Plant-Produced Nanoparticles Based on Artificial Self-Assembling Peptide Bearing the Influenza M2e Epitope
Abstract
1. Introduction
2. Results
2.1. Composition of SAP-M2e Fusion Protein
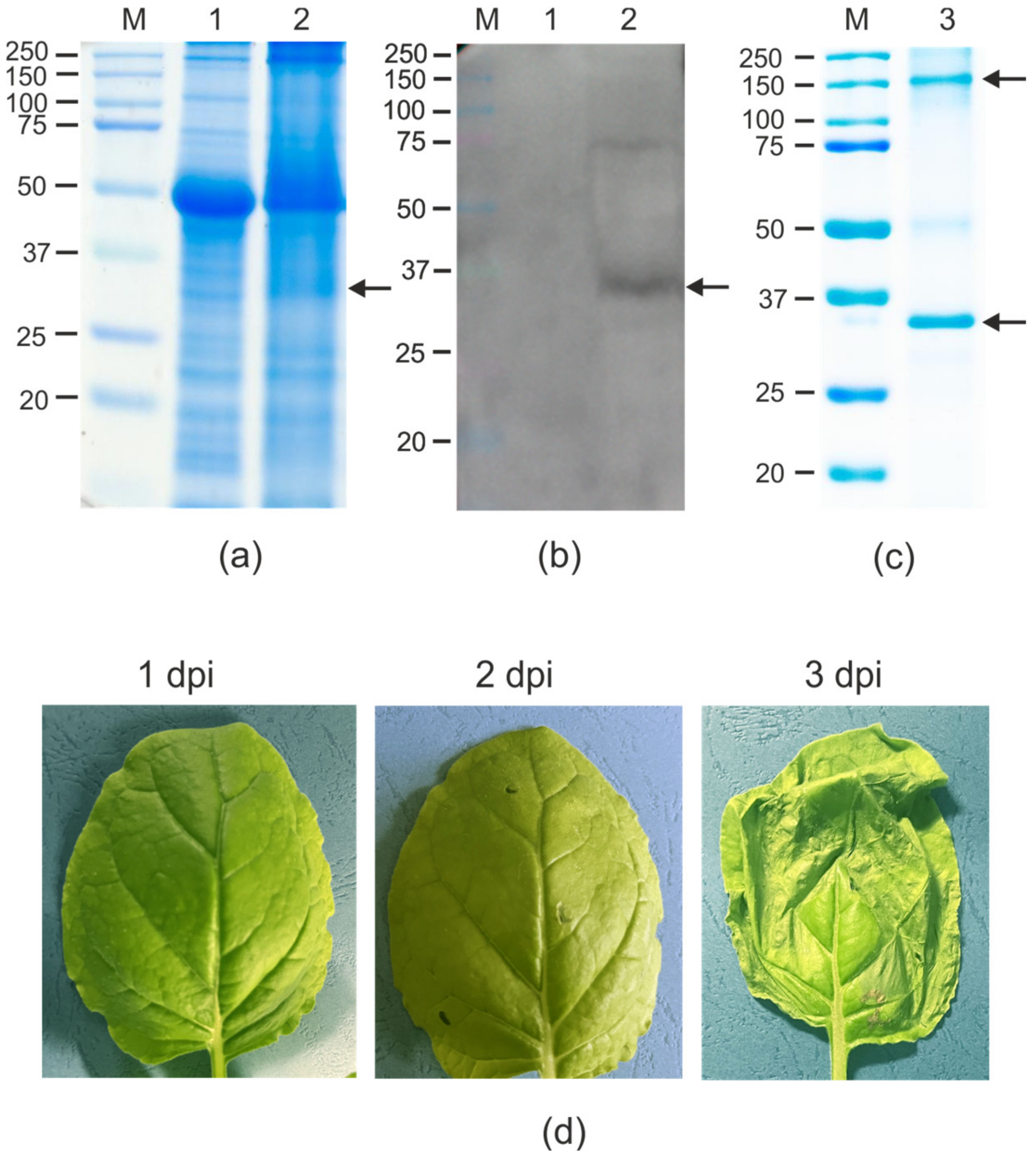
2.2. Expression and Purification of Recombinant 19S_SAP_Sp_4M2eh Protein
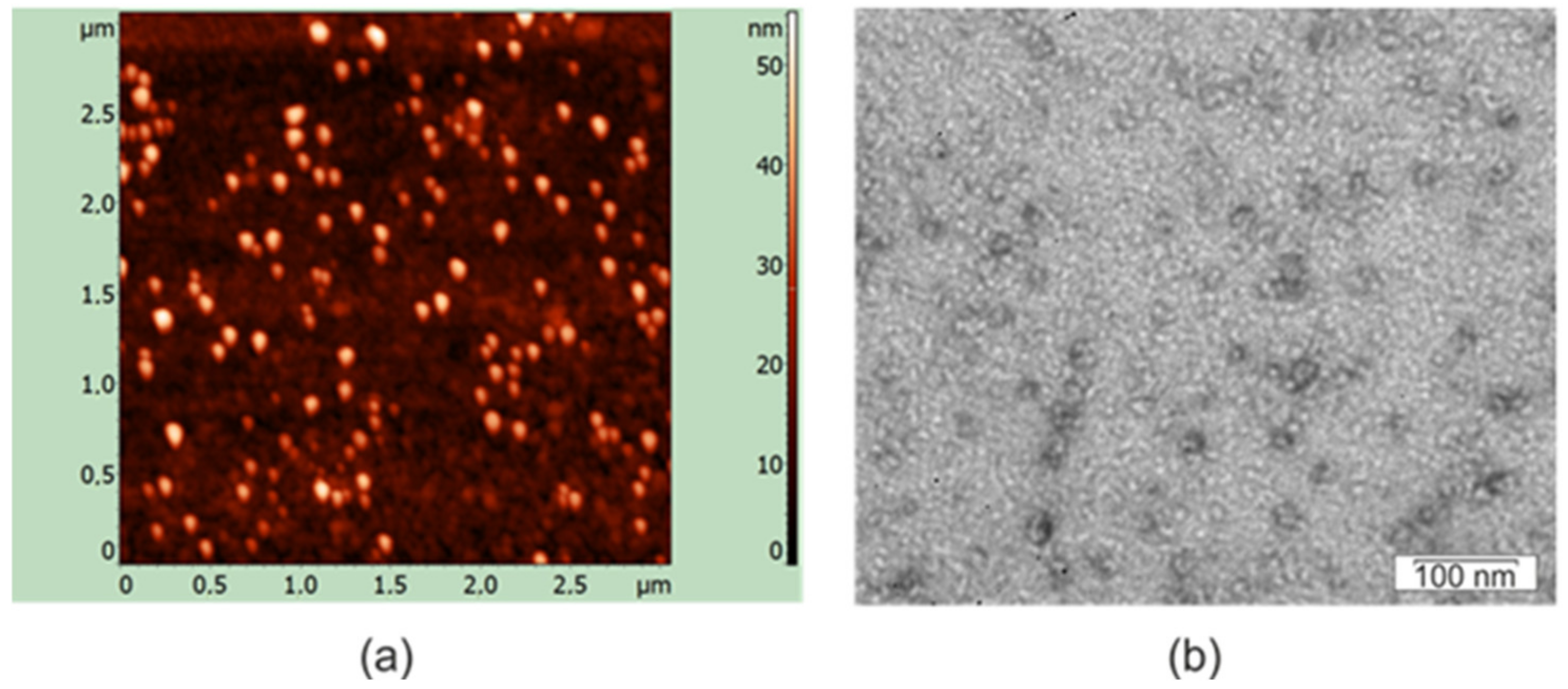
2.3. Protein Refolding and In Vitro Assembly of Nanoparticles
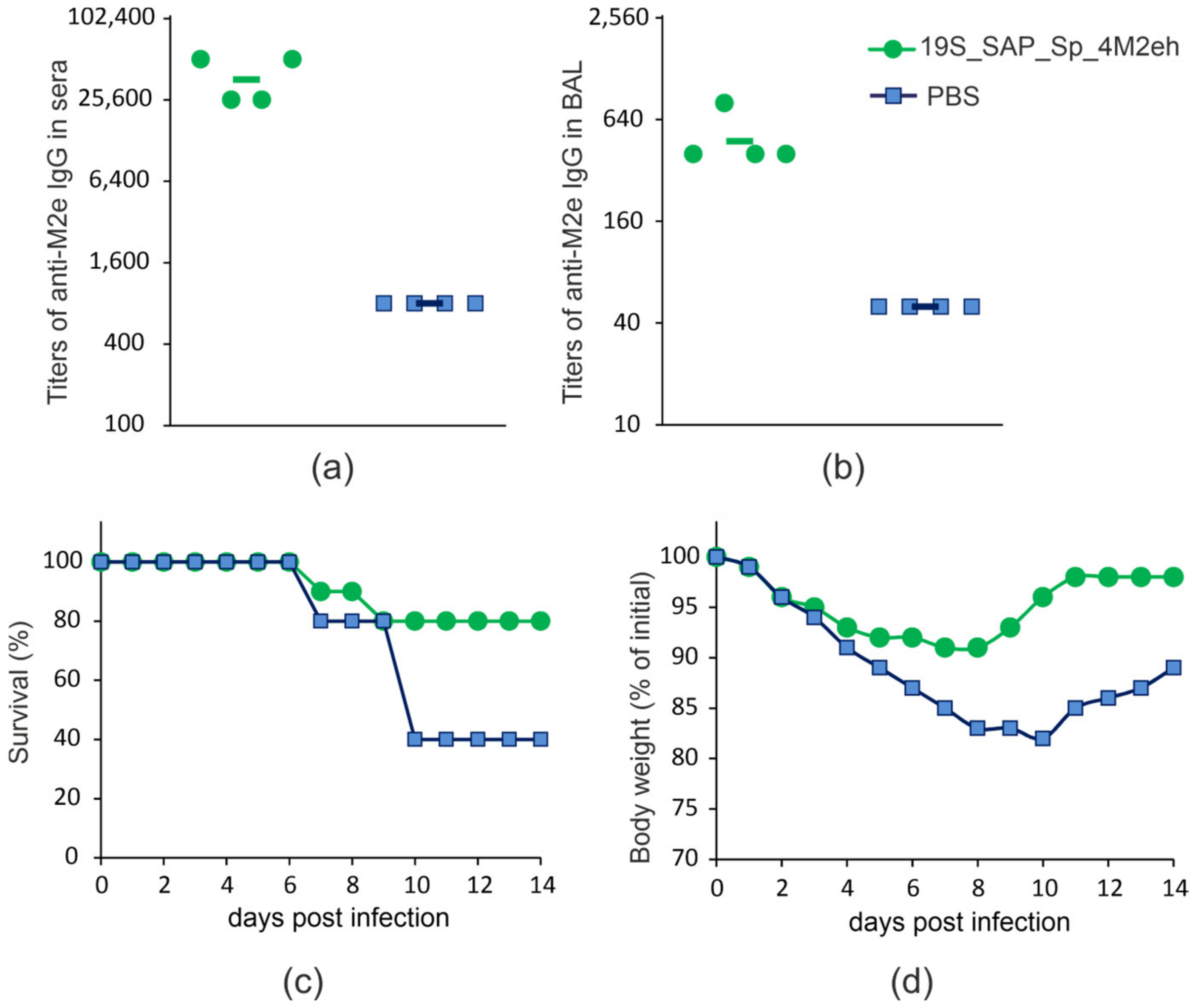
2.4. Immunogenicity and Protectivity of Plant-Produced 19S_SAP_Sp_4M2eh
3. Discussion
4. Materials and Methods
4.1. Structure of the SAP-Based Fusion Protein
4.2. Expression Vector
4.3. Agroinfiltration of N. benthamiana Leaves
4.4. SDS-PAGE and Western-Blotting of the Proteins Isolated from Plants
4.5. Isolation and Purification of Recombinant Proteins from Plant Biomass
4.6. Structural Analysis of Nanoparticles
4.7. Mouse Immunization
4.8. Antibody Detection by ELISA
4.9. Influenza Virus and Challenge
4.10. Ethics Statement
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chauhan, R.P.; Gordon, M.L. An overview of influenza A virus genes, protein functions, and replication cycle highlighting important updates. Virus Genes 2022, 58, 255–269. [Google Scholar] [CrossRef] [PubMed]
- Deng, L.; Cho, K.J.; Fiers, W.; Saelens, X. M2e-Based Universal Influenza A Vaccines. Vaccines 2015, 3, 105–136. [Google Scholar] [CrossRef] [PubMed]
- Nachbagauer, R.; Krammer, F. Universal influenza virus vaccines and therapeutic antibodies. Clin. Microbiol. Infect. 2017, 23, 222–228. [Google Scholar] [CrossRef] [PubMed]
- Fiers, W.; De Filette, M.; Birkett, A.; Neirynck, S.; Min Jou, W. A “universal” human influenza A vaccine. Virus Res. 2004, 103, 173–176. [Google Scholar] [CrossRef]
- Neirynck, S.; Deroo, T.; Saelens, X.; Vanlandschoot, P.; Jou, W.M.; Fiers, W. A universal influenza A vaccine based on the extracellular domain of the M2 protein. Nat. Med. 1999, 5, 1157–1163. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Zhang, M.; Mozdzanowska, K.; Zharikova, D.; Ho, H.; Wunner, W.; Couch, R.B.; Gerhard, W. Influenza A virus infection engenders a poor antibody response against the ectodomain of matrix protein 2. Virol. J. 2006, 3, 102. [Google Scholar] [CrossRef]
