Molecular Approaches to Overcome Self-Incompatibility in Diploid Potatoes
Abstract
1. Introduction
2. Molecular Basis of Self-Incompatibility (SI)
3. Self-Incompatibility in Potatoes
4. Phenotyping for Self-Compatibility (SC)
5. Molecular Analysis of the Essential Genes of SI
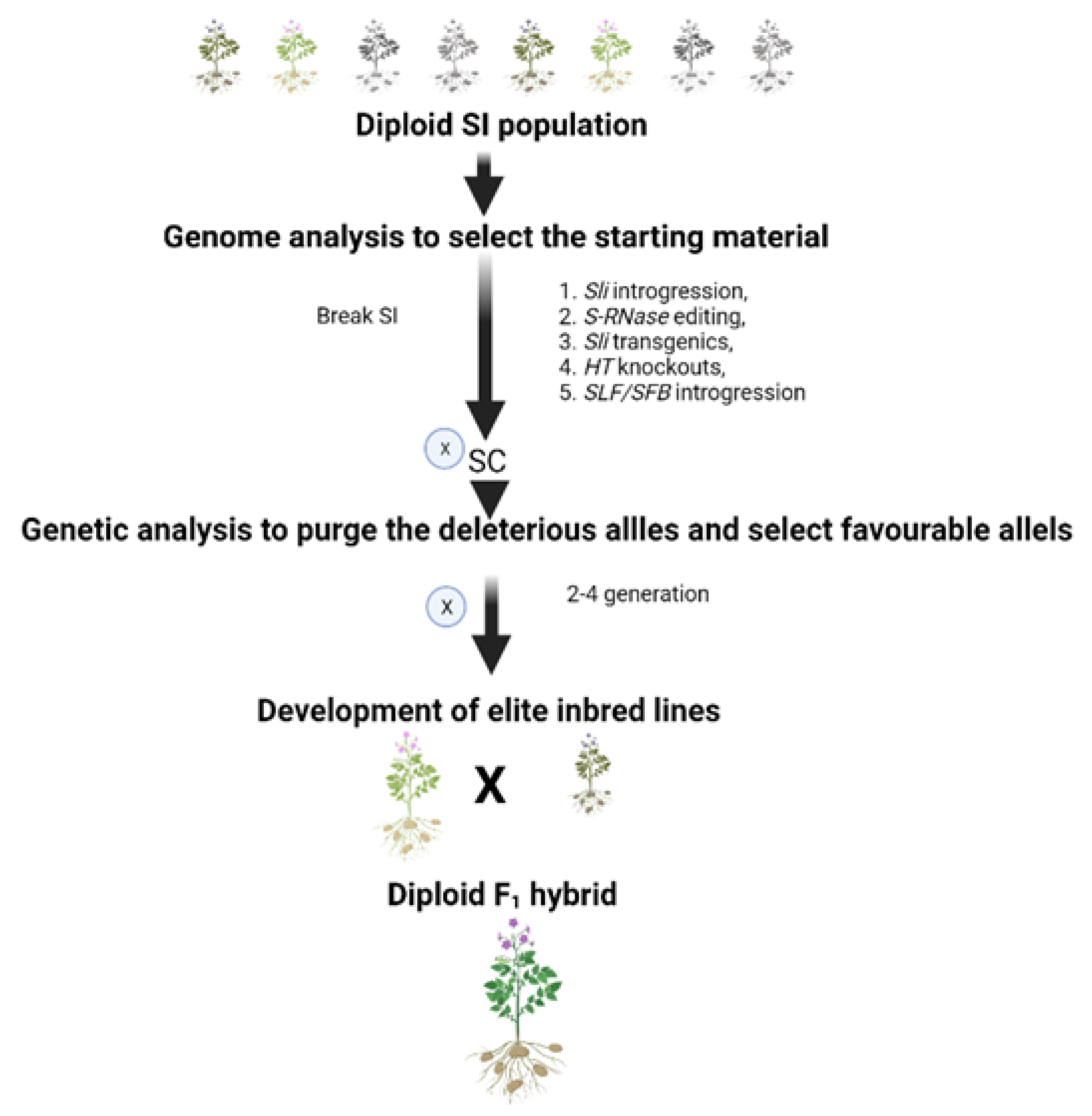
6. Molecular Approaches to Overcome SI
6.1. Inhibit the Function of the S-RNase Gene
6.2. Introduction of SLF/SFB
6.3. HT Knockouts
6.4. S-Locus Inhibitor Introgression
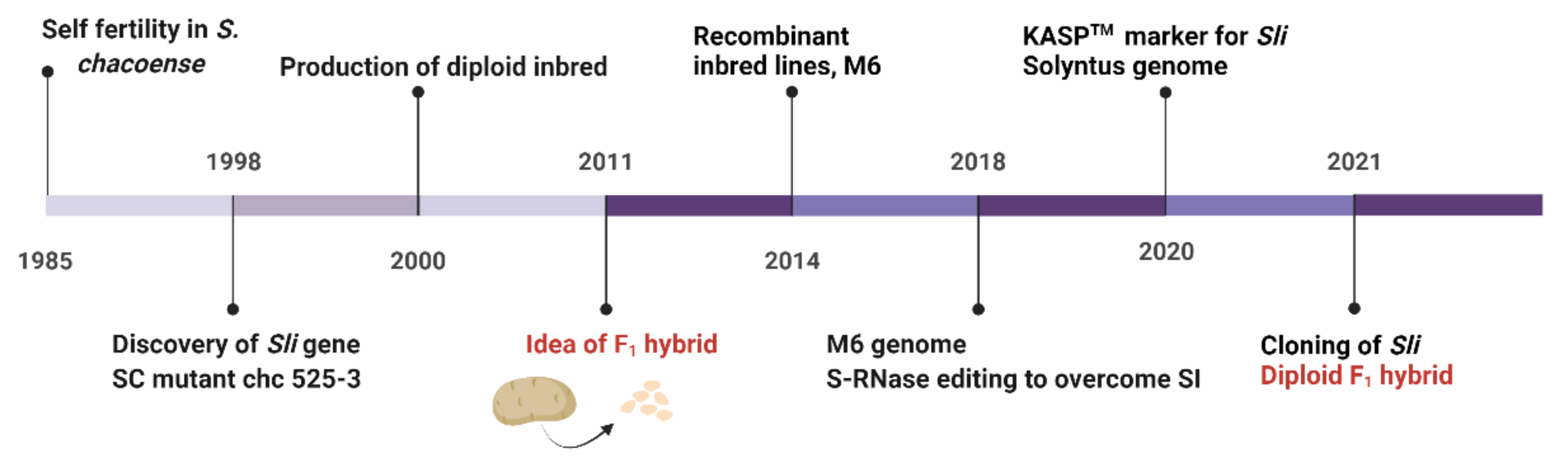
7. Sli Gene Identification and Its Function
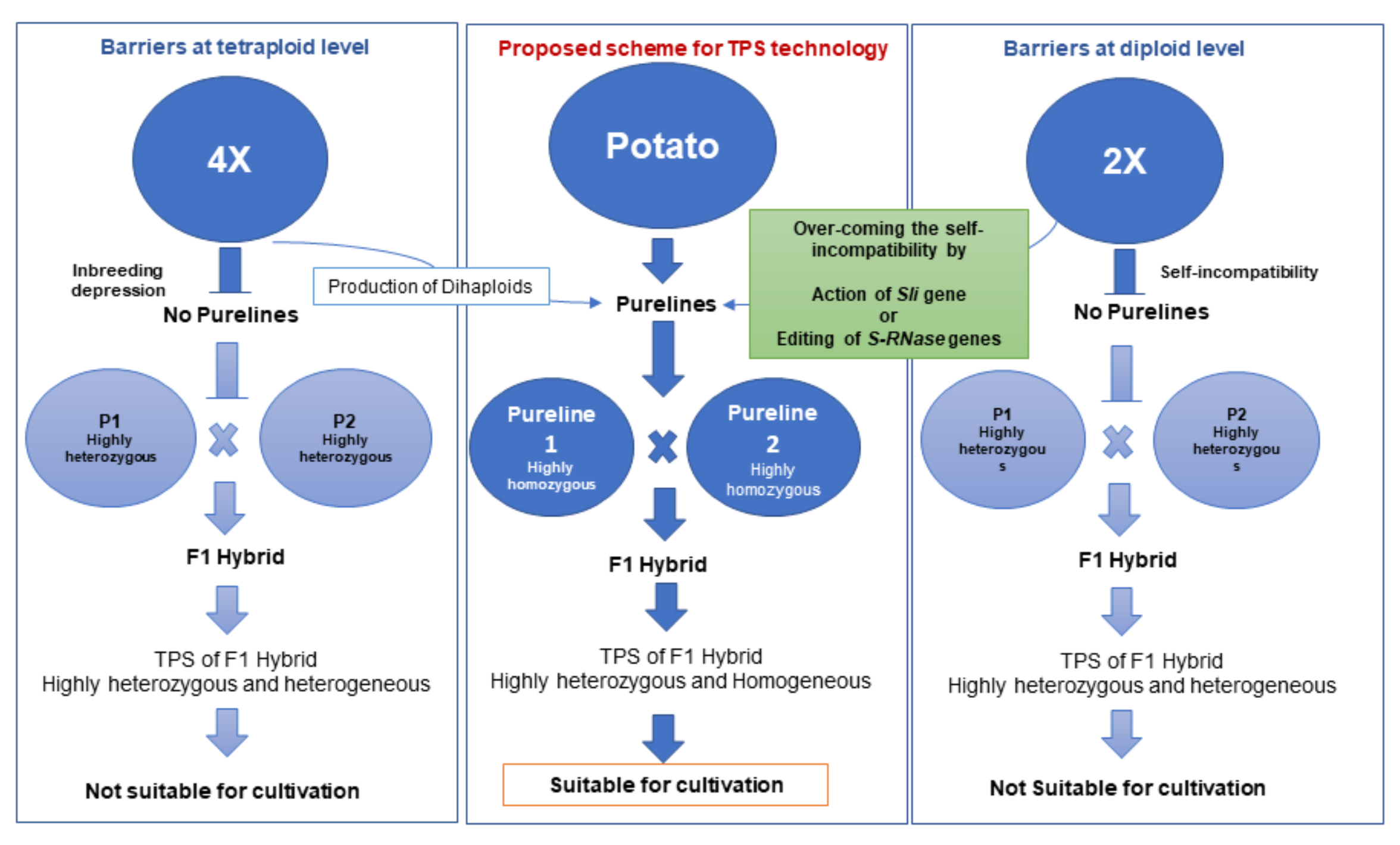
8. Developing Hybrid Potatoes
9. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lindhout, P.; Meijer, D.; Schotte, T.; Hutten, R.C.B.; Visser, R.G.F.; van Eck, H.J. Towards F1 Hybrid Seed Potato Breeding. Potato Res. 2011, 54, 301–312. [Google Scholar] [CrossRef]
- Jansky, S.H.; Chung, Y.S.; Kittipadukal, P. M6: A Diploid Potato Inbred Line for Use in Breeding and Genetics Research. J. Plant Regist. 2014, 8, 195–199. [Google Scholar] [CrossRef]
- Pushkarnath. Studies on Sterility in Potato. Euphytica 1953, 2, 49–58. [Google Scholar] [CrossRef]
- De Nettancourt, D. Incompatibility in Angiosperms. Sex. Plant Reprod. 1997, 10, 185–199. [Google Scholar] [CrossRef]
- Takayama, S.; Isogai, A. Self-Incompatibility in Plants. Annu. Rev. Plant Biol. 2005, 56, 467–489. [Google Scholar] [CrossRef]
- Hiscock, S.J.; McInnis, S.M. The Diversity of Self-Incompatibility Systems in Flowering Plants. Plant Biol. 2003, 5, 23–32. [Google Scholar] [CrossRef]
- McClure, B.; Cruz-García, F.; Romero, C. Compatibility and Incompatibility in S-RNase-Based Systems. Ann. Bot. 2011, 108, 647–658. [Google Scholar] [CrossRef] [PubMed]
- Kao, T.H.; Tsukamoto, T. The Molecular and Genetic Bases of S-RNase-Based Self-Incompatibility. Plant Cell 2004, 16, S72–S83. [Google Scholar] [CrossRef]
- Lee, H.S.; Huang, S.; Kao, T. S Proteins Control Rejection of Incompatible Pollen in Petunia Inflata. Nature 1994, 367, 560–563. [Google Scholar] [CrossRef]
- Murfett, J.; Atherton, T.L.; Mou, B.; Gassert, C.S.; McClure, B.A. S-RNase Expressed in Transgenic Nicotiana Causes S-Allele-Specific Pollen Rejection. Nature 1994, 367, 563–566. [Google Scholar] [CrossRef]
