Research Advances in AP2/ERF Transcription Factors in Rice Growth and Development
Abstract
1. Introduction
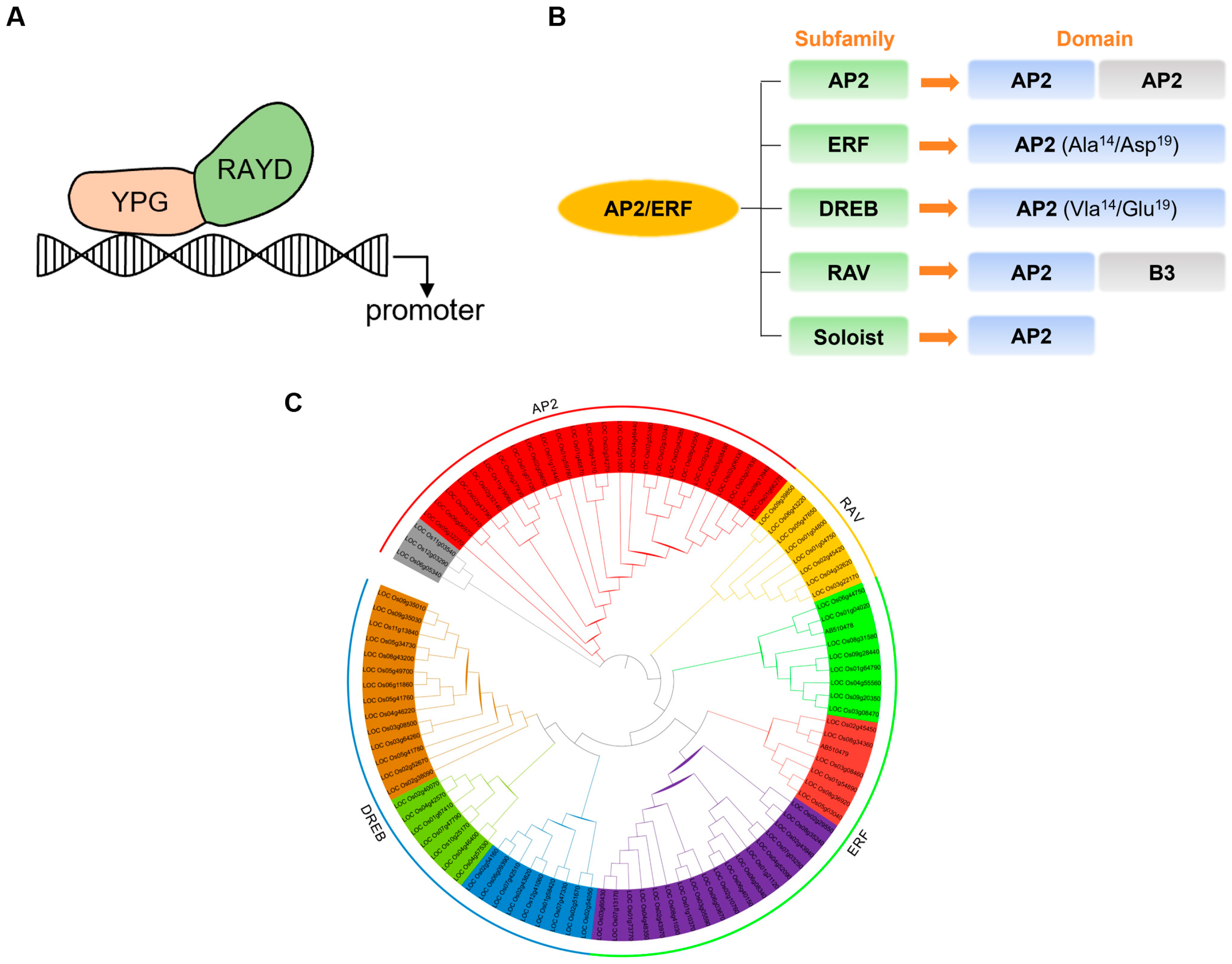
2. Structural Characteristics of AP2/ERF Transcription Factors
3. Research Progress on AP2/ERF Regulate Rice Growth and Development
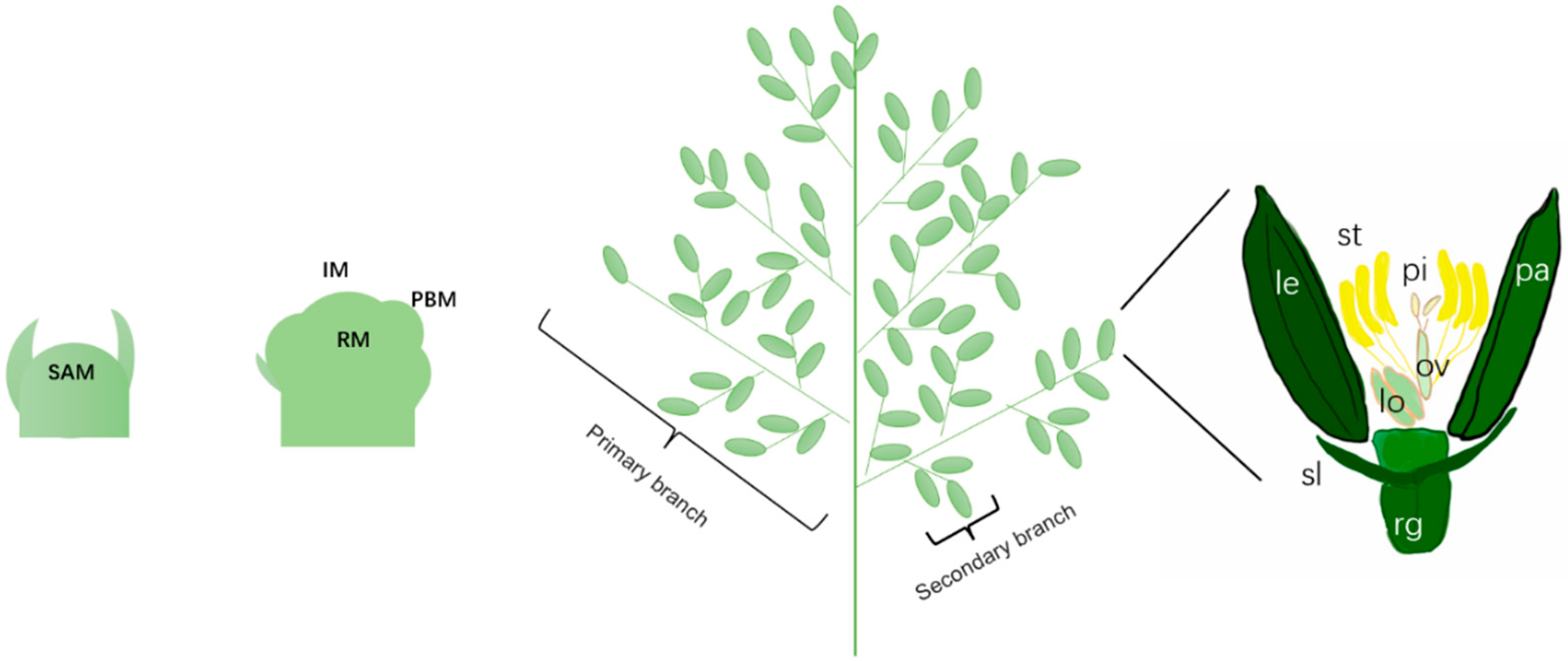
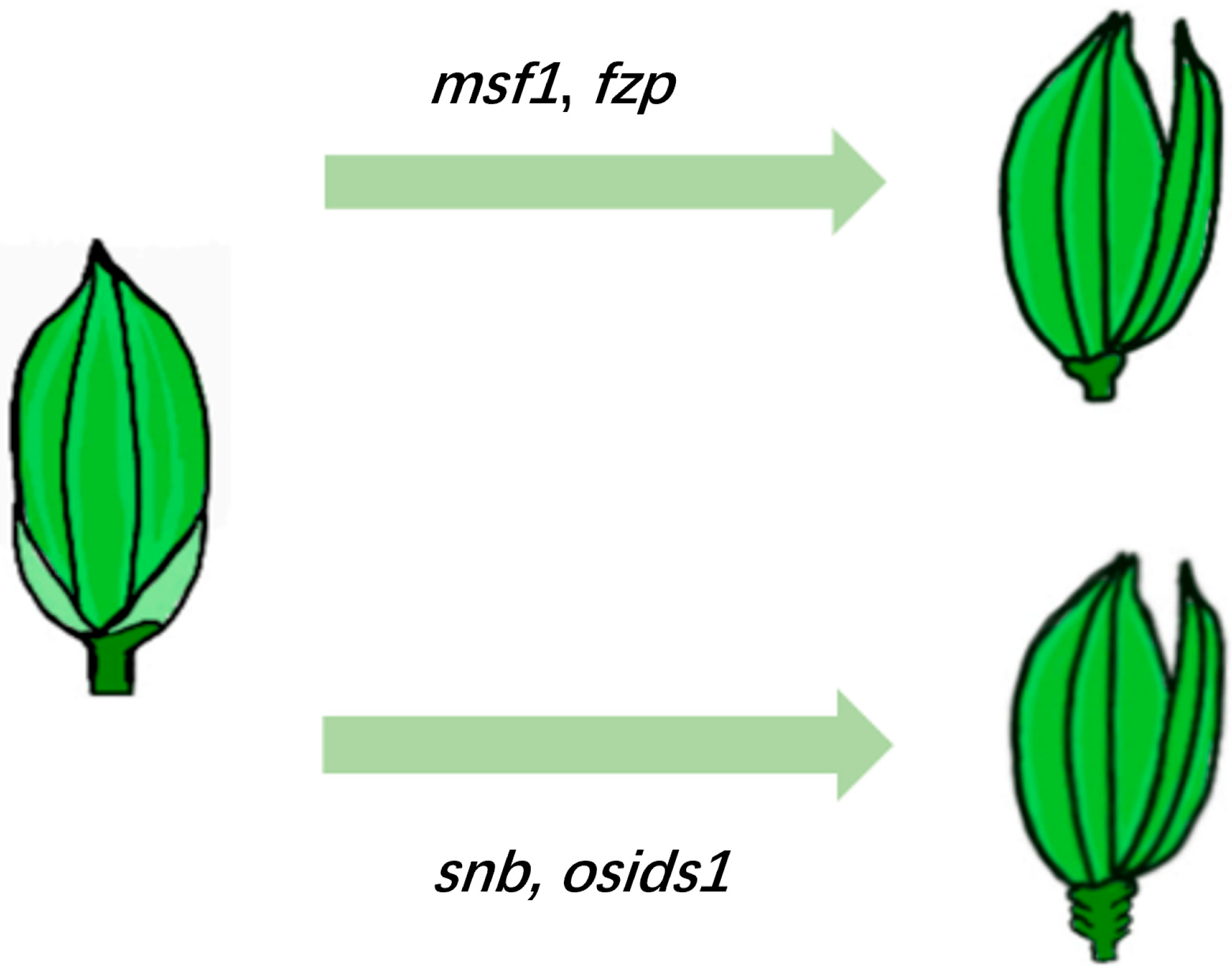
3.1. Development of Floral Organs in Rice
3.2. AP2/ERF Related to Seed Development
4. Research on AP2/ERF Transcription Factors Related to Root, Stem, and Leaf Development in Rice
4.1. AP2/ERF Related to Root Development
4.2. AP2/ERF Related to Stem Development
4.3. AP2/ERF Related to Leaf Development
5. Future Prospects
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| ABA | Abscisic acid |
| AP2 | APETALA 2 |
| BBM1 | Baby boom 1 |
| BR | Brassinosteroid |
| Crl5 | Crown rootless 5 |
| DRAP1 | Drought responsive AP2/EREBP 1 |
| ERF | Ethylene-response factor |
| FM | Floral meristem |
| FZP | Frizzy panicle |
| GA | Gibberellin |
| HL6 | Hairy leaf 6 |
| IAA | Indole-3-acetic acid |
| IM | Inflorescence meristem |
| LFS | Late flowering semi-dwarf |
| MFS1 | Multi-floret spikelet1 |
| OsIDS1 | Oryza sativa indeterminate spikelet 1 |
| OsLG3 | Oryza sativa leafy cotyledon 3 |
| qSH1 | Seed shattering 1 |
| RMD | Rice morphology determinant |
| RSR1 | Rice starch regulator 1 |
| SAM | Shoot apical meristem |
| SERF1 | Salt-responsive ERF 1 |
| SH5 | Seed shattering 5 |
| SHAT1 | Shattering abortion 1 |
| SM | Spikelet meristem |
| SNB | Supernumerary bract |
| SUB1A | Submergence 1A |
References
- Jofuku, K.D.; den Boer, B.G.; Van Montagu, M.; Okamuro, J.K. Control of Arabidopsis flower and seed development by the homeotic gene apetala2. Plant Cell 1994, 6, 1211–1225. [Google Scholar] [CrossRef]
- Auman, H.J.; Nottoli, T.; Lakiza, O.; Winger, Q.; Donaldson, S.; Williams, T. Transcription factor AP2 gamma is essential in the extraembryonic lineages for early postimplantation development. Development 2002, 129, 2733–2747. [Google Scholar] [CrossRef]
- Tsuwamoto, R.; Yokoi, S.; Takahata, Y. Arabidopsis embryomaker encoding an AP2 domain transcription factor plays a key role in developmental change from vegetative to embryonic phase. Plant Mol. Biol. 2010, 73, 481–492. [Google Scholar] [CrossRef]
- Lv, B.; Wei, K.; Hu, K.; Tian, T.; Zhang, F.; Yu, Z.; Zhang, D.; Su, Y.; Sang, Y.; Zhang, X.; et al. Mpk14-mediated auxin signaling controls lateral root development via ERF13 regulated very long chain fatty acid biosynthesis. Mol. Plant 2021, 14, 285–297. [Google Scholar] [CrossRef] [PubMed]
- Guyomarc’H, S.; Boutte, Y.; Laplaze, L. AP2/ERF transcription factors orchestrate very long chain fatty acid biosynthesis during Arabidopsis lateral root development. Mol. Plant 2021, 14, 205–207. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Cheng, S.; Song, Y.; Huang, Y.; Zhou, S.; Liu, X.; Zhou, D.X. The interaction between rice ERF3 and WOX11 promotes crown root development by regulating gene expression involved in cytokinin signaling. Plant Cell 2015, 27, 2469–2483. [Google Scholar] [CrossRef]