- Bachmann, M.F.; Jennings, G.T. Vaccine Delivery: A Matter of Size, Geometry, Kinetics and Molecular Patterns. Nat. Rev. Immunol. 2010, 10, 787–796. [Google Scholar] [CrossRef]
- Akagi, T.; Wang, X.; Uto, T.; Baba, M.; Akashi, M. Protein direct delivery to dendritic cells using nanoparticles based on amphiphilic poly(amino acid) derivatives. Biomaterials 2007, 28, 3427–3436. [Google Scholar] [CrossRef]
- Kalkanidis, M.; Pietersz, G.A.; Xiang, S.D.; Mottram, P.L.; Crimeen-Irwin, B.; Ardipradja, K.; Plebanski, M. Methods for nano-particle based vaccine formulation and evaluation of their immunogenicity. Methods 2006, 40, 20–29. [Google Scholar] [CrossRef]
- Minigo, G.; Scholzen, A.; Tang, C.K.; Hanley, J.C.; Kalkanidis, M.; Pietersz, G.A.; Apostolopoulos, V.; Plebanski, M. Poly-L-lysine-coated nanoparticles: A potent delivery system to enhance DNA vaccine efficacy. Vaccine 2006, 25, 1316–1327. [Google Scholar] [CrossRef]
- Mottram, P.L.; Leong, D.; Crimeen-Irwin, B.; Gloster, S.; Xiang, S.D.; Meanger, J.; Ghildyal, R.; Vardaxis, N.; Plebanski, M. Type 1 and 2 immunity following vaccination is influenced by nanoparticle size: Formulation of a model vaccine for respiratory syncytial virus. Mol. Pharm. 2007, 4, 73–84. [Google Scholar] [CrossRef] [PubMed]
- Reddy, S.T.; Rehor, A.; Schmoekel, H.G.; Hubbell, J.A.; Swartz, M.A. In vivo targeting of dendritic cells in lymph nodes with poly(propylene sulfide) nanoparticles. J. Control. Release 2006, 112, 26–34. [Google Scholar] [CrossRef] [PubMed]
- Jennings, G.T.; Bachmann, M.F. The coming of age of virus-like particle vaccines. Biol. Chem. 2008, 389, 521–536. [Google Scholar] [CrossRef] [PubMed]
- Kolpe, A.; Schepens, B.; Fiers, W.; Saelens, X. M2-based influenza vaccines: Recent advances and clinical potential. Expert. Rev. Vaccines 2017, 16, 123–136. [Google Scholar] [CrossRef]
- Mardanova, E.S.; Ravin, N.V. Plant-produced Recombinant Influenza A Vaccines Based on the M2e Peptide. Curr. Pharm. Des. 2018, 24, 1317–1324. [Google Scholar] [CrossRef]
