A Comprehensive Analysis of the Peanut SQUAMOSA Promoter Binding Protein-like Gene Family and How AhSPL5 Enhances Salt Tolerance in Transgenic Arabidopsis
Abstract
1. Introduction
2. Results
2.1. Identification of the AhSPL Gene Family in Peanuts
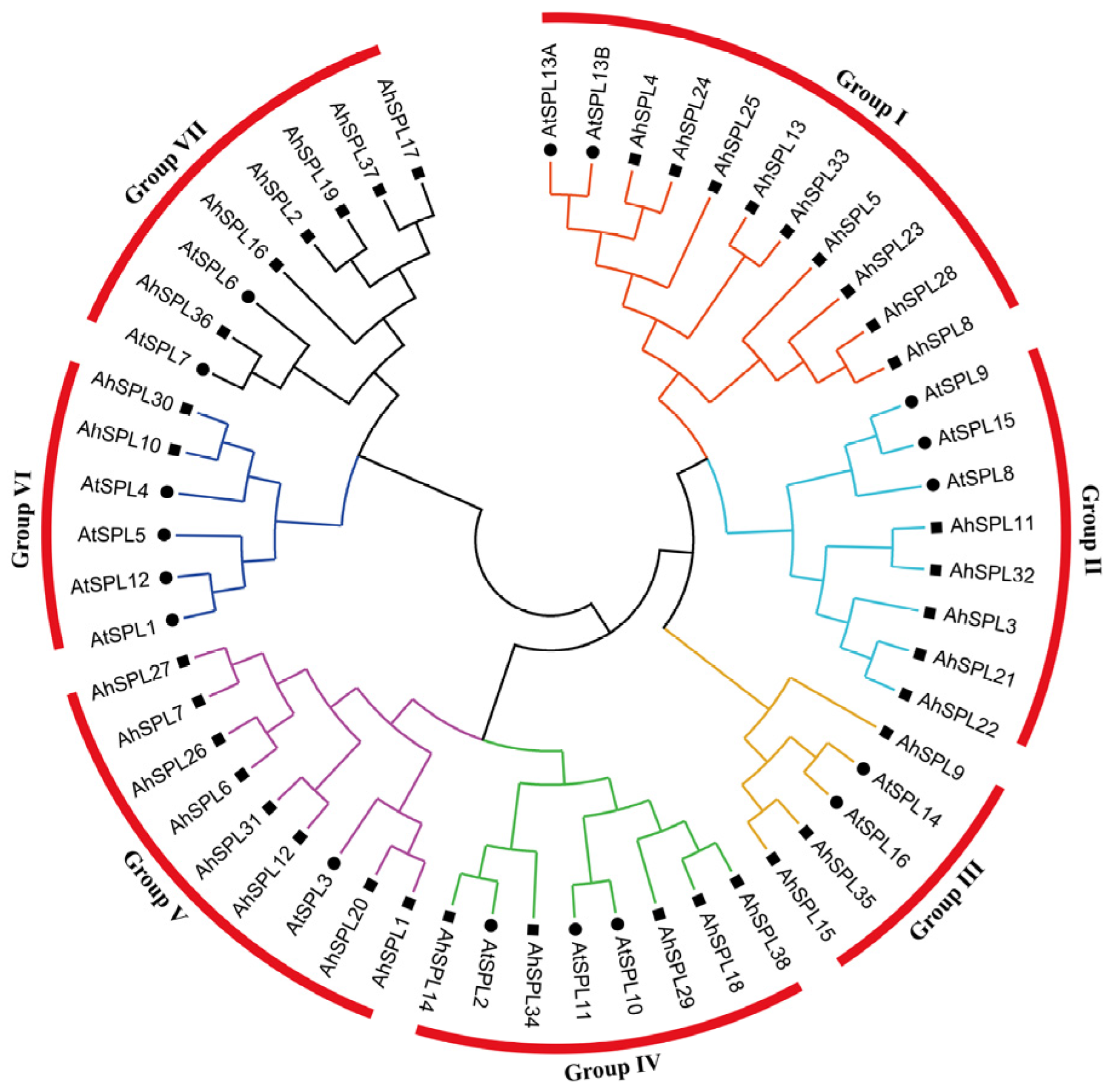
2.2. Phylogenetic Analysis
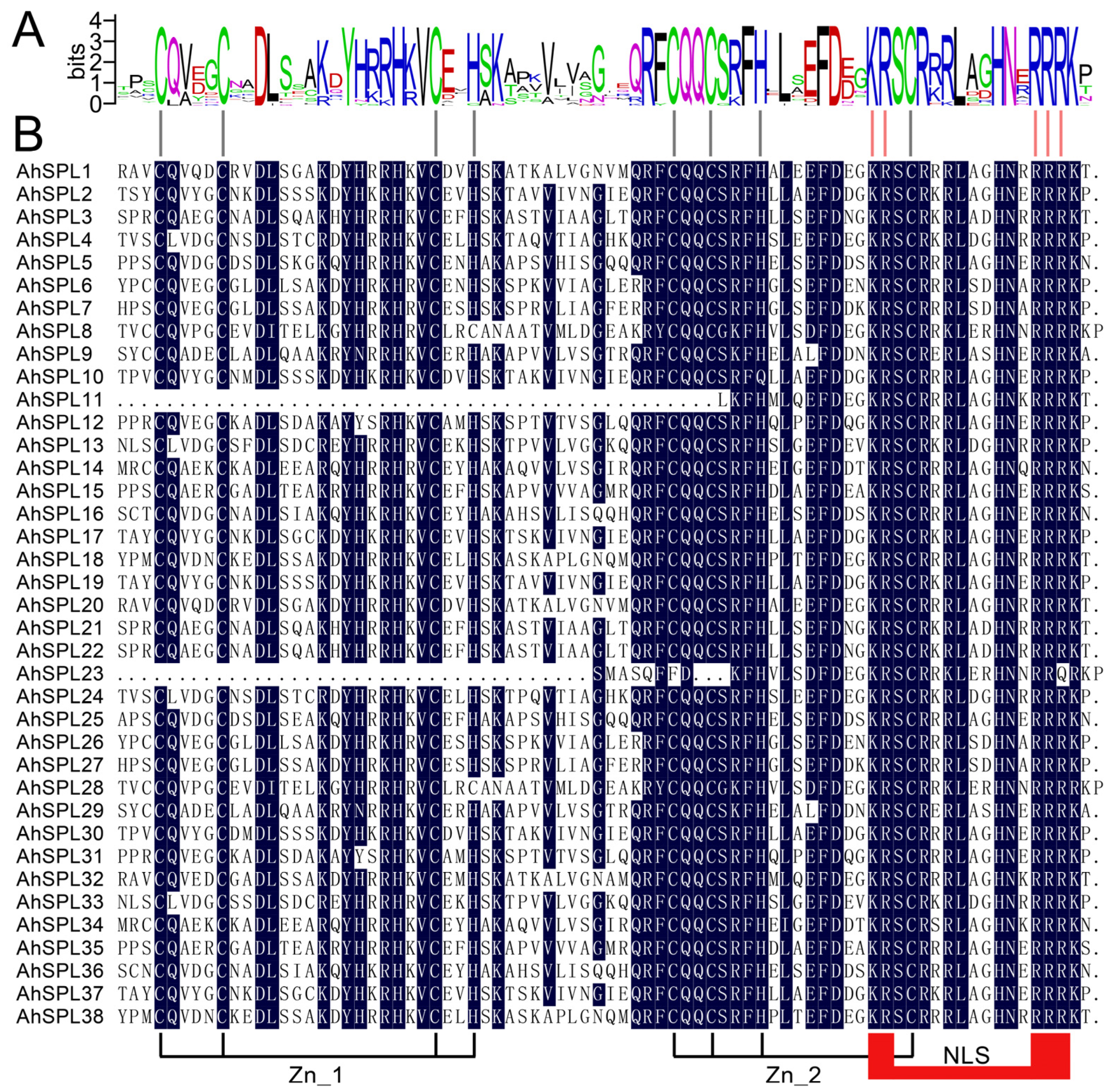
2.3. Conserved Motif and Gene Structure Analysis
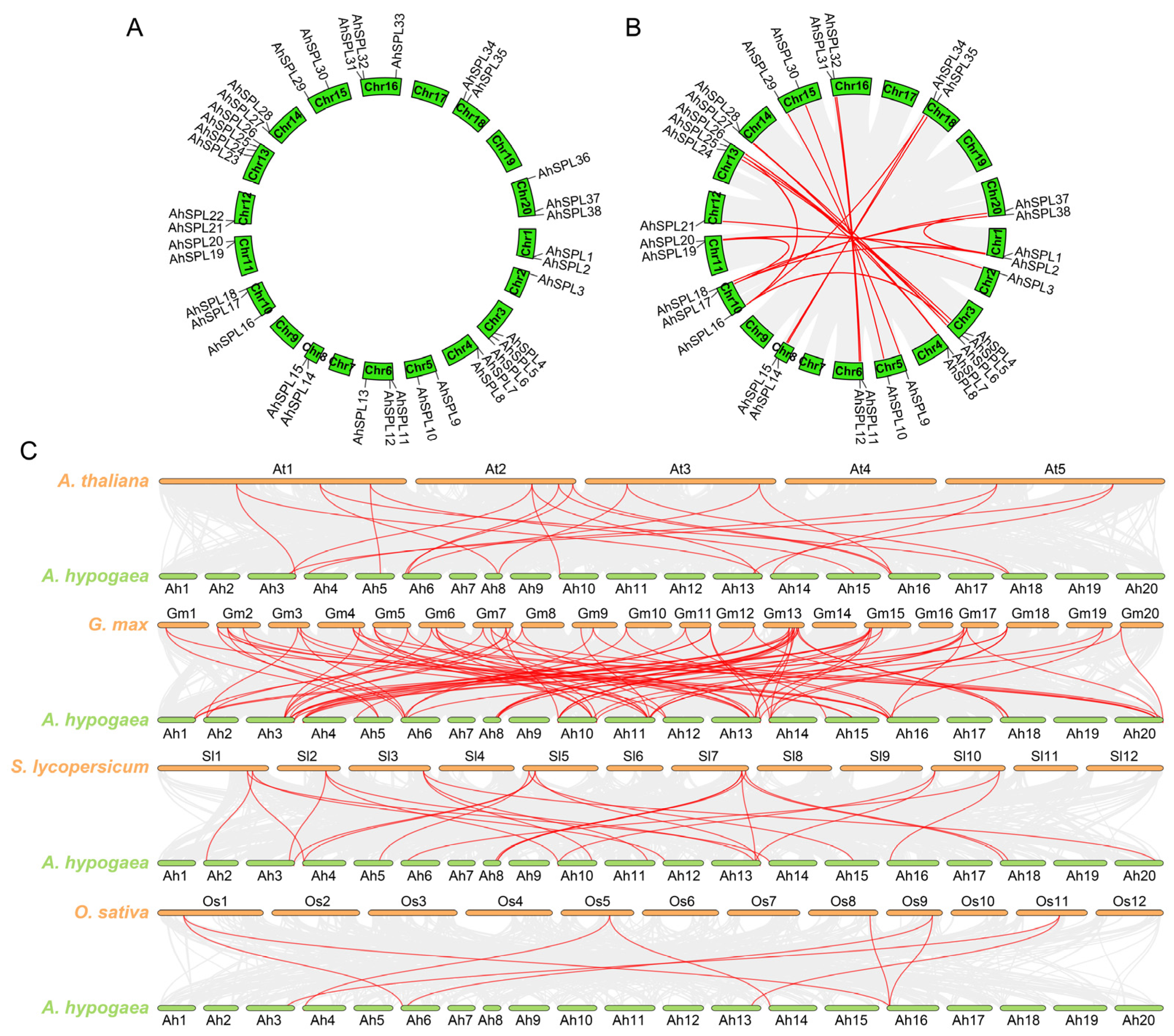
2.4. Chromosomal Localization, Duplication Events, and Syntenic Analysis
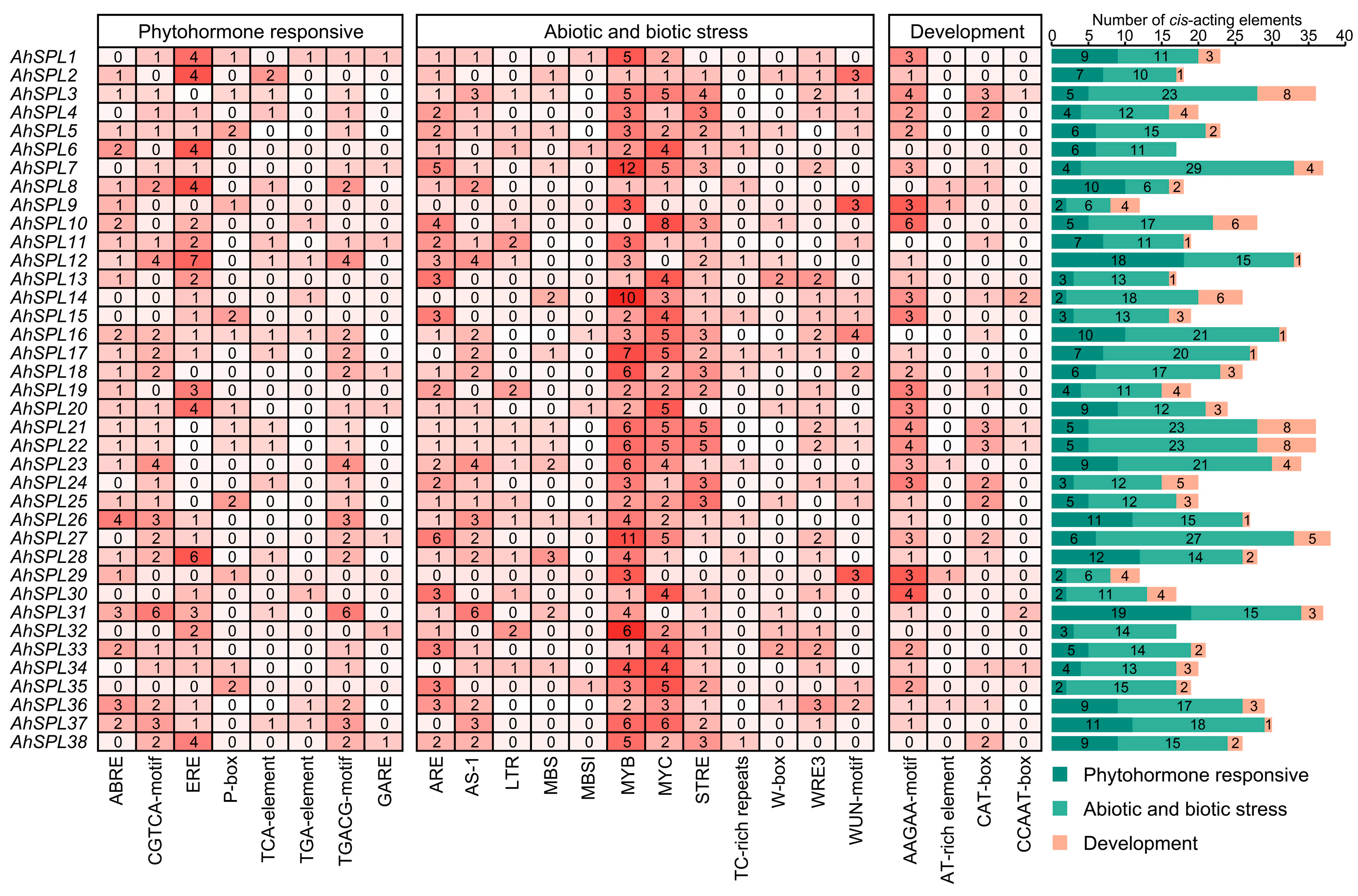
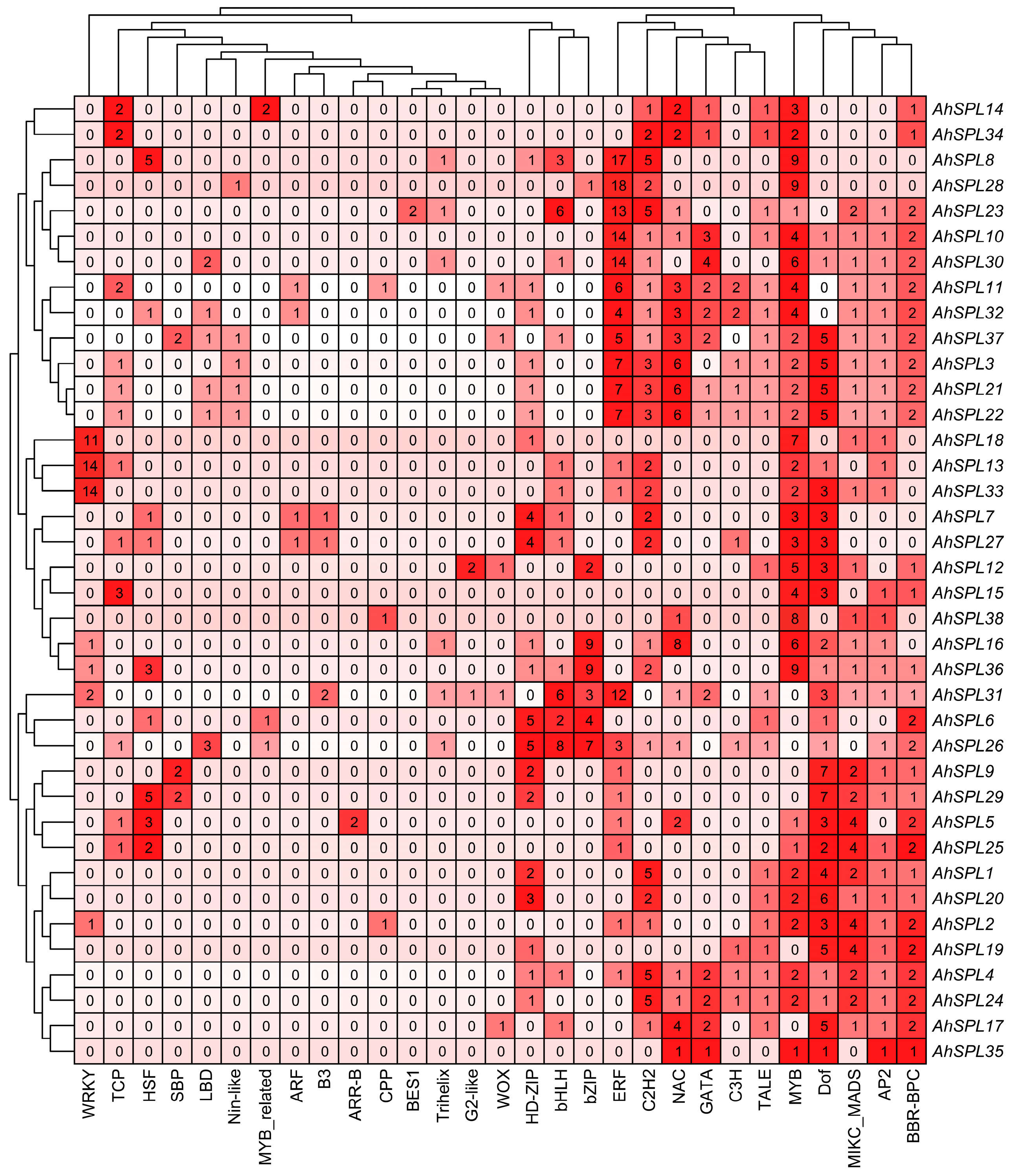
2.5. Cis-Acting Elements Analysis
2.6. Expression Pattern Analysis
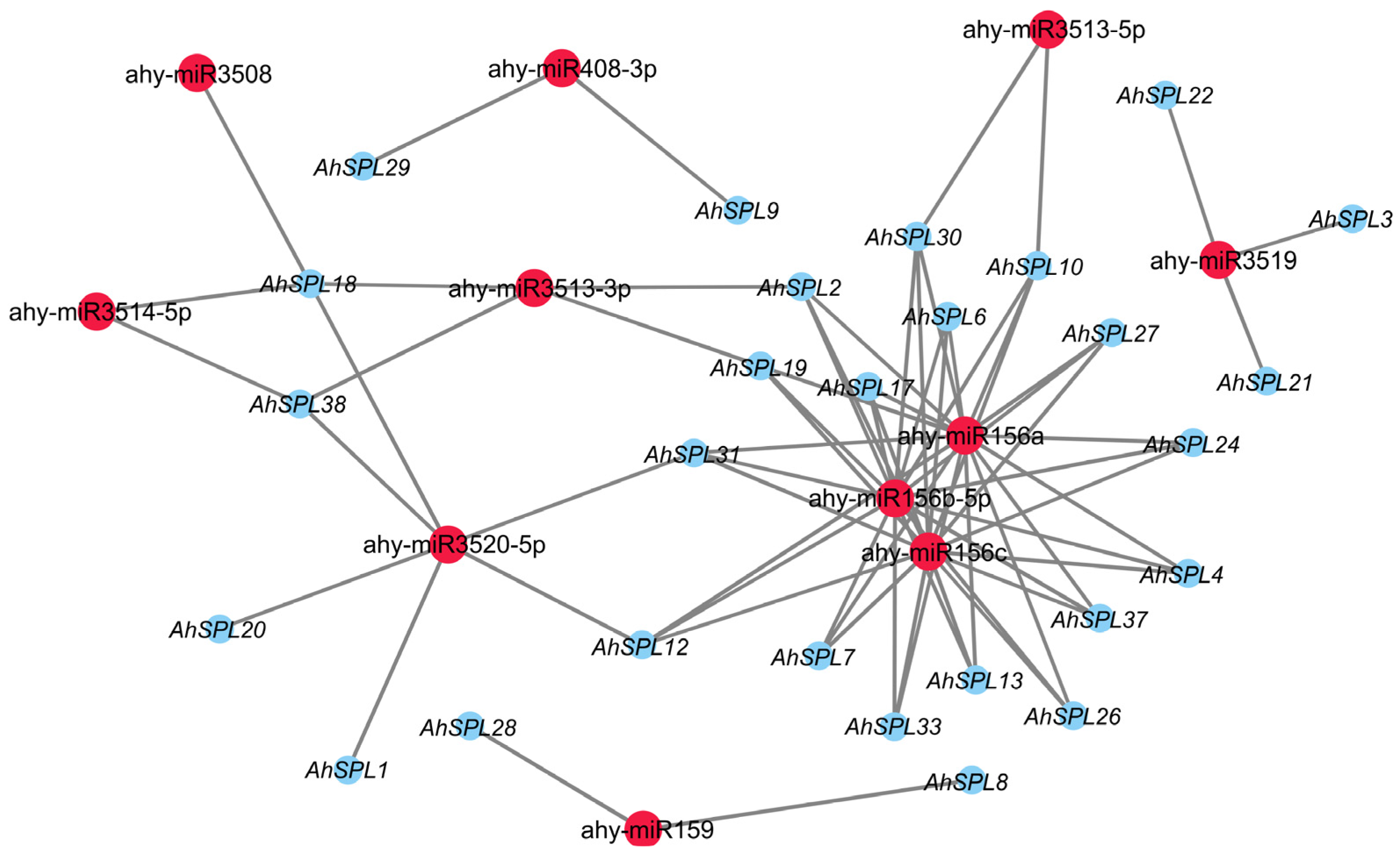
2.7. Prediction of Regulatory Network
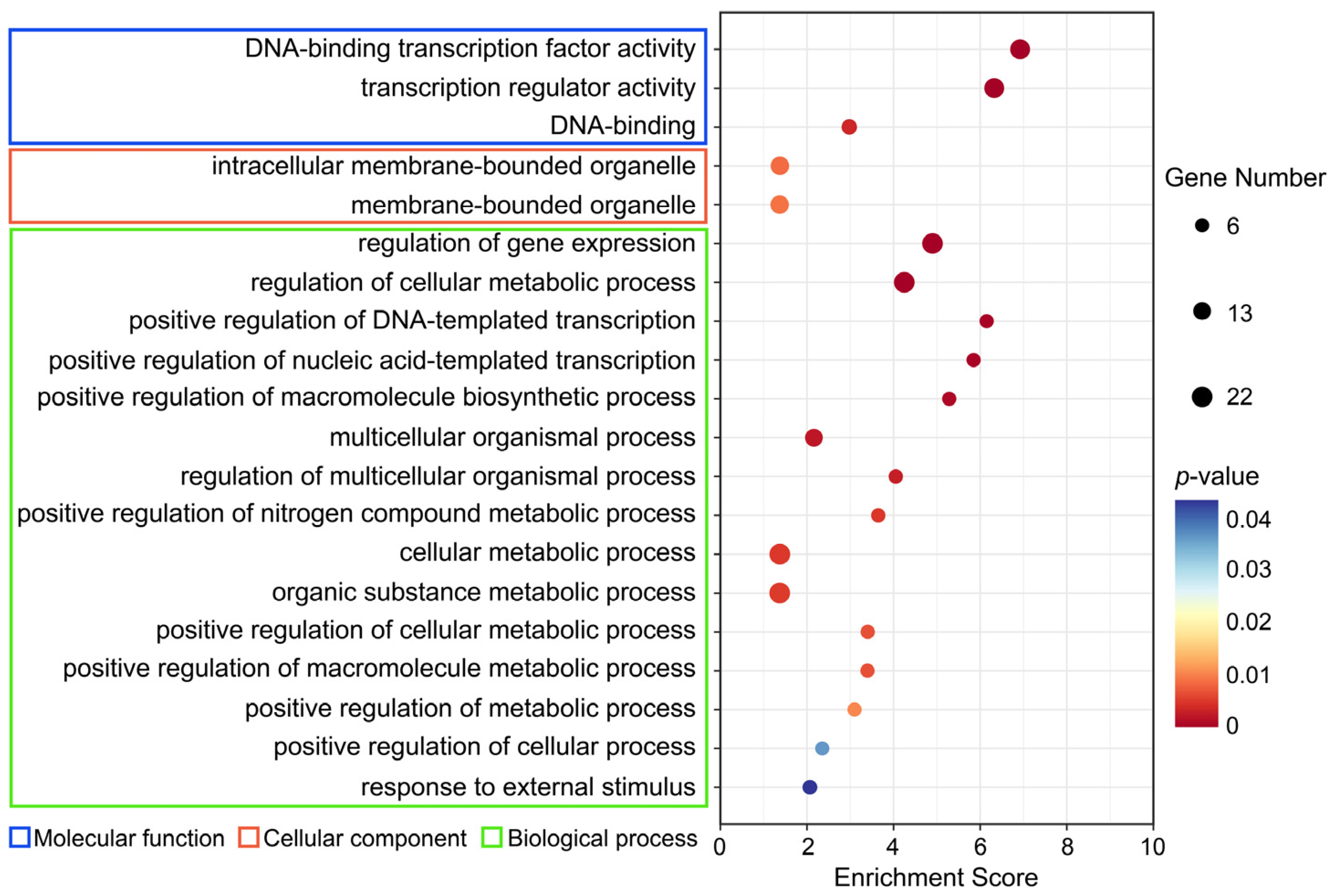
2.8. GO Enrichment Analysis
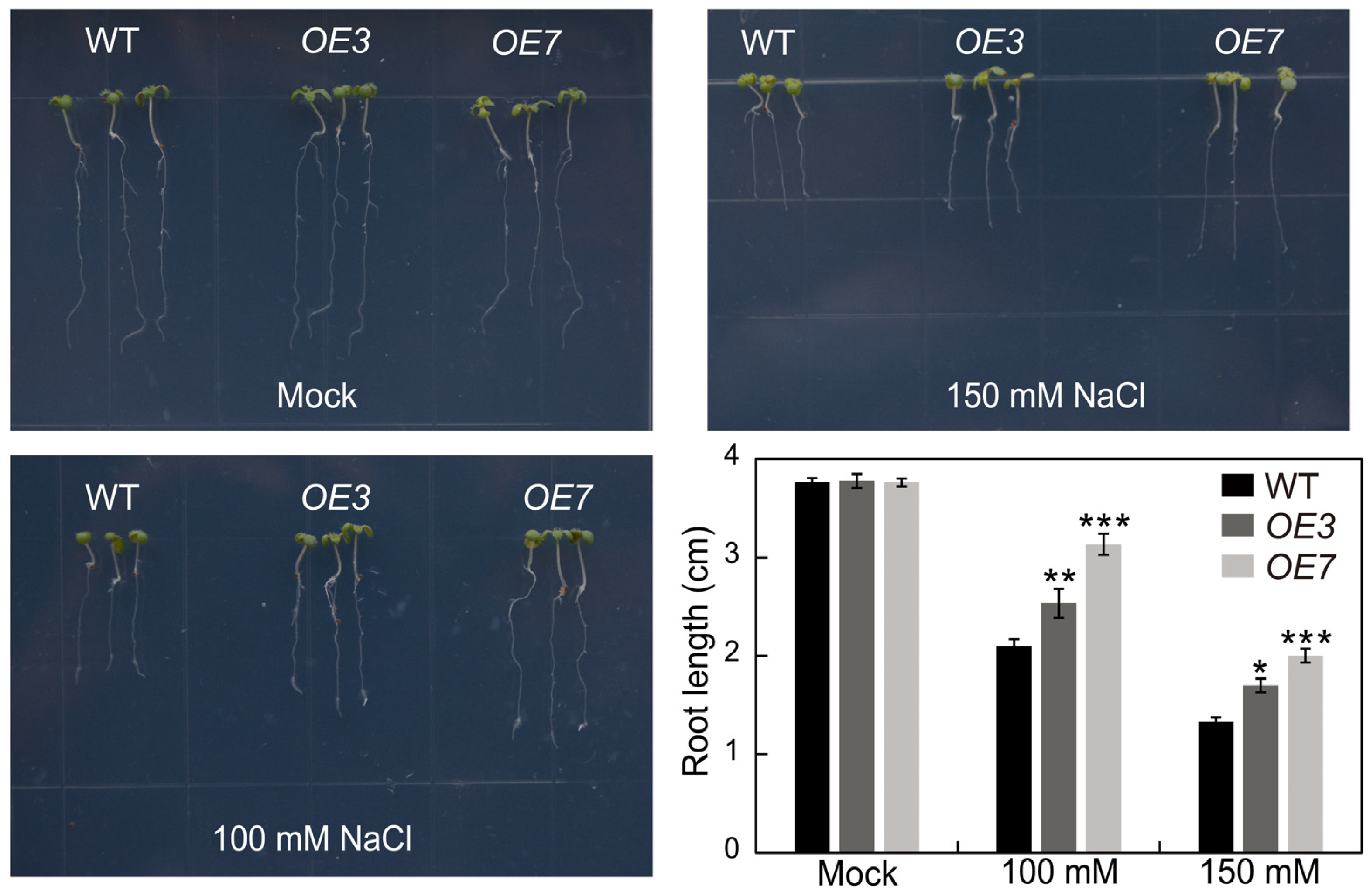
2.9. Overexpression of AhSPL5 Enhanced Salt Tolerance
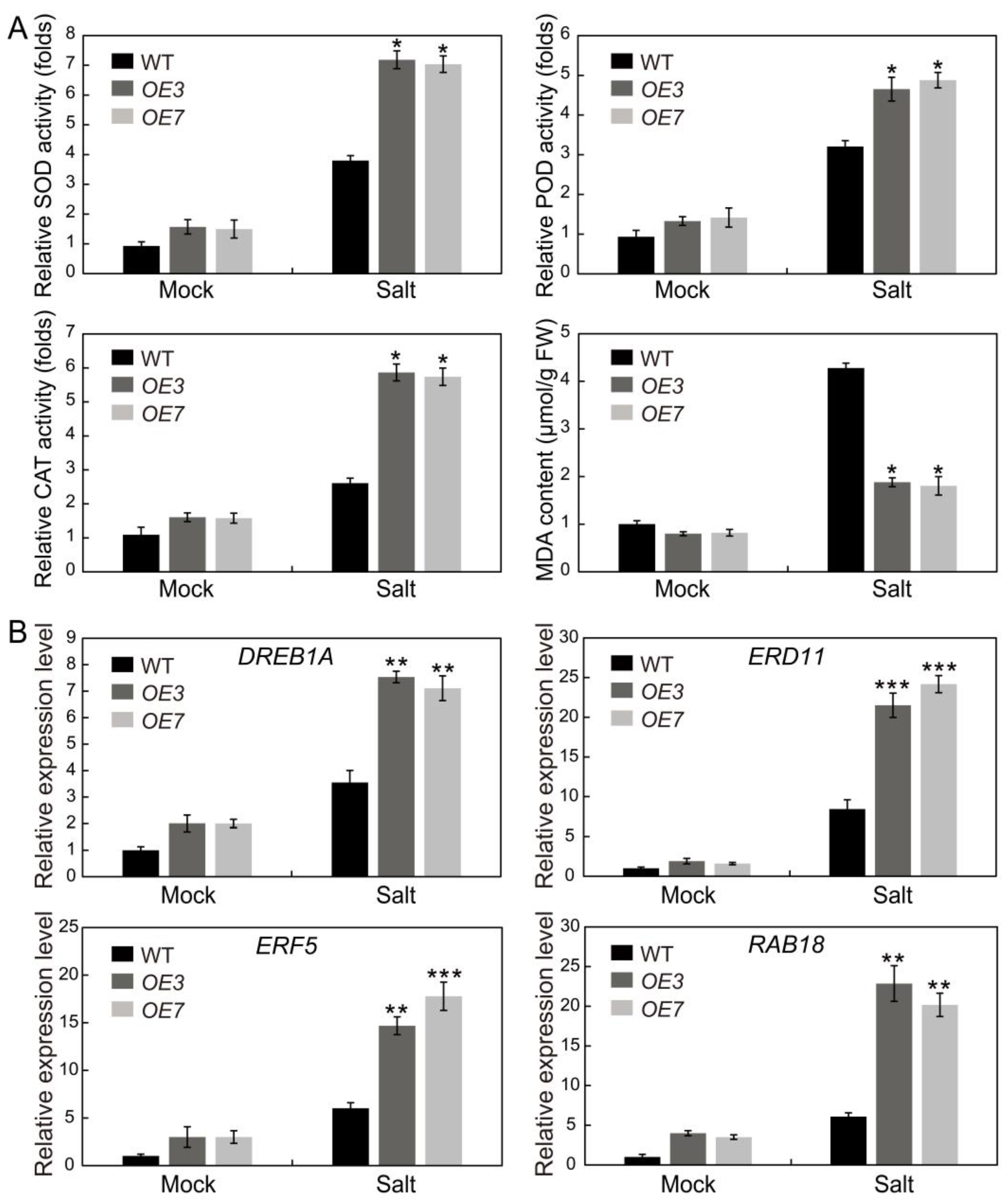
2.10. AhSPL5 Enhances ROS-Scavenging Capability and Regulates the Expression of Stress-Responsive Genes
3. Discussion
4. Materials and Methods
4.1. Identification and Annotation of SPL Transcription Factor Family in Peanuts
4.2. Multiple Sequence Alignment and Analysis of Phylogenetics
4.3. Gene Structure and Conserved Domains
4.4. Chromosomal Location and Gene Duplication
4.5. Analysis of Cis-Acting Elements
4.6. Expression Pattern Analysis
4.7. Prediction of Factors Involved in Regulating AhSPLs’ Expression
4.8. GO (Gene Ontology) Enrichment Analysis
4.9. RNA Extraction and qRT-PCR
4.10. Generation of AhSPL5 Transgenic Arabidopsis Plants
4.11. Seedling Growth Assays
4.12. Physiological Measurements
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Bertioli, D.J.; Jenkins, J.; Clevenger, J.; Dudchenko, O.; Gao, D.; Seijo, G.; Leal-Bertioli, S.C.M.; Ren, L.; Farmer, A.D.; Pandey, M.K.; et al. The genome sequence of segmental allotetraploid peanut Arachis hypogaea. Nat. Genet. 2019, 51, 877–884. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Lu, Q.; Liu, H.; Zhang, J.; Hong, Y.; Lan, H.; Li, H.; Wang, J.; Liu, H.; Li, S.; et al. Sequencing of Cultivated Peanut, Arachis hypogaea, Yields Insights into Genome Evolution and Oil Improvement. Mol. Plant 2019, 12, 920–934. [Google Scholar] [CrossRef]