- Sijacic, P.; Wang, X.; Skirpan, A.L.; Wang, Y.; Dowd, P.E.; McCubbin, A.G.; Huang, S.; Kao, T.-H. Identification of the Pollen Determinant of S-RNase-Mediated Self-Incompatibility. Nature 2004, 429, 302–305. [Google Scholar] [CrossRef]
- Kubo, K.-I.; Paape, T.; Hatakeyama, M.; Entani, T.; Takara, A.; Kajihara, K.; Tsukahara, M.; Shimizu-Inatsugi, R.; Shimizu, K.K.; Takayama, S. Gene Duplication and Genetic Exchange Drive the Evolution of S-RNase-Based Self-Incompatibility in Petunia. Nat. Plants 2015, 1, 14005. [Google Scholar] [CrossRef]
- Kao, T.H.; Mccubbin, A.G. How Flowering Plants Discriminate between Self and Non-Self Pollen to Prevent Inbreeding. Proc. Natl. Acad. Sci. USA 1996, 93, 12059–12065. [Google Scholar] [CrossRef]
- Nowak, M.D.; Davis, A.P.; Anthony, F.; Yoder, A.D. Expression and Trans-Specific Polymorphism of Self-Incompatibility Rnases in Coffea (Rubiaceae). PLoS ONE 2011, 6, e21019. [Google Scholar] [CrossRef]
- Asquini, E.; Gerdol, M.; Gasperini, D.; Igić, B.; Graziosi, G.; Pallavicini, A. S-RNase-like Sequences in Styles of Coffea (Rubiaceae). Evidence for S-RNase Based Gametophytic Self-Incompatibility? Trop. Plant Biol. 2011, 4, 237–249. [Google Scholar] [CrossRef]
- Kubo, K.; Entani, T.; Takara, A.; Wang, N.; Fields, A.M.; Hua, Z.; Toyoda, M.; Kawashima, S.; Ando, T.; Isogai, A.; et al. Collaborative Non-Self Recognition System in S-RNase-Based Self-Incompatibility. Science 2010, 330, 796–799. [Google Scholar] [CrossRef]
- Sun, L.; Williams, J.S.; Li, S.; Wu, L.; Khatri, W.A.; Stone, P.G.; Keebaugh, M.D.; Kaoa, T.H. S-Locus F-Box Proteins Are Solely Responsible for S-RNase-Based Self-Incompatibility of Petunia Pollen. Plant Cell 2018, 30, 2959–2972. [Google Scholar] [CrossRef]
- Franklin-Tong, N.; Franklin, C. Gametophytic Self-Incompatibility: Contrasting Mechanisms for Nicotiana and Papaver. Trends Cell Biol. 1993, 3, 340–345. [Google Scholar] [CrossRef]
- Wheeler, M.J.; de Graaf, B.H.J.; Hadjiosif, N.; Perry, R.M.; Poulter, N.S.; Osman, K.; Vatovec, S.; Harper, A.; Franklin, F.C.H.; Franklin-Tong, V.E. Identification of the Pollen Self-Incompatibility Determinant in Papaver Rhoeas. Nature 2009, 459, 992–995. [Google Scholar] [CrossRef]
- Cipar, M.S.; Peloquin, S.J.; Hougas, R.W. Variability in the Expression of Self-Incompatibility in Tuber-Bearing Diploid Solanum Species. Am. Potato J. 1964, 41, 155–162. [Google Scholar] [CrossRef]
