Bacterial Endophytes: The Hidden Actor in Plant Immune Responses against Biotic Stress
Abstract
1. Introduction
2. Diversity and Distribution of Bacterial Endophytes
3. Interaction of Bacterial Endophytes with the Host
3.1. Metabolites Implicated in the Interaction of Host–Bacterial Endophyte
3.2. Perception of Bacterial Endophytes and Modulation of Plant Immune System
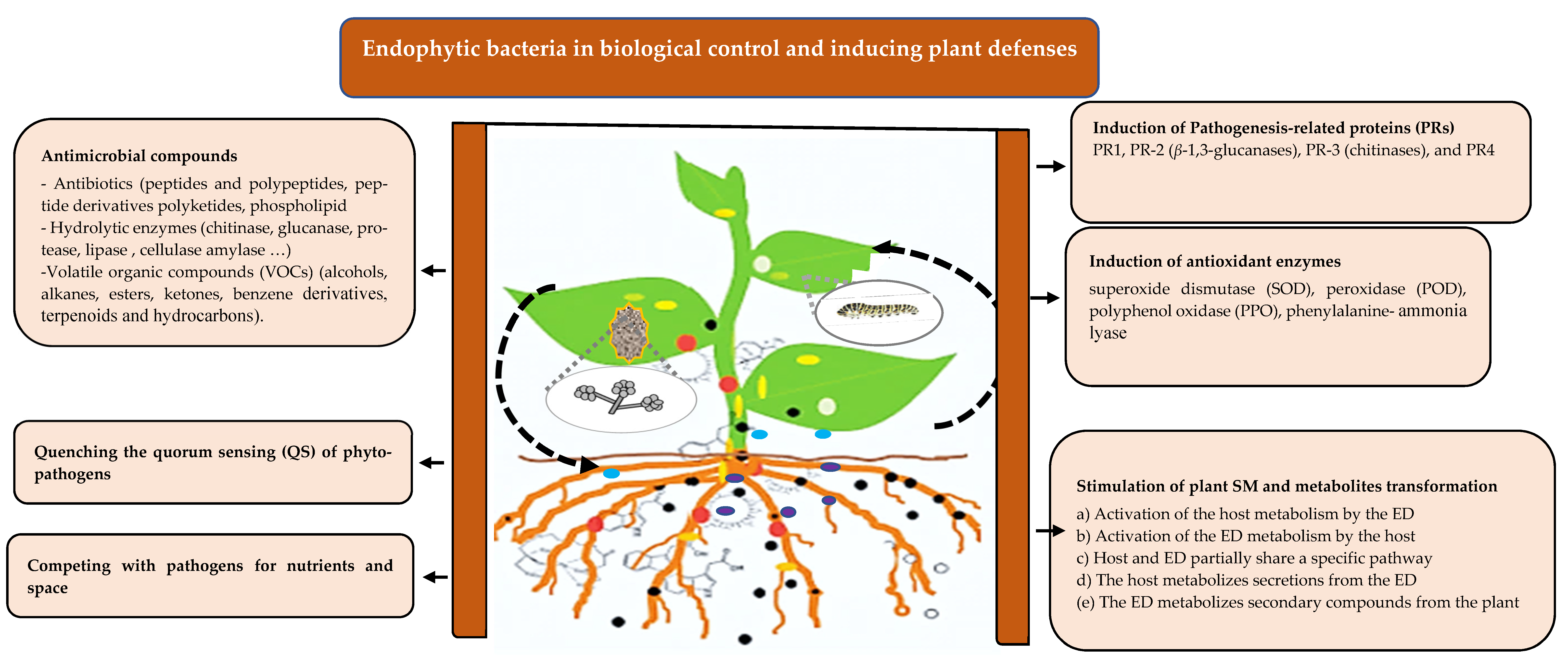
4. Extension of Plant Immunity by Endophytes
4.1. Direct Interactions in the Holobiont
4.2. Indirect Interactions in the Holobiont
4.2.1. Induction of Pathogenesis-Related Proteins (PRs) and Antioxidant Enzymes
4.2.2. Stimulation of Plant Secondary Metabolism
5. Conclusions and Perspective
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Yadav, A.N. Biodiversity and biotechnological applications of host specific endophytic fungi for sustainable agriculture and allied sectors. Acta Sci. Microbiol. 2018, 1, 1–5. [Google Scholar] [CrossRef]
- Santoyo, G.; Moreno-Hagelsiebb, G.; Orozco-Mosquedac, M.C.; Glick, B.R. Plant growth-promoting bacterial endophytes. Microbiol. Res. 2016, 183, 92–99. [Google Scholar] [CrossRef]
- Sessitsch, A.; Hardoim, P.; Döring, J.; Weilharter, A.; Krause, A.; Woyke, T.; Mitter, B.; Hauberg-Lotte, L.; Friedrich, F.; Rahalkar, M.; et al. Functional characteristics of an endophyte community colonizing rice roots as revealed by metagenomic analysis. Mol. Plant Microbe Interact. 2012, 25, 28–36. [Google Scholar] [CrossRef]
- Akinsanya, M.A.; Goha, J.K.; Lima, S.P.; Ting, A.S.Y. Metagenomics study of endophytic bacteria in Aloe vera using next-generation technology. Genom. Data 2015, 6, 159–163. [Google Scholar] [CrossRef]
- Puri, R.R.; Adachi, F.; Omichi, M.; Saeki, Y.; Yamamoto, A.; Hayashi, S.; Ali, M.A.; Itoh, K. Metagenomic study of endophytic bacterial community of sweet potato (Ipomoea batatas) cultivated in different soil and climatic conditions. World J. Microbiol. Biotechnol. 2019, 35, 176. [Google Scholar] [CrossRef]
- Ali, S.; Charles, T.C.; Glick, B.R. Delay of flower senescence by bacterial endophytes expressing 1-aminocyclopropane-1-carboxylate deaminase. J. Appl. Microbiol. 2012, 113, 1139–1144. [Google Scholar] [CrossRef] [PubMed]
- Coutinho, B.G.; Licastro, D.; Mendonc-Previato, L.; Cámara, M.; Venturi, V. Plant-influenced gene expression in the rice endophyte Burkholderia kururiensis M130. Mol. Plant-Microbe Interact. 2015, 28, 10–21. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Carvalhais, L.C.; Crawford, M.; Singh, E.; Dennis, P.G.; Pieterse, C.M.J.; Schenk, P.M. Inner plant values: Diversity, colonization and benefits from endophytic bacteria. Front. Microbiol. 2017, 8, 2552. [Google Scholar] [CrossRef]
- Lata, R.; Chowdhury, S.; Gond, S.K.; White, J.F.J. Induction of abiotic stress tolerance in plants by endophytic microbes. Lett. Appl. Microbiol. 2018, 66, 268–276. [Google Scholar] [CrossRef] [PubMed]
- Latha, P.; Karthikeyan, M.; Rajeswari, E. Endophytic Bacteria: Prospects and Applications for the Plant Disease Management. Plant Health Biotic Stress India 2019, 50. [Google Scholar] [CrossRef]
- Hardoim, P.R.; van Overbeek, L.S.; van Elsas, J.D. Properties of bacterial endophytes and their proposed role in plant growth. Trends Microbiol. 2008, 16, 463–471. [Google Scholar] [CrossRef] [PubMed]
- Rashid, S.; Charles, T.C.; Glick, B.R. Isolation and characterization of new plant growth-promoting bacterial endophytes. Appl. Soil Ecol. 2012, 61, 217–224. [Google Scholar] [CrossRef]
- Gary, S.; Bryn, D. Bioprospecting for microbial endophytes and their natural products. Microbiol. Mol. Biol. Rev. 2003, 67, 491–502. [Google Scholar] [CrossRef]
- Brader, G.; Company, S.; Vescio, K.; Mitter, B.; Trognitz, F.; Ma, L.J.; Sessitsch, A. Ecology and genome insights into plant-pathogenic and plant-nonpathogenic endophytes. Annu. Rev. Phytopathol. 2017, 55, 61–83. [Google Scholar] [CrossRef] [PubMed]
- Afzal, I.; Shinwari, Z.K.; Sikandar, S.; Shahzad, S. Plant beneficial endophytic bacteria: Mechanisms, diversity, host range and genetic determinants. Microbiol. Res. 2019, 221, 36–49. [Google Scholar] [CrossRef] [PubMed]
- Sturz, A.V.; Matheson, B.G. Populations of endophyti bacteria which influence host-resistance to Erwinia-induced bacterial soft rotin potato tubers. Plant Soil. 1996, 184, 265–271. [Google Scholar] [CrossRef]
- Pieterse, C.M.J.; Zamioudis, C.; Berendsen, R.L.; Weller, D.M.; Van Wees, S.C.M.; Bakker, P.A.H.M. Induced Systemic Resistance by Beneficial Microbes. Annu. Rev. Phytopathol. 2014, 52, 347–375. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Hidalgo, P.; García, J.M.; Pozo, M.J. Induced systemic resistance against Botrytis cinerea by Microspora strains isolated from root nodules. Front. Microbiol. 2015, 6, 922. [Google Scholar] [CrossRef]
- Oukala, N.; Pastor-Fernandez, J.; Sanmartín, N.; Aissat, K.; Pastor, V. Endophytic Bacteria from the Sahara Desert Protect Tomato Plants Against Botrytis cinerea Under Different Experimental Conditions. Curr. Microbiol. 2021. [Google Scholar] [CrossRef]
- Azevedo, J.L.; Maccheroni, J.W.; Pereira, J.O.; Araújo, W.L. Endophytic microorganisms: A review on insect control and recent advances on tropical plants. Electron. J. Biotechnol. 2000, 3, 40–65. [Google Scholar] [CrossRef]
