Phenotyping and Identification of Molecular Markers Associated with Leaf Rust Resistance in the Wheat Germplasm from Kazakhstan, CIMMYT and ICARDA
Abstract
1. Introduction
2. Results
2.1. Reaction of the Wheat Collection to Two Races of P. triticina at the Seedling Stage
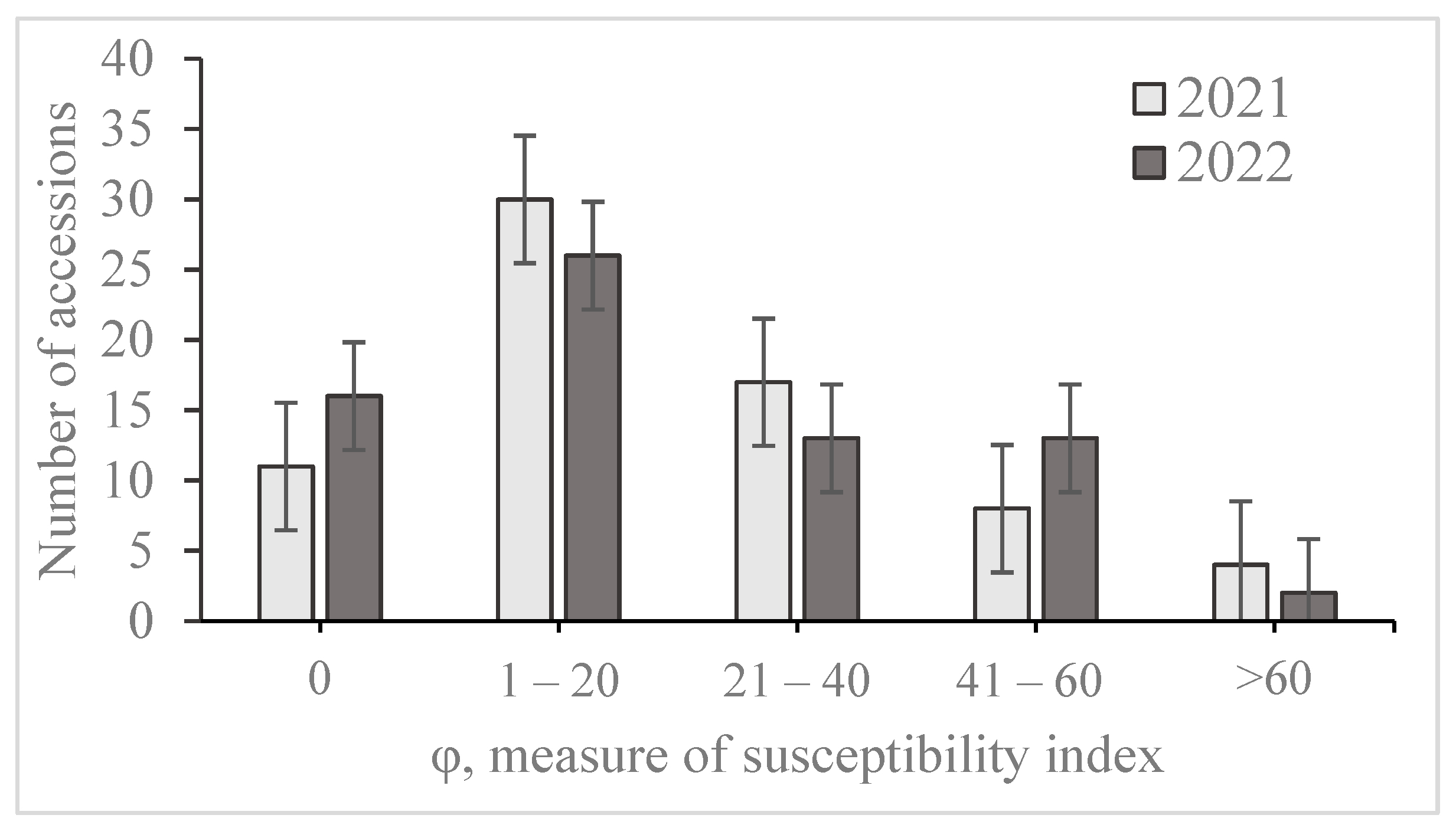
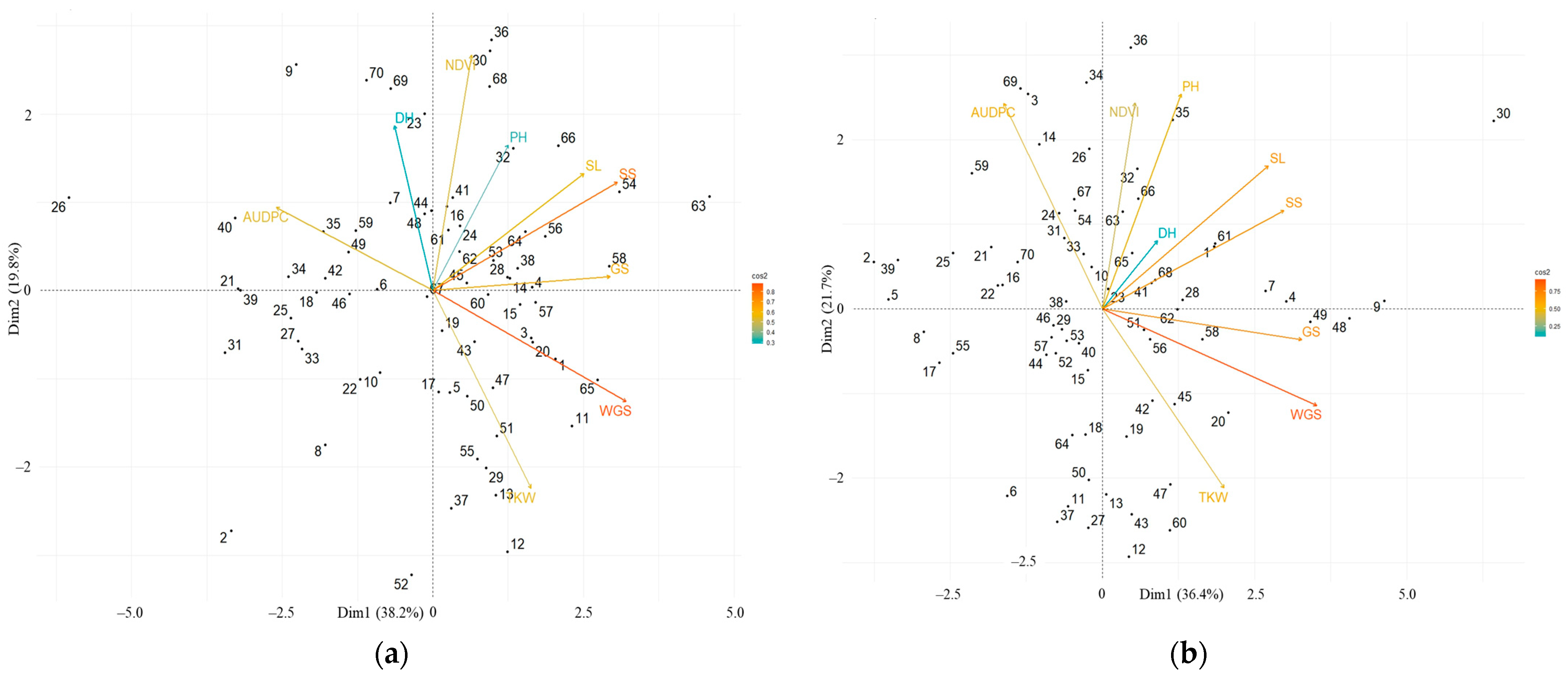
2.2. Field Evaluation
2.3. Identification of Lr Resistance Genes Using Molecular Markers
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Leaf Rust Spore Collection, Multiplication and Race Identification
4.3. Leaf Rust Evaluation at the Seedling Stage
4.4. Leaf Rust Evaluation at the Adult Plant Stage
4.5. Statistical Data Processing
4.6. DNA Extraction and Molecular Screening of Lr Resistance Genes
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Food and Agriculture (FAO). Organization of the United Nation. Available online: http://www.fao.org/faostat/en/#data/CC (accessed on 24 March 2023).
