Descriptive Genomic Analysis and Sequence Genotyping of the Two Papaya Species (Vasconcellea pubescens and Vasconcellea chilensis) Using GBS Tools
Abstract
:1. Introduction
2. Results
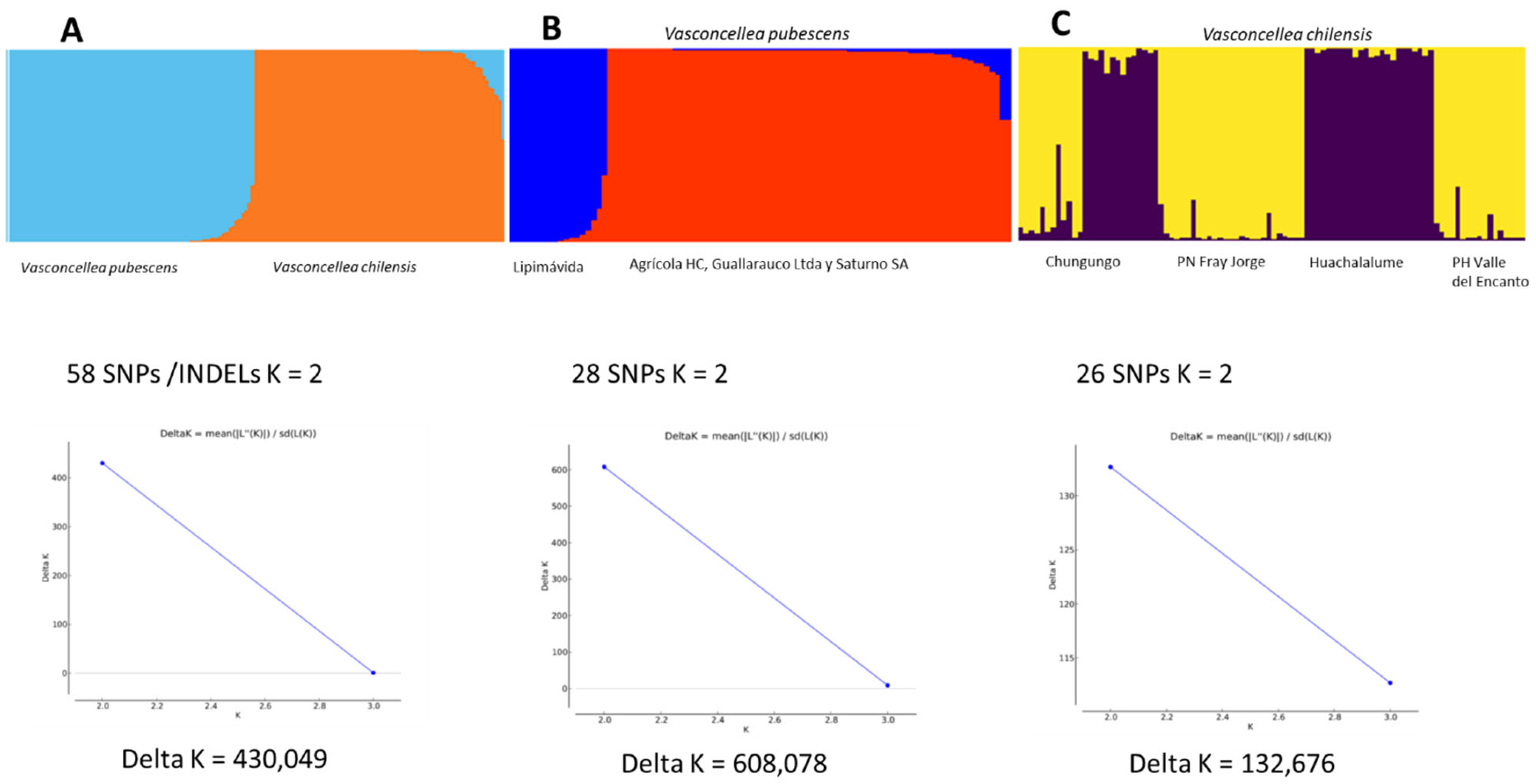
2.1. Genetic Diversity and Structure
2.2. Comparative Genomics
3. Discussion
4. Materials and Methods
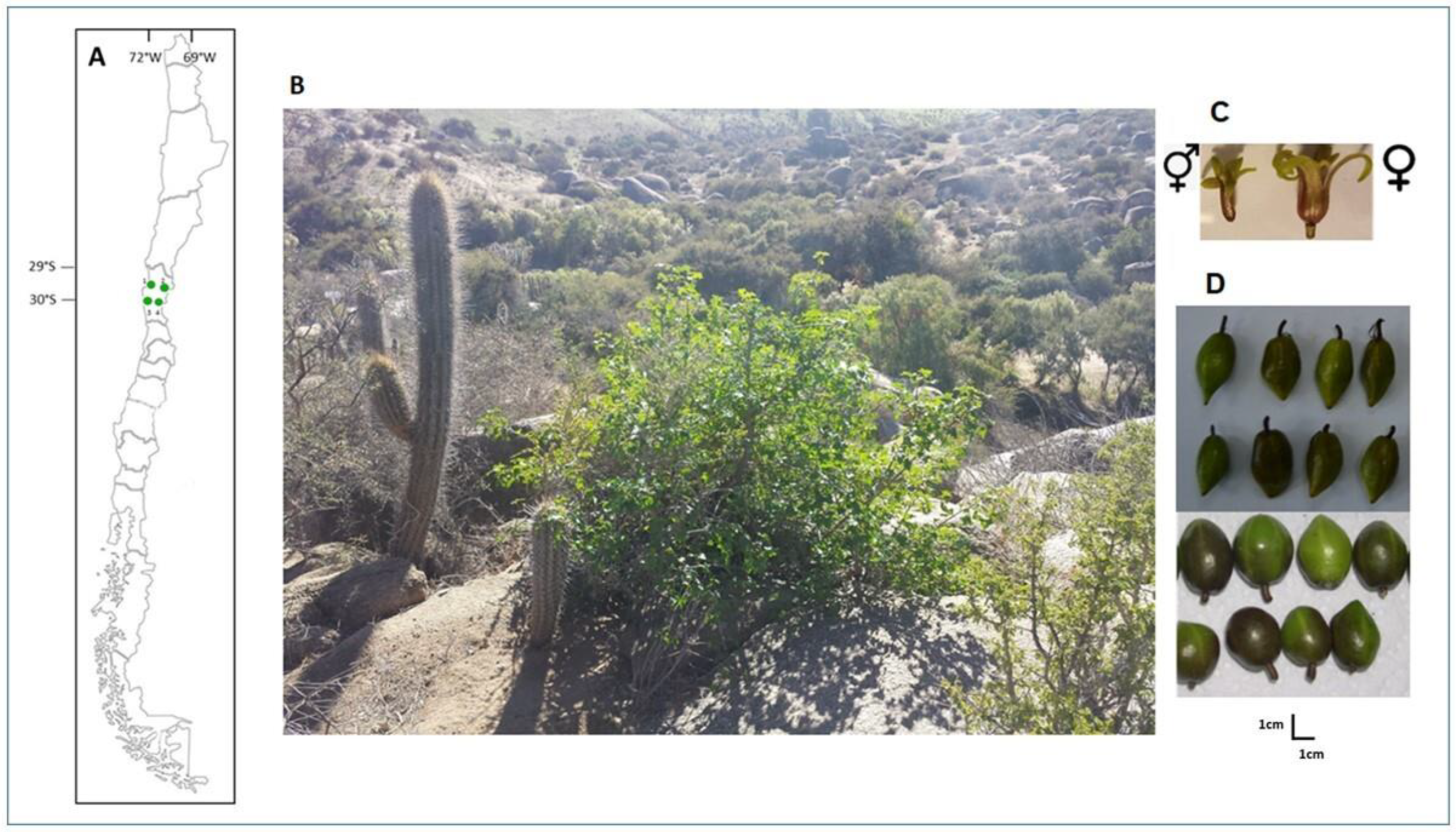
4.1. Plant Material
4.2. DNA Extraction and Genotyping by Sequencing (GBS)
4.3. Population Genetic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Moore, P.H.; Ming, R. Papaya Genome: A Model for Tropical Fruit Trees and Beyond. Trop. Plant Biol. 2008, 1, 179–180. [Google Scholar] [CrossRef]
- Badillo, V.M. Monografia de la Familia Caricaceae; Asociacion de Profesores: 1971. Badillo, V. M. Monografía de la Familia Caricaceae; Universidad Central de Venezuela: Maracay, Venezuela, 1971. [Google Scholar]
- Badillo, V.M. Caricaceae: Segundo Esquema; Revista de la Facultad de Agronomía (UCV), Alcance; Universidad Central de Venezuela, Facultad de Agronomía: Caracas, Venezuela, 1993; Volume 43, p. 111. [Google Scholar]
- Badillo, V.M.; Carica, L. Vasconcella St. Hil. (Caricaceae): Con La Rehabilitación de Este Último. Ernstia 2000, 10, 74–79. [Google Scholar]
- Badillo, V.M. Nota Correctiva Vasconcellea St. Hil. y No Vasconcella (Caricaceae). Ernstia 2001, 11, 75–76. [Google Scholar]
- Scheldeman, X.; Willemen, L.; Coppens d’Eeckenbrugge, G.; Romeijn-Peeters, E.; Restrepo, M.T.; Romero Motoche, J.; Jiménez, D.; Lobo, M.; Medina, C.I.; Reyes, C.; et al. Distribution, Diversity and Environmental Adaptation of Highland Papayas (Vasconcellea spp.) in Tropical and Subtropical America. In Plant Conservation and Biodiversity; Topics in Biodiversity and Conservation; Hawksworth, D.L., Bull, A.T., Eds.; Springer: Dordrecht, The Netherlands, 2007; pp. 293–310. ISBN 978-1-4020-6444-9. [Google Scholar]
- National Research Council. Lost Crops of the Incas: Little-Known Plants of the Andes with Promise for Worldwide Cultivation; The National Academies Press: Washington, DC, USA, 1999; ISBN 978-0-309-04264-2. [Google Scholar]
- Fabiane, R.C.; Pereira, T.N.S.; Hodnett, G.L.; Pereira, M.G.; Stelly, D.M. Fluorescent in Situ Hybridization of 18S and 5S RDNA in Papaya (Carica Papaya l.) and Wild Relatives. Caryologia 2008, 61, 411–416. [Google Scholar] [CrossRef]
- Salvatierra-González, M.A.; Jana-Ayala, C. Floral Expression and Pollen Germination Ability in Productive Mountain Papaya (Vasconcellea pubescens A.DC.) Orchards. Chil. J. Agric. Res. 2016, 76, 136–142. [Google Scholar] [CrossRef]
- Chong-Pérez, B.; Carrasco, B.; Silva, H.; Herrera, F.; Quiroz, K.; Garcia-Gonzales, R. Regeneration of Highland Papaya (Vasconcellea pubescens) from Anther Culture. Appl. Plant Sci. 2018, 6, e01182. [Google Scholar] [CrossRef] [PubMed]
- Correa, Q.J.E.; Yesid Bernal, H. Especies Vegetales Promisorias de Los Países Del Convenio Andrés Bello; Secretaría Ejecutiva del Convenio Andrés Bello: Bogota, Colombia, 1989. [Google Scholar]
- Latcham, R.E. La Agricultura Precolombina en Chile y los Países Vecinos; Ediciones de la Universidad de Chile: Santiago, Chile, 1936. [Google Scholar]
- Squeo, F.; Arancio, G.; Gutierrez, J. Libro Rojo de La Flora Nativa y de Los Sitios Prioritarios Para Su Conservación: Región de Coquimbo; Ediciones de la Universidad de La Serena: La Serena, Chile, 2001. [Google Scholar]
- Jordan, Z.M. In Vitro Morphogenic Responses of Vasconcellea chilensis Planch. Ex, A. DC (Caricaceae). Agron. Colomb. 2011, 29, 481–485. [Google Scholar]
