Genome-Wide Analysis of the bHLH Gene Family in Loropetalum chinense var. rubrum: Identification, Classification, Evolution, and Diversity of Expression Patterns under Cultivation
Abstract
1. Introduction
2. Results
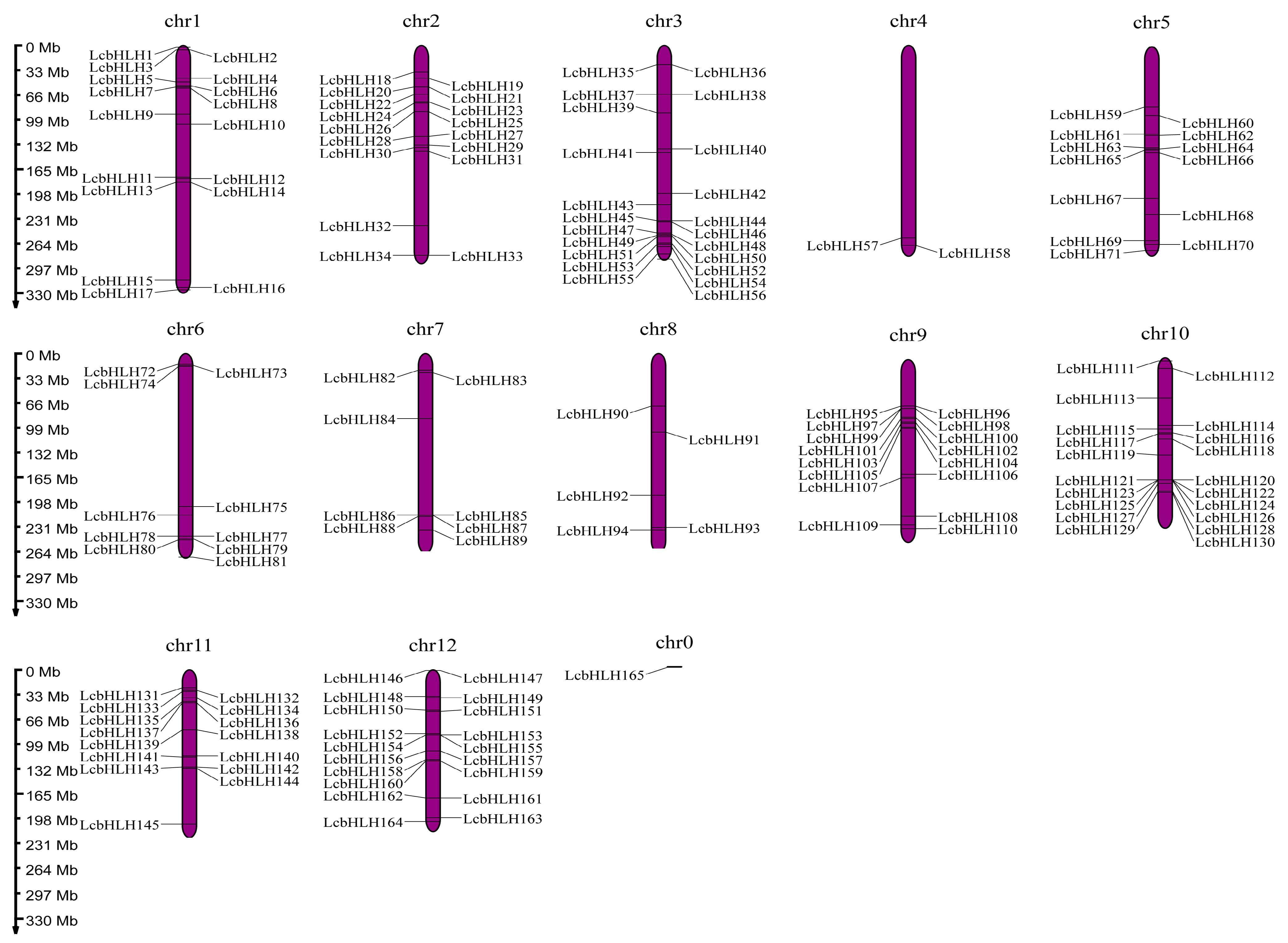
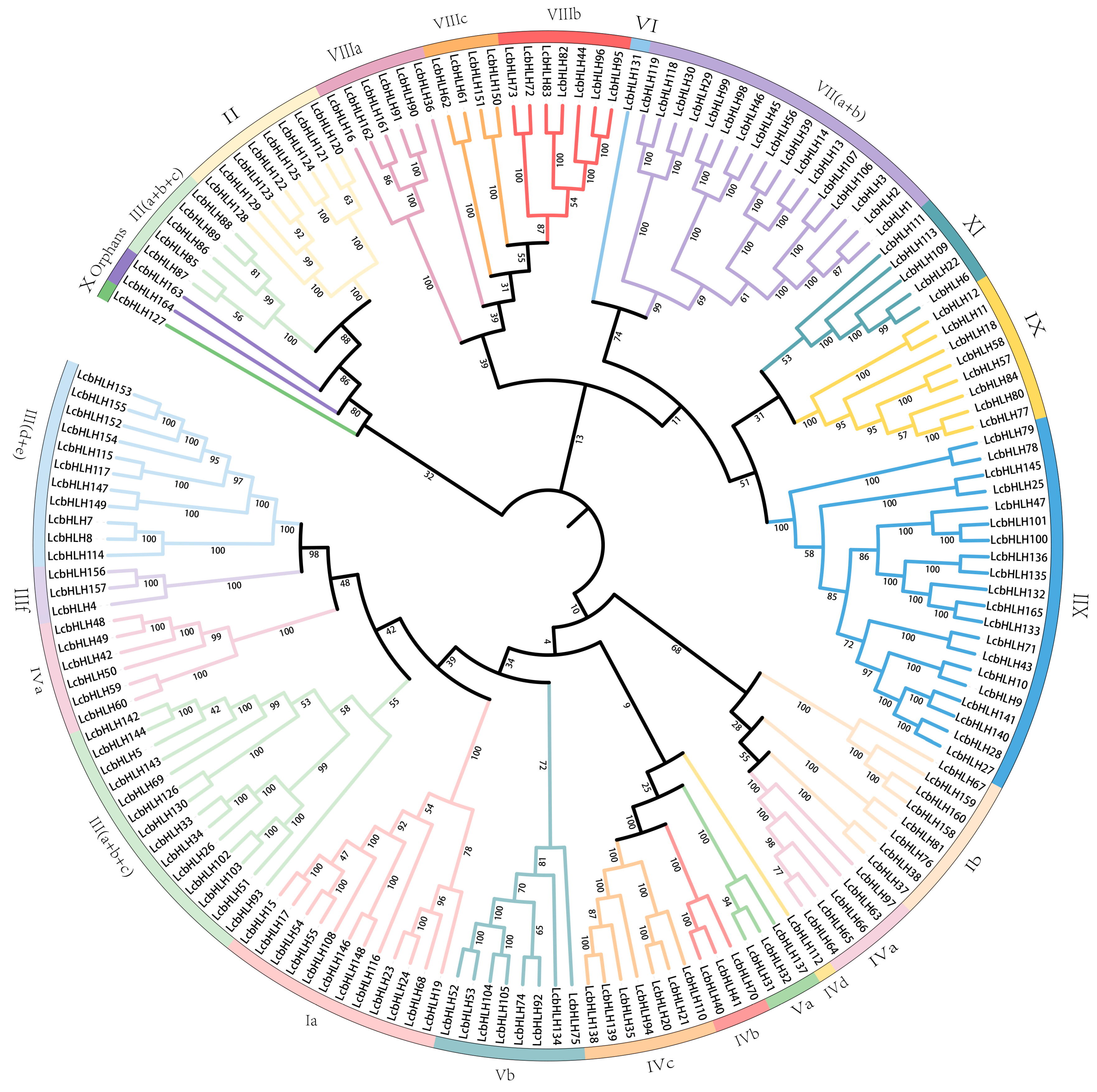
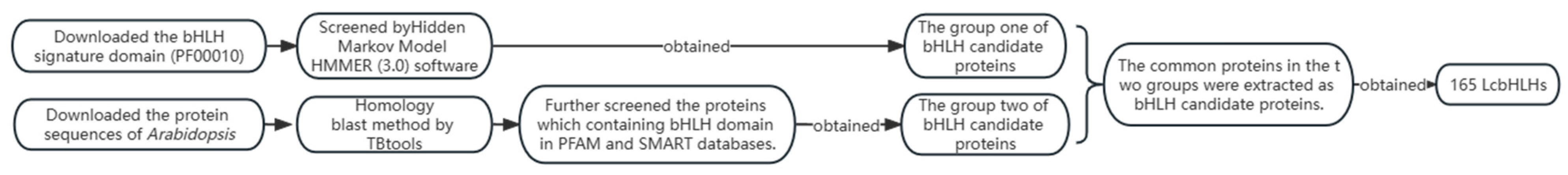
2.1. Identification of the LcbHLHs and Evolutionary Analysis
2.2. Multiple Sequence Alignment and Analyses of the Motifs, Domains, and Structure
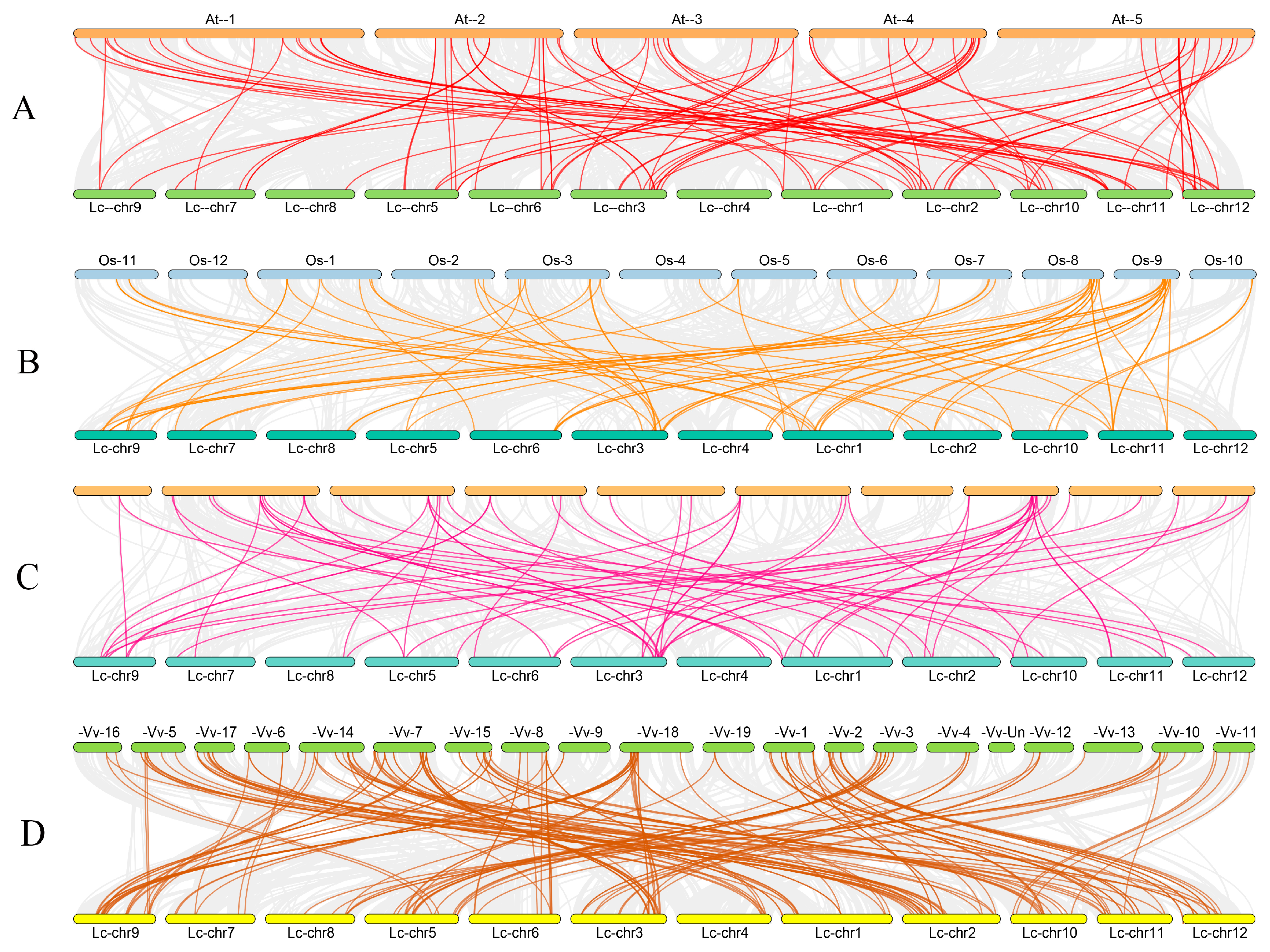
2.3. Analysis of Gene Duplication and Collinear Correlations
2.4. GO Annotation and Analyses of the Cis-Regulatory Elements
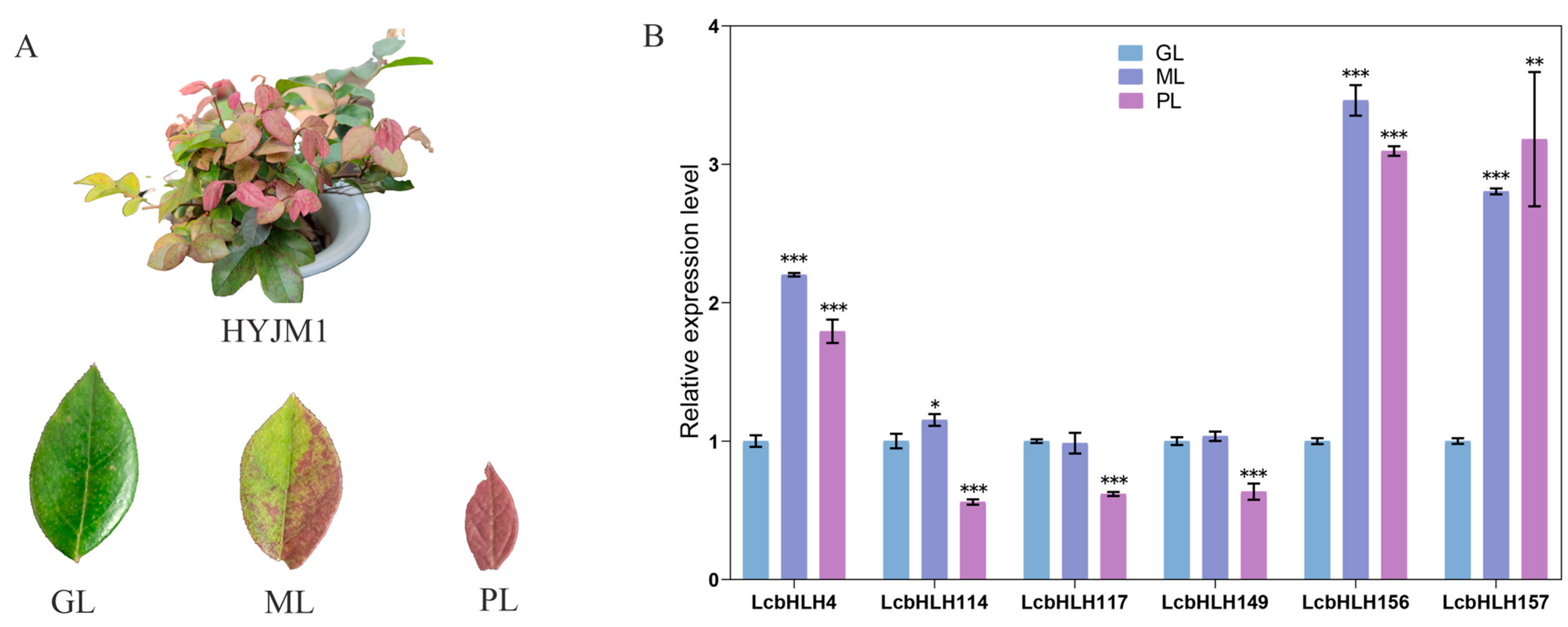
2.5. Expression Profiles of LcbHLHs and qRT–PCR Analysis of Gene Expression
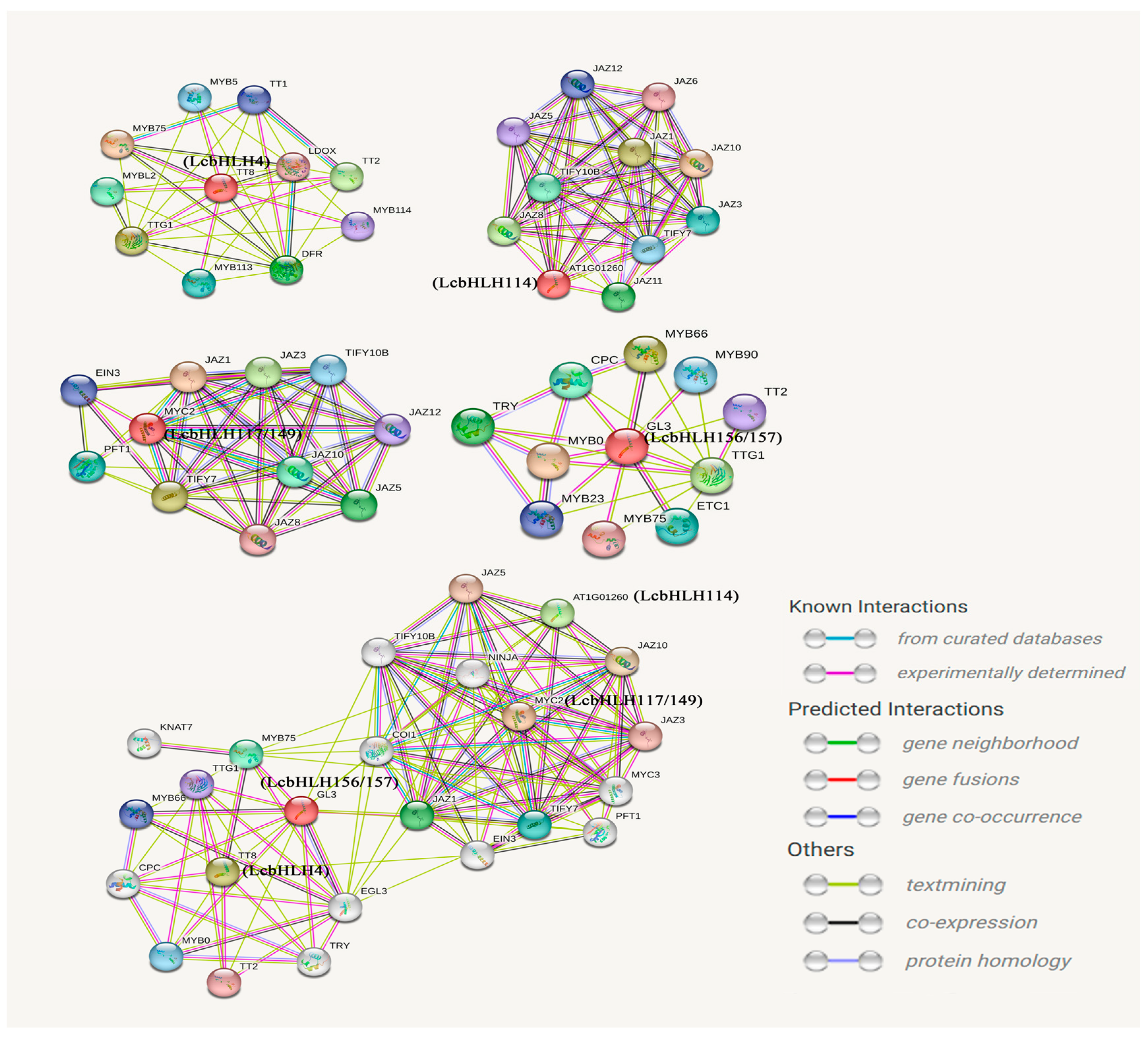
2.6. The Protein–Protein Interaction Network of the Candidate Genes
2.7. Overexpression of LcbHLH156
3. Discussion
3.1. Systematic and Comprehensive Genome-Wide Detection of LcbHLHs in L. chinense var. rubrum
3.2. Functional Prediction of LcbHLHs
4. Materials and Methods
4.1. Plant Materials and Data Resources
4.2. Identification and Physicochemical Characterization of the bHLH Gene Family
4.3. Chromosomal Localization, Tandem Duplication, and Collinearity of the bHLH Genes
4.4. Evolution and Multiple Sequence Alignment of LcbHLHs
4.5. Analysis of the Gene Structure, Conserved Motifs, and Family Structural Domains
4.6. GO Annotation, Analysis of the Cis-Acting Components, and the Protein–Protein Interactions of LcbHLHs
4.7. Gene Expression Patterns and Quantitative Real-Time PCR
4.8. Transient Expression of L. chinense var. rubrum and Color Measurement
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Riechmann, J.L.; Ratcliffe, O.J. A Genomic Perspective on Plant Transcription Factors. Curr. Opin. Plant Biol. 2000, 3, 423–434. [Google Scholar] [CrossRef] [PubMed]
- Phillips, S.E. Built by Association: Structure and Function of Helix-Loop-Helix DNA-Binding Proteins. Structure 1994, 2, 1–4. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Atchley, W.R.; Fitch, W.M. A Natural Classification of the Basic Helix–Loop–Helix Class of Transcription Factors. Proc. Natl. Acad. Sci. USA 1997, 94, 5172–5176. [Google Scholar] [CrossRef] [PubMed]
- Massari, M.E.; Murre, C. Helix-Loop-Helix Proteins: Regulators of Transcription in Eucaryotic Organisms. Mol. Cell. Biol. 2000, 20, 429–440. [Google Scholar] [CrossRef] [PubMed]
- Heim, M.A. The Basic Helix-Loop-Helix Transcription Factor Family in Plants: A Genome-Wide Study of Protein Structure and Functional Diversity. Mol. Biol. Evol. 2003, 20, 735–747. [Google Scholar] [CrossRef] [PubMed]
- Ledent, V.; Vervoort, M. The Basic Helix-Loop-Helix Protein Family: Comparative Genomics and Phylogenetic Analysis. Genome Res. 2001, 11, 754–770. [Google Scholar] [CrossRef] [PubMed]
- Carretero-Paulet, L.; Galstyan, A.; Roig-Villanova, I.; Martínez-García, J.F.; Bilbao-Castro, J.R.; Robertson, D.L. Genome-Wide Classification and Evolutionary Analysis of the bHLH Family of Transcription Factors in Arabidopsis, Poplar, Rice, Moss, and Algae. Plant Physiol. 2010, 153, 1398–1412. [Google Scholar] [CrossRef] [PubMed]