- Statista. Available online: https://www.statista.com/ (accessed on 13 March 2023).
- Statistics Committee. Ministry of National Economy of the Republic of Kazakhstan. Available online: https://new.stat.gov.kz/ (accessed on 20 April 2023).
- Kolmer, J. Leaf Rust of Wheat: Pathogen Biology, Variation and Host Resistance. Forests 2013, 4, 70–84. [Google Scholar] [CrossRef]
- Zhao, J.; Wang, M.; Chen, X.; Kang, Z. Role of Alternate Hosts in Epidemiology and Pathogen Variation of Cereal Rusts. Annu. Rev. Phytopathol. 2016, 54, 207–228. [Google Scholar] [CrossRef]
- Bolton, M.D.; Kolmer, J.A.; Garvin, D.F. Wheat leaf rust caused by Puccinia triticina. Mol. Plant Pathol. 2008, 9, 563–575. [Google Scholar] [CrossRef]
- Dinh, H.X.; Singh, D.; Periyannan, S.; Park, R.F.; Pourkheirandish, M. Molecular genetics of leaf rust resistance in wheat and barley. Theor. Appl. Genet. 2020, 133, 2035–2050. [Google Scholar] [CrossRef] [PubMed]
- Huerta-Espino, J.; Singh, R.P.; German, S. Global status of wheat leaf rust caused by Puccinia triticina. Euphytica 2011, 179, 143–160. [Google Scholar] [CrossRef]
- Pretorius, Z.A.; Visser, B.; Terefe, T. Races of Puccinia triticina detected on wheat in Zimbabwe, Zambia and Malawi and regional germplasm responses. Australas. Plant Pathol. 2015, 44, 217–224. [Google Scholar] [CrossRef]
- Roelfs, A.P. Barley stripe rust in Texas. Plant Dis. 1992, 76, 538. [Google Scholar] [CrossRef]
- Singh, R.P.; Huerta-Espino, J.; Pfeiffer, W.; Figueroa-Lopez, P. Occurrence and impact of a new leaf rust race on durum wheat in northwestern Mexico from 2001 to 2003. Plant Dis. 2004, 88, 703–708. [Google Scholar] [CrossRef] [PubMed]
- Riaz, M.; Wong, Y. Estimation of Yield Losses Due to Leaf Rust and Late Seeding on Wheat (Triticum aestivum L.) Variety Seher-06 in District Faisalabad, Punjab, Pakistan. Adv. Biotech. Micro. 2017, 5, 555657. [Google Scholar]
- Koyshibaev, M.K. Diseases of Wheat; FAO: Ankara, Turkey, 2018; p. 365. [Google Scholar]
- Keishilov, Z.; Kokhmetova, A.; Kumarbaeva, M.; Bolatbekova, A.; Malysheva, A.; Kokhmetova, A. Monitoring of leaf rust (Puccinia recondita) of spring wheat in Northern Kazakhstan 2019–2022. Herald Sci. KazATU 2022, 1, 930. [Google Scholar]
- Morgounov, A.; Akin, B.; Demir, L.; Keser, M.; Kokhmetova, A.; Martynov, S.; Yessimbekova, M. Yield gain due to fungicide application in varieties of winter wheat (Triticum aestivum) resistant and susceptible to leaf rust. Crop Pasture Sci. 2015, 66, 649. [Google Scholar] [CrossRef]
- Galymbek, K.; Kokhmetova, A.M.; Akan, K.; Madenova, A.K.; Atishova, M.N. Identification of germplasm of Wheat on leaf rust (Puccinia recondita Rob. Ex Desm. F.sp. Tritici). Ecol. Environ. Conserv. 2017, 23, 1211–1218. [Google Scholar]
- Kokhmetova, A.; Rsaliyev, S.; Atishova, M.; Kumarbayeva, M.; Malysheva, A.; Keishilov, Z.; Zhanuzak, D.; Bolatbekova, A. Evaluation of wheat germplasm for resistance to leaf rust (Puccinia triticina) and Identification of the Sources of Lr Resistance Genes Using Molecular Markers. Plants 2021, 10, 1484. [Google Scholar] [CrossRef]
- Kokhmetova, A.; Atishova, M.; Madenova, A.; Kumarbayeva, M. Genotyping of wheat germplasm for resistance to toxins of tan spot Pyrenophora tritici-repentis. J. Biotechnol. 2019, 305, 53. [Google Scholar] [CrossRef]
- Kokhmetova, A.; Kovalenko, N.; Kumarbaeva, M. Pyrenophora tritici-repentis population structure in the Republic of Kazakhstan and identification of wheat germplasm resistant to tan spot. Vavilov J. Genet. Breed. 2020, 24, 722–729. [Google Scholar] [CrossRef]
- Kokhmetova, A.; Kumarbayeva, M.; Atishova, M.; Nehe, A.; Riley, I.; Morgounov, A. Identification of high-yielding wheat genotypes resistant to Pyrenophora tritici-repentis (tan spot). Euphytica 2021, 217, 97. [Google Scholar] [CrossRef]
- Kumarbayeva, M.; Kokhmetova, A.; Kovalenko, N.; Atishova, M.; Keishilov, Z.; Aitymbetova, K. Characterization of Pyrenophora tritici-repentis (tan spot of wheat) races in Kazakhstan. Phytopathol. Mediterr. 2022, 61, 243–257. [Google Scholar] [CrossRef]
- Kokhmetova, A.; Sharma, R.; Rsaliyev, S.; Galymbek, K.; Baymagambetova, K.; Ziyaev, Z.; Morgounov, A. Evaluation of Central Asian wheat germplasm for stripe rust resistance. Plant Genet. Resour. 2018, 16, 178–184. [Google Scholar] [CrossRef]
- Kokhmetova, A.; Rsaliyev, A.; Malysheva, A.; Atishova, M.; Kumarbayeva, M.; Keishilov, Z. Identification of stripe rust resistance genes in common wheat cultivars and breeding lines from Kazakhstan. Plants 2021, 10, 2303. [Google Scholar] [CrossRef]
- Malysheva, A.; Kokhmetova, A.; Kumarbayeva, M.; Zhanuzak, D.; Bolatbekova, A.; Keishilov, Z.; Gultyaeva, E.; Kokhmetova, A.; Tsygankov, V.; Dutbayev, Y.; et al. Identification of Carriers of Puccinia Striiformis Resistance Genes in the Population of Recombinant Inbred Wheat Lines. Int. J. Biol. Chem. 2022, 15, 4–10. [Google Scholar] [CrossRef]
- Yessenbekova, G.; Kokhmetova, A.; Madenova, A.; Amanov, O.; Dutbayev, Y.; Kampitova, G. Identification of Lr34/Yr18 Gene in Wheat Germplasm in Kazakhstan. In Proceedings of the 2014 APS-CPS Joint Meeting, Minneapolis, MN, USA, 9–13 August 2014; p. 252. [Google Scholar]
- Kokhmetova, A.M.; Atishova, M.N. Identification of sources of resistance to wheat stem rust using molecular markers. Russ. J. Genet. Appl. Res. 2012, 2, 486–493. [Google Scholar] [CrossRef]
- Olivera, F.P.; Szabo, L.; Kokhmetova, A.; Morgunov, A.; Luster, D.G.; Jin, Y. Puccinia graminis f. sp. tritici population causing recent wheat stem rust epidemics in Kazakhstan is highly diverse and includes novel virulences. Phytopathology 2022, 112, 2403–2415. [Google Scholar] [CrossRef] [PubMed]