- Ali, S.; Charles, T.C.; Glick, B.R. Amelioration of high salinity stress damage by plant growth-promoting bacterial endophytes that contain ACC deaminase. Plant Physiol. Biochem. 2014, 80, 160–167. [Google Scholar] [CrossRef]
- Subramanian, P.; Mageswari, A.; Kim, K.; Lee, Y.; Sa, T. Psychrotolerant endophytic Pseudomonas sp. strains OB155 and OS261 induced chilling resistance in tomato plants (Solanum lycopersicum Mill.) by activation of their antioxidant capacity. Mol. Plant Microbe Interact. 2015, 28, 1073–1081. [Google Scholar] [CrossRef]
- Rajkumar, M.; Ae, N.; Freitas, H. Endophytic bacteria and their potential to enhance heavy metal phytoextraction. Chemosphere 2009, 77, 153–160. [Google Scholar] [CrossRef]
- Siciliano, S.D.; Fortin, N.; Mihoc, A.; Wisse, G.; Labelle, S.; Beaumier, D. Selection of specific endophytic bacterial genotypes by plants in response to soil contamination. J. Appl. Environ. Microbiol. 2001, 67, 2469–2475. [Google Scholar] [CrossRef]
- Dini-Andreote, F. Endophytes: The second layer of plant defense. Trends Plant Sci. 2020, 25, 319–322. [Google Scholar] [CrossRef]
- Kandel, S.; Joubert, P.; Doty, S. Bacterial endophyte colonization and distribution within plants. Microorganisms 2017, 5, 77. [Google Scholar] [CrossRef]
- Ali, S.; Duan, J.; Charles, T.C.; Glick, B.R. A bioinformatics approach to the determination of genes involved in endophytic behavior in Burkholderia spp. J. Theor. Biol. 2014, 343, 193–198. [Google Scholar] [CrossRef]
- Khare, E.; Mishra, J.; Arora, N. Multifaceted Interactions between Endophytes and Plant: Developments and Prospects. Front. Microbiol. 2018, 9, 2732. [Google Scholar] [CrossRef] [PubMed]
- Mercado-Blanco, J.; Lugtenberg, B. Biotechnological Applications of Bacterial Endophytes. Curr. Biotechnol. 2014, 3, 60–75. [Google Scholar] [CrossRef]
- Martínez-Medina, A.; Flors, V.; Heil, M.; Mauch-Mani, B.; Pieterse, C.M.J.; Pozo, M.J.; Ton, J.; van Dam, N.M.; Conrath, U. Recognizing Plant Defense Priming. Trends Plant Sci. 2016, 21, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Bulgarelli, D.; Schlaeppi, K.; Spaepen, S.; Ver Loren van Themaat, E.; Schulze-Lefert, P. Structure and functions of the bacterial microbiota of plants. Annu. Rev. Plant Biol. 2013, 64, 807–838. [Google Scholar] [CrossRef]
- Bulgarelli, D.; Rott, M.; Schlaeppi, K.; Ver Loren van Themaat, E.; Ahmadinejad, N.; Assenza, F.; Rauf, P.; Huettel, B.; Reinhardt, R.; Schmelzer, E.; et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 2012, 488, 91–95. [Google Scholar] [CrossRef]
- Mina, D.; Pereira, J.A.; Lino-Neto, T.; Baptista, P. Epiphytic and endophytic bacteria on olive tree phyllosphere: Exploring tissue and cultivar effect. Microb. Ecol. 2020, 80, 145–157. [Google Scholar] [CrossRef]
- Remus-Emsermann, M.N.P.; Lücker, S.; Müller, D.B.; Pottchoff, E.; Daims, H.; Vorholt, J.A. Spatial distribution analyses of natuiral phyllosphere-colonizing bacteria on Arabidopsis thaliana revealed by fluorescence in situ hybridization. Environ. Microbiol. 2014, 16, 2329–2340. [Google Scholar] [CrossRef]
- Lighthart, B. Mini-review of the concentration various found in the alfresco atmospheric bacterial populations. Aerobiologia 2000, 16, 7–16. [Google Scholar] [CrossRef]
- Hallmann, J.; Quadt-Hallmann, A.; Mahaffee, W.F.; Kloepper, J.W. Bacterial endophytes in agricultural crops. Can. J. Microbiol. 1997, 43, 895–914. [Google Scholar] [CrossRef]
- Sturz, A.V.; Nowak, J. Endophytic communities of rhizobacteria and the strategies required to create yield enhancing associations with crops. Appl. Soil Ecol. 2000, 15, 183–190. [Google Scholar] [CrossRef]
- Delmotte, N.; Knief, C.; Chaffron, S.; Innerebner, G.; Roschitzki, B.; Sclapbach, R.; von Mering, C.; Vorholt, J.A. Community proteogenomics reveals insights into the physiology of phyllosphere bacteria. Proc. Natl. Acad. Sci. USA 2009, 106, 16428–16433. [Google Scholar] [CrossRef] [PubMed]
- Frank, A.C.; Saldierma Guzmán, J.P.; Shay, J.E. Transmission of bacterial endophytes. Microorganisms 2017, 5, 70. [Google Scholar] [CrossRef] [PubMed]
- Dong, C.J.; Wang, L.L.; Li, Q.; Shang, Q.M. Bacterial communities in the rhizosphere, phyllosphere and endosphere of tomato plants. PLoS ONE 2019, 14, e223847. [Google Scholar] [CrossRef] [PubMed]
- Massoni, J.; Bortfeld-Miller, M.; Jardillier, L.; Salazar, G.; Sunagawa, S.; Vorholt, J.A. Consistent host and organ occupancy of phyllosphere bacteria in a community of wild herbaceous plant species. ISME J. 2020, 14, 245–258. [Google Scholar] [CrossRef] [PubMed]
- Brewer, T.; Handley, K.; Carini, P.; Gibert, J.; Fierer, N. Genome reduction in an abundant and ubiquitous soil bacterial lineage. Nat. Microbiol. 2016, 2, 16198. [Google Scholar] [CrossRef]
- Hottes, A.K.; Freddolino, P.L.; Khare, A.; Donnell, Z.N.; Liu, J.C.; Tavazoie, S. Bacterial adaptation through loss of function. PLoS Genet. 2013, 9, 1003617. [Google Scholar] [CrossRef] [PubMed]
- Song, H.; Hwang, J.; Yi, H.; Ulrich, R.L.; Yu, Y.; Nierman, W.C.; Kim, H.S. The early stage of bacterial genome-reductive evolution in the host. PLoS Pathog. 2010, 6, e1000922. [Google Scholar] [CrossRef] [PubMed]