- Ocampo, J.; Dambier, D.; Ollitrault, P.; D’eeckenbrugge, G.; Brottier, P.; Froelicher, Y.; Risterucci, A.-M. Microsatellite Markers in Carica Papaya L.: Isolation, Characterization and Transferability to Vasconcellea Species. Mol. Ecol. Notes 2006, 6, 212–217. [Google Scholar] [CrossRef]
- Chávez-Pesqueira, M.; Núñez-Farfán, J. Genetic Diversity and Structure of Wild Populations of Carica Papaya in Northern Mesoamerica Inferred by Nuclear Microsatellites and Chloroplast Markers. Ann. Bot. 2016, 118, 1293–1306. [Google Scholar] [CrossRef]
- Chávez-Pesqueira, M.; Núñez-Farfán, J. Domestication and Genetics of Papaya: A Review. Front. Ecol. Evol. 2017, 5, 155–163. [Google Scholar] [CrossRef]
- Chávez-Pesqueira, M.; Suárez-Montes, P.; Castillo, G.; Núñez-Farfán, J. Habitat Fragmentation Threatens Wild Populations of Carica Papaya (Caricaceae) in a Lowland Rainforest. Am. J. Bot. 2014, 101, 1092–1101. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Yu, Q.; Hou, S.; Li, Y.; Eustice, M.; Skelton, R.L.; Veatch, O.; Herdes, R.E.; Diebold, L.; Saw, J.; et al. Construction of a Sequence-Tagged High-Density Genetic Map of Papaya for Comparative Structural and Evolutionary Genomics in Brassicales. Genetics 2007, 177, 2481–2491. [Google Scholar] [CrossRef] [PubMed]
- Dai, F.; Miao, Q.; Zhou, B.; Yang, L.; Liu, Z.-L. Protective Effects of Flavonols and Their Glycosides against Free Radical-Induced Oxidative Hemolysis of Red Blood Cells. Life Sci. 2006, 78, 2488–2493. [Google Scholar] [CrossRef]
- Sengupta, S.; Das, B.; Acharyya, P.; Prasad, M.; Ghose, T.K. Genetic Diversity Analysis in a Set of Caricaceae Accessions Using Resistance Gene Analogues. BMC Genet. 2014, 15, 137. [Google Scholar] [CrossRef]
- Vidal, N.M.; Grazziotin, A.L.; Ramos, H.C.C.; Pereira, M.G.; Venancio, T.M. Development of a Gene-Centered Ssr Atlas as a Resource for Papaya (Carica Papaya) Marker-Assisted Selection and Population Genetic Studies. PLoS ONE 2014, 9, e112654. [Google Scholar] [CrossRef] [PubMed]
- Blas, A.L.; Yu, Q.; Veatch, O.J.; Paull, R.E.; Moore, P.H.; Ming, R. Genetic Mapping of Quantitative Trait Loci Controlling Fruit Size and Shape in Papaya. Mol. Breed. 2012, 29, 457–466. [Google Scholar] [CrossRef]
- Na, J.-K.; Wang, J.; Murray, J.E.; Gschwend, A.R.; Zhang, W.; Yu, Q.; Navajas-Pérez, R.; Feltus, F.A.; Chen, C.; Kubat, Z.; et al. Construction of Physical Maps for the Sex-Specific Regions of Papaya Sex Chromosomes. BMC Genom. 2012, 13, 176. [Google Scholar] [CrossRef]
- Wai, C.M.; Moore, P.H.; Paull, R.E.; Ming, R.; Yu, Q. An Integrated Cytogenetic and Physical Map Reveals Unevenly Distributed Recombination Spots along the Papaya Sex Chromosomes. Chromosom. Res. 2012, 20, 753–767. [Google Scholar] [CrossRef]
- Yu, Q.; Tong, E.; Skelton, R.L.; Bowers, J.E.; Jones, M.R.; Murray, J.E.; Hou, S.; Guan, P.; Acob, R.A.; Luo, M.-C.; et al. A Physical Map of the Papaya Genome with Integrated Genetic Map and Genome Sequence. BMC Genom. 2009, 10, 371. [Google Scholar] [CrossRef]
- Kyndt, T.; Droogenbroeck, B.V.; Haegeman, A.; Roldán-Ruiz, I.; Gheysen, G. Cross-Species Microsatellite Amplification in Vasconcellea and Related Genera and Their Use in Germplasm Classification. Genome 2006, 49, 786–798. [Google Scholar] [CrossRef]
