Genome-Wide Development of Polymorphic Microsatellite Markers and Genetic Diversity Analysis for the Halophyte Suaeda aralocaspica (Amaranthaceae)
Abstract
1. Introduction
2. Results
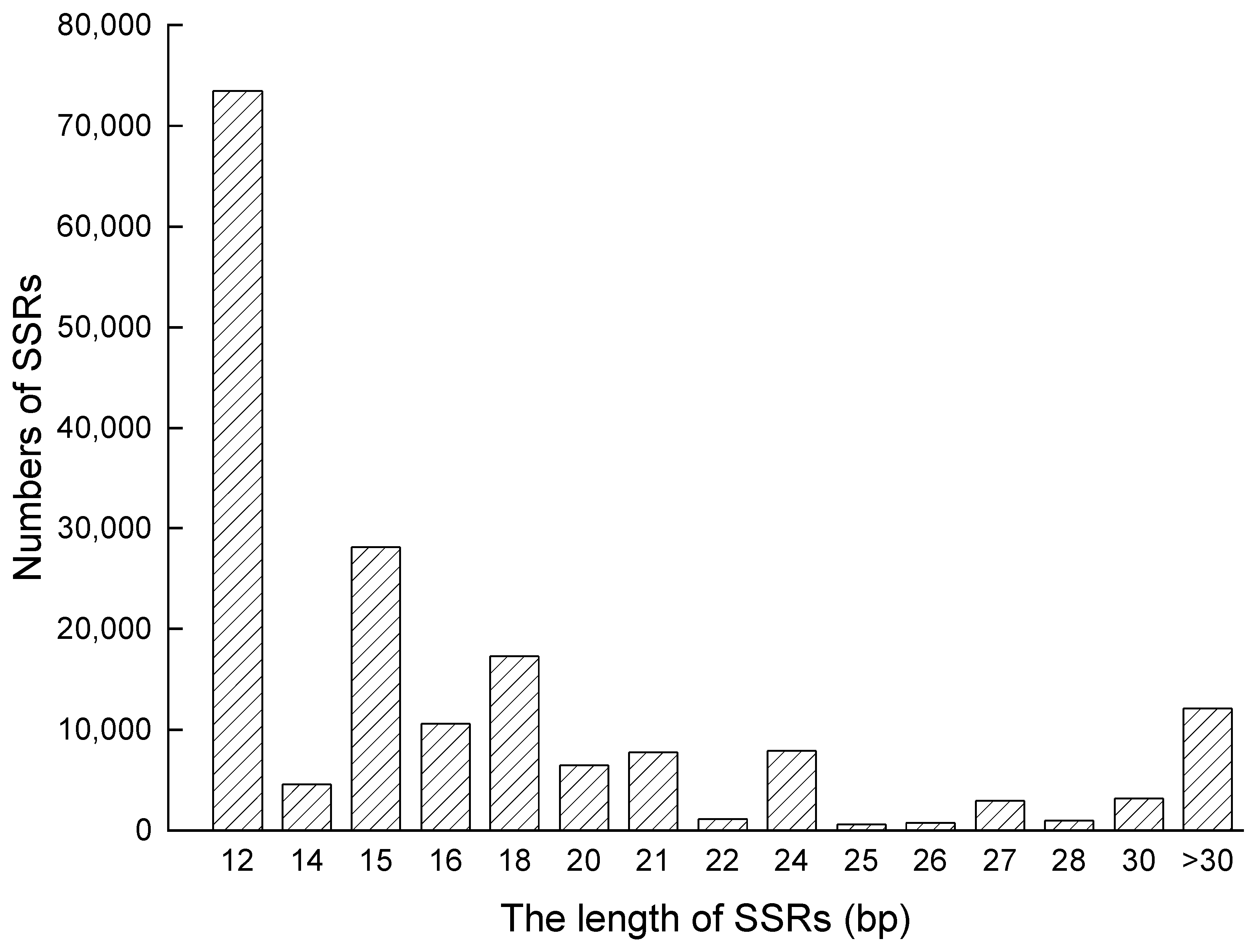
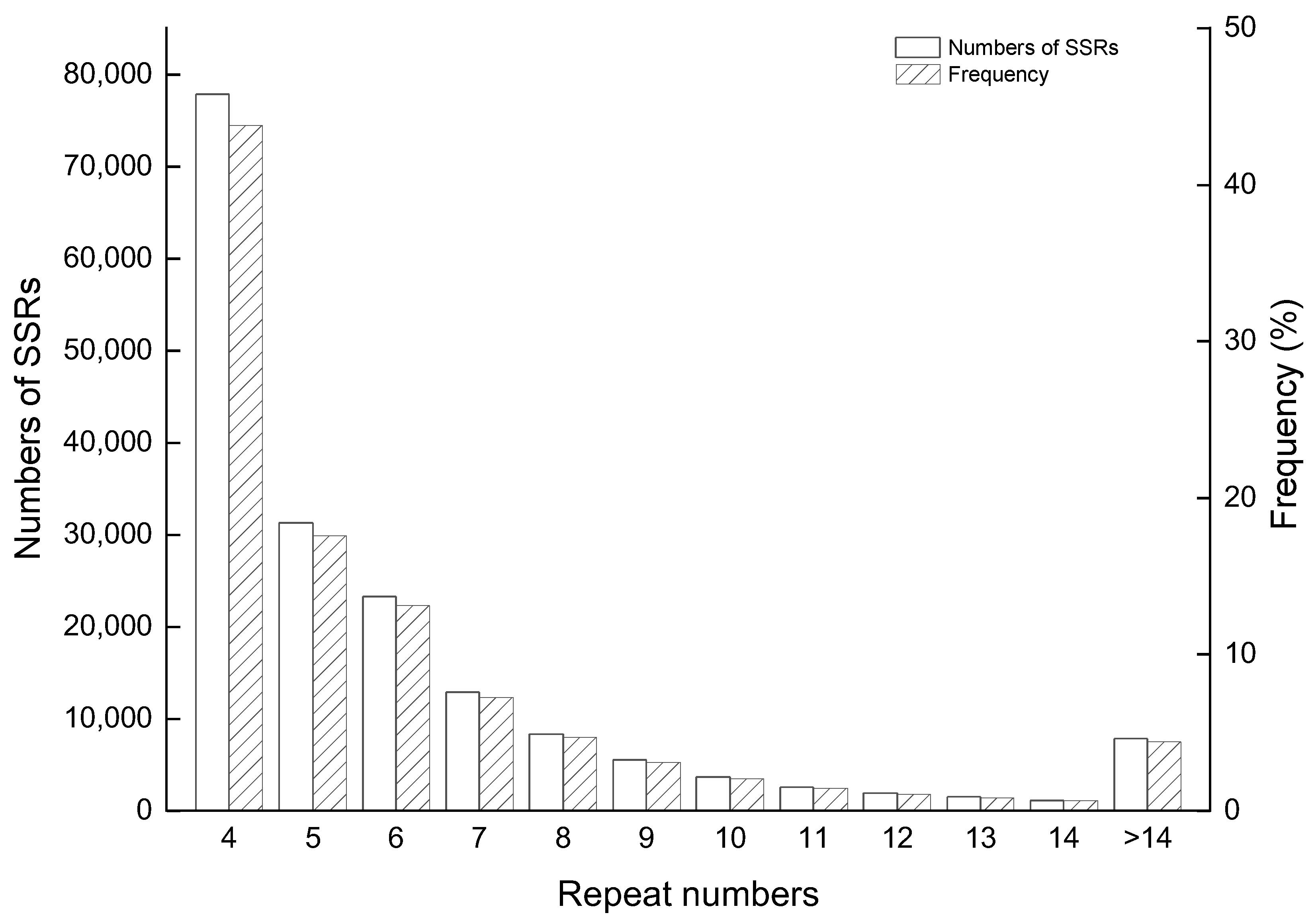
2.1. Analysis of the Distribution of SSRs in the Genome of S. aralocaspica
2.2. Development of Genome SSR Markers for S. aralocaspica
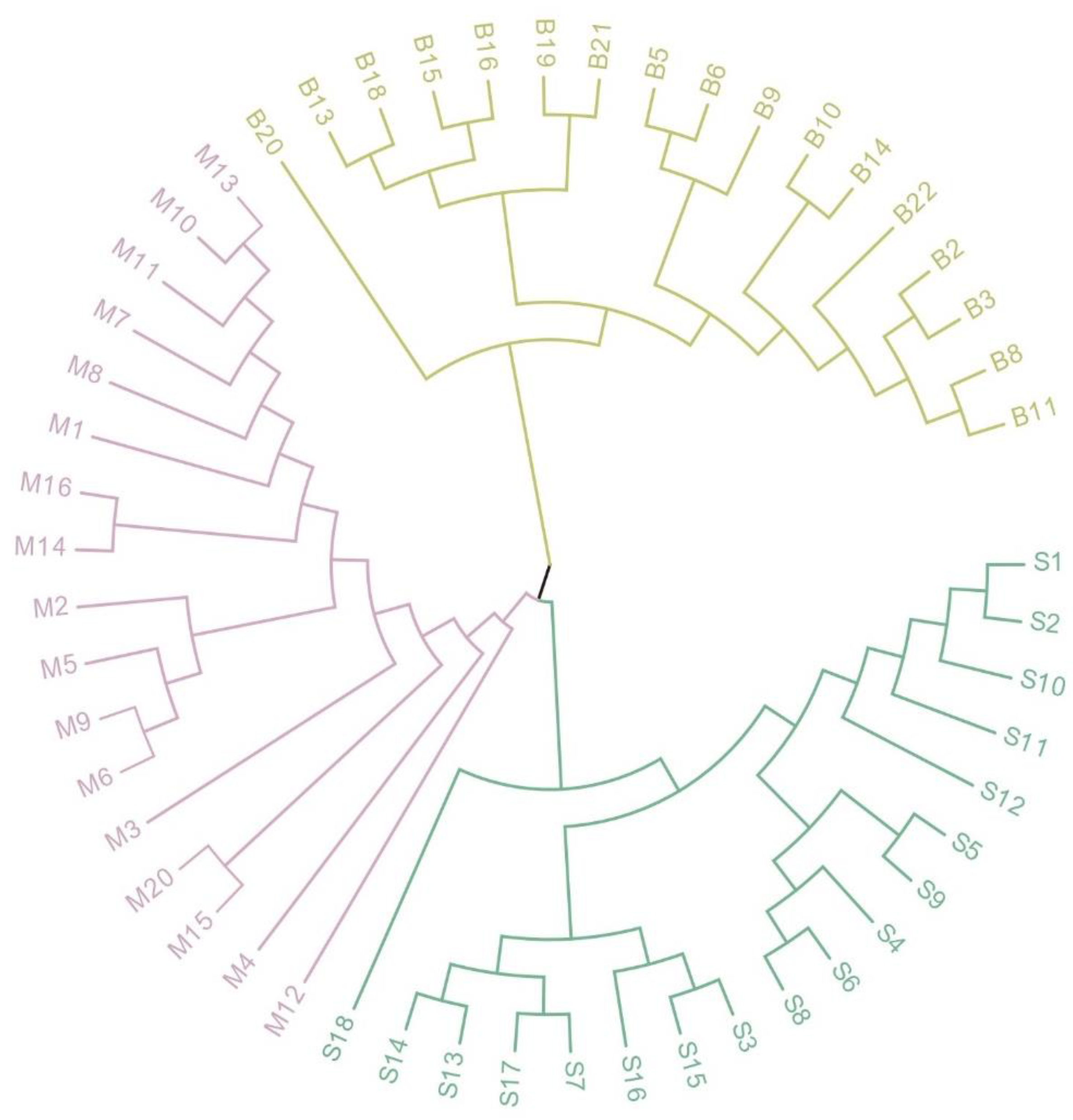
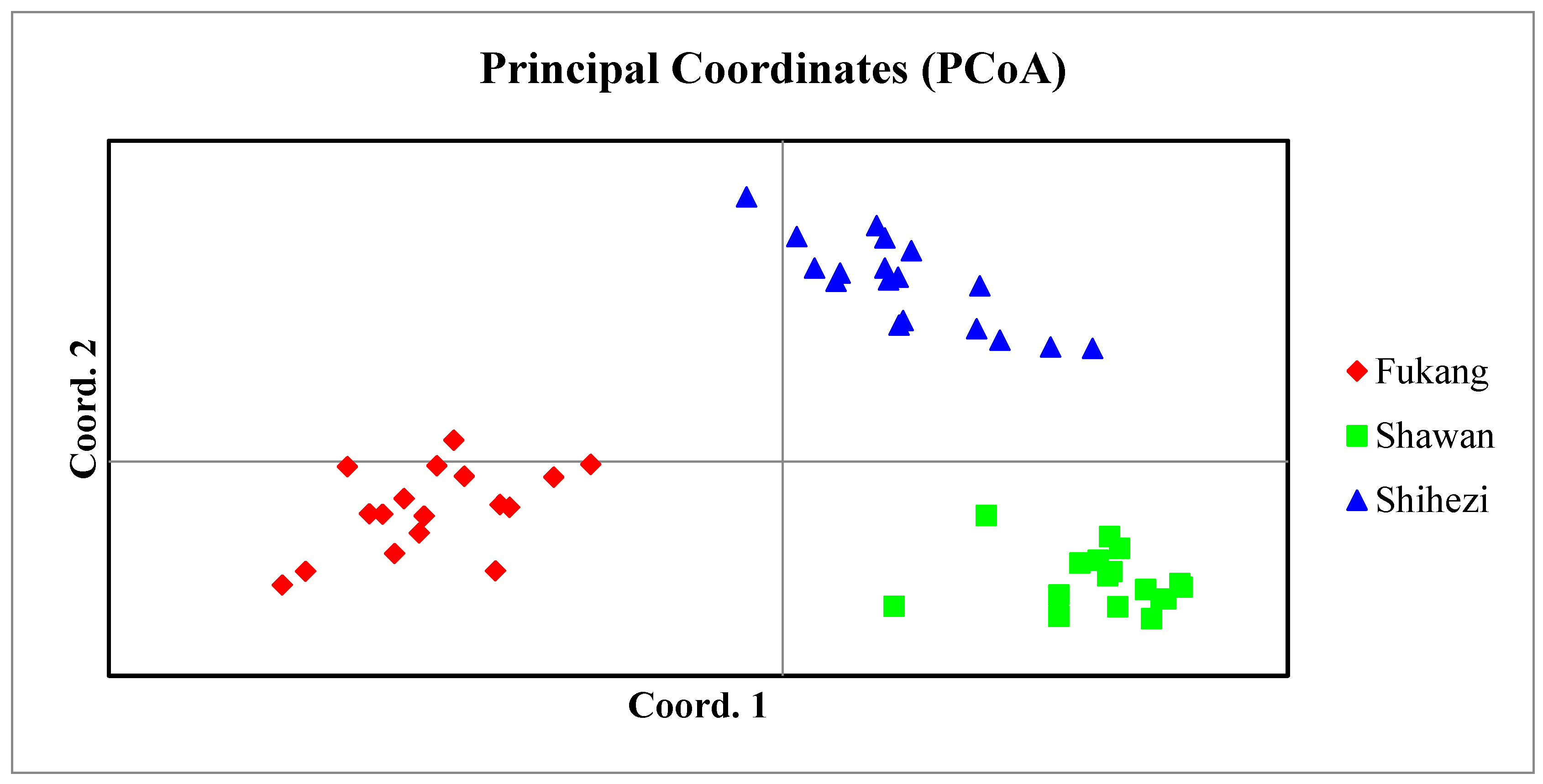
2.3. Genetic Diversity Analysis of S. aralocaspica Populations
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Genome SSR Identification and Development
4.3. PCR Amplification, and Electrophoresis Detection
4.4. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Huang, J.X.; Cheng, W.N.; Li, Y.L.; Yao, G. Reconstruction of phylogenetic relationships of Amaranthaceae based on multiple chloroplast gene sequence fragments. Chin. Bull. Botany 2022, 55, 457–467, (In Chinese with English Abstract). [Google Scholar]
- Wang, X.Y.; Shao, X.T.; Zhang, W.J.; Sun, T.; Ding, Y.L.; Lin, Z.; Li, Y. Genus Suaeda: Advances in phytology, chemistry, pharmacology and clinical application (1895–2021). Pharmacol. Res. 2022, 179, 106203. [Google Scholar] [CrossRef] [PubMed]
- Yadav, S.; Elansary, H.O.; Mattar, M.A.; Elhindi, K.M.; Alotaibi, M.A.; Mishra, A. Differential accumulation of metabolites in Suaeda species provides new insights into abiotic stress tolerance in C4-halophytic species in elevated CO2 conditions. Agronomy 2021, 11, 131. [Google Scholar] [CrossRef]
- Zhang, A.Q.; Pang, Q.Y.; Yan, A.F. Advances in salt-tolerance mechanisms of Suaeda plants. Acta Ecol. Sin. 2013, 33, 3575–3583, (In Chinese with English Abstract). [Google Scholar] [CrossRef]
- Wang, L.; Zhao, Z.Y.; Zhang, K.; Tian, C.Y. Oil content and fatty acid composition of dimorphic seeds of desert halophyte Suaeda aralocaspica. Afr. J. Agric. Res. 2012, 7, 1910–1914. [Google Scholar]
- Schütze, P.; Freitag, H.; Weising, K. An integrated molecular and morphological study of the subfamily Suaedoideae Ulbr. (Chenopodiaceae). Plant Syst. Evol. 2003, 239, 257–286. [Google Scholar] [CrossRef]
- Zhao, K.F.; Li, F.Z.; Fan, S.J.; Feng, L.T. Salt plants of China. Chin. Bull. Botany 1993, 3, 10–16, (In Chinese with English Abstract). [Google Scholar]
