Abstract
Phosphates are known to be essential for plant growth and development, with phosphorus compounds being involved in various physiological and biochemical reactions. Phosphates are known as one of the most important factors limiting crop yields. The problem of phosphorus deficiency in the soil has traditionally been solved by applying phosphate fertilizers. However, chemical phosphate fertilizers are considered ineffective compared to the organic fertilizers manure and compost. Therefore, increasing the bioavailability of phosphates for plants is one of the primary goals of sustainable agriculture. Phosphate-solubilizing soil microorganisms can make soil-insoluble phosphate bioavailable for plants through solubilization and mineralization. These microorganisms are currently in the focus of interest due to their advantages, such as environmental friendliness, low cost, and high biological efficiency. In this regard, the solubilization of phosphates by soil microorganisms holds strong potential in research, and inoculation of soils or crops with phosphate-solubilizing bacteria is a promising strategy to improve plant phosphate uptake. In this review, we analyze all the species of phosphate-solubilizing bacteria described in the literature to date. We discuss key mechanisms of solubilization of mineral phosphates and mineralization of organic phosphate-containing compounds: organic acids secreted by bacteria for the mobilization of insoluble inorganic phosphates, and the enzymes hydrolyzing phosphorus-containing organic compounds. We demonstrate that phosphate-solubilizing microorganisms have enormous potency as biofertilizers since they increase phosphorus bioavailability for the plant, promote sustainable agriculture, improve soil fertility, and raise crop yields. The use of phosphate-solubilizing microbes is regarded as a new frontier in increasing plant productivity.
1. Introduction
Phosphorus (P) is one of the most important macronutrients for the growth and development of plants [1,2]. It comprises 0.2 to 0.8% of the dry weight of plants [3] and is found in nucleic acids, enzymes, coenzymes, nucleotides, and phospholipids [4]. Phosphorus is the second most important macronutrient necessary for plants, after nitrogen. The average phosphorus content in soil is about 0.05% (by weight), with only 0.1% of this amount available to be used by plants [5]. Plant roots can absorb phosphorus in the form of orthophosphates H2PO4− or HPO42–, but the concentration of these ions in the soil is in the micromolar range [6]. In soil, most phosphates are present as insoluble iron, aluminum, and calcium phosphates.
Phosphates are involved in various physiological and biochemical reactions, including photosynthesis, root and stem development, flower and seed formation, crop maturation, nitrogen fixation in legumes, and resistance to plant diseases [3]. Phosphates are among the most important factors limiting crop yields [7].
The problem of phosphate deficiency in the soil is conventionally solved by using phosphorus fertilizers. However, with phosphate anions in chemical fertilizers being extremely reactive and quickly fixed due to interactions with Ca2+, Fe3+, and Al3+ in the soil, most of the phosphorus added to fertilizers is inaccessible to plants. Thus, the formation of insoluble complexes of corresponding phosphate salts leads to the efficiency of using chemical phosphate fertilizers of only 5–25% [8]. Also, the long-term application of phosphorus fertilizers causes soil acidification, water pollution, and eutrophication [9]. Therefore, increasing the bioavailability of soil-insoluble phosphate for the plants is one of the primary goals of agriculture and forestry development.
Soil microorganisms play a key role in plant nutrient uptake and are involved in a wide range of biological processes. Some of them can mineralize insoluble soil phosphorus, making them available for plant consumption. Aside from chemical fertilization, microbial phosphate solubilization and mineralization seem to be the only possible way to increase the amount of phosphorus accessible for plants. Numerous microorganisms in the soil and rhizosphere are known to effectively release phosphorus from the soil through solubilization and mineralization [10], and these are referred to as phosphate-solubilizing microorganisms.
Phosphate-solubilizing microorganisms are gaining attention due to their advantages, such as safety for the environment, low cost, and high efficiency [11,12]. The use of phosphate biofertilizers is a promising approach to improving food production and increasing crop yields. Phosphate-solubilizing microorganisms can promote plant growth by enhancing the efficiency of biological nitrogen fixation and phytohormone synthesis and increasing the availability of certain micronutrients, such as zinc and iron [13]. Phosphate-solubilizing microorganism inoculations have been proven to increase plant yield and phosphorus uptake both in pots and in the field [14].
Facilitating phosphate uptake and utilization by plants is critical for economic and environmental reasons. The solubilization of phosphates by soil microorganisms is regarded as a particularly prospective research object. Thus, the inoculation of soil or crops with phosphate-solubilizing microorganisms is considered a promising strategy for improving plant phosphorus uptake [15]. Here, we review the availability of soil phosphate, the diversity of phosphate-solubilizing soil microorganisms, the mechanisms and induction of phosphate solubilization and their role in plant growth, and the potential role of such microorganisms as natural biofertilizers in crop production.
2. Soil Phosphate-Solubilizing Microorganisms
Many types of soil bacteria and fungi can dissolve phosphates in vitro. Phosphate-solubilizing microorganisms increase the bioavailability of soil phosphorus for plants [5]. They dissolve insoluble inorganic (mineral) phosphorus and mineralize insoluble organic phosphates [3]. Salt-tolerant or halophilic soil microorganisms are able to dissolve insoluble phosphates, thereby contributing to the development of agriculture on saline and alkali soils [5].
The ability to solubilize phosphates was characterized in detail for mycorrhizal fungi [16] that are also proven to contribute to the crop nutrition by increasing the volume of rhizosphere and thus the volume of soil from which phosphates can be absorbed [17]. Other soil microorganisms are phosphate-solubilizing bacteria (PSB) that are generally associated with the plant rhizosphere [18]. Table 1 provides information on the genera of phosphate-solubilizing bacteria and fungi.

Table 1.
Genera of soil phosphate-solubilizing bacteria and fungi.
According to the literature data, PSBs account for 1–50% of the total microbial population, compared to the phosphate-solubilizing fungi accounting for 0.1–0.5%. Phosphate-solubilizing microorganisms are ubiquitous, with their species composition varying from soil to soil. Most phosphate-solubilizing microorganisms have been isolated from the rhizosphere of various plants, where they are metabolically active [37].
3. Biochemical Properties of the Soil Related to the Bioavailability of Phosphates
Soil phosphates exist both in inorganic and organic form. The inorganic (mineral) form of phosphate is represented by primary (apatites, strengite, and variscite) and secondary minerals of phosphorus (iron, aluminum, and calcium phosphates) [38]. The release of phosphate from these minerals is slow and regulated by several factors, particularly by soil pH [38,39]. Under acidic conditions, phosphates are adsorbed on Al/Fe oxides and hydroxides such as gibbsite and goethite. Under alkaline conditions, phosphates are precipitated with calcium. Additionally, phosphorus may be immobilized on the soil clay particles, a phenomenon strongly influenced by the type of ions adsorbed on the surface of clay minerals [40].
Soil microorganisms can immobilize phosphates from the soil. After absorption by microbial cells, phosphorus is incorporated into the cellular structures of microorganisms (e.g., nucleic acids, organic phosphorus esters, free inorganic phosphate, and coenzymes), with excess phosphorus likely to be stored as polyphosphates [41]. Microbial biomass is an important temporary immobilized phosphorus pool that can be mineralized and released into the soil solution as available phosphorus long-term [42,43].
In most microorganisms, the role of a phosphate reserve is played by inorganic polyphosphates, linear polymers of phosphoric acid containing between three and several hundred phosphate residues [44]. Many bacteria accumulate polyphosphates under unfavorable conditions [45]. For example, the accumulation of polyphosphates in E. coli cells occurs under conditions of amino acid deficiency [46]. Enzymes catalyzing the synthesis of polyphosphates are polyP kinases. The role of polyP as a phosphate reserve has been proven for many microorganisms belonging to different taxa, from archaea to fungi [45]. Polyphosphates are rich in “high energy” of anhydride bonds [47], and can be used as an energy source due to the Pi release [48].
Various enzymes are involved in the consumption of accumulated polyphosphates and the breakdown of phosphates outside the cell. Polyphosphate utilization and degradation is catalyzed by polypases, including exopolypase (PPX), and several polyphosphate-specific kinases, including polyP-glucokinase and polyP-fructokinase [45].
The biomass of soil microorganisms stores significant amounts of phosphorus, protecting it from plant adsorption [49,50]. During the biogeochemical recycling process, the phosphorus accumulated by bacteria and fungi is slowly released back into the soil and becomes available to plants, as evidenced by the correlation between phosphorus uptake by plants and phosphorus biomass in microorganisms [51].
The rate of phosphate release from microbial biomass likely depends on the amount of phosphorus available in soil [52], carbon availability [53], soil texture [54], and composition of microbial community [55]. The ability of microorganisms to balance between solubilization (mineralization) and immobilization processes determines the extent to which these bacteria and fungi can improve the availability of phosphorus in the soil–plant system. Figure 1 depicts several biochemical routes of phosphate solubilization by soil bacteria: synthesis of organic and inorganic acids; excretion of ammonia ions; synthesis and excretion of phosphatases and phytases; and synthesis of polysaccharides, cytokinins, and gibberellins.
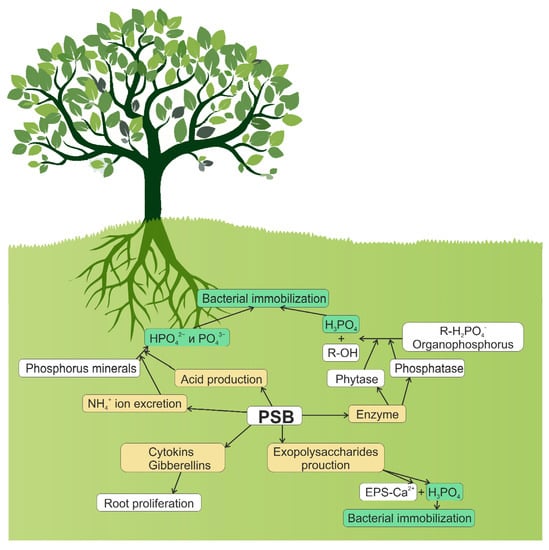
Figure 1.
Role of PSBs in the solubilization of phosphates: solubilization of inorganic phosphates due to the synthesis of organic and inorganic acids; NH4+-excretion; solubilization of organic phosphorus-containing compounds due to the synthesis and excretion of enzymes; synthesis of polysaccharides, cytokinins, and gibberellins.
4. Mechanisms of Inorganic Phosphate Solubilization
In recent years, numerous studies have been done on the characteristics and mechanisms of phosphorus solubilization processes. PSBs are thought to be able to dissolve insoluble phosphates by secreting low molecular weight organic acids [56]. In alkaline soils, phosphates can precipitate to form calcium phosphates, their solubility increasing with a decrease in soil pH. PSBs increase phosphorus availability by secreting organic acids that cause a decrease in soil pH [57,58]. The increase of soil pH leads to the appearance of divalent and trivalent forms of inorganic phosphorus: HPO42− and PO43− [37].
Organic acids are produced in the periplasmic space by direct oxidation [59]. The release of these organic acids into the environment is accompanied by a decrease in pH. Surprisingly, there is no correlation between pH and the amount of solubilized phosphorus [60]. The mechanism of acidification was suggested: the release of H+ is associated with the assimilation of cations. For example, NH4+ assimilation together with H+ excretion leads to phosphate solubilization [61]. An alternative mechanism for the solubilization of mineral phosphates for producing organic acids is the release of H+ to the extracellular surface via the exchange of cation or the ATPase activity with H+ translocation [18]. Also, assimilation of NH4+ in microbial cells was reported to be accompanied by proton release, leading to the solubilization of phosphorus without any organic acids being formed [3].
The efficiency of solubilization depends on the strength and nature of the acids. Tri- and dicarboxylic acids are more efficient than monobasic and aromatic acids, and aliphatic acids are more efficient in phosphate solubilization than phenolic, citric, and fumaric acids. Among the organic acids that solubilize phosphates are primarily citric, lactic, glicolic, 2-ketogluconic, oxalic, glyconic, acetic, malic, fumaric, succinic, tartaric, malonic, glutaric, propionic, butyric, glyoxalic, and adipic acids [37], with gluconic acid and 2-ketogluconic acid being the most common solubilizing agents for mineral phosphates. Listed in Table 2 are organic acids synthesized by PSBs.

Table 2.
Important phosphate-solubilizing microorganisms, their ecological niches, and organic acids produced.
Organic acids are synthetized as products of carbon metabolism [79,80], which is closely related to the concentration of soluble phosphate [81]. The production of acids such as gluconic acid and pyruvic acid by PSB was proven to be significantly affected by the concentration of soluble phosphate [82,83]. The conversion of glucose to the organic acids by the cells of PSBs in phosphate deficient conditions is higher than in phosphate sufficient conditions, and it is suggested that the effect of soluble phosphate on organic acid production underlies the regulation of soluble phosphate in PSB mediated mineral phosphate solubilization [82].
Low-molecular-weight organic acids are synthetized during glucose oxidation through direct periplasmic oxidation [79] and intracellular phosphorylation [80]. In the P. aeruginosa P4 strain, phosphate deficiency contributes to an increase in gluconic acid secretion, which is caused by the change of the predominant glucose metabolism pathway from phosphorylation to direct oxidation [82].
Gluconic acid was proven to be an effective organic acid component in mineral phosphate solubilization in Serratia marcescens [84], and E. herbicola [85]. B. multivorans WS-FJ9 solubilizes the phosphate in SP medium using only gluconic and pyruvic acids. However, gluconic acid production is not affected by changes in the concentration of soluble phosphate [86], which could be due to the constitutive nature of the direct oxidative pathway of glucose metabolism [82]. Soil fungi may be even more important than bacteria since they typically produce and excrete more acids, such as gluconic, citric, lactic, 2-ketogluconic, oxalic, tartaric, and acetic acid [3]. Other mechanisms of mineral phosphate solubilization by microorganisms are the production of inorganic acids (sulfuric, nitric, and carbonic acids) and the secretion of chelating agents. However, the effectiveness of inorganic acids and chelating agents in the phosphorus release into the soil is lower than that of organic acids [68]. Another mechanism of microbial phosphate solubilization was reported to be the secretion of enzymes [5].
