Embryo Rescue in Plant Breeding
Abstract
1. Introduction
2. The Origins of Embryo Rescue: A Brief Historical Overview
3. The Embryo Rescue Techniques
3.1. Pre-Zygotic Barriers in Hybridization and Techniques to Overcome Them
3.2. Post-Zygotic Barriers in Hybridization and Techniques to Overcome Them
3.3. Medium Composition and Environmental Factors Suitable for Embryo Rescue Techniques
4. Embryo Rescue Applications
4.1. Embryo Rescue in Hybrid Plants Obtained to Introduce Resistance Genes to Abiotic and Biotic Stresses and New Agronomic Traits into Crops
4.1.1. Dicot Species
| Crops | Traits | Introgressed from Wild/Crops | References |
|---|---|---|---|
| Brassica rapa | 1 CMS and Rfp | Brassica napus | [133] |
| Brassica napus | Tolerance to herbicides | Sinapis arvensis | [135] |
| Brassica juncea | Tolerance to herbicides | Brassica kaber (S. arvensis) | [134] |
| Brassica juncea | Tolerance to aphids | Brassica tournefortii | [154] |
| Brassica juncea | Tolerance to drought | Brassica tournefortii | [154] |
| Brassica rapa | Tolerance to herbicides | Brassica kaber (S. arvensis) | [134] |
| Brassica rapa subsp. pekinensis | Yellow/orange color of petals | Brassica juncea × B. rapa | [136] |
| Brassica napus | Clubroot-resistant gene, CRd | Brassica rapa subsp. pekinensis | [139,142] |
| Brassica napus | Resistance to blackleg disease | Brassica nigra | [141] |
| Brassica oleracea var. botrytis | Resistance to black rot disease | Brassica carinata | [146] |
| Brassica oleracea | Resistance to atrazine | Brassica napus | [147] |
| Brassica oleracea var. capitata | Resistance to Plasmodiophora brassicae | Brassica napus | [150] |
| Brassica oleracea var. botrytis | Resistance to powdery mildew | Brassica carinata | [149] |
| Brassica oleracea | Resistance to black rot disease | Brassica spp. | [151] |
| Brassica oleracea | Male sterility | Brassica napus | [152] |
| Brassica oleracea | CMS Tour | Brassica tournefortii | [153] |
| Phaseolus vulgaris | Resistance to Ascochyta blight | Phaseolus coccineus | [171] |
| Phaseolus vulgaris | Resistance to bean golden mosaic virus | Phaseolus polyanthus | [171] |
| Phaseolus vulgaris | Resistance to bean fly | Phaseolus coccineus | [171] |
| Solanum lycopersicum | Resistance to tobacco mosaic virus | Solanum peruvianum | [173] |
| Solanum lycopersicum | Resistance to tomato spotted wilt virus | Solanum peruvianum | [173] |
| Solanum lycopersicum | Resistance to root-knot nematode | Solanum peruvianum | [173] |
| Carica papaya | Resistance to papaya ringspot virus | Carica cauliflora | [174] |
| Vitis vinifera | Resistance to Plasmopora viticola | Chinese wild Vitis spp. | [193,195] |
| Vitis vinifera | Seedless | Chinese wild Vitis spp. | [194,195] |
| Vitis vinifera | Seedless | Vitis vinifera | [196] |
| Glycine max | Resistance to cyst nematode | Glycine tomentella | [185] |
| Gypsophila paniculata | Double flowers | Gypsophila manginii | [121] |
| Gentiana scabra | Flower color | Gentiana straminea | [118] |
| Gentiana triflora | Flower morphology | Gentiana paradoxa | [118] |
| Gentiana paradoxa (female) | Petaloid stamen | Gentiana septemfida (male) | [118] |
| Campanula carpatica | Flower shape | Campanula carpatica | [117] |
| Campanula medium | Tolerance to cold | Campanula formanekiana | [117] |
| Chrysanthemum × grandiflorum | Resistance to aphids | Chrysanthemum przewalskii | [15,112] |
| Chrysanthemum × grandiflorum | Tolerance to heat | Chrysanthemum makinoi | [15] |
| Chrysanthemum × grandiflorum | Tolerance to drought | Chrysanthemum indicum | [114] |
| Chrysanthemum × morifolium | Tolerance to salt | Artemisia japonica | [115] |
| Dendranthema crissum | Heterosis | Crossostephium chinense | [116] |
| Camellia japonica | Yellow color of petals | Camellia chrysantha | [120] |
4.1.2. Monocot Species
| Crops | Traits | Introgressed from Wild/Crops | References |
|---|---|---|---|
| Alstroemeria hybrid | Winter hardiness | Alstroemeria aurea | [216] |
| Alstroemeria hybrid | Fragrant flowers | Alstroemeria caryophyllea | [216] |
| Alstroemeria hybrid | Ever-blooming growth habit | Alstroemeria caryophyllea | [216] |
| Alstroemeria hybrid | Long flower stems | Alstroemeria caryophyllea | [216] |
| Alstroemeria hybrid | Flower color and morphology | Alstroemeria pelegrina | [217] |
| Alstroemeria hybrid | Flower color and morphology | Alstroemeria caryophyllea | [217] |
| Alstroemeria hybrid | Fragrant flowers | Alstroemeria caryophyllea | [217] |
| Hordeum vulgare | Resistance to powdery mildew | Hordeum bulbosum | [210,211] |
| Triticum aestivum | Resistance to powdery mildew | Secale cereale (addition line 1 ch. 6R) | [204,205] |
| Triticum aestivum | Multiple fungal disease resistance | Secale cereale (addition line 2 ch. 4R) | [207] |
| Elaeis guineensis | Resistance to fatal yellowing | Elaeis oleifera | [94] |
| Littonia modesta | Climbing habit | Sandersonia aurantiaca | [220] |
| Littonia modesta | Color of corolla | Sandersonia aurantiaca | [220] |
| Littonia modesta | Climbing habit | Gloriosa spp. | [220] |
| Littonia modesta | Pendulous flowers | Gloriosa spp. | [220] |
| Littonia modesta | Undulate and slightly reflexed tepals | Gloriosa spp. | [220] |
| Littonia modesta | Modification of corolla color | Gloriosa spp. | [220] |
| Sandersonia aurantiaca | Pendulous flowers | Gloriosa spp. | [220] |
| Sandersonia aurantiaca | Undulate tepals fused at the base | Gloriosa spp. | [220] |
| Tulipa gesneriana | Dark basic color of corolla | Tulipa kaufmanniana | [64] |
| Tulipa gesneriana | Early flowering time | Tulipa kaufmanniana | [64] |
| Tulipa gesneriana | Long pointed petals | Tulipa kaufmanniana | [64] |
| Tulipa gesneriana | Petals with carmine-red dorsal band | Tulipa kaufmanniana | [64] |
| Lilium hansonii (female) | Increase in flower spots | L. brownii var. colchesteri (male) | [65] |
| Lilium hansonii (female) | Increase in flower size | L. brownii var. colchesteri (male) | [65] |
| Lilium hansonii (female) | Increase in number of buds | L. brownii var. colchesteri (male) | [65] |
| Lilium hansonii (female) | Tepal size | L. brownii var. colchesteri (male) | [65] |
| Lilium hansonii (female) | Reduction in plant height | L. brownii var. colchesteri (male) | [65] |
| Lilium hansonii (female) | Color of pollen and stigma | L. brownii var. colchesteri (male) | [65] |
| Lilium hansonii (female) | Flower orientation | L. brownii var. colchesteri (male) | [65] |
4.2. Embryo Rescue to Recover Haploids, Dihaploids, Doubled Haploids, and Polyploid Embryos
4.2.1. Dicot Species
4.2.2. Monocot Species
4.2.3. Genome Editing to Induce Haploid Plants
| Crops | Methods | Ploidy Level | Main Objective | References |
|---|---|---|---|---|
| Cichorium intybus (2n = 2x = 18) | 1 IG cross with Cicerbita alpina | Gynogenetic haploids | 2 DHs | [242] |
| Lactuca sativa (2n = 2x = 18) | IG crosses with Helianthus spp. | Gynogenetic haploids | DHs | [243] |
| Rosa × hybrida (2n = 4x = 28) | 3 IS crosses with Rosa spp. (2n = 4x, 5x, 6x) | Triploids and tetraploids | Sterility and variability | [244] |
| Primula sinensis (2n = 2x = 24) (male) | IS cross with P. filchnerae (diploid) | Triploids and hexaploids | Genetic variability | [246] |
| Selenicereus megalanthus (2n = 4x = 44) | Gynogenesis | DHs, triploids, tetraploids | Cacti breeding | [247] |
| Hylocereus megalanthus (2n = 4x = 44) (male) | IS crosses with H. megalanthus and H. undatus (diploids) | Diploids, triploids, tetraploids | Cacti breeding | [248] |
| Gentiana spp. (2n = 2x = 26) | Gynogenesis | Haploids and diploids | DHs | [249] |
| Begonia semperflorens (2n = 2x) and (2n = 4x) | IS cross with B. “Orange Rubra” (2n = 2x) | Haploids, DHs, and others | Begonia spp. breeding | [251] |
| Dianthus caryophyllus (2n = 2x = 30) | Impollination with RX-treated pollen and gynogenesis | Haploids and DHs | Carnation breeding | [252] |
| Cyclamen persicum (2n = 2x = 48) | IS cross with C. hederifolium (2n = 2x = 34) | IS sterile hybrids | Fertile 4AD (2n = 82) | [255] |
| Cyclamen persicum (2n = 2x = 48) | IS cross with C. purpurascens (2n = 2x = 34) | IS sterile hybrids | Fertile AD | [257] |
| Cyclamen persicum (2n = 2x = 48) | IS cross with C. graecum (2n = 4x = 84) | IS low fertile hybrids | Cyclamen breeding | [256] |
| Cyclamen persicum (2n = 4x = 96) | IS cross with C. graecum (2n = 4x = 84) | IS fertile hybrids | Disease resistance | [256] |
| Jatropha curcas (2n = 2x = 22) | IG cross with Ricinus communis (2n = 2x = 20) | IG sterile hybrids | AD | [258] |
| Vitis vinifera × V. labrusca (2n = 4x = 78) | IG cross with V. vinifera diploid (2n = 2x = 38) | Haploids and triploids | DHs | [260] |
| Enarthrocarpus lyratus (2n = 2x = 20) | IG cross with B. oleracea (2n = 2x = 18) | IG sterile hybrids | Fertile AD (2n = 38) | [131] |
| Brassica rapa (2n = 2x = 16) | Androgenesis from 5 BC2 of B. rapa × B. napus | Haploids and DHs | Rfp gene cloning | [133] |
| Brassica oleracea ssp. acephala (2n = 2x = 18) | IS cross with B. rapa ssp. rapifera (2n = 2x = 20) | IS hybrids | Fertile AD (2n = 38) | [265] |
| Cucumis melo (2n = 2x = 24) | Irradiated pollen (Co60 γ- and X-rays) | Gynogenetic haploids | DHs | [269,270,271,272,273] |
| Citrus reticulata (2n = 2x = 18) | IS cross with C. sinensis (2n = 2x = 18) | Sterile triploids | Seedless | [278,279] |
| Citrus clementina (2n = 2x = 18) | IS cross with C. grandis (2n = 2x = 18) | Sterile triploids | Seedless | [277] |
| Citrus clementina (2n = 2x = 18) | IS with C. paradise × C. tangerine, tetraploid | Sterile triploids | Seedless | [278] |
| Glycine max (2n = 2x = 40) (male) | IS with G. tomentella (2n = 78) | Sterile hybrids (2n = 59) | AD (2n = 118) | [281] |
| Solanum lycopersicum (2n = 2x = 24) | Genome editing of the SlDMP gene | Gynogenetic haploids | DHs | [388] |
| Nicotiana tabacum (2n = 4x = 48) | Genome editing of the NtDMP gene | Gynogenetic haploids | DHs | [389] |
| Brassica napus (2n = 4x = 38) | Genome editing of the BnDMP gene | Gynogenetic haploids | DHs | [390] |
| Medicago truncatula (2n = 2x = 16) | Genome editing of the MtDMP gene | Gynogenetic haploids | DHs | [391] |
| Brassica oleracea (2n = 2x = 20) | Genome editing of the BoDMP gene | Gynogenetic haploids | DHs | [392] |
| Solanum tuberosum (2n = 4x = 48) | Genome editing of the StDMP gene | Gynogenetic haploids | DHs | [393] |
| Crops | Methods | Ploidy Level | Main Objective | References |
|---|---|---|---|---|
| Hemerocallis (2n = 2x = 22) | 1 IS cross with tetraploid cultivars | Triploids | Sterility | [289] |
| Zantedeschia elliottiana (2n = 2x = 32) | IS cross with tetraploid cultivars | Triploids | Sterility | [290] |
| Lilium auratum (2n = 2x = 24) | IS cross with L. henryi (2n = 2x = 24) | Hybrids producing 2n gametes | Polyploidy | [287] |
| Lilium spp. Orientalis (2n = 2x = 24) | IS cross with Lilium spp. (Asiatic hybrid) | Triploids | Variability | [286] |
| Aegilops ovate (2n = 4x = 28) | 2 IG cross with Secale cereale (2n =2x = 14) | Sterile triploids (2n = 3x = 21) | Fertile 3 AD | [302] |
| Avena sativa (2n = 6x = 42) | IG cross with Zea mays (2n = 2x = 20) (male) | 4 AL (2n = 6x = 42 +1, 2) | 5 OMA | [30,338] |
| Oryza sativa (2n = 2x = 24) | Androgenesis (anther culture) | Haploids | 6 DHs | [350,351,352,353,354,355,356,357,358,359,360,361,362,363,364,365,366,367,368,369,370,371,372,373,374,375,376,377,378,379] |
| Hordeum vulgare (2n = 2x = 14) | IS cross with H. bulbosum (2n = 2x =14) (male) | Gynogenetic haploids | DHs | [304,305,306,307,308] |
| Hordeum vulgare (2n = 2x = 14) | Cross with H. vulgare (genotype: hap/hap) (female) | Gynogenetic haploids | DHs | [304,314] |
| Triticum aestivum (2n = 6x = 42) | IG cross with H. vulgare (2n = 2x = 14) (male) | Gynogenetic haploids | DHs | [315] |
| Triticum aestivum (2n = 6x = 42) | IG cross with Z. mays (2n = 2x = 20) (male) | Gynogenetic haploids | DHs | [316,317,318,319,320,321,322,323,324,325] |
| Triticum aestivum (2n = 6x = 42) | IG cross with Pennisetum glaucum (2n = 2x = 14) (male) | Gynogenetic haploids | DHs | [326] |
| Triticum aestivum (2n = 6x = 42) | IG cross with Imperata cylindrical (2n = 2x = 20) (male) | Gynogenetic haploids | DHs | [327,328] |
| Triticum aestivum (2n = 6x = 42) | IG cross with Coix lacryma-jobi (2n = 2x = 20) (male) | Gynogenetic haploids | DHs | [329] |
| Triticum turgidum ssp. turgidum (2n = 4x =28) | IG cross with Z. mays (2n = 2x = 20) (male) | Gynogenetic haploids | DHs | [330] |
| Triticum turgidum (2n = 4x =28) | IG cross with Z. mays (2n = 2x = 20) (male) | Gynogenetic haploids | DHs | [333,334,335,336] |
| Triticum turgidum (2n = 4x =28) | IG cross with P. glaucum (2n = 2x = 14) (male) | Gynogenetic haploids | DHs | [336] |
| Avena sativa (2n = 6x = 42) | IG cross with Z. mays (2n = 2x = 20) (male) | Gynogenetic haploids | DHs | [31,337,338,339,340,341, 343,345] |
| Secale cereale (2n = 2x = 14) | IG cross with Z. mays (2n = 2x = 20) (male) | Gynogenetic haploids | DHs | [349] |
| Zea mays (2n = 2x = 20) (male) | Cross with Z. mays (genotype: cenh3/CENH3) | Androgenetic haploids | DHs | [307] |
| Triticum aestivum (2n = 6x = 42) | Genome editing of the MATRILINEAL (TaMTL) gene | Gynogenetic haploids | DHs | [383] |
| Zea mays (2n = 2x = 20) | Genome editing of the ZmMATL gene | Gynogenetic haploids | DHs | [384] |
| Oryza sativa (2n = 2x =24) | Genome editing of the OsMATL gene | Gynogenetic haploids | DHs | [385] |
| Oryza sativa (2n = 2x =24) | Genome editing of the OsECS1/ECS2 genes | Gynogenetic haploids | DHs | [387] |
| Setaria italica (2n = 2x = 18) | Genome editing of the SiMATL gene | Gynogenetic haploids | DHs | [386] |
4.3. Analysis of Embryogenesis with Embryo Rescue Technical Holder
4.4. Embryo Rescue to Overcome Seed Dormancy, Fruit Drops; to Propagate Rare Plant Species; and to Speed up the Selection Processes
5. Conclusions and Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Armenta-Medina, A.; Gillmor, C.S.; Gao, P.; Mora-Macias, J.; Kochian, L.V.; Xiang, D.; Datla, R. Developmental and genomic architecture of plant embryogenesis: From model plant to crops. Plant Commun. 2021, 2, 100136. [Google Scholar] [CrossRef] [PubMed]
- Soyars, C.L.; James, S.R.; Nimchuk, Z.L. Ready, aim, shoot: Stem cell regulation of the shoot apical meristem. Curr. Opin. Plant Biol. 2016, 29, 163–168. [Google Scholar] [CrossRef] [PubMed]