- Flor, H. Genetics of Pathogenicity in Melampsoralini. J. Agric. Res. 1946, 73, 335–357. [Google Scholar]
- McCallum, B.; Hiebert, C.; Cloutier, S. A review of wheat leaf rust research and the development of resistant cultivars in Canada. Can. J. Plant Pathol. 2016, 38, 1–18. [Google Scholar] [CrossRef]
- Park, R.; Golegaonkar, P.; Derevnina, L.; Sandhu, K.; Karaoglu, H.; Elmansour, H.; Dracatos, P.; Singh, D. Leaf rust of cultivated barley: Pathology and control. Annu. Rev. Phytopathol. 2015, 53, 565–589. [Google Scholar] [CrossRef]
- Chaves, M.; Martinelli, J.; Guterres, C.; Sganzerla, F. The Cereal Rust: An Overview. Pest Tech. 2008, 2, 38–55. [Google Scholar]
- McIntosh, R.; Dubcovsky, J.; Rogers, W.; Xia, X.; Raupp, W. Catalogue of Gene Symbols for Wheat: 2020 Supplement. Ann. Wheat Newslett. 2020, 66, 1–20. [Google Scholar]
- Sears, E. The transfer of leaf rust resistance from Aegilops umbellulata into wheat. Brookhaven Symp. Biol. 1956, 9, 1–21. [Google Scholar]
- Riley, R.; Chapman, V.; Johnson, R. The incorporation of yellow rust resistance of Aegilops comosa into wheat by genetically induced homoeologous recombination. Nature 1968, 217, 383–384. [Google Scholar] [CrossRef]
- Ellis, J.G.; Lagudah, E.S.; Spielmeyer, W.; Dodds, P.N. The past, present and future of breeding rust resistant wheat. Front. Plant Sci. 2014, 5, 641. [Google Scholar] [CrossRef]
- Parlevliet, J.; Ommeren, A. Partial resistance of barley to leaf rust, Puccinia hordei. II. Relationship between field trials, micro plot tests and latent period. Euphytica 1975, 24, 293–303. [Google Scholar] [CrossRef]
- Dodds, P.N.; Rathjen, J.P. Plant immunity: Towards an integrated view of plant–pathogen interactions. Nat. Rev. Genet. 2010, 11, 539–548. [Google Scholar] [CrossRef]
- Dyck, P.; Kerber, E. Resistance of the Race-Specific Type. In The Cereal Rusts; Roelfs, A., Bushnell, W.R., Eds.; Academic Press: Orlando, FL, USA, 1985; Volume 2, pp. 469–500. [Google Scholar]
- Park, R.; McIntosh, R. Adult plant resistances to Puccinia recondita f. sp. tritici in wheat. N. Z. J. Crop Hortic. Sci. 1994, 22, 151–158. [Google Scholar] [CrossRef][Green Version]
- Kolmer, J.; Singh, R.; Garvin, D.; Viccars, L.; William, H.; Huerta-Espino, J.; Obonnaya, F.; Raman, H.; Orford, S.; Bariana, H.; et al. Analysis of the Lr34/Yr18 rust resistance region in wheat germplasm. Crop Sci. 2008, 48, 1841–1852. [Google Scholar] [CrossRef]
- Dyck, P. The association of a gene for leaf rust resistance with the chromosome 7D suppressor of stem rust resistance in common wheat. Genome 1987, 29, 467–469. [Google Scholar] [CrossRef]
- Krattinger, S.; Lagudah, E.; Spielmeyer, W.; Singh, R.; Huerta-Espino, J.; McFadden, H.; Bossolini, E.; Selter, L.; Keller, B. A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 2009, 323, 1360. [Google Scholar] [CrossRef]
- Lagudah, E.; McFadden, H.; Singh, R.; Huerta-Espino, J.; Bariana, H.; Spielmeyer, W. Molecular genetic characterization of the Lr34/Yr18 slow rusting resistance gene region in wheat. Theor. Appl. Genet. 2006, 114, 21–30. [Google Scholar] [CrossRef]
- Helguera, M.; Khan, I.; Kolmer, J.; Lijavetzky, D.; Zhong-Qi, L.; Dubcovsky, J. PCR Assays for the Cluster of Rust Resistance Genes and Their Use to Develop Isogenic Hard Red Spring Wheat Lines. Crop Sci. 2003, 43, 1839–1847. [Google Scholar] [CrossRef]
- Moore, J.; Herrera-Foessel, S.; Lan, C.; Schnippenkoetter, W.; Ayliffe, M.; Huerta-Espino, J.; Lagudah, E. A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nat. Genet. 2015, 47, 1494–1498. [Google Scholar] [CrossRef]
- Kumar, S.; Bhardwaj, S.; Gangwar, O. Characterization of five new pathotypes of Puccinia triticina identified from Northeast India, Nepal, and Bangladesh. Aust. Plant Pathol. 2022, 51, 315–325. [Google Scholar] [CrossRef]
- Melchinger, A. Use of molecular markers in breeding for oligogenic disease resistance. Plant Breed. 1990, 104, 1–19. [Google Scholar] [CrossRef]
- Singh, A.; Pallavi, J.; Gupta, P.; Prabhu, K. Identification of microsatellite markers linked to leaf rust adult plant resistance (APR) gene 48 in wheat. Plant Breed. 2010, 130, 31–34. [Google Scholar] [CrossRef]
- Yue, Z.; Zai-Feng, L.; Xing, L.; Long, W.; Ye, Z.; Da-Qun, L. Molecular mapping for leaf rust resistance genes in wheat line Tian 95HF2. Acta Agron. Sin. 2010, 36, 1265–1269. [Google Scholar]
- Gultyaeva, E.; Kokhmetova, A.; Shreyder, E.; Shaydayuk, E.; Atishova, M.; Madenova, A.; Malysheva, A.; Galymbek, K. Genetic variability of perspective breeding material of spring bread wheat for resistance to leaf rust in Russia and Kazakhstan. Bull. NAS RK 2020, 3, 60–68. [Google Scholar] [CrossRef]
- Schachermayr, G.; Siedler, H.; Gale, M.; Winzeler, H.; Winzeler, M.; Keller, B. Identification and localization of molecular markers linked to the Lr9 leaf rust resistance gene of wheat. Theor. Appl. Genet. 1994, 88, 110–115. [Google Scholar] [CrossRef]
- Chelkowski, J.; Golka, L.; Stepien, L. Application of STS markers for leaf rust resistance genes in near–isogenic lines of spring wheat cv. Thatcher. J. Appl. Genet. 2003, 44, 323–338. [Google Scholar]
- Zhang, W.; Dubcovsky, J. Association between allelic variation at the Phytoene synthase 1 gene and yellow pigment content in the wheat grain. Theor. Appl. Genet. 2008, 116, 635–645. [Google Scholar] [CrossRef] [PubMed]