- Pinski, A.; Betekhtin, A.; Hupert-Kocurek, K.; Mur, L.A.J.; Hasterok, R. Defining the genetic basis of plant-endophytic bacteria interactions. Int. J. Mol. Sci. 2019, 20, 1947. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Droby, S.; Singh, V.K.; Singh, S.K.; White, J.F. Entry, colonization and distribution of endophytic microorganisms in plants in Microbial Endophytes. Funct. Plant Biol. 2019. [Google Scholar] [CrossRef]
- Straub, D.; Rothballer, M.; Hartmann, A.; Ludewig, U. The genome of the endophytic bacterium H. frisingense GSF30(T) identifies diverse strategies in the Herbaspirillum genus to interact with plants. Front. Microbiol. 2013, 4, 168. [Google Scholar] [CrossRef] [PubMed]
- Sessitsch, A.; Kuffner, M.; Kidd, P.; Vangronsveld, J.; Wenzel, W.W.; Fallmann, K.; Puschenreiter, M. The role of plant-associated bacteria in the mobilization and phytoextraction of trace elements in contaminated soils. Soil Biol. Biochem. 2013, 60, 182–194. [Google Scholar] [CrossRef]
- Reinhold-Hurek, B.; Maes, T.; Gemmer, S.; Van Montagu, M.; Hurek, T. An endoglucanase is involved in infection of rice roots by the not-cellulose metabolizing endophyte Azoarcus sp. strain BH72. Mol. Plant Microbe Interact. 2006, 19, 181–188. [Google Scholar] [CrossRef] [PubMed]
- Piromyou, P.; Songwattana, P.; Greetatorn, T.; Okubo, T.; Kakizaki, K.C.; Prakamhang, J.; Tittabutr, P.; Boonkerd, N.; Teaumroong, N.; Minamisawa, K. The Type III secretion system (T3SS) is a determinant for rice-endophyte colonization by non-photosynthetic Bradyrhizobium. Microbes Environ. 2015, 30, 291–300. [Google Scholar] [CrossRef]
- Vorholt, J.A. Microbial life in the phyllosphere. Nat. Rev. Microbiol. 2012, 10, 828–840. [Google Scholar] [CrossRef]
- Gupta, R.; Anand, G.; Gaur, R.; Yadav, D. Plant–microbiome interactions for sustainable agriculture: A review. Physiol. Mol. Biol. Plants 2021, 27, 165–179. [Google Scholar] [CrossRef]
- Verma, P.; Yadav, A.N.; Kumar, V.; Singh, D.P.; Sexena, A.K. Benefecial plant microbes interactions: Biodiversity of microbes from diverse extreme environment and its impact for crop improvement. In Plant Microbe Interactions in Agroecological Perspectives; Singh, D.P., Ed.; Springer: Singapore, 2017; pp. 543–580. [Google Scholar]
- Pouget, M. Les Relations Sol-Végétation Dans la Steppe Sud-Algéroise; Orstom: Paris, France, 1980; p. 569. [Google Scholar]
- Goudjal, Y.; Toumatia, O.; Yekkoura, A.; Sabaou, N.; Barakate, M.; Mathieu, F.; Zitouni, A. Biocontrol of Rhizoctonia solani damping-off and promotion of tomato plant growth by endophytic actinomycetes isolated from native plants of Algerian Sahara. Microbiol. Res. 2014, 169, 59–65. [Google Scholar] [CrossRef]
- Rodriguez-Blanco, A.; Sicardi, M.; Frioni, L. Plant genotype and nitrogen fertilization effects on abundance and diversity of diazotrophic bacteria associated with maize (Zea mays L.). Biol. Fertil. Soils. 2015, 51, 391–402. [Google Scholar] [CrossRef]
- Ding, T.; Melcher, U. Influences of plant species, season and location on leaf endophytic bacterial communities of non-cultivated plants. PLoS ONE 2016, 11, e0150895. [Google Scholar] [CrossRef]
- Bacon, C.W.; Glenn, A.E.; Yates, I.E. Fusarium verticillioides: Managing the endophytic association with maize for reduced fumonisins accumulation. Toxin Rev. 2008, 27, 411–446. [Google Scholar] [CrossRef]
- Walters, D.R.; Havis, N.D.; Oxley, S.J.P. Ramularia collo-cygni: The biology of an emerging pathogen of barley. FEMS Microbiol. Lett. 2008, 279, 1–7. [Google Scholar] [CrossRef]
- Badri, D.V.; Vivanco, J.M. Regulation and function of root exudates. Plant Cell Environ. 2009, 32, 666–681. [Google Scholar] [CrossRef] [PubMed]
- Mark, G.L.; Dow, J.M.; Kiely, P.D.; Higgins, H.; Haynes, J.; Baysse, C.; Abbas, A.; Foley, T.; Franks, A.; Morrissey, J.; et al. Transcriptome profiling of bacterial responses to root exudates identifies genes involved in microbe-plant interactions. Proc. Natl. Acad. Sci. USA 2005, 102, 17454–17459. [Google Scholar] [CrossRef]
- Yi, Y.; Jong, A.; Frenzel, E.; Kuipers, O.P. Comparative Transcriptomics of Bacillus mycoides Strains in Response to Potato-Root Exudates Reveals Different Genetic Adaptation of Endophytic and Soil Isolates. Front. Microbiol. 2017, 8, 1487. [Google Scholar] [CrossRef] [PubMed]
- Arora, N.K.; Mishra, J. Prospecting the roles of metabolites and additives in future bioformulations for sustainable agriculture. Appl. Soil Ecol. 2016, 107, 405–407. [Google Scholar] [CrossRef]
- Balachandar, D.; Sandhiya, G.S.; Sugitha, T.C.K.; Kumar, K. Flavonoids and growth hormones influence endophytic colonization and in planta nitrogen fixation by a diazotrophic Serratia sp. in rice. World J. Mic. Biotech. 2006, 22, 707–712. [Google Scholar] [CrossRef]
- Zhang, N.; Wang, D.; Liu, Y. Effects of different plant root exudates and their organic acid components on chemotaxis, biofilm formation and colonization by beneficial rhizosphere-associated bacterial strains. Plant Soil 2014, 374, 689–700. [Google Scholar] [CrossRef]
