Targeting TOR and SnRK1 Genes in Rice with CRISPR/Cas9
Abstract
:1. Introduction
2. Results
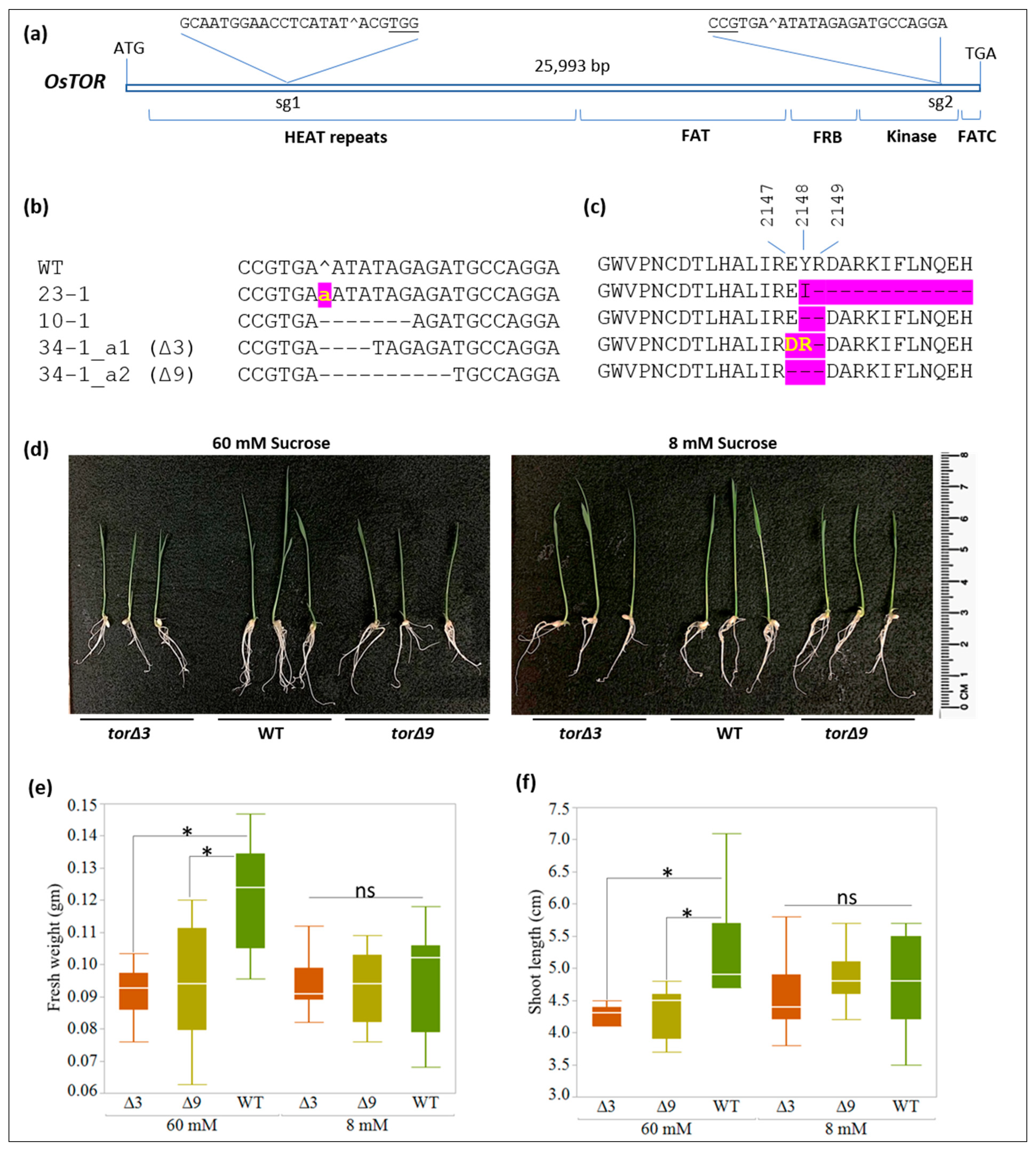
2.1. OsTOR Targeting
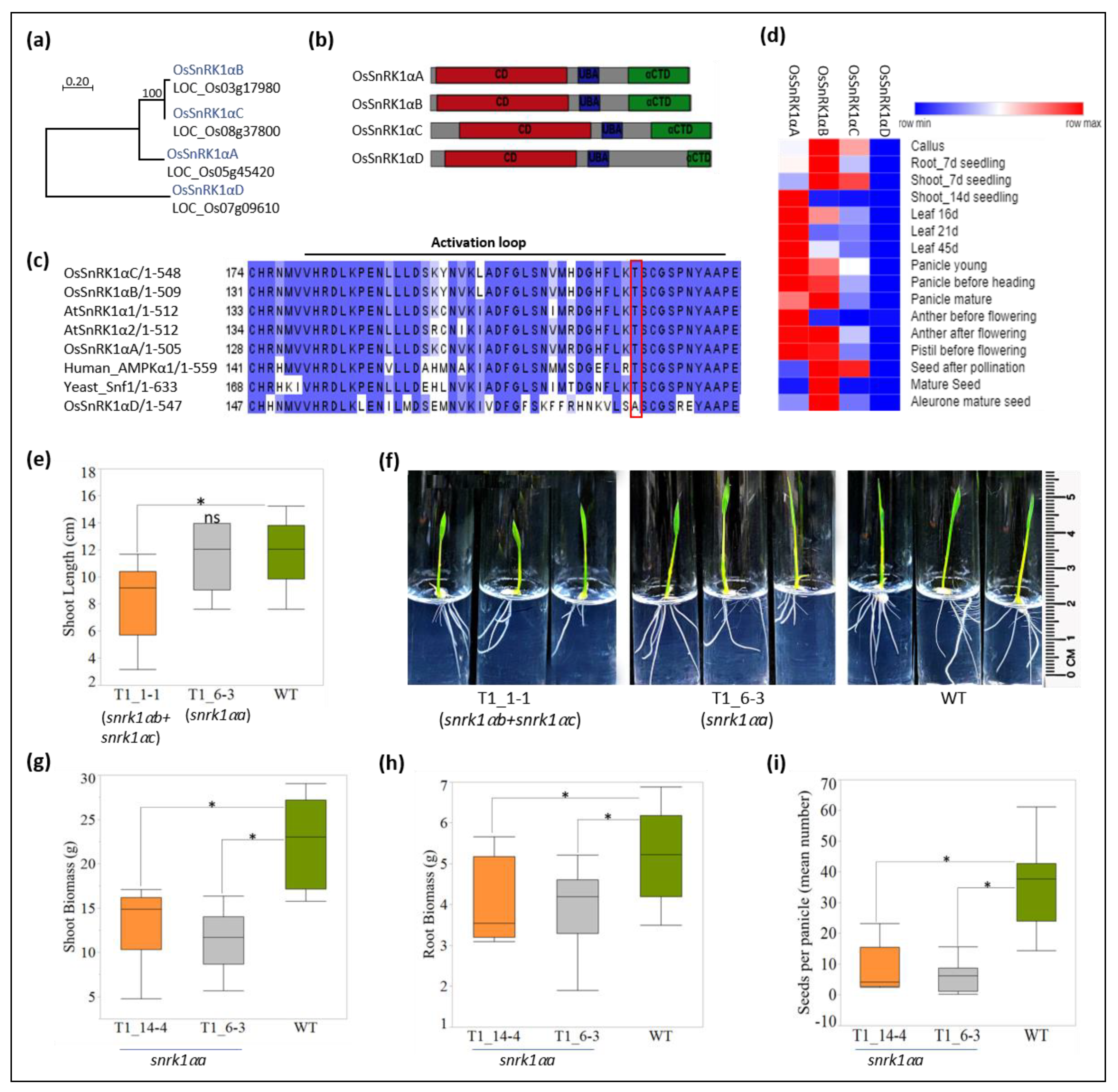
2.2. OsSnRK1α Targeting
2.3. Phenotyping Snrk1 Mutants
2.4. Testing the Efficacy of Inducible CRISPR/Cas9 on OsTOR
3. Discussion
4. Materials and Methods
4.1. Plant Lines
4.2. Molecular Analysis
4.3. Phenotypic Analysis
4.4. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cong, L.; Ran, F.A.; Cox, D.; Lin, S.; Barretto, R.; Habib, N.; Hsu, P.D.; Wu, X.; Jiang, W.; Marraffini, L.A.; et al. Multiplex genome engineering using CRISPR/Cas systems. Science 2013, 339, 819–823. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef] [PubMed]
- Mojica, F.J.; Díez-Villaseñor, C.; García-Martínez, J.; Almendros, C. Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology 2009, 155, 733–740. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nandy, S.; Pathak, B.; Zhao, S.; Srivastava, V. Heat-shock-inducible CRISPR/Cas9 system generates heritable mutations in rice. Plant Direct 2019, 3, 1–14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, H.; Zhang, J.; Wei, P.; Zhang, B.; Gou, F.; Feng, Z.; Mao, Y.; Yang, L.; Zhang, H.; Xu, N.; et al. The CRISPR/Cas9 system produces specific and homozygous targeted gene editing in rice in one generation. Plant Biotechnol. J. 2014, 12, 797–807. [Google Scholar] [CrossRef]
- Zhang, N.; Roberts, H.M.; Van Eck, J.; Martin, G.B. Generation and molecular characterization of CRISPR/Cas9-Induced mutations in 63 immunity-Associated genes in tomato reveals specificity and a range of gene modifications. Front. Plant Sci. 2020, 11, 10. [Google Scholar] [CrossRef] [Green Version]
- Jouanin, A.; Gilissen, L.J.W.J.; Schaart, J.G.; Leigh, F.J.; Cockram, J.; Wallington, E.J.; Boyd, L.A.; van den Broeck, H.C.; van der Meer, I.M.; America, A.H.P.; et al. CRISPR/Cas9 gene editing of gluten in wheat to reduce gluten content and exposure-reviewing methods to screen for coeliac safety. Front Nutr. 2021, 7, 51. [Google Scholar] [CrossRef]