- Bailey, P.C.; Martin, C.; Toledo-Ortiz, G.; Quail, P.H.; Huq, E.; Heim, M.A.; Jakoby, M.; Werber, M.; Weisshaar, B. Update on the Basic Helix-Loop-Helix Transcription Factor Gene Family in Arabidopsis Thaliana. Plant Cell 2003, 15, 2497–2502. [Google Scholar] [CrossRef]
- Toledo-Ortiz, G.; Huq, E.; Quail, P.H. The Arabidopsis Basic/Helix-Loop-Helix Transcription Factor Family. Plant Cell 2003, 15, 1749–1770. [Google Scholar] [CrossRef]
- Li, X.; Duan, X.; Jiang, H.; Sun, Y.; Tang, Y.; Yuan, Z.; Guo, J.; Liang, W.; Chen, L.; Yin, J.; et al. Genome-Wide Analysis of Basic/Helix-Loop-Helix Transcription Factor Family in Rice and Arabidopsis. Plant Physiol. 2006, 141, 1167–1184. [Google Scholar] [CrossRef]
- Hao, Y.; Zong, X.; Ren, P.; Qian, Y.; Fu, A. Basic Helix-Loop-Helix (bHLH) Transcription Factors Regulate a Wide Range of Functions in Arabidopsis. Int. J. Mol. Sci. 2021, 22, 7152. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Chen, C.-L.; Cui, M.; Zhou, W.-J.; Wu, H.-L.; Ling, H.-Q. Four IVa bHLH Transcription Factors Are Novel Interactors of FIT and Mediate JA Inhibition of Iron Uptake in Arabidopsis. Mol. Plant 2018, 11, 1166–1183. [Google Scholar] [CrossRef] [PubMed]
- Lu, R.; Zhang, J.; Wu, Y.-W.; Wang, Y.; Zhang, J.; Zheng, Y.; Li, Y.; Li, X.-B. bHLH Transcription Factors LP1 and LP2 Regulate Longitudinal Cell Elongation. Plant Physiol. 2021, 187, 2577–2591. [Google Scholar] [CrossRef] [PubMed]
- Sharma, N.; Xin, R.; Kim, D.-H.; Sung, S.; Lange, T.; Huq, E. No flowering in short day (NFL) is a bHLH transcription factor that promotes flowering specifically under short-day conditions in Arabidopsis. Development 2016, 143, 682–690. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Yang, W.; Jia, Q.; Wang, W.; Zhang, N.; Wang, X.; Wang, S. Pleurotus ostreatus bHLH Transcription Factors Regulate Plant Growth and Development When Expressed in Arabidopsis. Null 2017, 12, 542–549. [Google Scholar] [CrossRef]
- Sun, X.; Wang, Y.; Sui, N. Transcriptional Regulation of bHLH during Plant Response to Stress. Biochem. Biophys. Res. Commun. 2018, 503, 397–401. [Google Scholar] [CrossRef] [PubMed]
- Zhai, Y.; Zhang, L.; Xia, C.; Fu, S.; Zhao, G.; Jia, J.; Kong, X. The Wheat Transcription Factor, TabHLH39, Improves Tolerance to Multiple Abiotic Stressors in Transgenic Plants. Biochem. Biophys. Res. Commun. 2016, 473, 1321–1327. [Google Scholar] [CrossRef] [PubMed]
- Ren, Y.-R.; Yang, Y.-Y.; Zhao, Q.; Zhang, T.-E.; Wang, C.-K.; Hao, Y.-J.; You, C.-X. MdCIB1, an Apple bHLH Transcription Factor, Plays a Positive Regulator in Response to Drought Stress. Environ. Exp. Bot. 2021, 188, 104523. [Google Scholar] [CrossRef]
- Zhao, R.; Song, X.; Yang, N.; Chen, L.; Xiang, L.; Liu, X.-Q.; Zhao, K. Expression of the Subgroup IIIf bHLH Transcription Factor CpbHLH1 from Chimonanthus praecox (L.) in Transgenic Model Plants Inhibits Anthocyanin Accumulation. Plant Cell Rep. 2020, 39, 891–907. [Google Scholar] [CrossRef]
