad. Sci. USA 2010, 107, 12866–12871. [Google Scholar] [CrossRef] [PubMed]
- Diotallevi, F.; Mulder, B. The cellulose synthase complex: A polymerization driven supramolecular motor. Biophys. J. 2007, 92, 2666–2673. [Google Scholar] [CrossRef] [PubMed]
- DeBolt, S.; Gutierrez, R.; Ehrhardt, D.W.; Somerville, C. Nonmotile cellulose synthase subunits repeatedly accumulate within localized regions at the plasma membrane in Arabidopsis hypocotyl cells following 2,6-dichlorobenzonitrile treatment. Plant Physiol. 2007, 145, 334–338. [Google Scholar] [CrossRef] [PubMed]
- Paredez, A.R.; Persson, S.; Ehrhardt, D.W.; Somerville, C.R. Genetic evidence that cellulose synthase activity influences microtubule cortical array organization. Plant Physiol. 2008, 147, 1723–1734. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Li, S.; Pan, S.; Xin, X.; Gu, Y. CSI1, PATROL1, and exocyst complex cooperate in delivery of cellulose synthase complexes to the plasma membrane. Proc. Natl. Acad. Sci. USA 2018. [Google Scholar] [CrossRef] [PubMed]
- Persson, S.; Wei, H.R.; Milne, J.; Page, G.P.; Somerville, C.R. Identification of genes required for cellulose synthesis by regression analysis of public microarray data sets. Proc. Natl. Acad. Sci. USA 2005, 102, 8633–8638. [Google Scholar] [CrossRef] [PubMed]
- Brown, D.M.; Zeef, L.A.H.; Ellis, J.; Goodacre, R.; Turner, S.R. Identification of novel genes in Arabidopsis involved in secondary cell wall formation using expression profiling and reverse genetics. Plant Cell 2005, 17, 2281–2295. [Google Scholar] [CrossRef] [PubMed]
- Sato, S.; Kato, T.; Kakegawa, K.; Ishii, T.; Liu, Y.G.; Awano, T.; Takabe, K.; Nishiyama, Y.; Kuga, S.; Nakamura, Y.; et al. Role of the putative membrane-bound endo-1,4-beta-glucanase KORRIGAN in cell elongation and cellulose synthesis in Arabidopsis thaliana. Plant Cell Physiol. 2001, 42, 251–263. [Google Scholar] [CrossRef] [PubMed]
- Vain, T.; Crowell, E.F.; Timpano, H.; Biot, E.; Desprez, T.; Mansoori, N.; Trindade, L.M.; Pagant, S.; Robert, S.; Hofte, H.; et al. The Cellulase KORRIGAN Is Part of the Cellulose Synthase Complex. Plant Physiol. 2014, 165, 1521–1532. [Google Scholar] [CrossRef] [PubMed]
- Nicol, F.; His, I.; Jauneau, A.; Vernhettes, S.; Canut, H.; Hofte, H. A plasma membrane-bound putative endo-1,4-beta-d-glucanase is required for normal wall assembly and cell elongation in Arabidopsis. Embo J. 1998, 17, 5563–5576. [Google Scholar] [CrossRef] [PubMed]
- Lei, L.; Zhang, T.; Strasser, R.; Lee, C.M.; Gonneau, M.; Mach, L.; Vernhettes, S.; Kim, S.H.; Cosgrove, D.J.; Li, S.D.; et al. The jiaoyao1 Mutant Is an Allele of korrigan1 That Abolishes Endoglucanase Activity and Affects the Organization of Both Cellulose Microfibrils and Microtubules in Arabidopsis. Plant Cell 2014, 26, 2601–2616. [Google Scholar] [CrossRef] [PubMed]
- Mansoori, N.; Timmers, J.; Desprez, T.; Kamei, C.L.A.; Dees, D.C.T.; Vincken, J.P.; Visser, R.G.F.; Hofte, H.; Vernhettes, S.; Trindade, L.M. KORRIGAN1 Interacts Specifically with Integral Components of the Cellulose Synthase Machinery. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Li, S.D.; Lei, L.; Somerville, C.R.; Gu, Y. Cellulose synthase interactive protein 1 (CSI1) links microtubules and cellulose synthase complexes. Proc. Natl. Acad. Sci. USA 2012, 109, 185–190. [Google Scholar] [CrossRef] [PubMed]