- Kang, N.Y.; Lee, H.W.; Kim, J. The AP2/EREBP gene PUCHI co-acts with lBD16/ASL18 and IBD18/ASL20 downstream of ARF7 and ARF19 to regulate lateral root development in Arabidopsis. Plant Cell Physiol. 2013, 54, 1326–1334. [Google Scholar] [CrossRef]
- Wang, Z.; Song, G.; Zhang, F.; Shu, X.; Wang, N. Functional characterization of AP2/ERF transcription factors during flower development and anthocyanin biosynthesis related candidate genes in Lycoris. Int. J. Mol. Sci. 2023, 24, 14464. [Google Scholar] [CrossRef]
- Jiang, Q.; Wang, Z.; Hu, G.; Yao, X. Genome-wide identification and characterization of AP2/ERF gene superfamily during flower development in Actinidia eriantha. BMC Genom. 2022, 23, 650. [Google Scholar] [CrossRef] [PubMed]
- Nakano, T.; Fujisawa, M.; Shima, Y.; Ito, Y. The AP2/ERF transcription factor slERF52 functions in flower pedicel abscission in tomato. J. Exp. Bot. 2014, 65, 3111–3119. [Google Scholar] [CrossRef] [PubMed]
- Wollmann, H.; Mica, E.; Todesco, M.; Long, J.A.; Weigel, D. On reconciling the interactions between APETALA2, miR172 and AGAMOUS with the ABC model of flower development. Development 2010, 137, 3633–3642. [Google Scholar] [CrossRef]
- Meng, H.; Chen, Y.; Li, T.; Shi, H.; Yu, S.; Gao, Y.; Wang, Z.; Wang, X.; Zhu, J.K.; Hong, Y.; et al. APETALA2 is involved in ABA signaling during seed germination. Plant Mol. Biol. 2023, 112, 99–103. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Sheerin, D.J.; von Roepenack-Lahaye, E.; Stahl, M.; Hiltbrunner, A. The phytochrome interacting proteins ERF55 and ERF58 repress light-induced seed germination in Arabidopsis thaliana. Nat. Commun. 2022, 13, 1656. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Wang, H.; Zhang, J.; Chen, S. A seed-specific AP2-domain transcription factor from soybean plays a certain role in regulation of seed germination. Sci. China C Life Sci. 2008, 51, 336–345. [Google Scholar] [CrossRef] [PubMed]
- Teng, K.; Zhao, N.; Xie, Y.; Li, R.; Li, J. An AP2/ERF transcription factor controls generation of the twin-seedling rice. J. Adv. Res. 2024; in press. [Google Scholar] [CrossRef]
- Liu, C.; Ma, T.; Yuan, D.; Zhou, Y.; Long, Y.; Li, Z.; Dong, Z.; Duan, M.; Yu, D.; Jing, Y.; et al. The OsEIL1-OsERF115-target gene regulatory module controls grain size and weight in rice. Plant Biotechnol. J. 2022, 20, 1470–1486. [Google Scholar] [CrossRef]
- Harrop, T.; Mantegazza, O.; Luong, A.M.; Bethune, K.; Lorieux, M.; Jouannic, S.; Adam, H. A set of AP2-like genes is associated with inflorescence branching and architecture in domesticated rice. J. Exp. Bot. 2019, 70, 5617–5629. [Google Scholar] [CrossRef]
- Qi, W.; Sun, F.; Wang, Q.; Chen, M.; Huang, Y.; Feng, Y.Q.; Luo, X.; Yang, J. Rice ethylene-response AP2/ERF factor OsEATB restricts internode elongation by down-regulating a gibberellin biosynthetic gene. Plant Physiol. 2011, 157, 216–228. [Google Scholar] [CrossRef]
- Fu, F.F.; Xue, H.W. Coexpression analysis identifies rice starch regulator1, a rice AP2/EREBP family transcription factor, as a novel rice starch biosynthesis regulator. Plant Physiol. 2010, 154, 927–938. [Google Scholar] [CrossRef]