- Deng, J.; Li, J.; Su, M.; Lin, Z.; Chen, L.; Yang, P. A bHLH Gene NnTT8 of Nelumbo Nucifera Regulates Anthocyanin Biosynthesis. Plant Physiol. Biochem. 2021, 158, 518–523. [Google Scholar] [CrossRef]
- Arlotta, C.; Puglia, G.D.; Genovese, C.; Toscano, V.; Karlova, R.; Beekwilder, J.; De Vos, R.C.H.; Raccuia, S.A. MYB5-like and bHLH Influence Flavonoid Composition in Pomegranate. Plant Sci. 2020, 298, 110563. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Hou, H.; Jiang, X.; Wang, P.; Dai, X.; Chen, W.; Gao, L.; Xia, T. A WD40 Repeat Protein from Camellia sinensis Regulates Anthocyanin and Proanthocyanidin Accumulation through the Formation of MYB–bHLH–WD40 Ternary Complexes. Int. J. Mol. Sci. 2018, 19, 1686. [Google Scholar] [CrossRef] [PubMed]
- Shan, X.; Li, Y.; Yang, S.; Gao, R.; Zhou, L.; Bao, T.; Han, T.; Wang, S.; Gao, X.; Wang, L. A Functional Homologue of Arabidopsis TTG1 from Freesia Interacts with bHLH Proteins to Regulate Anthocyanin and Proanthocyanidin Biosynthesis in Both Freesia hybrida and Arabidopsis thaliana. Plant Physiol. Biochem. 2019, 141, 60–72. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Li, J.; Zhu, L.; Chang, P.; Li, L.; Zhang, L. Identification and Characterization of MYB-bHLH-WD40 Regulatory Complex Members Controlling Anthocyanidin Biosynthesis in Blueberry Fruits Development. Genes 2019, 10, 496. [Google Scholar] [CrossRef] [PubMed]
- Gawel, N.J.; Johnson, G.R.; Sauve, R. Identification of Genetic Diversity among Loropetalum chinense var. rubrum Introductions. J. Environ. Hortic. 1996, 14, 38–41. [Google Scholar] [CrossRef]
- Gao, Y.-F.; Zhao, D.-H.; Zhang, J.-Q.; Chen, J.-S.; Li, J.-L.; Weng, Z.; Rong, L.-P. De Novo Transcriptome Sequencing and Anthocyanin Metabolite Analysis Reveals Leaf Color of Acer Pseudosieboldianum in Autumn. BMC Genom. 2021, 22, 383. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Li, Y.; Zhao, M.; Hu, Y.; Meng, F.; Song, X.; Tigabu, M.; Chiang, V.L.; Sederoff, R.; Ma, W.; et al. Molecular and Metabolic Insights into Anthocyanin Biosynthesis for Leaf Color Change in Chokecherry (Padus virginiana). Int. J. Mol. Sci. 2021, 22, 10697. [Google Scholar] [CrossRef] [PubMed]
- Filyushin, M.A.; Shchennikova, A.V.; Kochieva, E.Z. Coexpression of Structural and Regulatory Genes of the Flavonoid Pathway Reveals the Characteristics of Anthocyanin Biosynthesis in Eggplant Organs (Solanum melongena L.). Russ J Plant Physiol 2023, 70, 27. [Google Scholar] [CrossRef]
- Wang, X.; Yang, D.; Xian, R.; Wang, G.; Ma, G.; Zhou, L. Effects of Drought Stress on Pigment and Photosynthetic Characteristics in Leaves of Loropetalum chinense var. rubrum. J. Sichuan For. Sci. Technol. 2018, 39, 82–86. [Google Scholar] [CrossRef]
- Fei, F.; Wang, H.; Tang, Q. Study on the Impact of Temperature on Loropetalum chinense var. rubrum Leaves Color. J. Hunan Inst. Sci. Technol. (Nat. Sci.) 2008, 21, 88–90. [Google Scholar]