- Sussex, I.M.; Kerk, N.M. The evolution of plant architecture. Curr. Opin. Plant Biol. 2001, 4, 33–37. [Google Scholar] [CrossRef]
- De Smet, I.; Lau, S.; Mayer, U.; Jürgens, G. Embryogenesis—The humble beginnings of plant life. Plant J. 2010, 61, 959–970. [Google Scholar] [CrossRef]
- Olsen, O.A. Nuclear endosperm development in cereals and Arabidopsis thaliana. Plant Cell 2004, 16, 214–228. [Google Scholar] [CrossRef]
- Xiang, D.; Quilichini, T.D.; Liu, Z.; Gao, P.; Pan, Y.; Li, Q.; Nilsen, K.T.; Venglat, P.; Esteban, E.; Pasha, A.; et al. The transcriptional landscape of polyploid wheats and their diploid ancestors during embryogenesis and grain development. Plant Cell 2019, 31, 2888–2911. [Google Scholar] [CrossRef]
- Itoh, J.-I.; Sato, Y.; Sato, Y.; Hibara, K.-I.; Sato, S.S.; Kobayashi, H.; Takehisa, H.; Sanguinet, K.A.; Namiki, N.; Nagamura, Y. Genome-wide analysis of spatiotemporal gene expression patterns during early embryogenesis in rice. Development 2016, 143, 1217–1227. [Google Scholar] [CrossRef]
- Meinke, D.W. Genome-wide identification of EMBRYO-DEFECTIVE (EMB) genes required for growth and development in Arabidopsis. New Phytol. 2020, 226, 306–325. [Google Scholar] [CrossRef]
- Basile, A.; Fambrini, M.; Pugliesi, C. The vascular plants: Open system of growth. Dev. Genes Evol. 2017, 227, 129–157. [Google Scholar] [CrossRef]
- Mayer, U.; Ruiz, R.; Berleth, T.; Miséra, S.; Jürgens, G. Mutations affecting body organization in the Arabidopsis embryo. Nature 1991, 353, 402–407. [Google Scholar] [CrossRef]
- Raghavan, V. Experimental Embryogenesis in Vascular Plants; Experimental Botany; Academic Press: London, UK, 1976; Volume 10, pp. 187–192. [Google Scholar]
- Raghavan, V. One hundred years of zygotic embryo culture investigations. Vitr. Cell Dev. Biol. Plant 2003, 39, 437–442. [Google Scholar] [CrossRef]
- Litz, R.E. Germplasm modification and its potential for finding new sources of resistance to diseases. J. Nematol. 1986, 18, 18–22. [Google Scholar] [PubMed]
- Sharma, D.R.; Kaur, R.; Kumar, K. Embryo rescue in plants: A review. Euphytica 1996, 89, 325–337. [Google Scholar] [CrossRef]
- Cheng, X.; Chen, S.M.; Chen, F.D.; Fang, W.M.; Deng, Y.M.; She, L.F. Interspecific hybrids between Dendranthema morifolium (Ramat.) Kitamura and D. nankingense (Nakai) Tzvel. achieved using ovary rescue and their cold tolerance characteristics. Euphytica 2010, 172, 101–108. [Google Scholar] [CrossRef]
- Deng, Y.; Teng, N.; Chen, S.; Guan, Z.; Song, A.; Chang, Q. Reproductive barriers in the intergeneric hybridization between Chrysanthemum grandiflorum (Ramat.) Kitam. and Ajania przewalskii Poljak. (Asteraceae). Euphytica 2010, 174, 41–50. [Google Scholar] [CrossRef]
- Shen, X.; Gmitter, F.G., Jr.; Grosser, J.W. Immature embryo rescue and culture. Methods Mol. Biol. 2011, 710, 75–92. [Google Scholar] [CrossRef]
- Loyola-Vargas, V.M.; Ochoa-Alejo, N. An introduction to plant cell culture: The future ahead. Methods Mol. Biol. 2012, 877, 1–8. [Google Scholar] [CrossRef]
- Krishna, R.; Khandagale, K.; Benke, A.P.; Soumia, P.S.; Manjunathagowda, D.C.; Ansari, W.A.; Mokat, D.N.; Gawande, S.J.; Ade, A.B.; Singh, M. Embryo rescue: A potential tool for improvement of economically important crops. In Advances in Plant Tissue Culture, Current Developments and Future Trends; Chapter 11; Rai, A.C., Kumar, A., Modi, A., Singh, M., Eds.; Academic Press: Cambridge, MA, USA, 2022; pp. 259–282. ISBN 9780323907958. [Google Scholar] [CrossRef]
- Schaal, B. Plants and people: Our shared history and future. Plant People Planet 2019, 1, 14–19. [Google Scholar] [CrossRef]
- Purugganan, M.D. Evolutionary insights into the nature of plant domestication. Curr. Biol. 2019, 29, R705–R714. [Google Scholar] [CrossRef]
- Zenkteler, M. In vitro pollination of angiosperm ovules with gymnosperm pollen grains. Vitr. Cell Dev. Biol. Plant 2000, 36, 125–127. [Google Scholar] [CrossRef]
- Rangaswamy, N.S. Application of in vitro pollination and in vitro fertilization. In Applied and Fundamental Aspects of Plant Cell Tissue, and Organ Culture; Reinert, J., Bajaj, Y.P.S., Eds.; Springer: Berlin/Heidelberg, Germany, 1977; pp. 412–425. [Google Scholar]
- Zenkteler, M. In vitro pollination of excised ovaries. Acta Biol. Cracoviensia Ser. Bot. 1999, 41, 31–38. [Google Scholar]
- Pierik, R.L.M. In Vitro Culture of Higher Plants; Martinus Nijhoff Publisher: Dordrecht, The Netherlands, 1987; p. 344. ISBN 9789024735310. [Google Scholar]
- Sahijram, L.; Rao, B.M. Hybrid embryo rescue in crop improvement. In Plant Biology and Biotechnology; Chapter 18; Bahadur, B., Venkat Rajam, M., Sahijram, L., Krishnamurthy, K., Eds.; Springer: New Delhi, India, 2015; pp. 363–384. [Google Scholar]
- Zulkarnain, Z.; Tapingkae, T.; Taji, A. Applications of in vitro techniques in plant breeding. In Advances in Plant Breeding Strategies: Breeding, Biotechnology and Molecular Tools; Chapter 10; Al-Khayri, J.M., Jain, S.M., Johnson, D.V., Eds.; Springer: Cham, Switzerland, 2015; pp. 293–328. [Google Scholar] [CrossRef]
- Pramanik, K.; Sahoo, J.P.; Mohapatra, P.P.; Acharya, L.K.; Jena, C. Insights into the embryo rescue—A modern in-vitro crop improvement approach in horticulture. Plant Cell Biotechnol. Mol. Biol. 2021, 22, 20–33. [Google Scholar]
- Mehbub, H.; Akter, A.; Akter, M.A.; Mandal, M.S.H.; Hoque, M.A.; Tuleja, M.; Mehraj, H. Tissue culture in ornamentals: Cultivation factors, propagation techniques, and its application. Plants 2022, 11, 3208. [Google Scholar] [CrossRef] [PubMed]
- Kynast, R.G.; Riera-Lizarazu, O.; Vales, M.I.; Okagaki, R.J.; Maquieira, S.B.; Chen, G.; Ananiev, E.V.; Odland, W.E.; Russell, C.D.; Stec, A.O.; et al. A complete set of maize individual chromosome additions to the oat genome. Plant Physiol. 2001, 125, 1216–1227. [Google Scholar] [CrossRef][Green Version]
- Skrzypek, E.; Warchoł, M.; Czyczyło-Mysza, I.; Juzoń, K.; Kinga Dziurka, K.; Marcińska, I. Oat doubled haploid production through wide hybridization with maize. Methods Mol. Biol. 2021, 2287, 323–332. [Google Scholar] [CrossRef]
- Hanning, E. Zur physiologie pflanzlicher embryonen. I. Ueber die cultur von cruciferen-embryonen ausserhalb des embrysacks. Bot. Ztg. 1904, 62, 45–80. [Google Scholar]
- Mehetre, S.S.; Aher, A.R. Embryo rescue: A tool to overcome incompatible interspecific hybridization in Gossypium Linn. A review. Ind. J. Biotechnol. 2004, 3, 29–36. [Google Scholar]
- Brown, H.T. On the culture of the exised embryos of barley on nutrient solutions containing nitrogen in different forms. Trans. Guinness Res. Lab. 1906, 1, 288–299. [Google Scholar]
- Dubard, M.; Urbain, J.-A. De l’influence de l’albumen sur le développment de l’embryonen. Compt. Rend. Acad. Sci. Paris 1913, 156, 1086–1089. [Google Scholar]
- Buckner, G.D.; Kastle, J.H. The growth of isolated plant embryos. J. Biol. Chem. 1917, 29, 209–213. [Google Scholar] [CrossRef]
- Andronescu, D.I. Germination and further development of the embryo of Zea mays separated from the endosperm. Am. J. Bot. 1919, 6, 443–452. [Google Scholar] [CrossRef]
- Dieterich, K. Über kultur von embryonen außerhalb des samens. Flora 1924, 117, 379–417. [Google Scholar] [CrossRef]
- Laibach, F. Das taubwerden von bastardsamen und die künstliche Aufzucht früh absterbender bastardembryonen. Z. Bot. 1925, 17, 417–459. [Google Scholar]
- Moyle, L.C.; Jewell, C.P.; Kostyun, J.L. Fertile approaches to dissecting mechanisms of premating and postmating prezygotic reproductive isolation. Curr. Opin. Plant Biol. 2014, 18, 16–23. [Google Scholar] [CrossRef]
- Stebbins, G.L. The inviability, weakness and sterility of interspecific hybrids. Adv. Genet. 1958, 9, 147–215. [Google Scholar] [CrossRef] [PubMed]
- De Jeu, M.J. In vitro techniques for ornamental breeding. Acta Hortic. 2000, 508, 55–60. [Google Scholar] [CrossRef]
- De Verna, J.W.; Myers, J.R.; Collins, G.B. Bypassing prefertilization barriers to hybridization in Nicotiana using in vitro pollination and fertilization. Theor. Appl. Genet. 1987, 73, 665–671. [Google Scholar] [CrossRef]
- Wilcock, C.; Neiland, R. Pollination failure in plants: Why it happens and when it matters. Trends Plant Sci. 2002, 7, 270–277. [Google Scholar] [CrossRef]
- Kinoshita, T. Reproductive barrier and genomic imprinting in the endosperm of flowering plants. Genes Genet. Syst. 2007, 82, 177–186. [Google Scholar] [CrossRef]
- Murthy, K.S.R.; Kondamudi, R.; Rao, P.V.C.; Pullaiah, T. In vitro flowering—A review. J. Agric. Technol. 2012, 8, 1517–1536. [Google Scholar]
- Bomblies, K.; Lempe, J.; Epple, P.; Warthmann, N.; Lanz, C.; Dangl, J.L.; Weigel, D. Autoimmune response as a mechanism for a Dobzhansky-Muller-type incompatibility syndrome in plants. PLoS Biol. 2007, 5, 1962–1972. [Google Scholar] [CrossRef]
- Bikard, D.; Patel, D.; Mette, C.L.; Giorgi, V.; Camilleri, C.; Bennett, M.J.; Loudet, O. Divergent evolution of duplicate genes leads to genetic incompatibilities within A. thaliana. Science 2009, 323, 623–626. [Google Scholar] [CrossRef]
- Yamamoto, E.; Takashi, T.; Moritaka, Y.; Lin, S.; Wu, J.; Matsumoto, T.; Kitano, H.; Matsuoka, M.; Ashikari, M. Gain of deleterious function causes an autoimmune response and Bateson-Dobzhansky-Muller incompatibility in rice. Mol. Genet. Genom. 2010, 283, 305–315. [Google Scholar] [CrossRef] [PubMed]
- Devreux, M. Effets de l’irradiation gamma cronique sur l’embryogénèse de Capsella bursa-pastoris Moench. In L’Energia Nucleare in Agricultura; 6th Congresso Nucleare di Roma; Vallecchi: Firenze, Italy, 1963; pp. 119–217. [Google Scholar]
- Yan, D.; Duermeyer, L.; Leoveanu, C.; Nambara, E. The functions of the endosperm during seed germination. Plant Cell Physiol. 2014, 55, 1521–1533. [Google Scholar] [CrossRef] [PubMed]
- Johnston, S.A.; Nijs, T.P.M.; Peloquin, S.J.; Hanneman, R.E. The significance of genic balance to endosperm development in interspecific crosses. Theor. Appl. Genet. 1980, 57, 5–9. [Google Scholar] [CrossRef] [PubMed]
- Nishiyama, I.; Yabuno, T. Causal relationships between the polar nuclei in double fertilization and interspecific cross-incompatibility in Avena. Cytologia 1978, 43, 453–466. [Google Scholar] [CrossRef]
- Dilkes, B.P.; Comai, L. A differential dosage hypothesis for parental effects in seed development. Plant Cell 2004, 16, 3174–3180. [Google Scholar] [CrossRef]
- Marasek-Ciolakowska, A.; Nishikawa, T.; Shea, D.J.; Okazaki, K. Breeding of lilies and tulips-interspecific hybridization and genetic background. Breed. Sci. 2018, 68, 35–52. [Google Scholar] [CrossRef]
- Zenkteler, M.E.; Nitzsche, W. In vitro culture of ovules of Triticum aestivum at early stages of embryogenesis. Plant Cell Rep. 1985, 4, 168–171. [Google Scholar] [CrossRef]
- Reed, S.M.; Collins, G.B. Interspecific hybrids in Nicotiana through in vitro culture of fertilized ovule. J. Hered. 1978, 69, 311–315. [Google Scholar] [CrossRef]
- Espinasse, A.; Volin, J.; Dybing, C.D.; Lay, C. Embryo rescue through in ovolo culture in Helianthus. Crop. Sci. 1991, 31, 102–108. [Google Scholar] [CrossRef]
- Momotaz, A.; Kato, M.; Kakihara, F. Production of intergeneric hybrids between Brassica and Sinapis species by means of embryo rescue techniques. Euphytica 1998, 103, 123–130. [Google Scholar] [CrossRef]
- Iwai, S.; Kishi, C.; Nakata, K.; Kawashima, N. Production of Nicotiana tabacum × Nicotiana acuminata hybrids by ovule colture. Plant Cell Rep. 1986, 5, 403–404. [Google Scholar] [CrossRef]
- Refaat, M.; Rossignol, L.; Demarly, Y. Interspecific hybrid Gossypium hirsutum L. × Gossypium barbadense L. via in vitro fertilization and ovule colture. Z. Pflanzenzücht. 1984, 93, 137–146. [Google Scholar]
- Hossain, M.M.; Inden, H.; Asahira, T. Intergeneric and interspecific hybrids through in vitro ovule culture in the Cruciferae. Plant Sci. 1988, 58, 121–128. [Google Scholar] [CrossRef]
- Kaneko, Y.; Bang, S.W. Interspecific and intergeneric hybridization and chromosomal engineering of Brassicaceae crops. Breed. Sci. 2014, 64, 14–22. [Google Scholar] [CrossRef]
- Custers, J.B.M.; Eikelboom, W.; Bergervoet, J.H.W.; van Eijk, J.P. Embryo-rescue in the genus Tulipa L.; successful direct transfer of T. kaufmanniana Regel germplasm into T. gesneriana. Euphytica 1995, 82, 253–261. [Google Scholar] [CrossRef]
- Kim, J.-Y.; Song, Y.-S.; Na, J.-K.; Kim, J.-H. Interspecific crossing between Lilium hansonii Leichtlin and L. brownie var. colchesteri for the breeding of new lily cultivars. Agronomy 2022, 12, 621. [Google Scholar] [CrossRef]
- Divakaran, M.; Babu, K.N.; Peter, K.V. Protocols for biotechnological interventions in improvement of vanilla (Vanilla planifolia Andrews.). Methods Mol. Biol. 2016, 1391, 47–63. [Google Scholar] [CrossRef]
- Eeckhaut, T.; De Keyser, E.; Van Huylenbroeck, J.; de Riek, J.; Van Bockstaele, E. Application of embryo rescue after interspecific crosses in the genus Rhododendron. Plant Cell Tissue Organ Cult. 2007, 89, 29–35. [Google Scholar] [CrossRef]
- Munoz, C.E.; Lyrene, P.M. In vitro attempts to overcome the cross-incompatibility between Vaccinium corymbosum L. and V. elliottii Chapm. Theor. Appl. Genet. 1985, 69, 591–596. [Google Scholar] [CrossRef]
- Pathirana, R.; Wiedow, C.; Pathirana, S.; Hedderley, D.; Morgan, E.; Scalzo, J.; Frew, T.; Timmerman-Vaughan, G. Ovule culture and embryo rescue facilitate interspecific hybridisation in blueberry (Vaccinium spp.). Acta Hortic. 2015, 1083, 123–133. [Google Scholar] [CrossRef]
- Erst, A.A.; Gorbunov, A.B.; Asbaganov, S.V.; Tomoshevich, M.A.; Banaev, E.V.; Erst, A.S. Applying biotechnology in the propagation and further selection of Vaccinium uliginosum × (V. corymbosum × V. angustifolium) hybrids. Plants 2021, 10, 1831. [Google Scholar] [CrossRef] [PubMed]
- Gamborg, O.L.; Miller, R.A.; Ojima, K. Nutrient requirements of suspension cultures of soybean root cells. Exp. Cell Res. 1968, 50, 151–158. [Google Scholar] [CrossRef] [PubMed]
- Murashige, T.; Skoog, F. A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol. Plant. 1962, 15, 473–497. [Google Scholar] [CrossRef]
- Nitsch, J.P.; Nitsch, C. Haploid plants from pollen grains. Science 1969, 163, 85–87. [Google Scholar] [CrossRef]
- Kobayashi, K.; Morii, T.; Kakihara, F.; Kato, M. Production of useful vegetables having A and C genomes, using CMS B. oleracea. Estimation of ovule culture conditions for F1 hybrid production. Breed. Sci. 1993, 43 (Suppl. S2), 330. [Google Scholar]
