Dynamics of H3K27me3 Modification on Plant Adaptation to Environmental Cues
Abstract
1. Introduction
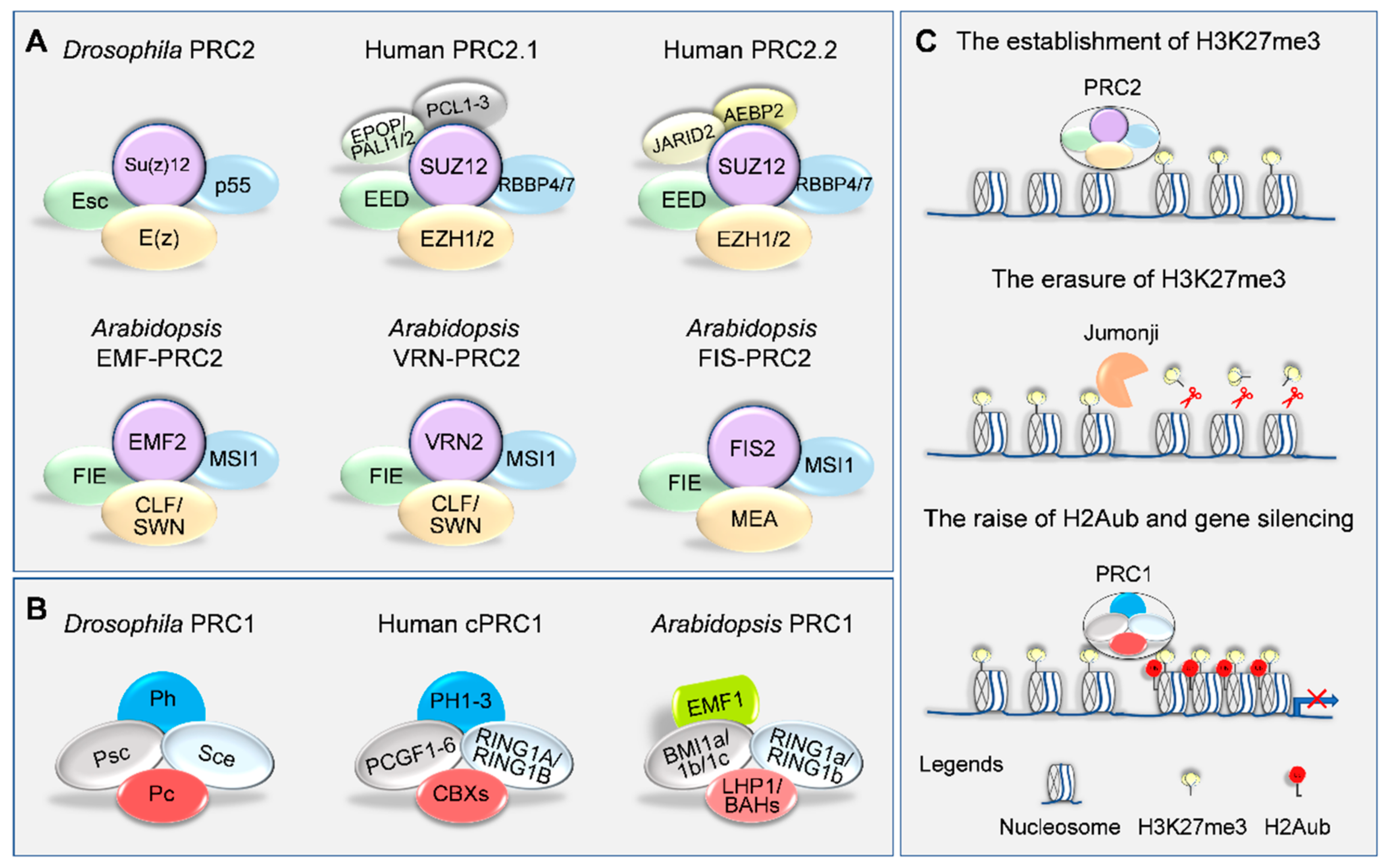
2. Writers and Erasers of H3K27me3 in Plants
3. PRC1-Mediated H3K27me3 Reading and Gene Silencing
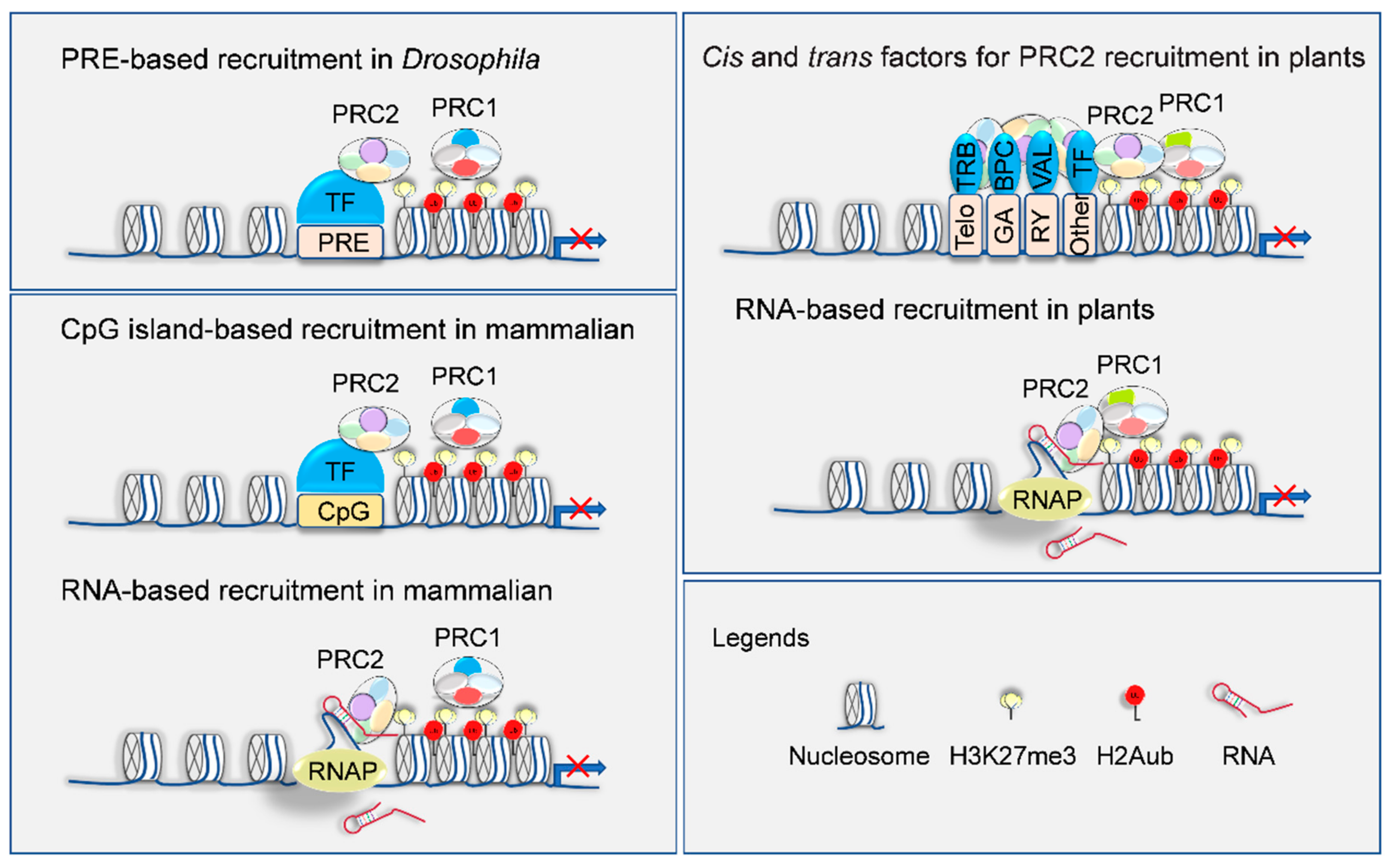
4. Targeting Mechanisms of H3K27me3 Deposition on Genome
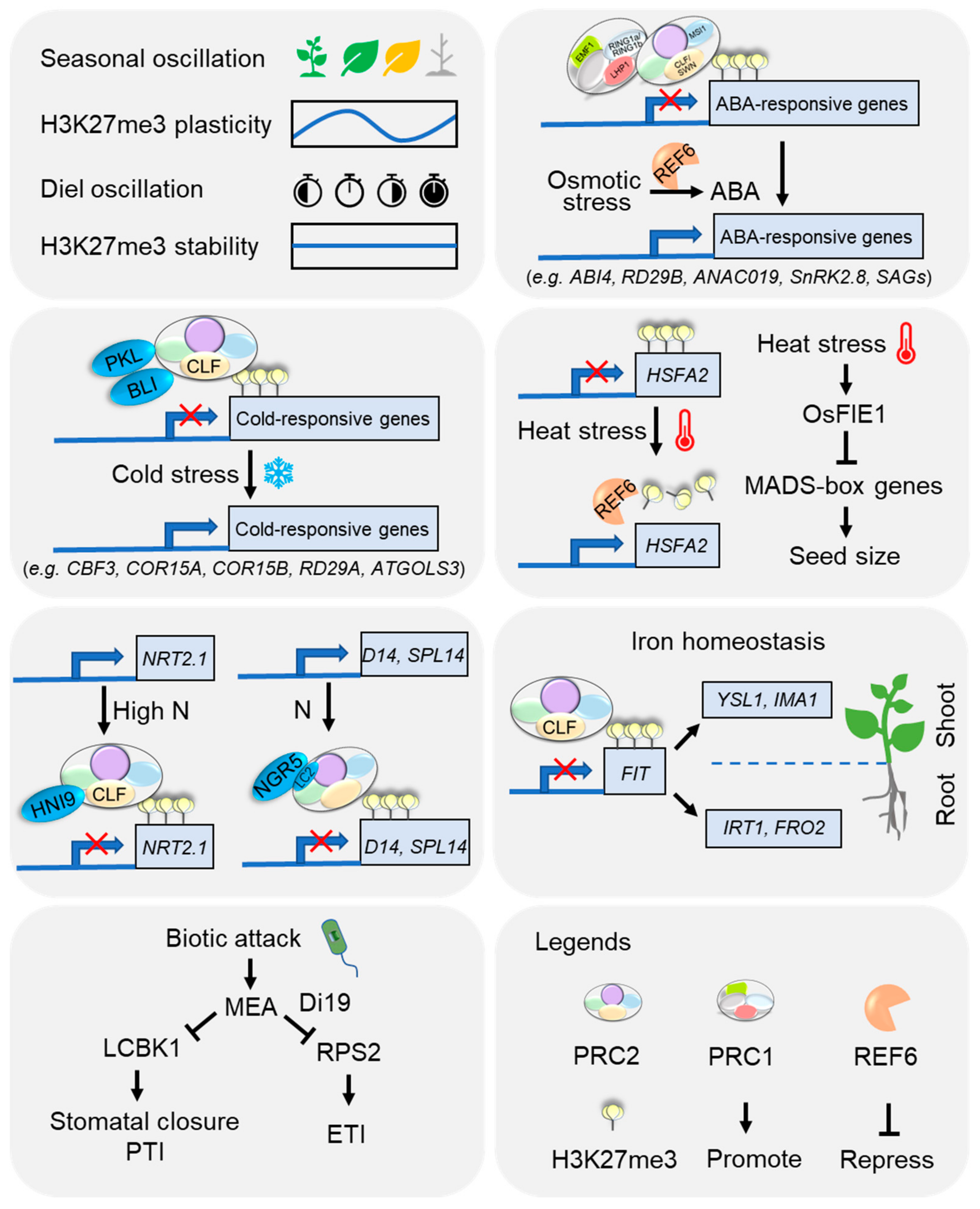
5. Emerging Roles of H3K27me3 in Plant Adaptation to Environmental Cues
5.1. Seasonal and Diel Oscillations
5.2. Extreme Temperature
5.3. Nutrients
5.4. Osmotic Stress
5.5. Biotic Stress
6. Conclusions and Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Mirouze, M.; Paszkowski, J. Epigenetic contribution to stress adaptation in plants. Curr. Opin. Plant Biol. 2011, 14, 267–274. [Google Scholar] [CrossRef]
- Chang, Y.N.; Zhu, C.; Jiang, J.; Zhang, H.; Zhu, J.K.; Duan, C.G. Epigenetic regulation in plant abiotic stress responses. J. Integr. Plant Biol. 2020, 62, 563–580. [Google Scholar] [CrossRef]
- Feng, S.; Jacobsen, S.E. Epigenetic modifications in plants: An evolutionary perspective. Curr. Opin. Plant Biol. 2011, 14, 179–186. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; He, Z. Small DNA Methylation, Big Player in Plant Abiotic Stress Responses and Memory. Front. Plant Sci. 2020, 11, 595603. [Google Scholar] [CrossRef]
- Meyer, P. Epigenetic variation and environmental change. J. Exp. Bot. 2015, 66, 3541–3548. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Lu, F.; Cui, X.; Cao, X. Histone Methylation in Higher Plants. Annu. Rev. Plant Biol. 2010, 61, 395–420. [Google Scholar] [CrossRef] [PubMed]
- Berr, A.; Shafiq, S.; Shen, W.H. Histone modifications in transcriptional activation during plant development. Biochim. Biophys. Acta 2011, 1809, 567–576. [Google Scholar] [CrossRef] [PubMed]
- Berger, S.L. The complex language of chromatin regulation during transcription. Nature 2007, 447, 407–412. [Google Scholar] [CrossRef]
- Turck, F.; Roudier, F.; Farrona, S.; Martin-Magniette, M.L.; Guillaume, E.; Buisine, N.; Gagnot, S.; Martienssen, R.A.; Coupland, G.; Colot, V. Arabidopsis TFL2/LHP1 specifically associates with genes marked by trimethylation of histone H3 lysine 27. PLoS Genet. 2007, 3, e86. [Google Scholar] [CrossRef]
- Lafos, M.; Kroll, P.; Hohenstatt, M.L.; Thorpe, F.L.; Clarenz, O.; Schubert, D. Dynamic regulation of H3K27 trimethylation during Arabidopsis differentiation. PLoS Genet. 2011, 7, e1002040. [Google Scholar] [CrossRef]
- Zhang, X.; Clarenz, O.; Cokus, S.; Bernatavichute, Y.V.; Pellegrini, M.; Goodrich, J.; Jacobsen, S.E. Whole-genome analysis of histone H3 lysine 27 trimethylation in Arabidopsis. PLoS Biol. 2007, 5, e129. [Google Scholar] [CrossRef] [PubMed]
- Margueron, R.; Reinberg, D. The Polycomb complex PRC2 and its mark in life. Nature 2011, 469, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Lanzuolo, C.; Orlando, V. Memories from the polycomb group proteins. Annu. Rev. Genet. 2012, 46, 561–589. [Google Scholar] [CrossRef] [PubMed]
- Simon, J.A.; Kingston, R.E. Occupying Chromatin: Polycomb Mechanisms for Getting to Genomic Targets, Stopping Transcriptional Traffic, and Staying Put. Mol. Cell 2013, 49, 808–824. [Google Scholar] [CrossRef] [PubMed]
- Czermin, B.; Melfi, R.; McCabe, D.; Seitz, V.; Imhof, A.; Pirrotta, V. Drosophila enhancer of Zeste/ESC complexes have a histone H3 methyltransferase activity that marks chromosomal Polycomb sites. Cell 2002, 111, 185–196. [Google Scholar] [CrossRef]
- Suganuma, T.; Pattenden, S.G.; Workman, J.L. Diverse functions of WD40 repeat proteins in histone recognition. Genes. Dev. 2008, 22, 1265–1268. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Birve, A.; Rasmuson-Lestander, A. In vivo analysis of Drosophila SU(Z)12 function. Mol. Genet. Genom. 2008, 279, 159–170. [Google Scholar] [CrossRef]
- Wen, P.; Quan, Z.; Xi, R. The biological function of the WD40 repeat-containing protein p55/Caf1 in Drosophila. Dev. Dyn. 2012, 241, 455–464. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Zhao, S.; Wang, G.G. Polycomb Gene Silencing Mechanisms: PRC2 Chromatin Targeting, H3K27me3 ’Readout’, and Phase Separation-Based Compaction. Trends Genet. 2021. [Google Scholar] [CrossRef] [PubMed]
- Vann, K.R.; Kutateladze, T.G. Architecture of PRC2 Holo Complexes. Trends Biochem. Sci. 2018, 43, 487–489. [Google Scholar] [CrossRef]
- Van Mierlo, G.; Veenstra, G.J.C.; Vermeulen, M.; Marks, H. The Complexity of PRC2 Subcomplexes. Trends Cell Biol. 2019, 29, 660–671. [Google Scholar] [CrossRef] [PubMed]
- Youmans, D.T.; Gooding, A.R.; Dowell, R.D.; Cech, T.R. Competition between PRC2.1 and 2.2 subcomplexes regulates PRC2 chromatin occupancy in human stem cells. Mol. Cell 2021, 81, 488–501.e489. [Google Scholar] [CrossRef]
- Healy, E.; Mucha, M.; Glancy, E.; Fitzpatrick, D.J.; Conway, E.; Neikes, H.K.; Monger, C.; Van Mierlo, G.; Baltissen, M.P.; Koseki, Y.; et al. PRC2.1 and PRC2.2 Synergize to Coordinate H3K27 Trimethylation. Mol. Cell 2019, 76, 437–452.e436. [Google Scholar] [CrossRef]
- Huang, Y.; Chen, D.H.; Liu, B.Y.; Shen, W.H.; Ruan, Y. Conservation and diversification of polycomb repressive complex 2 (PRC2) proteins in the green lineage. Brief Funct. Genom. 2017, 16, 106–119. [Google Scholar] [CrossRef]
- Butenko, Y.; Ohad, N. Polycomb-group mediated epigenetic mechanisms through plant evolution. Biochim. Biophys. Acta 2011, 1809, 395–406. [Google Scholar] [CrossRef]
- Pien, S.; Grossniklaus, U. Polycomb group and trithorax group proteins in Arabidopsis. Biochim. Biophys. Acta 2007, 1769, 375–382. [Google Scholar] [CrossRef]
- Kim, D.H.; Sung, S. Polycomb-mediated gene silencing in Arabidopsis thaliana. Mol. Cells 2014, 37, 841–850. [Google Scholar] [CrossRef] [PubMed]
- Kiyosue, T.; Ohad, N.; Yadegari, R.; Hannon, M.; Dinneny, J.; Wells, D.; Katz, A.; Margossian, L.; Harada, J.J.; Goldberg, R.B.; et al. Control of fertilization-independent endosperm development by the MEDEA polycomb gene in Arabidopsis. Proc. Natl. Acad. Sci. USA 1999, 96, 4186–4191. [Google Scholar] [CrossRef] [PubMed]
- Chaudhury, A.M.; Ming, L.; Miller, C.; Craig, S.; Dennis, E.S.; Peacock, W.J. Fertilization-independent seed development in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 1997, 94, 4223–4228. [Google Scholar] [CrossRef]
- Guitton, A.E.; Page, D.R.; Chambrier, P.; Lionnet, C.; Faure, J.E.; Grossniklaus, U.; Berger, F. Identification of new members of Fertilisation Independent Seed Polycomb Group pathway involved in the control of seed development in Arabidopsis thaliana. Development 2004, 131, 2971–2981. [Google Scholar] [CrossRef]
- Grossniklaus, U.; Vielle-Calzada, J.P.; Hoeppner, M.A.; Gagliano, W.B. Maternal control of embryogenesis by MEDEA, a polycomb group gene in Arabidopsis. Science 1998, 280, 446–450. [Google Scholar] [CrossRef]
- Ohad, N.; Margossian, L.; Hsu, Y.C.; Williams, C.; Repetti, P.; Fischer, R.L. A mutation that allows endosperm development without fertilization. Proc. Natl. Acad. Sci. USA 1996, 93, 5319–5324. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Kohler, C. Epigenetic processes in flowering plant reproduction. J Exp Bot 2017, 68, 797–807. [Google Scholar] [CrossRef]
- Chanvivattana, Y.; Bishopp, A.; Schubert, D.; Stock, C.; Moon, Y.H.; Sung, Z.R.; Goodrich, J. Interaction of Polycomb-group proteins controlling flowering in Arabidopsis. Development 2004, 131, 5263–5276. [Google Scholar] [CrossRef]