- Liu, W.; Stewart, C.J. Plant synthetic promoters and transcription factors. Curr. Opin. Biotechnol. 2016, 37, 36–44. [Google Scholar] [CrossRef] [PubMed]
- Franco-Zorrilla, J.M.; Solano, R. Identification of plant transcription factor target sequences. Biochim. Biophys. Acta Gene Regul. Mech. 2017, 1860, 21–30. [Google Scholar] [CrossRef]
- Feng, K.; Hou, X.L.; Xing, G.M.; Liu, J.X.; Duan, A.Q.; Xu, Z.S.; Li, M.Y.; Zhuang, J.; Xiong, A.S. Advances in AP2/ERF super-family transcription factors in plant. Crit. Rev. Biotechnol. 2020, 40, 750–776. [Google Scholar] [CrossRef]
- Shinshi, H.; Usami, S.; Ohme-Takagi, M. Identification of an ethylene-responsive region in the promoter of a tobacco class I chitinase gene. Plant Mol. Biol. 1995, 27, 923–932. [Google Scholar] [CrossRef] [PubMed]
- Ohme-Takagi, M.; Shinshi, H. Ethylene-inducible DNA binding proteins that interact with an ethylene-responsive element. Plant Cell 1995, 7, 173–182. [Google Scholar]
- Nakano, T.; Suzuki, K.; Fujimura, T.; Shinshi, H. Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol. 2006, 140, 411–432. [Google Scholar] [CrossRef] [PubMed]
- Weigel, D. The APETALA2 domain is related to a novel type of DNA binding domain. Plant Cell 1995, 7, 388–389. [Google Scholar]
- Licausi, F.; Ohme-Takagi, M.; Perata, P. Apetala2/ethylene responsive factor (AP2/ERF) transcription factors: Mediators of tress responses and developmental programs. New Phytol. 2013, 199, 639–649. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Yang, L.; Li, A.; Guo, J.; Wang, H.; Qi, H.; Li, M.; Yang, P.; Song, S. An AP2/ERF transcription factor confers chilling tolerance in rice. Sci. Adv. 2024, 10, eado4788. [Google Scholar] [CrossRef]
- Xie, Z.; Nolan, T.M.; Jiang, H.; Yin, Y. AP2/ERF transcription factor regulatory networks in hormone and abiotic stress responses in Arabidopsis. Front. Plant Sci. 2019, 10, 228. [Google Scholar] [CrossRef]
- Jofuku, K.D.; Omidyar, P.K.; Gee, Z.; Okamuro, J.K. Control of seed mass and seed yield by the floral homeotic gene APETALA2. Proc. Natl. Acad. Sci. USA 2005, 102, 3117–3122. [Google Scholar] [CrossRef]
- Maes, T.; Van de Steene, N.; Zethof, J.; Karimi, M.; D’Hauw, M.; Mares, G.; Van Montagu, M.; Gerats, T. PETUNIA AP2-like genes and their role in flower and seed development. Plant Cell 2001, 13, 229–244. [Google Scholar] [CrossRef]
- Zong, Y.; Hao, Z.; Tu, Z.; Shen, Y.; Zhang, C.; Wen, S.; Yang, L.; Ma, J.; Li, H. Genome-wide survey and identification of AP2/ERF genes involved in shoot and leaf development in liriodendron chinense. BMC Genom. 2021, 22, 807. [Google Scholar] [CrossRef]
- Woo, H.R.; Kim, J.H.; Kim, J.; Kim, J.; Lee, U.; Song, I.J.; Kim, J.H.; Lee, H.Y.; Nam, H.G.; Lim, P.O. The rav1 transcription factor positively regulates leaf senescence in Arabidopsis. J. Exp. Bot. 2010, 61, 3947–3957. [Google Scholar] [CrossRef]
- Zhu, X.; Yan, X.; Li, W.; Zhang, M.; Leng, J.; Yu, Q.; Liu, L.; Xue, D.; Zhang, D.; Ding, Z. GmERF13 mediates salt inhibition of nodulation through interacting with GmLBD16a in soybean. Nat. Commun. 2025, 16, 435. [Google Scholar] [CrossRef]