- Gebhardt, C.; Ritter, E.; Barone, A.; Debener, T.; Walkemeier, B.; Schachtschabel, U.; Kaufmann, H.; Thompson, R.D.; Bonierbale, M.W.; Ganal, M.W.; et al. RFLP Maps of Potato and Their Alignment with the Homoeologous Tomato Genome. Theor. Appl. Genet. 1991, 83, 49–57. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, J.M.E.; Van Eck, H.J.; Arens, P.; Verkerk-Bakker, B.; te Lintel Hekkert, B.; Bastiaanssen, H.J.M.; El-Kharbotly, A.; Pereira, A.; Jacobsen, E.; Stiekema, W.J. A Genetic Map of Potato (Solanum tuberosum) Integrating Molecular Markers, Including Transposons, and Classical Markers. Theor. Appl. Genet. 1995, 91, 289–300. [Google Scholar] [CrossRef]
- Hosaka, K.; Hanneman, R.E. Genetics of Self-Compatibility in a Self-Incompatible Wild Diploid Potato Species Solanum Chacoense. 1. Detection of an S Locus Inhibitor (Sli) Gene. Euphytica 1998, 99, 191–197. [Google Scholar] [CrossRef]
- Zhang, C.; Shaw, K.M.; Alsahlany, M.A. Dihaploid potato breeding at Michigan State University: Annual report of the Potato Association of America. Am. J. Potato Res. 2019, 94, 360. [Google Scholar] [CrossRef]
- Peterson, B.A.; Holt, S.H.; Laimbeer, F.P.E.; Doulis, A.G.; Coombs, J.; Douches, D.S.; Hardigan, M.A.; Buell, C.R.; Veilleux, R.E. Self-Fertility in a Cultivated Diploid Potato Population Examined with the Infinium 8303 Potato Single-Nucleotide Polymorphism Array. Plant Genome 2016, 9, plantgenome2016.01.0003. [Google Scholar] [CrossRef] [PubMed]
- De Jong, H.; Rowe, P.R. Inbreeding in Cultivated Diploid Potatoes. Potato Res. 1971, 14, 74–83. [Google Scholar] [CrossRef]
- Olsder, J.; Hermsen, J.G.T. Genetics of Self-Compatibility in Dihaploids of Solanum tuberosum L. I. Breeding Behaviour of Two Self-Compatible Dihaploids. Euphytica 1976, 25, 597–607. [Google Scholar] [CrossRef]
- Fulladolsa, A.C.; Charkowski, A.; Cai, X.; Whitworth, J.; Gray, S.; Jansky, S. Germplasm with Resistance to Potato Virus Y Derived from Solanum Chacoense: Clones M19 (39–7) and M20 (XD3). Am. J. Potato Res. 2019, 96, 390–395. [Google Scholar] [CrossRef]
- Kaiser, N.R.; Jansky, S.; Coombs, J.J.; Collins, P.; Alsahlany, M.; Douches, D.S. Assessing the Contribution of Sli to Self-Compatibility in North American Diploid Potato Germplasm Using KASPTM Markers. Am. J. Potato Res. 2021, 98, 104–113. [Google Scholar] [CrossRef]