5. Mineralization of Organic Phosphate-Containing Compounds
The content of organic phosphorus in the soil can reach 30–50% of the total amount, with soil organic phosphorus being found primarily in the form of inositol phosphate (soil phytate). Other organic phosphorus compounds were reported, including: phosphomonoesters, phosphodiesters, phospholipids, nucleic acids, and phosphotriesters [18]. Additionally, large quantities of xenobiotics (pesticides, detergents, antibiotics, and flame retardants) that are regularly released into the environment are also known to contain organic P. Most of these organic compounds are of high molecular weight and resistant to chemical hydrolysis. For plant uptake, these compounds must be converted into soluble ionic phosphate (Pi, HPO42−, H2PO4−) or low molecular weight organic phosphates [87].
Several groups of enzymes secreted by phosphate-solubilizing microorganisms are involved in the process of phosphate mineralization. The enzymes of the first group dephosphorylate phosphor-ester or phosphoanhydride bonds in organic compounds. These are non-specific acid phosphatases (NSAP). The most studied NSAP enzymes are phosphomonoesterases, also called phosphatases [88]. These enzymes can be acidic or alkaline phosphomonoesterases [89]. The pH of the soils possessing phosphatase activity was indicated to be from acidic to neutral, indicating that acid phosphatases play a major role in this process [18].
Phytase is another phosphate-solubilizing enzyme involved in the mineralization of organic P. This enzyme is responsible for releasing phosphorus from organic compounds in the soil (plant seeds and pollen) that are stored in the form of phytate (inositol polyphosphate). Phytase releases phosphates in a form that is available to plants. While plants cannot obtain phosphorus directly from the phytate, the presence of phosphate-solubilizing microbes in the rhizosphere can compensate for the inability of plants to obtain phosphates directly from phytate [43]. Phytates are synthesized by plants and constitute a significant amount (from 60 to 80%) of organic phosphorus in the soil [90]. However, the ester bonds in phytic acid are quite stable and their natural degradation is practically impossible [91]. Microbial mineralization of phytate by phytase plays an essential role in the process of phosphorus recycling. Phytate can be completely hydrolyzed with the formation of one inositol and six molecules of inorganic phosphate, or partially with the formation of lower isomers of inositol polyphosphate and inorganic phosphates [92].
Among the four types of phytases identified, β-propeller phytase (BPP, EC 3.1.3.8 or EC 3.1.3.26) differs from the other three phytases (histidine acid phosphatase, cysteine phytase, and purple acid phosphatase) in that it has a neutral ∼pH 7.0 rather than an acidic pH optimum. It has been indicated that BPP is the main class of phytate degrading enzymes in nature [90,93]. A typical BPP has a six-lobed propeller fold with two phosphate-binding sites (a cleavage site and an affinity site) and six calcium-binding sites, three of which are high-affinity binding sites responsible for enzyme stability and three of which are low-affinity sites, regulating the catalytic activity of the enzyme [94,95].
Thus far, only a small number of BPP have been isolated and studied, including Shewanella oneidensis MR-1 PhyS [96], Bacillus subtilis PhyC [97], Bacillus sp. DS11 Phy [98], B. subtilis 168 168PhyA [99], Bacillus licheniformis PhyL [99], Pedobacterobsis 5 MJ11 PhyP [100], and Janthinobacterium sp. TN115 PhyA115 [101], all of which are mesophilic or thermophilic.
Phosphate solubilization by acid phosphatases was reported for Pseudomonas sp. [102], Burkholderia cepacia [103], Enterobacter aerogenes, E. cloacae, Citrobacter freundi, Proteus mirabalis, and Serratia marcenscens [104]. Moreover, solubilization of organic phosphate by phytase activity was observed in Bacillus subtilis, Pseudomonas putida, and P. mendocina [105], and phosphatase activity was discovered in Klebsiella aerogenes [106] and P. fluorescens [107]. However, although P. sonchi SBR5 possesses some of these enzymes associated with phosphate solubilization, the activation of the corresponding genes was not observed when differential gene expression was analyzed under the phosphate solubilization conditions [108].
The production of phosphatases by the soil microbiome was proved to be tightly controlled by the availability of inorganic phosphorus and nitrogen. The addition of nitrogen increases the phosphatase activity [109,110], while the supply of inorganic phosphorus suppresses the production and activity of phosphatases due to the negative feedback mechanism [110].
Phytase-producing fungi are Aspergillus candidus, A. fumigatus, A. niger, A. parasiticus, A. rugulosus, A. terreus, Penicillium rubrum, P. simplicissimum, Pseudeurotium zonatum, Trichoderma harzianum, and Trichoderma viride. Soil Bacillus and Streptomyces spp. are able to mineralize complex organic phosphates by producing extracellular enzymes such as phosphoesterases, phosphodiesterases, phytases, and phospholipases [37].
6. Bacterial Influence on the Root System
Auxin, ethylene, abscisic acid, cytokinins, and gibberellins are well-known phytohormones. PSBs can mediate the changes in the plant root system through the prevention of signaling and auxin transport. For example, Phyllobacterium brassicacearum (STM196) causes changes in the distribution of auxin in the root of Arabidopsis [111].
PSB can also cause changes in plant roots by supplying them with a significant amount of auxin that will act synergistically with endogenous auxin [112] and alter the development of the plant root system [113]. Bacterial production of phytohormones such as auxins and cytokinins can influence cell proliferation, leading to changes in root system architecture through overproduction of lateral roots and root hairs, followed by increased nutrient and water uptake [114]. An increase in root area is associated with an increase in symbiotic interactions with PSB, and thus an increase in phosphorus mobilization. Increased auxin content and higher plant biomass were reported for wheat inoculated with Paenibacillus illinoisensis (IB 1087) and Pseudomonas extremaustralis (IB-Ki-13-1). Thereby, auxin is discovered to play an important role in stimulating root growth and nutrient uptake [113]. Similarly, in corn and sugar cane, inoculation with Paenibacillus sp. and Enterobacter cloacae led to increased biomass of both above-ground and below-ground plant parts [115].
The enzymes produced by PSBs, such as 1-aminocyclopropane-1-carboxylate deaminase, were indicated to contribute indirectly to architectural and functional root modifications triggered by the substrate cleavage [116]. For example, the root biomass of rice plants was strongly correlated with the level of deaminase produced by Alcaligenes sp. [117].
7. Factors Affecting the Microbial Phosphate Solubilization
The ability of phosphate-solubilizing microorganisms to convert insoluble organic and inorganic phosphates is related to the properties of soil. Phosphate-solubilizing microorganisms of the soils under extreme environmental conditions, such as saline-alkaline soils, nutrient-deficient soils or soils from extreme temperature environments, tend to solubilize more phosphate than phosphate-solubilizing microorganisms of soils under temperate conditions [5]. There are conflicting reports on the effect of temperature on phosphate solubilization, with some authors specifying the optimal solubilization temperature of 2025 °C [118], some indicating 28 °C [119], and the others suggesting 30 °C [120]. The ability of phosphate solubilization was demonstrated at an extreme temperature of 45 °C in desert soil [121,122] and at a low temperature of 10 °C [123].
Other factors influencing the microbial phosphate solubilization include the interaction with other soil microorganisms, the stage of plant vegetation, environmental conditions, types of soils of the climatic zone, plant types, agronomic practices, land use systems, and physico-chemical properties of the soil, including the amount of organic matter and pH [124]. Well-aerated soil is more conducive to phosphate solubilization compared to water-saturated moist soil. Among the phosphate-solubilizing bacteria, both anaerobic and aerobic are described. Bacteria of genera Agrobacterium, Bacillus, Burkholderia, Micrococcus, Pseudomonas are aerobic, genera Aeromonas, Enterobacter, Erwinia, Paenibacillus, Serratia are anaerobic of facultative anaerobic.
Adding a small amount of inorganic phosphate to the rhizosphere was indicated to stimulate the mineralization of phytic acid by bacteria and thereby improve the phosphorus nutrition of plants [125]. Also, it was reported that the physiological activity of PSBs is also affected by exogenous soluble phosphate: the growth rate of microorganisms depends on the level of available soluble phosphate [81]. The growth rate of the phosphate-solubilizing strain of Pseudomonas aeruginosa with an excess of phosphate was 25 times higher than in phosphate deficiency case [82]. The strain WS-FJ9 of B. multivorans grew better at high concentrations of soluble phosphate both in Petri dishes and in broth cultures [86], with the same results obtained for the P. aeruginosa P4 [82]. Phosphate-solubilizing activity of the B. multivorans WS-FJ9 strain was found to be significantly suppressed at a concentration of soluble phosphate of 5 mM and completely suppressed at a concentration of 20 mM [86] by a negative feedback mechanism [83]. Sensitivity to the soluble phosphate is found to be a major barrier to the widespread use of PSBs [86].
8. Solubilization of Mineral Phosphates: Molecular Genetics
The molecular genetics basis of mineral phosphate solubilization is not fully understood. With the production of organic acids considered to be the primary mechanism of mineral phosphate solubilization, it is believed that any gene involved in the synthesis of organic acids can influence the solubilization process [18].
It was indicated that the expression of pyrroloquinoline quinone synthase of Erwinia herbicola leads to the secretion of acid and solubilization of mineral phosphates [85]. This enzyme directs the synthesis of pyrroloquinoline quinone, a cofactor necessary for the formation of glucose dehydrogenase holoenzyme, which catalyzes the formation of gluconic acid from glucose by direct oxidation.
Similarly, the gene of mineral phosphate solubilization (gabY) was isolated in Pseudomonas cepacia [126]. Expression of the gabY gene allows the mineral phosphate solubilization phenotype to be induced through the production of gluconic acid. The gabY gene did not indicate a clear homology with the pyrroloquinoline quinone synthetase gene [85]. The gabY gene may play an alternative role in the expression and/or regulation of the direct oxidation pathway in Pseudomonas cepacia by acting as a functional in vivo mineral phosphate-solubilization gene. Thus far, very little is known about the genetic regulation of mineral phosphate-solubilization processes [127].
Most strains of Paenibacillus can solubilize phosphate through the production of gluconic acid. A study of 35 strains, including at least 18 species, indicated all but two strains to contain genes involved in the production of gluconic acid, encoding glucose-1-dehydrogenase and gluconic acid dehydrogenase. Strains apparently lacking these genes are P. beijingensis and P. terrae HPL-003. Strains carrying such genes include P. azotofixans, P. curdlanolyticus, P. dendritiformis, P. elgii, P. forsythia, P. graminis, P. lactis, P. massiliensis, P. mucilaginosus, P. peoriae, P. polymyxa, P. sabinae, P. sonchi, P. sophorae, P. vortex, and P. zanthoxyli. All of the genomes analyzed were found to contain genes for uptake and degradation of phosphonates, containing a highly stable C–P bond, as well as genes for a phosphate-specific transport system [128]. Phosphate solubilization was confirmed for P. elgii, P. kribbensis, P. macerans, P. mucilaginosus, P. polymyxa, P. xylanilyticus and several unclassified strains [129].
8.1. Mineralization of Organic Phosphate: Molecular Genetics
Different patterns of phosphatase activity are widespread in bacteria, especially in Enterobacteriaceae. Phosphatase production is often controlled by complex regulatory mechanisms, resulting in enzyme activity being detected only under certain environmental conditions. However, there is still a lack of full understanding of the properties, regulation, and role of these enzymes. Even in the most thoroughly studied bacteria, Escherichia coli and Salmonella typhimurium, only a few genes have been cloned and studied for their effect on the regulation of phosphatase activity [18].
The primary factor influencing the microbial production of phosphatases is the concentration of inorganic phosphate. This mechanism is best understood for E. coli alkaline phosphatase (PhoA) that is suddenly and completely induced when the inorganic phosphate concentration decreases from 100 mM to 0.16 mM [130]. The gene network includes the phosphate transport operon as a regulatory element, in addition to the sensor-activator operon. The regulation is due to the genes controlled by inorganic phosphate concentration and is activated by PhoB [18].
Another bacterial phosphatase repressed by inorganic phosphate is alkaline phosphatase from Morganella morganii, which is produced in conditions of low inorganic phosphate availability. This alkaline phosphatase is likely similar in its regulation and molecular weight of its polypeptide components to that in E. coli [131]. Pseudomonas fluorescens MF3, Providencia stuartii and P. rettgeri also have alkaline phosphatase activity that is also inhibited by inorganic phosphates [104]. Some studies suggest that the regulation of phosphatase gene expression in other families, such as those belonging to the Enterobacteriaceae family, may be like the E. coli pho genes. These data are based on a high degree of homology of the promotor structures between these genes [132].
Production of PhoN enzyme (acid phosphatase of class A) of Salmonella enterica serovar typhimurium is moderately induced by inorganic phosphate starvation [133]. This gene is under the control of the two-component regulatory system PhoP–PhoQ, promoting the transcription of PhoN and other PhoP genes that are activated at low concentrations of Mg2+ in the environment [134].
Most of the alkaline phosphatases found in the family Enterobacteriaceae are repressed by inorganic phosphate, while many acid phosphatases are unrepressed with phosphates. For some bacterial phosphatases, other regulatory systems were described. For example, the expression of the apo gene encoding the acid phosphatase enzyme in Pseudomonas fluorescens MF3 was determined to be regulated by temperature [135].