- Mago, R.; Spielmeyer, W.; Lawrence, G. Identification and mapping of molecular markers linked to rust resistance genes located on chromosome 1RS of rye using wheat-rye translocation lines. Theor. Appl. Genet. 2002, 104, 1317–1324. [Google Scholar] [CrossRef]
- Mago, R.; Miah, H.; Lawrence, G.J.; Wellings, C.R.; Spielmeyer, W.; Bariana, H.S.; Ellis, J.G. High-resolution mapping and mutation analysis separate the rust resistance genes Sr31, Lr26, and Yr9 on the short arm of rye chromosome 1. Theor. Appl. Genet. 2005, 112, 41–50. [Google Scholar] [CrossRef]
- Vikal, Y.; Chhuneja, P.; Singh, R.; Dhaliwal, H. Tagging of an Aegilops speltoides Derived Leaf Rust Resistance Gene Lr28 with a Microsatellite Marker in Wheat. J. Plant Biochem. Biotechnol. 2004, 13, 47–49. [Google Scholar] [CrossRef]
- McIntosh, R.; Wellings, C.; Park, R. Wheat Rusts: An Atlas of Resistance Genes; CSIRO Publishing: Melbourne, Australia, 1995. [Google Scholar] [CrossRef]
- William, M.; Singh, R.; Huerta-Espino, J.; Islas, S.; Hoisington, D. Molecular marker mapping of leaf rust resistance gene Lr46 and its association with stripe rust resistance gene Yr29 in wheat. Phytopathology 2003, 93, 153–159. [Google Scholar] [CrossRef]
- Tomkowiak, A.; Skowrońska, R.; Kwiatek, M.; Spychała, J.; Weigt, D.; Kurasiak-Popowska, D.; Niemann, J.; Mikołajczyk, S.; Nawracała, J.; Kowalczewski, P.; et al. Identification of leaf rust resistance genes Lr34 and Lr46 in common wheat (Triticum aestivum L. ssp. aestivum) lines of different origin using multiplex PCR. Open Life Sci. 2021, 16, 172–183. [Google Scholar] [CrossRef] [PubMed]
- Herrera-Foessel, S.; Singh, R.; Huerta-Espino, J.; Rosewarne, G.; Periyannan, S.; Viccars, L.; Calvo-Salazar, V.; Lan, C.; Lagudah, E. Lr68: A new gene conferring slow rusting resistance to leaf rust in wheat. Theor. Appl. Genet. 2012, 124, 1475–1486. [Google Scholar] [CrossRef] [PubMed]
- Wellings, C. Global status of stripe rust: A review of historical and current threats. Euphytica 2011, 179, 129–141. [Google Scholar] [CrossRef]
- Morgounov, A.; Rosseeva, L.; Koyshibayev, M. Leaf rust of spring wheat in Northern Kazakhstan and Siberia: Incidence, virulence, and breeding for resistance. Aust. J. Agric. Res. 2007, 58, 847. [Google Scholar] [CrossRef]
- Gultyaeva, E.; Kovalenko, N.; Shamanin, V.; Tyunin, V.; Shreyder, E.; Shaydayuk, E.; Morgunov, A. Population structure of leaf pathogens of common spring wheat in the West Asian regions of Russia and North Kazakhstan in 2017. Vavilov J. Genet. Breed. 2018, 22, 363–369. [Google Scholar] [CrossRef]
- Gultyaeva, E.; Shaydayuk, E.; Shamanin, V.; Akhmetova, A.; Tyunin, V.; Shreyder, E.; Kashina, I.; Eroshenko, L.; Sereda, G.; Morgunov, A. Genetic structure of Russian and Kazakhstani leaf rust causative agent Puccinia triticina Erikss. populations as assessed by virulence profiles and SSR markers. Agric. Biol. 2018, 53, 85–95. [Google Scholar] [CrossRef]
- Gultyaeva, E.; Shaydayuk, E.; Kazartsev, I.; Akhmetova, A.; Kosman, E. Microsatellite analysis of Puccinia triticina from Triticum and Aegilops hosts. Aust. Plant. Path. 2018, 47, 163–170. [Google Scholar] [CrossRef]
- Gad, M.; Hao-xing, L.; Ming-ju, L.; El-Orabey, W.; Hasan, M. Evaluation of wheat genotypes to rust diseases (Puccinia spp.) under agroclimatic conditions of Egypt and China. J. Agric. Crop Res. 2019, 7, 170–180. [Google Scholar] [CrossRef]
- Yan, X.; Li, Z.; Yang, H.; Zhang, H.; Gebrewahid, T.; Yao, Z.; Li, D.; Zhou, Y. Analysis of Wheat Leaf Rust Resistance genes in 30 Important Wheat Cultivars. Sci. Agric. Sin. 2017, 50, 272–285. [Google Scholar] [CrossRef]
- Kokhmetova, A.M.; Atishova, M.N.; Galymbek, K. Identification of wheat germplasm resistant to leaf, stripe and stem rust using molecular markers. Bull. NAS RK 2020, 2, 45–52. [Google Scholar] [CrossRef]
- Mallick, N.; Jha, S.K.; Agarwal, P.; Mall, A.; M., N.; Kumar, S.; Choudhary, M.K.; Bansal, S.; Saharan, M.S.; Sharma, J.B.; et al. Marker-Assisted Improvement of Bread Wheat Variety HD2967 for Leaf and Stripe Rust Resistance. Plants 2022, 11, 1152. [Google Scholar] [CrossRef] [PubMed]
- Hanzalová, A.; Zelba, O. Leaf rust (Puccinia triticina Eriks) resistance genes in wheat cultivars registered in the Czech Republic. J. Plant Dis. Prot. 2022, 129, 1087–1093. [Google Scholar] [CrossRef]
- Liu, Y.; Gebrewahid, T.; Zhang, P.; Li, Z.; Liu, D. Identification of leaf rust resistance genes in common wheat varieties from China and foreign countries. J. Integr. Agric. 2021, 20, 1302–1313. [Google Scholar] [CrossRef]
- Gao, P.; Zhou, Y.; Gebrewahid, T.; Zhang, P.; Yan, X.; Li, X.; Yao, Z.; Li, Z.; Liu, D. Identification of known leaf rust resistance genes in common wheat cultivars from Sichuan province in China. Crop Prot. 2019, 115, 122–129. [Google Scholar] [CrossRef]
- Liu, D.; Yuan, C.; Singh, R.; Randhawa, M.; Bhavani, S.; Kumar, U.; Huerta-Espino, J.; Lagudah, E.; Lan, C. Stripe rust and leaf rust resistance in CIMMYT wheat line “Mucuy” is conferred by combinations of race-specific and adult-plant resistance loci. Front. Plant Sci. 2022, 13, 880138. [Google Scholar] [CrossRef]
- Martinez, F.; Niks, R.; Singh, R.; Rubiales, D. Characterization of Lr46, a gene conferring partial resistance to wheat leaf rust. Hereditas 2001, 135, 111–114. [Google Scholar] [CrossRef]
- Zhang, P.; Qi, A.; Zhou, Y.; Xia, X.; He, Z.; Li, Z.; Liu, D. Quantitative trait loci mapping of adult-plant resistance to leaf rust in a Fundulea 900 × ‘Thatcher’ wheat cross. Plant Breed. 2017, 136, 1–7. [Google Scholar] [CrossRef]