- Zhuang, W.; Chen, H.; Yang, M.; Wang, J.; Pandey, M.K.; Zhang, C.; Chang, W.C.; Zhang, L.; Zhang, X.; Tang, R.; et al. The genome of cultivated peanut provides insight into legume karyotypes, polyploid evolution and crop domestication. Nat. Genet. 2019, 51, 865–876. [Google Scholar] [CrossRef] [PubMed]
- Riechmann, J.L.; Ratcliffe, O.J. A genomic perspective on plant transcription factors. Curr. Opin. Plant Biol. 2000, 3, 423–434. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Li, X.; Zhang, J.; Yi, D.; Li, F.; Wen, H.; Liu, W.; Wang, X. MsWRKY33 increases alfalfa (Medicago sativa L.) salt stress tolerance through altering the ROS scavenger via activating MsERF5 transcription. Plant Cell Environ. 2023, 46, 3887–3901. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.; Liu, S.; Zhang, C.; He, J.; Ma, D.; Wang, X.; Dong, T.; Guo, F.; Cai, J.; Long, T.; et al. The unique sweet potato NAC transcription factor IbNAC3 modulates combined salt and drought stresses. Plant Physiol. 2023, 191, 747–771. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Guo, C.; Li, Z.; Wang, G.; Yang, J.; Chen, L.; Hu, Z.; Sun, J.; Gao, J.; Yang, A.; et al. Deciphering the roles of tobacco MYB transcription factors in environmental stress tolerance. Front. Plant Sci. 2022, 13, 998606. [Google Scholar] [CrossRef]
- Lai, D.; Fan, Y.; Xue, G.; He, A.; Yang, H.; He, C.; Li, Y.; Ruan, J.; Yan, J.; Cheng, J. Genome-wide identification and characterization of the SPL gene family and its expression in the various developmental stages and stress conditions in foxtail millet (Setaria italica). BMC Genom. 2022, 23, 389. [Google Scholar] [CrossRef] [PubMed]
- Klein, J.; Saedler, H.; Huijser, P. A new family of DNA binding proteins includes putative transcriptional regulators of the Antirrhinum majus floral meristem identity gene SQUAMOSA. Mol. Gen. Genet. 1996, 250, 7–16. [Google Scholar]
- Yang, Z.; Wang, X.; Gu, S.; Hu, Z.; Xu, H.; Xu, C. Comparative study of SBP-box gene family in Arabidopsis and rice. Gene 2008, 407, 1–11. [Google Scholar] [CrossRef]
- Peng, X.; Wang, Q.; Zhao, Y.; Li, X.; Ma, Q. Comparative genome analysis of the SPL gene family reveals novel evolutionary features in maize. Genet. Mol. Biol. 2019, 42, 380–394. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Wang, Y.; Xiao, Y.; Zhang, X.; Du, B.; Turupu, M.; Wang, C.; Yao, Q.; Gai, S.; Huang, J.; et al. Genome-Wide Identification of the SQUAMOSA Promoter-Binding Protein-like (SPL) Transcription Factor Family in Sweet Cherry Fruit. Int. J. Mol. Sci. 2023, 24, 2880. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Cao, H.; Zhang, M.; Deng, S.; Li, T.; Xing, S. Genome-Wide Identification and Characterization of SPL Family Genes in Chenopodium quinoa. Genes 2022, 13, 1455. [Google Scholar] [CrossRef] [PubMed]