- Prinz, K.; Hensen, I.; Schie, S.; Debener, T.; Weising, K. Microsatellite markers for the tetraploid halophyte Suaeda maritima (L.) Dumort. (Chenopodiaceae) and cross-species amplification in related taxa. Mol. Ecol. Resour. 2009, 9, 1247–1249. [Google Scholar] [CrossRef]
- Park, J.; Okita, T.W.; Edwards, G.E. Salt tolerant mechanisms in single-cell C4 species Bienertia sinuspersici and Suaeda aralocaspica (Chenopodiaceae). Plant Sci. 2009, 176, 616–626. [Google Scholar] [CrossRef]
- Lara, M.V.; Offermann, S.; Smith, M.; Okita, T.W.; Andreo, C.S.; Edwards, G.E. Leaf development in the single-cell C4 system in Bienertia sinuspersici: Expression of genes and peptide levels for C4 metabolism in relation to chlorenchyma structure under different light conditions. Plant Physiol. 2008, 148, 593–610. [Google Scholar] [CrossRef]
- Edwards, G.E.; Voznesenskaya, E.V. Chapter 4 C4 photosynthesis: Kranz forms and single-cell C4 in terrestrial plants. C4 Photosynth. Relat. CO2 Conc. Mech. 2011, 32, 29–61. [Google Scholar]
- Sharpe, R.M.; Offermann, S. One decade after the discovery of single-cell C4 species in terrestrial plants: What did we learn about the minimal requirements of C4 photosynthesis? Photosynth Res. 2014, 119, 169–180. [Google Scholar] [CrossRef] [PubMed]
- Hughes, A.R.; Inouye, B.D.; Johnson, M.T.J.; Underwood, N.; Vellend, M. Ecological consequences of genetic diversity. Ecol. Lett. 2008, 11, 609–623. [Google Scholar] [CrossRef] [PubMed]
- Booy, G.; Hendriks, R.J.J.; Smulders, M.J.M.; Groenendael, J.M.V.; Vosman, B. Genetic diversity and the survival of populations. Plant Biol. 2000, 2, 379–395. [Google Scholar] [CrossRef]
- Frankham, R. Genetics and conservation biology. Comptes Rendus Biol. 2003, 326, s22–s29. [Google Scholar] [CrossRef]
- Allendorf, F.W.; Ryman, N. The role of genetics in population viability analysis. Popul. Viability Anal. 2002, 50, 85. [Google Scholar]
- Cheng, T.F.; Wang, H.; Zhou, D.W.; Chen, S.L.; Wang, J.L.; Shi, S.B.; Shen, J.W.; Lei, T.X. Progress in the study of genetic diversity of Gentiana macrophylla Radix. Chin. Tradit. Herb. Drugs 2019, 15, 3720–3728, (In Chinese with English Abstract). [Google Scholar]