- Kost, T.; Stopnisek, N.; Agnoli, K.; Eberl, L.; Weisskopf, L. Oxalotrophy, a widespread trait of plant-associated Burkholderia species, is involved in successful root colonization of lupin and maize by Burkholderia phytofirmans. Front. Microbiol. 2014, 4, 421. [Google Scholar] [CrossRef]
- Dubey, A.; Malla, M.A.; Kumar, A.; Dayanandan, S.; Khan, M.L. Plant endophytes: Unveiling hidden agenda for bioprospecting toward sustainable agricultur. Crit. Rev. Biotechnol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Trdá, L.; Fernandez, O.; Boutrot, F.; Héloir, M.C.; Kelloniemi, J.; Daire, X.; Adrian, M.; Clément, C.; Zipfel, C.; Dorey, S.; et al. The grapevine flagellin receptor VvFLS2 differentially recognizes flagellin-derived epitopes from the endophytic growth-promoting bacterium Burkholderia phytofirmans and plant pathogenic bacteria. New Phytol. 2014, 201, 1371–1384. [Google Scholar] [CrossRef]
- Deng, Y.; Chen, H.; Li, C.; Xu, J.; Qi, Q.; Xu, Y.; Zhu, Y.; Zheng, J.; Peng, D.; Ruan, L.; et al. Endophyte Bacillus subtilis evade plant defense by producing lantibiotic subtilomycin to mask self-produced flagellin. Commun. Biol. 2019, 2, 368. [Google Scholar] [CrossRef]
- Alquéres, S.; Meneses, C.; Rouws, L.; Rothballer, M.; Baldani, I.; Schmid, M.; Hartmann, A. The bacterial superoxide dismutase and glutathione reductase are crucial for endophytic colonization of rice roots by Gluconacetobacter diazotrophicus PAL5. Mol. Plant Microbe Interact. 2013, 26, 937–945. [Google Scholar] [CrossRef] [PubMed]
- Teixeira, P.J.P.L.; Colaianni, N.R.; Fitzpatrick, C.R.; Dangl, J.L. Beyond pathogens: Microbiota interactions with the plant immune system. Curr. Opin. Microb. 2019, 49, 7–17. [Google Scholar] [CrossRef]
- Plett, J.M.; Martin, F.M. Know your enemy, embrace your friend: Using omics to understand how plants respond differently to pathogenic and mutualistic microorganisms. Plant J. 2018, 93, 729–746. [Google Scholar] [CrossRef]
- Timm, C.M.; Campbell, A.G.; Utturkar, S.M.; Jun, S.R.; Parales, R.E.; Tan, W.A.; Robeson, M.S.; Lu, T.Y.; Jawdy, S.; Brown, S.D.; et al. Metabolic functions of Pseudomonas fluorescens strains from Populus deltoides depend on rhizosphere or endosphere isolation compartment. Front. Microbiol. 2015, 6, 1118. [Google Scholar] [CrossRef] [PubMed]
- Formey, D.; Sallet, E.; Lelandais-Briere, C.; Ben, C.; Bustos-Sanmamed, P.; Niebel, A.; Frugier, F.J.; Combier, P.; Debellé, F.; Hartmann, C.; et al. The small RNA diversity from Medicago truncatula roots under biotic interactions evidences the environmental plasticity of the miRNAome. Genome Biol. 2014, 15, 457. [Google Scholar] [CrossRef] [PubMed]
- Wu, P.; Wu, Y.; Liu, C.C.; Liu, L.W.; Ma, F.F.; Wu, X.Y.; Wu, M.; Hang, Y.Y.; Chen, J.Q.; Shao, Z.Q.; et al. Identification of arbuscular mycorrhiza (AM)-responsive microRNAs in tomato. Front. Plant Sci. 2016, 7, 429. [Google Scholar] [CrossRef]
- Kusajima, M.; Shima, S.; Fujita, M.; Minamisawa, K.; Che, F.-S.; Yamakawa, H.; Nakashita, H. Involvement of ethylene signaling in Azospirillum sp. B510-induced disease resistance in rice. Biosci. Biotechnol. Biochem. 2018, 82, 1522–1526. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhang, Q.; Zhang, J.; Wu, L.; Qi, Y.; Zhou, J.M. Identification of microRNAs involved in pathogen associated molecular pattern-triggered plant innate immunity. Plant Physiol. 2010, 152, 2222–2231. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, L.; Zou, Y.; Chen, L.; Cai, Z.; Zhang, S.; Zhao, F.; Tian, Y.; Jiang, Q.; Ferguson, B.J.P.; et al. Soybean miR172c targets the repressive AP2 transcription factor NNC1 to activate ENOD40 expression and regulate nodule initiation. Plant Cell 2014, 6, 4782–4801. [Google Scholar] [CrossRef]
- Holt, D.B.; Gupta, V.; Meyer, D.; Abel, N.B.; Andersen, S.U.; Stougaard, J.; Markmann, K. micro RNA 172 (miR172) signals epidermal infection and is expressed in cells primed for bacterial invasion in Lotus japonicus roots and nodules. New Phytol. 2015, 208, 241–256. [Google Scholar] [CrossRef]
- Fitzpatrick, C.R.; Salas-González, I.; Conway, J.M.; Finkel, O.M.; Gilbert, S.; Russ, D.; Teixeira, O.M.P.L.; Dangl, J.L. The plant microbiome: From ecology to reductionism and beyond. Annu. Rev. Microbiol. 2020, 74, 81–100. [Google Scholar] [CrossRef]
- Gunatilaka, A.L. Natural products from plant-associated microorganisms: Distribution, structural diversity, bioactivity, and implications of their occurrence. J. Nat. Prod. 2006, 69, 509–526. [Google Scholar] [CrossRef]
- Yu, H.; Zhang, L.; Li, L.; Zheng, C.; Guo, L.; Li, W.; Sun, P.; Qin, L. Recent developments and future prospects of antimicrobial metabolites produced by endophytes. Microbiol Res. 2010, 165, 437–449. [Google Scholar] [CrossRef]
- Brader, G.; Compant, S.; Mitter, B.; Trognitz, F.; Sessitsch, A. Metabolic potential of endophytic bacteria. Curr. Opin. Biotechnol. 2014, 27, 30–37. [Google Scholar] [CrossRef] [PubMed]
- Chung, J.H.; Song, G.C.; Ryu, C.M. Sweet scents from good bacteria: Case studies on bacterial volatile compounds for plant growth and immunity. Plant Mol. Biol. 2016, 90, 677–687. [Google Scholar] [CrossRef] [PubMed]