- Lei, L.; Singh, A.; Bashline, L.; Li, S.D.; Yingling, Y.G.; Gu, Y. CELLULOSE SYNTHASE INTERACTIVE1 Is Required for Fast Recycling of Cellulose Synthase Complexes to the Plasma Membrane in Arabidopsis. Plant Cell 2015, 27, 2926–2940. [Google Scholar] [CrossRef] [PubMed]
- Schindelman, G.; Morikami, A.; Jung, J.; Baskin, T.I.; Carpita, N.C.; Derbyshire, P.; McCann, M.C.; Benfey, P.N. COBRA encodes a putative GPI-anchored protein, which is polarly localized and necessary for oriented cell expansion in Arabidopsis. Genes Dev. 2001, 15, 1115–1127. [Google Scholar] [CrossRef] [PubMed]
- Benfey, P.N.; Linstead, P.J.; Roberts, K.; Schiefelbein, J.W.; Hauser, M.T.; Aeschbacher, R.A. Root Development In Arabidopsis—4 Mutants With Dramatically Altered Root Morphogenesis. Development 1993, 119, 57–70. [Google Scholar] [PubMed]
- Brady, S.M.; Orlando, D.A.; Lee, J.Y.; Wang, J.Y.; Koch, J.; Dinneny, J.R.; Mace, D.; Ohler, U.; Benfey, P.N. A high-resolution root spatiotemporal map reveals dominant expression patterns. Science 2007, 318, 801–806. [Google Scholar] [CrossRef] [PubMed]
- Ben-Tov, D.; Abraham, Y.; Stav, S.; Thompson, K.; Loraine, A.; Elbaum, R.; de Souza, A.; Pauly, M.; Kieber, J.J.; Harpaz-Saad, S. COBRA-LIKE2, a Member of the Glycosylphosphatidylinositol-Anchored COBRA-LIKE Family, Plays a Role in Cellulose Deposition in Arabidopsis Seed Coat Mucilage Secretory Cells. Plant Physiol. 2015, 167, 711–724. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.F.; Shang-Guan, K.K.; Zhang, B.C.; Liu, X.L.; Yan, M.X.; Zhang, L.J.; Shi, Y.Y.; Zhang, M.; Qian, Q.; Li, J.Y.; et al. Brittle Culm1, a COBRA-Like Protein, Functions in Cellulose Assembly through Binding Cellulose Microfibrils. PLoS Genet. 2013, 9. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.J.; Mao, Y.F.; Ha, S.; Liu, W.S.; Botella, J.R.; Zhu, J.K. A multiplex CRISPR/Cas9 platform for fast and efficient editing of multiple genes in Arabidopsis. Plant Cell Rep. 2016, 35, 1519–1533. [Google Scholar] [CrossRef] [PubMed]
- Sampathkumar, A.; Gutierrez, R.; McFarlane, H.E.; Bringmann, M.; Lindeboom, J.; Emons, A.M.; Samuels, L.; Ketelaar, T.; Ehrhardt, D.W.; Persson, S. Patterning and Lifetime of Plasma Membrane-Localized Cellulose Synthase Is Dependent on Actin Organization in Arabidopsis Interphase Cells. Plant Physiol. 2013, 162, 675–688. [Google Scholar] [CrossRef] [PubMed]
- Breuer, D.; Nowak, J.; Ivakov, A.; Somssich, M.; Persson, S.; Nikoloski, Z. System-wide organization of actin cytoskeleton determines organelle transport in hypocotyl plant cells. Proc. Natl. Acad. Sci. USA 2017, 114, E5741–E5749. [Google Scholar] [CrossRef] [PubMed]
- Nuhse, T.S.; Stensballe, A.; Jensen, O.N.; Peck, S.C. Phosphoproteomics of the arabidopsis plasma membrane and a new phosphorylation site database. Plant Cell 2004, 16, 2394–2405. [Google Scholar] [CrossRef] [PubMed]
- Nakagami, H.; Sugiyama, N.; Mochida, K.; Daudi, A.; Yoshida, Y.; Toyoda, T.; Tomita, M.; Ishihama, Y.; Shirasu, K. Large-Scale Comparative Phosphoproteomics Identifies Conserved Phosphorylation Sites in Plants. Plant Physiol. 2010, 153, 1161–1174. [Google Scholar] [CrossRef] [PubMed]
- Facette, M.R.; Shen, Z.X.; Bjornsdottir, F.R.; Briggs, S.P.; Smith, L.G. Parallel Proteomic and Phosphoproteomic Analyses of Successive Stages of Maize Leaf Development. Plant Cell 2013, 25, 2798–2812. [Google Scholar] [CrossRef] [PubMed]