- Li, S.; Lin, D.; Zhang, Y.; Deng, M.; Chen, Y.; Lv, B.; Li, B.; Lei, Y.; Wang, Y.; Zhao, L.; et al. Genome-edited powdery mildew resistance in wheat without growth penalties. Nature 2022, 602, 455–460. [Google Scholar] [CrossRef]
- Oliva, R.; Ji, C.; Atienza-Grande, G.; Huguet-Tapia, J.C.; Perez-Quintero, A.; Li, T.; Eom, J.S.; Li, C.; Nguyen, H.; Liu, B.; et al. Broad-spectrum resistance to bacterial blight in rice using genome editing. Nat. Biotechnol. 2019, 37, 1344–1350. [Google Scholar] [CrossRef] [Green Version]
- Devic, M. The importance of being essential: EMBRYO-DEFECTIVE genes in Arabidopsis. C. R. Biol. 2008, 331, 726–736. [Google Scholar] [CrossRef]
- Lloyd, J.P.; Seddon, A.E.; Moghe, G.D.; Simenc, M.C.; Shiu, S.H. Characteristics of plant essential genes allow for within- and between-species prediction of lethal mutant phenotypes. Plant Cell 2015, 27, 2133–2147. [Google Scholar] [CrossRef] [PubMed]
- Meinke, D.W. Genome-wide identification of EMBRYO-DEFECTIVE (EMB) genes required for growth and development in Arabidopsis. New Phytol. 2020, 226, 306–325. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Minkenberg, B.; Xie, K.; Yang, Y. Discovery of rice essential genes by characterizing a CRISPR-edited mutation of closely related rice MAP kinase genes. Plant J. 2017, 89, 636–648. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Monroe, J.G.; Srikant, T.; Carbonell-Bejerano, P.; Becker, C.; Lensink, M.; Exposito-Alonso, M.; Klein, M.; Hildebrandt, J.; Neumann, M.; Kliebenstein, D.; et al. Mutation bias reflects natural selection in Arabidopsis thaliana. Nature 2022, 602, 101–105. [Google Scholar] [CrossRef]
- Jamsheer, K.M.; Kumar, M.; Srivastava, V. SNF1-related protein kinase 1: The many-faced signaling hub regulating developmental plasticity in plants. J. Exp. Bot. 2021, 72, 6042–6065. [Google Scholar] [CrossRef]
- Margalha, L.; Confraria, A.; Baena-González, E. SnRK1 and TOR: Modulating growth–defense trade-offs in plant stress responses. J. Exp. Bot. 2019, 70, 2261–2274. [Google Scholar] [CrossRef]
- Wurzinger, B.; Nukarinen, E.; Nägele, T.; Weckwerth, W.; Teige, M. The SnRK1 kinase as central mediator of energy signaling between different organelles. Plant Physiol. 2018, 176, 1085–1094. [Google Scholar] [CrossRef] [Green Version]