- Williams, E.G.; Verry, I.M.; Williams, W.M. Use of embryo culture in interspecific hybridization. In Plant Improvement and Somatic Cell Genetics; Vasil, I.K., Scowcroft, W.R., Frey, K.J., Eds.; Academic Press: New York, NY, USA, 1982; pp. 119–128. [Google Scholar]
- Monnier, M. Survival of young immature Capsella embryos cultured in vitro. J. Plant Physiol. 1984, 115, 105–113. [Google Scholar] [CrossRef]
- Monnier, M. Culture of zygotic embryos. In In Vitro Embryogenesis in Plants; Thorpe, T.A., Ed.; Kluwer: Dordrecht, The Netherlands, 1995; pp. 117–153. [Google Scholar]
- Mòl, R.; Matthys-Rochon, E.; Dumas, C. Embryogeny and plant regeneration from maize zygotes by in vitro culture of fertilized embryo sacs. Plant Cell Rep. 1995, 14, 743–747. [Google Scholar] [CrossRef]
- Matthys-Rochon, E.; Piola, F.; Le Deunff, E.; Mòl, R.; Dumas, C. In vitro development of maize immature embryos: A tool for embryogenesis analysis. J. Exp. Bot. 1998, 49, 839–845. [Google Scholar] [CrossRef]
- Mòl, R.; Matthys-Rochon, E.; Dumas, C. In vitro culture of fertilized embryo sacs of maize: Zygotes and two celled proembryos can develop into plants. Planta 1993, 189, 213–217. [Google Scholar] [CrossRef]
- Chu, C.C.; Wang, C.C.; Sun, C.S. Establisment of an efficient medium for anther culture of rice through comparative experiments on the nitrogen sources. Sci. Sin. 1975, 18, 659–668. [Google Scholar]
- Podwyszyńska, M.; Marasek, A. Effect of thidiazuron and paclobutrazol on regeneration potential of flower stalk explants in vitro and subsequent shoot multiplication. Acta Soc. Bot. Pol. 2003, 72, 181–190. [Google Scholar] [CrossRef]
- Marasek-Ciolakowska, A.; Sochacki, D.; Marciniak, P. Breeding aspects of selected ornamental bulbous crops. Agronomy 2021, 11, 1709. [Google Scholar] [CrossRef]
- Custers, J.B.M.; Eikelboom, W.; Bergervoet, J.H.W.; van Eijk, J.P. In ovulo embryo culture of tulip (Tulipa L.); effects of culture conditions on seedling and bulblet formation. Sci. Hortic. 1992, 51, 111–122. [Google Scholar] [CrossRef]
- Kato, J.; Mii, M. Production of interspecific hybrids in ornamental plants. Methods Mol. Biol. 2012, 877, 233–245. [Google Scholar] [CrossRef]
- Mok, D.W.S.; Mok, M.C.; Rabakoarihanta, A.; Schii, C.T. Phaseolus wide hybridization through embryo culture. In Biotechnology in Agriculture and Forestry 2. Crops I; Bajaj, Y.P.S., Ed.; Springer: Berlin/Heidelberg, Germany, 1986; pp. 309–318. [Google Scholar]
- Sink, K.C.; Reynolds, F.J. Tomato (Lycopersicon esculentum L.). In Biotechnology in Agriculture and Forestry 2. Crops I; Bajaj, Y.P.S., Ed.; Springer: Berlin/Heidelberg, Germany, 1986; pp. 319–344. [Google Scholar]
- Jelanska, S. Cucurbits. In Biotechnology in Agriculture and Forestry 2. Crops I; Bajaj, Y.P.S., Ed.; Springer: Berlin/Heidelberg, Germany, 1986; pp. 371–386. [Google Scholar]
- Lentini, Z.; Restrepo, G.; Buitrago, M.E.; Tabares, E. Protocol for rescuing young cassava embryos. Front. Plant Sci. 2020, 11, 522. [Google Scholar] [CrossRef] [PubMed]
- Phillips, G.C.; Collins, G.B.; Taylor, N.L. Interspecific hybridization of red clover (Trifolium pratense L.) with T. sarosiense Hazsl. using in vitro embryo rescue. Theor. Appl. Genet. 1982, 62, 17–24. [Google Scholar] [CrossRef]
- Phillips, G.C.; Grosser, J.W.; Berger, S.; Taylor, N.L.; Collins, G.B. Interspecific hybridization between red clover and Trifolium alpestre using in vitro embryo rescue. Crop. Sci. 1992, 32, 1113–1115. [Google Scholar] [CrossRef]
- Roy, A.K.; Malaviya, D.R.; Kaushal, P. Generation of interspecific hybrids of Trifolium using embryo rescue techniques. Methods Mol. Biol. 2011, 710, 141–151. [Google Scholar] [CrossRef]
- Pellegrineschi, A.; Fatokun, C.A.; Thottappilly, G.; Adepoju, A.A. Cowpea embryo rescue. 1. Influence of culture media composition on plant recovery from isolated immature embryos. Plant Cell Rep. 1997, 17, 133–138. [Google Scholar] [CrossRef]
- Angelo, P.C.; Moraes, L.A.; Lopes, R.; Sousa, N.R.; da Cunha, R.N.; Quisen, R.C. In vitro rescue of interspecific embryos from Elaeis guineensis × E. oleifera (Arecaceae). Rev. Biol. Trop. 2011, 59, 1081–1088. [Google Scholar] [PubMed]
- Puangkrit, T.; Nontaswatsri, C. Intersubgeneric hybridization between Paracurcuma and Eucurcuma via embryo rescue. Acta Hortic. 2014, 1025, 37–42. [Google Scholar] [CrossRef]
- Nomura, Y.; Maeda, M.; Tsuchiya, T.; Makara, K. Efficient production of interspecific hybrids between Allium chinense and edible Allium spp. through ovary culture and pollen storage. Breed. Sci. 1994, 44, 151–155. [Google Scholar] [CrossRef]
- White, P.R. The Cultivation of Animal and Plant Cells, 2nd ed.; The Ronald Press Company: New York, NY, USA, 1963. [Google Scholar]
- Schenk, R.U.; Hildebrandt, A.C. Medium and techniques for induction and growth of monocotyledonous and dicotyledonous plant cell cultures. Can. J. Bot. 1972, 50, 199–204. [Google Scholar] [CrossRef]
- Lloyd, G.; McCown, B. Commercially feasible micropropagation of mountain laurel, Kalmia latifolia, by use of shoot-tip culture. Comb. Proc. Int. Plant Propagators Soc. 1980, 30, 421–427. [Google Scholar]
- Prasad, V.; Gupta, S.D. Applications and potentials of artificial neural networks in plant tissue culture. In Plant Tissue Culture Engineering; Springer: Dordrecht, The Netherlands, 2006; pp. 47–67. [Google Scholar] [CrossRef]
- Niedz, R.P.; Evens, T.J. Design of experiments (DOE)—History, concepts, and relevance to in vitro culture. Vitr. Cell Dev. Biol. Plant 2016, 52, 547–562. [Google Scholar] [CrossRef]
- Nezami-Alanagh, E.; Garoosi, G.A.; Landín, M.; Gallego, P.P. Combining DOE with neuro fuzzy logic for healthy mineral nutrition of pistachio rootstocks in vitro culture. Front. Plant Sci. 2018, 9, 1474. [Google Scholar] [CrossRef]
- Hameg, R.; Arteta, T.A.; Landin, M.; Gallego, P.P.; Barreal, M.E. Modeling and optimizing culture medium mineral composition for in vitro propagation of Actinidia arguta. Front. Plant Sci. 2020, 11, 554905. [Google Scholar] [CrossRef]
- Chandler, J.M.; Beard, B.H. Embryo culture of Helianthus hybrids. Crop. Sci. 1983, 23, 1004–1007. [Google Scholar] [CrossRef]
- Espinasse, A.; Lay, C.; Dybing, C.D. Factor controlling in vitro development of sunflower embryo. Agronomy 1985, 5, 825–832. [Google Scholar] [CrossRef]
- Krauter, R.; Friedt, W. Efficient interspecific hybridization in the genus Helianthus for genetic amelioration of sunflower. In Proceedings of the 12th EUCARPIA Congress, Part II, Gottingen, Germany, 27 February–4 March 1989; pp. 12–14. [Google Scholar]
- Bohorova, N.E.; Atanassov, A.; Georgieva-Todorova, J. In vitro organogenesis, androgenesis and embryogenesis in the genus Helianthus. Z. Pflanzenzücht. 1985, 95, 34–44. [Google Scholar]
- Krauter, R.; Steinmetz, A.; Friedt, W. Efficient interspecific hybridization in the genus Helianthus via “embryo rescue” and characterization of the hybrids. Theor. Appl. Genet. 1991, 82, 521–525. [Google Scholar] [CrossRef] [PubMed]
- Dahlhoff, M.; Kohler, H.; Nichterlein, K.; Friedt, W. Production of interspecific hybrids in the genus Helianthus by embryo-rescue and characterization of hybrids. In Genetic Manipulation in Plant Breeding; EUCARPIA Simposium: Terragona, Spain, 1991. [Google Scholar]
- Muraida, N.; Marubashi, W. Characterization of hybrid seedlings from crosses of Nicotiana stocktonii Brandegee × N. tabacum L. and N. stocktonii × progenitors of N. tabacum. Plant Biotechnol. 2015, 32, 139–147. [Google Scholar] [CrossRef]
- Subhashini, U.; Venkateswarlu, T.; Anjani, K.; Prasad, G.S. In vitro hybridization in an incompatible cross Nicotiana glutinosa × Nicotiana megalosiphon. Theor. Appl Genet. 1985, 71, 545–549. [Google Scholar] [CrossRef]
- Deng, Y.M.; Chen, S.M.; Lu, A.M.; Chen, F.D.; Tang, F.P.; Guan, Z.Y.; Teng, N.J. Production and characterisation of the intergeneric hybrids between Dendranthema morifolium and Artemisia vulgaris exhibiting enhanced resistance to chrysanthemum aphid (Macrosiphoniella sanbourni). Planta 2010, 231, 693–703. [Google Scholar] [CrossRef] [PubMed]
- Cheng, X.; Chen, S.; Chen, F.; Deng, Y.; Fang, W.; Tang, F.; Liu, Z.; Shao, W. Creating novel chrysanthemum germplasm via interspecific hybridization and backcrossing. Euphytica 2011, 177, 45–53. [Google Scholar] [CrossRef]
- Sun, C.Q.; Chen, F.D.; Teng, N.J.; Liu, Z.L.; Fang, W.M.; Hou, X.L. Interspecific hybrids between Chrysanthemum grandiflorum (Ramat.) Kitamura and C. indicum (L.) Des Moul. and their drought tolerance evaluation. Euphytica 2010, 174, 51–60. [Google Scholar] [CrossRef]
- Zhu, W.Y.; Jiang, J.F.; Chen, S.M.; Wang, L.; Xu, L.L.; Wang, H.B.; Li, P.L.; Guan, Z.Y.; Chen, F.D. Intergeneric hybrid between Chrysanthemum × morifolium and Artemisia japonica achieved via embryo rescue shows salt tolerance. Euphytica 2013, 191, 109–119. [Google Scholar] [CrossRef]
- Tang, F.; Chen, F.; Chen, S.; Teng, N.; Fang, W. Intergeneric hybridization and relationship of genera within the tribe Anthemideae Cass. (I. Dendranthema crissum (kitam.) kitam. × Crossostephium chinense (L.) Makino). Euphytica 2009, 169, 133–140. [Google Scholar] [CrossRef]
- Röper, A.C.; Lütken, H.; Hegelund, J.N.; Petersen, K.K.; Christensen, B.; Müller, R. Effect of different ovule isolation times on the embryo development of Campanula hybrids. Acta Hortic. 2012, 953, 161–166. [Google Scholar] [CrossRef]
- Takamura, Y.; Asano, C.; Hikage, T.; Hatakeyama, K.; Takahata, Y. Production of interspecific hybrids between Japanese gentians and wild species of Gentiana. Breed. Sci. 2019, 69, 680–687. [Google Scholar] [CrossRef]
- Nishimoto, S.I.; Shimizu, K.; Hashimoto, F.; Sakata, Y. Interspecific hybrids of Camellia chrysantha × C. japonica by ovule culture. J. Jpn. Soc. Hortic. Sci. 2003, 72, 236–242. [Google Scholar] [CrossRef]
- Nishimoto, S.-I.; Hashimoto, F.; Shimizu, K.; Sakata, Y. Petal coloration of interspecific hybrids between Camellia chrysantha × C. japonica. J. Jpn. Soc. Hortic. Sci. 2004, 73, 189–191. [Google Scholar] [CrossRef][Green Version]
- Kishi, F.; Kagami, Y.; Shinohara, M.; Hatano, S.; Tsurushima, H. Production of interspecific hybrid in Gypsophila by ovule-embryo culture. Euphytica 1994, 74, 85–90. [Google Scholar] [CrossRef]
- Nagaharu, U.; Nagaharu, N. Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jap. J. Bot. 1935, 7, 389–452. [Google Scholar]
- Nishi, S.; Kawada, J.; Toda, M. Studies on the embryo culture in vegetable crops. Part 2. Breeding of interspecific hybrids between cabbage varieties and Chinese cabbage varieties through the application of embryo culture techniques. Bull. Hort. Res. Sta. Jpn. Ser. 1962, A1, 111–156. [Google Scholar]
- Nishi, S.; Toda, M.; Toyoda, T. Studies on the embryos culture in vegetable crops. III. On the conditions affecting to embryo culture of interspecific hybrids between cabbage and Chinese cabbage. Bull. Hort. Res. Sta. Jpn. Ser. 1970, A9, 73–88. [Google Scholar]
- Inomata, N. Production of interspecific hybrids between Brassica campestris and Brassica oleracea by culture in vitro of excised ovaries. I. Effects of yeast extract and casein hydrolysate on the development of excised ovaries. Jpn. J. Breed. 1977, 27, 295–304. [Google Scholar] [CrossRef]
- Inomata, N. Production of interspecific hybrids between Brassica campestris and B. oleracea by culture in vitro of excised ovaries. II. Development of excised ovaries in various culture media with coconut milk and casein hydrolysate. Jpn. J. Genet. 1978, 53, 1–11. [Google Scholar] [CrossRef]
- Inomata, N. Production of interspecific hybrids between Brassica campestris and B. oleracea by culture in vitro of excised ovaries. III. Development of excised ovaries in the crosses of various cultivars. Jpn. J. Genet. 1978, 53, 161–173. [Google Scholar] [CrossRef]
- Matsuzawa, Y. Studies on the interspecific and intergeneric crossability in Brassica and Raphanus. Spec. Bull. Coll. Agric. Utsunomiya Univ. 1983, 39, 1–86. [Google Scholar]
- Bang, S.W.; Sugihara, K.; Jeong, B.H.; Kaneko, R.; Satake, E.; Kaneko, Y.; Matsuzawa, Y. Production and characterization of intergeneric hybrids between Brassica oleracea and a wild relative Moricandia arvensis. Plant Breed. 2007, 126, 101–103. [Google Scholar] [CrossRef]
- Tonosaki, K.; Michiba, K.; Bang, S.W.; Kitashiba, H.; Kaneko, Y.; Nishio, T. Genetic analysis of hybrid seed formation ability of Brassica rapa in intergeneric crossings with Raphanus sativus. Theor. Appl. Genet. 2013, 126, 837–846. [Google Scholar] [CrossRef]
- Gundimeda, H.R.; Prakash, S.; Shivanna, K.R. Intergeneric hybrids between Enarthrocarpus lyratus, a wild species, and crop brassicas. Theor. Appl. Genet. 1992, 83, 655–662. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Farooq, Z.; Chu, L.; Liu, J.; Wang, H.; Guo, J.; Ma, C.; Dai, C.; Wen, J.; Shen, J.; et al. High-generation near-isogenic lines combined with multi-omics to study the mechanism of polima cytoplasmic male sterility. BMC Plant Biol. 2021, 21, 130. [Google Scholar] [CrossRef]
- Formanová, N.; Li, X.Q.; Ferrie, A.M.; Depauw, M.; Keller, W.A.; Landry, B.; Brown, G.G. Towards positional cloning in Brassica napus: Generation and analysis of doubled haploid B. rapa possessing the B. napus pol CMS and Rfp nuclear restorer gene. Plant Mol. Biol. 2006, 61, 269–281. [Google Scholar] [CrossRef]
- Jugulam, M.; Ziauddin, A.; So, K.K.; Chen, S.; Hall, J.C. Transfer of dicamba tolerance from Sinapis arvensis to Brassica napus via embryo rescue and recurrent backcross breeding. PLoS ONE 2015, 10, e0141418. [Google Scholar] [CrossRef]
- Mithila, J.; Hall, J.C. Transfer of auxinic herbicide resistance from Brassica kaber to Brassica juncea and Brassica rapa through embryo rescue. Vitr. Cell Dev. Biol. Plant 2013, 49, 461–467. [Google Scholar] [CrossRef]
- Pen, S.; Kumar Nath, U.; Song, S.; Goswany, G.; Lee, H.-J.; Jung, H.-J.; Kim, H.-T.; Park, J.-I.; Nou, S.-I. Developmental stage and shape of embryo determine the efficacy of embryo rescue in introgressing orange/yellow color and anthocyanin genes of Brassica species. Plants 2018, 7, 99. [Google Scholar] [CrossRef]
- Ton, L.B.; Neik, T.X.; Batley, T. The use of genetic and gene technologies in shaping modern rapeseed cultivars (Brassica napus L.). Genes 2020, 11, 1161. [Google Scholar] [CrossRef] [PubMed]
- Bayer, P.E.; Golicz, A.A.; Tirnaz, S.; Chan, C.-K.K.; Edwards, D.; Batley, J. Variation in abundance of predicted resistance genes in the Brassica oleracea pangenome. Plant Biotechnol. J. 2019, 17, 789–800. [Google Scholar] [CrossRef]