- Yang, C.H.; Chen, L.J.; Sung, Z.R. Genetic regulation of shoot development in Arabidopsis: Role of the EMF genes. Dev. Biol. 1995, 169, 421–435. [Google Scholar] [CrossRef]
- Kim, S.Y.; Zhu, T.; Sung, Z.R. Epigenetic regulation of gene programs by EMF1 and EMF2 in Arabidopsis. Plant Physiol. 2010, 152, 516–528. [Google Scholar] [CrossRef] [PubMed]
- Gendall, A.R.; Levy, Y.Y.; Wilson, A.; Dean, C. The VERNALIZATION 2 gene mediates the epigenetic regulation of vernalization in Arabidopsis. Cell 2001, 107, 525–535. [Google Scholar] [CrossRef]
- De Lucia, F.; Crevillen, P.; Jones, A.M.; Greb, T.; Dean, C. A PHD-polycomb repressive complex 2 triggers the epigenetic silencing of FLC during vernalization. Proc. Natl. Acad. Sci. USA 2008, 105, 16831–16836. [Google Scholar] [CrossRef] [PubMed]
- Wood, C.C.; Robertson, M.; Tanner, G.; Peacock, W.J.; Dennis, E.S.; Helliwell, C.A. The Arabidopsis thaliana vernalization response requires a polycomb-like protein complex that also includes VERNALIZATION INSENSITIVE 3. Proc. Natl. Acad. Sci. USA 2006, 103, 14631–14636. [Google Scholar] [CrossRef]
- Ni, J.; Ma, X.; Feng, Y.; Tian, Q.; Wang, Y.; Xu, N.; Tang, J.; Wang, G. Updating and interaction of polycomb repressive complex 2 components in maize (Zea mays). Planta 2019, 250, 573–588. [Google Scholar] [CrossRef]
- Cheng, X.; Pan, M.; Zhiguo, E.; Zhou, Y.; Niu, B.; Chen, C. The maternally expressed polycomb group gene OsEMF2a is essential for endosperm cellularization and imprinting in rice. Plant Commun. 2021, 2, 100092. [Google Scholar] [CrossRef] [PubMed]
- Tonosaki, K.; Ono, A.; Kunisada, M.; Nishino, M.; Nagata, H.; Sakamoto, S.; Kijima, S.T.; Furuumi, H.; Nonomura, K.I.; Sato, Y.; et al. Mutation of the imprinted gene OsEMF2a induces autonomous endosperm development and delayed cellularization in rice. Plant Cell 2021, 33, 85–103. [Google Scholar] [CrossRef]
- Conrad, L.J.; Khanday, I.; Johnson, C.; Guiderdoni, E.; An, G.; Vijayraghavan, U.; Sundaresan, V. The polycomb group gene EMF2B is essential for maintenance of floral meristem determinacy in rice. Plant J. 2014, 80, 883–894. [Google Scholar] [CrossRef]
- Zhong, J.; Peng, Z.; Peng, Q.; Cai, Q.; Peng, W.; Chen, M.; Yao, J. Regulation of plant height in rice by the Polycomb group genes OsEMF2b, OsFIE2 and OsCLF. Plant Sci. 2018, 267, 157–167. [Google Scholar] [CrossRef]
- Black, J.C.; Van Rechem, C.; Whetstine, J.R. Histone lysine methylation dynamics: Establishment, regulation, and biological impact. Mol. Cell 2012, 48, 491–507. [Google Scholar] [CrossRef] [PubMed]
- Klose, R.J.; Kallin, E.M.; Zhang, Y. JmjC-domain-containing proteins and histone demethylation. Nat. Rev. Genet. 2006, 7, 715–727. [Google Scholar] [CrossRef]
- Hong, S.; Cho, Y.W.; Yu, L.R.; Yu, H.; Veenstra, T.D.; Ge, K. Identification of JmjC domain-containing UTX and JMJD3 as histone H3 lysine 27 demethylases. Proc. Natl. Acad. Sci. USA 2007, 104, 18439–18444. [Google Scholar] [CrossRef] [PubMed]
- Agger, K.; Cloos, P.A.; Christensen, J.; Pasini, D.; Rose, S.; Rappsilber, J.; Issaeva, I.; Canaani, E.; Salcini, A.E.; Helin, K. UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature 2007, 449, 731–734. [Google Scholar] [CrossRef]
- Li, G.; Margueron, R.; Ku, M.; Chambon, P.; Bernstein, B.E.; Reinberg, D. Jarid2 and PRC2, partners in regulating gene expression. Genes. Dev. 2010, 24, 368–380. [Google Scholar] [CrossRef] [PubMed]
- Pasini, D.; Cloos, P.A.; Walfridsson, J.; Olsson, L.; Bukowski, J.P.; Johansen, J.V.; Bak, M.; Tommerup, N.; Rappsilber, J.; Helin, K. JARID2 regulates binding of the Polycomb repressive complex 2 to target genes in ES cells. Nature 2010, 464, 306–310. [Google Scholar] [CrossRef]
- Crevillén, P. Histone Demethylases as Counterbalance to H3K27me3 Silencing in Plants. iScience 2020, 23, 101715. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Ou, Y.; Zhang, Z.; Li, J.; He, Y. Brassinosteroid Signaling Recruits Histone 3 Lysine-27 Demethylation Activity to FLOWERING LOCUS C Chromatin to Inhibit the Floral Transition in Arabidopsis. Mol. Plant 2018, 11, 1135–1146. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Li, L.; Li, L.; Guo, M.; Chory, J.; Yin, Y. Modulation of brassinosteroid-regulated gene expression by Jumonji domain-containing proteins ELF6 and REF6 in Arabidopsis. Proc. Natl. Acad. Sci. USA 2008, 105, 7618–7623. [Google Scholar] [CrossRef]
- Li, C.; Gu, L.; Gao, L.; Chen, C.; Wei, C.Q.; Qiu, Q.; Chien, C.W.; Wang, S.; Jiang, L.; Ai, L.F.; et al. Concerted genomic targeting of H3K27 demethylase REF6 and chromatin-remodeling ATPase BRM in Arabidopsis. Nat. Genet. 2016, 48, 687–693. [Google Scholar] [CrossRef]
- Noh, B.; Lee, S.H.; Kim, H.J.; Yi, G.; Shin, E.A.; Lee, M.; Jung, K.J.; Doyle, M.R.; Amasino, R.M.; Noh, Y.S. Divergent roles of a pair of homologous jumonji/zinc-finger-class transcription factor proteins in the regulation of Arabidopsis flowering time. Plant Cell 2004, 16, 2601–2613. [Google Scholar] [CrossRef]
- Lu, F.; Cui, X.; Zhang, S.; Jenuwein, T.; Cao, X. Arabidopsis REF6 is a histone H3 lysine 27 demethylase. Nat. Genet. 2011, 43, 715–719. [Google Scholar] [CrossRef]
- Tian, Z.; Li, X.; Li, M.; Wu, W.; Zhang, M.; Tang, C.; Li, Z.; Liu, Y.; Chen, Z.; Yang, M.; et al. Crystal structures of REF6 and its complex with DNA reveal diverse recognition mechanisms. Cell Discov. 2020, 6, 17. [Google Scholar] [CrossRef] [PubMed]
- Zheng, S.; Hu, H.; Ren, H.; Yang, Z.; Qiu, Q.; Qi, W.; Liu, X.; Chen, X.; Cui, X.; Li, S.; et al. The Arabidopsis H3K27me3 demethylase JUMONJI 13 is a temperature and photoperiod dependent flowering repressor. Nat. Commun. 2019, 10, 1303. [Google Scholar] [CrossRef]
- Gan, E.S.; Xu, Y.; Wong, J.Y.; Goh, J.G.; Sun, B.; Wee, W.Y.; Huang, J.; Ito, T. Jumonji demethylases moderate precocious flowering at elevated temperature via regulation of FLC in Arabidopsis. Nat. Commun. 2014, 5, 5098. [Google Scholar] [CrossRef]
- Li, T.; Chen, X.; Zhong, X.; Zhao, Y.; Liu, X.; Zhou, S.; Cheng, S.; Zhou, D.X. Jumonji C domain protein JMJ705-mediated removal of histone H3 lysine 27 trimethylation is involved in defense-related gene activation in rice. Plant Cell 2013, 25, 4725–4736. [Google Scholar] [CrossRef]
- Cheng, S.; Tan, F.; Lu, Y.; Liu, X.; Li, T.; Yuan, W.; Zhao, Y.; Zhou, D.X. WOX11 recruits a histone H3K27me3 demethylase to promote gene expression during shoot development in rice. Nucleic Acids Res. 2018, 46, 2356–2369. [Google Scholar] [CrossRef]
- Qian, Y.; Chen, C.; Jiang, L.; Zhang, J.; Ren, Q. Genome-wide identification, classification and expression analysis of the JmjC domain-containing histone demethylase gene family in maize. BMC Genom. 2019, 20, 256. [Google Scholar] [CrossRef]
- Cao, R.; Wang, L.; Wang, H.; Xia, L.; Erdjument-Bromage, H.; Tempst, P.; Jones, R.S.; Zhang, Y. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 2002, 298, 1039–1043. [Google Scholar] [CrossRef] [PubMed]
- Wani, A.H.; Boettiger, A.N.; Schorderet, P.; Ergun, A.; Münger, C.; Sadreyev, R.I.; Zhuang, X.; Kingston, R.E.; Francis, N.J. Chromatin topology is coupled to Polycomb group protein subnuclear organization. Nat. Commun. 2016, 7, 10291. [Google Scholar] [CrossRef] [PubMed]
- Eskeland, R.; Leeb, M.; Grimes, G.R.; Kress, C.; Boyle, S.; Sproul, D.; Gilbert, N.; Fan, Y.; Skoultchi, A.I.; Wutz, A.; et al. Ring1B compacts chromatin structure and represses gene expression independent of histone ubiquitination. Mol. Cell 2010, 38, 452–464. [Google Scholar] [CrossRef]