- Zhang, X. Cloning, Expression and Transformation of Lc CHI and Lc ANS Genes from Loropetalum chinense var. rubrum. Master Thesis, Hunan University of Technology, Zhuzhou, China, 2020. [Google Scholar]
- Sun, X.; Zhang, Z.; Li, J.; Zhang, H.; Peng, Y.; Li, Z. Uncovering Hierarchical Regulation among MYB-bHLH-WD40 Proteins and Manipulating Anthocyanin Pigmentation in Rice. Int. J. Mol. Sci. 2022, 23, 8203. [Google Scholar] [CrossRef]
- Wang, R.; Zhao, P.; Kong, N.; Lu, R.; Pei, Y.; Huang, C.; Ma, H.; Chen, Q. Genome-Wide Identification and Characterization of the Potato bHLH Transcription Factor Family. Genes 2018, 9, 54. [Google Scholar] [CrossRef]
- Cheng, X.; Xiong, R.; Liu, H.; Wu, M.; Chen, F.; Hanwei, Y.; Xiang, Y. Basic Helix-Loop-Helix Gene Family: Genome Wide Identification, Phylogeny, and Expression in Moso Bamboo. Plant Physiol. Biochem. 2018, 132, 104–119. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Su, L.; Gao, H.; Jiang, X.; Wu, X.; Li, Y.; Zhang, Q.; Wang, Y.; Ren, F. Genome-Wide Characterization of bHLH Genes in Grape and Analysis of Their Potential Relevance to Abiotic Stress Tolerance and Secondary Metabolite Biosynthesis. Front. Plant Sci. 2018, 9, 64. [Google Scholar] [CrossRef] [PubMed]
- Song, M.; Wang, H.; Wang, Z.; Huang, H.; Chen, S.; Ma, H. Genome-Wide Characterization and Analysis of bHLH Transcription Factors Related to Anthocyanin Biosynthesis in Fig (Ficus carica L.). Front. Plant Sci. 2021, 12, 730692. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.-J.; Wang, J.-R. Global Identification, Structural Analysis and Expression Characterization of bHLH Transcription Factors in Wheat. BMC Plant Biol. 2017, 17, 90. [Google Scholar] [CrossRef] [PubMed]
- Hurst, L.D. The Ka/Ks Ratio: Diagnosing the Form of Sequence Evolution. Trends Genet. 2002, 18, 486–487. [Google Scholar] [CrossRef] [PubMed]
- Yu, Q.; Hua, X.; Yao, H.; Zhang, Q.; He, J.; Peng, L.; Li, D.; Yang, Y.; Li, X. Abscisic Acid Receptors Are Involves in the Jasmonate Signaling in Arabidopsis. Plant Signal. Behav. 2021, 16, 1948243. [Google Scholar] [CrossRef]
- Fonseca, S.; Fernández-Calvo, P.; Fernández, G.M.; Díez-Díaz, M.; Gimenez-Ibanez, S.; López-Vidriero, I.; Godoy, M.; Fernández-Barbero, G.; Leene, J.V.; Jaeger, G.D.; et al. bHLH003, bHLH013 and bHLH017 Are New Targets of JAZ Repressors Negatively Regulating JA Responses. PLoS ONE 2014, 9, e86182. [Google Scholar] [CrossRef]
- Goossens, J.; Swinnen, G.; Vanden Bossche, R.; Pauwels, L.; Goossens, A. Change of a Conserved Amino Acid in the MYC 2 and MYC 3 Transcription Factors Leads to Release of JAZ Repression and Increased Activity. New Phytol. 2015, 206, 1229–1237. [Google Scholar] [CrossRef]
- Fernández-Calvo, P.; Chini, A.; Fernández-Barbero, G.; Chico, J.-M.; Gimenez-Ibanez, S.; Geerinck, J.; Eeckhout, D.; Schweizer, F.; Godoy, M.; Franco-Zorrilla, J.M.; et al. The Arabidopsis bHLH Transcription Factors MYC3 and MYC4 Are Targets of JAZ Repressors and Act Additively with MYC2 in the Activation of Jasmonate Responses. Plant Cell 2011, 23, 701–715. [Google Scholar] [CrossRef] [PubMed]