- Zhan, Z.; Shah, N.; Jia, R.; Li, X.; Zhang, C.; Piao, Z. Transferring of clubroot-resistant locus CRd from Chinese cabbage (Brassica rapa) to canola (Brassica napus) through interspecific hybridization. Breed. Sci. 2022, 72, 189–197. [Google Scholar] [CrossRef] [PubMed]
- Gong, Q.; Dai, C.-Y.; Zhang, X.-H.; Wang, X.-L.; Huang, Z.; Xu, A.-X.; Dong, J.-G.; Yu, C.-Y. Towards breeding of rapeseed (Brassica napus) with alien cytoplasm and powdery mildew resistance from Ethiopian mustard (Brassica carinata). Breed. Sci. 2020, 70, 387–395. [Google Scholar] [CrossRef] [PubMed]
- Gaebelein, R.; Alnajar, D.; Koopmann, B.; Mason, A.S. Hybrids between Brassica napus and B. nigra show frequent pairing between the B and A/C genomes and resistance to blackleg. Chromosome Res. 2019, 27, 221–236. [Google Scholar] [CrossRef]
- Liu, Y.; Xu, A.; Liang, F.; Yao, X.; Wang, Y.; Liu, X.; Zhang, Y.; Dalelhan, J.; Zhang, B.; Qin, M.; et al. Screening of clubroot-resistant varieties and transfer of clubroot resistance genes to Brassica napus using distant hybridization. Breed. Sci. 2018, 68, 258–267. [Google Scholar] [CrossRef]
- Vicente, J.G.; Holub, E.B. Xanthomonas campestris pv. Campestris (cause of black rot of crucifers) in the genomic era is still a worldwide threat to Brassica crops. Mol. Plant Pathol. 2013, 14, 2–18. [Google Scholar] [CrossRef]
- Pan, X.; Caldwell, C.D.; Falk, K.C.; Lada, R. The effect of cultivar, seeding rate and applied nitrogen on Brassica carinata seed yield and quality in contrasting environments. Can. J. Plant Sci. 2012, 92, 961–971. [Google Scholar] [CrossRef]
- Enjalbert, J.N.; Zheng, S.; Johnson, J.J.; Mullen, J.L.; Byrne, P.F.; McKay, J.K. Brassicaceae germplasm diversity for agronomic and seed quality traits under drought stress. Ind. Crop. Prod. 2013, 47, 176–185. [Google Scholar] [CrossRef]
- Sharma, B.B.; Kalia, P.; Singh, D.; Sharma, T.R. Introgression of black rot resistance from Brassica carinata to cauliflower (Brassica oleracea botrytis group) through embryo rescue. Front. Plant Sci. 2017, 8, 1255. [Google Scholar] [CrossRef]
- Ayotte, R.; Harney, P.M.; Souza, M.V. The transfer of atrazine resistance from Brassica napus L. to B. oleracea L. Production of F1 hybrids through embryo rescues. Euphytica 1987, 36, 615–624. [Google Scholar] [CrossRef]
- Weerakoon, S.R. Producing inter-specific hybrids between Brassica juncea (L.) Czern & Coss and B. oleracea (L.) to synthesize trigenomic (ABC) Brassica. J. Sci. Univ. Kelaniya 2011, 6, 13–34. [Google Scholar] [CrossRef]
- Tonguc, M.; Griffiths, P.D. Transfer of powdery mildew resistance from Brassica carinata to Brassica oleracea through embryo rescue. Plant Breed. 2004, 123, 587–589. [Google Scholar] [CrossRef]
- Chiang, M.S.; Chiang, B.Y.; Grant, W.F. Transfer of resistance to race 2 of Plasmodiophora brassicae from Brassica napus to cabbage (B. oleracea var. capitata). I. Interspecific hybridization between B. napus and B. oleracea var. capitata. Euphytica 1977, 26, 319–326. [Google Scholar] [CrossRef]
- Tonguc, M.; Griffiths, P.D. Development of black rot resistant interspecific hybrids between Brassica oleracea L. cultivars and Brassica accession A 19182, using embryo rescue. Euphytica 2004, 136, 313–318. [Google Scholar] [CrossRef]
- Chiang, M.S.; Crete, R. Cytoplasmic male sterility in Brassica oleracea induced by B. napus cytoplasm-female fertility and restoration of male fertility. Can. J. Plant Sci. 1987, 672, 891–897. [Google Scholar] [CrossRef]
- Bathia, R.; Dey, S.S.; Sharma, K.; Singh, S.; Kumar, S.; Pramarik, A.; Parkash, C.; Kumar, R. Back-cross introgression of ‘Tour’ cytoplasm from Brassica napus through in vitro embryo rescue reveals partial restoration of sterility in B. oleracea. Sci. Horticul. 2021, 282, 110014. [Google Scholar] [CrossRef]
- Kumar, R.; Chowdhury, J.B.; Jain, R.K. Interspecific hybridization in Brassica juncea and Brassica tournefortii through embryo rescue and their evaluation for biotic and abiotic stress tolerance. Indian J. Exp. Biol. 2001, 39, 911–915. [Google Scholar] [PubMed]
- Nuñez-Palenius, H.G.; Gomez-Lim, M.; Ochoa-Alejo, N.; Grumet, R.; Lester, G.; Cantliffe, D.J. Melon fruits: Genetic diversity, physiology, and biotechnology features. Crit. Rev. Biotechnol. 2008, 28, 13–55. [Google Scholar] [CrossRef]
- Nuñez-Palenius, H.G.; Ramírez-Malagón, R.; Ochoa-Alejo, N. Muskmelon embryo rescue techniques using in vitro embryo culture. Methods Mol. Biol. 2011, 710, 107–115. [Google Scholar] [CrossRef]
- Norton, J.D. Embryo culture of Cucumis species. HortScience 1981, 16, 433. [Google Scholar]
- Fassuliotis, G.; Nelson, B.V. Interspecific hybrids of Cucumis metuliferus × C. anguria obtained through embryo culture and somatic embryogenesis. Euphytica 1988, 37, 53–60. [Google Scholar] [CrossRef]
- Dabauza, M.; Gonzalez-Candelas, L.; Bordas, M.; Roig, L.A.; Ramon, D.; Moreno, V. Regeneration and characterization of Cucumis melo L. (+) Cucumis anguria L. var. longipes (Hook. fil.) Meeuse somatic hybrids. Plant Cell Tissue Organ Cult. 1998, 52, 123–131. [Google Scholar] [CrossRef]
- Van Der Maesen, L.J.G.; Pundir, R.P.S. Availability and use of wild Cicer germplasm. Plant Genet. Newsl. 1984, 57, 19–24. [Google Scholar]
- Ahmad, F.; Slinkard, A. The extent of embryo and endosperm growth following interspecific hybridization between Cicer arietinum L. and related annual wild species. Genet. Resour. Crop. Evol. 2004, 51, 765–772. [Google Scholar] [CrossRef]
- Singh, R.; Sharma, P.; Varshney, R.K.; Sharma, S.K.; Singh, N.K. Chickpea improvement: Role of wild species and genetic markers. Biotechnol. Genet. Eng. Rev. 2008, 25, 267–314. [Google Scholar] [CrossRef]
- Singh, R.K.; Singh, C.; Ambika; Chandana, B.S.; Mahto, R.K.; Patial, R.; Gupta, A.; Gahlaut, V.; Gayacharan; Hamwieh, A.; et al. Exploring chickpea germplasm diversity for broadening the genetic base utilizing genomic resourses. Front. Genet. 2022, 13, 905771. [Google Scholar] [CrossRef] [PubMed]
- Mallikarjuna, N.; Muehlbauer, F.J. Chickpea hybridization using in vitro techniques. Methods Mol. Biol. 2011, 710, 93–105. [Google Scholar] [CrossRef]
- Mercy, S.T.; Kakar, S.N. Barrier to interspecific crosses in Cicer. Proc. Ind. Natl. Sci. Acad. 1975, 41, 78–82. [Google Scholar]
- Badami, P.S.; Mallikarjuna, M.; Moss, J.P. Interspecific hybridization between Cicer arietinum and C. pinnatifidum. Plant Breed. 1997, 116, 393–395. [Google Scholar] [CrossRef]
- Bassiri, A.; Ahmad, F.; Slinkard, A.E. Pollen grain germination and pollen tube growth following in vivo and in vitro self and interspecific pollinations in annual Cicer species. Euphytica 1987, 36, 667–675. [Google Scholar] [CrossRef]
- Ahmad, F.; Slinkard, A.E.; Scoles, G.J. Investigations into the barrier(s) to interspecific hybridization between Cicer arietinum L. and eight other annual Cicer species. Plant Breed. 1988, 100, 193–198. [Google Scholar] [CrossRef]
- Singh, K.B.; Ocampo, B. Interspecific hybridization in annual Cicer species. J. Gen. Breed. 1993, 47, 199–204. [Google Scholar]
- Singh, K.B.; Ocampo, B. Exploitation of wild Cicer species for yield improvement in chickpea. Theor. Appl. Genet. 1997, 95, 418–423. [Google Scholar] [CrossRef]
- Geerts, P.; Toussaint, A.; Mergeai, G.; Baudoin, J.P. Phaseolus immature embryo rescue technology. Methods Mol. Biol. 2011, 710, 117–129. [Google Scholar] [CrossRef]
- Manzur, J.P.; Fita, A.; Prohens, J.; Rodríguez-Burruezo, A. Successful wide hybridization and introgression breeding in a diverse set of common peppers (Capsicum annuum) using different cultivated ají (C. baccatum) accessions as donor parents. PLoS ONE 2015, 10, e0144142. [Google Scholar] [CrossRef] [PubMed]
- Smith, P.G. Embryo culture of a tomato species hybrid. Proc. Am. Hort. Sci. 1944, 44, 413–416. [Google Scholar]
- Moore, G.A.; Litz, R.E. Biochemical markers for Carica papaya, C. cauliflora and plants from somatic embryos of their hybrid. J. Am. Soc. Hort. Sci. 1984, 109, 213–218. [Google Scholar] [CrossRef]
- Chung, G.; Singh, R.J. Broadening the genetic base of soybean: A multidisciplinary approach. Crit. Rev. Plant Sci. 2008, 27, 295–341. [Google Scholar] [CrossRef]
- Broué, P.; Douglass, J.; Grace, J.P.; Marshall, D.R. Interspecific hybridisation of soybeans and perennial Glycine species indigenous to Australia via embryo culture. Euphytica 1982, 31, 715–724. [Google Scholar] [CrossRef]
- Bodanese-Zanettini, M.H.; Lauxen, M.S.; Richter, S.N.C.; Cavalli-Molina, S.; Lange, C.E.; Wang, P.J.; Hu, C.Y. Wide hybridization between Brazilian soybean cultivars and wild perennial relatives. Theor. Appl. Genet. 1996, 93, 703–709. [Google Scholar] [CrossRef]
- Chung, G.H.; Kim, J.H. Production of interspecific hybrids between Glycine max and G. tomentella through embryo culture. Euphytica 1990, 48, 97–101. [Google Scholar] [CrossRef]
- Chung, G.H.; Kim, K.S. Obtaining intersubgeneric hybridization between Glycine max and G. latifolia through embryo culture. Korean J. Plant Tissue Cult. 1991, 18, 39–45. [Google Scholar]
- Newell, C.A.; Hymowitz, T. Successful wide hybridization between the soybean and a wild perennial relative, Glycine tomentella Hayata. Crop. Sci. 1982, 22, 1062–1065. [Google Scholar] [CrossRef]
- Newell, C.A.; Delannay, X.; Edge, M.E. Interspecific hybrids between the soybean and wild perennial relatives. J. Hered. 1987, 78, 301–306. [Google Scholar] [CrossRef]
- Schoen, D.J.; Burdon, J.J.; Brown, A.H.D. Resistance of Glycine tomentella to soybean leaf rust Phakopsora pachyrhizi in relation to ploidy level and geographical distribution. Theor. Appl. Genet. 1992, 83, 827–832. [Google Scholar] [CrossRef]
- Hartman, G.L.; Wang, T.C.; Hymowitz, T. Sources of resistance to soybean rust in perennial Glycine species. Plant Dis. 1992, 76, 396–399. [Google Scholar] [CrossRef]
- Shoemaker, R.C.; Heath, M.S.; Skorupksa, H.; Delannay, X.; Edge, M.; Newell, C.A. Fertile progeny of a hybridization between soybean [Glycine max (L.) Merr.] and G. tomentella Hayata. Theor. Appl. Genet. 1990, 80, 17–23. [Google Scholar] [CrossRef] [PubMed]
- Bauer, S.; Hymowitz, T.; Noel, G.R. Soybean cyst nematode resistance derived from Glycine tomentella in amphiploid (G. max × G. tomentella) hybrid lines. Nematropica 2007, 37, 277–285. [Google Scholar]
- Lim, S.M.; Hymowitz, T. Reactions of perennial wild species of genus Glycine to Septoria glycines. Plant Dis. 1987, 71, 891–893. [Google Scholar] [CrossRef]
- Horlock, C.M.; Teakle, D.S.; Jones, R.M. Natural infection of the native pasture legumes, Glycine latifolia, by alfalfa mosaic virus in Queensland. Aust. Plant Pathol. 1997, 26, 115–116. [Google Scholar] [CrossRef]
- Zheng, C.; Chen, P.; Hymowitz, T.; Wickizer, S.; Gergerich, R. Evaluation of Glycine species for resistance to Bean pod mottle virus. Crop. Prot. 2005, 24, 49–56. [Google Scholar] [CrossRef]
- Chang, S. Identification of Loci for Resistance to Sclerotinia Stem Rot (White Mold) in Accessions of Perennial Relative of Soybean (Glycine latifolia). Ph.D. Thesis, University of Illinois, Urbana, IL, USA, 2014. [Google Scholar]
- Hart, S.E.; Glenn, S.; Kenworthy, W.W. Tolerance and the basisfor selectivity to 2,4-D in perennial Glycine species. Weed Sci. 1991, 39, 535–539. [Google Scholar] [CrossRef]
- Pantalone, V.R.; Kenworthy, W.J.; Slaughter, L.H.; James, B.R. Chloride tolerance in soybean and perennial Glycine accessions. Euphytica 1997, 97, 235–239. [Google Scholar] [CrossRef]
- Singh, R.J.; Nelson, R.L. Intersubgeneric hybridization between Glycine max and G. tomentella: Production of F1, amphidiploid, BC1, BC2, BC3, and fertile soybean plants. Theor. Appl. Genet. 2015, 128, 1117–1136. [Google Scholar] [CrossRef]
- Polesani, M.; Bortesi, L.; Ferrarini, A.; Zamboni, A.; Fasoli, M.; Zadra, C.; Lovato, A.; Pezzotti, M.; Delledonne, M.; Polverari, A. General and species-specific transcriptional responses to downy mildew infection in a susceptible (Vitis vinifera) and a resistant (V. riparia) grapevine species. BMC Genom. 2010, 11, 117. [Google Scholar] [CrossRef]
- Li, G.R.; Ji, W.; Wang, G.; Zhang, J.X.; Wang, Y.J. An improved embryo-rescue protocol for hybrid progeny from seedless Vitis vinifera grapes × wild Chinese Vitis species. Vitr. Cell Dev. Biol. Plant 2014, 50, 110–120. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Li, Z.; Zhao, Y.; Zhao, J.; Luo, Q.; Wang, Y. New disease-resistant, seedless grapes are developed using embryo rescue and molecular markers. 3 Biotech 2020, 10, 4. [Google Scholar] [CrossRef] [PubMed]
- Giancaspro, A.; Mazzeo, A.; Carlomagno, A.; Gadaleta, A.; Somma, S.; Ferrara, G. Optimization of an in vitro embryo rescue protocol for breeding seedless table grapes (Vitis vinifera L.) in Italy. Horticulturae 2022, 8, 121. [Google Scholar] [CrossRef]
- Emershad, R.L.; Ramming, D.W. In-ovulo embryo culture of Vitis vinifera L. cv. ‘Thompson Seedless’. Hort. Sci. 1982, 17, 576. [Google Scholar]
- Wang, J.; Luo, M.-C.; Chen, Z.; You, F.M.; Wei, Y.; Zheng, Y.; Dvorak, J. Aegilops tauschii single nucleotide polymorphisms shed light on the origins of wheat D-genome genetic diversity and pinpoint the geographic origin of hexaploid wheat. New Phytol. 2013, 198, 925–937. [Google Scholar] [CrossRef] [PubMed]
- Mirzaghaderi, G.; Abdolmalaki, Z.; Ebrahimzadegan, R.; Bahmani, F.; Orooji, F.; Majdi, M.; Mozafari, A.A. Production of synthetic wheat lines to exploit the genetic diversity of emmer wheat and D genome containing Aegilops species in wheat breeding. Sci. Rep. 2020, 10, 19698. [Google Scholar] [CrossRef] [PubMed]
- Mirzaghaderi, G.; Mason, A.S. Broadening the bread wheat D genome. Theor. Appl. Genet. 2019, 132, 1295–1307. [Google Scholar] [CrossRef]
- Mujeeb-Kazi, A.; Rosas, V.; Roldan, S. Conservation of the genetic variation of Triticum tauschii (Coss.) Schmalh. (Aegilops squarrosa auct. non L.) in synthetic hexaploid wheats (T. turgidum L. s.lat. × T. tauschii; 2n = 6x = 42, AABBDD) and its potential utilization for wheat improvement. Gen. Resour. Crop. Evol. 1996, 43, 129–134. [Google Scholar] [CrossRef]
- Franke, R.; Meinel, A. History of the first fertile amphidiploid wheat × rye hybrid W. Rimpau’s triticale. Cereal Res. Commun. 1990, 18, 103–109. Available online: https://www.jstor.org/stable/23783083 (accessed on 2 May 2023).