- Mcintosh, R.A. Catalogue of Gene Symbols for Wheat; University Extension Press, University of Saskatchewan: Saskatoon, SK, Canada, 1998; Volume 5, pp. 119–120. [Google Scholar]
- Grover, A.; Sharma, P.C. Development and use of molecular markers: Past and present. Crit. Rev. Biotechnol. 2016, 36, 290–302. [Google Scholar] [CrossRef]
- Morgante, M.; Olivieri, A. PCR-amplified microsatellites as markers in plant genetics. Plant J. 1993, 3, 175–182. [Google Scholar] [CrossRef]
- Vieira, M.L.C.; Santini, L.; Diniz, A.L.; Munhozl, C.F. Microsatellite markers: What they mean and why they are so use-ful. Genet. Mol. Biol. 2016, 39, 312–328. [Google Scholar] [CrossRef]
- Park, J.S.; Takayama, K.; Suyama, Y.; Choi, B.H. Distinct phylogeographic structure of the halophyte Suaeda malacosperma (Chenopodiaceae/Amaranthaceae), endemic to Korea–Japan region, influenced by historical range shift dynamics. Plant Syst. Evol. 2019, 305, 193–203. [Google Scholar] [CrossRef]
- Prinz, K.; Weising, K.; Hensen, I. Genetic structure of coastal and inland populations of the annual halophyte Suaeda maritima (L.) dumort. in Central Europe, inferred from amplified fragment length polymorphism markers. Plant Biol. 2009, 11, 812–820. [Google Scholar] [CrossRef] [PubMed]
- Lomonosova, M.N.; Nikonova, D.E.; Kutsev, M.G.; Dorogina, O.V.; Korolyuk, A.Y. Genetic differentiation in the polyploid complex of Suaeda corniculata (CA Mey.) Bunge in Eastern Siberia. Russ. J. Genet. 2017, 53, 596–605. [Google Scholar] [CrossRef]
- Prinz, K.; Weising, K.; Hensen, I. Habitat fragmentation and recent bottlenecks influence genetic diversity and differentiation of the central European halophyte Suaeda maritima (Chenopodiaceae). Am. J. Bot. 2013, 100, 2210–2218. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Ma, G.L.; Wang, H.L.; Cheng, C.; Mu, S.Y.; Quan, W.L.; Jiang, L.; Zhao, Z.Y.; Zhang, Y.; Tian, C.Y.; et al. A draft genome assembly of halophyte Suaeda aralocaspica, a plant that performs C4 photosynthesis within individual cells. GigaScience 2019, 8, giz116. [Google Scholar] [CrossRef]
- Martina, M.; Acquadro, A.; Barchi, L.; Gulino, D.; Brusco, F.; Rabaglio, M.; Portis, F.; Portis, E.; Lanteri, S. Genome-wide survey and development of the first microsatellite markers database (AnCorDB) in Anemone coronaria L. Int. J. Mol. Sci. 2022, 23, 3126. [Google Scholar] [CrossRef]