- Lehmann, L.; Ferrari, R.; Vashisht, A.A.; Wohlschlegel, J.A.; Kurdistani, S.K.; Carey, M. Polycomb repressive complex 1 (PRC1) disassembles RNA polymerase II preinitiation complexes. J. Biol. Chem. 2012, 287, 35784–35794. [Google Scholar] [CrossRef]
- Yin, X.; Romero-Campero, F.J.; de Los Reyes, P.; Yan, P.; Yang, J.; Tian, G.; Yang, X.; Mo, X.; Zhao, S.; Calonje, M.; et al. H2AK121ub in Arabidopsis associates with a less accessible chromatin state at transcriptional regulation hotspots. Nat. Commun. 2021, 12, 315. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, Y.B.; Kahn, T.G.; Nix, D.A.; Li, X.Y.; Bourgon, R.; Biggin, M.; Pirrotta, V. Genome-wide analysis of Polycomb targets in Drosophila melanogaster. Nat. Genet. 2006, 38, 700–705. [Google Scholar] [CrossRef]
- Tolhuis, B.; de Wit, E.; Muijrers, I.; Teunissen, H.; Talhout, W.; van Steensel, B.; van Lohuizen, M. Genome-wide profiling of PRC1 and PRC2 Polycomb chromatin binding in Drosophila melanogaster. Nat. Genet. 2006, 38, 694–699. [Google Scholar] [CrossRef]
- Ku, M.; Koche, R.P.; Rheinbay, E.; Mendenhall, E.M.; Endoh, M.; Mikkelsen, T.S.; Presser, A.; Nusbaum, C.; Xie, X.; Chi, A.S.; et al. Genomewide analysis of PRC1 and PRC2 occupancy identifies two classes of bivalent domains. PLoS Genet. 2008, 4, e1000242. [Google Scholar] [CrossRef]
- Kim, S.Y.; Lee, J.; Eshed-Williams, L.; Zilberman, D.; Sung, Z.R. EMF1 and PRC2 cooperate to repress key regulators of Arabidopsis development. PLoS Genet. 2012, 8, e1002512. [Google Scholar] [CrossRef] [PubMed]
- Blackledge, N.P.; Farcas, A.M.; Kondo, T.; King, H.W.; McGouran, J.F.; Hanssen, L.L.P.; Ito, S.; Cooper, S.; Kondo, K.; Koseki, Y.; et al. Variant PRC1 complex-dependent H2A ubiquitylation drives PRC2 recruitment and polycomb domain formation. Cell 2014, 157, 1445–1459. [Google Scholar] [CrossRef]
- Zhou, Y.; Romero-Campero, F.J.; Gómez-Zambrano, Á.; Turck, F.; Calonje, M. H2A monoubiquitination in Arabidopsis thaliana is generally independent of LHP1 and PRC2 activity. Genome Biol. 2017, 18, 69. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Bratzel, F.; Hohmann, N.; Koch, M.; Turck, F.; Calonje, M. VAL- and AtBMI1-mediated H2Aub initiate the switch from embryonic to postgerminative growth in Arabidopsis. Curr. Biol. 2013, 23, 1324–1329. [Google Scholar] [CrossRef]
- Kalb, R.; Latwiel, S.; Baymaz, H.I.; Jansen, P.W.; Müller, C.W.; Vermeulen, M.; Müller, J. Histone H2A monoubiquitination promotes histone H3 methylation in Polycomb repression. Nat. Struct. Mol. Biol. 2014, 21, 569–571. [Google Scholar] [CrossRef]
- Tavares, L.; Dimitrova, E.; Oxley, D.; Webster, J.; Poot, R.; Demmers, J.; Bezstarosti, K.; Taylor, S.; Ura, H.; Koide, H.; et al. RYBP-PRC1 complexes mediate H2A ubiquitylation at polycomb target sites independently of PRC2 and H3K27me3. Cell 2012, 148, 664–678. [Google Scholar] [CrossRef] [PubMed]
- Kralemann, L.E.M.; Liu, S.; Trejo-Arellano, M.S.; Muñoz-Viana, R.; Köhler, C.; Hennig, L. Removal of H2Aub1 by ubiquitin-specific proteases 12 and 13 is required for stable Polycomb-mediated gene repression in Arabidopsis. Genome Biol. 2020, 21, 144. [Google Scholar] [CrossRef] [PubMed]
- Shao, Z.; Raible, F.; Mollaaghababa, R.; Guyon, J.R.; Wu, C.T.; Bender, W.; Kingston, R.E. Stabilization of chromatin structure by PRC1, a Polycomb complex. Cell 1999, 98, 37–46. [Google Scholar] [CrossRef]
- Brunk, B.P.; Martin, E.C.; Adler, P.N. Drosophila genes Posterior Sex Combs and Suppressor two of zeste encode proteins with homology to the murine bmi-1 oncogene. Nature 1991, 353, 351–353. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, L.; Oktaba, K.; Scheuermann, J.C.; Gambetta, M.C.; Ly-Hartig, N.; Müller, J. The role of the histone H2A ubiquitinase Sce in Polycomb repression. Development 2012, 139, 117–127. [Google Scholar] [CrossRef]
- Messmer, S.; Franke, A.; Paro, R. Analysis of the functional role of the Polycomb chromo domain in Drosophila melanogaster. Genes. Dev. 1992, 6, 1241–1254. [Google Scholar] [CrossRef][Green Version]
- Eissenberg, J.C. Structural biology of the chromodomain: Form and function. Gene 2012, 496, 69–78. [Google Scholar] [CrossRef] [PubMed]
- Narbonne, K.; Besse, F.; Brissard-Zahraoui, J.; Pret, A.M.; Busson, D. polyhomeotic is required for somatic cell proliferation and differentiation during ovarian follicle formation in Drosophila. Development 2004, 131, 1389–1400. [Google Scholar] [CrossRef][Green Version]
- Schoorlemmer, J.; Marcos-Gutiérrez, C.; Were, F.; Martínez, R.; García, E.; Satijn, D.P.; Otte, A.P.; Vidal, M. Ring1A is a transcriptional repressor that interacts with the Polycomb-M33 protein and is expressed at rhombomere boundaries in the mouse hindbrain. EMBO J. 1997, 16, 5930–5942. [Google Scholar] [CrossRef]
- Gao, Z.; Zhang, J.; Bonasio, R.; Strino, F.; Sawai, A.; Parisi, F.; Kluger, Y.; Reinberg, D. PCGF homologs, CBX proteins, and RYBP define functionally distinct PRC1 family complexes. Mol. Cell 2012, 45, 344–356. [Google Scholar] [CrossRef]
- Morey, L.; Pascual, G.; Cozzuto, L.; Roma, G.; Wutz, A.; Benitah, S.A.; Di Croce, L. Nonoverlapping functions of the Polycomb group Cbx family of proteins in embryonic stem cells. Cell Stem. Cell 2012, 10, 47–62. [Google Scholar] [CrossRef]
- Tonkin, E.; Hagan, D.M.; Li, W.; Strachan, T. Identification and characterisation of novel mammalian homologues of Drosophila polyhomeoticpermits new insights into relationships between members of the polyhomeotic family. Hum. Genet. 2002, 111, 435–442. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Shen, W.H. Polycomb silencing of KNOX genes confines shoot stem cell niches in Arabidopsis. Curr. Biol. 2008, 18, 1966–1971. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Pulido, L.; Devos, D.; Sung, Z.R.; Calonje, M. RAWUL: A new ubiquitin-like domain in PRC1 ring finger proteins that unveils putative plant and worm PRC1 orthologs. BMC Genom. 2008, 9, 308. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Molitor, A.; Liu, C.; Shen, W.H. The Arabidopsis PRC1-like ring-finger proteins are necessary for repression of embryonic traits during vegetative growth. Cell Res. 2010, 20, 1332–1344. [Google Scholar] [CrossRef]
- Bratzel, F.; López-Torrejón, G.; Koch, M.; Del Pozo, J.C.; Calonje, M. Keeping cell identity in Arabidopsis requires PRC1 RING-finger homologs that catalyze H2A monoubiquitination. Curr. Biol. 2010, 20, 1853–1859. [Google Scholar] [CrossRef]
- Li, W.; Wang, Z.; Li, J.; Yang, H.; Cui, S.; Wang, X.; Ma, L. Overexpression of AtBMI1C, a polycomb group protein gene, accelerates flowering in Arabidopsis. PLoS ONE 2011, 6, e21364. [Google Scholar] [CrossRef] [PubMed]
- Bratzel, F.; Yang, C.; Angelova, A.; López-Torrejón, G.; Koch, M.; del Pozo, J.C.; Calonje, M. Regulation of the new Arabidopsis imprinted gene AtBMI1C requires the interplay of different epigenetic mechanisms. Mol. Plant 2012, 5, 260–269. [Google Scholar] [CrossRef]
- Merini, W.; Romero-Campero, F.J.; Gomez-Zambrano, A.; Zhou, Y.; Turck, F.; Calonje, M. The Arabidopsis Polycomb Repressive Complex 1 (PRC1) Components AtBMI1A, B, and C Impact Gene Networks throughout All Stages of Plant Development. Plant Physiol. 2017, 173, 627–641. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Jiang, L.; Liu, B.Y.; Tan, C.F.; Chen, D.H.; Shen, W.H.; Ruan, Y. Evolution and conservation of polycomb repressive complex 1 core components and putative associated factors in the green lineage. BMC Genom. 2019, 20, 533. [Google Scholar] [CrossRef] [PubMed]