- Crespo-Herrera, L.A.; Garkava-Gustavsson, L.; Åhman, I. A systematic review of rye (Secale cereale L.) as a source of resistance to pathogens and pests in wheat (Triticum aestivum L.). Hereditas 2017, 154, 14. [Google Scholar] [CrossRef]
- An, D.; Zheng, Q.; Luo, Q.; Ma, P.; Zhang, H.; Li, L.; Ham, F.; Xu, H.; Xu, Y.; Zhang, X.; et al. Molecular cytogenetic identification of a new wheat-rye 6R chromosome disomic addition line with powdery mildew resistance. PLoS ONE 2015, 10, e0134534. [Google Scholar] [CrossRef]
- Han, G.; Yan, H.; Wang, J.; Cao, L.; Liu, S.; Li, X.; Zhou, Y.; Fan, J.; Li, L.; An, D. Molecular cytogenetic identification of a new wheat-rye 6R addition line and physical localization of its powdery mildew resistance gene. Front. Plant Sci. 2022, 13, 889494. [Google Scholar] [CrossRef]
- Mundt, C.C. Pyramiding for resistance durability: Theory and practice. Phytopathology 2018, 108, 792–802. [Google Scholar] [CrossRef]
- An, D.; Ma, P.; Zheng, Q.; Fu, S.; Li, L.; Han, F.; Han, G.; Wang, J.; Xu, Y.; Jin, Y.; et al. Development and molecular cytogenetic identification of a new wheat-rye 4R chromosome disomic addition line with resistances to powdery mildew, stripe rust and sharp eyespot. Theor. Appl. Genet. 2019, 132, 257–272. [Google Scholar] [CrossRef]
- Lapitan, N.L.V.; Sears, R.G.; Gill, B.S. Translocations and other karyotypic structural changes in wheat × rye hybrids regenerated from tissue culture. Theor. Appl. Genet. 1984, 68, 547–554. [Google Scholar] [CrossRef]
- D’Amato, F. Cytogenetics of differentiation in tissue and cell cultures. In Applied and Fundamental Aspects of Plant Cell, Tissue and Organ Culture; Reinert, J., Bajaj, Y.P.S., Eds.; Springer: Berlin/Heidelberg, Germany; New York, NY, USA, 1977; pp. 343–357. [Google Scholar]
- Konzak, C.F.; Reynolds, L.F.; Jensen, N.F. Embryo culture of barley species hybrids. Cytological studies of Hordeum sativum × Hordeum bulbosum. J. Hered. 1951, 42, 125–134. [Google Scholar] [CrossRef]
- Xu, J.; Kasha, K.J. Transfer of a dominant gene for powdery mildew resistance and DNA from Hordeum bulbosum into cultivated barley (H. vulgare). Theor. Appl. Genet. 1992, 84, 771–777. [Google Scholar] [CrossRef]
- Encina, C.L.; Regalado, J.J. Aspects of in vitro plant tissue culture and breeding of asparagus: A Review. Horticulturae 2022, 8, 439. [Google Scholar] [CrossRef]
- Marcellán, O.N.; Camadro, E.L. Preliminary results on embryo rescue for circumventing hybridization barriers in Asparagus. Biocell 2000, 24, 247–251. [Google Scholar]
- Corpas-Hervias, C.; Melero-Vara, J.M.; Molinero-Ruiz, M.L.; Zurera-Muñoz, C.; Basallote-Ureba, M.J. Characterization of isolates of Fusarium spp. Obtained from asparagus in Spain. Plant Dis. 2006, 90, 1441–1451. [Google Scholar] [CrossRef]
- Elmer, W.H. The economically important diseases of Asparagus in the United States. Plant Health Prog. 2001, 2, 13. [Google Scholar] [CrossRef]
- Bridgen, M.; Kollman, E.; Lu, C. Interspecific hybridization of Alstroemeria for the development of new, ornamentals. Acta Hortic. 2009, 836, 73–78. [Google Scholar] [CrossRef]
- Aros, D.; Suazo, M.; Rivas, C.; Zapata, P.; Úbeda, C.; Bridgen, M. Molecular and morphological characterization of new interspecific hybrids of alstroemeria originated from A. caryophylleae scented lines. Euphytica 2019, 215, 93. [Google Scholar] [CrossRef]
- Morgan, E.; Burge, G.; Seelye, J.; Hopping, M.E.; Grant, J.E.; Warren, A.G.F.; Brundell, D. Wide crosses in the Colchicaceae: Sandersonia aurantiaca (Hook.) × Littonia modesta (Hook.). Euphytica 2001, 121, 343–348. [Google Scholar] [CrossRef]
- Kao, K.N.; Michayluk, M.R. Nutritional requirements for growth of Vicia hastajana cells and protoplasts at a very low population density in liquid media. Planta 1975, 126, 105–110. [Google Scholar] [CrossRef]
- Amano, J.; Nakazawa, D.; Kuwuyama, S.; Mizuta, Y.; Okuno, H.; Watanabe, Y.; Godo, T.; Han, D.-S.; Nakano, M. Intergeneric hybridization among colchicaceous ornamentals, Gloriosa spp., Littonia modesta and Sandersonia aurantiaca via ovule culture. Plant Biotechnol. 2009, 26, 535–541. [Google Scholar] [CrossRef]
- Nakamura, T.; Kuwayama, S.; Tanaka, S.; Oomiya, T.; Saito, H.; Nakano, M. Production of intergeneric hybrid plants between Sandersonia aurantiaca and Gloriosa rothschildiana via ovule culture (Colchicaceae). Euphytica 2005, 142, 283–289. [Google Scholar] [CrossRef]
- Jauhar, P.P. Spontaneous haploids in durum wheat: Their cytogenetic characterization. Euphytica 2006, 148, 341–344. [Google Scholar] [CrossRef]
- D’Amato, F.; Baroncelli, S.; Durante, M. Genetica Vegetale; Bollati Boringhieri Editore s.p.a.: Torino, Italy, 1987; ISBN 88-339-5409-9. [Google Scholar]
- Guha, S.; Maheshwari, S. In vitro production of embryos from anthers of Datura. Nature 1964, 204, 497. [Google Scholar] [CrossRef]
- Portemer, V.; Renne, C.; Guillebaux, A.; Mercier, R. Large genetic screens for gynogenesis and androgenesis haploid inducers in Arabidopsis thaliana failed to identify mutants. Front. Plant Sci. 2015, 6, 147. [Google Scholar] [CrossRef]
- Ren, J.; Wu, P.; Trampe, B.; Tian, X.; Lübberstedt, T.; Chen, S. Novel technologies in doubled haploid line development. Plant Biotechnol. J. 2017, 15, 1361–1370. [Google Scholar] [CrossRef] [PubMed]
- Hale, B.; Ferrie, A.M.R.; Chellamma, S.; Samuel, J.P.; Phillips, G.C. Androgenesis-based doubled haploidy: Past, present, and future perspectives. Front. Plant Sci. 2022, 12, 751230. [Google Scholar] [CrossRef] [PubMed]
- San Noeum, L.H. Haploides d’Hordeum vulgare L. par culture in vitro d’ovaries non fecondes. Ann. Amelior. Plant. 1976, 26, 751–754. [Google Scholar]
- Joergensen, C.A. The experimental formation of heteroploidy plants in the genus Solanum. J. Genet. 1928, 19, 133–211. [Google Scholar] [CrossRef]
- Kimber, G.; Riley, R. Haploid angiosperm. Bot. Rev. 1963, 29, 480–531. Available online: https://www.jstor.org/stable/4353678 (accessed on 2 May 2023). [CrossRef]
- Kihara, H. Formation of haploids by means of delayed pollination in Triticum monococcum. Bot. Mag. Tokyo 1940, 54, 178–185. [Google Scholar] [CrossRef][Green Version]
- Gerassimova, H. Experimentally produced haploid plant in Crepis tectorum. Biol. J. 1936, 5, 895–900. [Google Scholar]
- Goodspeed, T.H.; Avery, P. The occurrence of a Nicotiana glutinosa haplont. Proc. Natl. Acad. Sci. USA 1929, 15, 502–504. [Google Scholar] [CrossRef] [PubMed]
- Kihara, H.; Tsunewaki, K. Use of alien cytoplasm as a new method of producing haploid. Jpn. J. Genet. 1962, 37, 310–313. [Google Scholar] [CrossRef]
- Huskins, C.L. The subdivision of the chromosomes and their multiplication in non-dividing tissues: Possible interpretations in terms of gene structure and gene action. Am. Nat. 1947, 81, 401–434. [Google Scholar] [CrossRef]
- Simantel, G.M.; Ross, J.G.; Huang, C.C.; Haensel, H.D. Colchicine-induced somatic reduction in Sorghum: II. Induction of plants homozygous for structural chromosome markers originally heterozygous in two pairs. J. Hered. 1963, 54, 221–228. [Google Scholar] [CrossRef] [PubMed]
- Lima de Faria, A.; Jaworska, H. Haplo-diploid chimaeras in Haplopappus gracilis. Hereditas 1964, 52, 119–122. [Google Scholar]
- Kalinowska, K.; Chamas, S.; Unkel, K.; Demidov, D.; Lermontova, I.; Dresselhaus, T.; Kumlehn, J.; Dunemann, F.; Houben, A. State-of-the-art and novel developments of in vivo haploid technologies. Theor. Appl. Genet. 2019, 132, 593–605. [Google Scholar] [CrossRef]
- Conner, J.A.; Mookkan, M.; Huo, H.; Ozias-Akins, P. A parthenogenesis gene of apomict origin elicits embryo formation from unfertilized eggs in a sexual plant. Proc. Natl. Acad. Sci. USA 2015, 112, 11205–11210. [Google Scholar] [CrossRef]
- Conner, J.A.; Podio, M.; Ozias-Akins, P. Haploid embryo production in rice and maize induced by PsASGR-BBML transgenes. Plant Reprod. 2017, 30, 41–52. [Google Scholar] [CrossRef]
- Zhang, Z.; Conner, J.; Guo, Y.; Ozias-Akins, P. Haploidy in tobacco induced by PsASGR-BBML transgenes via parthenogenesis. Genes 2020, 11, 1072. [Google Scholar] [CrossRef]
- Dore, C.; Prigent, J.; Desprez, B. In situ gynogenetic haploid plants of chicory (Cichorium intybus L.) after intergeneric hybridization with Cicerbita alpina Walbr. Plant Cell Rep. 1996, 15, 758–761. [Google Scholar] [CrossRef]
- Piosik, Ł.; Zenkteler, E.; Zenkteler, M. Development of haploid embryos and plants of Lactuca sativa induced by distant pollination with Helianthus annuus and H. tuberosus. Euphytica 2016, 208, 439–451. [Google Scholar] [CrossRef]
- Abdolmohammadi, M.; Kermani, M.J.; Zakizadeh, H.; Hamidoghli, Y. In vitro embryo germination and interploidy hybridization of rose (Rosa spp.). Euphytica 2014, 198, 255–264. [Google Scholar] [CrossRef]
- Kato, J.; Mii, M. Production of interspecific hybrid plants in Primula. Methods Mol. Biol. 2006, 318, 253–262. [Google Scholar] [CrossRef]
- Amano, J.; Kato, J.; Nakano, M.; Mii, M. Production of inter-section hybrids between Primula filchnerae and P. sinensis through ovule culture. Sci. Hortic. 2006, 110, 223–227. [Google Scholar] [CrossRef]
- Benega-Garcia, R.; Cisneros, A.; Schneider, B.; Tel-Zur, N. Gynogenesis in the vine cacti Hylocereus and Selenicereus (Cactaceae). Plant Cell Rep. 2009, 28, 719–726. [Google Scholar] [CrossRef]
- Cisneros, A.; Garcia, R.B.; Tel-Zur, N. Creation of novel interspecific-interploid Hylocereus hybrids (Cactaceae) via embryo rescue. Euphytica 2013, 189, 433–443. [Google Scholar] [CrossRef]
- Takamura, Y.; Takahashi, R.; Hikage, T.; Hatakeyama, K.; Takahata, Y. Production of haploids and doubled haploids from unfertilized ovule culture of various wild species of gentians (Gentiana spp.). Plant Cell Tissue Organ Cult. 2021, 146, 505–514. [Google Scholar] [CrossRef]
- Takahata, Y.; Keller, W.A. High frequency embryogenesis and plant regeneration in isolated 311 microspore culture of Brassica oleracea L. Plant Sci. 1991, 74, 235–242. [Google Scholar] [CrossRef]
- Chen, Y.M.; Mii, M. Inter-sectional hybrids obtained from reciprocal crosses between Begonia semperflorens (section Begonia) and B. ‘Orange Rubra’ (section Gaerdita × section Pritzelia). Breed. Sci. 2012, 62, 113–123. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sato, S.; Katoh, N.; Yoshida, H.; Iwai, S.; Hagimori, M. Production of doubled haploid plants of carnation (Dianthus caryophyllus L.) by pseudo fertilized ovule culture. Sci. Hortic. 2000, 83, 301–310. [Google Scholar] [CrossRef]
- Ishizaka, H. Interspecific hybridization by embryo rescue in the genus Cyclamen. Plant Biotechnol. 2008, 25, 511–519. [Google Scholar] [CrossRef][Green Version]
- Ishizaka, H. Breeding of fragrant cyclamen by interspecific hybridization and ion-beam irradiation. Breed. Sci. 2018, 68, 25–34. [Google Scholar] [CrossRef] [PubMed]
- Ishizaka, H.; Uematsu, J. Amphidiploids between Cyclamen persicum Mill. and C. hederifolium Aiton induced through colchicine treatment of ovules in vitro and plants. Breed. Sci. 1994, 44, 161–166. [Google Scholar] [CrossRef]
- Ishizaka, H. Cytogenetic studies in Cyclamen persicum, C. graecum (Primulaceae) and their hybrids. Plant Syst. Evol. 2003, 239, 1–14. [Google Scholar] [CrossRef]
- Ishizaka, H.; Uematsu, J. Interspecific hybrids of Cyclamen persicum Mill. and C. purpurascens Mill. produced by ovule culture. Euphytica 1995, 82, 31–37. [Google Scholar] [CrossRef]
- Premjet, D.; Obeng, A.K.; Kongbangkerd, A.; Premjet, S. Intergeneric hybrid from Jatropha curcas L. and Ricinus communis L.: Characterization and polyploid induction. Biology 2019, 8, 50. [Google Scholar] [CrossRef]
- Ji, W.; Li, G.R.; Luo, Y.X.; Ma, X.H.; Wang, M.; Ren, R. In vitro embryo rescue culture of F1 progenies from crosses between different ploidy grapes. Genet. Mol. Res. 2015, 14, 18616–18622. [Google Scholar] [CrossRef]
- Ji, W.; Li, Z.Q.; Zhou, Q.; Yao, W.K.; Wang, Y.J. Breeding new seedless grape by means of in vitro embryo rescue. Genet. Mol. Res. 2013, 12, 859–869. [Google Scholar] [CrossRef] [PubMed]
- Ji, W.; Wang, Y. Breeding for seedless grapes using Chinese wild Vitis spp. II. In vitro embryo rescue and plant development. J. Sci. Food. Agric. 2013, 93, 3870–3875. [Google Scholar] [CrossRef]
- Matsuzawa, Y.; Kaneko, Y.; Bang, S.W. Prospects of the wide cross for genetics and plant breeding in Brassicaceae. Bull. Coll. Agric. Utsunomiya Univ. 1996, 16, 5–10. [Google Scholar]
- Namai, H. Inducing cytogenetical alterations by means of interspecific and intergeneric hybridization in brassica crops. Gamma Field Symp. 1987, 26, 41–89. [Google Scholar]
- Ferrie, A.M.R.; Keller, W.A. Microspore culture for haploid plant production. In Fundamental Methods of Plant Cell, Tissue and Organ Culture and Laboratory Operations; Gamborg, O.L., Phillips, G.C., Eds.; Springer: Berlin/Heidelberg, Germany, 1995; pp. 155–164. [Google Scholar]
- Sosnowska, K.; Cegielska-Taras, T. Application of in vitro pollination of opened ovaries to obtain Brassica oleracea L. × B. rapa L. hybrids. Vitr. Cell. Dev. Biol. Plant 2014, 50, 257–262. [Google Scholar] [CrossRef]
- Zhou, Y.; Yang, M.; Zhao, S.; Shi, H.; Li, Y.; Gong, W.; Yang, J.; Wang, J.; Zou, Q.; Tao, L.; et al. Rapid creation of interspecific hybrid progeny to broaden genetic distance through Double Haploid (DH) inducer in Brassica napus. Plants 2022, 11, 695. [Google Scholar] [CrossRef]
- Fu, S.; Yin, L.; Xu, M.; Li, Y.; Wang, M.; Yang, J.; Fu, T.; Wang, J.; Shen, J.; Ali, A. Maternal doubled haploid production in interploidy hybridization between Brassica napus and Brassica allooctaploids. Planta 2017, 247, 113–125. [Google Scholar] [CrossRef]
- Li, C.; Sang, S.; Sun, M.; Yang, J.; Shi, Y.; Hu, X.; Li, Y.; Hao, M.; Chu, W.; Zhang, H.; et al. Direct modification of multiple gene homoeologs in Brassica oleracea and Brassica napus using doubled haploid inducer-mediated genome editing system. Plant Biotechnol. J. 2021, 19, 1889–1891. [Google Scholar] [CrossRef] [PubMed]
- Sauton, A.; Dumas de Vaulx, R. Induction of gynogenetic haploid plants in muskmelon (Cucumis melo L.) by use of irradiated pollen. Agronomie 1987, 7, 141–148. [Google Scholar] [CrossRef]
- Cuny, F.; Dumas de Vaulx, R.; Longhi, B.; Siadous, R. Analysis of muskmelon plants (Cucumis melo L.) obtained after pollination with gamma-irradiated pollen-effect of different doses. Agronomie 1992, 12, 623–630. [Google Scholar] [CrossRef]
- Cuny, F.; Grotte, M.; Dumas de Vaulx, R.; Rieu, A. Effects of gamma-irradiation of pollen on parthenogenetic haploid production in muskmelon (Cucumis melo L). Environ. Exp. Bot. 1993, 33, 301–312. [Google Scholar] [CrossRef]
- Yanmaz, R.; Ellialtioglu, S.; Taner, K.Y. The effects of gamma irradiation on pollen viability and haploid plant formation in snake cucumber (Cucumis melo L. var. flexuosus Naud.). In First International Symposium on Cucurbits; Abak, K., Büyükalaca, S., Eds.; ISHS: Adana, Turkey, 1999; pp. 307–310. [Google Scholar]
- Kuzuya, M.; Hosoy, K.; Masuya, Y.H.; Tomita, K.; Ezura, H. Histological observations of Powdery Mildew resistance in diploid and haploid melons. In The 7th EUCARPIA Meeting on Cucurbit Genetics & Breeding; Katzir, N., Paris, H.S., Eds.; ISHS: Ma’ale Ha Hamisha, Israel, 2000; pp. 71–75. [Google Scholar]
- Baktemur, G.; Taşkin, H.; Büyükalaca, S. Comparison of different methods for separation of haploid embryo induced through irradiated pollen and their economic analysis in Melon (Cucumis melo var. inodorus). Sci. World J. 2013, 2013, 529502. [Google Scholar] [CrossRef] [PubMed]