- Clot, C.R.; Polzer, C.; Prodhomme, C.; Schuit, C.; Engelen, C.J.M.; Hutten, R.C.B.; van Eck, H.J. The Origin and Widespread Occurrence of Sli-Based Self-Compatibility in Potato. Theor. Appl. Genet. 2020, 133, 2713–2728. [Google Scholar] [CrossRef]
- Vaser, R.; Adusumalli, S.; Leng, S.N.; Sikic, M.; Ng, P.C. SIFT Missense Predictions for Genomes. Nat. Protoc. 2016, 11, 1–9. [Google Scholar] [CrossRef]
- Van Lieshout, N.; van der Burgt, A.; de Vries, M.E.; ter Maat, M.; Eickholt, D.; Esselink, D.; van Kaauwen, M.P.W.; Kodde, L.P.; Visser, R.G.F.; Lindhout, P.; et al. Solyntus, the New Highly Contiguous Reference Genome for Potato (Solanum tuberosum). G3 Genes, Genomes, Genet. 2020, 10, 3489–3495. [Google Scholar] [CrossRef]
- Eggers, E.J.; van der Burgt, A.; van Heusden, S.A.W.; de Vries, M.E.; Visser, R.G.F.; Bachem, C.W.B.; Lindhout, P. Neofunctionalisation of the Sli Gene Leads to Self-Compatibility and Facilitates Precision Breeding in Potato. Nat. Commun. 2021, 12, 4141. [Google Scholar] [CrossRef]
- Ma, L.; Zhang, C.; Zhang, B.; Tang, F.; Li, F.; Liao, Q.; Tang, D.; Peng, Z.; Jia, Y.; Gao, M.; et al. A NonS-Locus F-Box Gene Breaks Self-Incompatibility in Diploid Potatoes. Nat. Commun. 2021, 12, 4142. [Google Scholar] [CrossRef]
- Johnson, M.A.; Harper, J.F.; Palanivelu, R. A Fruitful Journey: Pollen Tube Navigation from Germination to Fertilization. Annu. Rev. Plant Biol. 2019, 70, 809–837. [Google Scholar] [CrossRef]
- Ye, M.; Peng, Z.; Tang, D.; Yang, Z.; Li, D.; Xu, Y.; Zhang, C.; Huang, S. Generation of Self-Compatible Diploid Potato by Knockout of S-RNase. Nat. Plants 2018, 4, 651–654. [Google Scholar] [CrossRef]
- Dzidzienyo, D.K.; Bryan, G.J.; Wilde, G.; Robbins, T.P. Allelic Diversity of S-RNase Alleles in Diploid Potato Species. Theor. Appl. Genet. 2016, 129, 1985–2001. [Google Scholar] [CrossRef]
- Ioerger, T.R.; Gohlke, J.R.; Xu, B.; Kao, T.-H. Primary Structural Features of the Self-Incompatibility Protein in Solanaceae. Sex. Plant Reprod. 1991, 4, 81–87. [Google Scholar] [CrossRef]