- Krause, K.; Turck, F. Plant H3K27me3 has finally found its readers. Nat. Genet. 2018, 50, 1206–1208. [Google Scholar] [CrossRef]
- Kotake, T.; Takada, S.; Nakahigashi, K.; Ohto, M.; Goto, K. Arabidopsis TERMINAL FLOWER 2 gene encodes a heterochromatin protein 1 homolog and represses both FLOWERING LOCUS T to regulate flowering time and several floral homeotic genes. Plant Cell Physiol. 2003, 44, 555–564. [Google Scholar] [CrossRef]
- Gaudin, V.; Libault, M.; Pouteau, S.; Juul, T.; Zhao, G.; Lefebvre, D.; Grandjean, O. Mutations in LIKE HETEROCHROMATIN PROTEIN 1 affect flowering time and plant architecture in Arabidopsis. Development 2001, 128, 4847–4858. [Google Scholar] [CrossRef]
- Yang, Z.; Qian, S.; Scheid, R.N.; Lu, L.; Chen, X.; Liu, R.; Du, X.; Lv, X.; Boersma, M.D.; Scalf, M.; et al. EBS is a bivalent histone reader that regulates floral phase transition in Arabidopsis. Nat. Genet. 2018, 50, 1247–1253. [Google Scholar] [CrossRef]
- Li, Z.; Fu, X.; Wang, Y.; Liu, R.; He, Y. Polycomb-mediated gene silencing by the BAH-EMF1 complex in plants. Nat. Genet. 2018, 50, 1254–1261. [Google Scholar] [CrossRef]
- Qian, S.; Lv, X.; Scheid, R.N.; Lu, L.; Yang, Z.; Chen, W.; Liu, R.; Boersma, M.D.; Denu, J.M.; Zhong, X.; et al. Dual recognition of H3K4me3 and H3K27me3 by a plant histone reader SHL. Nat. Commun. 2018, 9, 2425. [Google Scholar] [CrossRef] [PubMed]
- Qian, F.; Zhao, Q.Y.; Zhang, T.N.; Li, Y.L.; Su, Y.N.; Li, L.; Sui, J.H.; Chen, S.; He, X.J. A histone H3K27me3 reader cooperates with a family of PHD finger-containing proteins to regulate flowering time in Arabidopsis. J. Integr. Plant Biol. 2021. [Google Scholar] [CrossRef]
- Zhang, Y.Z.; Yuan, J.; Zhang, L.; Chen, C.; Wang, Y.; Zhang, G.; Peng, L.; Xie, S.S.; Jiang, J.; Zhu, J.K.; et al. Coupling of H3K27me3 recognition with transcriptional repression through the BAH-PHD-CPL2 complex in Arabidopsis. Nat. Commun. 2020, 11, 6212. [Google Scholar] [CrossRef]
- López-González, L.; Mouriz, A.; Narro-Diego, L.; Bustos, R.; Martínez-Zapater, J.M.; Jarillo, J.A.; Piñeiro, M. Chromatin-dependent repression of the Arabidopsis floral integrator genes involves plant specific PHD-containing proteins. Plant Cell 2014, 26, 3922–3938. [Google Scholar] [CrossRef]
- Aubert, D.; Chen, L.; Moon, Y.H.; Martin, D.; Castle, L.A.; Yang, C.H.; Sung, Z.R. EMF1, a novel protein involved in the control of shoot architecture and flowering in Arabidopsis. Plant Cell 2001, 13, 1865–1875. [Google Scholar] [CrossRef] [PubMed]
- Moon, Y.H.; Chen, L.; Pan, R.L.; Chang, H.S.; Zhu, T.; Maffeo, D.M.; Sung, Z.R. EMF genes maintain vegetative development by repressing the flower program in Arabidopsis. Plant Cell 2003, 15, 681–693. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Cheng, J.C.; Castle, L.; Sung, Z.R. EMF genes regulate Arabidopsis inflorescence development. Plant Cell 1997, 9, 2011–2024. [Google Scholar] [CrossRef]
- Beltran, M.; Yates, C.M.; Skalska, L.; Dawson, M.; Reis, F.P.; Viiri, K.; Fisher, C.L.; Sibley, C.R.; Foster, B.M.; Bartke, T.; et al. The interaction of PRC2 with RNA or chromatin is mutually antagonistic. Genome Res. 2016, 26, 896–907. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Sun, B.K.; Erwin, J.A.; Song, J.J.; Lee, J.T. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 2008, 322, 750–756. [Google Scholar] [CrossRef]
- Kassis, J.A.; Brown, J.L. Polycomb group response elements in Drosophila and vertebrates. Adv. Genet. 2013, 81, 83–118. [Google Scholar] [CrossRef] [PubMed]
- Laugesen, A.; Højfeldt, J.W.; Helin, K. Molecular Mechanisms Directing PRC2 Recruitment and H3K27 Methylation. Mol. Cell 2019, 74, 8–18. [Google Scholar] [CrossRef]
- Rinn, J.L.; Kertesz, M.; Wang, J.K.; Squazzo, S.L.; Xu, X.; Brugmann, S.A.; Goodnough, L.H.; Helms, J.A.; Farnham, P.J.; Segal, E.; et al. Functional Demarcation of Active and Silent Chromatin Domains in Human HOX Loci by Noncoding RNAs. Cell 2007, 129, 1311–1323. [Google Scholar] [CrossRef] [PubMed]
- Ballaré, C.; Lange, M.; Lapinaite, A.; Martin, G.M.; Morey, L.; Pascual, G.; Liefke, R.; Simon, B.; Shi, Y.; Gozani, O.; et al. Phf19 links methylated Lys36 of histone H3 to regulation of Polycomb activity. Nat. Struct. Mol. Biol. 2012, 19, 1257–1265. [Google Scholar] [CrossRef] [PubMed]
- Cai, L.; Rothbart, S.B.; Lu, R.; Xu, B.; Chen, W.-Y.; Tripathy, A.; Rockowitz, S.; Zheng, D.; Patel, D.J.; Allis, C.D.; et al. An H3K36 Methylation-Engaging Tudor Motif of Polycomb-like Proteins Mediates PRC2 Complex Targeting. Mol. Cell 2013, 49, 571–582. [Google Scholar] [CrossRef]
- Jermann, P.; Hoerner, L.; Burger, L.; Schübeler, D. Short sequences can efficiently recruit histone H3 lysine 27 trimethylation in the absence of enhancer activity and DNA methylation. Proc. Natl. Acad. Sci. USA 2014, 111, E3415. [Google Scholar] [CrossRef] [PubMed]
- Lynch, M.D.; Smith, A.J.; De Gobbi, M.; Flenley, M.; Hughes, J.R.; Vernimmen, D.; Ayyub, H.; Sharpe, J.A.; Sloane-Stanley, J.A.; Sutherland, L.; et al. An interspecies analysis reveals a key role for unmethylated CpG dinucleotides in vertebrate Polycomb complex recruitment. EMBO J. 2012, 31, 317–329. [Google Scholar] [CrossRef]
- Hecker, A.; Brand, L.H.; Peter, S.; Simoncello, N.; Kilian, J.; Harter, K.; Gaudin, V.; Wanke, D. The Arabidopsis GAGA-Binding Factor BASIC PENTACYSTEINE6 Recruits the POLYCOMB-REPRESSIVE COMPLEX1 Component LIKE HETEROCHROMATIN PROTEIN1 to GAGA DNA Motifs. Plant Physiol. 2015, 168, 1013–1024. [Google Scholar] [CrossRef]
- Deng, W.; Buzas, D.M.; Ying, H.; Robertson, M.; Taylor, J.; Peacock, W.J.; Dennis, E.S.; Helliwell, C. Arabidopsis Polycomb Repressive Complex 2 binding sites contain putative GAGA factor binding motifs within coding regions of genes. BMC Genom. 2013, 14, 593. [Google Scholar] [CrossRef]
- Mu, Y.; Zou, M.; Sun, X.; He, B.; Xu, X.; Liu, Y.; Zhang, L.; Chi, W. BASIC PENTACYSTEINE Proteins Repress ABSCISIC ACID INSENSITIVE4 Expression via Direct Recruitment of the Polycomb-Repressive Complex 2 in Arabidopsis Root Development. Plant Cell Physiol. 2017, 58, 607–621. [Google Scholar] [CrossRef] [PubMed]
- Xiao, J.; Jin, R.; Yu, X.; Shen, M.; Wagner, J.D.; Pai, A.; Song, C.; Zhuang, M.; Klasfeld, S.; He, C.; et al. Cis and trans determinants of epigenetic silencing by Polycomb repressive complex 2 in Arabidopsis. Nat. Genet. 2017, 49, 1546–1552. [Google Scholar] [CrossRef]
- Theune, M.L.; Bloss, U.; Brand, L.H.; Ladwig, F.; Wanke, D. Phylogenetic Analyses and GAGA-Motif Binding Studies of BBR/BPC Proteins Lend to Clues in GAGA-Motif Recognition and a Regulatory Role in Brassinosteroid Signaling. Front. Plant Sci. 2019, 10, 466. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Mohamed, D.; Dowhanik, S.; Petrella, R.; Gregis, V.; Li, J.; Wu, L.; Gazzarrini, S. Spatiotemporal Restriction of FUSCA3 Expression by Class I BPCs Promotes Ovule Development and Coordinates Embryo and Endosperm Growth. Plant Cell 2020, 32, 1886–1904. [Google Scholar] [CrossRef]
- Zhou, Y.; Hartwig, B.; James, G.V.; Schneeberger, K.; Turck, F. Complementary Activities of TELOMERE REPEAT BINDING Proteins and Polycomb Group Complexes in Transcriptional Regulation of Target Genes. Plant Cell 2016, 28, 87–101. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Wang, Y.; Krause, K.; Yang, T.; Dongus, J.A.; Zhang, Y.; Turck, F. Telobox motifs recruit CLF/SWN-PRC2 for H3K27me3 deposition via TRB factors in Arabidopsis. Nat. Genet. 2018, 50, 638–644. [Google Scholar] [CrossRef]