The literature data indicate that the regulation of phosphatases is a complex system requiring considerable additional research. In any case, the existing knowledge about bacterial phosphatases provides the basis for better understanding and further investigation of the phosphatase expression in soil bacteria [18].
Nonspecific acid phosphatases (phosphohydrolases or NSAP) released by phosphate-solubilizing microorganisms and dephosphorylating phosphodiester or phosphoanhydride bonds in P-containing substances play an important role in the mineralization of phosphates in most soils [136]. These NSAPs can be both acidic or alkaline phosphomonoesterases [137]. Several NSAP genes were isolated and characterized [136]. For example, the olpA gene of Chryseobacterium meningosepticum encodes a wide range of phosphatases that efficiently hydrolyze monophosphates and phosphorylated sugars [138]. PhoN gene of Deinococcus radiodurans and PhoK gene of E. coli are able to encode periplasmic acid phosphatase and extracellular alkaline phosphatase to enhance the biomineralization of toxic ions in contaminated soils [139]. The cyanobacteria Microcystis aeruginosa contain genes encoding an extracellular alkaline phosphatase that solubilized various inorganic and/or organic phosphates [140]. The phytase enzyme encoded by the appA and phyA genes is another important enzyme for phosphate mineralization responsible for phosphorus release from phytate in the soil [137].
The transcriptome analysis of the phosphate-solubilizing strain of B. multivorans WS-FJ9 isolated from the rhizosphere of Pinus elliotii Engelm indicated the global activation of genes involved in the growth of WS-FJ9 stimulated by high concentrations of soluble phosphate. Among these genes, there are genes encoding the as flagellar MS-ring protein, flagellar hook protein FlgE, and Fe-S protein HscA. However, the phosphorylating pathway of glucose metabolism in B. multivorans WS-FJ9 is also affected by soluble phosphate. The expression of the glycerate kinase gene and the 2-oxoglutarate dehydrogenase gene increased continuously with a decrease in soluble phosphate; phosphate deficiency contributed to the activation of these genes [86]. Glycerate kinase and 2-oxoglutarate dehydrogenase are rate-limiting enzymes in both the Entner–Doudoroff and the tricarboxylic acid cycle pathways [141]. These two pathways of glucose metabolism are successive ones of the phosphorylating pathway [142]. Additionally, pyruvate generated in the glycolysis pathway was found in B. multivorans WS-FJ9, and its level was inversely proportional corresponding to the concentration of soluble phosphate, indicating the repression of the phosphorylating pathway by soluble phosphate [86].
In the Pseudomonas putida CSV86 strain, the phosphorylation pathway was found to be regulated by the glucose metabolizing enzymes and/or the glucose transport process [143]. Genes involved in the ABC-type sugar transporter system were overexpressed in phosphate deficient B. multivorans WS-FJ9 [86], which was considered an important transport system for glucose uptake and transport [144]. Therefore, soluble phosphate was found to negatively regulate the glucose phosphorylation pathway in B. multivorans WS-FJ9, which may affect the organic acid secretion and, consequently, phosphate-solubilization activity.
Additionally, the gene of the histidine protein kinase PhoR was found to be activated by the low concentration of soluble phosphate [86]. PhoR is considered an important part of the two-component regulatory operon PhoB–PhoR (Pho Regulon) [145]. This two-component transcription factor regulates the expression of several Pho regulon genes, such as the PhoA and PhoB alkaline phosphatase genes [146] and the gene of APase-alkaline diphosphoesterase PhoD [147] encoding enzymes involved in the mineralization of organic phosphates.
In addition to the genes involved in phosphate solubilization, the expression levels of many transcriptional regulatory genes and genes involved in signal transduction are regulated at various concentrations of soluble phosphate. The transcription regulator of the LysR family negatively regulates the set of genes, including those involved in bacterial virulence, metabolism, and motility [148]. TonB-dependent regulatory systems perceive signals outside the bacterial cell and transmit them to the cytoplasm, where the TonB-dependent receptor is involved in both transport [149] and signal conversion [150]. The expression levels of these genes are regulated by soluble phosphate. These genes both directly and indirectly contribute to the response to exogenous soluble phosphate.
The transcriptome analysis revealed a complex response of B. multivorans WS-FJ9 to the various levels of exogenous soluble phosphate. This finding may help in understanding the molecular mechanism behind the regulation of soluble phosphate on the physiological activity of PSB, especially on phosphate solubilization. This knowledge is expected to provide the basis for a molecular engineering strategy to reduce the sensitivity of PSB strains to soluble phosphate, which could improve their relevant biological activity.
8.2. Regulation of Inorganic Phosphate in Bacterial Cells
The phosphate (Pho) regulon is a global regulatory mechanism involved in the control of bacterial inorganic phosphate, which was first described for Escherichia coli and later for many other bacterial species [151]. Pho-regulon activates extracellular enzymes generating inorganic phosphate from organic ones, phosphate-specific carriers, and enzymes involved in phosphate storage and preservation. The most conserved member of the Pho regulon in the bacteria is the Pi-specific transporter (Pst) [152]. The most common enzymes induced in response to inorganic phosphate starvation in bacteria are alkaline phosphatases (PhoA), phospholipases (PhoD), glycerophosphodiester phosphodiesterases, phytases, and 5’-nucleotidase [151]. To store inorganic phosphates, most bacteria induce the expression of polyphosphate kinases, which are able to accumulate polyphosphate and, if necessary, reuse it [153]. To conserve nutrients, some bacteria can replace teichoic acids that are rich in inorganic phosphate polymers found in the cell wall of Gram-positive bacteria with teichuronic acids not containing phosphates [154].
The Pho regulon is controlled by a two-component regulatory system that includes an inner membrane histidine kinase sensor protein and a regulator of the cytoplasmic transcriptional response. These proteins were named differently in some bacteria, for example, PhoR–PhoB in E. coli [155], PhoR–PhoP in Bacillus subtilis [156], PnpR–PnpS in Streptococcus pneumoniae [157], etc. In all cases of inorganic phosphate deficiency, the response regulator is phosphorylated at the aspartic acid residue by the sensor kinase. The phosphorylated response regulator can bind to the specific DNA sequences and activate or inhibit gene transcription. These specific sequences, known as PHO boxes, were first characterized in E. coli [158] as the sum of two 11-nt direct repeat units, each consisting of seven highly-conserved and four less-conserved nucleotides [159]. Similar features were observed in many other bacteria, including B. subtilis [156], Sinorhizobium meliloti [160], Corynebacterium glutamicum, and Streptomyces coelicolor [10.1128/JB.00121-07]. The consensus sequence of the PHO box is found to significantly vary between bacteria [159].
The depletion of inorganic phosphate in the medium is important for the activation of the Pho regulon in bacteria [161]. In E. coli, in addition to the two-component PhoR–PhoB system, the inorganic phosphate uptake pathway requires five more proteins, four of them being the components of the Pst and one a component of the PhoU metal-binding protein. When concentration of inorganic phosphate is limited, PhoB is activated by PhoR acting as a kinase, but under conditions of excess free phosphate, PhoB activation is interrupted by PhoR acting as a phosphatase [162]. PhoU is required for PhoB dephosphorylation under phosphate-rich conditions [163].
Although PhoU is found in many bacterial genomes, this gene is absent in B. subtilis. The inorganic phosphate-signaling network in this bacterium includes a positive feedback loop between the PhoP-PhoR and ResD-ResE two-component systems [164]. ResD does not bind to the phoPR operon and appears to transfer its control through the expression of terminal oxidases [165].
8.3. Interaction of PSB with the Plant Roots
Most plants express two types of inorganic phosphate transporters to maintain intracellular phosphate concentrations required for optimal cell function. These genes (PHT) are divided into two families: (1) high-affinity PHT1 transporters expressed in the roots and (2) low-affinity PHT2 transporters responsible for the intracellular transport of phosphates in plant shoots [166]. Several PSB species have been identified as regulating the expression of specific plant genes. PHT1 gene expression in plant roots has been found to be modulated by Pseudomonas putida (MTCC 5279) which is known to increase the expression of PT1, the member of the PHT1 gene family in Arabidopsis thaliana roots during phosphate deficiency and salt stress [167]. Under the combined stress and in the presence of P. putida, the expression of PT2 also increased. However, the expression of PHO2 (the gene responsible for the accumulation of phosphate in shoots) was suppressed due to the PHO2 gene playing an important role in phosphate-signaling under low phosphate conditions.
Similarly, increased phosphorus uptake in wheat inoculated with a consortium of Bacillus species was found to result from increased expression of the PHT1 transporter gene in the roots [168]. Thus, rhizospheric PSB can trigger the changes in phosphate transporter gene expression either directly by altering plant metabolism associated with phytohormone production or indirectly by increasing phosphate availability. These molecular changes in roots (up- and down-regulation of phosphate transporters) have now been partially proven to be induced by PSB, positively influencing the intake of phosphate and subsequently reflecting a positive effect on plant growth and yield.
9. PSB Co-Inoculation of Plants and PSB Synergy with the Use of Phosphate Fertilizers
Inoculation with a mixture of PSBs belonging to Bacillus, Pseudomonas, and Streptomyces genera demonstrated better results in terms of plant growth and establishment than a single inoculation. Co-inoculation of wheat with phosphate-solubilizing and auxin-producing rhizobacteria Pseudomonas fluorescens Ms-01 and Azosprillum brasilense DSM1690 increased the biomass of roots and shoots compared with a single inoculation with the same bacteria [169]. Similarly, inoculation of amaranth plants with a consortium of three bacteria [Bacillus firmus (KUCr1), Cellulosimicrobium cellulans (KUCr3), and Pseudomonas aeruginosa (KUCd1)] increased the total root length and overall plant growth compared to mono- and double inoculation with different strains [170].
The studies indicate that the synergism of combining PSBs and phosphate fertilizers may increase the agronomic efficiency of the fertilizers in soils [171,172]. For example, maize growth was greatly improved by PSB inoculation in combination with various types of inorganic and organic phosphate fertilizers: livestock manure, bird droppings, single superphosphate, and rock phosphates [173]. The improvement of maize growth parameters was observed in response to inoculation with phosphate-solubilizing Enterobacter sakazakii J129 in combination with NPK fertilizer [174]. Also, increased phosphate uptake and utilization efficiency were reported for the wheat inoculated with Pseudomonas sp. or Enterobacter sp. along with the diammonium phosphate- based fertilizer [175].
The use of PSBs is indicating promise as a biotechnological intervention to improve the agronomic efficiency of rock phosphates as direct phosphate fertilizers. Enterobacter sp. was proven to be capable of increasing the solubilization of the phosphate from rock-phosphate up to 17.5% [176]. Also, PSBs combined with rock phosphate were proven to significantly increase the availability of phosphate in the soil (27%) compared to the release from rock phosphate alone (4%) [177]. A single inoculation of wheat plants with five different strains of Pseudomonas (P. plecoglossicida, P. reinekei, P. koreensis, P. japonica, and P. frederiksbergensis) in combination with rock-phosphate fertilizer was observed to increase phosphate bioavailability, nutrient uptake, chlorophyll content, morphological characteristics, and root structure associated with higher phosphate uptake compared to non-inoculated plants [178]. Moreover, using a consortium of three strains (Pseudomonas corrugata SP77, P. koreensis LT62 and P. frederiksbergensis G62) was also identified as having positive effects on plant growth, e.g., in enhancing the growth of Medicago truncatula when combined with rock-phosphate [179].
Not only are PSBs known to increase the agronomic efficiency of mineral phosphorus fertilizers, but also combinations of these bacteria with organic P-fertilizers have been reported. For example, the combined use of bioorganic phosphorus (a mixture of compost, biogas residues) and Bacillus sp. was found to increase the availability of phosphate in the soil and the PSB population [180]. Similarly, growth parameters of chili plants were proven to improve in response to PSB inoculation with simultaneous application of organic phosphate fertilizers (eggshell, tea powder, animal bone waste) [181]. Moreover, partial replacement of mineral fertilizers with organic fertilizers can increase the availability of phosphate in the soil and change the shape of the PSB community in the soil, leading to improved phosphate mineralization and its uptake by rice plants [182].
Thus, the inoculation of plants with PSBs in combination with mineral or organic fertilizers is regarded as a promising integrated strategy to increase the availability of phosphates by increasing the agronomic efficiency of using phosphate fertilizers to improve soil fertility and support agriculture.
Symbiotic Interactions between Phosphate-Solubilizing and Nitrogen-Fixing Microorganisms
Some microorganisms at the soil–root interface establish the nitrogen-fixing symbiosis with legumes through the formation of nodules on roots or stems [183]. Biological nitrogen fixation is a symbiotic process [184] during which plants provide carbon as an energy source for symbiotic microorganisms in exchange for bacteria-derived nitrogen compounds that can be easily assimilated by the host plant [185]. Almost 80% of the nitrogen available to plants originates from a symbiotic group, including Rhizobium which invades plants to form nodules. The non-symbiotic group consists of free-living microorganisms, such as Bacillus, Azotobacter, Azospirillum, and Herbaspirillum, and others contributing approximately 5–10% of the biological input of nitrogen to the soil [186].
The effectiveness of the biological fixation of nitrogen in cereal legumes was proven to be significantly limited by phosphate availability [187]. A significant correlation between nodal phosphate content and nitrogen fixation was identified. An adequate supply of phosphates to plants was found to regulate several metabolic processes associated with nitrogen fixation, such as the assimilation of ammonium into amino acids and ureides [188] and the synthesis of mitochondrial and symbiosomal membranes for nitrogen-fixing nodules [189]. In general, the formation and functioning of nodules and the energy costs of nitrogen fixation in legumes are highly dependent on the phosphate status of plants and in nodule tissues, and inadequate phosphate status reduces the potential contribution of nitrogen fixation [171].