- Carrasco, B.; Avila, P.; Perez-Diaz, J.; Muñoz, P.; Garcia-Gonzales, R.; Lavandero, B.; Zurita-Silva, A.; Retamales, J.; Caligari, P. Genetic Structure of Highland Papayas (Vasconcellea pubescens (Lenn, et C. Koch) Badillo) Cultivated along a Geographic Gradient in Chile as Revealed by Inter Simple Sequence Repeats (ISSR). Genet. Resour. Crop Evol. 2008, 56, 331–337. [Google Scholar] [CrossRef]
- Carrasco, B.; García-Gonzáles, R.; Díaz, C.; Ávila, P.; Cáceres, P.; Lobos, G.A.; Silva, H.; Caligari, P.D.S. Genetic and Morphological Characterization of the Endangered Austral Papaya Vasconcellea chilensis (Planch. Ex, A. DC.) Solms. Genet. Resour. Crop. Evol. 2014, 61, 1423–1432. [Google Scholar] [CrossRef]
- Devey, D. Ecological Genetics: Design, Analysis and Application. Lowe, A., Harris S, Ashton, P. 2004. Oxford: Blackwell Publishing. £29.99 (Softback). 344 Pp. Ann. Bot. 2005, 95, 705. [Google Scholar] [CrossRef]
- Nadeem, M.A.; Nawaz, M.A.; Shahid, M.Q.; Doğan, Y.; Comertpay, G.; Yıldız, M.; Hatipoğlu, R.; Ahmad, F.; Alsaleh, A.; Labhane, N.; et al. DNA Molecular Markers in Plant Breeding: Current Status and Recent Advancements in Genomic Selection and Genome Editing. Biotechnol. Biotechnol. Equip. 2018, 32, 261–285. [Google Scholar] [CrossRef]
- Goodstein, D.M.; Shu, S.; Howson, R.; Neupane, R.; Hayes, R.D.; Fazo, J.; Mitros, T.; Dirks, W.; Hellsten, U.; Putnam, N.; et al. Phytozome: A Comparative Platform for Green Plant Genomics. Nucleic Acids Res. 2012, 40, D1178–D1186. [Google Scholar] [CrossRef] [PubMed]
- Michael, T.P.; VanBuren, R. Progress, Challenges and the Future of Crop Genomes. Curr. Opin. Plant Biol. 2015, 24, 71–81. [Google Scholar] [CrossRef]
- Ming, R.; Hou, S.; Feng, Y.; Yu, Q.; Dionne-Laporte, A.; Saw, J.H.; Senin, P.; Wang, W.; Ly, B.V.; Lewis, K.L.T.; et al. The Draft Genome of the Transgenic Tropical Fruit Tree Papaya (Carica Papaya Linnaeus). Nature 2008, 452, 991–996. [Google Scholar] [CrossRef]
- Batley, J.; Edwards, D. SNP Applications in Plants. In Association Mapping in Plants; Oraguzie, N.C., Rikkerink, E.H.A., Gardiner, S.E., De Silva, H.N., Eds.; Springer: New York, NY, USA, 2007; pp. 95–102. ISBN 978-0-387-36011-9. [Google Scholar]
- Sonah, H.; Bastien, M.; Iquira, E.; Tardivel, A.; Légaré, G.; Boyle, B.; Normandeau, É.; Laroche, J.; Larose, S.; Jean, M.; et al. An Improved Genotyping by Sequencing (GBS) Approach Offering Increased Versatility and Efficiency of SNP Discovery and Genotyping. PLoS ONE 2013, 8, e54603. [Google Scholar] [CrossRef]
- Elshire, R.J.; Glaubitz, J.C.; Sun, Q.; Poland, J.A.; Kawamoto, K.; Buckler, E.S.; Mitchell, S.E. A Robust, Simple Genotyping-by-Sequencing (GBS) Approach for High Diversity Species. PLoS ONE 2011, 6, e19379. [Google Scholar] [CrossRef] [PubMed]
- Glaubitz, J.C.; Casstevens, T.M.; Lu, F.; Harriman, J.; Elshire, R.J.; Sun, Q.; Buckler, E.S. TASSEL-GBS: A High Capacity Genotyping by Sequencing Analysis Pipeline. PLoS ONE 2014, 9, e90346. [Google Scholar] [CrossRef]
- Truong, H.T.; Ramos, A.M.; Yalcin, F.; de Ruiter, M.; van der Poel, H.J.A.; Huvenaars, K.H.J.; Hogers, R.C.J.; van Enckevort, L.J.G.; Janssen, A.; van Orsouw, N.J.; et al. Sequence-Based Genotyping for Marker Discovery and Co-Dominant Scoring in Germplasm and Populations. PLoS ONE 2012, 7, e37565. [Google Scholar] [CrossRef] [PubMed]
- Voss-Fels, K.; Snowdon, R.J. Understanding and Utilizing Crop Genome Diversity via High-Resolution Genotyping. Plant Biotechnol. J. 2016, 14, 1086–1094. [Google Scholar] [CrossRef]
- Poland, J.A.; Rife, T.W. Genotyping-by-Sequencing for Plant Breeding and Genetics. Plant Genome 2012, 5, 92–102. [Google Scholar] [CrossRef]