- de Vleesschauwer, D.; Filipe, O.; Hoffman, G.; Seifi, H.S.; Haeck, A.; Canlas, P.; van Bockhaven, J.; de Waele, E.; Demeestere, K.; Ronald, P.; et al. Target of rapamycin signaling orchestrates growth-defense trade-offs in plants. New Phytol. 2018, 217, 305–319. [Google Scholar] [CrossRef] [Green Version]
- Jamsheer, K.M.; Jindal, S.; Laxmi, A. Evolution of TOR–SnRK dynamics in green plants and its integration with phytohormone signaling networks. J. Exp. Bot. 2019, 70, 2239–2259. [Google Scholar] [CrossRef]
- Wang, W.; Lu, Y.; Li, J.; Zhang, X.; Hu, F.; Zhao, Y.; Zhou, D.-X. SnRK1 stimulates the histone H3K27me3 demethylase JMJ705 to regulate a transcriptional switch to control energy homeostasis. Plant Cell 2021, 33, 3721–3742. [Google Scholar] [CrossRef]
- Yip, C.K.; Murata, K.; Walz, T.; Sabatini, D.M.; Kang, S.A. Structure of the human mTOR Complex I and its implications for rapamycin inhibition. Mol. Cell 2010, 38, 768–774. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, H.; Rudge, D.G.; Koos, J.D.; Vaidialingam, B.; Yang, H.J.; Pavletich, N.P. mTOR kinase structure, mechanism and regulation. Nature 2013, 497, 217–223. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moreau, M.; Azzopardi, M.; Clément, G.; Dobrenel, T.; Marchive, C.; Renne, C.; Martin-Magniette, M.L.; Taconnat, L.; Renou, J.P.; Robaglia, C.; et al. Mutations in the Arabidopsis homolog of LST8/GβL, a partner of the target of Rapamycin kinase, impair plant growth, flowering, and metabolic adaptation to long days. Plant Cell 2012, 24, 463–481. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gangloff, Y.G.; Mueller, M.; Dann, S.G.; Svoboda, P.; Sticker, M.; Spetz, J.F.; Um, S.H.; Brown, E.J.; Cereghini, S.; Thomas, G.; et al. Disruption of the mouse mTOR gene leads to early post-implantation lethality and prohibits embryonic stem cell development. Mol. Cell. Biol. 2004, 24, 9508–9516. [Google Scholar] [CrossRef] [Green Version]