- Hu, Z.; Wang, X.; Wei, L.; Wansee, S.; Rabbani, N.H.; Chen, L.; Kang, Z.; Wang, J. Taap2-10, an AP2/ERF transcription factor, contributes to wheat resistance against stripe rust. J. Plant Physiol. 2023, 288, 154078. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Gao, M.; Zhao, Y.; Chen, Y.; Wu, L.; Yin, H.; Xiong, S.; Wang, S.; Wang, J.; Yang, Y.; et al. LcERF19, an AP2/ERF transcription factor from Litsea cubeba, positively regulates geranial and neral biosynthesis. Hortic. Res. 2022, 9, uhac093. [Google Scholar] [CrossRef] [PubMed]
- Xue, G.P.; Loveridge, C.W. Hvdrf1 is involved in abscisic acid-mediated gene regulation in barley and produces two forms of AP2 transcriptional activators, interacting preferably with a CT-rich element. Plant J. 2004, 37, 326–339. [Google Scholar] [CrossRef] [PubMed]
- Ray, D.K.; Mueller, N.D.; West, P.C.; Foley, J.A. Yield trends are insufficient to double global crop production by 2050. PLoS ONE 2013, 8, e66428. [Google Scholar] [CrossRef]
- Zhang, D.; Yuan, Z. Molecular control of grass inflorescence development. Annu. Rev. Plant Biol. 2014, 65, 553–578. [Google Scholar] [CrossRef]
- Thompson, B.E.; Hake, S. Translational biology: From Arabidopsis flowers to grass inflorescence architecture. Plant Physiol. 2009, 149, 38–45. [Google Scholar] [CrossRef]
- Wang, S.S.; Chung, C.L.; Chen, K.Y.; Chen, R.K. A novel variation in the Frizzle Panicle (FZP) gene promoter improves grain number and yield in rice. Genetics 2020, 215, 243–252. [Google Scholar] [CrossRef]
- Xing, H.; Wang, H.; Huang, Y.; Ma, X.; Wu, S.; Li, Y.; Sun, C.; Sun, H. FZP modulates tillering via OsMADS57 in rice. Plant Biotechnol. J. 2025, 23, 1202–1212. [Google Scholar] [CrossRef]
- Wang, W.; Chen, W.; Wang, J. Frizzle Panicle (FZP) regulates rice spikelets development through modulating cytokinin metabolism. BMC Plant Biol. 2023, 23, 650. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.S.; Chen, R.K.; Chen, K.Y.; Liu, C.Y.; Kao, S.M.; Chung, C.L. Genetic mapping of the qSBN7 locus, a QTL controlling secondary branch number per panicle in rice. Breed. Sci. 2017, 67, 340–347. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.Y.; Lee, J.; Moon, S.; Park, S.Y.; An, G. The rice heterochronic gene supernumerary bract regulates the transition from spikelet meristem to floral meristem. Plant J. 2007, 49, 64–78. [Google Scholar] [CrossRef]
- Lee, D.Y.; An, G. Two AP2 family genes, Supernumerary Bract (SNB) and OsIndeterminate Spikelet 1 (osIDS1), synergistically control inflorescence architecture and floral meristem establishment in rice. Plant J. 2012, 69, 445–461. [Google Scholar] [CrossRef]
- Lee, Y.S.; Lee, D.Y.; Cho, L.H.; An, G. Rice miR172 induces flowering by suppressing OsIDS1 and SNB, two AP2 genes that negatively regulate expression of Ehd1 and florigens. Rice 2014, 7, 31. [Google Scholar] [CrossRef]
- Ji, H.; Han, C.D.; Lee, G.S.; Jung, K.H.; Kang, D.Y.; Oh, J.; Oh, H.; Cheon, K.S.; Kim, S.L.; Choi, I.; et al. Mutations in the microRNA172 binding site of Supernumerary Bract (SNB) suppress internode elongation in rice. Rice 2019, 12, 62. [Google Scholar] [CrossRef]