- Romay, M.C.; Millard, M.J.; Glaubitz, J.C.; Peiffer, J.A.; Swarts, K.L.; Casstevens, T.M.; Elshire, R.J.; Acharya, C.B.; Mitchell, S.E.; Flint-Garcia, S.A.; et al. Comprehensive Genotyping of the USA National Maize Inbred Seed Bank. Genome Biol. 2013, 14, R55. [Google Scholar] [CrossRef] [PubMed]
- Spindel, J.; Wright, M.; Chen, C.; Cobb, J.; Gage, J.; Harrington, S.; Lorieux, M.; Ahmadi, N.; McCouch, S. Bridging the Genotyping Gap: Using Genotyping by Sequencing (GBS) to Add High-Density SNP Markers and New Value to Traditional Bi-Parental Mapping and Breeding Populations. Theor. Appl. Genet. 2013, 126, 2699–2716. [Google Scholar] [CrossRef] [PubMed]
- Jarquín, D.; Kocak, K.; Posadas, L.; Hyma, K.; Jedlicka, J.; Graef, G.; Lorenz, A. Genotyping by Sequencing for Genomic Prediction in a Soybean Breeding Population. BMC Genom. 2014, 15, 740. [Google Scholar] [CrossRef]
- Uitdewilligen, J.G.A.M.L.; Wolters, A.-M.A.; D’hoop, B.B.; Borm, T.J.A.; Visser, R.G.F.; van Eck, H.J. A Next-Generation Sequencing Method for Genotyping-by-Sequencing of Highly Heterozygous Autotetraploid Potato. PLoS ONE 2013, 8, e62355. [Google Scholar] [CrossRef]
- Gürcan, K.; Teber, S.; Ercisli, S.; Yilmaz, K.U. Genotyping by Sequencing (GBS) in Apricots and Genetic Diversity Assessment with GBS-Derived Single-Nucleotide Polymorphisms (SNPs). Biochem. Genet. 2016, 54, 854–885. [Google Scholar] [CrossRef]
- Cao, K.; Li, Y.; Deng, C.H.; Gardiner, S.E.; Zhu, G.; Fang, W.; Chen, C.; Wang, X.; Wang, L. Comparative Population Genomics Identified Genomic Regions and Candidate Genes Associated with Fruit Domestication Traits in Peach. Plant Biotechnol. J. 2019, 17, 1954–1970. [Google Scholar] [CrossRef]
- Chong, X.; Zhang, F.; Wu, Y.; Yang, X.; Zhao, N.; Wang, H.; Guan, Z.; Fang, W.; Chen, F. A SNP-Enabled Assessment of Genetic Diversity, Evolutionary Relationships and the Identification of Candidate Genes in Chrysanthemum. Genome Biol. Evol. 2016, 8, 3661–3671. [Google Scholar] [CrossRef]
- Carrasco, B.; González, M.; Gebauer, M.; García-González, R.; Maldonado, J.; Silva, H. Construction of a Highly Saturated Linkage Map in Japanese Plum (Prunus Salicina, L.) Using GBS for SNP Marker Calling. PLoS ONE 2018, 13, e0208032. [Google Scholar] [CrossRef] [PubMed]
- Koepke, T.; Schaeffer, S.; Harper, A.; Dicenta, F.; Edwards, M.; Henry, R.J.; Møller, B.L.; Meisel, L.; Oraguzie, N.; Silva, H.; et al. Comparative Genomics Analysis in Prunoideae to Identify Biologically Relevant Polymorphisms. Plant Biotechnol. J. 2013, 11, 883–893. [Google Scholar] [CrossRef] [PubMed]
- Tajima, F. Statistical Method for Testing the Neutral Mutation Hypothesis by DNA Polymorphism. Genetics 1989, 123, 585–595. [Google Scholar] [CrossRef] [PubMed]
- Jobin-Decor, M.P.; Graham, G.C.; Henry, R.J.; Drew, R.A. RAPD and Isozyme Analysis of Genetic Relationships between Carica Papaya and Wild Relatives. Genet. Resour. Crop Evol. 1997, 44, 471–477. [Google Scholar] [CrossRef]
- Kanupriya, K.; Shobhana, M.; Vasugi, C.; Aswath, C.; Radhika, V.; Reddy, L.; Dinesh, M. Genetic Relationship among Papaya (Carica Papaya) and Wild Papaya (Vasconcellea Species) Using RAPD and ISSR Markers. Indian J. Agric. Sci. 2012, 82, 366–369. [Google Scholar]
- Rodríguez, J.; Rodríguez, P.; González, M.E.; Martínez-Gómez, P. Molecular Characterization of Cuban Endemism Carica Cubensis Solms Using Random Amplified Polymorphic DNA (RAPD) Markers. Agric. Sci. 2010, 1, 95. [Google Scholar] [CrossRef]