- Wen, J.; Li, Y.; Qi, T.; Gao, H.; Liu, B.; Zhang, M.; Huang, H.; Song, S. The C-Terminal Domains of Arabidopsis GL3/EGL3/TT8 Interact with JAZ Proteins and Mediate Dimeric Interactions. Plant Signal. Behav. 2018, 13, e1422460. [Google Scholar] [CrossRef] [PubMed]
- Song, S.; Liu, B.; Song, J.; Pang, S.; Song, T.; Gao, S.; Zhang, Y.; Huang, H.; Qi, T. A Molecular Framework for Signaling Crosstalk between Jasmonate and Ethylene in Anthocyanin Biosynthesis, Trichome Development, and Defenses against Insect Herbivores in Arabidopsis. J. Integr. Plant Biol. 2022, 64, 1770–1788. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Lv, W.; Zhang, H.; Ma, L.; Li, P.; Ge, L.; Li, G. Genome-Wide Analysis of the Basic Helix-Loop-Helix (bHLH) Transcription Factor Family in Maize. BMC Plant Biol. 2018, 18, 235. [Google Scholar] [CrossRef] [PubMed]
- Flagel, L.E.; Wendel, J.F. Gene Duplication and Evolutionary Novelty in Plants. New Phytol. 2009, 183, 557–564. [Google Scholar] [CrossRef] [PubMed]
- Wicker, T.; Buchmann, J.P.; Keller, B. Patching Gaps in Plant Genomes Results in Gene Movement and Erosion of Colinearity. Genome Res. 2010, 20, 1229–1237. [Google Scholar] [CrossRef] [PubMed]
- Qian, W.; Liao, B.-Y.; Chang, A.Y.-F.; Zhang, J. Maintenance of Duplicate Genes and Their Functional Redundancy by Reduced Expression. Trends Genet. 2010, 26, 425–430. [Google Scholar] [CrossRef] [PubMed]
- Oda, A.; Fujiwara, S.; Kamada, H.; Coupland, G.; Mizoguchi, T. Antisense Suppression of the Arabidopsis PIF3 Gene Does Not Affect Circadian Rhythms but Causes Early Flowering and Increases FT Expression. FEBS Lett. 2004, 557, 259–264. [Google Scholar] [CrossRef] [PubMed]
- Lin, L.; Liu, X.; Yin, R. PIF3 Integrates Light and Low Temperature Signaling. Trends Plant Sci. 2018, 23, 93–95. [Google Scholar] [CrossRef]
- Oh, E.; Yamaguchi, S.; Kamiya, Y.; Bae, G.; Chung, W.-I.; Choi, G. Light Activates the Degradation of PIL5 Protein to Promote Seed Germination through Gibberellin in Arabidopsis. Plant J. 2006, 47, 124–139. [Google Scholar] [CrossRef]
- Oh, E.; Yamaguchi, S.; Hu, J.; Yusuke, J.; Jung, B.; Paik, I.; Lee, H.-S.; Sun, T.; Kamiya, Y.; Choi, G. PIL5, a Phytochrome-Interacting bHLH Protein, Regulates Gibberellin Responsiveness by Binding Directly to the GAI and RGA Promoters in Arabidopsis Seeds. Plant Cell 2007, 19, 1192–1208. [Google Scholar] [CrossRef] [PubMed]
- Renau-Morata, B.; Nebauer, S.G.; García-Carpintero, V.; Cañizares, J.; Gómez Minguet, E.; de los Mozos, M.; Molina, R.V. Flower Induction and Development in Saffron: Timing and Hormone Signalling Pathways. Ind. Crops Prod. 2021, 164, 113370. [Google Scholar] [CrossRef]
- Liu, X.-J.; An, X.-H.; Liu, X.; Hu, D.-G.; Wang, X.-F.; You, C.-X.; Hao, Y.-J. MdSnRK1.1 Interacts with MdJAZ18 to Regulate Sucrose-Induced Anthocyanin and Proanthocyanidin Accumulation in Apple. J. Exp. Bot. 2017, 68, 2977–2990. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.-B.; Li, S.; Zhang, R.-F.; Zhao, J.; Chen, Y.-C.; Zhao, Q.; Yao, Y.-X.; You, C.-X.; Zhang, X.-S.; Hao, Y.-J. The bHLH Transcription Factor MdbHLH3 Promotes Anthocyanin Accumulation and Fruit Colouration in Response to Low Temperature in Apples. Plant Cell Environ. 2012, 35, 1884–1897. [Google Scholar] [CrossRef]