- Han, B.; Wang, C.B.; Tang, Z.H.; Ren, Y.K.; Li, Y.L.; Zhang, D.Y.; Dong, Y.H.; Zhao, X.H. Genome-wide analysis of microsatellite markers based on sequenced database in Chinese spring wheat (Triticum aestivum L.). PLoS ONE 2015, 10, e0141540. [Google Scholar] [CrossRef]
- Portis, E.; Lanteri, S.; Barchi, L.; Portis, F.; Valente, L.; Toppino, L.; Rotino, G.L.; Acquadro, A. Comprehensive characterization of simple sequence repeats in eggplant (Solanum melongena L.) genome and construction of a web resource. Front. Plant Sci. 2018, 9, 401. [Google Scholar] [CrossRef]
- Hou, S.; Ren, X.M.; Yang, Y.; Wang, D.H.; Du, W.; Wang, X.F.; Li, H.Y.; Han, Y.H.; Liu, L.L.; Sun, Z.X. Genome-wide development of polymorphic microsatellite markers and association analysis of major agronomic traits in core germplasm resources of tartary buckwheat. Front. Plant Sci. 2022, 13, 357. [Google Scholar] [CrossRef]
- Shiga, T.; Yokogawa, M.; Kaneko, S.; Isagi, Y. Genetic diversity and population structure of Nuphar submersa (Nymphaeaceae), a critically endangered aquatic plant endemic to Japan, and implications for its conservation. J. Plant Res. 2017, 130, 83–93. [Google Scholar] [CrossRef]
- Li, W.J.; Su, Z.H.; Li, A.R.; Guan, K.Y.; Feng, Y. Isolation and characterization of 18 microsatellites for the invasive native Pedicularis kansuensis (Orobanchaceae). Grassl. Sci. 2019, 65, 135–138. [Google Scholar] [CrossRef]
- Meloni, M.; Reid, A.; Caujapé-Castells, J.; Moisés, S.; Fernández-Palacios, J.M.; Conti, E. High genetic diversity and population structure in the endangered Canarian endemic Ruta oreojasme (Rutaceae). Genetica 2015, 143, 571–580. [Google Scholar] [CrossRef]
- Yamashiro, T.; Yamashiro, A.; Inoue, M.; Maki, M. Genetic diversity and divergence in populations of the threatened grassland perennial Vincetoxicum atratum (Apocynaceae-Asclepiadoideae) in Japan. J. Hered. 2016, 107, 455–462. [Google Scholar] [CrossRef] [PubMed]
- Vieira, Â.F.; Dias, E.F.; Moura, M. Geography, geology and ecology influence population genetic diversity and structure in the endangered endemic Azorean Ammi (Apiaceae). Plant Syst. Evol. 2018, 304, 163–176. [Google Scholar] [CrossRef]