- Van Droogenbroeck, B.; Breyne, P.; Goetghebeur, P.; Romeijn-Peeters, E.; Kyndt, T.; Gheysen, G. AFLP Analysis of Genetic Relationships among Papaya and Its Wild Relatives (Caricaceae) from Ecuador. Theor. Appl. Genet. 2002, 105, 289–297. [Google Scholar] [CrossRef]
- Brown, J.E.; Bauman, J.M.; Lawrie, J.F.; Rocha, O.J.; Moore, R.C. The Structure of Morphological and Genetic Diversity in Natural Populations of Carica Papaya (Caricaceae) in Costa Rica. Biotropica 2012, 44, 179–188. [Google Scholar] [CrossRef]
- Singh, N.; Choudhury, D.R.; Singh, A.K.; Kumar, S.; Srinivasan, K.; Tyagi, R.K.; Singh, N.K.; Singh, R. Comparison of SSR and SNP Markers in Estimation of Genetic Diversity and Population Structure of Indian Rice Varieties. PLoS ONE 2013, 8, e84136. [Google Scholar] [CrossRef]
- Arias, D.M.; Albarrán-Lara, A.L.; González-Rodríguez, A.; Peñaloza-Ramírez, J.; Dorado, O.; Leyva, E. Genetic Diversity and Structure of Wild Populations of the Tropical Dry Forest Tree Jacaratia Mexicana (Brassicales: Caricaceae) at a Local Scale in Mexico. Rev. Biol. Trop. 2012, 60, 1–10. [Google Scholar] [CrossRef]
- Causse, M.; Desplat, N.; Pascual, L.; Le Paslier, M.-C.; Sauvage, C.; Bauchet, G.; Bérard, A.; Bounon, R.; Tchoumakov, M.; Brunel, D.; et al. Whole Genome Resequencing in Tomato Reveals Variation Associated with Introgression and Breeding Events. BMC Genom. 2013, 14, 791. [Google Scholar] [CrossRef] [PubMed]
- Foster, J.T.; Allan, G.J.; Chan, A.P.; Rabinowicz, P.D.; Ravel, J.; Jackson, P.J.; Keim, P. Single Nucleotide Polymorphisms for Assessing Genetic Diversity in Castor Bean (Ricinus Communis). BMC Plant Biol. 2010, 10, 13. [Google Scholar] [CrossRef] [PubMed]
- Abdullah, N.; Bakar, N.A.; Bakar, U.K.A.; Abidin, R.A.Z.; Yusof, M.F. Sequence Information on Single Nucleotide Polymorphism (SNP) through Genome Sequencing Analysis of Carica Papaya Variety Eksotika and Sekaki. J. Trop. Agric. Food Sci. 2016, 44, 219–228. [Google Scholar]
- Ueno, H.; Urasaki, N.; Natsume, S.; Yoshida, K.; Tarora, K.; Shudo, A.; Terauchi, R.; Matsumura, H. Genome Sequence Comparison Reveals a Candidate Gene Involved in Male-Hermaphrodite Differentiation in Papaya (Carica Papaya) Trees. Mol. Genet. Genom. 2015, 290, 661–670. [Google Scholar] [CrossRef] [PubMed]
- VanBuren, R.; Wai, C.M.; Zhang, J.; Han, J.; Arro, J.; Lin, Z.; Liao, Z.; Yu, Q.; Wang, M.-L.; Zee, F.; et al. Extremely Low Nucleotide Diversity in the X-Linked Region of Papaya Caused by a Strong Selective Sweep. Genome Biol. 2016, 17, 230. [Google Scholar] [CrossRef]
- Lee, C.-Y.; Lin, H.-J.; Viswanath, K.K.; Lin, C.-P.; Chang, B.C.-H.; Chiu, P.-H.; Chiu, C.-T.; Wang, R.-H.; Chin, S.-W.; Chen, F.-C. The Development of Functional Mapping by Three Sex-Related Loci on the Third Whorl of Different Sex Types of Carica Papaya L. PLoS ONE 2018, 13, e0194605. [Google Scholar] [CrossRef]
- Nybom, H. Comparison of Different Nuclear DNA Markers for Estimating Intraspecific Genetic Diversity in Plants. Mol. Ecol. 2004, 13, 1143–1155. [Google Scholar] [CrossRef]
- Aranzana, M.J.; Illa, E.; Howad, W.; Arús, P. A First Insight into Peach [Prunus Persica (L.) Batsch] SNP Variability. Tree Genet. Genomes 2012, 8, 1359–1369. [Google Scholar] [CrossRef]
- Emanuelli, F.; Lorenzi, S.; Grzeskowiak, L.; Catalano, V.; Stefanini, M.; Troggio, M.; Myles, S.; Martinez-Zapater, J.M.; Zyprian, E.; Moreira, F.M.; et al. Genetic Diversity and Population Structure Assessed by SSR and SNP Markers in a Large Germplasm Collection of Grape. BMC Plant Biol. 2013, 13, 39. [Google Scholar] [CrossRef]