- Qi, T.; Wang, J.; Huang, H.; Liu, B.; Gao, H.; Liu, Y.; Song, S.; Xie, D. Regulation of Jasmonate-Induced Leaf Senescence by Antagonism between bHLH Subgroup IIIe and IIId Factors in Arabidopsis. Plant Cell 2015, 27, 1634–1649. [Google Scholar] [CrossRef] [PubMed]
- Abe, H.; Urao, T.; Ito, T.; Seki, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) Function as Transcriptional Activators in Abscisic Acid Signaling. Plant Cell 2003, 15, 63–78. [Google Scholar] [CrossRef] [PubMed]
- Song, S.; Qi, T.; Fan, M.; Zhang, X.; Gao, H.; Huang, H.; Wu, D.; Guo, H.; Xie, D. The bHLH Subgroup IIId Factors Negatively Regulate Jasmonate-Mediated Plant Defense and Development. PLOS Genet. 2013, 9, e1003653. [Google Scholar] [CrossRef] [PubMed]
- Qi, T.; Song, S.; Ren, Q.; Wu, D.; Huang, H.; Chen, Y.; Fan, M.; Peng, W.; Ren, C.; Xie, D. The Jasmonate-ZIM-Domain Proteins Interact with the WD-Repeat/bHLH/MYB Complexes to Regulate Jasmonate-Mediated Anthocyanin Accumulation and Trichome Initiation in Arabidopsis thaliana. Plant Cell 2011, 23, 1795–1814. [Google Scholar] [CrossRef]
- Nemie-Feyissa, D.; Heidari, B.; Blaise, M.; Lillo, C. Analysis of Interactions between Heterologously Produced bHLH and MYB Proteins That Regulate Anthocyanin Biosynthesis: Quantitative Interaction Kinetics by Microscale Thermophoresis. Phytochemistry 2015, 111, 21–26. [Google Scholar] [CrossRef]
- Li, Y.; Shan, X.; Gao, R.; Yang, S.; Wang, S.; Gao, X.; Wang, L. Two IIIf Clade-bHLHs from Freesia Hybrida Play Divergent Roles in Flavonoid Biosynthesis and Trichome Formation When Ectopically Expressed in Arabidopsis. Sci Rep 2016, 6, 30514. [Google Scholar] [CrossRef]
- Baudry, A.; Heim, M.A.; Dubreucq, B.; Caboche, M.; Weisshaar, B.; Lepiniec, L. TT2, TT8, and TTG1 Synergistically Specify the Expression of BANYULS and Proanthocyanidin Biosynthesis in Arabidopsis thaliana. Plant J. 2004, 39, 366–380. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, A.; Zhao, M.; Leavitt, J.M.; Lloyd, A.M. Regulation of the Anthocyanin Biosynthetic Pathway by the TTG1/bHLH/Myb Transcriptional Complex in Arabidopsis Seedlings. Plant J. 2008, 53, 814–827. [Google Scholar] [CrossRef] [PubMed]
- Payne, C.T.; Zhang, F.; Lloyd, A.M. GL3 Encodes a bHLH Protein That Regulates Trichome Development in Arabidopsis Through Interaction with GL1 and TTG1. Genetics 2000, 156, 1349–1362. [Google Scholar] [CrossRef] [PubMed]
- Bouyer, D.; Geier, F.; Kragler, F.; Schnittger, A.; Pesch, M.; Wester, K.; Balkunde, R.; Timmer, J.; Fleck, C.; Hülskamp, M. Two-Dimensional Patterning by a Trapping/Depletion Mechanism: The Role of TTG1 and GL3 in Arabidopsis Trichome Formation. PLOS Biol. 2008, 6, e141. [Google Scholar] [CrossRef]