- Long, J.M.; Liu, C.Y.; Feng, M.Q.; Liu, Y.; Wu, X.M.; Guo, W.W. miR156-SPL modules regulate induction of somatic embryogenesis in citrus callus. J. Exp. Bot. 2018, 69, 2979–2993. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Zhang, Z.; Liu, D.; Zhang, K.; Li, A.; Mao, L. SQUAMOSA promoter-binding protein-like transcription factors: Star players for plant growth and development. J. Integr. Plant Biol. 2010, 52, 946–951. [Google Scholar] [CrossRef] [PubMed]
- Liu, N.; Tu, L.; Wang, L.; Hu, H.; Xu, J.; Zhang, X. MicroRNA 157-targeted SPL genes regulate floral organ size and ovule production in cotton. BMC Plant Biol. 2017, 17, 7. [Google Scholar] [CrossRef] [PubMed]
- Huijser, P.; Schmid, M. The control of developmental phase transitions in plants. Development 2011, 138, 4117–4129. [Google Scholar] [CrossRef] [PubMed]
- Xing, S.; Salinas, M.; Höhmann, S.; Berndtgen, R.; Huijser, P. miR156-targeted and nontargeted SBP-box transcription factors act in concert to secure male fertility in Arabidopsis. Plant Cell 2010, 22, 3935–3950. [Google Scholar] [CrossRef] [PubMed]
- Miura, K.; Ikeda, M.; Matsubara, A.; Song, X.J.; Ito, M.; Asano, K.; Matsuoka, M.; Kitano, H.; Ashikari, M. OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat. Genet. 2010, 42, 545–549. [Google Scholar] [CrossRef]
- Jiao, Y.; Wang, Y.; Xue, D.; Wang, J.; Yan, M.; Liu, G.; Dong, G.; Zeng, D.; Lu, Z.; Zhu, X.; et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 2010, 42, 541–544. [Google Scholar] [CrossRef]
- Wang, S.; Wu, K.; Yuan, Q.; Liu, X.; Liu, Z.; Lin, X.; Zeng, R.; Zhu, H.; Dong, G.; Qian, Q.; et al. Control of grain size, shape and quality by OsSPL16 in rice. Nat. Genet. 2012, 44, 950–954. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Li, S.; Liu, Q.; Wu, K.; Zhang, J.; Wang, S.; Wang, Y.; Chen, X.; Zhang, Y.; Gao, C.; et al. The OsSPL16-GW7 regulatory module determines grain shape and simultaneously improves rice yield and grain quality. Nat. Genet. 2015, 47, 949–954. [Google Scholar] [CrossRef] [PubMed]