- Lagudah, E.; Krattinger, S.; Herrera-Foessel, S.; Singh, R.; Huerta-Espino, J.; Spielmeyer, W.; Brown-Guedira, G.; Selter, L.; Keller, B. Gene-specific markers for the wheat gene Lr34/Yr18/Pm38 which confers resistance to multiple fungal pathogens. Theor. Appl. Genet. 2009, 119, 889–898. [Google Scholar] [CrossRef]
- Pal, D.; Bhardwaj, S.; Sharma, P.; Sharma, D.; Khan, H.; Prabhu, K. Molecular marker aided selection for developing rust resistant genotypes by pyramiding Lr19/Sr25 and Yr15 in wheat (Triticum aestivum L.). Australas. Plant Pathol. 2020, 49, 631–640. [Google Scholar] [CrossRef]
- Zhang, P.; Gebrewahid, T.; Zhou, Y.; Li, Q.; Li, Z.; Liu, D. Seedling and adult plant resistance to leaf rust in 46 Chinese bread wheat landraces and 39 wheat lines with known Lr genes. J. Integr. Agric. 2019, 18, 1014–1023. [Google Scholar] [CrossRef]
- Kumar, K.; Jan, I.; Saripalli, G.; Sharma, P.; Mir, R.; Balyan, H.; Gupta, P. An update on resistance genes and their use in the development of leaf rust-resistant cultivars in wheat. Front. Genet. 2022, 13, 816057. [Google Scholar] [CrossRef] [PubMed]
- Hanzalová, A.; Šliková, S.; Hudcovicová, M. Virulence of wheat leaf rust (Puccinia triticina Eriks.) and Lr resistance genes in wheat cultivars in the Slovak Republic in the years 2016–2019. Cereal Res. Commun. 2022, 50, 281–286. [Google Scholar] [CrossRef]
- Nemati, Z.; Mostowfizadeh-Ghalamfarsa, R.; Dadkhodaie, A.; Mehrabi, R.; Steffenson, B. Virulence of Leaf Rust Physiological Races in Iran from 2010 to 2017. Plant Dis. 2020, 104, 363–372. [Google Scholar] [CrossRef]
- Goyeau, H.; Berder, J.; Czerepak, C.; Gautier, A.; Lanen, C.; Lannou, C. Low diversity and fast evolution in the population of Puccinia triticina causing durum wheat leaf rust in France from 1999 to 2009, as revealed by an adapted differential set. Plant Pathol. 2012, 61, 761–772. [Google Scholar] [CrossRef]
- Imbaby, I.; Mahmoud, M.; Hassan, M.; Abd-El-Aziz, A. Identification of Leaf Rust Resistance Genes in Selected Egyptian Wheat Cultivars by Molecular Markers. Sci. World J. 2014, 2014, 574285. [Google Scholar] [CrossRef]
- Gultyaeva, E.; Gannibal, P.; Shaydayuk, E. Long-Term Studies of Wheat Leaf Rust in the North-Western Region of Russia. Agriculture 2023, 13, 255. [Google Scholar] [CrossRef]
- Ivanova, V. Physiological specialization of wheat leaf rust (Puccinia triticina Eriks.) in Bulgaria. Rom. Agric. Res. 2022, 39, 366–374. [Google Scholar] [CrossRef]
- Kolmer, J.; Hughes, M. Physiologic Specialization of Puccinia triticina on wheat in the United States in 2016. Plant Dis. 2018, 102, 1066–1071. [Google Scholar] [CrossRef]
- Tomkowiak, A.; Bobrowska, R.; Kwiatek, M.; Spychala, J.; Kuczynski, J.; Tyczewska, A.; Kowalczewski, P.; Weigt, D.; Kosiada, T. Analysis of miRNA expression associated with gene Lr34 responsible for resistance mechanisms to wheat leaf rust. Pak. J. Bot. 2023, 55, 379–385. [Google Scholar] [CrossRef]
- Tomkowiak, A.; Jędrzejewski, T.; Spychała, J.; Kuczyński, J.; Kwiatek, M.; Tyczewska, A.; Skowrońska, R.; Twardowski, T. Analysis of miRNA expression associated with the Lr46 gene responsible for APR resistance in wheat (Triticum aestivum L.). J. Appl. Genet. 2020, 61, 503–511. [Google Scholar] [CrossRef] [PubMed]
- Mikhailova, L.A.; Gultyaeva, E.I.; Mironenko, N.V. Methods for studying the structure of populations of the leaf rust causative agent. In Collection of Guidelines on Plant Protection; All-Russia Institute of Plant Protection: St. Petersburg, Russia, 1998; pp. 105–126. (In Russian) [Google Scholar]
- Lind, V.; Gultyaeva, E. Virulence of Puccinia triticina on winter wheat in Germany and the European regions of Russian Federation. J. Phytopathol. 2007, 155, 13–21. [Google Scholar] [CrossRef]
- Roelfs, A.; Singh, R.; Saari, E. Rust Diseases of Wheat: Concept and Methods of Disease Management; CIMMYT: Veracruz, Mexico, 1992. [Google Scholar]
- Long, D.; Kolmer, J. A North American System of Nomenclature for Puccinia triticina. Phytopathology 1989, 79, 525–529. [Google Scholar] [CrossRef]
- Gultyaeva, E.; Shaydayuk, E.; Kosman, E. Regional and temporal differentiation of virulence phenotypes of Puccinia triticina from common wheat in Russia during the period 2001–2018. Plant. Pathol. 2020, 69, 860–871. [Google Scholar] [CrossRef]
- Mains, E.; Jackson, H. Physiologic specialization in the leaf rust of wheat, Puccinia triticina. Phytopathology 1926, 16, 89–120. [Google Scholar]
- Dospekhov, B.A. Methods of Field Experience (With the Basics of Statistical Processing of Research Results), 5th ed.; Kolos: Kovalivka, Ukraine, 1985. [Google Scholar]
- Zadoks, J.C.; Chang, T.T.; Konzak, C.F. A decimal code for the growth stages of cereals. Weed Res. 1974, 14, 415–421. [Google Scholar] [CrossRef]
- Peterson, R.F.; Campbell, A.; Hannah, A. A Diagrammatic Scale for Estimating Rust Intensity on Leaves and Stems of Cereals. Can. J. Res. 1948, 26, 496–500. [Google Scholar] [CrossRef]
- AACC (American Association of Cereal Chemists). International Approved Methods of the American Association of Cereal Chemists, 10th ed.; The Association of Cereal Chemists: St. Paul, MN, USA, 2000. [Google Scholar]
- Saari, E.; Wilcoxson, R. Plant disease situation of high-yielding durum wheat in Asia and Africa. Annu. Rev. Phytopathol. 1974, 2, 49–68. [Google Scholar] [CrossRef]
- Wilcoxson, R.; Skovmand, B.; Atif, A. Evaluation of wheat cultivars ability to retard development of stem rust. Ann Appl Biol. 1974, 80, 275–281. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; The R Foundation for Statistical Computing: Vienna, Austria, 2018; Available online: http://www.R-project.org/ (accessed on 21 April 2023).