- Ren, D.; Li, Y.; Zhao, F.; Sang, X.; Shi, J.; Wang, N.; Guo, S.; Ling, Y.; Zhang, C.; Yang, Z.; et al. Multi-Floret Spikelet1, which encodes an AP2/ERF protein, determines spikelet meristem fate and sterile lemma identity in rice. Plant Physiol. 2013, 162, 872–884. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Ali, A.; Wang, J.; Song, J.; Fang, Y.; Zhou, T.; Luo, Y.; Zhang, H.; Chen, X.; Liao, Y.; et al. A homologous gene of OsREL2/ASP1, ASP-LSL regulates pleiotropic phenotype including long sterile lemma in rice. BMC Plant Biol. 2021, 21, 390. [Google Scholar] [CrossRef]
- Song, S.; Wang, G.; Hu, Y.; Liu, H.; Bai, X.; Qin, R.; Xing, Y. OsMFT1 increases spikelets per panicle and delays heading date in rice by suppressing EHD1, FZP and SEPALLATA-like genes. J. Exp. Bot. 2018, 69, 4283–4293. [Google Scholar] [CrossRef]
- Shim, Y.; Lim, C.; Seong, G.; Choi, Y.; Kang, K.; Paek, N.C. The AP2/ERF transcription factor late flowering semi-dwarf suppresses long-day-dependent repression of flowering. Plant Cell Environ. 2022, 45, 2446–2459. [Google Scholar] [CrossRef]
- Yu, J.; Xiong, H.; Zhu, X.; Zhang, H.; Li, H.; Miao, J.; Wang, W.; Tang, Z.; Zhang, Z.; Yao, G.; et al. OsLG3 contributing to rice grain length and yield was mined by ho-lamap. BMC Biol. 2017, 15, 28. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Ma, X.; Zhao, S.; Tang, Y.; Liu, F.; Gu, P.; Fu, Y.; Zhu, Z.; Cai, H.; Sun, C.; et al. The APETALA2-like transcription factor supernumerary bract controls rice seed shattering and seed size. Plant Cell 2019, 31, 17–36. [Google Scholar] [CrossRef] [PubMed]
- Ren, D.; Hu, J.; Xu, Q.; Cui, Y.; Zhang, Y.; Zhou, T.; Rao, Y.; Xue, D.; Zeng, D.; Zhang, G.; et al. FZP determines grain size and sterile lemma fate in rice. J. Exp. Bot. 2018, 69, 4853–4866. [Google Scholar] [CrossRef]
- Schmidt, R.; Schippers, J.H.; Mieulet, D.; Watanabe, M.; Hoefgen, R.; Guiderdoni, E.; Mueller-Roeber, B. Salt-responsive ERF1 is a negative regulator of grain filling and gibberellin-mediated seedling establishment in rice. Mol. Plant 2014, 7, 404–421. [Google Scholar] [CrossRef]
- Khanday, I.; Santos-Medellin, C.; Sundaresan, V. Somatic embryo initiation by rice BABY BOOM1 involves activation of zygote-expressed auxin biosynthesis genes. New Phytol. 2023, 238, 673–687. [Google Scholar] [CrossRef]
- Chahal, L.S.; Conner, J.A.; Ozias-Akins, P. Phylogenetically distant baby boom genes from setaria italica induce parthenogenesis in rice. Front. Plant Sci. 2022, 13, 863908. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.J.; Zhang, X.F.; Xue, H.W. Rice aleurone layer specific OsNF-YB1 regulates grain filling and endosperm development by interacting with an erf transcription factor. J. Exp. Bot. 2016, 67, 6399–6411. [Google Scholar] [CrossRef]
- Zhang, Y.L.; Xia, Q.Y.; Jiang, X.Q.; Hu, W.; Ye, X.X.; Huang, Q.X.; Yu, S.B.; Guo, A.P.; Lu, B.R. Reducing seed shattering in weedy rice by editing SH4 and qSH1 genes: Implications in environmental biosafety and weed control through transgene mitigation. Biology 2022, 11, 1823. [Google Scholar] [CrossRef]