- Jerome Jeyakumar, J.M.; Ali, A.; Wang, W.M.; Thiruvengadam, M. Characterizing the Role of the miR156-SPL Network in Plant Development and Stress Response. Plants 2020, 9, 1206. [Google Scholar] [CrossRef] [PubMed]
- Dong, H.; Yan, S.; Jing, Y.; Yang, R.; Zhang, Y.; Zhou, Y.; Zhu, Y.; Sun, J. MIR156-Targeted SPL9 Is Phosphorylated by SnRK2s and Interacts with ABI5 to Enhance ABA Responses in Arabidopsis. Front. Plant Sci. 2021, 12, 708573. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Schwarz, S.; Saedler, H.; Huijser, P. SPL8, a local regulator in a subset of gibberellin-mediated developmental processes in Arabidopsis. Plant Mol. Biol. 2007, 63, 429–439. [Google Scholar] [CrossRef] [PubMed]
- Padmanabhan, M.S.; Ma, S.; Burch-Smith, T.M.; Czymmek, K.; Huijser, P.; Dinesh-Kumar, S.P. Novel positive regulatory role for the SPL6 transcription factor in the N TIR-NB-LRR receptor-mediated plant innate immunity. PLoS Pathog. 2013, 9, e1003235. [Google Scholar] [CrossRef] [PubMed]
- Chao, L.M.; Liu, Y.Q.; Chen, D.Y.; Xue, X.Y.; Mao, Y.B.; Chen, X.Y. Arabidopsis Transcription Factors SPL1 and SPL12 Confer Plant Thermotolerance at Reproductive Stage. Mol. Plant. 2017, 10, 735–748. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Shi, M.; Yu, J.; Guo, C. SPL9 mediates freezing tolerance by directly regulating the expression of CBF2 in Arabidopsis thaliana. BMC Plant Biol. 2022, 22, 59. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; He, J.; Liu, M.; Miao, J.; Ma, C.; Feng, Y.; Qian, J.; Li, H.; Bi, H.; Liu, W. The SPL transcription factor TaSPL6 negatively regulates drought stress response in wheat. Plant Physiol. Biochem. 2024, 206, 108264. [Google Scholar] [CrossRef]
- Li, Y.; Han, S.; Sun, X.; Khan, N.U.; Zhong, Q.; Zhang, Z.; Zhang, H.; Ming, F.; Li, Z.; Li, J. Variations in OsSPL10 confer drought tolerance by directly regulating OsNAC2 expression and ROS production in rice. J. Integr. Plant Biol. 2023, 65, 918–933. [Google Scholar] [CrossRef]
- Yue, E.; Liu, Z.; Li, C.; Li, Y.; Liu, Q.; Xu, J.H. Overexpression of miR529a confers enhanced resistance to oxidative stress in rice (Oryza sativa L.). Plant Cell Rep. 2017, 36, 1171–1182. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Xue, H.; Zhang, F.; Jiang, Q.; Yang, S.; Yue, P.; Wang, F.; Zhang, Y.; Li, L.; He, P.; et al. The miR156/SPL module regulates apple salt stress tolerance by activating MdWRKY100 expression. Plant Biotechnol. J. 2021, 19, 311–323. [Google Scholar] [CrossRef]