- Micheletti, D.; Troggio, M.; Zharkikh, A.; Costa, F.; Malnoy, M.; Velasco, R.; Salvi, S. Genetic Diversity of the Genus Malus and Implications for Linkage Mapping with SNPs. Tree Genet. Genomes 2011, 7, 857–868. [Google Scholar] [CrossRef]
- Silva, J.a.T.; Rashid, Z.; Nhut, D.T.; Sivakumar, D.; Gera, A.; Souza, M.T.; Tennant, P. Papaya (Carica Papaya L.) Biology and Biotechnology. Tree For. Sci. Biotechnol. 2007, 1, 47–73. [Google Scholar]
- Sousa, E.; Kost, B.; Malhó, R. Arabidopsis Phosphatidylinositol-4-Monophosphate 5-Kinase 4 Regulates Pollen Tube Growth and Polarity by Modulating Membrane Recycling. Plant Cell 2008, 20, 3050–3064. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Yang, H.; Yu, J.; Chen, C.; Zhang, G.; Bao, J.; Du, Y.; Kibukawa, M.; Li, Z.; Wang, J.; et al. Genomic Organization, Transcript Variants and Comparative Analysis of the Human Nucleoporin 155 (NUP155) Gene. Gene 2002, 288, 9–18. [Google Scholar] [CrossRef]
- Yuan, Y.; Qi, L.; Yu, J.; Wang, X.; Huang, L. Transcriptome-Wide Analysis of SAMe Superfamily to Novelty Phosphoethanolamine N-Methyltransferase Copy in Lonicera Japonica. Int. J. Mol. Sci. 2014, 16, 521–534. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, W.; Liu, E.; Ishii, H.; Matsunaga, S.; Schlögelhofer, P.; Kurumizaka, H. Homologous Pairing Activities of Arabidopsis Thaliana RAD51 and DMC1. J. Biochem. 2019, 165, 289–295. [Google Scholar] [CrossRef] [PubMed]
- Pradillo, M.; López, E.; Linacero, R.; Romero, C.; Cuñado, N.; Sánchez-Morán, E.; Santos, J.L. Together Yes, but Not Coupled: New Insights into the Roles of RAD51 and DMC1 in Plant Meiotic Recombination. Plant J. 2012, 69, 921–933. [Google Scholar] [CrossRef] [PubMed]
- Ashraf, M.A.; Rahman, A. Cold Stress Response in Arabidopsis Thaliana Is Mediated by GNOM ARF-GEF. Plant J. 2019, 97, 500–516. [Google Scholar] [CrossRef] [PubMed]
- Kopczak, S.D.; Haas, N.A.; Hussey, P.J.; Silflow, C.D.; Snustad, D.P. The Small Genome of Arabidopsis Contains at Least Six Expressed Alpha-Tubulin Genes. Plant Cell 1992, 4, 539–547. [Google Scholar] [CrossRef]
- Krupková, E.; Immerzeel, P.; Pauly, M.; Schmülling, T. The TUMOROUS SHOOT DEVELOPMENT2 Gene of Arabidopsis Encoding a Putative Methyltransferase Is Required for Cell Adhesion and Co-Ordinated Plant Development. Plant J. 2007, 50, 735–750. [Google Scholar] [CrossRef]
- Sun, K.; Wolters, A.-M.A.; Loonen, A.E.H.M.; Huibers, R.P.; van der Vlugt, R.; Goverse, A.; Jacobsen, E.; Visser, R.G.F.; Bai, Y. Down-Regulation of Arabidopsis DND1 Orthologs in Potato and Tomato Leads to Broad-Spectrum Resistance to Late Blight and Powdery Mildew. Transgenic Res. 2016, 25, 123–138. [Google Scholar] [CrossRef]
- Katano, K.; Kataoka, R.; Fujii, M.; Suzuki, N. Differences between Seedlings and Flowers in Anti-ROS Based Heat Responses of Arabidopsis Plants Deficient in Cyclic Nucleotide Gated Channel 2. Plant Physiol. Biochem. 2018, 123, 288–296. [Google Scholar] [CrossRef] [PubMed]
- Jarsch, I.K.; Ott, T. Perspectives on Remorin Proteins, Membrane Rafts, and Their Role during Plant-Microbe Interactions. Mol. Plant Microbe Interact. 2011, 24, 7–12. [Google Scholar] [CrossRef] [PubMed]
- Raffaele, S.; Bayer, E.; Mongrand, S. Upregulation of the Plant Protein Remorin Correlates with Dehiscence and Cell Maturation. Plant Signal. Behav. 2009, 4, 915–919. [Google Scholar] [CrossRef] [PubMed]