- Marión, R.M.; Montero, J.J.; López de Silanes, I.; Graña-Castro, O.; Martínez, P.; Schoeftner, S.; Palacios-Fábrega, J.A.; Blasco, M.A. TERRA regulate the transcriptional landscape of pluripotent cells through TRF1-dependent recruitment of PRC2. Elife 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Qüesta, J.I.; Song, J.; Geraldo, N.; An, H.; Dean, C. Arabidopsis transcriptional repressor VAL1 triggers Polycomb silencing at FLC during vernalization. Science 2016, 353, 485–488. [Google Scholar] [CrossRef]
- Jing, Y.; Guo, Q.; Lin, R. The B3-Domain Transcription Factor VAL1 Regulates the Floral Transition by Repressing FLOWERING LOCUS T. Plant Physiol. 2019, 181, 236–248. [Google Scholar] [CrossRef]
- Chen, N.; Wang, H.; Abdelmageed, H.; Veerappan, V.; Tadege, M.; Allen, R.D. HSI2/VAL1 and HSL1/VAL2 function redundantly to repress DOG1 expression in Arabidopsis seeds and seedlings. New Phytol. 2020, 227, 840–856. [Google Scholar] [CrossRef]
- Chen, N.; Veerappan, V.; Abdelmageed, H.; Kang, M.; Allen, R.D. HSI2/VAL1 Silences AGL15 to Regulate the Developmental Transition from Seed Maturation to Vegetative Growth in Arabidopsis. Plant Cell 2018, 30, 600–619. [Google Scholar] [CrossRef]
- Xie, Y.; Zhang, Y.; Han, J.; Luo, J.; Li, G.; Huang, J.; Wu, H.; Tian, Q.; Zhu, Q.; Chen, Y.; et al. The Intronic cis Element SE1 Recruits trans-Acting Repressor Complexes to Repress the Expression of ELONGATED UPPERMOST INTERNODE1 in Rice. Mol. Plant 2018, 11, 720–735. [Google Scholar] [CrossRef] [PubMed]
- Roscoe, T.J.; Vaissayre, V.; Paszkiewicz, G.; Clavijo, F.; Kelemen, Z.; Michaud, C.; Lepiniec, L.C.; Dubreucq, B.; Zhou, D.X.; Devic, M. Regulation of FUSCA3 Expression During Seed Development in Arabidopsis. Plant Cell Physiol. 2019, 60, 476–487. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Song, X.; Zhang, L.; Yu, Y.; Liang, Z.; Lei, Y.; Ruan, J.; Tan, B.; Liu, J.; Li, C. The transcriptional repressors VAL1 and VAL2 recruit PRC2 for genome-wide Polycomb silencing in Arabidopsis. Nucleic Acids Res. 2021, 49, 98–113. [Google Scholar] [CrossRef]
- Sasnauskas, G.; Kauneckaite, K.; Siksnys, V. Structural basis of DNA target recognition by the B3 domain of Arabidopsis epigenome reader VAL1. Nucleic Acids Res. 2018, 46, 4316–4324. [Google Scholar] [CrossRef]
- Derkacheva, M.; Steinbach, Y.; Wildhaber, T.; Mozgová, I.; Mahrez, W.; Nanni, P.; Bischof, S.; Gruissem, W.; Hennig, L. Arabidopsis MSI1 connects LHP1 to PRC2 complexes. EMBO J. 2013, 32, 2073–2085. [Google Scholar] [CrossRef]
- Kuwabara, A.; Gruissem, W. Arabidopsis RETINOBLASTOMA-RELATED and Polycomb group proteins: Cooperation during plant cell differentiation and development. J. Exp. Bot. 2014, 65, 2667–2676. [Google Scholar] [CrossRef] [PubMed]
- Del Olmo, I.; López, J.A.; Vázquez, J.; Raynaud, C.; Piñeiro, M.; Jarillo, J.A. Arabidopsis DNA polymerase ϵ recruits components of Polycomb repressor complex to mediate epigenetic gene silencing. Nucleic Acids Res. 2016, 44, 5597–5614. [Google Scholar] [CrossRef] [PubMed]
- Bloomer, R.H.; Hutchison, C.E.; Bäurle, I.; Walker, J.; Fang, X.; Perera, P.; Velanis, C.N.; Gümüs, S.; Spanos, C.; Rappsilber, J.; et al. The Arabidopsis epigenetic regulator ICU11 as an accessory protein of Polycomb Repressive Complex 2. Proc. Natl. Acad. Sci. USA 2020, 117, 16660–16666. [Google Scholar] [CrossRef] [PubMed]
- Saleh, A.; Al-Abdallat, A.; Ndamukong, I.; Alvarez-Venegas, R.; Avramova, Z. The Arabidopsis homologs of trithorax (ATX1) and enhancer of zeste (CLF) establish ’bivalent chromatin marks’ at the silent AGAMOUS locus. Nucleic Acids Res. 2007, 35, 6290–6296. [Google Scholar] [CrossRef]
- Zhou, J.X.; Liu, Z.W.; Li, Y.Q.; Li, L.; Wang, B.; Chen, S.; He, X.J. Arabidopsis PWWP domain proteins mediate H3K27 trimethylation on FLC and regulate flowering time. J. Integr. Plant Biol. 2018, 60, 362–368. [Google Scholar] [CrossRef]
- Lodha, M.; Marco, C.F.; Timmermans, M.C. The ASYMMETRIC LEAVES complex maintains repression of KNOX homeobox genes via direct recruitment of Polycomb-repressive complex2. Genes. Dev. 2013, 27, 596–601. [Google Scholar] [CrossRef]
- Yoon, J.; Cho, L.H.; Lee, S.; Pasriga, R.; Tun, W.; Yang, J.; Yoon, H.; Jeong, H.J.; Jeon, J.S.; An, G. Chromatin Interacting Factor OsVIL2 Is Required for Outgrowth of Axillary Buds in Rice. Mol. Cells 2019, 42, 858–868. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Cao, X.; Liu, Y.; Li, J.; Li, Y.; Li, D.; Zhang, K.; Gao, C.; Dong, A.; Liu, X. A chromatin loop represses WUSCHEL expression in Arabidopsis. Plant J. 2018, 94, 1083–1097. [Google Scholar] [CrossRef]
- Zhou, Y.; Tergemina, E.; Cui, H.; Förderer, A.; Hartwig, B.; Velikkakam James, G.; Schneeberger, K.; Turck, F. Ctf4-related protein recruits LHP1-PRC2 to maintain H3K27me3 levels in dividing cells in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2017, 114, 4833–4838. [Google Scholar] [CrossRef]
- Schatlowski, N.; Stahl, Y.; Hohenstatt, M.L.; Goodrich, J.; Schubert, D. The CURLY LEAF interacting protein BLISTER controls expression of polycomb-group target genes and cellular differentiation of Arabidopsis thaliana. Plant Cell 2010, 22, 2291–2305. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zheng, M.; Wang, Y.; Wang, Y.; Wang, C.; Ren, Y.; Lv, J.; Peng, C.; Wu, T.; Liu, K.; Zhao, S.; et al. DEFORMED FLORAL ORGAN1 (DFO1) regulates floral organ identity by epigenetically repressing the expression of OsMADS58 in rice (Oryza sativa). New Phytol. 2015, 206, 1476–1490. [Google Scholar] [CrossRef]
- Velanis, C.N.; Perera, P.; Thomson, B.; de Leau, E.; Liang, S.C.; Hartwig, B.; Förderer, A.; Thornton, H.; Arede, P.; Chen, J.; et al. The domesticated transposase ALP2 mediates formation of a novel Polycomb protein complex by direct interaction with MSI1, a core subunit of Polycomb Repressive Complex 2 (PRC2). PLoS Genet. 2020, 16, e1008681. [Google Scholar] [CrossRef]
- Johnston, A.J.; Matveeva, E.; Kirioukhova, O.; Grossniklaus, U.; Gruissem, W. A dynamic reciprocal RBR-PRC2 regulatory circuit controls Arabidopsis gametophyte development. Curr. Biol. 2008, 18, 1680–1686. [Google Scholar] [CrossRef] [PubMed]
- Calonje, M.; Sanchez, R.; Chen, L.; Sung, Z.R. EMBRYONIC FLOWER1 participates in polycomb group-mediated AG gene silencing in Arabidopsis. Plant Cell 2008, 20, 277–291. [Google Scholar] [CrossRef]
- Zeng, X.; Gao, Z.; Jiang, C.; Yang, Y.; Liu, R.; He, Y. HISTONE DEACETYLASE 9 Functions with Polycomb Silencing to Repress FLOWERING LOCUS C Expression. Plant Physiol. 2020, 182, 555–565. [Google Scholar] [CrossRef]
- Sun, B.; Zhou, Y.; Cai, J.; Shang, E.; Yamaguchi, N.; Xiao, J.; Looi, L.S.; Wee, W.Y.; Gao, X.; Wagner, D.; et al. Integration of Transcriptional Repression and Polycomb-Mediated Silencing of WUSCHEL in Floral Meristems. Plant Cell 2019, 31, 1488–1505. [Google Scholar] [CrossRef]
- Li, Z.; Luo, X.; Ou, Y.; Jiao, H.; Peng, L.; Fu, X.; Macho, A.P.; Liu, R.; He, Y. JASMONATE-ZIM DOMAIN proteins engage Polycomb chromatin modifiers to modulate Jasmonate signaling in Arabidopsis. Mol. Plant 2021. [Google Scholar] [CrossRef]