- Wang, Q.; Zhang, Z.; Guo, C.; Zhao, X.; Li, Z.; Mou, Y.; Sun, Q.; Wang, J.; Yuan, C.; Li, C.; et al. Hsf transcription factor gene family in peanut (Arachis hypogaea L.): Genome-wide characterization and expression analysis under drought and salt stresses. Front. Plant Sci. 2023, 14, 1214732. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Shan, S. Deciphering the roles of peanut (Arachis hypogaea L.) type-one protein phosphatase (TOPP) family in abiotic stress tolerance. Agronomy 2023, 13, 2444. [Google Scholar] [CrossRef]
- Wang, Q.; Zhao, X.; Sun, Q.; Mou, Y.; Wang, J.; Yan, C.; Yuan, C.; Li, C.; Shan, S. Genome-wide identification of the LRR-RLK gene family in peanut and functional characterization of AhLRR-RLK265 in salt and drought stresses. Int. J. Biol. Macromol. 2024, 254, 127829. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Zhao, S.Z.; Zhao, C.Z.; Zhang, Y.; Xia, H.; Lopez-Baltazar, J.; Wan, S.B.; Wang, X.J. Cloning and characterization of SPL-family genes in the peanut (Arachis hypogaea L.). Genet. Mol. Res. 2016, 15, gmr.15017344. [Google Scholar] [CrossRef] [PubMed]
- Zheng, C.; Ye, M.; Sang, M.; Wu, R. A Regulatory Network for miR156-SPL Module in Arabidopsis thaliana. Int. J. Mol. Sci. 2019, 20, 6166. [Google Scholar] [CrossRef] [PubMed]
- Kavas, M.; Kızıldoğan, A.K.; Abanoz, B. Comparative genome-wide phylogenetic and expression analysis of SBP genes from potato (Solanum tuberosum). Comput. Biol. Chem. 2017, 67, 131–140. [Google Scholar] [CrossRef] [PubMed]
- Salinas, M.; Xing, S.; Höhmann, S.; Berndtgen, R.; Huijser, P. Genomic organization, phylogenetic comparison and differential expression of the SBP-box family of transcription factors in tomato. Planta 2012, 235, 1171–1184. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, R.K.; Goel, R.; Kumari, S.; Dahuja, A. Genomic organization, phylogenetic comparison, and expression profiles of the SPL family genes and their regulation in soybean. Dev. Genes Evol. 2017, 227, 101–119. [Google Scholar] [CrossRef]
- Zhou, Q.; Zhang, S.; Chen, F.; Liu, B.; Wu, L.; Li, F.; Zhang, J.; Bao, M.; Liu, G. Genome-wide identification and characterization of the SBP-box gene family in Petunia. BMC Genom. 2018, 19, 193. [Google Scholar] [CrossRef] [PubMed]