- Broothaerts, W.; Vanvinckenroye, P.; Decock, B.; Van Damme, J.; Vendrig, J.C. Petunia Hybrida S-Proteins: Ribonuclease Activity and the Role of Their Glycan Side Chains in Self-Incompatibility. Sex. Plant Reprod. 1991, 4, 258–266. [Google Scholar] [CrossRef]
- Karunanandaa, B.; Huang, S.; Kao, T. Carbohydrate Moiety of the Petunia Inflata S3 Protein Is Not Required for Self-Incompatibility Interactions between Pollen and Pistil. Plant Cell 1994, 6, 1933–1940. [Google Scholar] [CrossRef][Green Version]
- Ioerger, T.R.; Clark, A.G.; Kao, T.H. Polymorphism at the Self-Incompatibility Locus in Solanaceae Predates Speciation. Proc. Natl. Acad. Sci. USA 1990, 87, 9732–9735. [Google Scholar] [CrossRef]
- Igic, B.; Kohn, J.R. Evolutionary Relationships among Self-Incompatibility RNases. Proc. Natl. Acad. Sci. USA 2001, 98, 13167–13171. [Google Scholar] [CrossRef] [PubMed]
- Sutherland, B.G.; Tobutt, K.R.; Robbins, T.P. Trans-Specific S-RNase and SFB Alleles in Prunus Self-Incompatibility Haplotypes. Mol. Genet. Genomics 2008, 279, 95–106. [Google Scholar] [CrossRef]
- Richman, A.D.; Kohn, J.R. Evolutionary Genetics of Self-Incompatibility in the Solanaceae. Plant Mol. Biol. 2000, 42, 169–179. [Google Scholar] [CrossRef]
- Ikeda, K.; Igic, B.; Ushijima, K.; Yamane, H.; Hauck, N.R.; Nakano, R.; Sassa, H.; Iezzoni, A.F.; Kohn, J.R.; Tao, R. Primary Structural Features of the S Haplotype-Specific F-Box Protein, SFB, in Prunus. Sex. Plant Reprod. 2004, 16, 235–243. [Google Scholar] [CrossRef]
- Qiao, H.; Wang, H.; Zhao, L.; Zhou, J.; Huang, J.; Zhang, Y.; Xue, Y. The F-Box Protein AhSLF-S2 Physically Interacts with S-RNases That May Be Inhibited by the Ubiquitin/26S Proteasome Pathway of Protein Degradation during Compatible Pollination in Antirrhinum. Plant Cell 2004, 16, 582–595. [Google Scholar] [CrossRef] [PubMed]
- McClure, B. New Views of S-RNase-Based Self-Incompatibility. Curr. Opin. Plant Biol. 2006, 9, 639–646. [Google Scholar] [CrossRef]
- Hua, Z.; Kao, T. Identification and Characterization of Components of a Putative Petunia S-Locus F-Box–Containing E3 Ligase Complex Involved in S-RNase–Based Self-Incompatibility. Plant Cell 2006, 18, 2531–2553. [Google Scholar] [CrossRef]
- Huang, J.; Zhao, L.; Yang, Q.; Xue, Y. AhSSK1, a Novel SKP1-like Protein That Interacts with the S-Locus F-Box Protein SLF. Plant J. 2006, 46, 780–793. [Google Scholar] [CrossRef]