Nitrogen fixation, as one of the key biological processes of the rhizosphere, is known to correlate with phosphate availability, with phosphate deficiency severely limiting the activity of diazotrophic (nitrogen-fixing bacteria) and reducing the symbiotic partnership between the host plant and rhizobia, in addition to the nitrogen fixation process itself [190]. As a rule, phosphate deficiency impairs the formation, development, and function of nodules by reducing the energy available to support the metabolic activity of nodules [191]. Plants living in symbiosis with nitrogen-fixing bacteria have a higher phosphorus demand than plants that do not have such symbionts [192]. The increased phosphorus demand in plants is generally due to the phosphates required for the nitrogen fixation processes [188,193]. However, other studies suggest that the high phosphate requirement is due to the needs of nitrogen-fixing bacteria, whose phosphorus requirements were found to be higher than those of non-nitrogen-fixing bacteria [172]. The availability of various phospholipids (such as phosphatidylserine, phosphatidylcholine, and phosphatidylethanolamine) may be crucial in maintaining the symbiosis between legumes and rhizobia [194]. These compounds are required to create the correct rhizobia membrane architecture to effectively participate in BNF [195,196].
Therefore, introducing phosphate-solubilizing microbes to nutrient-deficient soils, where the nutrients are stored in biologically inaccessible forms, could help farmers improve crop yields and obtain economic benefits through the rational use of phosphate fertilizers and the increase of nitrogen fixation. For example, the combination of nitrogen-fixing Rhizobium and phosphate-solubilizing Bacillus megatherium was observed to increase the yield and quality of Vicia faba seeds and the yield, nodule biomass, shoot biomass, and nitrogen and phosphorus content of Lablab purpureus [197].
The synergy between soil phosphate-solubilizing bacteria and rhizobial strains of nodule legumes is an important rhizosphere process that is worth investigating. Several studies have reported that the application of phosphate-solubilizing microorganisms, including PSB, to the soil, individually or in the consortia, increased plant growth by solubilization of insoluble phosphates and subsequent increase of nitrogen fixation [196]. Phosphate-solubilizing Serratia, combined with nitrogen-fixing Mesorhizobium ciceri and arbuscular mycorrhiza fungus Glomus fasciculatum, increased the growth of cereal legumes [191]. Similarly, co-inoculation of wheat with a diazotrophic Paenibacillus beijingensis BJ-18) and phosphate-solubilizing Paenibacillus sp. B1 significantly increased plant growth and the content of phosphates and nitrogen in the plant (roots and shoots) and the soil [198]. Furthermore, the inoculation of chickpeas with nitrogen-fixing Mesorhizobium ciceri, phosphate-solubilizing fungus (Penicillium WF6), and/or PSB Serratia T1 resulted in increased availability and uptake of phosphates and positively affected nitrogen fixation [199].
According to several studies, phosphate solubilization may be one of the main mechanisms by which PSBs enhance nitrogen fixation [200]. Bioavailable phosphates positively affect the symbiosis of legumes and rhizobia. Legumes treated with biofertilizers containing consortia of three PSBs: Pseudomonas sp., Burkholderia sp., and Enterobacter sp. in combination with rock phosphate had increased phosphate content both in the soil and plants, providing favorable conditions for nodulation, which in turn increased the efficiency of the nitrogen fixation [171].
Despite numerous reports highlighting interactions between phosphate-solubilizing and nitrogen-fixing bacteria, the mechanisms underlying these associations are not yet fully understood.
10. Perspectives
To enhance the use of PSBs as effective and important components in sustainable soil management systems, more data in needed on the molecular mechanisms they use to increase the bioavailability of phosphates. Consumers pay attention to the health, quality, and nutritional value of agricultural products. Thus, applying phosphate-solubilizing microorganisms as biofertilizers is one option to increase food production without posing a health risk, while saving natural sources of phosphate fertilizers and developing sustainable agriculture. It is important for researchers to continue studying phosphate-solubilizing microorganisms and translate this knowledge into a form that can be easily used by farmers [137].
11. Conclusions
The papers published in the recent years indicate that the efficiency of using phosphates in agriculture could be improved by inoculation with phosphate-solubilizing bacteria, which increase the availability of phosphate without disturbing the biochemical composition of the soil. These potential biofertilizers are universal since they can be used for various crops and generally are not specific for plants. Conversely, the development of personalized, plant-specific consortia of phosphate-solubilizers is likely to increase productivity further. Inoculation of phosphate-solubilizing microorganisms into the soil appears to be an effective way to convert insoluble phosphate compounds into bioavailable forms, resulting in better plant growth, yield, and quality. In this review, we demonstrate that the bacteria of genera Bacillus, Pseudomonas, Rhizobium, Aspergillus, and Penicillium genera are regarded as the most effective phosphate-solubilizers for increasing the phosphate bioavailability in soil. PSBs cause immediate plant growth by providing phosphates in an easily absorbable form. Additionally, phosphate-solubilizing microorganisms support plant growth by increasing the efficiency of nitrogen fixation. Two main mechanisms increasing free phosphate in the soil can be distinguished: the mobilization of inorganic and organic phosphates. In general, the mobilization of inorganic phosphate is performed primarily due to the release of organic acids by bacteria, and enzymes released into the extracellular environment perform the mineralization of organic phosphates.
While most farmers rely on inorganic sources of phosphates with which to avoid nutrient deficiencies, significant amounts of these fertilizers are lost from the soil through various mechanisms and are unavailable for plant uptake. In legume cropping systems, phosphate deficiency can also lead to nitrogen deficiency and reduced crop yields. Since the PSB-based biofertilizers have indicated promising effects on plant growth and yield, we assume that the phosphate-solubilizing microorganisms may be potential substitutes for inorganic phosphate fertilizers as a method to meet plant requirements and consequently increase the yields in sustainable agriculture. Their application is an environmentally and economically sound approach.
In this review, we described the molecular mechanisms of use phosphate-solubilizing microorganisms as biofertilizers. Solubilization of inorganic phosphate in the soil is found to increase its bioavailability for the plant, thus using PSBs in the soil is to promote sustainable agriculture, improve soil fertility, and increase crop yields. We consider use of PSB as microbial inoculants as the new frontier for increasing plant productivity. This technology can contribute to low-cost farming systems and a cleaner environment.
Author Contributions
Conceptualization, A.T., M.G. and S.S.; writing—original draft preparation, A.T.; writing—review and editing, S.S.; visualization, A.T.; funding acquisition, M.G. and S.S. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Ministry of Science and Higher Education of the Russian Federation, agreement No. 075-15-2021-1085.
Institutional Review Board Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Maharajan, T.; Ceasar, S.A.; Krishna, T.P.A.; Ramakrishnan, M.; Duraipandiyan, V.; Abdulla, A.-D.N.; Ignacimuthu, S. Utilization of molecular markers for improving the phosphorus efficiency in crop plants. Plant Breed. 2018, 137, 10–26. [Google Scholar] [CrossRef]
- Santana, E.B.; Marques, E.L.S.; Dias, J.C.T. Effects of phosphate-solubilizing bacteria, native microorganisms, and rock dust on Jatropha curcas L. growth. Genet. Mol. Res. 2016, 15, 1–18. [Google Scholar] [CrossRef]
- Sharma, S.B.; Sayyed, R.Z.; Trivedi, M.H.; Gobi, T.A. Phosphate solubilizing microbes: Sustainable approach for managing phosphorus deficiency in agricultural soils. Springerplus 2013, 2, 587. [Google Scholar] [CrossRef]
- Nesme, T.; Metson, G.S.; Bennett, E.M. Global phosphorus flows through agricultural trade. Glob. Environ. Chang. 2018, 50, 133–141. [Google Scholar] [CrossRef]
- Zhu, F.; Qu, L.; Hong, X.; Sun, X. Isolation and Characterization of a Phosphate-Solubilizing Halophilic Bacterium Kushneriasp. YCWA18 from Daqiao Saltern on the Coast of Yellow Sea of China. Evid. Based Complementary Altern. Med. 2011, 2011, 615032. [Google Scholar] [CrossRef]
- Ai, P.; Sun, S.; Zhao, J.; Fan, X.; Xin, W.; Guo, Q.; Yu, L.; Shen, Q.; Wu, P.; Miller, A.J.; et al. Two rice phosphate transporters, OsPht1;2 and OsPht1;6, have different functions and kinetic properties in uptake and translocation. Plant J. 2009, 57, 798–809. [Google Scholar] [CrossRef] [PubMed]
- Azziz, G.; Bajsa, N.; Haghjou, T.; Taulé, C.; Valverde, Á.; Igual, J.M.; Arias, A. Abundance, diversity and prospecting of culturable phosphate solubilizing bacteria on soils under crop–pasture rotations in a no-tillage regime in Uruguay. Appl. Soil Ecol. 2012, 61, 320–326. [Google Scholar] [CrossRef]
- Schnug, E.; Haneklaus, S.H. The Enigma of Fertilizer Phosphorus Utilization. In Phosphorus in Agriculture: 100% Zero; Springer: Dordrecht, The Netherlands, 2016; pp. 7–26. [Google Scholar]
- Chaney, R.L. Food Safety Issues for Mineral and Organic Fertilizers. Adv. Agron. 2012, 117, 51–116. [Google Scholar] [CrossRef]
- Bhattacharyya, P.N.; Jha, D.K. Plant growth-promoting rhizobacteria (PGPR): Emergence in agriculture. World J. Microbiol. Biotechnol. 2012, 28, 1327–1350. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.S.; Zaidi, A.; Wani, P.A. Role of phosphate-solubilizing microorganisms in sustainable agriculture—A review. Agron. Sustain. Dev. 2007, 27, 29–43. [Google Scholar] [CrossRef]
- Owen, D.; Williams, A.P.; Griffith, G.W.; Withers, P.J.A. Use of commercial bio-inoculants to increase agricultural production through improved phosphrous acquisition. Appl. Soil Ecol. 2015, 86, 41–54. [Google Scholar] [CrossRef]
- Wani, P.; Khan, M.; Zaidi, A. Co-inoculation of nitrogen-fixing and phosphate-solubilizing bacteria to promote growth, yield and nutrient uptake in chickpea. Acta Agron. Hung. 2007, 55, 315–323. [Google Scholar] [CrossRef]
- Wang, H.-Y.; Liu, S.; Zhai, L.-M.; Zhang, J.-Z.; Ren, T.-Z.; Fan, B.-Q.; Liu, H.-B. Preparation and utilization of phosphate biofertilizers using agricultural waste. J. Integr. Agric. 2015, 14, 158–167. [Google Scholar] [CrossRef]
- Alori, E.; Fawole, O.; Afolayan, A. Characterization of Arbuscular Mycorrhizal Spores Isolated from Southern Guinea Savanna of Nigeria. J. Agric. Sci. 2012, 4, 13. [Google Scholar] [CrossRef]