- Ek-Ramos, M.J.; Gomez-Flores, R.; Orozco-Flores, A.A.; Rodríguez-Padilla, C.; González-Ochoa, G.; Tamez-Guerra, P. Bioactive Products from Plant-Endophytic Gram-Positive Bacteria. Front. Microbiol. 2019, 10, 463. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.B.; Bassler, B.L. Quorum sensing in bacteria. Annu. Rev. Microbiol. 2001, 55, 165–199. [Google Scholar] [CrossRef]
- Villarreal-Delgado, M.F.; Villa-Rodríguez, E.D.; Cira-Chávez, L.A.; Estrada-Alvarado, M.I.; Parra-Cota, F.I.; de los Santos-Villalobos, S. The genus bacillus as a biological control agent and its implications in the agricultural biosecurity. Mex. J. Phytopathol. 2018, 6, 95–130. [Google Scholar] [CrossRef]
- Ongena, M.; Jacques, P. Bacillus lipopeptides: Versatile weapons for plant disease biocontrol. Trends Microbiol. 2008, 16, 115–125. [Google Scholar] [CrossRef]
- Miller, R.V.; Miller, C.M.; Kinney, D.G.; Redgrave, B.; Sears, J.; Condron, M.; Strobel, T. Ecomycins, unique antimycotics from Pseudomonas viridiflava. J. Appl. Microbiol. 1998, 84, 937–944. [Google Scholar] [CrossRef]
- Stein, T. Bacillus subtilis antibiotics: Structures, syntheses and specific functions. Mol. Microbiol. 2005, 56, 845–857. [Google Scholar] [CrossRef]
- Grady, E.N.; MacDonald, J.; Liu, L.; Richman, A.; Yuan, Z.C. Current knowledge and perspectives of Paenibacillus. Microb. Cell Fact. 2016, 15, 203. [Google Scholar] [CrossRef]
- Fadiji, A.E.; Babalola, O.O. Elucidating Mechanisms of Endophytes Used in Plant Protection and Other Bioactivities with multifunctional prospects. Front. BioEng. BioTechnol. 2020, 8, 467. [Google Scholar] [CrossRef]
- Haggag, W.M.; Abdallh, E.G. Purification and Characterization of Chitinase Produced by Endophytic Streptomyces hygroscopicus Against Some Phytopathogens. Microbiol. Res. 2012, 2, 145–151. [Google Scholar] [CrossRef]
- Pleban, S.; Chernin, L.; Chet, I. Chitinolytic activity of an endophytic strain of Bacillus cereus. Lett. Appl. Microbiol. 1997, 25, 284–288. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Wang, Z.; Qiao, X.; Li, Z.; Li, F.; Chen, M.; Wang, Y.; Huang, Y.; Cui, H. Antifungal activity of volatile organic compounds from Streptomyces alboflavus TD-1. FEMS Microbiol. Lett. 2013, 341, 45–51. [Google Scholar] [CrossRef]
- Sheoran, N.; Valiya Nadakkakath, A.; Munjal, V.; Kundu, A.; Subaharan, K.; Venugopal, V.; Rajamma, S.; Eapen, S.J.; Kumar, A. Genetic analysis of plant endophytic Pseudomonas putida BP25 and chemo-profiling of its antimicrobial volatile organic compounds. Microbiol. Res. 2015, 173, 66–78. [Google Scholar] [CrossRef]
- Fernando, W.G.D.; Ramarathnam, R.; Krishnamoorthy, A.S.; Savchuk, S.C. Identification and use of potential bacterial organic antifungal volatiles in biocontrol. Soil Biol. Biochem. 2005, 37, 955–964. [Google Scholar] [CrossRef]
- Li, Y.; Mao, Z.C.; Wu, Y.X.; Ho, H.H.; He, Y.Q. Comprehensive volatile organic compounds profiling of Bacillus species with biocontrol properties by head space solid phase microextraction with gas chromatography-mass spectrometry. Biocontrol. Sci. Techn. 2014, 25, 132–143. [Google Scholar] [CrossRef]
- Munjal, V.; Nadakkakath, A.V.; Sheoran, N.; Kundu, A.; Venugopal, V.; Subaharan, K.; Rajamma, S.; Eapen, S.J.; Kumar, A. Genotyping and identification of broad spectrum antimicrobial volatiles in black pepper root endophytic biocontrol agent, Bacillus megaterium BP17. Biol. Control 2016, 92, 66–76. [Google Scholar] [CrossRef]
- Van Loon, l.C.; Geraats, B.P.J.; Linthrost, H.J.M. Ethylen as a modulator of disease resistance in plants. Trends Plant Sci. 2006, 11, 184–191. [Google Scholar] [CrossRef]
- Gamalero, E.; Glick, B. Bacterial modulation of plant ethylen levels. Plant Physiol. 2015, 169, 13–22. [Google Scholar] [CrossRef]
- Hosni, T.; Moretti, C.; Devescovi, G.; Suarez-Moreno, Z.R.; Fatmi, M.B.; Guarnaccia, C.; Pongor, S.; Onofri, A.; Buonaurio, R.; Venturi, V. Sharing of quorum-sensing signals and role of interspecies communities in a bacterial plant disease. ISME J. 2011, 5, 1857–1870. [Google Scholar] [CrossRef]
- Kusari, P.; Kusari, S.; Lamshöft, M.; Sezgin, S.; Spiteller, M.; Kayser, O. Quorum quenching is an antivirulence strategy employed by endophytic bacteria. Appl. Microbiol. Biotechnol. 2014, 98, 7173–7183. [Google Scholar] [CrossRef] [PubMed]
- Achari, G.A.; Ramesh, R. Characterization of bacteria degrading 3-hydroxy palmitic acid methyl ester (3OH-PAME), a quorum sensing molecule of Ralstonia solanacearum. Lett. Appl. Microbiol. 2014, 60, 447–455. [Google Scholar] [CrossRef]
- Tamehiro, N.; Okamot-Hosova, Y.; Okamoto, S.; Ubukata, M.; Hamada, M.; Naganawa, H.; Ochi, K. Bacilysocin, a novel phospholipid antibiotic produced by Bacillus subtilis 168. Antimicrob. Agents Chemother. 2002, 46, 315–320. [Google Scholar] [CrossRef]
- Espinasse, S.; Gohar, M.; Lereclus, D.; Sanchis, V. An ABC transporter from Bacillus thuringiensis is essential for betaexotoxin I production. J. Bacteriol. 2002, 184, 5848–5854. [Google Scholar] [CrossRef]
- Castillo, U.F.; Strobel, G.A.; Ford, E.J.; Hess, W.M.; Porter, H.; Jensen, J.B.; Albert, H.; Robison, R.; Condron, M.A.M.; Teplow, D.B.; et al. Munumbicins, wide-spectrum antibiotics produced by Streptomyces NRRL30562, endophytic on Kennedia nigriscans. Microbiology 2002, 148, 2675–2685. [Google Scholar] [CrossRef]