- Enciso-Rodriguez, F.; Manrique-Carpintero, N.C.; Nadakuduti, S.S.; Buell, C.R.; Zarka, D.; Douches, D. Overcoming Self-Incompatibility in Diploid Potato Using CRISPR-Cas9. Front. Plant Sci. 2019, 10, 376. [Google Scholar] [CrossRef]
- Kondo, K.; Yamamoto, M.; Itahashi, R.; Sato, T.; Egashira, H.; Hattori, T.; Kowyama, Y. Insights into the Evolution of Self-Compatibility in Lycopersicon from a Study of Stylar Factors. Plant J. 2002, 30, 143–153. [Google Scholar] [CrossRef]
- O’Brien, M.; Kapfer, C.; Major, G.; Laurin, M.; Bertrand, C.; Kondo, K.; Kowyama, Y.; Matton, D.P. Molecular Analysis of the Stylar-Expressed Solanum Chacoense Small Asparagine-Rich Protein Family Related to the HT Modifier of Gametophytic Self-Incompatibility in Nicotiana. Plant J. 2002, 32, 985–996. [Google Scholar] [CrossRef]
- Hosaka, K.; Hanneman, R.E. Genetics of Self-Compatibility in a Self-Incompatible Wild Diploid Potato Species Solanum Chacoense. 2. Localization of an S Locus Inhibitor (Sli) Gene on the Potato Genome Using DNA Markers. Euphytica 1998, 103, 265–271. [Google Scholar] [CrossRef]
- Gardner, K.M.; Douglass, K.; De Jong, H.; De Koeyer, D.; Tai, H.H. Genetic mapping of self-compatibility in diploid potato using genotyping by sequencing. In Proceedings of the PAG Conference XXVII, San Diego, CA, USA, 12–16 January 2019; p. PE0940. [Google Scholar]
- Hardigan, M.A.; Laimbeer, F.P.E.; Newton, L.; Crisovan, E.; Hamilton, J.P.; Vaillancourt, B.; Wiegert-Rininger, K.; Wood, J.C.; Douches, D.S.; Farré, E.M.; et al. Genome Diversity of Tuber-Bearing Solanum Uncovers Complex Evolutionary History and Targets of Domestication in the Cultivated Potato. Proc. Natl. Acad. Sci. USA 2017, 114, E9999–E10008. [Google Scholar] [CrossRef]
- Hermsen, J.G.T. Genetics of Self-Compatibility in Dihaploids of Solanum tuberosum L. 3. Lethality of S-Bearing Translocation Homozygotes. Euphytica 1978, 27, 13–17. [Google Scholar] [CrossRef]
- Endelman, J.B.; Jansky, S.H. Genetic Mapping with an Inbred Line-Derived F2 Population in Potato. Theor. Appl. Genet. 2016, 129, 935–943. [Google Scholar] [CrossRef]
- Van Os, H.; Andrzejewski, S.; Bakker, E.; Barrena, I.; Bryan, G.J.; Caromel, B.; Ghareeb, B.; Isidore, E.; De Jong, W.; Van Koert, P.; et al. Construction of a 10,000-Marker Ultradense Genetic Recombination Map of Potato: Providing a Framework for Accelerated Gene Isolation and a Genomewide Physical Map. Genetics 2006, 173, 1075–1087. [Google Scholar] [CrossRef]