- Ollitrault, P.; Dambier, D.; Luro, F.; Froelicher, Y. Ploidy manipulation for breeding seedless triploid citrus. In Plant Breeding Reviews; Janick, J., Ed.; Wiley: Hoboken, NJ, USA, 2008; pp. 323–352. [Google Scholar]
- Geraci, G.; Esen, A.; Soost, R.K. Triploid progenies of citrus cultivars from 2x × 2x crosses. J. Hered. 1975, 66, 177–178. [Google Scholar] [CrossRef]
- Navarro, L.; Juárez, J.; Aleza, P.; Pina, J.A. Recovery of triploid seedless mandarin hybrids from 2n × 2n and 2n × 4n crosses by embryo rescue and flow cytometry. In Plant Biotechnology 2002 and Beyond; Vasil, I.K., Ed.; Springer: Dordrecht, The Netherlands, 2003; pp. 541–544. [Google Scholar]
- Aleza, P.; Juárez, J.; Cuenca, J.; Ollitrault, P.; Navarro, L. Recovery of citrus triploid hybrids by embryo rescue and flow cytometry from 2x × 2x sexual hybridisation and its application to extensive breeding programs. Plant Cell Rep. 2010, 29, 1023–1034. [Google Scholar] [CrossRef] [PubMed]
- Cimen, B.; Yesiloglu, T.; Donmez, D.; Kacar, Y.K.; Ercisly, S. Recovering triploid citrus hybrids from 2x × 2x sexual crosses with the aid of embryo rescue and flow cytometry in Turkey. Mol. Biol. Rep. 2022, 49, 5625–5634. [Google Scholar] [CrossRef]
- Aleza, P.; Juárez, J.; Ollitrault, P.; Navarro, L. Production of tetraploid plants of non apomictic citrus genotypes. Plant Cell Rep. 2009, 28, 1837–1846. [Google Scholar] [CrossRef]
- Singh, R.J.; Nelson, R.L. Methodology for creating alloplasmic soybean lines by using Glycine tomentella as a maternal parent. Plant Breed. 2014, 133, 624–631. [Google Scholar] [CrossRef]
- Bakhshaie, M.; Khosravi, S.; Azadi, P.; Bagheri, H.; van Tuyl, J.M. Biotechnological advances in Lilium. Plant Cell Rep. 2016, 35, 1799–1826. [Google Scholar] [CrossRef] [PubMed]
- Van Tuyl, J.M.; Meijer, B.; Van Dien, M.P. The use of oryzalin as an alternative for colchicine in in-vitro chromosome doubling of Lilium. Acta Hortic. 1992, 325, 625–630. [Google Scholar] [CrossRef]
- Takamura, T.; Lim, K.B.; Van Tuyl, J.M. Effect of a new compound on the mitotic polyploidization of Lilium longiflorum and Oriental hybrid lilies. Acta Hortic. 2002, 572, 37–42. [Google Scholar] [CrossRef]
- Ming, S.; Li, X.-F.; Kong, Y.; Shi, J.-F.; Zang, Q.-X. Polyploidy induction of three Lilium species endemic to china (Lilium pumilum, L. sargentiae, L. tsingtauense). Acta Hortic. 2012, 935, 83–90. [Google Scholar] [CrossRef]
- Lim, K.B.; Barba-Gonzalez, R.; Zhou, S.; Ramanna, M.S.; Van Tuyl, J.M. Meiotic polyploidization with homoeologous recombination induced by caffeine treatment in interspecific lily hybrids. Korean J. Genet. 2005, 27, 219–226. [Google Scholar]
- Chung, M.Y.; Chung, J.D.; Ramanna, M.; van Tuyl, J.M.; Lim, K.B. Production of polyploids and unreduced gametes in Lilium auratum × L. henryi hybrid. Int. J. Biol. Sci. 2013, 9, 693–701. [Google Scholar] [CrossRef]
- Barba-Gonzalez, R.; Lokker, A.C.; Lim, K.B.; Ramanna, M.S.; Van Tuyl, J.M. Use of 2n gametes for the production of sexual polyploids from sterile Oriental × Asiatic hybrids of lilies (Lilium). Theor. Appl. Genet. 2004, 109, 1125–1132. [Google Scholar] [CrossRef]
- Li, Z.; Pinkham, L.; Campbell, N.F.; Espinosa, A.C.; Conev, R. Development of triploid daylily (Hemerocallis) germplasm by embryo rescue. Euphytica 2009, 169, 313–318. [Google Scholar] [CrossRef]
- Yao, J.L.; Cohen, D. Production of triploid Zantedeschia hybrids using embryo rescue. N. Z. J. Crop. Hortic. Sci. 1996, 24, 297–301. [Google Scholar] [CrossRef]
- Moreno, R.; Espejo, J.A.; Cabrera, A.; Millan, T.; Gil, J. Ploidic and molecular analysis of ‘Morado de Huetor’ asparagus (Asparagus officinalis L.) population: A Spanish tetraploid landrace. Genet. Resour. Crop. Evol. 2006, 53, 729–736. [Google Scholar] [CrossRef]
- Moreno, R.; Espejo, J.A.; Cabrera, A.; Gil, J. Origin of tetraploid cultivated asparagus landraces inferred from nuclear ribosomal DNA internal transcribed spacers’ polymorphisms. Ann. Appl. Biol. 2008, 153, 233–241. [Google Scholar] [CrossRef]
- Doré, C. Chromosome doubling of asparagus (Asparagus officinalis) haploids by in vitro culture of colchicine treated meristems. Ann. L’Amelioration Plantes 1976, 26, 647–653. [Google Scholar]
- Skiebe, K.; Stein, M.; Gottwald, J.; Wolterstorff, B. Breeding of polyploidy asparagus (Asparagus officinalis L.). Plant Breed. 1991, 106, 99–106. [Google Scholar] [CrossRef]
- Nakashima, T.; Kunitake, H.; Tanaka, M. Intercrossing between diploid, triploid and tetraploid of Asparagus officinalis L. Jpn. J. Breed. 1992, 42, 462–463. [Google Scholar]
- Castro, P.; Rubio, J.; Gil, J.; Moreno, R. Introgression of new germplasm in current diploid cultivars of garden asparagus from tetraploid Spanish landrace “Morado de Huetor”. Sci. Hortic. 2014, 168, 157–160. [Google Scholar] [CrossRef]
- Carmona-Martin, E.; Regalado, J.J.; Raghavan, L.; Encina, C.L. In vitro induction of autooctoploid asparagus genotypes. Plant Cell Tissue Organ Cult. 2015, 121, 249–254. [Google Scholar] [CrossRef]
- Regalado, J.J.; Carmona-Martin, E.; Madrid, E.; Moreno, R.; Gil, J.; Encina, C.L. Production of “super-males” of asparagus by anther culture and its detection with SSR-ESTs. Plant Cell Tissue Organ Cult. 2016, 124, 119–135. [Google Scholar] [CrossRef]
- Moreno, R.; Espejo, J.A.; Gil, J. Development of triploid hybrids in Asparagus breeding employing a tetraploid landrace. Euphytica 2010, 173, 369–375. [Google Scholar] [CrossRef]
- Kunitake, H.; Nakashima, T.; Mori, K.; Tanaka, M. Somaclonal and chromosomal effects of genotype, ploidy and culture duration in Asparagus officinalis L. Euphytica 1998, 102, 309–316. [Google Scholar] [CrossRef]
- Ozaki, Y.; Narikiyo, K.; Fujita, C.; Okubo, H. Ploidy variation of progenies from intra- and inter-ploidy crosses with regard to trisomic production in asparagus (Asparagus officinalis L.). Sex. Plant Reprod. 2004, 17, 157–164. [Google Scholar] [CrossRef]
- Wojciechowska, B.; Pudelska, H. Hybrids and amphiploids of Aegilops ovata L. with Secale cereale L.: Production, morphology and fertility. J. Appl. Genet. 2002, 43, 415–421. [Google Scholar] [PubMed]
- Broughton, S.; Sidhu, P.K.; Davies, P.A. In vitro culture for doubled haploids: Tools for molecular breeding. Methods Mol. Biol. 2014, 1145, 167–189. [Google Scholar] [CrossRef]
- Patial, M.; Chauhan, R.; Chaudhary, H.K.; Pramanick, K.K.; Shukla, A.K.; Kumar, V.; Verma, R.P.S. Au-courant and novel technologies for efficient doubled haploid development in barley (Hordeum vulgare L.). Crit. Rev. Biotechnol. 2023, 43, 575–593. [Google Scholar] [CrossRef]
- Satpathy, P.; de la Fuente, S.A.; Ott, V.; Müller, A.; Büchner, H.; Daghma, D.E.S.; Kumlehn, J. Generation of doubled haploid barley by interspecific pollination with Hordeum bulbosum. In Doubled Haploid Technology. Methods in Molecular Biology; Segui-Simarro, J.M., Ed.; Humana: New York, NY, USA, 2021; Volume 2287, pp. 215–226. [Google Scholar] [CrossRef]
- Kasha, K.J.; Kao, K.N. High frequency haploid production in barley (Hordeum vulgare L.). Nature 1970, 225, 874–876. [Google Scholar] [CrossRef]
- Fedak, G. Haploids from barley × rye crosses. Can. J. Genet. Cytol. 1977, 19, 15–19. [Google Scholar] [CrossRef]
- Fedak, G. Hordecale (H. vulgare × Secale cereale L.). In Biotechnology in Agriculture and Forestry 2. Crops I; Bajaj, Y.P.S., Ed.; Springer: Berlin/Heidelberg, Germany, 1986; pp. 544–555. [Google Scholar]
- Sanei, M.; Pickering, R.; Kumke, K.; Nasuda, S.; Houben, A. Loss of centromeric histone H3 (CENH3) from centromeres precedes uniparental chromosome elimination in interspecific barley hybrids. Proc. Natl Acad. Sci. USA 2011, 108, 498–505. [Google Scholar] [CrossRef]
- Britt, A.B.; Kuppu, S. Cenh3: An emerging player in haploid induction technology. Front. Plant Sci. 2016, 7, 357. [Google Scholar] [CrossRef]
- Wang, N.; Gent, J.I.; Kelly, D.R. Haploid induction by a maize cenh3 null mutant. Sci. Adv. 2021, 7, 2229. [Google Scholar] [CrossRef]
- Lv, J.; Yu, K.; Wei, J.; Gui, H.; Liu, C.; Liang, D.; Wang, Y.; Zhou, H.; Carlin, R.; Rich, R.; et al. Generation of paternal haploids in wheat by genome editing of the centromeric histone CENH3. Nat. Biotechnol. 2020, 38, 1397–1401. [Google Scholar] [CrossRef]
- Widiez, T. Haploid embryos: Being like mommy or like daddy? Trends Plant Sci. 2021, 26, 425–427. [Google Scholar] [CrossRef]
- Hagberg, A.; Hagberg, G. High frequency of spontaneous haploids in the progeny of an induced mutation in barley. Hereditas 2009, 93, 341–343. [Google Scholar] [CrossRef]
- Polgári, D.; Mihók, E.; Sági, L. Composition and random elimination of paternal chromosomes in a large population of wheat × barley (Triticum aestivum L. × Hordeum vulgare L.) hybrids. Plant Cell Rep. 2019, 38, 767–775. [Google Scholar] [CrossRef]
- Bains, N.S.; Ravi, J.S.; Gosal, S.S. Production of wheat haploids through embryo rescue from wheat × maize crosses. Curr. Sci. 1995, 69, 621–623. Available online: https://www.jstor.org/stable/24097186 (accessed on 2 May 2023).
- Jauhar, P.P.; Xu, S.S.; Baenziger, P.S. Haploidy in cultivated wheats: Induction and utility in basic and applied research. Crop. Sci. 2009, 49, 737–755. [Google Scholar] [CrossRef]
- Deveax, P. Production of wheat doubled haploids through intergeneric hybridization with maize. Methods Mol. Biol. 2021, 2287, 267–279. [Google Scholar] [CrossRef]
- Eliby, S.; Bekkuzhina, S.; Kishchenko, O.; Iskakova, G.; Kylyshbayeva, G.; Jatayev, S.; Soole, K.; Langridge, P.; Borisjuk, N.; Shavrukov, Y. Developments and prospects for doubled haploid wheat. Biotechnol. Adv. 2022, 60, 108007. [Google Scholar] [CrossRef]
- Santra, M.; Hong Wang, H.; Seifert, S.; Haley, S. Doubled haploid laboratory protocol for wheat using wheat-maize wide hybridization. Methods Mol. Biol. 2017, 1679, 235–249. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Friebel, B.; Raupp, W.J.; Harrison, S.A.; Gill, B.S. Wheat embryogenesis and haploid production in wheat × maize hybrids. Euphytica 1996, 90, 315–324. [Google Scholar] [CrossRef]
- Guzy-Wróbelska, J.; Szarejko, I. Molecular and agronomic evaluation of wheat doubled haploid lines obtained through maize pollination and anther culture methods. Plant Breed. 2003, 122, 305–313. [Google Scholar] [CrossRef]
- Inagaki, M.N. Doubled haploid production in wheat through wide hybridization. In Doubled Haploid Production in Crop Plants: A Manual; Maluszynski, M., Ed.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2003; pp. 53–58. [Google Scholar]
- Bakos, F.; Jäger, K.; Barnabás, B. Regeneration of haploid plants after distant pollination of wheat via zygote rescue. Acta Biol. Cracov. Ser. Bot. 2005, 47, 167–171. [Google Scholar]
- Bidmeshkipour, A.; Thengane, R.J.; Bahagvat, M.D.; Ghaffari, S.M.; Rau, V.S. Production of haploid wheat via maize pollination. J. Sci. Islam. Repub. Iran 2007, 18, 5–11. [Google Scholar]
- Gernand, D.; Rutten, T.; Varshney, A.; Rubtsova, M.; Prodanovi, S.; Brüß, C.; Kumlehn, J.; Matzk, F.; Houben, A. Uniparental chromosome elimination at mitosis and interphase in wheat and pearl millet crosses involves micronucleus formation, progressive heterochromatinization, and DNA fragmentation. Plant Cell 2005, 17, 2431–2438. [Google Scholar] [CrossRef]
- Chaudhary, H.K.; Sethi, G.S.; Singh, S.; Pratap, A.; Sharma, S. Efficient haploid induction in wheat by using pollen of Imperata cylindrica. Plant Breed. 2005, 124, 96–98. [Google Scholar] [CrossRef]
- Komeda, N.; Chaudhary, H.K.; Suzuki, G. Cytological evidence for chromosome elimination in wheat × Imperata cylindrica hybrids. Genes Genet. Syst. 2007, 82, 241–248. [Google Scholar] [CrossRef][Green Version]
- Mochida, K.; Tsujimoto, H. Production of wheat doubled haploids by pollination with Job’s tears (Coix lachryma-jobi L.). J. Hered. 2001, 92, 81–83. [Google Scholar] [CrossRef] [PubMed]
- O’Donoughue, L.S.; Bennett, M.D. Durum wheat haploid production using maize wide-crossing. Theor. Appl. Genet. 1994, 89, 559–566. [Google Scholar] [CrossRef] [PubMed]
- Doğramaci-Altuntepe, M.; Jauhar, P.P. Production of durum wheat substitution haploids from durum × maize crosses and their cytological characterization. Genome 2001, 44, 137–142. [Google Scholar] [CrossRef]
- Joppa, L.R.; Williams, N.D. Langdon durum disomic substitution lines and aneuploid analysis in tetraploid wheat. Genome 1988, 30, 222–228. [Google Scholar] [CrossRef]
- Almouslem, A.B.; Jauhar, P.P.; Peterson, T.S.; Bommineni, V.R.; Rao, M.B. Haploid durum wheat production via hybridization with maize. Crop. Sci. 1998, 38, 1080–1087. [Google Scholar] [CrossRef]
- Jauhar, P.P. Haploid and doubled haploid production in durum wheat by wide hybridization. In Doubled Haploid Production in Crop Plants: A Manual; Maluszynski, M., Ed.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2003; pp. 161–167. [Google Scholar]
- García-Llamas, C.; Martín, A.; Ballesteros, J. Differences among auxin treatments on haploid production in durum wheat × maize crosses. Plant Cell Rep. 2004, 23, 46–49. [Google Scholar] [CrossRef] [PubMed]
- García-Llamas, C.; Ramírez, M.C.; Ballesteros, J. Effect of pollinator on haploid production in durum wheat crossed with maize and pearl millet. Plant Breed. 2004, 123, 201–203. [Google Scholar] [CrossRef]
- Davies, P.A.; Sidhu, P.K. Oat doubled haploids following maize pollination. Methods Mol. Biol. 2017, 1536, 23–30. [Google Scholar] [CrossRef]
- Skrzypek, E.; Warzecha, T.; Noga, A.; Warchoł, M.; Czyczyło-Mysza, I.; Dziurka, K.; Marcińska, I.; Kapłoniak, K.; Sutkowska, A.; Nita, Z.; et al. Complex characterization of oat (Avena sativa L.) lines obtained by wide crossing with maize (Zea mays L.). PeerJ 2018, 6, e5107. [Google Scholar] [CrossRef]
- Warchoł, M.; Skrzypek, E.; Nowakowska, A.; Marcińska, I.; Czyczyło-Mysza, I.; Dziurka, K.; Juzoń, K.; Cyganek, K. The effect of auxin and genotype on the production of Avena sativa L. doubled haploid lines. Plant Growth Regul. 2016, 78, 155–165. [Google Scholar] [CrossRef]
- Skrzypek, E.; Warchoł, M.; Czyczyło-Mysza, I.; Marcińska, I.; Nowakowska, A.; Dziurka, K.; Juzoń, K.; Noga, A. The effect of light intensity on the production of oat (Avena sativa L.) doubled haploids through oat × maize crosses. Cereal Res. Commun. 2016, 44, 490–500. [Google Scholar] [CrossRef][Green Version]
- Noga, A.; Skrzypek, E.; Warchoł, M.; Czyczyło-Mysza, I.; Dziurka, K.; Marcińska, I.; Juzoń, K.; Warzecha, T.; Sutkowska, A.; Nita, Z.; et al. Conversion of oat (Avena sativa L.) haploid embryos into plants in relation to embryo developmental stage and regeneration media. Vitr. Cell Dev. Biol. Plant 2016, 52, 590–597. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kynast, R.G.; Riera-Lizarazu, O. Development and use of oat-maize chromosome additions and radiation hybrids. Methods Mol. Biol. 2011, 701, 259–284. [Google Scholar] [CrossRef]
- Warchoł, M.; Czyczyło-Mysza, I.; Marcińska, I.; Dziurka, K.; Noga, A.; Skrzypek, E. The effect of genotype, media composition, pH and sugar concentrations on oat (Avena sativa L.) doubled haploid production through oat × maize crosses. Acta Physiol. Plant. 2018, 40, 93. [Google Scholar] [CrossRef]
- Zhuang, J.J.; Xu, J. Increasing differentiation frequencies in wheat pollen callus. In Cell and Tissue Culture Techniques for Cereal Crop Improvement; Hu, H., Vega, M.R., Eds.; Science Press: Beijing, China, 1983; p. 431. [Google Scholar]