- Castillo, U.; Harper, J.K.; Strobel, G.A.; Sears, J.; Alesi, K.; Ford, E.; Hess, W.M.; Porter, H.; Jensen, J.B.; Albert, H.; et al. Kakadumycins, novel antibiotics from Streptomyces sp. NRRL 30566, an endophyte of Grevillea pteridifolia. FEMS Microbiol. Lett. 2003, 224, 183–190. [Google Scholar] [CrossRef]
- Fiedler, H.P.; Bruntner, C.; Riedlinger, J.; Bull, A.T.; Knutsen, G.; Goodfellow, M.; Jones, A.; Maldonado, L.; Pathom-aree, W.; Beil, W.; et al. Proximicin A, B and C, novel aminofuran antibiotic and anticancer compounds isolated from marine strains of the actinomycete Verrucosispora. J. Antibiot. 2008, 61, 158–163. [Google Scholar] [CrossRef] [PubMed]
- Ding, L.; Maier, A.; Fiebig, H.H.; Lin, W.H.; Hertweck, C. A family of multicyclic indolsesquiterpenes from a bacterial endophyte. Organ. Biomol. Chem. 2011, 9, 4029–4031. [Google Scholar] [CrossRef] [PubMed]
- Bae, M.; Chung, B.; Oh, K.B.; Shin, J.; Oh, D.C. Hormaomycins B and C: New antibiotic cyclic depsipeptides from a marine mud flat derived Streptomyces sp. Mar. Drugs 2015, 13, 5187–5200. [Google Scholar] [CrossRef] [PubMed]
- Singh, M.; Kumar, A.; Singh, R.; Pandey, K.D. Endophytic bacteria: A new source of bioactive compounds. Biotech 2017, 7, 315. [Google Scholar] [CrossRef]
- Mohamad, A.; Abdalla, O.; Li, L.; Ma, J.; Hatab, S.R.; Xu, L.; Guo, J.W.; Rasulov, B.A.; Liu, Y.H.; Hedlund, B.P.; et al. Evaluation of the antimicrobial activity of endophytic bacterial populations from Chinese traditional medicinal plant licorice and characterization of the bioactive secondary metabolites produced by Bacillus atrophaeus against Verticillium dahliae. Front. Microbiol. 2018, 9, 924. [Google Scholar] [CrossRef] [PubMed]
- Diale, M.O.; Ubomba-Jaswa, E.; Serepa-Dlaini, M.H. The antibacterial activity of bacterial endophytes isolated from Combretum molle. Afr. J. Biotechnol. 2018, 17, 255–262. [Google Scholar] [CrossRef]
- Xu, J.X.; Li, Z.Y.; Yan, H.; Zhou, G.Y.; Cao, L.X.; Yang, Q.; He, Y.H. Isolation and characterization of Bacillus subtilis strain 1-L-29, an endophytic bacteria from Camellia oleifera with antimicrobial activity and efficient plant-root colonization. PLoS ONE 2020. [Google Scholar] [CrossRef] [PubMed]
- Mends, M.T.; Yu, E.; Strobel, G.A.; Riyaz-Ul-Hassan, S.; Booth, E.; Geary, B.; Sears, J.; Taatjes, C.A.; Hadi, M.Z. An Endophytic Nodulisporium sp. Producing Volatile Organic Compounds Having Bioactivity and Fuel Potential. J. Pet. Environ. Biotechnol. 2012, 3, 117. [Google Scholar] [CrossRef]
- D’Alessandro, M.; Erb, M.; Ton, J.; Brandenburg, A.; Karlen, D.; Zopfi, J.; Turlings, T.C.J. Volatiles produced by soil-borne endophytic bacteria increase plant pathogen resistance and affect tritrophic interactions. Plant Cell Environ. 2014, 37, 813–826. [Google Scholar] [CrossRef]
- Gao, Z.; Zhang, B.; Liu, H.; Han, J.; Zhang, Y. Identification of endophytic Bacillus velezensis ZSY-1 strain and antifungal activity of its volatile compounds against Alternaria solani and Botrytis cinerea. Biol. Control 2017. [Google Scholar] [CrossRef]
- Massawe, V.C.; Hanif, A.; Farzand, A.; Mburu, D.K.; Ochola, S.O.; Wu, L.; Tahir, H.A.S.; Gu, Q.; Wu, H.; Gao, X. Volatile Compounds of Endophytic Bacillus spp. have Biocontrol Activity Against Q, 1 Sclerotinia sclerotiorum. Phytopathology 2018, 1–13. [Google Scholar] [CrossRef]
- Xie, S.; Liu, J.; Gu, S.; Chen, X.; Jiang, H.; Ding, T. Antifungal activity of volatile compounds produced by endophytic Bacillus subtilis DZSY21 against Curvularia lunata. Ann. Microbiol. 2020, 70, 2. [Google Scholar] [CrossRef]
- Palumbo, J.D.; Yuen, G.Y.; Jochum, C.C.; Tatum, K.; Kobayashi, D.Y. Mutagenesis of beta-1,3-Glucanase Genes in Lysobacter enzymogenes Strain C3 Results in Reduced Biological Control Activity Toward Bipolaris Leaf Spot of Tall Fescue and Pythium Damping-Off of Sugar Beet. Phytopathology 2005, 95, 701–707. [Google Scholar] [CrossRef]
- Ren, J.H.; Li, H.; Wanga, Y.F.; Ye, J.R.; Yan, A.Q.; Wua, X.Q. Biocontrol potential of an endophytic Bacillus pumilus JK-SX001 against poplar canker. Biol. Control 2013, 67, 421–430. [Google Scholar] [CrossRef]
- El-Deeb, B.; Fayez, K.; Gherbawy, Y. Isolation and characterization of endophytic bacteria from Plectranthus tenuiflorus medicinal plant in Saudi Arabia desert and their antimicrobial activities. J. Plant Interact. 2013, 8, 56–64. [Google Scholar] [CrossRef]
- Zhao, L.; Xu, Y.; Lai, X.H.; Shan, C.Z.; Deng Yuliang, J. Screening and characterization of endophytic Bacillus and Paenibacillus strains from medicinal plant Lonicera japonica for use as potential plant growth promoters. Braz. J. Microbiol. 2015, 46, 977–989. [Google Scholar] [CrossRef]
- De Kesel, J.; Conrath, U.; Flors, V.; Luna, E.; Mageroy, M.H.; Mauch-Mani, B.; Pastor, V.; Pozo, M.J.; Pieterse, C.M.J.; Ton, J.; et al. The induced resistance lexicon: Do’s and don’ts. TIPS 2021. [Google Scholar] [CrossRef]
- Pieterse, C.M.J.; Van Wees, S.C.M.; Van Pelt, J.A.; Knoester, M.; Laan, R.; Weisbeek, P.J.; van Loon, L.C. A novel signaling pathway controlling induced systemic resistance in Arabidopsis. Plant Cell 1998, 10, 1571–1580. [Google Scholar] [CrossRef]