- Carascal, M.B.; Pavon, R.D.N.; Rivera, W.L. Recent Progress in Recombinant Influenza Vaccine Development Toward Heterosubtypic Immune Response. Front. Immunol. 2022, 13, 878943. [Google Scholar] [CrossRef]
- Raman, S.; Machaidze, G.; Lustig, A.; Aebi, U.; Burkhard, P. Structure-based design of peptides that self-assemble into regular polyhedral nanoparticles. Nanomedicine 2006, 2, 95–102. [Google Scholar] [CrossRef]
- Malashkevich, V.N.; Kammerer, R.A.; Efimov, V.P.; Schulthess, T.; Engel, J. The crystal structure of a five-stranded coiled coil in COMP: A prototype ion channel? Science 1996, 274, 761–765. [Google Scholar] [CrossRef]
- Burkhard, P.; Meier, M.; Lustig, A. Design of a minimal protein oligomerization domain by a structural approach. Protein Sci. 2000, 9, 2294–2301. [Google Scholar] [CrossRef]
- Kaba, S.A.; Brando, C.; Guo, Q.; Mittelholzer, C.; Raman, S.; Tropel, D.; Aebi, U.; Burkhard, P.; Lanar, D.E. A nonadjuvanted polypeptide nanoparticle vaccine confers long-lasting protection against rodent malaria. J. Immunol. 2009, 183, 7268–7277. [Google Scholar] [CrossRef]
- Kaba, S.A.; McCoy, M.E.; Doll, T.A.; Brando, C.; Guo, Q.; Dasgupta, D.; Yang, Y.; Mittelholzer, C.; Spaccapelo, R.; Crisanti, A.; et al. Protective antibody and CD8+ T-cell responses to the Plasmodium falciparum circumsporozoite protein induced by a nanoparticle vaccine. PLoS ONE 2012, 7, e48304. [Google Scholar] [CrossRef] [PubMed]
- Babapoor, S.; Neef, T.; Mittelholzer, C.; Girshick, T.; Garmendia, A.; Shang, H.; Khan, M.I.; Burkhard, P. A Novel Vaccine Using Nanoparticle Platform to Present Immunogenic M2e against Avian Influenza Infection. Influenza Res. Treat. 2011, 2011, 126794. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Helal, Z.; Ladman, B.; Karch, C.; Gelb, J.; Burkhard, P.; Khan, M.I. Nanoparticle Vaccine for Avian Influenza Virus: A Challenge Study against Highly Pathogenic H5N2 Subtype. J. Virol. Antivir. Res. 2018, 7, 1. [Google Scholar] [CrossRef]
- Pimentel, T.A.; Yan, Z.; Jeffers, S.A.; Holmes, K.V.; Hodges, R.S.; Burkhard, P. Peptide nanoparticles as novel immunogens: Design and analysis of a prototypic severe acute respiratory syndrome vaccine. Chem. Biol. Drug. Des. 2009, 73, 53–61. [Google Scholar] [CrossRef]