- Hofmann, N.R. SHAT1, a new player in seed shattering of rice. Plant Cell 2012, 24, 839. [Google Scholar] [CrossRef] [PubMed]
- Kitomi, Y.; Ito, H.; Hobo, T.; Aya, K.; Kitano, H.; Inukai, Y. The auxin responsive AP2/ERF transcription factor crown rootless5 is involved in crown root initiation in rice through the induction of OsRR1, a type-a response regulator of cytokinin signaling. Plant J. 2011, 67, 472–484. [Google Scholar] [CrossRef] [PubMed]
- Lavarenne, J.; Gonin, M.; Guyomarc’H, S.; Rouster, J.; Champion, A.; Sallaud, C.; Laplaze, L.; Gantet, P.; Lucas, M. Inference of the gene regulatory network acting downstream of crown rootless 1 in rice reveals a regulatory cascade linking genes involved in auxin signaling, crown root initiation, and root meristem specification and maintenance. Plant J. 2019, 100, 954–968. [Google Scholar] [CrossRef] [PubMed]
- Neogy, A.; Singh, Z.; Mushahary, K.; Yadav, S.R. Dynamic cytokinin signaling and function of auxin in cytokinin responsive domains during rice crown root development. Plant Cell Rep. 2021, 40, 1367–1375. [Google Scholar] [CrossRef]
- Lee, D.K.; Yoon, S.; Kim, Y.S.; Kim, J.K. Rice OsERF71 mediated root modification affects shoot drought tolerance. Plant Signal. Behav. 2017, 12, e1268311. [Google Scholar] [CrossRef]
- Lee, D.K.; Jung, H.; Jang, G.; Jeong, J.S.; Kim, Y.S.; Ha, S.H.; Do, C.Y.; Kim, J.K. Overexpression of the OsERF71 transcription factor alters rice root structure and drought resistance. Plant Physiol. 2016, 172, 575–588. [Google Scholar] [CrossRef]
- Jung, H.; Chung, P.J.; Park, S.H.; Redillas, M.; Kim, Y.S.; Suh, J.W.; Kim, J.K. Overexpression of OsERF48 causes regulation of OsCML16, a calmodulin-like protein gene that enhances root growth and drought tolerance. Plant Biotechnol. J. 2017, 15, 1295–1308. [Google Scholar] [CrossRef]
- Zhai, H.; Zhou, C.; Zhang, Y.; Wang, Y.; Wang, M.; Wei, S.; Li, T. Mechanism analysis of OsBHLH34-OsERF34 mediated regulation of rice resistance to sheath blight. Int. J. Mol. Sci. 2025, 26, 2249. [Google Scholar] [CrossRef]
- Zhang, J.; Liu, Z.; Sakamoto, S.; Mitsuda, N.; Ren, A.; Persson, S.; Zhang, D. Ethylene response factor 34 promotes secondary cell wall thickening and strength of rice peduncles. Plant Physiol. 2022, 190, 1806–1820. [Google Scholar] [CrossRef]
- Ma, Z.; Wu, T.; Huang, K.; Jin, Y.M.; Li, Z.; Chen, M.; Yun, S.; Zhang, H.; Yang, X.; Chen, H.; et al. A novel AP2/ERF transcription factor, OsRPH1, negatively regulates plant height in rice. Front. Plant Sci. 2020, 11, 709. [Google Scholar] [CrossRef]
- Jung, S.E.; Bang, S.W.; Kim, S.H.; Seo, J.S.; Yoon, H.B.; Kim, Y.S.; Kim, J.K. Overexpression of OsERF83, a vascular tissue-specific transcription factor gene, confers drought tolerance in rice. Int. J. Mol. Sci. 2021, 22, 7656. [Google Scholar] [CrossRef]
- Tezuka, D.; Kawamata, A.; Kato, H.; Saburi, W.; Mori, H.; Imai, R. The rice ethylene response factor OsERF83 positively regulates disease resistance to magnaporthe oryzae. Plant Physiol. Biochem. 2019, 135, 263–271. [Google Scholar] [CrossRef]