- Riede, C.; Anderson, J. Linkage of RFLP markers to an aluminum tolerance gene in wheat. Crop Sci. 1996, 36, 905–909. [Google Scholar] [CrossRef]
- Chen, X.; Line, R.; Leung, H. Genome scanning for resistance-gene analogs in rice, barley, and wheat by high-resolution electrophoresis. Theor. Appl. Genet. 1998, 97, 345–355. [Google Scholar] [CrossRef]
- Schachermayr, G.; Feuillet, C.; Keller, B. Molecular markers for the detection of the wheat leaf rust resistance gene Lr10 in diverse genetic backgrounds. Mol. Breed. 1997, 3, 65–74. [Google Scholar] [CrossRef]
- Wu, H.; Kang, Z.; Li, X.; Li, Y.; Li, Y.; Wang, S.; Liu, D. Identification of Wheat Leaf Rust Resistance Genes in Chinese Wheat Cultivars and the Improved Germplasms. Plant Dis. 2020, 104, 2669–2680. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Yan, X.; Gebrewahid, T.; Zhou, Y.; Yang, E.; Xia, X.; He, Z.; Li, Z.; Liu, D. Genome-wide association mapping of leaf rust and stripe rust resistance in wheat accessions using the 90K SNP array. Theor. Appl. Genet. 2021, 134, 1233–1251. [Google Scholar] [CrossRef] [PubMed]

| Factor | SS | df | MS | F-Value | hb2, % |
|---|---|---|---|---|---|
| Genotype | 171.84 | 69 | 2.49 | 13.19 *** | 0.90 |
| Race | 4.46 | 1 | 4.46 | 23.63 *** | |
| Residuals | 13.03 | 69 | 0.19 | ||
| Total | 189.34 | 139 |
| Cultivar Name | Leaf Rust Severity 2021 | AUDPC | φ, % | Leaf Rust Severity 2022 | AUDPC | φ, % | Reaction to Infection with Races P. triticina | Lr Gene (s) Detected Based on Linked Markers | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1st Score | 2nd Score | 3rd Score | 1st Score | 2nd Score | 3rd Score | MKTKQ | TJTTR | ||||||
| Alatau | 0 | 5R | 10MR | 30 | 3 | 0 | 10MR | 20MS | 120 | 10 | 2 | 3 | Lr46 |
| Almaly | 10MR | 15MR | 20MR | 120 | 12 | 10MS | 20MS | 50S | 450 | 38 | 2 | 3 | Lr34 |
| Aliya | 5MR | 10MR | 20MS | 130 | 13 | 30MS | 50S | 70S | 970 | 83 | 2 | 2 | Lr10, Lr28, Lr37 |
| Azharly | 5MR | 5MR | 30MS | 150 | 15 | 0 | 0 | 10MR | 20 | 2 | 0 | 0 | Lr46 |
| Akbidaj | 5MR | 15MR | 30MS | 190 | 19 | 10R | 30MS | 50S | 500 | 43 | 3 | 4 | Lr46 |
| Batyr | 0 | 10MR | 30MS | 160 | 16 | 0 | 10MR | 10MR | 60 | 5 | 3 | 3 | Lr46 |
| Egemen 20 | 10MR | 20MS | 30MS | 300 | 29 | 0 | 0 | 0 | 0 | 0 | 2 | 2 | - |
| Farabi | 10MR | 20MS | 30MS | 300 | 29 | 0 | 10MS | 30MS | 200 | 17 | 3 | 3 | Lr46 |
| Mataj | 10MR | 30MS | 40S | 460 | 45 | 0 | 5R | 20MS | 90 | 8 | 2 | 2 | Lr9, Lr37, Lr46 |
| Kyzyl bidaj | 0 | 10MR | 30MS | 160 | 16 | 20MS | 30MS | 40S | 520 | 44 | 4 | 4 | Lr37, Lr19 |
| Keremet | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | Lr19, Lr26, Lr37, Lr46 |
| Koksu | 10MR | 10MR | 30MS | 180 | 18 | 0 | 0 | 10MS | 40 | 3 | 2 | 2 | Lr46 |
| Kokbidaj | 5R | 5R | 20MS | 95 | 9 | 0 | 0 | 10MR | 20 | 2 | 2 | 3 | Lr37 |
| Karaspan | 5R | 20MS | 40S | 365 | 36 | 20MS | 30MS | 50S | 570 | 49 | 4 | 4 | Lr34 |
| Karlygash | 10MR | 10MR | 40MS | 220 | 22 | 0 | 0 | 0 | 0 | 0 | 2 | 3 | Lr34 |
| Kazakhstanskaya 10 | 5MR | 20MS | 30MS | 290 | 28 | 0 | 5MR | 30MS | 140 | 12 | 2 | 3 | - |
| Progress | 10MS | 30MS | 50S | 530 | 52 | 5R | 20MS | 40S | 365 | 31 | 4 | 4 | - |
| Prezident | 10MS | 30MS | 30MS | 400 | 39 | 0 | 10MS | 20MS | 160 | 14 | 2 | 3 | - |
| Raminal | 10R | 10MR | 30MS | 170 | 17 | 0 | 10R | 10R | 30 | 3 | 2 | 2 | Lr9, Lr34 |
| Rasad | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 2 | Lr34, Lr37, Lr46 |
| Rausin | 20MS | 50S | 70S | 930 | 91 | 20MS | 40S | 40S | 680 | 58 | 4 | 4 | - |
| Reke | 10MR | 20MR | 30MS | 220 | 22 | 10MR | 20MS | 50S | 430 | 37 | 4 | 4 | Lr28, Lr37, Lr46 |
| Zhalyn | 10MS | 30MS | 50S | 530 | 52 | 10MR | 20MS | 30MS | 300 | 26 | 4 | 4 | - |
| Yuzhnaya 12 | 0 | 20MS | 40S | 360 | 35 | 20MS | 30MS | 50S | 570 | 49 | 3 | 4 | Lr9 |
| Pirotriks 50 | 10MS | 30MS | 40MS | 440 | 43 | 10MS | 20MS | 40S | 400 | 34 | 3 | 4 | Lr28 |
| Daulet | 20MS | 60S | 60S | 980 | 96 | 10MS | 30MS | 50S | 530 | 45 | 4 | 4 | - |