- Taylor, N.G. Identification of cellulose synthase AtCesA7 (IRX3) in vivo phosphorylation sites—A potential role in regulating protein degradation. Plant Mol. Biol. 2007, 64, 161–171. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Rodriguez, C.; Ketelaar, K.; Schneider, R.; Villalobos, J.A.; Somerville, C.R.; Persson, S.; Wallace, I.S. BRASSINOSTEROID INSENSITIVE2 negatively regulates cellulose synthesis in Arabidopsis by phosphorylating cellulose synthase 1. Proc. Natl. Acad. Sci. USA 2017, 114, 3533–3538. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.L.; Ehrhardt, D.W.; Somerville, C.R. Mutations of cellulose synthase (CESA1) phosphorylation sites modulate anisotropic cell expansion and bidirectional mobility of cellulose synthase. Proc. Natl. Acad. Sci. USA 2010, 107, 17188–17193. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.L.; Jia, H.L.; Zhao, H.Y.; Liu, D.; Liu, Y.M.; Liu, B.Y.; Bauer, S.; Somerville, C.R. Anisotropic Cell Expansion Is Affected through the Bidirectional Mobility of Cellulose Synthase Complexes and Phosphorylation at Two Critical Residues on CESA3. Plant Physiol. 2016, 171, 242–250. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, Y.; Meents, M.J.; McDonnell, L.M.; Barkwill, S.; Sampathkumar, A.; Cartwright, H.N.; Demura, T.; Ehrhardt, D.W.; Samuels, A.L.; Mansfield, S.D. Visualization of cellulose synthases in Arabidopsis secondary cell walls. Science 2015, 350, 198–203. [Google Scholar] [CrossRef] [PubMed]
- Bischoff, V.; Desprez, T.; Mouille, G.; Vernhettes, S.; Gonneau, M.; Hofte, H. Phytochrome Regulation of Cellulose Synthesis in Arabidopsis. Curr. Biol. 2011, 21, 1822–1827. [Google Scholar] [CrossRef] [PubMed]
- Fagard, M.; Desnos, T.; Desprez, T.; Goubet, F.; Refregier, G.; Mouille, G.; McCann, M.; Rayon, C.; Vernhettes, S.; Hofte, H. PROCUSTE1 encodes a cellulose synthase required for normal cell elongation specifically in roots and dark-grown hypocotyls of arabidopsis. Plant Cell 2000, 12, 2409–2423. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.W.; Wang, Z.Y. Brassinosteroid Signal Transduction from Receptor Kinases to Transcription Factors. Annu. Rev. Plant Biol. 2010, 61, 681–704. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Rodriguez, C.; Rubio-Somoza, I.; Sibout, R.; Persson, S. Phytohormones and the cell wall in Arabidopsis during seedling growth. Trends Plant Sci. 2010, 15, 291–301. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.L.; Zhang, J.; Yuan, M.; Ehrhardt, D.W.; Wang, Z.Y.; Mao, T.L. Arabidopsis MICROTUBULE DESTABILIZING PROTEIN40 Is Involved in Brassinosteroid Regulation of Hypocotyl Elongation. Plant Cell 2012, 24, 4012–4025. [Google Scholar] [CrossRef] [PubMed]
- Bishop, G.J.; Koncz, C. Brassinosteroids and plant steroid hormone signaling. Plant Cell 2002, 14, S97–S110. [Google Scholar] [CrossRef] [PubMed]
- Li, J.M.; Nam, K.H. Regulation of brassinosteroid signaling by a GSK3/SHAGGY-like kinase. Science 2002, 295, 1299–1301. [Google Scholar] [PubMed]
- Koivomagi, M.; Ord, M.; Iofik, A.; Valk, E.; Venta, R.; Faustova, I.; Kivi, R.; Balog, E.R.M.; Rubin, S.M.; Loog, M. Multisite phosphorylation networks as signal processors for Cdk1. Nat. Struct. Mol. Biol. 2013, 20, 1415–1424. [Google Scholar] [CrossRef] [PubMed]