- Lin, C.C.; Lee, W.J.; Zeng, C.Y.; Chou, M.Y.; Lin, T.J.; Lin, C.S.; Ho, M.C.; Shih, M.C. Sub1a-1 anchors a regulatory cascade for epigenetic and transcriptional controls of submergence tolerance in rice. PNAS Nexus 2023, 2, pgad229. [Google Scholar] [CrossRef] [PubMed]
- Fukao, T.; Yeung, E.; Bailey-Serres, J. The submergence tolerance regulator sub1a mediates crosstalk between submergence and drought tolerance in rice. Plant Cell 2011, 23, 412–427. [Google Scholar] [CrossRef] [PubMed]
- Lim, C.; Kang, K.; Shim, Y.; Sakuraba, Y.; An, G.; Paek, N.C. Rice ethylene response factor 101 promotes leaf senescence through jasmonic acid-mediated regulation of OsNAP and OsMYC2. Front. Plant Sci. 2020, 11, 1096. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Gao, D.; Xiong, Y.; Tang, X.; Xiao, X.; Wang, C.; Yu, S. Hairy leaf 6, an AP2/ERF transcription factor, interacts with OsWOX3B and regulates trichome formation in rice. Mol. Plant 2017, 10, 1417–1433. [Google Scholar] [CrossRef]
- Li, J.; Tang, B.; Li, Y.; Li, C.; Guo, M.; Chen, H.; Han, S.; Li, J.; Lou, Q.; Sun, W.; et al. Rice SPL10 positively regulates trichome development through expression of HL6 and auxin-related genes. J. Integr. Plant Biol. 2021, 63, 1521–1537. [Google Scholar] [CrossRef]
| Genes | Gene ID | Function Description | References |
|---|---|---|---|
| FZP | 4344233 | Control the number of spikelets, delay the transition from spikelets to flowers, and affect yield and quality. | [41,42,43,44] |
| SNB | 4342787 | Positive regulation of spikelet to flower transition affects inflorescence structure and floral meristem establishment. | [45,46,47,48] |
| OsIDS1 | 4334582 | Collaborate with SNB to control inflorescence structure and floral meristem, affecting bract formation. | [47,48] |
| MFS1 | 4339208 | Participate in the establishment of spikelet meristematic tissue and the formation of floral organ characteristics, regulating the transition time from spikelet to flower. | [49,50] |
| LFS | 4345697 | Promote flowering under long day conditions by inhibiting OsLFL1 expression. | [53] |
| Genes | Gene ID | Function Description | References |
|---|---|---|---|
| OsLG3 | 4331845 | Positive regulation of rice grain length which regulates BR signal transduction by encoding phosphatase, increasing grain length and yield. | [53] |
| OsERF115 | 4346073 | Positively regulates grain size and weight, promoting elongation and division of glume cells, and increasing grain weight. | [16] |
| OsSNB | 4342787 | Participates in the regulation of brassinosteroid and auxin signaling, negatively regulate grain size, and its mutants increase grain length and weight. | [54] |
| FZP | 4344233 | Regulates the determinacy of spikelet and positively regulating seed size. | [42,55] |
| SERF1 | 107277887 | Responds to salt stress, regulating RPBF expression, affecting grain filling and starch biosynthesis. | [56] |
| RSR1 | 4337654 | Negatively regulates starch synthesis gene expression in seeds, affecting seed size and increasing amylose content. | [19] |
| BBM1, BBM2, BBM3 | 4350315 | Plays a critical role in early embryonic development and is an inducer of embryonic development. | [57,58] |