- Menand, B.; Desnos, T.; Nussaume, L.; Berger, F.; Bouchez, D.; Meyer, C.; Robaglia, C. Expression and disruption of the Arabidopsis TOR (target of rapamycin) gene. Proc. Natl. Acad. Sci. USA 2002, 99, 6422–6427. [Google Scholar] [CrossRef] [Green Version]
- Jamsheer, K.M.; Jindal, S.; Sharma, M.; Awasthi, P.; Sreejath, S.; Sharma, M.; Thomas Mannully, C.; Laxmi, A. A negative feedback loop of TOR signaling balances growth and stress-response trade-offs in plants. Cell Rep. 2022, 39, 110631. [Google Scholar] [CrossRef]
- Xiong, Y.; McCormack, M.; Li, L.; Hall, Q.; Xiang, C.; Sheen, J. Glucose-TOR signalling reprograms the transcriptome and activates meristems. Nature 2013, 496, 181–186. [Google Scholar] [CrossRef] [Green Version]
- Broeckx, T.; Hulsmans, S.; Rolland, F. The plant energy sensor: Evolutionary conservation and divergence of SnRK1 structure, regulation, and function. J. Exp. Bot. 2016, 67, 6215–6252. [Google Scholar] [CrossRef]
- Ramon, M.; Dang, T.V.T.; Broeckx, T.; Hulsmans, S.; Crepin, N.; Sheen, J.; Rolland, F. Default Activation and Nuclear Translocation of the Plant Cellular Energy Sensor SnRK1 Regulate Metabolic Stress Responses and Development. Plant Cell 2019, 31, 1614–1632. [Google Scholar] [CrossRef]
- Baena-Gonzalez, E.; Rolland, F.; Thevelein, J.M.; Sheen, J. A central integrator of transcription networks in plant stress and energy signalling. Nature 2007, 448, 938–942. [Google Scholar] [CrossRef]
- Xia, L.; Zou, D.; Sang, J.; Xu, X.; Yin, H.; Li, M.; Wu, S.; Hu, S.; Hao, L.; Zhang, Z. Rice Expression Database (RED): An integrated RNA-Seq-derived gene expression database for rice. J. Genet. Genom. 2017, 44, 235–241. [Google Scholar] [CrossRef] [PubMed]
- Pedrotti, L.; Weiste, C.; Nägele, T.; Wolf, E.; Lorenzin, F.; Dietrich, K.; Mair, A.; Weckwerth, W.; Teige, M.; Baena-González, E.; et al. Snf1-RELATED KINASE1-controlled C/S1-bZIP signaling activates alternative mitochondrial metabolic pathways to ensure plant survival in extended darkness. Plant Cell 2018, 30, 495–509. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, Y.; Liu, J.; Lin, Y.; Xu, X.; Xia, Y.; Bai, J.; Yu, Y.; Xiao, F.; Ding, Y.; Ding, C.; et al. Sucrose non-fermenting-1-related protein kinase 1 regulates sheath-to-panicle transport of non-structural carbohydrates during rice grain filling. Plant Physiol. 2022, kiac124. [Google Scholar] [CrossRef]