- Zhang, C.; Wang, P.; Tang, D.; Yang, Z.; Lu, F.; Qi, J.; Tawari, N.R.; Shang, Y.; Li, C.; Huang, S. The Genetic Basis of Inbreeding Depression in Potato. Nat. Genet. 2019, 51, 374–378. [Google Scholar] [CrossRef]
- Marand, A.P.; Jansky, S.H.; Gage, J.L.; Hamernik, A.J.; de Leon, N.; Jiang, J. Residual Heterozygosity and Epistatic Interactions Underlie the Complex Genetic Architecture of Yield in Diploid Potato. Genetics 2019, 212, 317–332. [Google Scholar] [CrossRef]
- Hanneman, R.E., Jr. Self fertility in Solanum chacoense. Am. Potato J. 1985, 62, 428–429. [Google Scholar]
- Birhman, R.K.; Hosaka, K. Production of Inbred Progenies of Diploid Potatoes Using an S-Locus Inhibitor (Sli) Gene, and Their Chacterization. Genome 2000, 43, 495–502. [Google Scholar] [CrossRef]
- Leisner, C.P.; Hamilton, J.P.; Crisovan, E.; Manrique-Carpintero, N.C.; Marand, A.P.; Newton, L.; Pham, G.M.; Jiang, J.; Douches, D.S.; Jansky, S.H.; et al. Genome Sequence of M6, a Diploid Inbred Clone of the High-Glycoalkaloid-Producing Tuber-Bearing Potato Species Solanum Chacoense, Reveals Residual Heterozygosity. Plant J. 2018, 94, 562–570. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Yang, Z.; Tang, D.; Zhu, Y.; Wang, P.; Li, D.; Zhu, G.; Xiong, X.; Shang, Y.; Li, C.; et al. Genome Design of Hybrid Potato. Cell 2021, 184, 3873–3883. [Google Scholar] [CrossRef] [PubMed]
- Fujii, S.; Kubo, K.I.; Takayama, S. Non-Self- and Self-Recognition Models in Plant Self-Incompatibility. Nat. Plants 2016, 2, 16130. [Google Scholar] [CrossRef] [PubMed]
| Clone Name | Species | Ploidy 1 | Remark | Reference |
|---|---|---|---|---|
| chc 525-3 | S. chacoense | 2x | Sli donor in Tuberosum | [23] |
| M6 | S7 S. chacoense | 2x | Male donor for introgression of SC, developed from chc 523-3 | [2] |
| 524–8 | S7 S. chacoense | 2x | Inbred S. chacoense clone from M6 breeding program | [2] |
| 39-7 | S. chacoense | 2x | PI 275138 | [28] |
| PI 133664–40 | S. chacoense | 2x | Genes segregate for SC in a 1:1 ratio | [29] |
| CIP 705468 | S. stenotomum | 2x | Landrace, Huasa Amarilla | [24] |
| US-W4 | S. tuberosum | DH | Parthenogenetically produced from clone ‘20–20-34′ | [30] |