- Zhou, X.J.; Ren, X.L.; Liu, W.Z. Genetic diversity of SSR markers in wild populations of Tapiscia sinensis, an endangered tree species. Biochem. Syst. Ecol. 2016, 69, 1–5. [Google Scholar] [CrossRef]
- Endler, J.A. Geographic Variation, Speciation, and Clines; Princeton University Press, Princeton University: Princeton, NJ, USA, 1977; pp. 97–139. [Google Scholar]
- Freeland, J.R. Molecular Ecology; John Wiley & Sons: New York, NY, USA, 2020; pp. 457–458. [Google Scholar]
- Zhang, D.Y.; Jiang, X.H. Evolution of plant mating systems, resource allocation responses and genetic diversity. Acta Phytoecol. Sin. 2001, 2, 130–143, (In Chinese with English abstract). [Google Scholar]
- Nybom, H. Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants. Mol. Ecol. 2004, 13, 1143–1155. [Google Scholar] [CrossRef]
- Reed, D.H.; Briscoe, D.A.; Frankham, R. Inbreeding and extinction: The effect of environmental stress and lineage. Conserv. Genet. 2002, 3, 301–307. [Google Scholar] [CrossRef]
- Vít, P.; Krak, K.; Douda, J.; Mandák, B. Microsatellite markers for Anthericum ramosum: Development, characterization, and cross-species amplification. Appl. Plant Sci. 2020, 8, e11323. [Google Scholar] [CrossRef]
- Krak, K.; Vít, P.; Douda, J.; Mandák, B. Development of 18 microsatellite markers for Salvia pratensis. Appl. Plant Sci. 2020, 8, e11316. [Google Scholar] [CrossRef]
- Su, X.N.; Wang, H.M.; Huang, X.X.; Xie, X.Y.; Zhang, X.T.; Niu, L.; Dong, L.; Jiang, S.L.; Wu, J.L.; Jiang, S.M. Analysis of genetic diversity and genetic structure of wild mizuna in Northeast China using SSR markers. Acta Agric. Univ. Jiangxiensis 2018, 40, 579–589, (In Chinese with English Abstract). [Google Scholar]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research—An update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Fan, W.Q.; Gai, H.M.; Sun, X.; Yang, A.G.; Zhang, Z.F.; Ren, M. SSR data format conversion software DataFormater. Mol. Plant Breed. 2016, 14, 265–270, (In Chinese with English Abstract). [Google Scholar]
| Items | Numbers |
|---|---|
| Total size of genome (Mb) | 452 |
| Total number of identified SSRs | 177,805 |
| Total length of SSRs (bp) | 3,465,418 |