- Wang, J.; Hu, J.; Qian, Q.; Xue, H.W. LC2 and OsVIL2 promote rice flowering by photoperoid-induced epigenetic silencing of OsLF. Mol. Plant 2013, 6, 514–527. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Jiang, D.; Axelsson, E.; Lorković, Z.J.; Montgomery, S.; Holec, S.; Pieters, B.; Al Temimi, A.H.K.; Mecinović, J.; Berger, F. LHP1 Interacts with ATRX through Plant-Specific Domains at Specific Loci Targeted by PRC2. Mol. Plant 2018, 11, 1038–1052. [Google Scholar] [CrossRef]
- Yang, J.; Lee, S.; Hang, R.; Kim, S.R.; Lee, Y.S.; Cao, X.; Amasino, R.; An, G. OsVIL2 functions with PRC2 to induce flowering by repressing OsLFL1 in rice. Plant J. 2013, 73, 566–578. [Google Scholar] [CrossRef]
- Yoon, H.; Yang, J.; Liang, W.; Zhang, D.; An, G. OsVIL2 Regulates Spikelet Development by Controlling Regulatory Genes in Oryza sativa. Front. Plant Sci. 2018, 9, 102. [Google Scholar] [CrossRef]
- Roy, S.; Gupta, P.; Rajabhoj, M.P.; Maruthachalam, R.; Nandi, A.K. The Polycomb-Group Repressor MEDEA Attenuates Pathogen Defense. Plant Physiol. 2018, 177, 1728–1742. [Google Scholar] [CrossRef] [PubMed]
- Hohenstatt, M.L.; Mikulski, P.; Komarynets, O.; Klose, C.; Kycia, I.; Jeltsch, A.; Farrona, S.; Schubert, D. PWWP-DOMAIN INTERACTOR OF POLYCOMBS1 Interacts with Polycomb-Group Proteins and Histones and Regulates Arabidopsis Flowering and Development. Plant Cell 2018, 30, 117–133. [Google Scholar] [CrossRef]
- Liu, C.; Xi, W.; Shen, L.; Tan, C.; Yu, H. Regulation of floral patterning by flowering time genes. Dev. Cell 2009, 16, 711–722. [Google Scholar] [CrossRef] [PubMed]
- Jullien, P.E.; Mosquna, A.; Ingouff, M.; Sakata, T.; Ohad, N.; Berger, F. Retinoblastoma and its binding partner MSI1 control imprinting in Arabidopsis. PLoS Biol. 2008, 6, e194. [Google Scholar] [CrossRef] [PubMed]
- Duan, Y.; Chen, Y.; Li, W.; Pan, M.; Qu, X.; Shi, X.; Cai, Z.; Liu, H.; Zhao, F.; Kong, L.; et al. RETINOBLASTOMA-RELATED Genes Specifically Control Inner Floral Organ Morphogenesis and Pollen Development in Rice. Plant Physiol. 2019, 181, 1600–1614. [Google Scholar] [CrossRef]
- Zhang, P.; Zhu, C.; Geng, Y.; Wang, Y.; Yang, Y.; Liu, Q.; Guo, W.; Chachar, S.; Riaz, A.; Yan, S.; et al. Rice and Arabidopsis homologs of yeast CHROMOSOME TRANSMISSION FIDELITY PROTEIN 4 commonly interact with Polycomb complexes but exert divergent regulatory functions. Plant Cell 2021. [Google Scholar] [CrossRef]
- Lindner, M.; Simonini, S.; Kooiker, M.; Gagliardini, V.; Somssich, M.; Hohenstatt, M.; Simon, R.; Grossniklaus, U.; Kater, M.M. TAF13 interacts with PRC2 members and is essential for Arabidopsis seed development. Dev. Biol. 2013, 379, 28–37. [Google Scholar] [CrossRef]
- Li, Z.; Li, B.; Liu, J.; Guo, Z.; Liu, Y.; Li, Y.; Shen, W.H.; Huang, Y.; Huang, H.; Zhang, Y.; et al. Transcription factors AS1 and AS2 interact with LHP1 to repress KNOX genes in Arabidopsis. J. Integr. Plant Biol. 2016, 58, 959–970. [Google Scholar] [CrossRef]
- Berger, N.; Dubreucq, B.; Roudier, F.; Dubos, C.; Lepiniec, L. Transcriptional regulation of Arabidopsis LEAFY COTYLEDON2 involves RLE, a cis-element that regulates trimethylation of histone H3 at lysine-27. Plant Cell 2011, 23, 4065–4078. [Google Scholar] [CrossRef]
- Wu, K.; Wang, S.; Song, W.; Zhang, J.; Wang, Y.; Liu, Q.; Yu, J.; Ye, Y.; Li, S.; Chen, J.; et al. Enhanced sustainable green revolution yield via nitrogen-responsive chromatin modulation in rice. Science 2020, 367. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Luo, X.; Liu, X.; Wu, W.; Cui, X.; He, Y.; Huang, J. Arabidopsis PEAPODs function with LIKE HETEROCHROMATIN PROTEIN1 to regulate lateral organ growth. J. Integr. Plant Biol. 2020, 62, 812–831. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.Y.; Jiang, H.; Wang, G.F. PHERES1 Controls Endosperm Gene Imprinting and Seed Development. Trends Plant Sci 2020, 25, 517–519. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.H.; Xi, Y.; Sung, S. Modular function of long noncoding RNA, COLDAIR, in the vernalization response. PLoS Genet. 2017, 13, e1006939. [Google Scholar] [CrossRef]
- Wu, H.W.; Deng, S.; Xu, H.; Mao, H.Z.; Liu, J.; Niu, Q.W.; Wang, H.; Chua, N.H. A noncoding RNA transcribed from the AGAMOUS (AG) second intron binds to CURLY LEAF and represses AG expression in leaves. New Phytol. 2018, 219, 1480–1491. [Google Scholar] [CrossRef] [PubMed]
- Ariel, F.; Jegu, T.; Latrasse, D.; Romero-Barrios, N.; Christ, A.; Benhamed, M.; Crespi, M. Noncoding transcription by alternative RNA polymerases dynamically regulates an auxin-driven chromatin loop. Mol. Cell 2014, 55, 383–396. [Google Scholar] [CrossRef]
- Tian, Y.; Zheng, H.; Zhang, F.; Wang, S.; Ji, X.; Xu, C.; He, Y.; Ding, Y. PRC2 recruitment and H3K27me3 deposition at FLC require FCA binding of COOLAIR. Sci. Adv. 2019, 5, eaau7246. [Google Scholar] [CrossRef]
- Heo, J.B.; Sung, S. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science 2011, 331, 76–79. [Google Scholar] [CrossRef]
- Kim, D.H.; Sung, S. Vernalization-Triggered Intragenic Chromatin Loop Formation by Long Noncoding RNAs. Dev. Cell 2017, 40, 302–312.e304. [Google Scholar] [CrossRef]
- Molitor, A.M.; Latrasse, D.; Zytnicki, M.; Andrey, P.; Houba-Hérin, N.; Hachet, M.; Battail, C.; Del Prete, S.; Alberti, A.; Quesneville, H.; et al. The Arabidopsis hnRNP-Q Protein LIF2 and the PRC1 Subunit LHP1 Function in Concert to Regulate the Transcription of Stress-Responsive Genes. Plant Cell 2016, 28, 2197–2211. [Google Scholar] [CrossRef]
- Baulcombe, D.C.; Dean, C. Epigenetic regulation in plant responses to the environment. Cold Spring Harb Perspect Biol. 2014, 6, a019471. [Google Scholar] [CrossRef] [PubMed]
- Sani, E.; Herzyk, P.; Perrella, G.; Colot, V.; Amtmann, A. Hyperosmotic priming of Arabidopsis seedlings establishes a long-term somatic memory accompanied by specific changes of the epigenome. Genome Biol. 2013, 14, R59. [Google Scholar] [CrossRef] [PubMed]
- Liu, N.; Fromm, M.; Avramova, Z. H3K27me3 and H3K4me3 chromatin environment at super-induced dehydration stress memory genes of Arabidopsis thaliana. Mol. Plant 2014, 7, 502–513. [Google Scholar] [CrossRef] [PubMed]
- Payá-Milans, M.; Poza-Viejo, L.; Martín-Uriz, P.S.; Lara-Astiaso, D.; Wilkinson, M.D.; Crevillén, P. Genome-wide analysis of the H3K27me3 epigenome and transcriptome in Brassica rapa. Gigascience 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Shu, J.; Chen, C.; Thapa, R.K.; Bian, S.; Nguyen, V.; Yu, K.; Yuan, Z.C.; Liu, J.; Kohalmi, S.E.; Li, C.; et al. Genome-wide occupancy of histone H3K27 methyltransferases CURLY LEAF and SWINGER in Arabidopsis seedlings. Plant Direct. 2019, 3, e00100. [Google Scholar] [CrossRef]
- Kleinmanns, J.A.; Schubert, D. Polycomb and Trithorax group protein-mediated control of stress responses in plants. Biol. Chem. 2014, 395, 1291–1300. [Google Scholar] [CrossRef]
- Kleinmanns, J.A.; Schatlowski, N.; Heckmann, D.; Schubert, D. BLISTER Regulates Polycomb-Target Genes, Represses Stress-Regulated Genes and Promotes Stress Responses in Arabidopsis thaliana. Front. Plant Sci. 2017, 8, 1530. [Google Scholar] [CrossRef] [PubMed]
- Ramirez-Prado, J.S.; Latrasse, D.; Rodriguez-Granados, N.Y.; Huang, Y.; Manza-Mianza, D.; Brik-Chaouche, R.; Jaouannet, M.; Citerne, S.; Bendahmane, A.; Hirt, H.; et al. The Polycomb protein LHP1 regulates Arabidopsis thaliana stress responses through the repression of the MYC2-dependent branch of immunity. Plant J. 2019, 100, 1118–1131. [Google Scholar] [CrossRef]
- Gupta, P.; Roy, S.; Nandi, A.K. MEDEA-interacting protein LONG-CHAIN BASE KINASE 1 promotes pattern-triggered immunity in Arabidopsis thaliana. Plant Mol. Biol. 2020, 103, 173–184. [Google Scholar] [CrossRef]