- Gao, C.; Guo, Y.; Wang, J.; Li, D.; Liu, K.; Qi, S.; Jin, C.; Duan, S.; Gong, J.; Li, Z.; et al. Brassica napusGLABRA3-1 Promotes Anthocyanin Biosynthesis and Trichome Formation in True Leaves When Expressed in Arabidopsis Thaliana. Plant Biol. 2018, 20, 3–9. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Wang, X.; Zhu, D.; Cui, S.; Li, X.; Cao, Y.; Ma, L. A Single Amino Acid Substitution in IIIf Subfamily of Basic Helix-Loop-Helix Transcription Factor AtMYC1 Leads to Trichome and Root Hair Patterning Defects by Abolishing Its Interaction with Partner Proteins in Arabidopsis. J. Biol. Chem. 2012, 287, 14109–14121. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, L.; Zhang, D.; Su, D.; Li, W.; Wang, X.; Chen, Q.; Cai, W.; Xu, L.; Cao, F.; et al. Comprehensive Analysis of Metabolome and Transcriptome Reveals the Mechanism of Color Formation in Different Leave of Loropetalum chinense var. rubrum. BMC Plant Biol. 2023, 23, 133. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, L.; Zhang, D.; Liu, Y.; Lin, L.; Xiong, X.; Zhang, D.; Sun, M.; Cai, M.; Yu, X.; et al. Transcriptomic and Metabolomic Profiling Provides Insights into Flavonoid Biosynthesis and Flower Coloring in Loropetalum chinense and Loropetalum chinense var. rubrum. Agronomy 2023, 13, 1296. [Google Scholar] [CrossRef]
- Liu, Y.; Zhang, Y.; Liu, Y.; Lin, L.; Xiong, X.; Zhang, D.; Li, S.; Yu, X.; Li, Y. Genome-Wide Identification and Characterization of WRKY Transcription Factors and Their Expression Profile in Loropetalum chinense var. rubrum. Plants 2023, 12, 2131. [Google Scholar] [CrossRef]
- Bateman, A.; Birney, E.; Durbin, R.; Eddy, S.R.; Howe, K.L.; Sonnhammer, E.L.L. The Pfam Protein Families Database. Nucleic Acids Res. 2000, 28, 263–266. [Google Scholar] [CrossRef]
- Potter, S.C.; Luciani, A.; Eddy, S.R.; Park, Y.; Lopez, R.; Finn, R.D. HMMER Web Server: 2018 Update. Nucleic Acids Res. 2018, 46, W200–W204. [Google Scholar] [CrossRef] [PubMed]
- Salih, H.; Tan, L.; Htet, N.N.W. Genome-Wide Identification, Characterization of bHLH Transcription Factors in Mango. Trop. Plant Biol. 2021, 14, 72–81. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A Toolkit for Detection and Evolutionary Analysis of Gene Synteny and Collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef] [PubMed]
- Conesa, A.; Gotz, S.; Garcia-Gomez, J.M.; Terol, J.; Talon, M.; Robles, M. Blast2GO: A Universal Tool for Annotation, Visualization and Analysis in Functional Genomics Research. Bioinformatics 2005, 21, 3674–3676. [Google Scholar] [CrossRef] [PubMed]
- Galili, T.; O’Callaghan, A.; Sidi, J.; Sievert, C. Heatmaply: An R Package for Creating Interactive Cluster Heatmaps for Online Publishing. Bioinformatics 2018, 34, 1600–1602. [Google Scholar] [CrossRef] [PubMed]
- Hu, K. Become Competent in Generating RNA-Seq Heat Maps in One Day for Novices Without Prior R Experience. In Nuclear Reprogramming: Methods and Protocols; Hu, K., Ed.; Methods in Molecular Biology; Springer US: New York, NY, USA, 2021; pp. 269–303. ISBN 978-1-07-161084-8. [Google Scholar]
- Li, M.; Sun, L.; Gu, H.; Cheng, D.; Guo, X.; Chen, R.; Wu, Z.; Jiang, J.; Fan, X.; Chen, J. Genome-Wide Characterization and Analysis of bHLH Transcription Factors Related to Anthocyanin Biosynthesis in Spine Grapes (Vitis davidii). Sci. Rep. 2021, 11, 6863. [Google Scholar] [CrossRef] [PubMed]