- Wahome, N.; Pfeiffer, T.; Ambiel, I.; Yang, Y.; Keppler, O.T.; Bosch, V.; Burkhard, P. Conformation-specific display of 4E10 and 2F5 epitopes on self-assembling protein nanoparticles as a potential HIV vaccine. Chem. Biol. Drug. Des. 2012, 80, 349–357. [Google Scholar] [CrossRef]
- Karch, C.P.; Paquin-Proulx, D.; Eller, M.A.; Matyas, G.R.; Burkhard, P.; Beck, Z. Impact of the expression system on the immune responses to self-assembling protein nanoparticles (SAPNs) displaying HIV-1 V1V2 loop. Nanomedicine 2020, 29, 102255. [Google Scholar] [CrossRef]
- El Bissati, K.; Zhou, Y.; Dasgupta, D.; Cobb, D.; Dubey, J.P.; Burkhard, P.; Lanar, D.E.; McLeod, R. Effectiveness of a novel immunogenic nanoparticle platform for Toxoplasma peptide vaccine in HLA transgenic mice. Vaccine 2014, 32, 3243–3248. [Google Scholar] [CrossRef]
- El Bissati, K.; Zhou, Y.; Paulillo, S.M.; Raman, S.K.; Karch, C.P.; Reed, S.; Estes, A.; Estes, A.; Lykins, J.; Burkhard, P.; et al. Engineering and characterization of a novel Self Assembling Protein for Toxoplasma peptide vaccine in HLA-A*11:01, HLA-A*02:01 and HLA-B*07:02 transgenic mice. Sci. Rep. 2020, 10, 16984. [Google Scholar] [CrossRef]
- Li, J.; Helal, Z.H.; Karch, C.P.; Mishra, N.; Girshick, T.; Garmendia, A.; Burkhard, P.; Khan, M.I. A self-adjuvanted nanoparticle based vaccine against infectious bronchitis virus. PLoS ONE 2018, 13, e0203771. [Google Scholar] [CrossRef]
- Zykova, A.A.; Blokhina, E.A.; Stepanova, L.A.; Shuklina, M.A.; Tsybalova, L.M.; Kuprianov, V.V.; Ravin, N.V. Nanoparticles based on artificial self-assembling peptide and displaying M2e peptide and stalk HA epitopes of influenza A virus induce potent humoral and T-cell responses and protect against the viral infection. Nanomedicine 2022, 39, 102463. [Google Scholar] [CrossRef]
- Liu, H.; Timko, M.P. Improving Protein Quantity and Quality-The Next Level of Plant Molecular Farming. Int. J. Mol. Sci. 2022, 23, 1326. [Google Scholar] [CrossRef]