| Frequency (SSRs/Mb) | 393.37 |
| Density (bp/Mb) | 7666.85 |
| Total content of genome SSRs(%) | 0.77 |
| Repeat Type | Predominant Type | Number | Proportion (%) | Frequency (SSRs/Mb) | Total Length (bp) | Average Length (bp) |
|---|---|---|---|---|---|---|
| Di | AT | 25,256 | 14.20 | 55.88 | 473,878 | 18.76 |
| Tri | AAC/GTT | 134,663 | 75.74 | 297.93 | 2,383,446 | 17.70 |
| Tetra | AAAT/ATTT | 11,150 | 6.27 | 24.67 | 337,516 | 30.27 |
| Penta | AAAAT/ATTTT | 3771 | 2.12 | 8.34 | 113,540 | 30.11 |
| Hexa | AACAAT/ATTGTT | 2965 | 1.67 | 6.56 | 157,038 | 52.96 |
| Total | 177,805 | 100 | 393.37 | 3,465,418 | 19.49 |
| The Motif of Repeat | Repeat Numbers | Total | Percentage (%) | |||||
|---|---|---|---|---|---|---|---|---|
| 4 | 5 | 6 | 7 | 8 | >8 | |||
| AT/AT | 0 | 0 | 4595 | 3065 | 2511 | 7751 | 17,922 | 10.08 |
| AG/CT | 0 | 0 | 1594 | 913 | 549 | 1359 | 4415 | 2.48 |
| AC/GT | 0 | 0 | 1110 | 625 | 401 | 748 | 2884 | 1.62 |
| AAC/GTT | 34,600 | 18,327 | 10,383 | 5476 | 3172 | 6728 | 78,686 | 44.25 |
| AAT/ATT | 10,890 | 3893 | 2108 | 1243 | 904 | 5491 | 24,529 | 13.80 |
| ACC/GGT | 5994 | 2217 | 966 | 438 | 248 | 155 | 10,018 | 5.63 |
| AAG/CTT | 6119 | 1338 | 439 | 165 | 70 | 317 | 8448 | 4.75 |
| ATC/ATG | 4297 | 1245 | 481 | 219 | 107 | 207 | 6556 | 3.69 |
| ACT/AGT | 1229 | 481 | 273 | 130 | 73 | 237 | 2423 | 1.36 |
| AGC/CTG | 1429 | 277 | 76 | 30 | 9 | 20 | 1841 | 1.04 |
| AGG/CCT | 1068 | 262 | 95 | 47 | 31 | 47 | 1550 | 0.87 |
| AAAT/ATTT | 2861 | 1037 | 457 | 220 | 111 | 375 | 5061 | 2.85 |
| AATC/ATTG | 962 | 142 | 34 | 8 | 4 | 4 | 1154 | 0.65 |
| AATT/AATT | 562 | 151 | 47 | 28 | 2 | 4 | 794 | 0.45 |
| AAAAT/ATTTT | 709 | 182 | 38 | 10 | 7 | 6 | 952 | 0.54 |
| Total | 70,720 | 29,552 | 22,696 | 12,617 | 8199 | 23,449 | 167,233 | 94.05 |
| Percentage (%) | 39.77 | 16.62 | 12.76 | 7.10 | 4.61 | 13.19 | 94.05 | |
| Locus | N | Na | Ne | I | Ho | He | uHe | Fst | PIC |
|---|---|---|---|---|---|---|---|---|---|
| SA-di-11 | 52 | 4 | 2.116 | 1.004 | 0.000 | 0.527 | 0.532 | 0.243 | 0.488 |
| SA-di-13 | 52 | 4 | 2.392 | 1.007 | 0.808 | 0.582 | 0.588 | 0.100 | 0.517 |
| SA-tri-24 | 52 | 7 | 4.989 | 1.755 | 0.038 | 0.800 | 0.807 | 0.390 | 0.773 |
| SA-tri-26 | 52 | 6 | 3.462 | 1.354 | 0.385 | 0.711 | 0.718 | 0.287 | 0.658 |