- Dziurka, K.; Dziurka, M.; Muszyńska, E.; Czyczyło-Mysza, I.; Warchoł, M.; Juzoń, K.; Laskoś, K.; Skrzypek, E. Anatomical and hormonal factors determining the development of haploid and zygotic embryos of oat (Avena sativa L.). Sci. Rep. 2022, 12, 548. [Google Scholar] [CrossRef]
- Möller, B.; Weijers, W. Auxin control of embryo patterning. Cold Spring Harb. Perspect. Biol. 2009, 1, a001545. [Google Scholar] [CrossRef]
- Terceros, G.C.; Resentini, F.; Cucinotta, M.; Manrique, S.; Colombo, L.; Mendes, M.A. The importance of cytokinins during reproductive development in Arabidopsis and beyond. Int. J. Mol. Sci. 2020, 21, 8161. [Google Scholar] [CrossRef]
- Hays, D.B.; Yeung, E.C.; Pharis, R.P. The role of gibberellins in embryo axis development. J. Exp. Bot. 2002, 53, 1747–1751. [Google Scholar] [CrossRef] [PubMed]
- Marcińska, I.; Czyczyło-Mysza, I.; Skrzypek, E.; Warchoł, M.; Zieliński, K.; Dubas, E. Obtaining of winter rye (Secale cereale L. ssp. cereale) haploid embryos through hybridization with maize (Zea mays L.). Cereal Res. Commun. 2018, 46, 521–532. [Google Scholar] [CrossRef]
- Ma, L.; Yang, C.; Zeng, D.; Cai, J.; Li, X.; Ji, Z.; Xia, Y.; Qian, Q.; Bao, J. Mapping QTLs for heading synchrony in a doubled haploid population of rice in two environments. J. Genet. Genom. 2009, 36, 297–304. [Google Scholar] [CrossRef]
- Lee, G.-H.; Kang, I.-K.; Kim, K.-M. Mapping of novel QTL regulating grain shattering using doubled haploid population in rice (Oryza sativa L.). Int. J. Genom. 2016, 2016, 2128010. [Google Scholar] [CrossRef]
- Kulkarni, S.R.; Balachandran, S.M.; Ulaganathan, K.; Balakrishnan, D.; Prasad, A.S.H.; Rekha, G.; Kousik, M.B.V.N.; Hajira, S.K.; Kale, R.R.; Aleena, D.; et al. Mapping novel QTLs for yield related traits from a popular rice hybrid KRH-2 derived doubled haploid (DH) population. 3 Biotech 2021, 11, 513. [Google Scholar] [CrossRef] [PubMed]
- Rao, Y.; Dong, G.; Zeng, D.; Hu, J.; Zeng, L.; Gao, Z.; Zhang, G.; Guo, L.; Quia, Q. Genetic analysis of leaffolder resistance in rice. J. Genet. Genom. 2010, 37, 325–331. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Chen, X.; Zhang, X.; Lin, Z.; Shang, J.; Xu, J.; Zhai, W.; Zhu, L. Isolation and identification of a gene in response to rice blast disease in rice. Plant Mol. Biol. 2004, 54, 99–109. [Google Scholar] [CrossRef] [PubMed]
- Du, X.X.; Park, J.R.; Wang, X.H.; Jan, R.; Lee, G.S.; Kim, K.M. Genotype and phenotype interaction between OsWKRYq6 and BLB after Xanthomonas oryzae pv. Oryzae inoculation in the field. Plants 2022, 11, 287. [Google Scholar] [CrossRef]
- Zhao, D.-D.; Son, J.-H.; Farooq, M.; Kim, K.-M. Identification of candidate gene for internode length in rice to enhance resistance to lodging using QTL analysis. Plants 2021, 10, 1369. [Google Scholar] [CrossRef]
- Kim, T.H.; Hur, Y.J.; Han, S.I.; Cho, J.H.; Kim, K.M.; Lee, J.H.; Song, Y.C.; Kwon, Y.U.; Shin, D. Drought-tolerant QTL qVDT11 leads to stable tiller formation under drought stress conditions in rice. Plant Sci. 2017, 256, 131–138. [Google Scholar] [CrossRef]
- Zhao, D.-D.; Jang, Y.-H.; Kim, E.-G.; Park, J.-R.; Jan, R.; Lubna; Asaf, S.; Asif, S.; Farooq, M.; Chung, H.; et al. Identification of a major locus for lodging resistance to typhoons using QTL analysis in rice. Plants 2023, 12, 449. [Google Scholar] [CrossRef]
- Swamy, B.P.M.; Descalsota, G.I.L.; Nha, C.T.; Amparado, A.; Inabangan-Asilo, M.A.; Manito, C.; Tesoro, F.; Reinke, R. Identification of genomic regions associated with agronomic and biofortification traits in DH populations of rice. PLoS ONE 2018, 13, e0201756. [Google Scholar] [CrossRef]
- Li, Z.K.; Yu, S.B.; Lafitte, H.R.; Huang, N.; Courtois, B.; Hittalmani, S.; Vijayakumar, C.H.M.; Liu, G.F.; Wang, G.C.; Shashidhar, H.E.; et al. QTL × environment interactions in rice. I. heading date and plant height. Theor. Appl. Genet. 2003, 108, 141–153. [Google Scholar] [CrossRef]
- Zhao, D.-D.; Park, J.-R.; Jang, Y.-H.; Kim, E.-G.; Du, X.-X.; Farooq, M.; Yun, B.-J.; Kim, K.-M. Identification of one major QTL and a novel gene OsIAA17q5 associated with tiller number in rice using QTL analysis. Plants 2022, 11, 538. [Google Scholar] [CrossRef] [PubMed]
- Park, J.R.; Kim, E.G.; Jang, Y.H.; Kim, K.M. Screening and identification of genes affecting grain quality and spikelet fertility during high-temperature treatment in grain filling stage of rice. BMC Plant Biol. 2021, 21, 263. [Google Scholar] [CrossRef]
- d’Hooghvorst, I.; Ferreres, I.; Nogués, S. Anther culture and chromosome doubling in Mediterranean japonica rice. Methods Mol. Biol. 2021, 2287, 333–341. [Google Scholar] [CrossRef]
- Tripathy, S.K. High-throughput doubled haploid production for indica rice breeding. Methods Mol. Biol. 2021, 2287, 343–360. [Google Scholar]
- Ali, J.; Nicolas, K.L.C.; Akther, S.; Torabi, A.; Ebadi, A.A.; Marfori-Nazarea, C.M.; Mahender, A. Improved anther culture media for enhanced callus formation and plant regeneration in rice (Oryza sativa L.). Plants 2021, 10, 839. [Google Scholar] [CrossRef] [PubMed]
- Rout, P.; Naik, N.; Ngangkham, U.; Verma, R.L.; Katara, J.L.; Singh, O.N.; Samantaray, S. Doubled Haploids generated through anther culture from an elite long duration rice hybrid, CRHR32: Method optimization and molecular characterization. Plant Biotechnol. 2016, 33, 177–186. [Google Scholar] [CrossRef]
- Naik, N.; Rout, P.; Umakanta, N.; Verma, R.L.; Katara, J.L.; Sahoo, K.K.; Singh, O.N.; Samantaray, S. Development of doubled haploids from an elite indica rice hybrid (BS6444G) using anther culture. Plant Cell Tissue Organ Cult. 2017, 128, 679–689. [Google Scholar] [CrossRef]
- Serrat, X.; Cardona, M.; Gil, J.; Brito, A.M.; Moysset, L.; Nogues, S.; Lalanne, E. A mediterranean japonica rice (Oryza sativa) cultivar improvement through anther culture. Euphytica 2013, 195, 31–44. [Google Scholar] [CrossRef][Green Version]
- Datta, K.; Sahoo, G.; Krishnan, S.; Ganguly, M.; Datta, S.K. Genetic stability developed for β-Carotene synthesis in BR29 rice line using dihaploid homozygosity. PLoS ONE 2014, 9, e100212. [Google Scholar] [CrossRef]
- d’Hooghvorst, I.; Ramos-Fuentes, E.; López-Cristofannini, C.; Ortega, M.; Vidal, R.; Serrat, X.; Nogués, S. Antimitotic and hormone effects on green double haploid plant production through anther culture of Mediterranean japonica rice. Plant Cell Tissue Organ Cult. 2018, 134, 205–215. [Google Scholar] [CrossRef]
- Bishnoi, U.; Jain, R.; Rohilla, J.; Chowdhury, V.; Gupta, K.; Chowdhury, J. Anther culture of recalcitrant indica × basmati rice hybrids. Euphytica 2000, 114, 93–101. [Google Scholar] [CrossRef]
- Cha-Um, S.; Srianan, B.; Pichakum, A.; Kirdmanee, C. An efficient procedure for embryogenic callus induction and double haploid plant regeneration through anther culture of Thai aromatic rice (Oryza sativa L. subsp. indica). Vitr. Cell Dev. Biol. Plant 2009, 45, 171–179. [Google Scholar] [CrossRef]
- Yi, G.; Lee, H.-S.; Kim, K.-M. Improved marker-assisted selection efficiency of multi-resistance in doubled haploid rice plants. Euphytica 2014, 203, 421–428. [Google Scholar] [CrossRef]
- Ferreres, I.; Ortega, M.; López-Cristoffanini, C.; Nogués, S.; Serrat, X. Colchicine and osmotic stress for improving anther culture efficiency on long grain temperate and tropical japonica rice genotypes. Plant Biotechnol. 2019, 36, 269–273. [Google Scholar] [CrossRef] [PubMed]
- Lantos, C.; Jancsó, M.; Székely, Á.; Nagy, É.; Szalóki, T.; Pauk, J. Improvement of anther culture to integrate doubled haploid technology in temperate rice (Oryza sativa L.) breeding. Plants 2022, 11, 3446. [Google Scholar] [CrossRef]
- Lantos, C.; Jancsó, M.; Székely, Á.; Szalóki, T.; Venkatanagappa, S.; Pauk, J. Development of in vitro anther culture for doubled haploid plant production in indica rice (Oryza sativa L.) genotypes. Plants 2023, 12, 1774. [Google Scholar] [CrossRef]
- Dash, B.; Bhuyan, S.S.; Kumar Singh, S.; Chandravani, M.; Swain, N.; Rout, P.; Katara, J.L.; Parameswaran, C.; Devanna, B.N.; Samantaray, S. Androgenesis in indica rice: A comparative competency in development of doubled haploids. PLoS ONE 2022, 17, e0267442. [Google Scholar] [CrossRef]
- Pattnaik, S.S.; Dash, B.; Bhuyan, S.S.; Katara, J.L.; Parameswaran, C.; Verma, R.; Ramesh, N.; Samantaray, S. Anther culture efficiency in quality hybrid rice: A comparison between hybrid rice and its ratooned plants. Plants 2020, 9, 1306. [Google Scholar] [CrossRef]
- López-Cristoffanini, C.; Serrat, X.; Ramos-Fuentes, E.; Hooghvorst, I.; Llaó, R.; López-Carbonell, M.; Nogués, S. An improved anther culture procedure for obtaining new commercial Mediterranean temperate japonica rice (Oryza sativa) genotypes. Plant Biotechnol. 2018, 35, 161–166. [Google Scholar] [CrossRef] [PubMed]
- Tai, H.K.; Li, K.T.; Charng, Y.C. A one-time inducible transposon for terminating selectable markers in transgenic plants. Bot. Stud. 2011, 52, 375–381. [Google Scholar]
- Yang, H.-C.; Charng, Y.-C. Application of an inducible transposon with anther culture in generation of di-haploid homologous mutants. Bot Stud. 2014, 55, 27. [Google Scholar] [CrossRef] [PubMed]
- Gilles, L.M.; Martinant, J.-P.; Rogowsky, P.M.; Widiez, T. Haploid induction in plants. Curr. Biol. 2017, 27, R1089–R1107. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Wang, K.; Jia, Z.; Gong, Q.; Lin, Z.; Du, L.; Pei, X.; Ye, X. Efficient induction of haploid plants in wheat by editing of TaMTL using an optimized Agrobacterium-mediated CRISPR system. J. Exp. Bot. 2020, 71, 1337–1349. [Google Scholar] [CrossRef] [PubMed]
- Kelliher, T.; Starr, D.; Richbourg, L.; Chintamanani, S.; Delzer, B.; Nuccio, M.L.; Green, J.; Chen, Z.; McCuiston, J.; Wang, W.; et al. MATRILINEAL, a sperm-specific phospholipase, triggers maize haploid induction. Nature 2017, 542, 105–109. [Google Scholar] [CrossRef] [PubMed]
- Yao, L.; Zhang, Y.; Liu, C.; Liu, Y.; Wang, Y.; Liang, D.; Liu, J.; Sahoo, G.; Kelliher, T. OsMATL mutation induces haploid seed formation in indica rice. Nat. Plants 2018, 4, 530–533. [Google Scholar] [CrossRef]
- Cheng, Z.; Sun, Y.; Yang, S.; Zhi, H.; Yin, T.; Ma, X.; Zhang, H.; Diao, X.; Guo, Y.; Li, X.; et al. Establishing in planta haploid inducer line by edited SiMTL in foxtail millet (Setaria italica). Plant Biotechnol. J. 2021, 19, 1089–1091. [Google Scholar] [CrossRef]
- Zhang, X.; Shi, C.; Li, S.; Zhang, B.; Luo, P.; Peng, X.; Zhao, P.; Dresselhaus, T.; Sun, M.-X. A female in vivo haploid-induction system via mutagenesis of egg cell-specific peptidases. Mol. Plant. 2023, 16, 471–480. [Google Scholar] [CrossRef]
- Zhong, Y.; Chen, B.; Wang, D.; Zhu, X.; Li, M.; Zhang, J.; Chen, M.; Wang, M.; Riksen, T.; Liu, J.; et al. In vivo maternal haploid induction in tomato. Plant Biotechnol. J. 2022, 20, 250–252. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, L.; Zhang, J.; Jia, M.; Cao, L.; Yu, J.; Zhao, D. Haploid induction in allotetraploid tobacco using DMPs mutation. Planta 2022, 255, 98. [Google Scholar] [CrossRef]
- Li, Y.; Li, D.; Xiao, Q.; Wang, H.; Wen, J.; Tu, J.; Shen, J.; Fu, T.; Yi, B. An in planta haploid induction system in Brassica napus. J. Integr. Plant Biol. 2022, 64, 1140–1144. [Google Scholar] [CrossRef] [PubMed]
- Wang, N.; Xia, X.; Jiang, T.; Li, L.; Zhang, P.; Niu, L.; Cheng, H.; Wang, K.; Lin, H. In planta haploid induction by genome editing of DMP in the model legume Medicago truncatula. Plant Biotechnol. J. 2022, 20, 22–24. [Google Scholar] [CrossRef]
- Zhao, X.; Yuan, K.; Liu, Y.; Zhang, N.; Yang, L.; Zhang, Y.; Wang, Y.; Ji, J.; Fang, Z.; Han, F.; et al. In vivo maternal haploid induction based on genome editing of DMP in Brassica oleracea. Plant Biotechnol. J. 2022, 20, 2242–2244. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Yin, J.; Luo, J.; Tang, D.; Zhu, X.; Wang, J.; Liu, Z.; Wang, P.; Zhong, Y.; Liu, C.; et al. Construction of homozygous diploid potato through maternal haploid induction. 3 Biotech 2022, 3, 163–168. [Google Scholar] [CrossRef]
- Li, J.; Yu, X.; Zhang, C.; Li, N.; Zhao, J. The application of CRISPR/Cas technologies to Brassica crops: Current progress and future perspectives. aBIOTECH 2022, 3, 146–161. [Google Scholar] [CrossRef]
- Chaikam, V.; Molenaar, W.; Melchinger, A.E.; Boddupalli, P.M. Doubled haploid technology for line development in maize: Technical advances and prospects. Theor. Appl. Genet. 2019, 132, 3227–3243. [Google Scholar] [CrossRef]
- Wang, D.; Zhong, Y.; Feng, B.; Qi, X.; Yan, T.; Liu, J.; Guo, S.; Wang, Y.; Liu, Z.; Cheng, D.; et al. The RUBY reporter enables efficient haploid identification in maize and tomato. Plant Biotechnol. J. 2023, 21, 1707–1715. [Google Scholar] [CrossRef] [PubMed]
- Clement, J.S.; Mabry, T.J. Pigment evolution in the Caryophyllales: A systematic overview. Bot. Acta 1996, 109, 360–367. [Google Scholar] [CrossRef]
- Polturak, G.; Breitel, D.; Grossman, N.; Sarrion-Perdigones, A.; Weithorn, E.; Pliner, M.; Orzaez, D.; Granell, A.; Rogachev, I.; Aharoni, A. Elucidation of the first committed step in betalain biosynthesis enables the heterologous engineering of betalain pigments in plants. New Phytol. 2016, 210, 269–283. [Google Scholar] [CrossRef]
- He, Y.; Zhang, T.; Sun, H.; Zhan, H.; Zhao, Y. A reporter for noninvasively monitoring gene expression and plant transformation. Hortic. Res. 2020, 7, 152. [Google Scholar] [CrossRef] [PubMed]
- Kumlehn, J.; Schieder, O.; Lörz, H. In vitro development of wheat (Triticum aestivum L.) from zygote to plant via ovule culture. Plant Cell Rep. 1997, 16, 663–667. [Google Scholar] [CrossRef] [PubMed]
- Bakos, F.; Szabó, L.; Olmedilla, A.; Barnabás, B. Histological comparison between wheat embryos developing in vitro from isolated zygotes and those developing in vivo. Sex. Plant Reprod. 2009, 22, 15–25. [Google Scholar] [CrossRef]
- Kranz, E.; Lörz, H. In vitro fertilisation with isolated, single gametes results in zygotic embryogenesis and fertile maize plants. Plant Cell 1993, 5, 739–746. [Google Scholar] [CrossRef] [PubMed]
- Leduc, N.; Matthys-Rochon, E.; Rougier, M.; Mogensen, L.; Holm, P.B.; Magnard, J.L.; Dumas, C. Isolated maize zygotes mimic in vivo embryonic development and express microinjected genes when cultured in vitro. Dev. Biol. 1996, 177, 190–203. [Google Scholar] [CrossRef]
- Holm, P.B.; Knudsen, S.; Mouritzen, P.; Negri, D.; Olsen, F.L.; Roué, C. Regeneration of fertile barley plants from mechanically isolated protoplasts of the fertilized egg cell. Plant Cell 1994, 6, 531–543. [Google Scholar] [CrossRef][Green Version]
- Kumlehn, J.; Lörz, H.; Kranz, E. Differentiation of isolated wheat zygotes into embryos and normal plants. Planta 1998, 205, 327–333. [Google Scholar] [CrossRef]
- Kumlehn, J.; Lörz, H.; Kranz, E. Monitoring individual development of isolated wheat zygotes: A novel approach to study early embryogenesis. Protoplasma 1999, 208, 156–162. [Google Scholar] [CrossRef]
- Bakos, F.; Darkó, É.; Pónya, Z.; Barnabás, B. Regeneration of fertile wheat (Triticum aestivum L.) plants from isolated zygotes using wheat microspore culture as nurse cells. Plant Cell Tissue Organ Cult. 2003, 74, 243–247. [Google Scholar] [CrossRef]
- Bakos, F.; Darkó, É.; Pónya, Z.; Barnabás, B. Application of wheat (Triticum aestivum L.) microspore culture and ovaries to raise wheat zygotes in vitro. Acta Biol. Crac. Bot. 2003, 45, 107–110. [Google Scholar]