- Zahmanova, G.; Takova, K.; Valkova, R.; Toneva, V.; Minkov, I.; Andonov, A.; Lukov, G.L. Plant-Derived Recombinant Vaccines against Zoonotic Viruses. Life 2022, 12, 156. [Google Scholar] [CrossRef] [PubMed]
- Shanmugaraj, B.; Bulaon, C.J.I.; Phoolcharoen, W. Plant Molecular Farming: A Viable Platform for Recombinant Biopharmaceutical Production. Plants 2020, 9, 842. [Google Scholar] [CrossRef] [PubMed]
- Mahmood, N.; Nasir, S.B.; Hefferon, K. Plant-Based Drugs and Vaccines for COVID-19. Vaccines 2020, 9, 15. [Google Scholar] [CrossRef] [PubMed]
- Dubey, K.K.; Luke, G.A.; Knox, C.; Kumar, P.; Pletschke, B.I.; Singh, P.K.; Shukla, P. Vaccine and Antibody Production in Plants: Developments and Computational Tools. Brief Funct. Genomics 2018, 17, 295–307. [Google Scholar] [CrossRef]
- Bhuiyan, F.R.; Howlader, S.; Raihan, T.; Hasan, M. Plants Metabolites: Possibility of Natural Therapeutics Against the COVID-19 Pandemic. Front. Med. 2020, 7, 444. [Google Scholar] [CrossRef] [PubMed]
- Schillberg, S.; Finnern, R. Plant molecular farming for the production of valuable proteins—Critical evaluation of achievements and future challenges. J. Plant. Physiol. 2021, 258–259, 153359. [Google Scholar] [CrossRef]
- Fischer, R.; Buyel, J.F. Molecular farming—The slope of enlightenment. Biotechnol. Adv. 2020, 40, 107519. [Google Scholar] [CrossRef]
- Sainsbury, F. Innovation in plant-based transient protein expression for infectious disease prevention and preparedness. Curr. Opin. Biotechnol. 2020, 61, 110–115. [Google Scholar] [CrossRef]
- Thuenemann, E.C.; Byrne, M.J.; Peyret, H.; Saunders, K.; Castells-Graells, R.; Ferriol, I.; Santoni, M.; Steele, J.F.C.; Ranson, N.A.; Avesani, L.; et al. A Replicating Viral Vector Greatly Enhances Accumulation of Helical Virus-Like Particles in Plants. Viruses 2021, 13, 885. [Google Scholar] [CrossRef]
- Nosaki, S.; Miura, K. Transient expression of recombinant proteins in plants. Methods Enzymol. 2021, 660, 193–203. [Google Scholar] [CrossRef] [PubMed]
- Mardanova, E.S.; Blokhina, E.A.; Tsybalova, L.M.; Peyret, H.; Lomonossoff, G.P.; Ravin, N.V. Efficient transient expression of recombinant proteins in plants by the novel pEff vector based on the genome of potato virus X. Front. Plant. Sci. 2017, 8, 247. [Google Scholar] [CrossRef] [PubMed]