- Zhu, T.; Liu, Y.; Ma, L.; Wang, X.; Zhang, D.; Han, Y.; Ding, Q.; Ma, L. Genome-wide identification, phylogeny and expression analysis of the SPL gene family in wheat. BMC Plant Biol. 2020, 20, 420. [Google Scholar] [CrossRef] [PubMed]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef] [PubMed]
- Perin, E.C.; da Silva Messias, R.; Borowski, J.M.; Crizel, R.L.; Schott, I.B.; Carvalho, I.R.; Rombaldi, C.V.; Galli, V. ABA-dependent salt and drought stress improve strawberry fruit quality. Food Chem. 2019, 271, 516–526. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Liu, J.; Liu, E.; Qiao, K.; Gong, S.; Wang, J.; Zhou, A.; Zhang, J. Arabidopsis cold-regulated plasma membrane protein Cor413pm1 is a regulator of ABA response. Biochem. Biophys. Res. Commun. 2021, 561, 88–92. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.C.; Niu, C.Y.; Yang, C.R.; Jinn, T.L. The heat stress factor HSFA6b connects ABA signaling and ABA-mediated heat responses. Plant Physiol. 2016, 172, 1182–1199. [Google Scholar] [CrossRef]
- Olive, M.R.; Walker, J.C.; Singh, K.; Dennis, E.S.; Peacock, W.J. Functional properties of the anaerobic responsive element of the maize Adh1 gene. Plant Mol. Biol. 1990, 15, 593–604. [Google Scholar] [CrossRef]
- Yin, H.; Hong, G.; Li, L.; Zhang, X.; Kong, Y.; Sun, Z.; Li, J.; Chen, J.; He, Y. miR156/SPL9 regulates reactive oxygen species accumulation and immune response in Arabidopsis thaliana. Phytopathology 2019, 109, 632–642. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Ruan, Q.; Zhu, X.; Wang, B.; Wei, B.; Wei, X. Identification of Alfalfa SPL gene family and expression analysis under biotic and abiotic stresses. Sci. Rep. 2023, 13, 84. [Google Scholar] [CrossRef]
- He, B.; Gao, S.; Lu, H.; Yan, J.; Li, C.; Ma, M.; Wang, X.; Chen, X.; Zhan, Y.; Zeng, F. Genome-wide analysis and molecular dissection of the SPL gene family in Fraxinus mandshurica. BMC Plant Biol. 2022, 22, 451. [Google Scholar] [CrossRef]
- Wang, Q.; Li, X.; Guo, C.; Wen, L.; Deng, Z.; Zhang, Z.; Li, W.; Liu, T.; Guo, Y. Senescence-related receptor kinase 1 functions downstream of WRKY53 in regulating leaf senescence in Arabidopsis. J. Exp. Bot. 2023, 74, 5140–5152. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Wang, Q.; Yan, C.; Sun, Q.; Wang, J.; Li, C.; Yuan, C.; Mou, Y.; Shan, S. The bHLH transcription factor AhbHLH121 improves salt tolerance in peanut. Int. J. Biol. Macromol. 