- Deprost, D.; Yao, L.; Sormani, R.; Moreau, M.; Leterreux, G.; Nicolaï, M.; Bedu, M.; Robaglia, C.; Meyer, C. The Arabidopsis TOR kinase links plant growth, yield, stress resistance and mRNA translation. EMBO Rep. 2007, 8, 864–870. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ren, M.; Qiu, S.; Venglat, P.; Xiang, D.; Feng, L.; Selvaraj, G.; Datla, R. Target of rapamycin regulates development and ribosomal RNA expression through kinase domain in Arabidopsis. Plant Physiol. 2011, 155, 1367–1382. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aznar, N.R.; Consolo, V.F.; Salerno, G.L.; Martínez-Noël, G.M.A. TOR signaling downregulation increases resistance to the cereal killer Fusarium graminearum. Plant Signal. Behav. 2018, 13, e1414120. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lu, C.A.; Lin, C.C.; Lee, K.W.; Chen, J.L.; Huang, L.F.; Ho, S.L.; Liu, H.J.; Hsing, Y.I.; Yu, S.M. The SnRK1A protein kinase plays a key role in sugar signaling during germination and seedling growth of rice. Plant Cell 2007, 19, 2484–2499. [Google Scholar] [CrossRef] [Green Version]
- Yang, H.; Wu, J.J.; Tang, T.; Liu, K.D.; Dai, C. CRISPR/Cas9-mediated genome editing efficiently creates specific mutations at multiple loci using one sgRNA in Brassica napus. Sci. Rep. 2017, 7, 7489. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Zhang, S.; Zhang, R.; Gao, J.; Qi, Y.; Song, G.; Li, W.; Li, Y.; Li, G. Efficient multiplex genome editing by CRISPR/Cas9 in common wheat. Plant Biotechnol. J. 2021, 19, 427–429. [Google Scholar] [CrossRef]
- Filipe, O.; De Vleesschauwer, D.; Haeck, A.; Demeestere, K.; Höfte, M. The energy sensor OsSnRK1a confers broad-spectrum disease resistance in rice. Sci. Rep. 2018, 8, 3864. [Google Scholar] [CrossRef] [Green Version]