- Hager, K.J.; Perez Marc, G.; Gobeil, P.; Diaz, R.S.; Heizer, G.; Llapur, C.; Makarkov, A.I.; Vasconcellos, E.; Pillet, S.; Riera, F.; et al. Efficacy and safety of a recombinant plant-based adjuvanted COVID-19 vaccine. N. Engl. J. Med. 2022, 386, 2084–2096. [Google Scholar] [CrossRef]
- Ravin, N.V.; Blokhina, E.A.; Kuprianov, V.V.; Stepanova, L.A.; Shaldjan, A.A.; Kovaleva, A.A.; Tsybalova, L.M.; Skryabin, K.G. Development of a candidate influenza vaccine based on virus-like particles displaying influenza M2e peptide into the immunodominant loop region of hepatitis B core antigen: Insertion of multiple copies of M2e increases immunogenicity and protective efficiency. Vaccine 2015, 33, 3392–3397. [Google Scholar] [CrossRef]
- De Filette, M.; Min Jou, W.; Birkett, A.; Lyons, K.; Schultz, B.; Tonkyro, A.; Resch, S.; Fiers, W. Universal influenza A vaccine: Optimization of M2-based constructs. Virology 2005, 337, 149–161. [Google Scholar] [CrossRef]
- Arai, R.; Ueda, H.; Kitayama, A.; Kamiya, N.; Nagamune, T. Design of the linkers which effectively separate domains of a bifunctional fusion protein. Protein Eng. 2001, 14, 529–532. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef] [PubMed]
- Guan, Y.; Zhu, Q.; Huang, D.; Zhao, S.; Jan Lo, L.; Peng, J. An equation to estimate the difference between theoretically predicted and SDS PAGE-displayed molecular weights for an acidic peptide. Sci. Rep. 2015, 5, 13370. [Google Scholar] [CrossRef]
- Lomonossoff, G.P.; DAoust, M.-A. Plant-Produced Biopharmaceuticals: A Case of Technical Developments Driving Clinical Deployment. Science 2016, 353, 1237–1240. [Google Scholar] [CrossRef]
- Rybicki, E.P. Plant-Based Vaccines against Viruses. Virol. J. 2014, 11, 205. [Google Scholar] [CrossRef]
- Reed, J.; Osbourn, A. Engineering Terpenoid Production through Transient Expression in Nicotiana benthamiana. Plant. Cell. Rep. 2018, 37, 1431–1441. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, T.; Hoshikawa, K.; Ezura, K.; Okazawa, R.; Fujita, S.; Takaoka, M.; Mason, H.S.; Ezura, H.; Miura, K. Improvement of the transient expression system for production of recombinant proteins in plants. Sci. Rep. 2018, 8, 4755. [Google Scholar] [CrossRef] [PubMed]