- Zhang, J.; Dong, W.H.; Galli, A.; Potrykus, I. Regeneration of fertile plants from isolated zygotes of rice (Oryza sativa). Plant Cell Rep. 1999, 19, 128–132. [Google Scholar] [CrossRef]
- Uchiumi, T.; Komatsu, S.; Koshiba, T.; Okamoto, T. Isolation of gametes and central cells from Oryza sativa L. Sex. Plant Reprod. 2006, 19, 37–45. [Google Scholar] [CrossRef]
- Uchiumi, T.; Uemura, I.; Okamoto, T. Establishment of an in vitro fertilization system in rice (Oryza sativa L.). Planta 2007, 226, 581–589. [Google Scholar] [CrossRef]
- Koiso, N.; Toda, E.; Ichikawa, M.; Kato, N.; Okamoto, T. Development of gene expression system in egg cells and zygotes isolated from rice and maize. Plant Direct 2017, 1, e00010. [Google Scholar] [CrossRef]
- He, Y.C.; Sun, M.X.; Yang, H.Y. Regeneration of fertile plants from isolated tobacco zygotes by in vitro culture. Chin. Sci. Bull. 2004, 49, 810–814. [Google Scholar] [CrossRef]
- Toda, E.; Kiba, T.; Kato, N.; Okamoto, T. Isolation of gametes and zygotes from Setaria viridis. J. Plant Res. 2022, 135, 627–633. [Google Scholar] [CrossRef] [PubMed]
- Sprunck, S.; Baumann, U.; Edwards, K.; Langridge, P.; Dresselhaus, T. The transcript composition of egg cells changes significantly following fertilization in wheat (Triticum aestivum L.). Plant J. 2005, 41, 660–672. [Google Scholar] [CrossRef]
- Okamoto, T.; Scholten, S.; Lörz, H.; Kranz, E. Identification of genes that are up- or down-regulated in the apical or basal cell of maize two-celled embryos and monitoring their expression during zygote development by a cell manipulation- and PCR-based approach. Plant Cell Physiol. 2005, 46, 332–338. [Google Scholar] [CrossRef]
- Okamoto, T.; Kranz, E. In vitro fertilization—A tool to dissect cell specification from a higher plant zygote. Curr. Sci. 2005, 89, 1861–1869. [Google Scholar]
- Yang, H.P.; Kaur, N.; Kiriakopolos, S.; McCormick, S. EST generation and analyses towards identifying female gametophyte-specific genes in Zea mays L. Planta 2006, 224, 1004–1014. [Google Scholar] [CrossRef]
- Ning, J.; Peng, X.B.; Qu, L.H.; Xin, H.P.; Yan, T.T.; Sun, M. Differential gene expression in egg cells and zygotes suggests that the transcriptome is restructed before the first zygotic division in tobacco. FEBS Lett. 2006, 580, 1747–1752. [Google Scholar] [CrossRef]
- Uchiumi, T.; Shinkawa, T.; Isobe, T.; Okamoto, T. Identification of the major protein components of rice egg cells. J. Plant Res. 2007, 120, 575–579. [Google Scholar] [CrossRef]
- Toda, E.; Okamoto, T. Gene expression and genome editing systems by direct delivery of macromolecules into rice egg cells and zygotes. Bio-Protocol 2020, 10, e3681. [Google Scholar] [CrossRef]
- Fambrini, M.; Pugliesi, C.; Vernieri, P.; Giuliano, G.; Baroncelli, S. Characterization of a sunflower (Helianthus annuus L.) mutant deficient in carotenoid synthesis and abscisic-acid content induced by in vitro tissue culture. Theor. Appl. Genet. 1993, 87, 65–69. [Google Scholar] [CrossRef]
- Conti, A.; Pancaldi, S.; Fambrini, M.; Michelotti, V.; Bonora, S.; Salvini, M.; Pugliesi, C. A deficiency at the gene coding for ζ-carotene desaturase characterizes the sunflower non dormant-1 mutant. Plant Cell Physiol. 2004, 45, 445–455. [Google Scholar] [CrossRef]
- Lo Schiavo, F.; Giuliano, G.; De Vries, S.C.; Genga, A.; Bollini, R.; Pitto, L.; Cozzani, G.; Nuti-Ronchi, V.; Terzi, M. A carrot cell variant temperature sensitive for somatic embryogenesis reveals a defect in the glycosylation of extracellular proteins. Mol. Gen. Genet. 1990, 223, 385–393. [Google Scholar] [CrossRef]
- De Jong, A.J.; Cordewener, J.; Lo Schiavo, F.; Terzi, M.; Vandekerckhove, J.; Van Kammen, A.; De Vries, S.C. A carrot somatic embryo mutant is rescued by chitinase. Plant Cell 1992, 4, 425–433. [Google Scholar] [CrossRef]
- Raboy, V.J. Progress in breeding low phytate crops. J. Nutr. 2002, 132, 503S–505S. [Google Scholar] [CrossRef]
- Liu, Q.L.; Xu, X.H.; Ren, X.L.; Fu, H.W.; Wu, D.X.; Shu, Q.Y. Generation and characterization of low phytic acid germplasm in rice (Oryza sativa L.). Theor. Appl. Genet. 2007, 114, 803–814. [Google Scholar] [CrossRef]
- Larson, S.R.; Rutger, J.N.; Young, K.A.; Raboy, V. Isolation and genetic mapping of a non-lethal rice (Oryza sativa L.) low phytic acid 1 mutation. Crop. Sci. 2000, 40, 1397–1405. [Google Scholar] [CrossRef]
- Raboy, V.; Gerbasi, P.F.; Young, K.A.; Stoneberg, S.D.; Pickett, S.G.; Bauman, A.T.; Murthy, P.P.N.; Sheridan, W.F.; Ertl, D.S. Origin and seed phenotype of maize low phytic acid 1–1 and low phytic acid 2–1. Plant Physiol. 2000, 124, 355–368. [Google Scholar] [CrossRef]
- Pilu, R.; Panzeri, D.; Gavazzi, G.; Rasmussen, S.K.; Consonni, G.; Nielsen, E. Phenotypic, genetic and molecular characterization of a maize low phytic acid mutant (lpa 241). Theor. Appl. Genet. 2003, 107, 980–987. [Google Scholar] [CrossRef] [PubMed]
- Meis, S.J.; Fehr, W.R.; Schnebly, S.R. Seed source effect on field emergence of soybean lines with reduced phytate and raffinose saccharides. Crop. Sci. 2003, 43, 1336–1339. [Google Scholar] [CrossRef]
- Dolfini, S.; Consonni, G.; Viotti, C.; Dal Prà, M.; Saltini, G.; Giulini, A.; Pilu, R.; Malgioglio, A.; Gavazzi, G. A mutational approach to the study of seed development in maize. J. Exp. Bot. 2007, 58, 1197–1205. [Google Scholar] [CrossRef] [PubMed]
- Consonni, G.; Aspesi, C.; Barbante, A.; Dolfini, S.; Giuliani, C.; Giulini, A.; Hansen, S.; Brettschneider, R.; Pilu, R.; Gavazzi, G. Analysis of four maize mutants arrested in early embryogenesis reveals an irregular pattern of cell division. Sex. Plant Reprod. 2003, 15, 281–290. [Google Scholar] [CrossRef]
- Gu, J.; Dempsey, S.; Newton, K.J. Rescue of a maize mitochondrial cytochrome oxidase mutant by tissue culture. Plant J. 1994, 6, 787–794. [Google Scholar] [CrossRef] [PubMed]
- Le Page-Degivry, M.T.; Garello, G. In situ abscisic acid synthesis: A requirement for induction of embryo dormancy in Helianthus annuus. Plant Physiol. 1992, 98, 1386–1390. [Google Scholar] [CrossRef]
- Pugliesi, C.; Fambrini, M.; Vernieri, P.; Baroncelli, S. Characterization of a wilty sunflower (Helianthus annuus L.) mutant I. Abscisic acid content, light-dark changes in the stomatal conductance and genetic analysis. J. Exp. Bot. 1994, 45, 533–538. [Google Scholar] [CrossRef]
- Giordani, T.; Natali, L.; D’Ercole, A.; Pugliesi, C.; Fambrini, M.; Vernieri, P.; Vitagliano, C.; Cavallini, A. Expression of a dehydrin gene during embryo development and drought stress in ABA-deficient mutants of sunflower (Helianthus annuus L.). Plant Mol. Biol. 1999, 39, 739–748. [Google Scholar] [CrossRef]
- Ochatt, S.J.; Sangwan, R.S.; Marget, P.; Ndong, Y.A.; Rancillac, M.; Perney, P. New approaches towards the shortening of generation cycles for faster breeding of protein legumes. Plant Breed. 2002, 121, 436–440. [Google Scholar] [CrossRef]
- Alissa, A.; Jonard, R.; Serieys, H.; Vincourt, P. La culture d’embryons isolés in vitro dans un programme d’amélioration du tournesol. C.R. Acad Sci. Paris T. Série III 1986, 302, 161–164. [Google Scholar]
- Azpiroz, H.S.; Vincourt, P.; Serieys, H.; Gallais, A. La culture in vitro des embryons immature dans acceleration du cycle de selection des lignées de tournesol et ses effets morphovégétatifs. Helia 1987, 10, 35–38. [Google Scholar]
- Caser, M.; Dente, F.; Ghione, G.G.; Mansuino, A.; Giovannini, A.; Scariot, V. Shortening of selection time of Rosa hybrida by in vitro culture of isolated embryos and immature seeds. Propag. Ornam. Plants 2014, 14, 139–144. [Google Scholar]
- Yuan, M.S.; Wu, M.C.; Shii, C.T. Shortening breeding cycles of spider lilies (Lycoris spp.) through embryo culture and dikaryotype hybridization between Lycoris aurea and “a” karyotype species. Acta Hortic. 2003, 620, 345–352. [Google Scholar] [CrossRef]
- Bermejo, C.; Gatti, I.; Cointry, E. In vitro embryo culture to shorten the breeding cycle in lentil (Lens culinaris Medik). Plant Cell Tiss Organ Cult. 2016, 127, 585–590. [Google Scholar] [CrossRef]
- Surma, M.; Adamski, T.; Święcicki, W.; Barzyk, P.; Kaczmarek, Z.; Kuczyńska, A.; Krystkowiak, K.; Mikofajczak, K.; Ogrodowicz, P. Preliminary results of in vitro culture of pea and lupin embryos for the reduction of generation cycles in single seed descent technique. Acta Soc. Bot. Pol. 2013, 82, 231–236. [Google Scholar] [CrossRef]
- Ochatt, S.J.; Sangwan, R.S. In vitro shortening of generation time in Arabidopsis thaliana. Plant Cell Tissue Organ Cult. 2008, 93, 133–137. [Google Scholar] [CrossRef]
- Rodríguez-Garay, B.; Rodríguez-Domínguez, J.M. Micropropagation of Agave Species. Methods Mol. Biol. 2018, 1815, 151–159. [Google Scholar] [CrossRef]
- Barsberg, S.; Rasmussen, H.N.; Kodahl, N. Composition of Cypripedium calceolus (Orchidaceae) seeds analyzed by attenuated total reflectance IR spectroscopy: In search of understanding longevity in the ground. Am. J. Bot. 2013, 100, 2066–2073. [Google Scholar] [CrossRef]
- Merritt, D.J.; Hay, F.R.; Swarts, N.D.; Sommerville, K.D.; Dixon, K.W. Ex situ conservation and cryopreservation of orchid germplasm. Int. J. Plant Sci. 2014, 175, 46–58. [Google Scholar] [CrossRef]
- Kendon, J.P.; Rajaovelona, L.; Sandford, H.; Fang, R.; Bell, J.; Sarasan, V. Collecting near mature and immature orchid seeds for ex situ conservation: ‘in vitro collecting’ as a case study. Bot. Stud. 2017, 58, 34. [Google Scholar] [CrossRef]
- Iyer, C.P.A.; Subramanyam, M.D. Possible role of embryo culture in mango breeding. Indian J. Hortic. 1971, 29, 135–136. [Google Scholar]
- Krishna, H.; Singh, S.K. Biotechnological advances in mango (Mangifera indica L.) and their future implication in crop improvement: A review. Biotechnol. Adv. 2007, 25, 223–243. [Google Scholar] [CrossRef] [PubMed]
- Chandra, R.; Padaria, J.C.; Srivastava, S. Factors influencing in vitro establishment of mango shoot buds. Indian J. Plant Physiol. 2004, 9, 136–144. [Google Scholar]
- Sahijram, L.; Bollamma, K.T.; Naren, A.; Soneji, J.R.; Dinesh, M.R.; Halesh, G.K. In vitro embryo rescue in mango (Mangifera indica L.) breeding. Ind. J. Hortic. 2005, 62, 235–237. [Google Scholar]
- Pérez-Hernández, J.B.; Grajal-Martín, M.J. In vitro culture of immature zygotic mango embryos and plantlet development. Hortscience 2011, 46, 1528–1532. [Google Scholar] [CrossRef]
- Souza, F.V.D.; RebouÇas, C.C.; Souza, E.H.; Peixoto, C.P. In vitro conservation of mango (Mangifera indica L.) Ubá and Carlota cvs. through culturing immature embryos. An. Acad. Bras. Cienc. 2020, 92, e20190400. [Google Scholar] [CrossRef]
- Chen, Y.C.; Huang, Y.Y.; Wang, C.M.; Chiu, C.A.; Lin, H.L.; Lee, P.F.; Cheng, Y.M.; Chang, C. Re-emergence of Lilium callosum Sieb. et Zucc. in Taiwan after a fire allows propagation and renews the possibility of conservation. Bot. Stud. 2017, 58, 47. [Google Scholar] [CrossRef]
- Vovides, A.P.; Iglesias, C. Seed germination of Magnolia dealbata Zucc. (Magnoliaecea), an endangered species from Mexico. Hortscience 1996, 31, 877. [Google Scholar] [CrossRef]
- IUCN. The IUCN Red List of Threatened Species. Version 2019-2. Available online: http://www.iucnredlist.org (accessed on 22 May 2023).
- Mata-Rosas, M.; Jiménez-Rodríguez, A.; Chávez-Avila, V.M. Somatic embryogenesis and organogenesis in Magnolia dealbata Zucc. (Magnoliaceae), an endangered, endemic Mexican species. Hortscience 2006, 41, 1325–1329. [Google Scholar] [CrossRef]
- Meinke, D.W.; Sussex, I.M. Embryo-lethal mutants of Arabidopsis thaliana. A model system for genetic analysis of plant embryo development. Dev. Biol. 1979, 72, 50–61. [Google Scholar] [CrossRef]
- Gutiérrez-Marcos, J.F.; Dal Prà, M.; Giulini, A.; Costa, L.M.; Gavazzi, G.; Cordelier, S.; Sellam, O.; Tatout, C.; Paul, W.; Perez, P.; et al. empty pericarp4 encodes a mitochondrion-targeted pentatricopeptide repeat protein necessary for seed development and plant growth in maize. Plant Cell 2007, 19, 196–210. [Google Scholar] [CrossRef]
- Candela, H.; Pérez-Pérez, J.M.; Micol, J.L. Uncovering the post-embryonic functions of gametophytic- and embryonic-lethal genes. Trends Plant Sci. 2011, 16, 336–345. [Google Scholar] [CrossRef] [PubMed]
- Burger, D.W.; Liu, L.; Zary, K.W.; Lee, C.I. Organogenesis and plant regeneration from immature embryos of Rosa hybrida L. Plant Cell Tissue Organ Cult. 1990, 21, 147–152. [Google Scholar] [CrossRef]
- Kim, S.W.; In, D.S.; Choi, P.S.; Liu, J.R. Plant regeneration from immature zygotic embryo-derived embryogenic calluses and cell suspension cultures of Catharanthus roseus. Plant Cell Tissue Organ Cult. 2004, 76, 131–135. [Google Scholar] [CrossRef]
- Azad, M.A.; Rabbani, M.G.; Amin, L. Plant regeneration and somatic embryogenesis from immature embryos derived through interspecific hybridization among different Carica species. Int. J. Mol. Sci. 2012, 13, 7065–7076. [Google Scholar] [CrossRef] [PubMed]
- Backiyarani, S.; Uma, S.; Saranya, S.; Durai, P.; Perianayagaraj, S.E.; Selvaraj, V.; Saraswathi, M.S.; Karthic, R.; Kalpana, S. Multiple shoot induction in zygotic embryos: A strategy for acceleration of banana breeding. Plant Cell Tissue Organ Cult. 2021, 147, 339–350. [Google Scholar] [CrossRef]
- Toda, E.; Kolso, N.; Takebayashi, A.; Ichikawa, M.; Kiba, T.; Osakabe, K.; Osakabe, Y.; Sakakibara, H.; Kato, N.; Okamoto, T. An efficient DNA- and selectable-marker-free genome-editing system using zygotes in rice. Nat. Plants 2019, 5, 363–368. [Google Scholar] [CrossRef]
- Kang, M.; Lee, K.; Finley, T.; Chappell, H.; Veena, V.; Wang, K. An improved Agrobacterium-mediated transformation and genome-editing method for maize inbred B104 using a ternary vector system and immature embryos. Front. Plant Sci. 2022, 13, 860971. [Google Scholar] [CrossRef]
- Darqui, F.S.; Radonic, L.M.; Baracochea, V.C.; Hopp, H.E.; Bilbao, M.L. Peculiarities of the transformation of Asteraceae family species: The cases of sunflower and lettuce. Front. Plant Sci. 2021, 12, 767459. [Google Scholar] [CrossRef]
- Fambrini, M.; Usai, G.; Pugliesi, C. Induction of somatic embryogenesis in plants: Different players and focus on WUSCHEL and WUS-RELATED HOMEOBOX (WOX) transcription factors. Int. J. Mol. Sci. 2022, 23, 15950. [Google Scholar] [CrossRef]
- Saha, S.; Tullu, A.; Lulsdorf, M.M.; Vandenberg, A. Improvement of embryo rescue technique using 4-chloroindole-3 acetic acid in combination with in vivo grafting to overcome barriers in lentil interspecific crosses. Plant Cell Tissue Organ Cult. 2015, 120, 109–116. [Google Scholar] [CrossRef]
- Li, S.; Liu, K.; Yu, S.; Jia, S.; Chen, S.; Fu, Y.; Sun, F.; Luo, Q.; Wang, Y. The process of embryo abortion of stenospermocarpic grape and it develops into plantlet in vitro using embryo rescue. Plant Cell Tissue Organ Cult. 2020, 143, 389–409. [Google Scholar] [CrossRef]

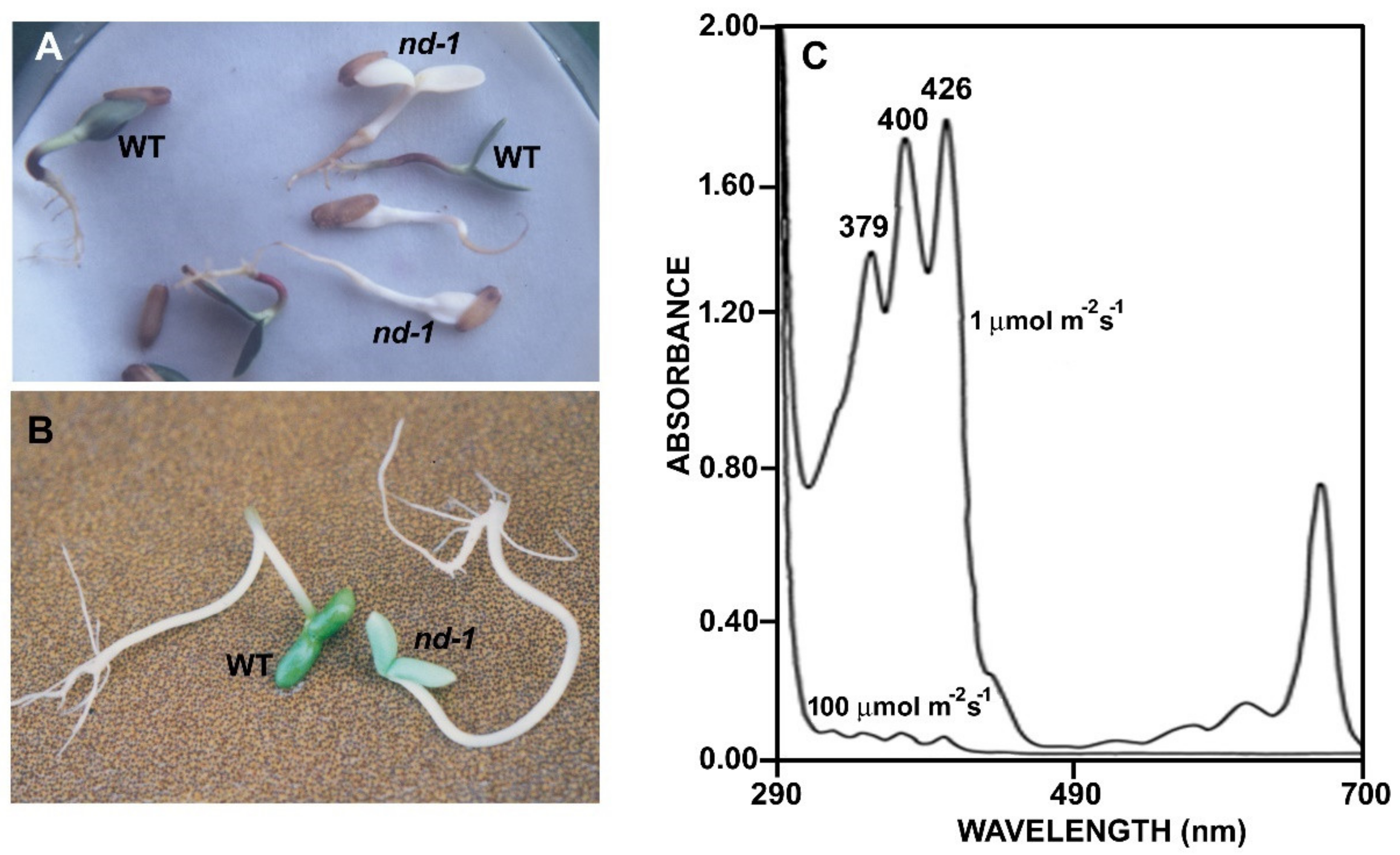
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rogo, U.; Fambrini, M.; Pugliesi, C. Embryo Rescue in Plant Breeding. Plants 2023, 12, 3106. https://doi.org/10.3390/plants12173106
Rogo U, Fambrini M, Pugliesi C. Embryo Rescue in Plant Breeding. Plants. 2023; 12(17):3106. https://doi.org/10.3390/plants12173106
Chicago/Turabian StyleRogo, Ugo, Marco Fambrini, and Claudio Pugliesi. 2023. "Embryo Rescue in Plant Breeding" Plants 12, no. 17: 3106. https://doi.org/10.3390/plants12173106
APA StyleRogo, U., Fambrini, M., & Pugliesi, C. (2023). Embryo Rescue in Plant Breeding. Plants, 12(17), 3106. https://doi.org/10.3390/plants12173106