- Delavaux, C.S.; Smith-Ramesh, L.M.; Kuebbing, S.E. Beyond nutrients: A meta-analysis of the diverse effects of arbuscular mycorrhizal fungi on plants and soils. Ecology 2017, 98, 2111–2119. [Google Scholar] [CrossRef]
- Browne, P.; Rice, O.; Miller, S.H.; Burke, J.; Dowling, D.N.; Morrissey, J.P.; O’Gara, F. Superior inorganic phosphate solubilization is linked to phylogeny within the Pseudomonas fluorescens complex. Appl. Soil Ecol. 2009, 43, 131–138. [Google Scholar] [CrossRef]
- Rodríguez, H.; Fraga, R. Phosphate solubilizing bacteria and their role in plant growth promotion. Biotechnol. Adv. 1999, 17, 319–339. [Google Scholar] [CrossRef]
- Jha, A.; Sharma, D.; Saxena, J. Effect of single and dual phosphate-solubilizing bacterial strain inoculations on overall growth of mung bean plants. Arch. Agron. Soil Sci. 2012, 58, 967–981. [Google Scholar] [CrossRef]
- Srinivasan, R.; Yandigeri, M.S.; Kashyap, S.; Alagawadi, A.R. Effect of salt on survival and P-solubilization potential of phosphate solubilizing microorganisms from salt affected soils. Saudi J. Biol. Sci. 2012, 19, 427–434. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Bauddh, K.; Barman, S.C.; Singh, R.P. Amendments of microbial biofertilizers and organic substances reduces requirement of urea and DAP with enhanced nutrient availability and productivity of wheat (Triticum aestivum L.). Ecol. Eng. 2014, 71, 432–437. [Google Scholar] [CrossRef]
- Jiang, Z.; Zhang, X.; Wang, Z.; Cao, B.; Deng, S.; Bi, M.; Zhang, Y. Enhanced biodegradation of atrazine by Arthrobacter sp. DNS10 during co-culture with a phosphorus solubilizing bacteria: Enterobacter sp. P1. Ecotoxicol. Environ. Saf. 2019, 172, 159–166. [Google Scholar] [CrossRef] [PubMed]
- Jahan, M.; Mahallati, M.N.; Amiri, M.B.; Ehyayi, H.R. Radiation absorption and use efficiency of sesame as affected by biofertilizers inoculation in a low input cropping system. Ind. Crop. Prod. 2013, 43, 606–611. [Google Scholar] [CrossRef]
- Zhao, L.; Zhang, Y.-Q. Effects of phosphate solubilization and phytohormone production of Trichoderma asperellum Q1 on promoting cucumber growth under salt stress. J. Integr. Agric. 2015, 14, 1588–1597. [Google Scholar] [CrossRef]
- Postma, J.; Nijhuis, E.H.; Someus, E. Selection of phosphorus solubilizing bacteria with biocontrol potential for growth in phosphorus rich animal bone charcoal. Appl. Soil Ecol. 2010, 46, 464–469. [Google Scholar] [CrossRef]
- Mamta; Rahi, P.; Pathania, V.; Gulati, A.; Singh, B.; Bhanwra, R.K.; Tewari, R. Stimulatory effect of phosphate-solubilizing bacteria on plant growth, stevioside and rebaudioside-A contents of Stevia rebaudiana Bertoni. Appl. Soil Ecol. 2010, 46, 222–229. [Google Scholar] [CrossRef]
- Chakraborty, U.; Chakraborty, B.N.; Basnet, M.; Chakraborty, A.P. Evaluation of Ochrobactrum anthropic TRS-2 and its talc based formulation for enhancement of growth of tea plants and management of brown root rot disease. J. Appl. Microbiol. 2009, 107, 625–634. [Google Scholar] [CrossRef]
- Dastager, S.G.; Deepa, C.K.; Pandey, A. Isolation and characterization of novel plant growth promoting Micrococcus sp. NII-0909 and its interaction with cowpea. Plant Physiol. Biochem. 2010, 48, 987–992. [Google Scholar] [CrossRef]
- Bidondo, L.F.; Silvani, V.; Colombo, R.; Pérgola, M.; Bompadre, J.; Godeas, A. Pre-symbiotic and symbiotic interactions between Glomus intraradices and two Paenibacillus species isolated from AM propagules. In Vitro and in vivo assays with soybean (AG043RG) as plant host. Soil Biol. Biochem. 2011, 43, 1866–1872. [Google Scholar] [CrossRef]
- Oteino, N.; Lally, R.D.; Kiwanuka, S.; Lloyd, A.; Ryan, D.; Germaine, K.J.; Dowling, D.N. Plant growth promotion induced by phosphate solubilizing endophytic Pseudomonas isolates. Front. Microbiol. 2015, 6, 745. [Google Scholar] [CrossRef]
- Paul, D.; Sinha, S.N. Isolation and characterization of phosphate solubilizing bacterium Pseudomonas aeruginosa KUPSB12 with antibacterial potential from river Ganga, India. Ann. Agrar. Sci. 2017, 15, 130–136. [Google Scholar] [CrossRef]
- Tajini, F.; Trabelsi, M.; Drevon, J.-J. Combined inoculation with Glomus intraradices and Rhizobium tropici CIAT899 increases phosphorus use efficiency for symbiotic nitrogen fixation in common bean (Phaseolus vulgaris L.). Saudi J. Biol. Sci. 2012, 19, 157–163. [Google Scholar] [CrossRef] [PubMed]
- Korir, H.; Mungai, N.W.; Thuita, M.; Hamba, Y.; Masso, C.; Tola, Y.H. Co-inoculation Effect of Rhizobia and Plant Growth Promoting Rhizobacteria on Common Bean Growth in a Low Phosphorus Soil. Front. Plant Sci. 2017, 8, 141. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Bai, T.; Dai, L.; Wang, F.; Tao, J.; Meng, S.; Hu, Y.; Wang, S.; Hu, S. A study of organic acid production in contrasts between two phosphate solubilizing fungi: Penicillium oxalicum and Aspergillus niger. Sci. Rep. 2016, 6, 25313. [Google Scholar] [CrossRef]
- Behera, B.C.; Yadav, H.; Singh, S.K.; Mishra, R.R.; Sethi, B.K.; Dutta, S.K.; Thatoi, H.N. Phosphate solubilization and acid phosphatase activity of Serratia sp. isolated from mangrove soil of Mahanadi river delta, Odisha, India. J. Genet. Eng. Biotechnol. 2017, 15, 169–178. [Google Scholar] [CrossRef] [PubMed]
- Thampi, A.; Bhai, R.S. Rhizosphere actinobacteria for combating Phytophthora capsici and Sclerotium rolfsii, the major soil borne pathogens of black pepper (Piper nigrum L.). Biol. Control 2017, 109, 1–13. [Google Scholar] [CrossRef]
- Kalayu, G. Phosphate Solubilizing Microorganisms: Promising Approach as Biofertilizers. Int. J. Agron. 2019, 2019, 4917256. [Google Scholar] [CrossRef]
- Arai, Y.; Sparks, D. Phosphate Reaction Dynamics in Soils and Soil Components: A Multiscale Approach. Adv. Agron. 2007, 94, 135–179. [Google Scholar] [CrossRef]
- Devau, N.; Le Cadre, E.; Hinsinger, P.; Jaillard, B.; Gérard, F. Soil pH controls the environmental availability of phosphorus: Experimental and mechanistic modelling approaches. Appl. Geochem. 2009, 24, 2163–2174. [Google Scholar] [CrossRef]
- Mnkeni, P.N.S.; MacKenzie, A.F. Retention of Ortho- and Polyphosphates in Some Quebec Soils as Affected by Added Organic Residues and Calcium Carbonate. Can. J. Soil Sci. 1985, 65, 575–585. [Google Scholar] [CrossRef]
- Hallama, M.; Pekrun, C.; Lambers, H.; Kandeler, E. Hidden miners—The roles of cover crops and soil microorganisms in phosphorus cycling through agroecosystems. Plant Soil 2019, 434, 7–45. [Google Scholar] [CrossRef]
- Ehlers, K.; Bakken, L.R.; Frostegård, Å.; Frossard, E.; Bünemann, E.K. Phosphorus limitation in a Ferralsol: Impact on microbial activity and cell internal P pools. Soil Biol. Biochem. 2010, 42, 558–566. [Google Scholar] [CrossRef]
- Richardson, A.E.; Simpson, R.J. Soil Microorganisms Mediating Phosphorus Availability Update on Microbial Phosphorus. Plant Physiol. 2011, 156, 989–996. [Google Scholar] [CrossRef] [PubMed]
- Kulaev, I.S.; Vagabov, V.M. Polyphosphate Metabolism in Micro-Organisms. Adv. Microb. Physiol. 1983, 24, 83–171. [Google Scholar] [CrossRef] [PubMed]
- Hirota, R.; Kuroda, A.; Kato, J.; Ohtake, H. Bacterial phosphate metabolism and its application to phosphorus recovery and industrial bioprocesses. J. Biosci. Bioeng. 2010, 109, 423–432. [Google Scholar] [CrossRef] [PubMed]
- Kuroda, A.; Murphy, H.; Cashel, M.; Kornberg, A. Guanosine Tetra- and Pentaphosphate Promote Accumulation of Inorganic Polyphosphate in Escherichia coli. J. Biol. Chem. 1997, 272, 21240–21243. [Google Scholar] [CrossRef]
- Moradali, M.F.; Rehm, B.H.A. Bacterial biopolymers: From pathogenesis to advanced materials. Nat. Rev. Microbiol. 2020, 18, 195–210. [Google Scholar] [CrossRef]
- Karl, D.M. Microbially Mediated Transformations of Phosphorus in the Sea: New Views of an Old Cycle. Annu. Rev. Mar. Sci. 2014, 6, 279–337. [Google Scholar] [CrossRef]
- Seeling, B.; Zasoski, R.J. Microbial effects in maintaining organic and inorganic solution phosphorus concentrations in a grassland topsoil. Plant Soil 1993, 148, 277–284. [Google Scholar] [CrossRef]
- Khan, K.S.; Joergensen, R.G. Changes in microbial biomass and P fractions in biogenic household waste compost amended with inorganic P fertilizers. Bioresour. Technol. 2009, 100, 303–309. [Google Scholar] [CrossRef]
- Sugito, T.; Yoshida, K.; Takebe, M.; Shinano, T.; Toyota, K. Soil microbial biomass phosphorus as an indicator of phosphorus availability in a Gleyic Andosol. Soil Sci. Plant Nutr. 2010, 56, 390–398. [Google Scholar] [CrossRef]
- Spohn, M.; Widdig, M. Turnover of carbon and phosphorus in the microbial biomass depending on phosphorus availability. Soil Biol. Biochem. 2017, 113, 53–59. [Google Scholar] [CrossRef]
- Malik, M.A. Phosphorus fractions, microbial biomass and enzyme activities in some alkaline calcareous subtropical soils. Afr. J. Biotechnol. 2012, 11, 4773–4781. [Google Scholar] [CrossRef]
- Chen, G.C.; He, Z.L. Microbial biomass phosphorus turnover in variable-charge soils in China. Commun. Soil Sci. Plant Anal. 2002, 33, 2101–2117. [Google Scholar] [CrossRef]
- Dinh, M.-V.; Guhr, A.; Spohn, M.; Matzner, E. Release of phosphorus from soil bacterial and fungal biomass following drying/rewetting. Soil Biol. Biochem. 2017, 110, 1–7. [Google Scholar] [CrossRef]
- Patel, D.K.; Archana, G.; Kumar, G.N. Variation in the Nature of Organic Acid Secretion and Mineral Phosphate Solubilization by Citrobacter sp. DHRSS in the Presence of Different Sugars. Curr. Microbiol. 2008, 56, 168–174. [Google Scholar] [CrossRef]
- Vijayakumar, A.; Rajasekharan, R. Distinct Roles of Alpha/Beta Hydrolase Domain Containing Proteins. Biochem. Mol. Biol. J. 2016, 2, 1–3. [Google Scholar] [CrossRef]
- Son, H.-J.; Park, G.-T.; Cha, M.-S.; Heo, M.-S. Solubilization of insoluble inorganic phosphates by a novel salt- and pH-tolerant Pantoea agglomerans R-42 isolated from soybean rhizosphere. Bioresour. Technol. 2006, 97, 204–210. [Google Scholar] [CrossRef]
- Zhao, K.; Penttinen, P.; Zhang, X.; Ao, X.; Liu, M.; Yu, X.; Chen, Q. Maize rhizosphere in Sichuan, China, hosts plant growth promoting Burkholderia cepacia with phosphate solubilizing and antifungal abilities. Microbiol. Res. 2014, 169, 76–82. [Google Scholar] [CrossRef]
- Asea, P.; Kucey, R.; Stewart, J. Inorganic phosphate solubilization by two Penicillium species in solution culture and soil. Soil Biol. Biochem. 1988, 20, 459–464. [Google Scholar] [CrossRef]
- Illmer, P.; Schinner, F. Solubilization of inorganic calcium phosphates—Solubilization mechanisms. Soil Biol. Biochem. 1995, 27, 257–263. [Google Scholar] [CrossRef]
- Kouas, S.; Labidi, N.; Debez, A.; Abdelly, C. Effect of P on nodule formation and N fixation in bean. Agron. Sustain. Dev. 2005, 25, 389–393. [Google Scholar] [CrossRef]
- Taha, S.M.; Mahmoud, S.A.Z.; El-Damaty, A.H.; El-Hafez, A.M. Activity of phosphate-dissolving bacteria in Egyptian soils. Plant Soil 1969, 31, 149–160. [Google Scholar] [CrossRef]
- Bajpai, P.D.; Rao, W.V.B.S. Phosphate solubilising bacteria. Soil Sci. Plant Nutr. 1971, 17, 46–53. [Google Scholar] [CrossRef]
- Banik, S.; Dey, B.K. Phosphate-Solubilizing Potentiality of the Microorganisms Capable of Utilizing Aluminium Phosphate as a Sole Phosphate Source. Zentralbl. Mikrobiol. 1983, 138, 17–23. [Google Scholar] [CrossRef]
- Singal, R.; Gupta, R.; Saxena, R.K. Rock phosphate solubilization under alkaline conditions by Aspergillus japonicus and A. foetidus. Folia Microbiol. 1994, 39, 33–36. [Google Scholar] [CrossRef] [PubMed]