- Zischewski, J.; Sack, M.; Fischer, R. Overcoming low yields of plant-made antibodies by a protein engineering approach. Biotechnol. J. 2016, 11, 107–116. [Google Scholar] [CrossRef]
- Mardanova, E.S.; Kotlyarov, R.Y.; Ravin, N.V. Rapid Transient Expression of Receptor-Binding Domain of SARS-CoV-2 and the Conserved M2e Peptide of Influenza A Virus Linked to Flagellin in Nicotiana benthamiana Plants Using Self-Replicating Viral Vector. Plants 2022, 11, 3425. [Google Scholar] [CrossRef]
- Saunders, K.; Thuenemann, E.C.; Shah, S.N.; Peyret, H.; Kristianingsih, R.; Lopez, S.G.; Richardson, J.; Lomonossoff, G.P. The Use of a Replicating Virus Vector for in Planta Generation of Tobacco Mosaic Virus Nanorods Suitable for Metallization. Front. Bioeng. Biotechnol. 2022, 10, 877361. [Google Scholar] [CrossRef] [PubMed]
- Zahmanova, G.G.; Mazalovska, M.; Takova, K.H.; Toneva, V.T.; Minkov, I.N.; Mardanova, E.S.; Ravin, N.V.; Lomonossoff, G.P. Rapid High-Yield Transient Expression of Swine Hepatitis E ORF2 Capsid Proteins in Nicotiana benthamiana Plants and Production of Chimeric Hepatitis E Virus-Like Particles Bearing the M2e Influenza Epitope. Plants 2020, 9, 29. [Google Scholar] [CrossRef]
- Mardanova, E.S.; Kotlyarov, R.Y.; Kuprianov, V.V.; Stepanova, L.A.; Tsybalova, L.M.; Lomonosoff, G.P.; Ravin, N.V. Rapid high-yield expression of a candidate influenza vaccine based on the ectodomain of M2 protein linked to flagellin in plants using viral vectors. BMC Biotechnol. 2015, 15, 42. [Google Scholar] [CrossRef]
- Mardanova, E.S.; Kotlyarov, R.Y.; Stuchinskaya, M.D.; Nikolaeva, L.I.; Zahmanova, G.; Ravin, N.V. High-Yield Production of Chimeric Hepatitis E Virus-Like Particles Bearing the M2e Influenza Epitope and Receptor Binding Domain of SARS-CoV-2 in Plants Using Viral Vectors. Int. J. Mol. Sci. 2022, 23, 15684. [Google Scholar] [CrossRef]