- Liu, J.; Feng, L.; Gu, X.; Deng, X.; Qiu, Q.; Li, Q.; Zhang, Y.; Wang, M.; Deng, Y.; Wang, E.; et al. An H3K27me3 demethylase-HSFA2 regulatory loop orchestrates transgenerational thermomemory in Arabidopsis. Cell Res. 2019, 29, 379–390. [Google Scholar] [CrossRef]
- Nishio, H.; Buzas, D.M.; Nagano, A.J.; Suzuki, Y.; Sugano, S.; Ito, M.; Morinaga, S.; Kudoh, H. From the laboratory to the field: Assaying histone methylation at FLOWERING LOCUS C in naturally growing Arabidopsis halleri. Genes Genet. Syst. 2016, 91, 15–26. [Google Scholar] [CrossRef]
- Nishio, H.; Buzas, D.M.; Nagano, A.J.; Iwayama, K.; Ushio, M.; Kudoh, H. Repressive chromatin modification underpins the long-term expression trend of a perennial flowering gene in nature. Nat. Commun. 2020, 11, 2065. [Google Scholar] [CrossRef] [PubMed]
- Nishio, H.; Nagano, A.J.; Ito, T.; Suzuki, Y.; Kudoh, H. Seasonal plasticity and diel stability of H3K27me3 in natural fluctuating environments. Nat. Plants 2020, 6, 1091–1097. [Google Scholar] [CrossRef]
- Berry, S.; Dean, C. Environmental perception and epigenetic memory: Mechanistic insight through FLC. Plant J. 2015, 83, 133–148. [Google Scholar] [CrossRef]
- Menon, G.; Schulten, A.; Dean, C.; Howard, M. Digital paradigm for Polycomb epigenetic switching and memory. Curr. Opin. Plant Biol. 2021, 61, 102012. [Google Scholar] [CrossRef] [PubMed]
- Whittaker, C.; Dean, C. The FLC Locus: A Platform for Discoveries in Epigenetics and Adaptation. Annu. Rev. Cell Dev. Biol. 2017, 33, 555–575. [Google Scholar] [CrossRef]
- Costa, S.; Dean, C. Storing memories: The distinct phases of Polycomb-mediated silencing of Arabidopsis FLC. Biochem. Soc. Trans. 2019, 47, 1187–1196. [Google Scholar] [CrossRef]
- Zeng, Z.; Zhang, W.; Marand, A.P.; Zhu, B.; Buell, C.R.; Jiang, J. Cold stress induces enhanced chromatin accessibility and bivalent histone modifications H3K4me3 and H3K27me3 of active genes in potato. Genome Biol. 2019, 20, 123. [Google Scholar] [CrossRef]
- Carter, B.; Bishop, B.; Ho, K.K.; Huang, R.; Jia, W.; Zhang, H.; Pascuzzi, P.E.; Deal, R.B.; Ogas, J. The Chromatin Remodelers PKL and PIE1 Act in an Epigenetic Pathway That Determines H3K27me3 Homeostasis in Arabidopsis. Plant Cell 2018, 30, 1337–1352. [Google Scholar] [CrossRef] [PubMed]
- Yang, R.; Hong, Y.; Ren, Z.; Tang, K.; Zhang, H.; Zhu, J.-K.; Zhao, C. A Role for PICKLE in the Regulation of Cold and Salt Stress Tolerance in Arabidopsis. Front. Plant Sci. 2019, 10, 900. [Google Scholar] [CrossRef] [PubMed]
- Kwon, C.S.; Lee, D.; Choi, G.; Chung, W.I. Histone occupancy-dependent and -independent removal of H3K27 trimethylation at cold-responsive genes in Arabidopsis. Plant J. 2009, 60, 112–121. [Google Scholar] [CrossRef]
- Bouché, F.; Detry, N.; Périlleux, C. Heat can erase epigenetic marks of vernalization in Arabidopsis. Plant Signal. Behav. 2015, 10, e990799. [Google Scholar] [CrossRef][Green Version]
- Huh, J.H.; Bauer, M.J.; Hsieh, T.-F.; Fischer, R. Endosperm gene imprinting and seed development. Curr. Opin. Genet. Dev. 2007, 17, 480–485. [Google Scholar] [CrossRef] [PubMed]
- Folsom, J.J.; Begcy, K.; Hao, X.; Wang, D.; Walia, H. Rice fertilization-Independent Endosperm1 regulates seed size under heat stress by controlling early endosperm development. Plant Physiol. 2014, 165, 238–248. [Google Scholar] [CrossRef] [PubMed]
- Dhatt, B.K.; Paul, P.; Sandhu, J.; Hussain, W.; Irvin, L.; Zhu, F.; Adviento-Borbe, M.A.; Lorence, A.; Staswick, P.; Yu, H.; et al. Allelic variation in rice Fertilization Independent Endosperm 1 contributes to grain width under high night temperature stress. New Phytol. 2021, 229, 335–350. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.-Y.; Cheng, Y.-H.; Chen, K.-E.; Tsay, Y.-F. Nitrate Transport, Signaling, and Use Efficiency. Annu. Rev. Plant Biol. 2018, 69, 85–122. [Google Scholar] [CrossRef]
- Widiez, T.; El Kafafi, E.S.; Girin, T.; Berr, A.; Ruffel, S.; Krouk, G.; Vayssières, A.; Shen, W.-H.; Coruzzi, G.M.; Gojon, A.; et al. High nitrogen insensitive 9 (HNI9)-mediated systemic repression of root NO3- uptake is associated with changes in histone methylation. Proc. Natl. Acad. Sci. USA 2011, 108, 13329–13334. [Google Scholar] [CrossRef] [PubMed]
- Bellegarde, F.; Herbert, L.; Séré, D.; Caillieux, E.; Boucherez, J.; Fizames, C.; Roudier, F.; Gojon, A.; Martin, A. Polycomb Repressive Complex 2 attenuates the very high expression of the Arabidopsis gene NRT2.1. Sci. Rep. 2018, 8, 7905. [Google Scholar] [CrossRef] [PubMed]
- Park, E.Y.; Tsuyuki, K.M.; Hu, F.; Lee, J.; Jeong, J. PRC2-Mediated H3K27me3 Contributes to Transcriptional Regulation of FIT-Dependent Iron Deficiency Response. Front. Plant Sci. 2019, 10, 627. [Google Scholar] [CrossRef]
- Park, E.Y.; Tsuyuki, K.M.; Parsons, E.M.; Jeong, J. PRC2-mediated H3K27me3 modulates shoot iron homeostasis in Arabidopsis thaliana. Plant Signal. Behav. 2020, 15, 1784549. [Google Scholar] [CrossRef]
- Widiez, T.; Symeonidi, A.; Luo, C.; Lam, E.; Lawton, M.; Rensing, S.A. The chromatin landscape of the moss Physcomitrella patens and its dynamics during development and drought stress. Plant J. 2014, 79, 67–81. [Google Scholar] [CrossRef] [PubMed]
- Liu, N.; Ding, Y.; Fromm, M.; Avramova, Z. Different gene-specific mechanisms determine the ’revised-response’ memory transcription patterns of a subset of A. thaliana dehydration stress responding genes. Nucleic Acids Res. 2014, 42, 5556–5566. [Google Scholar] [CrossRef]
- González, R.M.; Ricardi, M.M.; Iusem, N.D. Epigenetic marks in an adaptive water stress-responsive gene in tomato roots under normal and drought conditions. Epigenetics 2013, 8, 864–872. [Google Scholar] [CrossRef]
- Alexandre, C.; Möller-Steinbach, Y.; Schönrock, N.; Gruissem, W.; Hennig, L. Arabidopsis MSI1 is required for negative regulation of the response to drought stress. Mol. Plant 2009, 2, 675–687. [Google Scholar] [CrossRef] [PubMed]
- Han, B.; Xu, W.; Ahmed, N.; Yu, A.; Wang, Z.; Liu, A. Changes and Associations of Genomic Transcription and Histone Methylation with Salt Stress in Castor Bean. Plant Cell Physiol. 2020, 61, 1120–1133. [Google Scholar] [CrossRef]
- Sun, L.; Song, G.; Guo, W.; Wang, W.; Zhao, H.; Gao, T.; Lv, Q.; Yang, X.; Xu, F.; Dong, Y.; et al. Dynamic Changes in Genome-Wide Histone3 Lysine27 Trimethylation and Gene Expression of Soybean Roots in Response to Salt Stress. Front. Plant Sci. 2019, 10, 1031. [Google Scholar] [CrossRef]
- Zheng, D.; Wang, L.; Chen, L.; Pan, X.; Lin, K.; Fang, Y.; Wang, X.E.; Zhang, W. Salt-Responsive Genes are Differentially Regulated at the Chromatin Levels Between Seedlings and Roots in Rice. Plant Cell Physiol. 2019, 60, 1790–1803. [Google Scholar] [CrossRef]
- Pu, L.; Liu, M.S.; Kim, S.Y.; Chen, L.F.; Fletcher, J.C.; Sung, Z.R. EMBRYONIC FLOWER1 and ULTRAPETALA1 Act Antagonistically on Arabidopsis Development and Stress Response. Plant Physiol. 2013, 162, 812–830. [Google Scholar] [CrossRef]
- Mehdi, S.; Derkacheva, M.; Ramström, M.; Kralemann, L.; Bergquist, J.; Hennig, L. The WD40 Domain Protein MSI1 Functions in a Histone Deacetylase Complex to Fine-Tune Abscisic Acid Signaling. Plant Cell 2016, 28, 42–54. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.K. Abiotic Stress Signaling and Responses in Plants. Cell 2016, 167, 313–324. [Google Scholar] [CrossRef]