- Xie, K.; Minkenberg, B.; Yang, Y. Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc. Natl. Acad. Sci. USA 2015, 112, 3570–3575. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stemmer, M.; Thumberger, T.; del Sol Keyer, M.; Wittbrodt, J.; Mateo, J.L. CCTop: An intuitive, flexible and reliable CRISPR/Cas9 target prediction tool. PLoS ONE 2015, 10, e0124633. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dehairs, J.; Talebi, A.; Cherifi, Y.; Swinnen, J.V. CRISP-ID: Decoding CRISPR mediated indels by Sanger sequencing. Sci. Rep. 2016, 6, 28973. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hill, J.T.; Demarest, B.L.; Bisgrove, B.W.; Su, Y.C.; Smith, M.; Yost, H.J. Poly peak parser: Method and software for identification of unknown indels using sanger sequencing of polymerase chain reaction products. Dev. Dyn. 2014, 243, 1632–1636. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Gene | CRISPR Construct | No. Of Plates | T0 Lines 1 | Targeted Lines 2 | Survivors 3 | % Transf. 4 | % Targeting 5 | Heritable Mutation |
|---|---|---|---|---|---|---|---|---|
| OsTOR | pNS71 | 10 | 21 | 3 | 1 | 52.5 | 14 | In-frame deletion |
| OsSnRK1αB OsSnRK1αC | pNS72 | 10 | 10 | 9 | 3 | 25 | 90 | Indels/in-frame deletion |
| OsSnRK1αA | pNS73 | 10 | 40 | 30 | 10 | 100 | 75 | Indels |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pathak, B.; Maurya, C.; Faria, M.C.; Alizada, Z.; Nandy, S.; Zhao, S.; Jamsheer K, M.; Srivastava, V. Targeting TOR and SnRK1 Genes in Rice with CRISPR/Cas9. Plants 2022, 11, 1453. https://doi.org/10.3390/plants11111453
Pathak B, Maurya C, Faria MC, Alizada Z, Nandy S, Zhao S, Jamsheer K M, Srivastava V. Targeting TOR and SnRK1 Genes in Rice with CRISPR/Cas9. Plants. 2022; 11(11):1453. https://doi.org/10.3390/plants11111453
Chicago/Turabian StylePathak, Bhuvan, Chandan Maurya, Maria C. Faria, Zahra Alizada, Soumen Nandy, Shan Zhao, Muhammed Jamsheer K, and Vibha Srivastava. 2022. "Targeting TOR and SnRK1 Genes in Rice with CRISPR/Cas9" Plants 11, no. 11: 1453. https://doi.org/10.3390/plants11111453
APA StylePathak, B., Maurya, C., Faria, M. C., Alizada, Z., Nandy, S., Zhao, S., Jamsheer K, M., & Srivastava, V. (2022). Targeting TOR and SnRK1 Genes in Rice with CRISPR/Cas9. Plants, 11(11), 1453. https://doi.org/10.3390/plants11111453