| Konditerskaya | 10MR | 10MS | 40MS | 260 | 25 | 0 | 10MS | 30MS | 200 | 17 | 2 | 3 | Lr28 |
| 428/MK-122A-1 | 5MR | 10MS | 40MS | 250 | 25 | 0 | 10MS | 30MS | 200 | 17 | 3 | 4 | - |
| Steklovidnaya 24 | 0 | 0 | 30MS | 120 | 12 | 0 | 20MS | 30S | 310 | 26 | 3 | 2 | - |
| Sultan 2 | 5R | 20MS | 60S | 465 | 46 | 0 | 20MS | 40S | 360 | 31 | 4 | 4 | - |
| Naz/Immun 78 | 20MS | 40S | 60S | 780 | 76 | 20MS | 50S | 70S | 930 | 79 | 4 | 4 | - |
| Naz/GF 55-2 | 5R | 5R | 5R | 20 | 2 | 0 | 0 | 0 | 0 | 0 | 2 | 1 | Lr34 |
| Naz/GF 55-3 | 0 | 10MR | 30MS | 160 | 16 | 10MR | 30MS | 50S | 510 | 44 | 4 | 3 | Lr26, Lr68 |
| Naz/GF 55-5 | 0 | 20MS | 30MS | 280 | 27 | 10MR | 40S | 40S | 620 | 53 | 3 | 3 | - |
| Yr/Octyabrina | 10MR | 30MS | 40MS | 420 | 41 | 20MS | 20MS | 50S | 490 | 42 | 4 | 4 | Lr10 |
| 425/Obri | 0 | 10MR | 40MS | 200 | 20 | 10MR | 30MS | 50S | 510 | 44 | 3 | 4 | Lr28 |
| Alihan | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 2 | Lr37, Lr68 |
| Anar | 0 | 5R | 30MS | 130 | 13 | 0 | 10R | 30MS | 140 | 12 | 2 | 3 | Lr9 |
| Derbes | 20MS | 40S | 50S | 730 | 72 | 20MS | 40S | 40S | 680 | 58 | 4 | 4 | - |
| Krasnovodapadskaya 210 | 0 | 30MS | 40MS | 400 | 39 | 10MS | 30MS | 30S | 430 | 37 | 3 | 4 | Lr37 |
| Almatinskaya polukarlikovaya/Progress | 10MS | 30MS | 30MS | 400 | 39 | 0 | 0 | 10R | 10 | 1 | 2 | 2 | - |
| Almaly/Orbij | 5MR | 10MS | 30MS | 210 | 21 | 0 | 0 | 0 | 0 | 0 | 1 | 2 | Lr9, Lr34, Lr68 |
| Gozgon | 0 | 0 | 5R | 5 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 2 | - |
| Bunyodkor | 0 | 20MS | 30S | 310 | 30 | 10MR | 20MS | 30S | 330 | 28 | 1 | 1 | Lr26 |
| Faravon | 0 | 5MR | 30MS | 140 | 14 | 0 | 0 | 5MR | 10 | 1 | 2 | 3 | - |
| Hazrati Bashir | 15MR | 30MS | 40S | 470 | 46 | 5MR | 20MS | 40S | 370 | 32 | 3 | 3 | Lr10 |
| Hisorok | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | Lr9, Lr10, Lr19, Lr37 |
| Layagatlii 80 | 0 | 10MS | 30MS | 200 | 20 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | Lr10, Lr26 |
| Shafag 2 | 0 | 10MS | 60S | 380 | 37 | 0 | 10MS | 30MS | 200 | 17 | 0 | 0 | Lr26 |
| Egana | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 1 | Lr9, Lr10, Lr34 |
| 415-SP-2 | 0 | 5MR | 5MR | 21 | 2 | 0 | 0 | 10MR | 14 | 1 | 2 | 2 | - |
| 416-SP-2 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 2 | Lr34, Lr26 |
| 2-CP | 0 | 10MR | 20MS | 84 | 8 | 10MR | 10MS | 30MS | 154 | 13 | 4 | 4 | Lr34 |
| 3-CP | 0 | 0 | 20MS | 56 | 5 | 0 | 10MR | 20MS | 84 | 7 | 3 | 4 | Lr37 |
| 4-CP | 0 | 10MS | 10MS | 84 | 8 | 10MR | 30MS | 30MS | 266 | 23 | 3 | 3 | Lr34 |
| 5-CP | 0 | 5R | 10MS | 35 | 3 | 0 | 20MS | 30MS | 196 | 17 | 3 | 3 | Lr34 |
| 6-CP | 0 | 10MS | 10MS | 84 | 8 | 10MR | 10MS | 20MS | 126 | 11 | 1 | 2 | - |
| 7-CP | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | Lr37 |
| 8-CP | 0 | 20MS | 30S | 217 | 21 | 20MS | 40S | 60S | 546 | 47 | 3 | 4 | - |
| 9-CP | 10MS | 10MS | 30MS | 168 | 16 | 0 | 0 | 0 | 0 | 0 | 1 | 2 | Lr26, Lr37 |
| 10-CP | 0 | 0 | 10MS | 28 | 3 | 0 | 0 | 0 | 0 | 0 | 2 | 3 | - |
| 11-CP | 0 | 20MS | 30MS | 196 | 19 | 10MS | 10MS | 30MS | 168 | 14 | 4 | 4 | Lr37 |
| 12-CP | 0 | 0 | 0 | 0 | 0 | 0 | 20MS | 30MS | 196 | 17 | 3 | 3 | - |
| 13-CP | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | - |
| 14-CP | 0 | 10MS | 20MS | 112 | 11 | 0 | 10MR | 10MS | 56 | 5 | 2 | 2 | Lr26, Lr37 |
| 15-CP | 0 | 0 | 10MS | 28 | 3 | 0 | 20MS | 40MS | 224 | 19 | 3 | 3 | Lr34, Lr37 |
| 16-CP | 0 | 5R | 10MS | 35 | 3 | 10MS | 30MS | 40MS | 308 | 26 | 3 | 4 | Lr34 |
| 17-CP | 0 | 0 | 0 | 0 | 0 | 0 | 10MR | 20MR | 56 | 5 | 2 | 2 | Lr34, Lr37 |
| 18-CP | 10MS | 40S | 50S | 483 | 47 | 20MS | 50S | 50S | 581 | 50 | 4 | 4 | - |
| 19-CP | 10MS | 10MS | 30MS | 168 | 16 | 10MS | 20MS | 40MS | 252 | 22 | 3 | 4 | Lr34 |
| Controls | |||||||||||||
| Morocco | 30MS | 50S | 80S | 1020 | 100 | 30MS | 60S | 90S | 1170 | 100 | - | ||
| Transfer/6*TC | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 10MR | 20 | 2 | Lr9 | ||
| TC*6/Exchange | 0 | 20MS | 40MS | 320 | 31 | 0 | 5R | 20MS | 90 | 8 | Lr10 | ||