| CD-320-20 | S. tuberosum | DH | Clone derived from the US-W4 | [30] |
| XD3 | S. tuberosum | 2x | Cross between US-W4 and 39–7 | [28] |
| RH89-039-16 (RH) | S. tuberosum | 2x | SC Sli haplotype at the distal end of chromosome | [25] |
| DMRH-89 | S. tuberosum | 2x | Cross between RH and S. tuberosum Group Phureja DM 1–3516 R44 (DM) | [25] |
| 1S1 | S. tuberosum (Phureja) | [29] | ||
| G254 | S. tuberosum | DH | Dihaploid extracted from Gineke | [27] |
| IVP07-1001-4 | S. chacoense | 2x | Sli/Sli, WUR Plant Breeding | [23] |
| 16HP01-66 | S. chacoense | 2x | Sli/sli, SC, Solynta breeding program | [31] |
| 17SC25-8 | S. chacoense | 2x | Sli/sli, SC, Solynta breeding program | [31] |
| Solyntus | S. chacoense | 2x | Sli/Sli, Solynta breeding program | [32] |
| 16BL5033-2702 | S. chacoense | 2x | Sli/Sli, Solynta breeding program | [33] |
| 18SC12-151 | S. chacoense | 2x | Sli/sli, SC, Solynta breeding program | |
| 18SC11-19 | S. chacoense | 2x | Sli/sli, SC, Solynta breeding program | |
| 17SC11-1157 | S. chacoense | 2x | Sli/sli, SC, Solynta breeding program | |
| 17SC11-4016 | S. chacoense | 2x | Sli/sli, SC, Solynta breeding program | |
| 320-02 | S. chacoense | Sli/sli, USDA | ||
| 17SC100-18 | S. chacoense | 2x | Sli/Sli, Solynta breeding program | |
| 17SC100-2 | S. chacoense | 2x | Sli/Sli, Solynta breeding program | |
| E172 | S. chacoense | 2x | Cross between SI, E and chc 525-3 | [34] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kardile, H.B.; Yilma, S.; Sathuvalli, V. Molecular Approaches to Overcome Self-Incompatibility in Diploid Potatoes. Plants 2022, 11, 1328. https://doi.org/10.3390/plants11101328
Kardile HB, Yilma S, Sathuvalli V. Molecular Approaches to Overcome Self-Incompatibility in Diploid Potatoes. Plants. 2022; 11(10):1328. https://doi.org/10.3390/plants11101328
Chicago/Turabian StyleKardile, Hemant Balasaheb, Solomon Yilma, and Vidyasagar Sathuvalli. 2022. "Molecular Approaches to Overcome Self-Incompatibility in Diploid Potatoes" Plants 11, no. 10: 1328. https://doi.org/10.3390/plants11101328
APA StyleKardile, H. B., Yilma, S., & Sathuvalli, V. (2022). Molecular Approaches to Overcome Self-Incompatibility in Diploid Potatoes. Plants, 11(10), 1328. https://doi.org/10.3390/plants11101328