- Liepman, A.H.; Nairn, C.J.; Willats, W.G.T.; Sorensen, I.; Roberts, A.W.; Keegstra, K. Functional genomic analysis supports conservation of function among cellulose synthase-like a gene family members and suggests diverse roles of mannans in plants. Plant Physiol. 2007, 143, 1881–1893. [Google Scholar] [CrossRef] [PubMed]
- Richmond, T.A.; Somerville, C.R. The cellulose synthase superfamily. Plant Physiol. 2000, 124, 495–498. [Google Scholar] [CrossRef] [PubMed]
- Youngs, L.H.; Hamann, T.; Osborne, E.; Somerville, C. The Cellulose Synthase Superfamily. In Cellulose: Molecular and Structural Biology; Springer: Dordrecht, The Netherlands, 2007; pp. 35–48. ISBN 978-1-4020-5380-1. [Google Scholar]
- Burton, R.A.; Wilson, S.M.; Hrmova, M.; Harvey, A.J.; Shirley, N.J.; Stone, B.A.; Newbigin, E.J.; Bacic, A.; Fincher, G.B. Cellulose synthase-like CslF genes mediate the synthesis of cell wall (1,3;1,4)-beta-d-glucans. Science 2006, 311, 1940–1942. [Google Scholar] [CrossRef] [PubMed]
- Vega-Sanchez, M.E.; Verhertbruggen, Y.; Christensen, U.; Chen, X.W.; Sharma, V.; Varanasi, P.; Jobling, S.A.; Talbot, M.; White, R.G.; Joo, M.; et al. Loss of Cellulose Synthase-Like F6 Function Affects Mixed-Linkage Glucan Deposition, Cell Wall Mechanical Properties, and Defense Responses in Vegetative Tissues of Rice. Plant Physiol. 2012, 159, 56–69. [Google Scholar] [CrossRef] [PubMed]
- Dhugga, K.S.; Barreiro, R.; Whitten, B.; Stecca, K.; Hazebroek, J.; Randhawa, G.S.; Dolan, M.; Kinney, A.J.; Tomes, D.; Nichols, S.; et al. Guar seed beta-mannan synthase is a member of the cellulose synthase super gene family. Science 2004, 303, 363–366. [Google Scholar] [CrossRef] [PubMed]
- Liepman, A.H.; Wilkerson, C.G.; Keegstra, K. Expression of cellulose synthase-like (Csl) genes in insect cells reveals that CslA family members encode mannan synthases. Proc. Natl. Acad. Sci. USA 2005, 102, 2221–2226. [Google Scholar] [CrossRef] [PubMed]
- Cocuron, J.C.; Lerouxel, O.; Drakakaki, G.; Alonso, A.P.; Liepman, A.H.; Keegstra, K.; Raikhel, N.; Wilkerson, C.G. A gene from the cellulose synthase-like C family encodes a beta-1,4 glucan synthase. Proc. Natl. Acad. Sci. USA 2007, 104, 8550–8555. [Google Scholar] [CrossRef] [PubMed]
- Gu, F.W.; Bringmann, M.; Combs, J.R.; Yang, J.Y.; Bergmann, D.C.; Nielsen, E. Arabidopsis CSLD5 Functions in Cell Plate Formation in a Cell Cycle-Dependent Manner. Plant Cell 2016, 28, 1722–1737. [Google Scholar] [CrossRef] [PubMed]
- Park, S.; Szumlanski, A.L.; Gu, F.; Guo, F.; Nielsen, E. A role for CSLD3 during cell-wall synthesis in apical plasma membranes of tip-growing root-hair cells. Nat. Cell Biol. 2011, 13, 973–980. [Google Scholar] [CrossRef] [PubMed]
- Bernal, A.J.; Yoo, C.M.; Mutwil, M.; Jensen, J.K.; Hou, G.; Blaukopf, C.; Sorensen, I.; Blancaflor, E.B.; Scheller, H.V.; Willats, W.G.T. Functional Analysis of the Cellulose Synthase-Like Genes CSLD1, CSLD2, and CSLD4 in Tip-Growing Arabidopsis Cells. Plant Physiol. 2008, 148, 1238–1253. [Google Scholar] [CrossRef] [PubMed]
- Yin, L.; Verhertbruggen, Y.; Oikawa, A.; Manisseri, C.; Knierim, B.; Prak, L.; Jensen, J.K.; Knox, J.P.; Auer, M.; Willats, W.G.; et al. The cooperative activities of CSLD2, CSLD3, and CSLD5 are required for normal Arabidopsis development. Mol. Plant 2011, 4, 1024–1037. [Google Scholar] [CrossRef] [PubMed]