- Mauch-Mani, B.; Baccelli, I.; Una, E.; Flors, V. Defense priming: An adaptative part of induced resistance. Annu. Rev. Plant Biol. 2017, 68, 485–512. [Google Scholar] [CrossRef] [PubMed]
- Ward, E.R.; Uknes, S.J.; Williams, S.C.; Dincher, S.S.; Wiederhold, D.L.; Alexander, D.C.; Ahl-Goy, P.; Métraux, J.P.; Ryals, J.A. Coordinate gene activity in response to agents that induce systemic acquired resistance. Plant Cell 1991, 3, 1085–1094. [Google Scholar] [CrossRef] [PubMed]
- Van Loon, L.C.; Van Strien, E.A. The families of pathogenesis-related proteins, their activities, and comparative analysis of PR-1 type proteins. Physiol. Mol. Plant. Path. 1999, 55, 85–97. [Google Scholar] [CrossRef]
- Van Oosten, V.R.; Bodenhausen, N.; Rymond, P.; Van Pelt, J.A.; van Loon, L.C.; Dicke, M.; Pieterse, C.M. Differential effectiveness of microbially induced resistance against herbivorous insects in Arabidopsis. Mol. Plant Microbe Interact. 2008, 21, 919–930. [Google Scholar] [CrossRef]
- Gao, F.K.; Dai, C.C.; Liu, X.Z. Mechanisms of fungal endophytes in plant protection against pathogens. Afr. J. Microbiol. Res. 2010, 4, 1346–1351. [Google Scholar]
- Niu, D.D.; Liu, H.X.; Jiang, C.H.; Wang, Y.P.; Wang, Q.Y.; Jin, H.L.; Guo, J.H. The plant growth-promoting rhizobacterium Bacillus cereus AR156 induces systemic resistance in Arabidopsis thaliana by simultaneously activating salicylate- and jasmonate/ethylene-dependent signaling pathways. Mol. Plant Microbe Interact. 2011, 24, 533–542. [Google Scholar] [CrossRef]
- Maurhofer, M.; Hase, C.; Meuwly, P.; Metraux, J.P.; Defago, G. Induction of systemic resistance of tobacco to tobacco necrosis virus by the root-colonizing Pseudomonas fluorescens strain CHAO: Influence of the gacA gene and of pyoverdine production. Phytopathology 1994, 84, 139–146. [Google Scholar] [CrossRef]
- Pandey, S.S.; Singh, S.; Babu, C.S.; Shanker, K.; Srivastava, N.K.; Kalra, A. Endophytes of opium poppy differentially modulate host plant productivity and genes for the biosynthetic pathway of benzylisoquinoline alkaloids. Planta 2016, 43, 1097–1114. [Google Scholar] [CrossRef]
- Pandey, S.S.; Singh, S.; Babu, C.S.; Shanker, K.; Srivastava, N.K.; Shukla, A.K. Fungal endophytes of Catharanthus roseus enhance vindoline content by modulating structural and regulatory genes related to terpenoid indole alkaloid biosynthesis. Sci. Rep. 2016, 6, 26583. [Google Scholar] [CrossRef]
- Sreekanth, D.; Kristin, I.M.; Brett, A.N. Endophytic fungi from Cathranthus roseus: A potential resource for the discovery of antimicrobial polyketides. Nat. Prod. Chem. Res. 2017, 5, 256. [Google Scholar] [CrossRef]
- Gond, S.K.; Bergena, M.S.; Torresa, M.S.; White, J.F. Endophytic Bacillus spp. produce antifungal lipopeptides and induce host defense gene expression in maize. Microbiol. Res. 2015, 17, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Audenaert, K.; Pattery, T.; Cornelis, P.; Höfte, M. Induction of systemic resistance to Botrytis cinerea in tomato by Pseudomonas aeruginosa 7NSK2: Role of salicylic acid, pyochelin and pyocyanin. Mol. Plant Microbe Interact. 2002, 15, 1147–1156. [Google Scholar] [CrossRef] [PubMed]
- Ryu, C.M.; Farag, M.A.; Hu, C.H.; Reddy, M.S.; Kloepper, J.W.; Paré, P.W. Bacterial volatiles induce systemic resistance in Arabidopsis. Plant Physiol. 2004, 134, 1017–1026. [Google Scholar] [CrossRef] [PubMed]
- Schuhegger, R.; Ihring, A.; Gantner, S.; Bahnweg, G.; Knaooe, C.; Vogg, G.; Hutzler, P.; Schmid, M.; Breusegem, F.V.; Eberl, L.; et al. Induction of systemic resistance in tomato by N-acyl-L-homoserine lactone-producing rhizosphere bacteria. Plant Cell Environ. 2006, 29, 909–918. [Google Scholar] [CrossRef]
- Pérez-García, A.; Romero, D.; de Vicente, A. Plant protection and growth stimulation by microorganisms: Biotechnological application of Bacilli in agriculture. Curr. Opin. Biotechnol. 2011, 22, 187–193. [Google Scholar] [CrossRef]
- Benhamou, N.; Gagné, S.; Quéré, D.L.; Dehbi, L. Bacterial-Mediated Induced Resistance in Cucumber: Beneficial Effect of the Endophytic Bacterium Serratia plymuthica on the Protection against Infection by Pythium ultimum. Biochem. Cell Biol. 2000, 90, 45–56. [Google Scholar] [CrossRef]
- Dalal, J.M.; Kulkarni, N.S.; Bodhankar, M.G. Utilization of Indigenous Endophytic Microbes for Induction of Systemic Resistance (ISR) in Soybean (Glycine Max (L) Merril) Against Challenge Inoculation with F. oxysporum. Res. Biotechnol. 2015, 6, 10–25. [Google Scholar]
- Alvin, A.; Miller, K.I.; Neilan, B.A. Exploring the potential of endophytes from medicinal plants as sources of antimycobacterial compounds. Microbiol. Res. 2014, 169, 483–495. [Google Scholar] [CrossRef] [PubMed]
- Fallahzadeh-Mamaghani, V.; Ahmadzadeh, M.; Sharifi, R. Screening systemic resistance-inducing fluorescent pseudomonads for control of bacterial blight of cotton caused by Xanthomonas campestris pv. malvacearum. Plant Pathol. 2009, 91, 663–670. [Google Scholar] [CrossRef]
- Chandrasekaran, M.; Chun, S. Induction of defense related enzymes in tomato (Solanum lycopersicum) plants treated with Bacillus subtilis CBR05 against Xanthomonas campestris pv. Vesicatoria. Biocontrol Sci. Technol. 2016, 26, 1366–1378. [Google Scholar] [CrossRef]