| SA-te-29 | 52 | 3 | 1.702 | 0.730 | 0.173 | 0.413 | 0.417 | 0.261 | 0.369 |
| SA-di-34 | 50 | 4 | 2.417 | 1.057 | 0.060 | 0.586 | 0.592 | 0.158 | 0.526 |
| SA-di-35 | 52 | 6 | 2.973 | 1.294 | 0.019 | 0.664 | 0.670 | 0.548 | 0.609 |
| SA-di-39 | 52 | 5 | 2.818 | 1.276 | 0.038 | 0.645 | 0.651 | 0.278 | 0.605 |
| SA-di-40 | 52 | 3 | 2.000 | 0.791 | 0.038 | 0.500 | 0.505 | 0.198 | 0.408 |
| SA-tri-41 | 52 | 6 | 3.485 | 1.400 | 0.231 | 0.713 | 0.720 | 0.096 | 0.664 |
| SA-tri-42 | 52 | 4 | 2.094 | 0.942 | 0.019 | 0.522 | 0.527 | 0.342 | 0.469 |
| SA-tri-43 | 51 | 4 | 1.676 | 0.689 | 0.020 | 0.403 | 0.407 | 0.152 | 0.343 |
| SA-tri-46 | 52 | 8 | 3.634 | 1.553 | 0.212 | 0.725 | 0.732 | 0.314 | 0.69 |
| SA-te-50 | 52 | 8 | 2.917 | 1.331 | 0.212 | 0.657 | 0.664 | 0.258 | 0.597 |
| SA-di-53 | 52 | 10 | 4.650 | 1.809 | 0.154 | 0.785 | 0.793 | 0.415 | 0.757 |
| SA-di-54 | 52 | 5 | 2.528 | 1.093 | 0.038 | 0.604 | 0.610 | 0.407 | 0.528 |
| SA-di-59 | 52 | 5 | 2.367 | 1.063 | 0.077 | 0.577 | 0.583 | 0.528 | 0.516 |
| SA-di-61 | 51 | 9 | 3.315 | 1.433 | 0.961 | 0.698 | 0.705 | 0.127 | 0.648 |
| SA-di-62 | 50 | 7 | 2.244 | 1.177 | 0.080 | 0.554 | 0.560 | 0.456 | 0.524 |
| SA-di-63 | 51 | 12 | 3.605 | 1.655 | 0.412 | 0.723 | 0.730 | 0.209 | 0.69 |
| SA-di-64 | 52 | 6 | 5.017 | 1.685 | 0.692 | 0.801 | 0.808 | 0.129 | 0.771 |
| SA-di-65 | 52 | 4 | 1.405 | 0.596 | 0.135 | 0.288 | 0.291 | 0.160 | 0.272 |
| SA-di-67 | 52 | 8 | 4.404 | 1.674 | 0.000 | 0.773 | 0.780 | 0.469 | 0.74 |
| SA-di-69 | 52 | 8 | 4.265 | 1.697 | 1.000 | 0.766 | 0.773 | 0.027 | 0.734 |
| SA-di-73 | 52 | 6 | 2.041 | 1.109 | 0.058 | 0.510 | 0.515 | 0.258 | 0.487 |
| SA-di-74 | 52 | 6 | 1.992 | 1.014 | 0.038 | 0.498 | 0.503 | 0.338 | 0.464 |
| SA-tri-77 | 52 | 6 | 3.453 | 1.373 | 1.000 | 0.710 | 0.717 | 0.096 | 0.664 |
| SA-tri-78 | 51 | 7 | 2.876 | 1.419 | 0.078 | 0.652 | 0.659 | 0.350 | 0.623 |
| SA-tri-83 | 52 | 7 | 4.765 | 1.649 | 0.019 | 0.790 | 0.798 | 0.265 | 0.757 |
| SA-tri-85 | 52 | 10 | 5.131 | 1.827 | 0.135 | 0.805 | 0.813 | 0.398 | 0.778 |
| SA-tri-86 | 52 | 9 | 4.122 | 1.671 | 0.173 | 0.757 | 0.765 | 0.297 | 0.727 |
| SA-tri-88 | 52 | 4 | 2.687 | 1.122 | 0.981 | 0.628 | 0.634 | 0.063 | 0.559 |
| SA-te-92 | 52 | 6 | 2.098 | 0.964 | 0.654 | 0.523 | 0.528 | 0.162 | 0.456 |