- Whitelaw, M.A.; Harden, T.J.; Helyar, K.R. Phosphate solubilisation in solution culture by the soil fungus Penicillium radicum. Soil Biol. Biochem. 1999, 31, 655–665. [Google Scholar] [CrossRef]
- Kim, K.Y.; McDonald, G.A.; Jordan, D. Solubilization of hydroxyapatite by Enterobacter agglomerans and cloned Escherichia coli in culture medium. Biol. Fertil. Soils 1997, 24, 347–352. [Google Scholar] [CrossRef]
- Vazquez, P.; Holguin, G.; Puente, M.E.; Lopez-Cortes, A.; Bashan, Y. Phosphate-solubilizing microorganisms associated with the rhizosphere of mangroves in a semiarid coastal lagoon. Biol. Fertil. Soils 2000, 30, 460–468. [Google Scholar] [CrossRef]
- Reyes, I.; Baziramakenga, R.; Bernier, L.; Antoun, H. Solubilization of phosphate rocks and minerals by a wild-type strain and two UV-induced mutants of Penicillium rugulosum. Soil Biol. Biochem. 2001, 33, 1741–1747. [Google Scholar] [CrossRef]
- Kim, K.Y.; Hwangbo, H.; Park, R.D.; Kim, Y.W.; Rim, Y.S.; Park, K.H.; Kim, T.H.; Suh, J.S. 2-Ketogluconic Acid Production and Phosphate Solubilization by Enterobacter intermedium. Curr. Microbiol. 2003, 47, 87–92. [Google Scholar] [CrossRef]
- Rashid, M.; Khalil, S.; Ayub, N.; Alam, S.; Latif, F. Organic Acids Production and Phosphate Solubilization by Phosphate Solubilizing Microorganisms (PSM) Under in vitro Conditions. Pak. J. Biol. Sci. 2004, 7, 187–196. [Google Scholar] [CrossRef]
- Fankem, H.; Nwaga, D.; Deube, A.; Dieng, L.; Merbach, W.; Etoa, F. Occurrence and functioning of phosphate solubilizing microorganisms from oil palm tree (Elaeis guineensis) rhizosphere in Cameroon. Afr. J. Biotechnol. 2006, 5, 2450–2460. [Google Scholar]
- Chuang, C.-C.; Kuo, Y.-L.; Chao, C.-C.; Chao, W.-L. Solubilization of inorganic phosphates and plant growth promotion by Aspergillus niger. Biol. Fertil. Soils 2007, 43, 575–584. [Google Scholar] [CrossRef]
- Vyas, P.; Gulati, A. Organic acid production in vitro and plant growth promotion in maize under controlled environment by phosphate-solubilizing fluorescent Pseudomonas. BMC Microbiol. 2009, 9, 174. [Google Scholar] [CrossRef] [PubMed]
- Puente, M.E.; Li, C.Y.; Bashan, Y. Microbial Populations and Activities in the Rhizoplane of Rock-Weathering Desert Plants. II. Growth Promotion of Cactus Seedlings. Plant Biol. 2004, 6, 643–650. [Google Scholar] [CrossRef]
- Puente, M.E.; Li, C.Y.; Bashan, Y. Rock-degrading endophytic bacteria in cacti. Environ. Exp. Bot. 2009, 66, 389–401. [Google Scholar] [CrossRef]
- Lopez, B.R.; Bashan, Y.; Bacilio, M. Endophytic bacteria of Mammillaria fraileana, an endemic rock-colonizing cactus of the southern Sonoran Desert. Arch. Microbiol. 2011, 193, 527–541. [Google Scholar] [CrossRef]
- Oubrie, A.; Rozeboom, H.J.; Kalk, K.H.; Olsthoorn, A.J.; Duine, J.A.; Dijkstra, B.W. Structure and mechanism of soluble quinoprotein glucose dehydrogenase. EMBO J. 1999, 18, 5187–5194. [Google Scholar] [CrossRef] [PubMed]
- Lessie, T.G.; Phibbs, P.V. Alternative Pathways of Carbohydrate Utilization in Pseudomonads. Annu. Rev. Microbiol. 1984, 38, 359–388. [Google Scholar] [CrossRef]
- Thomas, L.; Hodgson, D.A.; Wentzel, A.; Nieselt, K.; Ellingsen, T.E.; Moore, J.; Morrissey, E.R.; Legaie, R.; The STREAM Consortium; Wohlleben, W.; et al. Metabolic Switches and Adaptations Deduced from the Proteomes of Streptomyces coelicolor Wild Type and phoP Mutant Grown in Batch Culture. Mol. Cell. Proteom. 2012, 11, M111.013797. [Google Scholar] [CrossRef]
- Buch, A.; Archana, G.; Kumar, G.N. Metabolic channeling of glucose towards gluconate in phosphate-solubilizing Pseudomonas aeruginosa P4 under phosphorus deficiency. Res. Microbiol. 2008, 159, 635–642. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Q.; Wu, X.; Wen, X. Effects of Soluble Phosphate on Phosphate-Solubilizing Characteristics and Expression of gcd Gene in Pseudomonas frederiksbergensis JW-SD2. Curr. Microbiol. 2016, 72, 198–206. [Google Scholar] [CrossRef]
- Ben Farhat, M.; Fourati, A.; Chouayekh, H. Coexpression of the Pyrroloquinoline Quinone and Glucose Dehydrogenase Genes from Serratia marcescens CTM 50650 Conferred High Mineral Phosphate-Solubilizing Ability to Escherichia coli. Appl. Biochem. Biotechnol. 2013, 170, 1738–1750. [Google Scholar] [CrossRef]
- Liu, S.T.; Lee, L.Y.; Tai, C.Y.; Hung, C.-H.; Chang, Y.S.; Wolfram, J.H.; Rogers, R.; Goldstein, A.H. Cloning of an Erwinia herbicola gene necessary for gluconic acid production and enhanced mineral phosphate solubilization in Escherichia coli HB101: Nucleotide sequence and probable involvement in biosynthesis of the coenzyme pyrroloquinoline quinone. J. Bacteriol. 1992, 174, 5814–5819. [Google Scholar] [CrossRef][Green Version]
- Zeng, Q.; Wu, X.; Wang, J.; Ding, X. Phosphate Solubilization and Gene Expression of Phosphate-Solubilizing Bacterium Burkholderia multivorans WS-FJ9 under Different Levels of Soluble Phosphate. J. Microbiol. Biotechnol. 2017, 27, 844–855. [Google Scholar] [CrossRef] [PubMed]
- Peix, A.; Mateos, P.F.; Rodriguez-Barrueco, C.; Martínez-Molina, E.; Velazquez, E. Growth promotion of common bean (Phaseolus vulgaris L.) by a strain of Burkholderia cepacia under growth chamber conditions. Soil Biol. Biochem. 2001, 33, 1927–1935. [Google Scholar] [CrossRef]
- Nannipieri, P.; Giagnoni, L.; Landi, L.; Renella, G. Role of Phosphatase Enzymes in Soil. In Phosphorus in Action; Bünemann, E.K., Oberson, A., Frossard, E., Eds.; Springer: Berlin/Heidelberg, Germany, 2011; pp. 215–243. [Google Scholar] [CrossRef]
- Jorquera, M.A.; Crowley, D.E.; Marschner, P.; Greiner, R.; Fernández, M.T.; Romero, D.; Menezes-Blackburn, D.; De La Luz Mora, M. Identification of β-propeller phytase-encoding genes in culturable Paenibacillus and Bacillus spp. from the rhizosphere of pasture plants on volcanic soils. FEMS Microbiol. Ecol. 2011, 75, 163–172. [Google Scholar] [CrossRef] [PubMed]
- Lim, B.L.; Yeung, P.; Cheng, T.; Hill, J.E. Distribution and diversity of phytate-mineralizing bacteria. ISME J. 2007, 1, 321–330. [Google Scholar] [CrossRef]
- Turner, B.L.; Papházy, M.J.; Haygarth, P.M.; McKelvie, I.D. Inositol phosphates in the environment. Philos. Trans. R. Soc. London. Ser. B Biol. Sci. 2002, 357, 449–469. [Google Scholar] [CrossRef]
- Rao, D.E.C.S.; Rao, K.V.; Reddy, T.P.; Reddy, V.D. Molecular characterization, physicochemical properties, known and potential applications of phytases: An overview. Crit. Rev. Biotechnol. 2009, 29, 182–198. [Google Scholar] [CrossRef]
- Huang, H.; Shi, P.; Wang, Y.; Luo, H.; Shao, N.; Wang, G.; Yang, P.; Yao, B. Diversity of Beta-Propeller Phytase Genes in the Intestinal Contents of Grass Carp Provides Insight into the Release of Major Phosphorus from Phytate in Nature. Appl. Environ. Microbiol. 2009, 75, 1508–1516. [Google Scholar] [CrossRef] [PubMed]
- Kim, O.-H.; Kim, Y.-O.; Shim, J.-H.; Jung, Y.-S.; Jung, W.-J.; Choi, W.-C.; Lee, H.; Lee, S.-J.; Kim, K.-K.; Auh, J.-H.; et al. β-Propeller Phytase Hydrolyzes Insoluble Ca2+-Phytate Salts and Completely Abrogates the Ability of Phytate to Chelate Metal Ions. Biochemistry 2010, 49, 10216–10227. [Google Scholar] [CrossRef]
- Fu, S.; Sun, J.; Qian, L. Effect of Ca2+ on beta-propeller phytases. Protein Pept. Lett. 2008, 15, 39–42. [Google Scholar] [CrossRef]
- Cheng, C.; Lim, B.L. Beta-propeller phytases in the aquatic environment. Arch. Microbiol. 2006, 185, 1–13. [Google Scholar] [CrossRef]
- Kerovuo, J.; Lauraeus, M.; Nurminen, P.; Kalkkinen, N.; Apajalahti, J. Isolation, Characterization, Molecular Gene Cloning, and Sequencing of a Novel Phytase from Bacillus subtilis. Appl. Environ. Microbiol. 1998, 64, 2079–2085. [Google Scholar] [CrossRef]
- Kim, Y.-O.; Lee, J.-K.; Kim, H.-K.; Yu, J.-H.; Oh, T.-K. Cloning of the thermostable phytase gene (phy) from Bacillus sp. DS11 and its overexpression in Escherichia coli. FEMS Microbiol. Lett. 1998, 162, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Tye, A.; Siu, F.; Leung, T.; Lim, B. Molecular cloning and the biochemical characterization of two novel phytases from B. subtilis 168 and B. licheniformis. Appl. Microbiol. Biotechnol. 2002, 59, 190–197. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Shao, N.; Wang, Y.; Luo, H.; Yang, P.; Zhou, Z.; Zhan, Z.; Yao, B. A novel beta-propeller phytase from Pedobacter nyackensis MJ11 CGMCC 2503 with potential as an aquatic feed additive. Appl. Microbiol. Biotechnol. 2009, 83, 249–259. [Google Scholar] [CrossRef]
- Zhang, R.; Yang, P.; Huang, H.; Yuan, T.; Shi, P.; Meng, K.; Yao, B. Molecular and biochemical characterization of a new alkaline β-propeller phytase from the insect symbiotic bacterium Janthinobacterium sp. TN115. Appl. Microbiol. Biotechnol. 2011, 92, 317–325. [Google Scholar] [CrossRef]
- Jorquera, M.A.; Hernández, M.T.; Rengel, Z.; Marschner, P.; de la Luz Mora, M. Isolation of culturable phosphobacteria with both phytate-mineralization and phosphate-solubilization activity from the rhizosphere of plants grown in a volcanic soil. Biol. Fertil. Soils 2008, 44, 1025–1034. [Google Scholar] [CrossRef]
- Rodríguez, H.; Rossolini, G.M.; Gonzalez, T.; Li, J.P.; Glick, B.R. Isolation of a Gene from Burkholderia cepacia IS-16 Encoding a Protein That Facilitates Phosphatase Activity. Curr. Microbiol. 2000, 40, 362–366. [Google Scholar] [CrossRef] [PubMed]
- Thaller, M.C.; Berlutti, F.; Schippa, S.; Iori, P.; Passariello, C.; Rossolini, G.M. Heterogeneous Patterns of Acid Phosphatases Containing Low-Molecular-Mass Polypeptides in Members of the Family Enterobacteriaceae. Int. J. Syst. Bacteriol. 1995, 45, 255–261. [Google Scholar] [CrossRef]
- Richardson, A.E.; Hadobas, P.A. Soil isolates of Pseudomonas spp. that utilize inositol phosphates. Can. J. Microbiol. 1997, 43, 509–516. [Google Scholar] [CrossRef] [PubMed]
- Ohtake, H.; Wu, H.; Imazu, K.; Anbe, Y.; Kato, J.; Kuroda, A. Bacterial phosphonate degradation, phosphite oxidation and polyphosphate accumulation. Resour. Conserv. Recycl. 1996, 18, 125–134. [Google Scholar] [CrossRef]
- Lidbury, I.D.E.A.; Murphy, A.R.J.; Scanlan, D.J.; Bending, G.D.; Jones, A.M.E.; Moore, J.D.; Goodall, A.; Hammond, J.P.; Wellington, E.M.H. Comparative genomic, proteomic and exoproteomic analyses of three Pseudomonas strains reveals novel insights into the phosphorus scavenging capabilities of soil bacteria. Environ. Microbiol. 2016, 18, 3535–3549. [Google Scholar] [CrossRef]
- Brito, L.F.; Gil López, M.; Straube, L.; Passaglia, L.M.P.; Wendisch, V.F. Inorganic Phosphate Solubilization by Rhizosphere Bacterium Paenibacillus sonchi: Gene Expression and Physiological Functions. Front. Microbiol. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Heuck, C.; Smolka, G.; Whalen, E.D.; Frey, S.; Gundersen, P.; Moldan, F.; Fernandez, I.J.; Spohn, M. Effects of long-term nitrogen addition on phosphorus cycling in organic soil horizons of temperate forests. Biogeochemistry 2018, 141, 167–181. [Google Scholar] [CrossRef]
- Marklein, A.R.; Houlton, B.Z. Nitrogen inputs accelerate phosphorus cycling rates across a wide variety of terrestrial ecosystems. New Phytol. 2012, 193, 696–704. [Google Scholar] [CrossRef] [PubMed]