- Raffaele, S.; Bayer, E.; Lafarge, D.; Cluzet, S.; German Retana, S.; Boubekeur, T.; Leborgne-Castel, N.; Carde, J.-P.; Lherminier, J.; Noirot, E.; et al. Remorin, a Solanaceae Protein Resident in Membrane Rafts and Plasmodesmata, Impairs Potato Virus X Movement. Plant Cell 2009, 21, 1541–1555. [Google Scholar] [CrossRef]
- Costa, A.; Luoni, L.; Marrano, C.A.; Hashimoto, K.; Köster, P.; Giacometti, S.; De Michelis, M.I.; Kudla, J.; Bonza, M.C. Ca2+-Dependent Phosphoregulation of the Plasma Membrane Ca2+-ATPase ACA8 Modulates Stimulus-Induced Calcium Signatures. J. Exp. Bot. 2017, 68, 3215–3230. [Google Scholar] [CrossRef] [PubMed]
- Bakshi, A.; Wilson, R.L.; Lacey, R.F.; Kim, H.; Wuppalapati, S.K.; Binder, B.M. Identification of Regions in the Receiver Domain of the ETHYLENE RESPONSE1 Ethylene Receptor of Arabidopsis Important for Functional Divergence. Plant Physiol. 2015, 169, 219–232. [Google Scholar] [CrossRef]
- Carretero-Paulet, L.; Cairó, A.; Talavera, D.; Saura, A.; Imperial, S.; Rodríguez-Concepción, M.; Campos, N.; Boronat, A. Functional and Evolutionary Analysis of DXL1, a Non-Essential Gene Encoding a 1-Deoxy-D-Xylulose 5-Phosphate Synthase like Protein in Arabidopsis Thaliana. Gene 2013, 524, 40–53. [Google Scholar] [CrossRef]
- Floss, D.S.; Hause, B.; Lange, P.R.; Küster, H.; Strack, D.; Walter, M.H. Knock-down of the MEP Pathway Isogene 1-Deoxy-D-Xylulose 5-Phosphate Synthase 2 Inhibits Formation of Arbuscular Mycorrhiza-Induced Apocarotenoids, and Abolishes Normal Expression of Mycorrhiza-Specific Plant Marker Genes. Plant J. 2008, 56, 86–100. [Google Scholar] [CrossRef]
- Kazenwadel, C.; Klebensberger, J.; Richter, S.; Pfannstiel, J.; Gerken, U.; Pickel, B.; Schaller, A.; Hauer, B. Optimized Expression of the Dirigent Protein AtDIR6 in Pichia Pastoris and Impact of Glycosylation on Protein Structure and Function. Appl. Microbiol. Biotechnol. 2013, 97, 7215–7227. [Google Scholar] [CrossRef]
- Riera, M.; Figueras, M.; Lopez, C.; Goday, A.; Pages, M. Protein Kinase CK2 Modulates Developmental Functions of the Abscisic Acid Responsive Protein Rab17 from Maize. Proc. Natl. Acad. Sci. USA 2004, 101, 9879–9884. [Google Scholar] [CrossRef]
- Chen, D.; Molitor, A.M.; Xu, L.; Shen, W.-H. Arabidopsis PRC1 Core Component AtRING1 Regulates Stem Cell-Determining Carpel Development Mainly through Repression of Class I KNOX Genes. BMC Biol. 2016, 14, 112. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and Accurate Short Read Alignment with Burrows-Wheeler Transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Bradbury, P.J.; Zhang, Z.; Kroon, D.E.; Casstevens, T.M.; Ramdoss, Y.; Buckler, E.S. TASSEL: Software for Association Mapping of Complex Traits in Diverse Samples. Bioinformatics 2007, 23, 2633–2635. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic Analysis in Excel. Population Genetic Software for Teaching and Research—An Update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef]
- Yeh, F.; Yang, R.; Boyle, T. Microsoft Windows-Based Freeware for Populations Genetic Analysis; POPGENE Version 1.32, Undefined; University of Alberta: Edmonton, AB, Canada, 1999. [Google Scholar]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Gracia, A. DnaSP 6: DNA Sequence Polymorphism Analysis of Large Data Sets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Hull-Sanders, H.M.; Eubanks, M.D.; Carr, D.E. Inbreeding Depression and Selfing Rate of Ipomoea Hederacea Var. Integriuscula (Convolvulaceae). Am. J. Bot. 2005, 92, 1871–1877. [Google Scholar] [CrossRef] [PubMed]