- Ravin, N.V.; Kotlyarov, R.Y.; Mardanova, E.S.; Kuprianov, V.V.; Migunov, A.I.; Stepanova, L.A.; Tsybalova, L.M.; Kiselev, O.I.; Skryabin, K.G. Plant-produced recombinant influenza vaccine based on virus-like HBc particles carrying an extracellular domain of M2 protein. Biochemistry 2012, 77, 33–40. [Google Scholar] [CrossRef]
- Robinson, C.R.; Sauer, R.T. Optimizing the Stability of Single-Chain Proteins by Linker Length and Composition Mutagenesis. Proc. Natl. Acad. Sci. USA 1998, 95, 5929–5934. [Google Scholar] [CrossRef]
- Kapila, J.; De Rycke, R.; Van Montagu, M.; Angenon, G. An Agrobacterium-mediated transient gene expression system for intact leaves. Plant. Sci. 1997, 122, 101–108. [Google Scholar] [CrossRef]
- Schroeder, A.B.; Dobson, E.T.A.; Rueden, C.T.; Tomancak, P.; Jug, F.; Eliceiri, K.W. The ImageJ ecosystem: Open-source software for image visualization, processing, and analysis. Protein Sci. 2021, 30, 234–249. [Google Scholar] [CrossRef] [PubMed]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Blokhina, E.A.; Mardanova, E.S.; Zykova, A.A.; Stepanova, L.A.; Shuklina, M.A.; Tsybalova, L.M.; Ravin, N.V. Plant-Produced Nanoparticles Based on Artificial Self-Assembling Peptide Bearing the Influenza M2e Epitope. Plants 2023, 12, 2228. https://doi.org/10.3390/plants12112228
Blokhina EA, Mardanova ES, Zykova AA, Stepanova LA, Shuklina MA, Tsybalova LM, Ravin NV. Plant-Produced Nanoparticles Based on Artificial Self-Assembling Peptide Bearing the Influenza M2e Epitope. Plants. 2023; 12(11):2228. https://doi.org/10.3390/plants12112228
Chicago/Turabian StyleBlokhina, Elena A., Eugenia S. Mardanova, Anna A. Zykova, Liudmila A. Stepanova, Marina A. Shuklina, Liudmila M. Tsybalova, and Nikolai V. Ravin. 2023. "Plant-Produced Nanoparticles Based on Artificial Self-Assembling Peptide Bearing the Influenza M2e Epitope" Plants 12, no. 11: 2228. https://doi.org/10.3390/plants12112228
APA StyleBlokhina, E. A., Mardanova, E. S., Zykova, A. A., Stepanova, L. A., Shuklina, M. A., Tsybalova, L. M., & Ravin, N. V. (2023). Plant-Produced Nanoparticles Based on Artificial Self-Assembling Peptide Bearing the Influenza M2e Epitope. Plants, 12(11), 2228. https://doi.org/10.3390/plants12112228