- Little, A.; Schwerdt, J.G.; Shirley, N.J.; Khor, S.F.; Neumann, K.; O’Donovan, L.A.; Lahnstein, J.; Collins, H.M.; Henderson, M.; Fincher, G.B.; et al. Revised phylogeny of the Cellulose Synthase gene superfamily: Insights into cell wall evolution. Plant Physiol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Zulawski, M.; Braginets, R.; Schulze, W.X. PhosPhAt goes kinases-searchable protein kinase target information in the plant phosphorylation site database PhosPhAt. Nucleic Acids Res. 2013, 41, D1176–D1184. [Google Scholar] [CrossRef] [PubMed]
- Schulze, W.X.; Yao, Q.M.; Xu, D. Databases for Plant Phosphoproteomics. Plant Phosphoproteomics Methods Protoc. 2015, 1306, 207–216. [Google Scholar] [CrossRef]
- Hohmann, U.; Lau, K.; Hothorn, M. The Structural Basis of Ligand Perception and Signal Activation by Receptor Kinases. Annu. Rev. Plant Biol. 2017, 68, 109–137. [Google Scholar] [CrossRef] [PubMed]
- Ellis, C.; Karafyllidis, I.; Wasternack, C.; Turner, J.G. The Arabidopsis mutant cev1 links cell wall signaling to jasmonate and ethylene responses. Plant Cell 2002, 14, 1557–1566. [Google Scholar] [CrossRef] [PubMed]
- Narvaez-Vasquez, J.; Pearce, G.; Ryan, C.A. The plant cell wall matrix harbors a precursor of defense signaling peptides. Proc. Natl. Acad. Sci. USA 2005, 102, 12974–12977. [Google Scholar] [CrossRef] [PubMed]
- Hematy, K.; Sado, P.E.; Van Tuinen, A.; Rochange, S.; Desnos, T.; Balzergue, S.; Pelletier, S.; Renou, J.P.; Hofte, H. A receptor-like kinase mediates the response of Arabidopsis cells to the inhibition of cellulose synthesis. Curr. Biol. 2007, 17, 922–931. [Google Scholar] [CrossRef] [PubMed]
- Der Does, D.; Boutrot, F.; Engelsdorf, T.; Rhodes, J.; McKenna, J.F.; Vernhettes, S.; Koevoets, I.; Tintor, N.; Veerabagu, M.; Miedes, E.; et al. The Arabidopsis leucine-rich repeat receptor kinase MIK2/LRR-KISS connects cell wall integrity sensing, root growth and response to abiotic and biotic stresses. PLoS Genet. 2017, 13. [Google Scholar] [CrossRef] [PubMed]
- Shih, H.-W.; Miller, N.D.; Dai, C.; Spalding, E.P.; Monshausen, G.B. The receptor-like kinase FERONIA is required for mechanical signal transduction in Arabidopsis seedlings. Curr. Biol. 2014, 24, 1887–1892. [Google Scholar] [CrossRef] [PubMed]
- Feng, W.; Kita, D.; Peaucelle, A.; Cartwright, H.N.; Doan, V.; Duan, Q.; Liu, M.-C.; Maman, J.; Steinhorst, L.; Schmitz-Thom, I.; et al. The FERONIA Receptor Kinase Maintains Cell-Wall Integrity during Salt Stress through Ca(2+) Signaling. Curr. Biol. 2018, 28, 666–675.e5. [Google Scholar] [CrossRef] [PubMed]
- Cheung, A.Y.; Wu, H.-M. THESEUS 1, FERONIA and relatives: A family of cell wall-sensing receptor kinases? Curr. Opin. Plant Biol. 2011, 14, 632–641. [Google Scholar] [CrossRef] [PubMed]
- Deslauriers, S.D.; Larsen, P.B. FERONIA is a key modulator of brassinosteroid and ethylene responsiveness in Arabidopsis hypocotyls. Mol. Plant 2010, 3, 626–640. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.; Li, L.; Ye, H.; Yu, X.; Algreen, A.; Yin, Y. Three related receptor-like kinases are required for optimal cell elongation in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2009, 106, 7648–7653. [Google Scholar] [CrossRef] [PubMed]
- Decreux, A.; Messiaen, J. Wall-associated kinase WAK1 interacts with cell wall pectins in a calcium-induced conformation. Plant Cell Physiol. 2005, 46, 268–278. [Google Scholar] [CrossRef] [PubMed]