2024, 256, 128492. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Hu, T.; Zhao, J.; Park, M.Y.; Earley, K.W.; Wu, G.; Yang, L.; Poethig, R.S. Developmental functions of miR156-regulated SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) genes in Arabidopsis thaliana. PLoS Genet. 2016, 12, e1006263. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Lu, S. Molecular characterization of the SPL gene family in Populus trichocarpa. BMC Plant Biol. 2014, 14, 131. [Google Scholar] [CrossRef] [PubMed]
- Quevillon, E.; Silventoinen, V.; Pillai, S.; Harte, N.; Mulder, N.; Apweiler, R.; Lopez, R. InterProScan: Protein domains identifier. Nucleic Acids Res. 2005, 33, W116–W120. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. 20 years of the SMART protein domain annotation resource. Nucleic Acids Res. 2018, 46, D493–D496. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.C.; Williams, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein identification and analysis tools in the ExPASy server. Methods Mol. Biol. 1999, 112, 531–552. [Google Scholar] [PubMed]
- Wang, Q.; Guo, C.; Li, Z.; Sun, J.; Deng, Z.; Wen, L.; Li, X.; Guo, Y. Potato NAC transcription factor StNAC053 enhances salt and drought tolerance in transgenic Arabidopsis. Int. J. Mol. Sci. 2021, 22, 2568. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, W202–W208. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant. 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef]
- Librado, P.; Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Clevenger, J.; Chu, Y.; Scheffler, B.; Ozias-Akins, P. A Developmental transcriptome map for allotetraploid Arachis hypogaea. Front. Plant Sci. 2016, 7, 1446. [Google Scholar] [CrossRef]
- Zhang, H.; Zhao, X.; Sun, Q.; Yan, C.; Wang, J.; Yuan, C.; Li, C.; Shan, S.; Liu, F. Comparative transcriptome analysis reveals molecular defensive mechanism of Arachis hypogaea in response to salt stress. Int. J. Genom. 2020, 2020, 6524093. [Google Scholar] [CrossRef]
- Zhao, X.; Li, C.; Wan, S.; Zhang, T.; Yan, C.; Shan, S. Transcriptomic analysis and discovery of genes in the response of Arachis hypogaea to drought stress. Mol. Biol. Rep. 2018, 45, 119–131. [Google Scholar] [CrossRef]