- Chen, H.; Tong, J.; Fu, W.; Liang, Z.; Ruan, J.; Yu, Y.; Song, X.; Yuan, L.; Xiao, L.; Liu, J.; et al. The H3K27me3 Demethylase RELATIVE OF EARLY FLOWERING6 Suppresses Seed Dormancy by Inducing Abscisic Acid Catabolism. Plant Physiol. 2020, 184, 1969–1978. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Ichihashi, Y.; Suzuki, T.; Shibata, A.; Shirasu, K.; Yamaguchi, N.; Ito, T. Abscisic acid-dependent histone demethylation during postgermination growth arrest in Arabidopsis. Plant Cell Environ. 2019, 42, 2198–2214. [Google Scholar] [CrossRef]
- Liu, C.; Cheng, J.; Zhuang, Y.; Ye, L.; Li, Z.; Wang, Y.; Qi, M.; Xu, L.; Zhang, Y. Polycomb repressive complex 2 attenuates ABA-induced senescence in Arabidopsis. Plant J. 2019, 97, 368–377. [Google Scholar] [CrossRef]
- Zhu, Y.; Hu, X.; Duan, Y.; Li, S.; Wang, Y.; Rehman, A.U.; He, J.; Zhang, J.; Hua, D.; Yang, L.; et al. The Arabidopsis Nodulin Homeobox Factor AtNDX Interacts with AtRING1A/B and Negatively Regulates Abscisic Acid Signaling. Plant Cell 2020, 32, 703–721. [Google Scholar] [CrossRef] [PubMed]
- Meller, B.; Kuźnicki, D.; Arasimowicz-Jelonek, M.; Deckert, J.; Floryszak-Wieczorek, J. BABA-Primed Histone Modifications in Potato for Intergenerational Resistance to Phytophthora infestans. Front. Plant Sci. 2018, 9, 1228. [Google Scholar] [CrossRef]
- Crespo-Salvador, Ó.; Escamilla-Aguilar, M.; López-Cruz, J.; López-Rodas, G.; González-Bosch, C. Determination of histone epigenetic marks in Arabidopsis and tomato genes in the early response to Botrytis cinerea. Plant Cell Rep. 2018, 37, 153–166. [Google Scholar] [CrossRef]
- Yuan, M.; Ngou, B.P.M.; Ding, P.; Xin, X.-F. PTI-ETI crosstalk: An integrative view of plant immunity. Curr. Opin. Plant Biol. 2021, 62, 102030. [Google Scholar] [CrossRef] [PubMed]
- Saripalli, G.; Singh, K.; Gautam, T.; Kumar, S.; Raghuvanshi, S.; Prasad, P.; Jain, N.; Sharma, P.K.; Balyan, H.S.; Gupta, P.K. Genome-wide analysis of H3K4me3 and H3K27me3 modifications due to Lr28 for leaf rust resistance in bread wheat (Triticum aestivum). Plant Mol. Biol. 2020, 104, 113–136. [Google Scholar] [CrossRef]
- Atighi, M.R.; Verstraeten, B.; De Meyer, T.; Kyndt, T. Genome-wide shifts in histone modifications at early stage of rice infection with Meloidogyne graminicola. Mol. Plant Pathol. 2021, 22, 440–455. [Google Scholar] [CrossRef] [PubMed]
- Dvořák Tomaštíková, E.; Hafrén, A.; Trejo-Arellano, M.S.; Rasmussen, S.R.; Sato, H.; Santos-González, J.; Köhler, C.; Hennig, L.; Hofius, D. Polycomb Repressive Complex 2 and KRYPTONITE regulate pathogen-induced programmed cell death in Arabidopsis. Plant Physiol. 2021. [Google Scholar] [CrossRef] [PubMed]
| Drosophila | Characteristic Domains | Human | Arabidopsis | Rice | Maize |
|---|---|---|---|---|---|
| Polycomb Repressive Complex 2 (PRC2) | |||||
| E(z) | SET, CXC and SANT | EZH1, EZH2 | CLF, SWN, MEA | OsCLF, OsiEZ1 | ZmMEZ1, ZmMEZ2, ZmMEZ3 |
| Esc | WD40 | EED | FIE | OsFIE1, OsFIE2 | ZmFIE1, ZmFIE2 |
| Su(z)12 | Zinc-finger and VEFS | SUZ12 | FIS2, VRN2, EMF2 | OsEMF2a, OsEMF2b | ZmEMF2a, ZmEMF2b |
| p55 | WD40 | RBBP4, RBBP7 | MSI1–5 | OsMSI1, OsMSI3, OsMSI4 | ZmMSI1a/b/c/d; ZmMSI3; ZmMSI4a/b |
| Polycomb Repressive Complex 1 (PRC1) | |||||
| Sce | RING | RING1A, RING1B | AtRING1A/B | OsRING1A/B | ZmRING1A/B/C/D |
| Psc | RING | PCGF1–6 | AtBMI1A/B/C | OsBMI1A/B/C | ZmBMI1A/B/C/D/E/F |
| Pc | CHROMO | CBX2/4/6/7/8 | – | – | – |
| Ph | SAM and Zinc-finger | PH1–3 | – | – | – |
| CHROMO | LHP1 | OsLHP1 | ZmLHP1 | ||
| EMF1 | – | – | |||
| Trans-Acting Factor | Interacting PcG Protein | Cis-Acting Factor | Targeted Loci | Biological Function | Reference |
|---|---|---|---|---|---|
| Transcription factors | |||||
| BPC1/2/4/6 | SWN | N | ABI4 | Lateral root development | [119] |
| BPC class I subfamily | FIE | GAGA motif | N | N | [118,120] |
| BPC6 | LHP1, VRN2 | GAGA motif | Homeotic genes | Vegetative growth and flowering | [117] |
| BPC1/2/3 | FIE, MEA, FIS2, MSI1 | GAGA motif | FUSCA3 | Seed development | [122] |
| N | N | W-box, RY motif | FUSCA3 | Seed development | [131] |
| BBR/BPC | N | GAGA motif | N | Brassinosteroid signaling | [121] |
| TRB1/2/3 | CLF, SWN | Telobox | N | Vegetative growth and flowering | [123,124] |
| C2H2 ZnF family | FIE | Telobox | N | N | [120] |
| AP2 subfamily | FIE | PRE-like | N | N | [120] |
| VAL1/2 | CLF, SWN | RY repeat | N | Somatic embryonic calli | [132] |
| VAL1/2 | MSI1, LHP1 | N | FT, FLC | Flowering | [126,127] |
| VAL1/2 | CLF, MSI1, LHP1 | RY repeat | DOG1, AGL15 | Seed germination and vegetative growth | [128,129] |
| N | N | RLE | LEC2 | Embryo development | [164] |
| AS1/2 | CLF, FIE, EMF2, LHP1 | N | BP, KNAT2 | Leaf differentiation | [140,163] |
| PWO1 | CLF, SWN, MEA | N | N | Vegetative growth and flowering | [157] |
| PDP1/2/3 | MSI5 | N | FLC | Flowering | [138] |
| JAZ1/4/8/10/3/6/9 | EMF2, LHP | N | DYT1, AMS, MS1, JAZ1 | Jasmonate signaling | [151] |
| PPD1/2 | LHP1 | N | D3-type cyclins and HMGA | Lateral organ growth | [166] |
| NGR5 | LC2 | GCCGCC motif | D14, SPL14 | Tillering from nitrogen regulation | [165] |
| AG | LHP1 | Chromatin loop | WUS | Meristem maintenance and determinacy | [142] |
| KNU | FIE | N | WUS | Floral meristems determinacy | [150] |
| Di19 | MEA | N | RPS2 | Pathogen defense | [156] |
| RBR1 | FIE, MSI | N | MET1 | Female gametogenesis | [139,147,159] |
| OsRBR1/2 | OsMSI1 | N | N | Floral development | [160] |
| ESD7 | CLF, EMF2, MSI1 | N | FT, SOC1 | Vegetative growth and flowering | [134] |
| Chromatin remodelers | |||||
| BLI | CLF | N | Homeotic genes, FLC, FT | Vegetative growth and flowering | [144] |
| ICU11 | CLF, SWN, EMF2, MSI1, FIE | N | FLC | Flowering | [135] |
| ALP2 | MSI1 | N | N | Vegetative growth and flowering | [146] |
| EOL1 | CLF, SWN, LHP1 | N | N | H3K27me3 inheritance | [143] |
| OsCTF4 | OsCLF, OsLHP1, OsSWN | N | KRP1, KRP5 | Cell cycle and vegetative growth | [161] |
| ATX1 | CLF | N | AG | Vegetative development | [136] |
| TAF13 | MEA, SWN | N | PHE1, FUS3 and AtFH5 | Seed development | [162] |
| SVP | LHP1 | N | SEP3 | Floral patterning | [158] |
| ATRX | LHP1 | N | FLC | Flowering | [153] |
| PcG-associated proteins | |||||
| VIN3, VEL1, VRN5 | VRN2 | N | FLC | Flowering | [38] |
| OsVIL3 | OsVIL2 | N | OsLF | Rice flowering | [152] |
| OsVIL2 | OsEMF2b | N | OsLFL1, OsTB1 | Rice flowering | [141,154,155] |
| DFO1 | OsMSI1, OsiEZ1 | N | OsMADS58 | Floral organ identity | [145] |
| ncRNAs | |||||
| N | CLF | AG-incRNA4 | AG | Tissue specification | [169] |
| N | LHP1 | APOLO | PID | Auxin signaling | [170] |
| FCA | CLF | COOLAIR | FLC | Flowering | [171] |
| N | CLF | COLDAIR | FLC | Flowering | [168,172] |
| N | CLF | COLDWRAP | FLC | Flowering | [173] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shen, Q.; Lin, Y.; Li, Y.; Wang, G. Dynamics of H3K27me3 Modification on Plant Adaptation to Environmental Cues. Plants 2021, 10, 1165. https://doi.org/10.3390/plants10061165
Shen Q, Lin Y, Li Y, Wang G. Dynamics of H3K27me3 Modification on Plant Adaptation to Environmental Cues. Plants. 2021; 10(6):1165. https://doi.org/10.3390/plants10061165
Chicago/Turabian StyleShen, Qingwen, Yisheng Lin, Yingbo Li, and Guifeng Wang. 2021. "Dynamics of H3K27me3 Modification on Plant Adaptation to Environmental Cues" Plants 10, no. 6: 1165. https://doi.org/10.3390/plants10061165
APA StyleShen, Q., Lin, Y., Li, Y., & Wang, G. (2021). Dynamics of H3K27me3 Modification on Plant Adaptation to Environmental Cues. Plants, 10(6), 1165. https://doi.org/10.3390/plants10061165