- Contesto, C.; Desbrosses, G.; Lefoulon, C.; Bena, G.; Borel, F.; Galland, M.; Gamet, L.; Varoquaux, F.; Touraine, B. Effects of rhizobacterial ACC deaminase activity on Arabidopsis indicate that ethylene mediates local root responses to plant growth-promoting rhizobacteria. Plant Sci. 2008, 175, 178–189. [Google Scholar] [CrossRef]
- Haidar, B.; Ferdous, M.; Fatema, B.; Ferdous, A.S.; Islam, M.R.; Khan, H. Population diversity of bacterial endophytes from jute (Corchorus olitorius) and evaluation of their potential role as bioinoculants. Microbiol. Res. 2018, 208, 43–53. [Google Scholar] [CrossRef]
- Kudoyarova, G.R.; Vysotskaya, L.B.; Arkhipova, T.N.; Kuzmina, L.Y.; Galimsyanova, N.F.; Sidorova, L.V.; Gabbasova, I.M.; Melentiev, A.I.; Veselov, S.Y. Effect of auxin producing and phosphate solubilizing bacteria on mobility of soil phosphorus, growth rate, and P acquisition by wheat plants. Acta Physiol. Plant. 2017, 39, 253. [Google Scholar] [CrossRef]
- Ortíz-Castro, R.; Contreras-Cornejo, H.A.; Macías-Rodríguez, L.; López-Bucio, J. The role of microbial signals in plant growth and development. Plant Signal. Behav. 2009, 4, 701–712. [Google Scholar] [CrossRef]
- Li, Y.; Liu, X.; Hao, T.; Chen, S. Colonization and Maize Growth Promotion Induced by Phosphate Solubilizing Bacterial Isolates. Int. J. Mol. Sci. 2017, 18, 1253. [Google Scholar] [CrossRef]
- Saraf, M.; Jha, C.K.; Patel, D. The Role of ACC Deaminase Producing PGPR in Sustainable Agriculture. In Plant Growth and Health Promoting Bacteria; Springer: Berlin/Heidelberg, Germany, 2010; pp. 365–385. [Google Scholar] [CrossRef]
- Bal, H.B.; Nayak, L.; Das, S.; Adhya, T.K. Isolation of ACC deaminase producing PGPR from rice rhizosphere and evaluating their plant growth promoting activity under salt stress. Plant Soil 2012, 366, 93–105. [Google Scholar] [CrossRef]
- White, C.; Sayer, J.; Gadd, G. Microbial solubilization and immobilization of toxic metals: Key biogeochemical processes for treatment of contamination. FEMS Microbiol. Rev. 1997, 20, 503–516. [Google Scholar] [CrossRef]
- Kang, J.; Amoozegar, A.; Hesterberg, D.; Osmond, D. Phosphorus leaching in a sandy soil as affected by organic and inorganic fertilizer sources. Geoderma 2011, 161, 194–201. [Google Scholar] [CrossRef]
- Rosado, A.S.; De Azevedo, F.S.; Da Cruz, D.W.; Van Elsas, J.D.; Seldin, L. Seldin Phenotypic and genetic diversity of Paenibacillus azotofixans strains isolated from the rhizoplane or rhizosphere soil of different grasses. J. Appl. Microbiol. 1998, 84, 216–226. [Google Scholar] [CrossRef]
- Nahas, E. Factors determining rock phosphate solubilization by microorganisms isolated from soil. World J. Microbiol. Biotechnol. 1996, 12, 567–572. [Google Scholar] [CrossRef]
- Nautiyal, C.S.; Bhadauria, S.; Kumar, P.; Lal, H.; Mondal, R.; Verma, D. Stress induced phosphate solubilization in bacteria isolated from alkaline soils. FEMS Microbiol. Lett. 2000, 182, 291–296. [Google Scholar] [CrossRef]
- Johri, J.K.; Surange, S.; Nautiyal, C.S. Occurrence of Salt, pH, and Temperature-tolerant, Phosphate-solubilizing Bacteria in Alkaline Soils. Curr. Microbiol. 1999, 39, 89–93. [Google Scholar] [CrossRef]
- Seshachala, U.; Tallapragada, P. Phosphate Solubilizers from the Rhizospher of Piper nigrum L. in Karnataka, India. Chil. J. Agric. Res. 2012, 72, 397–403. [Google Scholar] [CrossRef]
- Zhang, L.; Ding, X.; Chen, S.; He, X.; Zhang, F.; Feng, G. Reducing carbon: Phosphorus ratio can enhance microbial phytin mineralization and lessen competition with maize for phosphorus. J. Plant Interactions 2014, 9, 850–856. [Google Scholar] [CrossRef]
- Babu-Khan, S.; Yeo, T.C.; Martin, W.L.; Duron, M.R.; Rogers, R.D.; Goldstein, A.H. Cloning of a mineral phosphate-solubilizing gene from Pseudomonas cepacia. Appl. Environ. Microbiol. 1995, 61, 972–978. [Google Scholar] [CrossRef] [PubMed]
- Van Schie, B.J.; Hellingwerf, K.J.; van Dijken, J.P.; Elferink, M.G.; van Dijl, J.M.; Kuenen, J.G.; Konings, W.N. Energy transduction by electron transfer via a pyrrolo-quinoline quinone-dependent glucose dehydrogenase in Escherichia coli, Pseudomonas aeruginosa, and Acinetobacter calcoaceticus (var. lwoffi). J. Bacteriol. 1985, 163, 493–499. [Google Scholar] [CrossRef]
- Xie, J.; Shi, H.; Du, Z.; Wang, T.; Liu, X.; Chen, S. Comparative genomic and functional analysis reveal conservation of plant growth promoting traits in Paenibacillus polymyxa and its closely related species. Sci. Rep. 2016, 6, 21329. [Google Scholar] [CrossRef]
- Grady, E.N.; Macdonald, J.; Liu, L.; Richman, A.; Yuan, Z.-C. Current knowledge and perspectives of Paenibacillus: A review. Microb. Cell Factories 2016, 15, 203. [Google Scholar] [CrossRef]
- Rosenberg, H. Phosphate Transport in Prokaryotes. In Ion Transport in Prokaryotes; Elsevier: Amsterdam, The Netherlands, 1987; pp. 205–248. [Google Scholar]
- Thaller, M.C.; Berlutti, F.; Schippa, S.; Lombardi, G.; Rossolini, G.M. Characterization and sequence of PhoC, the principal phosphate-irrepressible acid phosphatase of Morganella morganii. Microbiology 1994, 140, 1341–1350. [Google Scholar] [CrossRef]
- Pond, J.L.; Eddy, C.K.; Mackenzie, K.F.; Conway, T.; Borecky, D.J.; Ingram, L.O. Cloning, sequencing, and characterization of the principal acid phosphatase, the phoC+ product, from Zymomonas mobilis. J. Bacteriol. 1989, 171, 767–774. [Google Scholar] [CrossRef]
- Kier, L.D.; Weppelman, R.; Ames, B.N. Resolution and purification of three periplasmic phosphatases of Salmonella typhimurium. J. Bacteriol. 1977, 130, 399–410. [Google Scholar] [CrossRef]
- Miller, S.I.; Kukral, A.M.; Mekalanos, J.J. A two-component regulatory system (phoP phoQ) controls Salmonella typhimurium virulence. Proc. Natl. Acad. Sci. USA 1989, 86, 5054–5058. [Google Scholar] [CrossRef]
- Burini, J.-F.; Gügi, B.; Merieau, A.; Guespin-Michel, J.F. Lipase and acidic phosphatase from the psychrotrophic bacterium Pseudomonas fluorescens: Two enzymes whose synthesis is regulated by the growth temperature. FEMS Microbiol. Lett. 1994, 122, 13–18. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez, H.; Fraga, R.; Gonzalez, T.; Bashan, Y. Genetics of phosphate solubilization and its potential applications for improving plant growth-promoting bacteria. Plant Soil 2006, 287, 15–21. [Google Scholar] [CrossRef]
- Alori, E.T.; Glick, B.R.; Babalola, O.O. Microbial Phosphorus Solubilization and Its Potential for Use in Sustainable Agriculture. Front. Microbiol. 2017, 8, 971. [Google Scholar] [CrossRef]
- Passariello, C.; Schippa, S.; Iori, P.; Berlutti, F.; Thaller, M.C.; Rossolini, G.M. The molecular class C acid phosphatase of Chryseobacterium meningosepticum (OlpA) is a broad-spectrum nucleotidase with preferential activity on 5′-nucleotides. Biochim. Biophys. Acta—Proteins Proteom. 2003, 1648, 203–209. [Google Scholar] [CrossRef]
- Jiang, L.; Liu, X.; Yin, H.; Liang, Y.; Liu, H.; Miao, B.; Peng, Q.; Meng, D.; Wang, S.; Yang, J.; et al. The utilization of biomineralization technique based on microbial induced phosphate precipitation in remediation of potentially toxic ions contaminated soil: A mini review. Ecotoxicol. Environ. Saf. 2020, 191, 110009. [Google Scholar] [CrossRef]
- Xie, E.; Su, Y.; Deng, S.; Kontopyrgou, M.; Zhang, D. Significant influence of phosphorus resources on the growth and alkaline phosphatase activities of Microcystis aeruginosa. Environ. Pollut. 2020, 268, 115807. [Google Scholar] [CrossRef] [PubMed]
- Reher, M.; Bott, M.; Schönheit, P. Characterization of glycerate kinase (2-phosphoglycerate forming), a key enzyme of the nonphosphorylative Entner-Doudoroff pathway, from the thermoacidophilic euryarchaeon Picrophilus torridus. FEMS Microbiol. Lett. 2006, 259, 113–119. [Google Scholar] [CrossRef] [PubMed]
- Conway, T. The Entner-Doudoroff pathway: History, physiology and molecular biology. FEMS Microbiol. Lett. 1992, 103, 1–28. [Google Scholar] [CrossRef] [PubMed]
- Basu, A.; Phale, P.S. Inducible uptake and metabolism of glucose by the phosphorylative pathway in Pseudomonas putida CSV86. FEMS Microbiol. Lett. 2006, 259, 311–316. [Google Scholar] [CrossRef] [PubMed]
- Elferink, M.G.L.; Albers, S.-V.; Konings, W.N.; Driessen, A.J.M. Sugar transport in Sulfolobus solfataricus is mediated by two families of binding protein-dependent ABC transporters. Mol. Microbiol. 2001, 39, 1494–1503. [Google Scholar] [CrossRef]
- Lamarche, M.G.; Wanner, B.L.; Crépin, S.; Harel, J. The phosphate regulon and bacterial virulence: A regulatory network connecting phosphate homeostasis and pathogenesis. FEMS Microbiol. Rev. 2008, 32, 461–473. [Google Scholar] [CrossRef]
- Hulett, F.M.; Lee, J.; Shi, L.; Sun, G.; Chesnut, R.; Sharkova, E.; Duggan, M.F.; Kapp, N. Sequential action of two-component genetic switches regulates the PHO regulon in Bacillus subtilis. J. Bacteriol. 1994, 176, 1348–1358. [Google Scholar] [CrossRef]
- Eder, S.; Shi, L.; Jensen, K.; Yamane, K.; Hulett, F.M. A Bacillus subtilis secreted phosphodiesterase/alkaline phosphatase is the product of a Pho regulon gene, phoD. Microbiology 1996, 142, 2041–2047. [Google Scholar] [CrossRef] [PubMed]
- Maddocks, S.E.; Oyston, P.C.F. Structure and function of the LysR-type transcriptional regulator (LTTR) family proteins. Microbiology 2008, 154, 3609–3623. [Google Scholar] [CrossRef] [PubMed]
- Heller, K.J.; Kadner, R.J.; Günther, K. Suppression of the btuB451 mutation by mutations in the tonB gene suggests a direct interaction between TonB and TonB-dependent receptor proteins in the outer membrane of Escherichia coli. Gene 1988, 64, 147–153. [Google Scholar] [CrossRef]
- Braun, V.; Mahren, S.; Ogierman, M. Regulation of the FecI-type ECF sigma factor by transmembrane signalling. Curr. Opin. Microbiol. 2003, 6, 173–180. [Google Scholar] [CrossRef]
- Wanner, B.L.; Chang, B.D. The phoBR operon in Escherichia coli K-12. J. Bacteriol. 1987, 169, 5569–5574. [Google Scholar] [CrossRef]
- Santos-Beneit, F.; Rodríguez-García, A.; Franco-Domínguez, E.; Martín, J.F. Phosphate-dependent regulation of the low- and high-affinity transport systems in the model actinomycete Streptomyces coelicolor. Microbiology 2008, 154, 2356–2370. [Google Scholar] [CrossRef] [PubMed]
- Ghorbel, S.; Smirnov, A.; Chouayekh, H.; Sperandio, B.; Esnault, C.; Kormanec, J.; Virolle, M.-J. Regulation of ppk Expression and In Vivo Function of Ppk in Streptomyces lividans TK24. J. Bacteriol. 2006, 188, 6269–6276. [Google Scholar] [CrossRef]
- Liu, W.; Eder, S.; Hulett, F.M. Analysis of Bacillus subtilis tagAB and tagDEF Expression during Phosphate Starvation Identifies a Repressor Role for PhoP∼P. J. Bacteriol. 1998, 180, 753–758. [Google Scholar] [CrossRef]
- Tommassen, J.; de Geus, P.; Lugtenberg, B.; Hackett, J.; Reeves, P. Regulation of the pho regulon of Escherichia coli K-12. J. Mol. Biol. 1982, 157, 265–274. [Google Scholar] [CrossRef]
- Hulett, F.M. The signal-transduction network for Pho regulation in Bacillus subtilis. Mol. Microbiol. 1996, 19, 933–939. [Google Scholar] [CrossRef] [PubMed]
- Novak, R.; Cauwels, A.; Charpentier, E.; Tuomanen, E. Identification of a Streptococcus pneumoniae Gene Locus Encoding Proteins of an ABC Phosphate Transporter and a Two-Component Regulatory System. J. Bacteriol. 1999, 181, 1126–1133. [Google Scholar] [CrossRef] [PubMed]