- Nei, M.; Li, W.H. Mathematical Model for Studying Genetic Variation in Terms of Restriction Endonucleases. Proc. Natl. Acad. Sci. USA 1979, 76, 5269–5273. [Google Scholar] [CrossRef]
- Pritchard, J.K.; Stephens, M.; Donnelly, P.J. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the Number of Clusters of Individuals Using the Software STRUCTURE: A Simulation Study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- Earl, D.A.; vonHoldt, B.M. STRUCTURE HARVESTER: A Website and Program for Visualizing STRUCTURE Output and Implementing the Evanno Method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]
- Francis, R. Pophelper: An R Package and Web App to Analyse and Visualize Population Structure. Mol. Ecol. Resour. 2016, 17, 27–32. [Google Scholar] [CrossRef] [PubMed]
- Ramasamy, R.K.; Ramasamy, S.; Bindroo, B.B.; Naik, V.G. STRUCTURE PLOT: A Program for Drawing Elegant STRUCTURE Bar Plots in User Friendly Interface. Springerplus 2014, 3, 431. [Google Scholar] [CrossRef] [PubMed]
- Excoffier, L.; Smouse, P.E.; Quattro, J.M. Analysis of Molecular Variance Inferred from Metric Distances among DNA Haplotypes: Application to Human Mitochondrial DNA Restriction Data. Genetics 1992, 131, 479–491. [Google Scholar] [CrossRef] [PubMed]

| Species | N | L | Ho | π (10−3) | D |
|---|---|---|---|---|---|
| V. pubescens | 92 | 4675 | 0.04 | 3.2 | −2.281 |
| Guallarauco Ltd.a | 24 | 0.05 | 3.5 | ||
| Saturno SA | 23 | 0.04 | 3.2 | ||
| Sociedad Agrícola HC | 24 | 0.04 | 3.0 | ||
| Frutos de Lipimávida | 21 | 0.05 | 3.1 | ||
| V. chilensis | 94 | 4451 | 0.12 | 15.4 | −1.065 |
| Chungungo | 26 | 0.12 | 17.0 | ||
| Fray Jorge | 27 | 0.13 | 17.8 | ||
| Huachalalume | 24 | 0.10 | 15.2 | ||
| Valle del Encanto | 17 | 0.12 | 11.4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Carrasco, B.; Arévalo, B.; Perez-Diaz, R.; Rodríguez-Alvarez, Y.; Gebauer, M.; Maldonado, J.E.; García-Gonzáles, R.; Chong-Pérez, B.; Pico-Mendoza, J.; Meisel, L.A.; et al. Descriptive Genomic Analysis and Sequence Genotyping of the Two Papaya Species (Vasconcellea pubescens and Vasconcellea chilensis) Using GBS Tools. Plants 2022, 11, 2151. https://doi.org/10.3390/plants11162151
Carrasco B, Arévalo B, Perez-Diaz R, Rodríguez-Alvarez Y, Gebauer M, Maldonado JE, García-Gonzáles R, Chong-Pérez B, Pico-Mendoza J, Meisel LA, et al. Descriptive Genomic Analysis and Sequence Genotyping of the Two Papaya Species (Vasconcellea pubescens and Vasconcellea chilensis) Using GBS Tools. Plants. 2022; 11(16):2151. https://doi.org/10.3390/plants11162151
Chicago/Turabian StyleCarrasco, Basilio, Bárbara Arévalo, Ricardo Perez-Diaz, Yohaily Rodríguez-Alvarez, Marlene Gebauer, Jonathan E. Maldonado, Rolando García-Gonzáles, Borys Chong-Pérez, José Pico-Mendoza, Lee A. Meisel, and et al. 2022. "Descriptive Genomic Analysis and Sequence Genotyping of the Two Papaya Species (Vasconcellea pubescens and Vasconcellea chilensis) Using GBS Tools" Plants 11, no. 16: 2151. https://doi.org/10.3390/plants11162151
APA StyleCarrasco, B., Arévalo, B., Perez-Diaz, R., Rodríguez-Alvarez, Y., Gebauer, M., Maldonado, J. E., García-Gonzáles, R., Chong-Pérez, B., Pico-Mendoza, J., Meisel, L. A., Ming, R., & Silva, H. (2022). Descriptive Genomic Analysis and Sequence Genotyping of the Two Papaya Species (Vasconcellea pubescens and Vasconcellea chilensis) Using GBS Tools. Plants, 11(16), 2151. https://doi.org/10.3390/plants11162151