| OsSNB (ssh1 allele) | 4342787 | Regulates qSH1 and SH5, affecting the deposition of lignin in the detachment zone and the normal development of the detachment layer, and reducing grain size. | [54] |
| SH4 | 9266435 | Key transcription factors that control seed drop and affect the formation of detached cells. | [60,61] |
| SHAT1 | 9269072 | Participates in delamination development and is crucial for genetic regulation of rice seed drop. | [61] |
| Genes | Gene ID | Function Description | References |
|---|---|---|---|
| Crl5 | 4342308 | Induced by auxin, upregulation of OsRR1 inhibits cytokinin signaling and promotes crown root formation. | [62,63] |
| ERF3 | 4331843 | Interacts with WOX11 to regulate coronal root development, controlling coronal root initiation through RR2. | [6,64] |
| OsAP2/ERF-40 | 4324418 | Specifically expressed in adventitious root primordia, it affects root development by regulating the OsERF3-WOX11-RR2 pathway. | [64] |
| OsERF71 | 4340383 | Expressed in root meristem tissue, affecting cell wall relaxation and lignin synthesis, enhancing root adaptability. | [65,66] |
| OsERF48/ OsDRAP1 | 4345541 | Regulates OsCML16 expression to promote root growth, including primary and lateral roots. | [67] |
| Genes | Gene ID | Function Description | References |
|---|---|---|---|
| OsERF34 | 9266374 | Promotes the expression of RMD, positively regulates secondary cell wall synthesis and mechanical strength. | [68,69] |
| OsRPH1 | 4339670 | Regulates stem development and is closely related to the gibberellin signaling pathway. | [70] |
| OsERF83 | 107276031 | Overexpression enhances drought resistance but affects stem and spike length, leading to a decrease in yield. | [71,72] |
| Genes | Gene ID | Function Description | References |
|---|---|---|---|
| SUB1A | 4352338 | Inhibits ethylene accumulation, weakens the effects of jasmonic acid and salicylic acid on leaf senescence, and enhances stress tolerance. | [73,74] |
| OsERF101 | 4335707 | Combines OsNAP and OsMYC2 promoters to promote leaf senescence. | [75] |
| HL6 | 4341722 | Affects the distribution and morphology of trichomes on rice leaves and other epidermal parts. | [76,77] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, Y.; Li, R.; Li, D.; Fan, X.; Chen, X.; Zhuang, C.; Li, J. Research Advances in AP2/ERF Transcription Factors in Rice Growth and Development. Plants 2025, 14, 2673. https://doi.org/10.3390/plants14172673
He Y, Li R, Li D, Fan X, Chen X, Zhuang C, Li J. Research Advances in AP2/ERF Transcription Factors in Rice Growth and Development. Plants. 2025; 14(17):2673. https://doi.org/10.3390/plants14172673
Chicago/Turabian StyleHe, Ying, Ruiqi Li, Dike Li, Xingyi Fan, Xiaoyuan Chen, Chuxiong Zhuang, and Jing Li. 2025. "Research Advances in AP2/ERF Transcription Factors in Rice Growth and Development" Plants 14, no. 17: 2673. https://doi.org/10.3390/plants14172673
APA StyleHe, Y., Li, R., Li, D., Fan, X., Chen, X., Zhuang, C., & Li, J. (2025). Research Advances in AP2/ERF Transcription Factors in Rice Growth and Development. Plants, 14(17), 2673. https://doi.org/10.3390/plants14172673