| TC*7/Tr | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | Lr19 | ||
| TC*6/ST-1-25 | 0 | 10MR | 30MS | 160 | 16 | 0 | 0 | 20MS | 80 | 7 | Lr26 | ||
| CS2D-2M | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | Lr28 | ||
| TC*6/PI58548 | 0 | 0 | 20MS | 80 | 8 | 0 | 10MR | 20MS | 120 | 10 | Lr34 | ||
| TC*6/VPM | 0 | 20MS | 30MS | 280 | 27 | 0 | 0 | 10MR | 20 | 2 | Lr37 | ||
| Pavon 76 | 10MR | 20MS | 40MS | 340 | 33 | 0 | 20MS | 30MS | 280 | 24 | Lr46 | ||
| Parula | 0 | 0 | 10MR | 20 | 2 | 0 | 0 | 0 | 0 | 0 | Lr68 | ||
| Race | Virulence Formula (Avirulent/Virulent) | Response of Lr Genes (%) | |
|---|---|---|---|
| R | S | ||
| TJTTR | Lr9, Lr19, Lr26/Lr1, Lr2a, Lr2c, Lr3, Lr16, Lr24, Lr3ka, Lr11, Lr17, Lr30, Lr2b, Lr3bg, Lr14a, Lr14b, Lr15, Lr18, Lr20 | 15 | 85 |
| MKTKQ | Lr2a, Lr2b, Lr2c, Lr9, Lr19, Lr20/Lr1, Lr3, Lr16, Lr24, Lr26, Lr3ka, Lr11, Lr17, Lr30, Lr3bg, Lr14a, Lr14b, Lr15, Lr18 | 30 | 70 |
| Year | Month | Temperature (°C) | Monthly Rainfalls (mm) | Average Relative Humidity (%) |
|---|---|---|---|---|
| 2021 | April | 12.5 | 54 | 50 |
| May | 19.5 | 70 | 51 | |
| June | 23.0 | 20 | 38 | |
| 2022 | April | 16.7 | 45 | 54 |
| May | 19.0 | 142 | 65 | |
| June | 24.3 | 36 | 49 |
| Gen | Chr | Type of Marker | Primer Name | Sequence of Primers 5′-3′ | Anneling t°C | Fragmet Size, b.p. | Reference |
|---|---|---|---|---|---|---|---|
| Lr9 | 6BL | STS | J13-1 | 5′-CCACACTACCCCAAAGAGACG-3′ | 62 | 1100 | [51] |
| J13-2 | 5′-TCCTTTTATTCCGCACGCCGG-3′ | ||||||
| Lr10 | 1AS | STS | F1.2245 | 5′-GTGTAATGCATGCAGGTTCC-3′ | 57 | 310 | [104] |
| Lr10-6/r2 | 5′-AGGTGTGAGTGAGTTATGTT-3′ | ||||||
| Lr19 | 7AL | STS | Psy1-EF2 | 5′-CAAGTTCCCCATAGATATTCAG-3′ | 63 | 191 | [53] |
| Psy1-ER4 | 5′-AGAGAAAACCATTGCATCTGTA-3′ | ||||||
| Lr26 | 1BL | STS | Iag 95 | 5′-CTCTGTGGATAGTTACTTGATCGA-3′ 5′-CCTAGAACATGCATGGCTGTTACA-3′ | 55 | 1100 | [55] |
| Lr28 | 4AL | SSR | WMC 313 | 5′-CCCGGCATAAGTCTATGGTT-3 5′-CAATGAATGAGATACGTGAA-3′ | 51 | 320 | [56] |
| Lr34 | 7DS | STS | csLV34 | 5′-GTTGGTTAAGACTGGTGATGG-3′ 5′-TGCTTGCTATTGCTGAATAGT-3′ | 55 | +150 | [43] |
| −229 | |||||||
| Lr37 | 2AS | CAPS | Uric | 5′-GGTCGCCCTGGCTTGCACCT-3′ | 64 | 285 | [44] |
| Ln2 | 5′-TGCAGCTACAGCAGTATGTACACAAAA-3′ | ||||||
| Lr46 | 1BL | SSR | Wmc44 | 5′-GGT CTT CTG GGC TTT GAT CCT G-3′ 5′-GTT GCT AGG GAC CCG TAG TGG-3′ | 61 | 242 | [58] |
| Lr68 | 7BL | STS | csGS | 5′-AAG ATT GTT CAC AGA TCC ATG TCA-3′ 5′-GAG TAT TCC GGC TCA AAA AGG-3′ | 60 | 385 | [60] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Malysheva, A.; Kokhmetova, A.; Urazaliev, R.; Kumarbayeva, M.; Keishilov, Z.; Nurzhuma, M.; Bolatbekova, A.; Kokhmetova, A. Phenotyping and Identification of Molecular Markers Associated with Leaf Rust Resistance in the Wheat Germplasm from Kazakhstan, CIMMYT and ICARDA. Plants 2023, 12, 2786. https://doi.org/10.3390/plants12152786
Malysheva A, Kokhmetova A, Urazaliev R, Kumarbayeva M, Keishilov Z, Nurzhuma M, Bolatbekova A, Kokhmetova A. Phenotyping and Identification of Molecular Markers Associated with Leaf Rust Resistance in the Wheat Germplasm from Kazakhstan, CIMMYT and ICARDA. Plants. 2023; 12(15):2786. https://doi.org/10.3390/plants12152786
Chicago/Turabian StyleMalysheva, Angelina, Alma Kokhmetova, Rakhym Urazaliev, Madina Kumarbayeva, Zhenis Keishilov, Makpal Nurzhuma, Ardak Bolatbekova, and Assiya Kokhmetova. 2023. "Phenotyping and Identification of Molecular Markers Associated with Leaf Rust Resistance in the Wheat Germplasm from Kazakhstan, CIMMYT and ICARDA" Plants 12, no. 15: 2786. https://doi.org/10.3390/plants12152786
APA StyleMalysheva, A., Kokhmetova, A., Urazaliev, R., Kumarbayeva, M., Keishilov, Z., Nurzhuma, M., Bolatbekova, A., & Kokhmetova, A. (2023). Phenotyping and Identification of Molecular Markers Associated with Leaf Rust Resistance in the Wheat Germplasm from Kazakhstan, CIMMYT and ICARDA. Plants, 12(15), 2786. https://doi.org/10.3390/plants12152786