- Makino, K.; Amemura, M.; Kim, S.K.; Nakata, A.; Shinagawa, H. Role of the sigma 70 subunit of RNA polymerase in transcriptional activation by activator protein PhoB in Escherichia coli. Genes Dev. 1993, 7, 149–160. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Blanco, A.G.; Solà, M.; Gomis-Ruth, F.X.; Coll, M. Tandem DNA Recognition by PhoB, a Two-Component Signal Transduction Transcriptional Activator. Structure 2002, 10, 701–713. [Google Scholar] [CrossRef]
- Yuan, Z.-C.; Zaheer, R.; Morton, R.; Finan, T.M. Genome prediction of PhoB regulated promoters in Sinorhizobium meliloti and twelve proteobacteria. Nucleic Acids Res. 2006, 34, 2686–2697. [Google Scholar] [CrossRef]
- Auesukaree, C.; Homma, T.; Tochio, H.; Shirakawa, M.; Kaneko, Y.; Harashima, S. Intracellular Phosphate Serves as a Signal for the Regulation of the PHO Pathway in Saccharomyces cerevisiae. J. Biol. Chem. 2004, 279, 17289–17294. [Google Scholar] [CrossRef]
- Carmany, D.O.; Hollingsworth, K.; McCleary, W.R. Genetic and Biochemical Studies of Phosphatase Activity of PhoR. J. Bacteriol. 2003, 185, 1112–1115. [Google Scholar] [CrossRef]
- Steed, P.M.; Wanner, B.L. Use of the rep technique for allele replacement to construct mutants with deletions of the pstSCAB-phoU operon: Evidence of a new role for the PhoU protein in the phosphate regulon. J. Bacteriol. 1993, 175, 6797–6809. [Google Scholar] [CrossRef]
- Schau, M.; Eldakak, A.; Hulett, F.M. Terminal Oxidases Are Essential to Bypass the Requirement for ResD for Full Pho Induction in Bacillus subtilis. J. Bacteriol. 2004, 186, 8424–8432. [Google Scholar] [CrossRef]
- Birkey, S.M.; Liu, W.; Zhang, X.; Duggan, M.F.; Hulett, F.M. Pho signal transduction network reveals direct transcriptional regulation of one two-component system by another two-component regulator: Bacillus subtilis PhoP directly regulates production of ResD. Mol. Microbiol. 1998, 30, 943–953. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Zhou, J.; Zhou, H.-J.; Wang, M.-M.; Liu, M.-M.; Ke, Y.-Z.; Li, P.-F.; Li, J.-N.; Du, H. Global Survey and Expressions of the Phosphate Transporter Gene Families in Brassica napus and Their Roles in Phosphorus Response. Int. J. Mol. Sci. 2020, 21, 1752. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, S.; Srivastava, S. Prescience of endogenous regulation in Arabidopsis thaliana by Pseudomonas putida MTCC 5279 under phosphate starved salinity stress condition. Sci. Rep. 2020, 10, 5855. [Google Scholar] [CrossRef] [PubMed]
- Saia, S.; Rappa, V.; Ruisi, P.; Abenavoli, M.R.; Sunseri, F.; Giambalvo, D.; Frenda, A.S.; Martinelli, F. Soil inoculation with symbiotic microorganisms promotes plant growth and nutrient transporter genes expression in durum wheat. Front. Plant Sci. 2015, 6, 815. [Google Scholar] [CrossRef]
- Kadmiri, I.M.; Chaouqui, L.; Azaroual, S.E.; Sijilmassi, B.; Yaakoubi, K.; Wahby, I. Phosphate-Solubilizing and Auxin-Producing Rhizobacteria Promote Plant Growth Under Saline Conditions. Arab. J. Sci. Eng. 2018, 43, 3403–3415. [Google Scholar] [CrossRef]
- Chatterjee, S.; Sau, G.B.; Sinha, S.; Mukherjee, S.K. Effect of co-inoculation of plant growth-promoting rhizobacteria on the growth of amaranth plants. Arch. Agron. Soil Sci. 2012, 58, 1387–1397. [Google Scholar] [CrossRef]
- Bargaz, A.; Lyamlouli, K.; Chtouki, M.; Zeroual, Y.; Dhiba, D. Soil Microbial Resources for Improving Fertilizers Efficiency in an Integrated Plant Nutrient Management System. Front. Microbiol. 2018, 9, 1606. [Google Scholar] [CrossRef]
- Mahanta, D.; Rai, R.K.; Dhar, S.; Varghese, E.; Raja, A.; Purakayastha, T.J. Modification of root properties with phosphate solubilizing bacteria and arbuscular mycorrhiza to reduce rock phosphate application in soybean-wheat cropping system. Ecol. Eng. 2018, 111, 31–43. [Google Scholar] [CrossRef]
- Adnan, M.; Fahad, S.; Zamin, M.; Shah, S.; Mian, I.A.; Danish, S.; Zafar-Ul-Hye, M.; Battaglia, M.L.; Naz, R.M.M.; Saeed, B.; et al. Coupling Phosphate-Solubilizing Bacteria with Phosphorus Supplements Improve Maize Phosphorus Acquisition and Growth under Lime Induced Salinity Stress. Plants 2020, 9, 900. [Google Scholar] [CrossRef]
- Javeed, H.M.R.; Qamar, R.; Rehman, A.U.; Ali, M.; Rehman, A.; Farooq, M.; Zamir, S.I.; Nadeem, M.; Cheema, M.A.; Shehzad, M.; et al. Improvement in Soil Characteristics of Sandy Loam Soil and Grain Quality of Spring Maize by Using Phosphorus Solublizing Bacteria. Sustainability 2019, 11, 7049. [Google Scholar] [CrossRef]
- Suleman, M.; Yasmin, S.; Rasul, M.; Yahya, M.; Atta, B.M.; Mirza, M.S. Phosphate solubilizing bacteria with glucose dehydrogenase gene for phosphorus uptake and beneficial effects on wheat. PLoS ONE 2018, 13, e0204408. [Google Scholar] [CrossRef] [PubMed]
- Park, J.H.; Bolan, N.; Megharaj, M.; Naidu, R. Concomitant rock phosphate dissolution and lead immobilization by phosphate solubilizing bacteria (Enterobacter sp.). J. Environ. Manag. 2011, 92, 1115–1120. [Google Scholar] [CrossRef]
- Manzoor, M.; Abbasi, M.K.; Sultan, T. Isolation of Phosphate Solubilizing Bacteria from Maize Rhizosphere and Their Potential for Rock Phosphate Solubilization–Mineralization and Plant Growth Promotion. Geomicrobiol. J. 2017, 34, 81–95. [Google Scholar] [CrossRef]
- Elhaissoufi, W.; Khourchi, S.; Ibnyasser, A.; Ghoulam, C.; Rchiad, Z.; Zeroual, Y.; Lyamlouli, K.; Bargaz, A. Phosphate Solubilizing Rhizobacteria Could Have a Stronger Influence on Wheat Root Traits and Aboveground Physiology Than Rhizosphere P Solubilization. Front. Plant Sci. 2020, 11, 979. [Google Scholar] [CrossRef] [PubMed]
- Ben Zineb, A.; Trabelsi, D.; Ayachi, I.; Barhoumi, F.; Aroca, R.; Mhamdi, R. Inoculation with Elite Strains of Phosphate-Solubilizing Bacteria Enhances the Effectiveness of Fertilization with Rock Phosphates. Geomicrobiol. J. 2020, 37, 22–30. [Google Scholar] [CrossRef]
- Tahir, M.; Khalid, U.; Ijaz, M.; Shah, G.M.; Naeem, M.A.; Shahid, M.; Mahmood, K.; Ahmad, N.; Kareem, F. Combined application of bio-organic phosphate and phosphorus solubilizing bacteria (Bacillus strain MWT 14) improve the performance of bread wheat with low fertilizer input under an arid climate. Braz. J. Microbiol. 2018, 49, 15–24. [Google Scholar] [CrossRef]
- Jayakumar, N.; Paulraj, P.; Sajeesh, P.; Sajna, K.; Zinneera, A. Application of native phosphate solubilizing bacteria for the use of cheap organic and inorganic phosphate source in agricultural practise of Capsicum annum (Chili)—A pilot scale field study. Mater. Today Proc. 2019, 16, 1630–1639. [Google Scholar] [CrossRef]
- Bi, Q.-F.; Li, K.-J.; Zheng, B.-X.; Liu, X.-P.; Li, H.-Z.; Jin, B.-J.; Ding, K.; Yang, X.-R.; Lin, X.-Y.; Zhu, Y.-G. Partial replacement of inorganic phosphorus (P) by organic manure reshapes phosphate mobilizing bacterial community and promotes P bioavailability in a paddy soil. Sci. Total Environ. 2020, 703, 134977. [Google Scholar] [CrossRef]
- Skorupska, A.; Wielbo, J.; Kidaj, D.; Marek-Kozaczuk, M. Enhancing Rhizobium–Legume Symbiosis Using Signaling Factors. In Microbes for Legume Improvement; Springer: Vienna, Austria, 2010; pp. 27–54. [Google Scholar] [CrossRef]
- Lindström, K.; Mousavi, S.A. Effectiveness of nitrogen fixation in rhizobia. Microb. Biotechnol. 2020, 13, 1314–1335. [Google Scholar] [CrossRef]
- Lazali, M.; Zaman-Allah, M.; Amenc, L.; Ounane, G.; Abadie, J.; Drevon, J.-J. A phytase gene is overexpressed in root nodules cortex of Phaseolus vulgaris–rhizobia symbiosis under phosphorus deficiency. Planta 2013, 238, 317–324. [Google Scholar] [CrossRef]
- Arfarita, N.; Muhibuddin, A.; Imai, T. Exploration of indigenous free nitrogen-fixing bacteria from rhizosphere of Vigna radiata for agricultural land treatment. J. Degrad. Min. Lands Manag. 2019, 6, 1617–1623. [Google Scholar] [CrossRef]
- Nasto, M.K.; Alvarez-Clare, S.; Lekberg, Y.; Sullivan, B.W.; Townsend, A.R.; Cleveland, C.C. Interactions among nitrogen fixation and soil phosphorus acquisition strategies in lowland tropical rain forests. Ecol. Lett. 2014, 17, 1282–1289. [Google Scholar] [CrossRef]
- Mitran, T.; Meena, R.S.; Lal, R.; Layek, J.; Kumar, S.; Datta, R. Role of Soil Phosphorus on Legume Production. In Legumes for Soil Health and Sustainable Management; Springer: Singapore, 2018; pp. 487–510. [Google Scholar]
- Si, Z.; Yang, Q.; Liang, R.; Chen, L.; Chen, D.; Li, Y. Digalactosyldiacylglycerol Synthase Gene MtDGD1 Plays an Essential Role in Nodule Development and Nitrogen Fixation. Mol. Plant-Microbe Interact. 2019, 32, 1196–1209. [Google Scholar] [CrossRef]
- Aziz, A.; Ahiabor, B.; Opoku, A.; Abaidoo, R. Contributions of Rhizobium Inoculants and Phosphorus Fertilizer to Biological Nitrogen Fixation, Growth and Grain Yield of Three Soybean Varieties on a Fluvic Luvisol. Am. J. Exp. Agric. 2016, 10, 1–11. [Google Scholar] [CrossRef]
- Zaidi, A.; Ahemad, M.; Oves, M.; Ahmad, E.; Khan, M.S. Role of Phosphate-Solubilizing Bacteria in Legume Improvement. In Microbes for Legume Improvement; Springer: Vienna, Austria, 2010; pp. 273–292. [Google Scholar] [CrossRef]
- Sanz-Saez, A.; Morales, F.; Arrese-Igor, C.; Aranjuelo, I. P Deficiency: A Major Limiting Factor for Rhizobial Symbiosis. In Legume Nitrogen Fixation in Soils with Low Phosphorus Availability; Springer International Publishing: Cham, Switzerland, 2017; pp. 21–39. [Google Scholar]
- Van Gerven, L.P.A.; Kuiper, J.J.; Mooij, W.M.; Janse, J.H.; Paerl, H.W.; de Klein, J.J.M. Nitrogen fixation does not axiomatically lead to phosphorus limitation in aquatic ecosystems. Oikos 2019, 128, 563–570. [Google Scholar] [CrossRef]
- Tewari, S.; Pooniya, V.; Sharma, S. Next generation bioformulation prepared by amalgamating Bradyrhizobium, cell free culture supernatant, and exopolysaccharides enhances the indigenous rhizospheric rhizobial population, nodulation, and productivity of pigeon pea. Appl. Soil Ecol. 2020, 147, 103363. [Google Scholar] [CrossRef]
- Batterman, S.A.; Wurzburger, N.; Hedin, L.O. Nitrogen and phosphorus interact to control tropical symbiotic N2fixation: A test in Inga punctata. J. Ecol. 2013, 101, 1400–1408. [Google Scholar] [CrossRef]
- Singh, O.; Gupta, M.; Mittal, V.; Kiran, S.; Nayyar, H.; Gulati, A.; Tewari, R. Novel phosphate solubilizing bacteria ‘Pantoea cypripedii PS1′ along with Enterobacter aerogenes PS16 and Rhizobium ciceri enhance the growth of chickpea (Cicer arietinum L.). Plant Growth Regul. 2014, 73, 79–89. [Google Scholar] [CrossRef]
- Püschel, D.; Janoušková, M.; Voříšková, A.; Gryndlerová, H.; Vosátka, M.; Jansa, J. Arbuscular Mycorrhiza Stimulates Biological Nitrogen Fixation in Two Medicago spp. through Improved Phosphorus Acquisition. Front. Plant Sci. 2017, 8, 390. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, Q.; Guan, G.; Chen, S. Phosphate solubilizing bacteria stimulate wheat rhizosphere and endosphere biological nitrogen fixation by improving phosphorus content. PeerJ 2020, 8, e9062. [Google Scholar] [CrossRef] [PubMed]
- Zaidi, A.; Khan, M.S. Stimulatory effects of dual inoculation with phosphate solubilising microorganisms and arbuscular mycorrhizal fungus on chickpea. Aust. J. Exp. Agric. 2007, 47, 1016–1022. [Google Scholar] [CrossRef]
- Wani, P.A.; Khan, M.S.; Zaidi, A. Synergistic effects of the inoculation with nitrogen-fixing and phosphate-solubilizing rhizobacteria on the performance of field-grown chickpea. J. Plant Nutr. Soil Sci. 2007, 170, 283–287. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).