- Tian, F.; Yang, D.C.; Meng, Y.Q.; Jin, J.; Gao, G. PlantRegMap: Charting functional regulatory maps in plants. Nucleic Acids Res. 2020, 48, D1104–D1113. [Google Scholar] [CrossRef]
- Dai, X.; Zhuang, Z.; Zhao, P.X. psRNATarget: A plant small RNA target analysis server (2017 release). Nucleic Acids Res. 2018, 46, W49–W54. [Google Scholar] [CrossRef] [PubMed]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Powell, S.; Forslund, K.; Szklarczyk, D.; Trachana, K.; Roth, A.; Huerta-Cepas, J.; Gabaldón, T.; Rattei, T.; Creevey, C.; Kuhn, M.; et al. eggNOG v4.0: Nested orthology inference across 3686 organisms. Nucleic Acids Res. 2014, 42, D231–D239. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Henriques, R.; Lin, S.S.; Niu, Q.W.; Chua, N.H. Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat. Protoc. 2006, 1, 641–646. [Google Scholar] [CrossRef]
- He, K.; Zhao, X.; Chi, X.; Wang, Y.; Jia, C.; Zhang, H.; Zhou, G.; Hu, R. A novel Miscanthus NAC transcription factor MlNAC10 enhances drought and salinity tolerance in transgenic Arabidopsis. J. Plant Physiol. 2019, 233, 84–93. [Google Scholar] [CrossRef]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sun, X.; Zhang, L.; Xu, W.; Zheng, J.; Yan, M.; Zhao, M.; Wang, X.; Yin, Y. A Comprehensive Analysis of the Peanut SQUAMOSA Promoter Binding Protein-like Gene Family and How AhSPL5 Enhances Salt Tolerance in Transgenic Arabidopsis. Plants 2024, 13, 1057. https://doi.org/10.3390/plants13081057
Sun X, Zhang L, Xu W, Zheng J, Yan M, Zhao M, Wang X, Yin Y. A Comprehensive Analysis of the Peanut SQUAMOSA Promoter Binding Protein-like Gene Family and How AhSPL5 Enhances Salt Tolerance in Transgenic Arabidopsis. Plants. 2024; 13(8):1057. https://doi.org/10.3390/plants13081057
Chicago/Turabian StyleSun, Xiaohui, Lili Zhang, Weihua Xu, Jianpeng Zheng, Meiling Yan, Ming Zhao, Xinyu Wang, and Yan Yin. 2024. "A Comprehensive Analysis of the Peanut SQUAMOSA Promoter Binding Protein-like Gene Family and How AhSPL5 Enhances Salt Tolerance in Transgenic Arabidopsis" Plants 13, no. 8: 1057. https://doi.org/10.3390/plants13081057
APA StyleSun, X., Zhang, L., Xu, W., Zheng, J., Yan, M., Zhao, M., Wang, X., & Yin, Y. (2024). A Comprehensive Analysis of the Peanut SQUAMOSA Promoter Binding Protein-like Gene Family and How AhSPL5 Enhances Salt Tolerance in Transgenic Arabidopsis. Plants, 13(8), 1057. https://doi.org/10.3390/plants13081057