- Shi, Q.; Li, X.; Du, J.; Li, X. Anthocyanin Synthesis and the Expression Patterns of bHLH Transcription Factor Family during Development of the Chinese Jujube Fruit (Ziziphus jujuba Mill.). Forests 2019, 10, 346. [Google Scholar] [CrossRef]




| Subfamilies | Number of LcbHLHs | Number of AtbHLHs | Number of FcbHLHs | Number of OsbHLHs |
|---|---|---|---|---|
| Ia | 12 | 10 | 8 | 15 |
| Ib | 8 | 13 | 7 | 7 |
| II | 8 | 4 | 0 | 2 |
| III(a + b + c) | 19 | 10 | 9 | 10 |
| III(d + e) | 11 | 8 | 6 | 5 |
| IIIf | 3 | 4 | 1 | 7 |
| IVa | 11 | 4 | 4 | 6 |
| IVb | 3 | 3 | 4 | 4 |
| IVc | 7 | 4 | 3 | 4 |
| IVd | 1 | 2 | 1 | 9 |
| Va | 3 | 3 | 3 | 3 |
| Vb | 8 | 5 | 6 | 9 |
| VI | 1 | 2 | 2 | 0 |
| VII(a + b) | 17 | 15 | 7 | 14 |
| VIIIa | 6 | 4 | 3 | 1 |
| VIII(b + c) | 11 | 11 | 14 | 18 |
| IX | 8 | 6 | 5 | 5 |
| X | 1 | 10 | 14 | 14 |
| XI | 5 | 5 | 4 | 7 |
| XII | 20 | 17 | 13 | 18 |
| XIII | 0 | 3 | 4 | 3 |
| XIV | 0 | 3 | 0 | 3 |
| XV | 0 | 4 | 0 | 6 |
| Orphans | 2 | 9 | 0 | 3 |
| Motif | Sequence |
|---|---|
| 1 | SHSLAERRRRERJNERFKALRSLVPNCSK |
| 2 | MDKASMLDEAIEYVKELQRQVQELSMKLE |
| 3 | EEPKSDYIHVRARRGQATD |
| 4 | QVMSFEQSNWDASVHEIQGMTSFEHPHNQDQQLHLLHEMQQNGHHHPQSF |
| 5 | FVQKPANFQTSLGFLGDLPTPDNASASSVLYDPLFHLNLPPQPPLFRDLF |
| 6 | YNLPASRTASLFGGGIDEKEGSGGVYQNGVATQFDNGVLEFTGDIGGMGK |
| 7 | RJVSALEKLGLDIVHANVSTF |
| 8 | ERAKLAKSAGIRTLVCIPTASGVVELGSTEIIKEDLGLIQLIKSLF |
| 9 | NSSTLPDTSPYINNPPTQHLLNLFHLPRCSPSSNLLPNSSI |
| 10 | DNIRLSMEELSYHQNPHQEDDAALEQHLGFDMENCYNINNN |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Y.; Lin, L.; Liu, Y.; Mo, Q.; Zhang, D.; Li, W.; Xiong, X.; Yu, X.; Li, Y. Genome-Wide Analysis of the bHLH Gene Family in Loropetalum chinense var. rubrum: Identification, Classification, Evolution, and Diversity of Expression Patterns under Cultivation. Plants 2023, 12, 3392. https://doi.org/10.3390/plants12193392
Liu Y, Lin L, Liu Y, Mo Q, Zhang D, Li W, Xiong X, Yu X, Li Y. Genome-Wide Analysis of the bHLH Gene Family in Loropetalum chinense var. rubrum: Identification, Classification, Evolution, and Diversity of Expression Patterns under Cultivation. Plants. 2023; 12(19):3392. https://doi.org/10.3390/plants12193392
Chicago/Turabian StyleLiu, Yang, Ling Lin, Yang Liu, Qiong Mo, Damao Zhang, Weidong Li, Xingyao Xiong, Xiaoying Yu, and Yanlin Li. 2023. "Genome-Wide Analysis of the bHLH Gene Family in Loropetalum chinense var. rubrum: Identification, Classification, Evolution, and Diversity of Expression Patterns under Cultivation" Plants 12, no. 19: 3392. https://doi.org/10.3390/plants12193392
APA StyleLiu, Y., Lin, L., Liu, Y., Mo, Q., Zhang, D., Li, W., Xiong, X., Yu, X., & Li, Y. (2023). Genome-Wide Analysis of the bHLH Gene Family in Loropetalum chinense var. rubrum: Identification, Classification, Evolution, and Diversity of Expression Patterns under Cultivation. Plants, 12(19), 3392. https://doi.org/10.3390/plants12193392