| SA-te-93 | 52 | 5 | 2.155 | 0.904 | 0.173 | 0.536 | 0.541 | 0.350 | 0.444 |
| SA-te-94 | 52 | 12 | 3.532 | 1.702 | 0.077 | 0.717 | 0.724 | 0.306 | 0.691 |
| SA-te-96 | 52 | 4 | 1.528 | 0.691 | 0.019 | 0.346 | 0.349 | 0.105 | 0.323 |
| SA-te-97 | 52 | 4 | 1.648 | 0.756 | 0.058 | 0.393 | 0.397 | 0.337 | 0.359 |
| SA-te-98 | 51 | 3 | 1.922 | 0.711 | 0.020 | 0.480 | 0.484 | 0.128 | 0.374 |
| Pop | N | Na | Ne | I | Ho | He | uHe | F | |
|---|---|---|---|---|---|---|---|---|---|
| Fukang | Mean | 16.895 | 4.447 | 2.667 | 1.085 | 0.299 | 0.573 | 0.591 | 0.515 |
| SE | 0.063 | 0.269 | 0.155 | 0.062 | 0.053 | 0.027 | 0.028 | 0.078 | |
| Shawan | Mean | 16.974 | 2.921 | 1.794 | 0.642 | 0.207 | 0.365 | 0.376 | 0.602 |
| SE | 0.026 | 0.265 | 0.113 | 0.066 | 0.061 | 0.035 | 0.036 | 0.109 | |
| Shihezi | Mean | 17.895 | 2.947 | 1.920 | 0.712 | 0.228 | 0.416 | 0.428 | 0.571 |
| SE | 0.063 | 0.196 | 0.114 | 0.056 | 0.058 | 0.031 | 0.032 | 0.098 |
| Fukang | Shawan | Shihezi | |
|---|---|---|---|
| Fukang | 1 | ||
| Shawan | 0.438 | 1 | |
| Shihezi | 0.548 | 0.654 | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, W.; Wang, J.; Tian, C.; Shi, W.; Wang, L. Genome-Wide Development of Polymorphic Microsatellite Markers and Genetic Diversity Analysis for the Halophyte Suaeda aralocaspica (Amaranthaceae). Plants 2023, 12, 1865. https://doi.org/10.3390/plants12091865
Xu W, Wang J, Tian C, Shi W, Wang L. Genome-Wide Development of Polymorphic Microsatellite Markers and Genetic Diversity Analysis for the Halophyte Suaeda aralocaspica (Amaranthaceae). Plants. 2023; 12(9):1865. https://doi.org/10.3390/plants12091865
Chicago/Turabian StyleXu, Wei, Jiancheng Wang, Changyan Tian, Wei Shi, and Lei Wang. 2023. "Genome-Wide Development of Polymorphic Microsatellite Markers and Genetic Diversity Analysis for the Halophyte Suaeda aralocaspica (Amaranthaceae)" Plants 12, no. 9: 1865. https://doi.org/10.3390/plants12091865
APA StyleXu, W., Wang, J., Tian, C., Shi, W., & Wang, L. (2023). Genome-Wide Development of Polymorphic Microsatellite Markers and Genetic Diversity Analysis for the Halophyte Suaeda aralocaspica (Amaranthaceae). Plants, 12(9), 1865. https://doi.org/10.3390/plants12091865