- Wagner, T.A.; Kohorn, B.D. Wall-associated kinases are expressed throughout plant development and are required for cell expansion. Plant Cell 2001, 13, 303–318. [Google Scholar] [CrossRef] [PubMed]
- Delteil, A.; Gobbato, E.; Cayrol, B.; Estevan, J.; Michel-Romiti, C.; Dievart, A.; Kroj, T.; Morel, J.-B. Several wall-associated kinases participate positively and negatively in basal defense against rice blast fungus. BMC Plant Biol. 2016, 16, 17. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Huang, H.-J.; Ren, S.-T.; Li, J.-J.; Sun, Y.; Sun, D.-Y.; Zhang, S.-Q. The rice wall-associated receptor-like kinase gene OsDEES1 plays a role in female gametophyte development. Plant Physiol. 2012, 160, 696–707. [Google Scholar] [CrossRef] [PubMed]
- Kohorn, B.D.; Kohorn, S.L.; Todorova, T.; Baptiste, G.; Stansky, K.; McCullough, M. A dominant allele of Arabidopsis pectin-binding wall-associated kinase induces a stress response suppressed by MPK6 but not MPK3 mutations. Mol. Plant 2012, 5, 841–851. [Google Scholar] [CrossRef] [PubMed]
- Kohorn, B.D.; Hoon, D.; Minkoff, B.B.; Sussman, M.R.; Kohorn, S.L. Rapid Oligo-Galacturonide Induced Changes in Protein Phosphorylation in Arabidopsis. Mol. Cell. Proteom. 2016, 15, 1351–1359. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.L.; Rahman, A.; Baskin, T.I.; Kieber, J.J. Two Leucine-Rich Repeat Receptor Kinases Mediate Signaling, Linking Cell Wall Biosynthesis and ACC Synthase in Arabidopsis. Plant Cell 2008, 20, 3065–3079. [Google Scholar] [CrossRef] [PubMed]
- Basu, D.; Tian, L.; Debrosse, T.; Poirier, E.; Emch, K.; Herock, H.; Travers, A.; Showalter, A.M. Glycosylation of a Fasciclin-Like Arabinogalactan-Protein (SOS5) Mediates Root Growth and Seed Mucilage Adherence via a Cell Wall Receptor-Like Kinase (FEI1/FEI2) Pathway in Arabidopsis. PLoS ONE 2016, 11, e0145092. [Google Scholar] [CrossRef] [PubMed]
- Dai, Y.J.; Chen, B.L.; Meng, Y.L.; Zhao, W.Q.; Zhou, Z.G.; Oosterhuis, D.M.; Wang, Y.H. Effects of elevated temperature on sucrose metabolism and cellulose synthesis in cotton fibre during secondary cell wall development. Funct. Plant Biol. 2015, 42, 909–919. [Google Scholar] [CrossRef]
- Ivakov, A.; Flis, A.; Apelt, F.; Funfgeld, M.; Scherer, U.; Stitt, M.; Kragler, F.; Vissenberg, K.; Persson, S.; Suslov, D. Cellulose Synthesis and Cell Expansion Are Regulated by Different Mechanisms in Growing Arabidopsis Hypocotyls. Plant Cell 2017, 29, 1305–1315. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Speicher, T.L.; Li, P.Z.; Wallace, I.S. Phosphoregulation of the Plant Cellulose Synthase Complex and Cellulose Synthase-Like Proteins. Plants 2018, 7, 52. https://doi.org/10.3390/plants7030052
Speicher TL, Li PZ, Wallace IS. Phosphoregulation of the Plant Cellulose Synthase Complex and Cellulose Synthase-Like Proteins. Plants. 2018; 7(3):52. https://doi.org/10.3390/plants7030052
Chicago/Turabian StyleSpeicher, Tori L., Patrick Ziqiang Li, and Ian S. Wallace. 2018. "Phosphoregulation of the Plant Cellulose Synthase Complex and Cellulose Synthase-Like Proteins" Plants 7, no. 3: 52. https://doi.org/10.3390/plants7030052
APA StyleSpeicher, T. L., Li, P. Z., & Wallace, I. S. (2018). Phosphoregulation of the Plant Cellulose Synthase Complex and Cellulose Synthase-Like Proteins. Plants, 7(3), 52. https://doi.org/10.3390/plants7030052