- Harish, S.; Kavino, M.; Kumar, N.; Samiyappan, R. Biopriming Banana with Plant Growth-Promoting Endophytic Bacteria Induces Systemic Resistance against Banana bunchy top virus. Acta Hortic. 2009, 828, 30. [Google Scholar] [CrossRef]
- Beris, D.; Theologidis, I.; Skandalis, N.; Vassilakos, N. Bacillus amyloliquefaciens strain MBI600 induces salicylic acid dependent resistance in tomato plants against Tomato spotted wilt virus and Potato virus Y. Sci. Rep. 2018, 8, 10320. [Google Scholar] [CrossRef]
- Manohar Jebakumar, R.; Selvarajan, R. Biopriming of microprapagated banana plants at pre- or post-BBTV inoculation stage with rhizosphere and endophytic bacteria determines their ability to induce systemic resistance against BBTV in cultivar Grand Naine. Biocontrol Sci. Technol. 2018, 28, 1074–1090. [Google Scholar] [CrossRef]
- Munif, A.; Hallmann, J.; Sicora, R.A. Induced systemic resistance of selected endophytic bacteria against Meloidogyne incognita on tomato. Meded. Rijksuniv. Gent. Fak. Landbouwkd. Toegep. Biol. Wet. 2001, 66, 663–669. [Google Scholar]
- Li, H.; Soares, M.A.; Torres, M.S.; Bergen, M.; White, J.F., Jr. Endophytic bacterium, Bacillus amyloliquefaciens, enhances ornamental hosta resistance to diseases and insect pests. J. Plant Interact. 2015, 10, 224–229. [Google Scholar] [CrossRef]
- Rashid, H.O.; Chung, Y.R. Induction of Systemic Resistance against Insect Herbivores in Plants by Beneficial Soil Microbes. Front. Plant Sci. 2017, 8, 1816. [Google Scholar] [CrossRef] [PubMed]
- Singh, M.; Srivastava, M.; Kumar, A.; Singh, A.K.; Pandey, K.D. Endophytic bacteria in plant disease management in Microbial Endophytes: Prospects for Sustainable Agriculture. Woodhead Publ. Ser. Food Sci. Technol. Nutr. 2020, 61–89. [Google Scholar] [CrossRef]
- Delaney, T.P. Genetic dissection of acquired resistance to disease. Plant Physiol. 1997, 113, 5–12. [Google Scholar] [CrossRef] [PubMed]
- Golshani, F.; Fakheri, B.A.; Behshad, E.; Vashvaei, R.M. PRs proteins and their mechanism in plants. Biol. Forum Int. J. 2015, 7, 477–495. [Google Scholar]
- Cai, X.Q.; Lin, N.; Chen, W.; Hu, F.P. Control effects on litchi downy blight disease by endophytic bacterial strain Tb2 and its pathogenesis-related proteins. Acta Hortic. 2010, 863, 631–636. [Google Scholar] [CrossRef]
- Nie, P.; Li, X.; Wang, S.; Guo, J.; Zhao, H.; Niu, D. Induced systemic resistance against Botrytis cinerea by Bacillus cereus AR156 through a JA/ET- and NPR1- dependent signaling pathway and activates PAMP-triggered immunity in Arabidopsis. Front. Plant Sci. 2017, 8, 238. [Google Scholar] [CrossRef] [PubMed]
- Mishra, A.; Singh, S.P.; Mahfooz, S.; Singh, S.P.; Bhattacharya, A.; Mishra, N.; Nautiyal, C.S. Endophyte-Mediated Modulation of Defense-Related Genes and Systemic Resistance in Withania somnifera (L.) Dunal under Alternaria alternata Stress. Appl. Environ. Microbiol. 2018, 84, 8. [Google Scholar] [CrossRef] [PubMed]
- Benhamou, N.; Kloepper, J.W.; Tuzun, S. Induction of resistance against Fusarium wilt of tomato by combination of chitosan with an endophytic bacterial strain: Ultrastructure and cytochemistry of the host response. Planta 1998, 204, 153–168. [Google Scholar] [CrossRef]
- Conn, V.M.; Walker, A.R.; Franco, C.M.M. Endophytic Actinobacteria Induce Defense Pathways. Mol. Plant Microbe Interact. 2008, 21, 208–218. [Google Scholar] [CrossRef]
- Karthikeyan, M.; Radhika, K.; Mathiyazhagan, S.; Bhaskaran, R.; Samiyappan, R.; Velazhahan, R. Induction of phenolics and defense related enzymes in coconut (Cocos nucifera L.) roots treated with biocontrol agents. Braz. J. Plant Physiol. 2006, 18, 367–377. [Google Scholar] [CrossRef]
- Gajanayaka, G.M.D.R.; Prasannath, K.; De Costa, D.M. Variation of chitinase and β-1,3 glucanase activities in tomato and chilli tissues grown under different crop management practices and agroecological regions. In Proceedings of the Peradeniya University International Research Sessions, Sri Lanka, Jaffna, 4–5 July 2014; Volume 18, p. 519. [Google Scholar]
- Prasannath, K.; De Costa, D.M. Induction of peroxidase activity in tomato leaf tissues treated with two crop management systems across a temperature gradient. In Proceedings of the International Conference on Dry Zone Agriculture 2015; Faculty of Agriculture, University of Jaffna: Jaffna, Sri Lanka, 2015; pp. 34–35. [Google Scholar]
- Ting, A.S.Y.; Meon, S.; Kadir, J.; Radu, S.; Singh, G. Induction of host defense enzymes by the endophytic bacterium Serratia marcescens, in banana plantlets. Int. J. Pest Manag. 2010, 56, 183–188. [Google Scholar] [CrossRef]
- Ludwig-Meuller, J. Plants and endophytes: Equal partners in secondary metabolite production? Biotechnol. Lett. 2015, 37, 1325–1334. [Google Scholar] [CrossRef]
- Ogbe, A.A.; Finnie, J.F.; Van Staden, J. The role of endophytes in secondary metabolites accumulation in medicinal plants under abiotic stress. S. Afr. J. Bot. 2020, 8, 46. [Google Scholar] [CrossRef]
- Pedras, M.S.C.; Zheng, Q.-A.; Gadagi, R.S.; Rimmer, S.R. Phytoalexins and polar metabolites from the oil seeds canola and rapeseed: Differential metabolic responses to the biotroph Albugo candida and to abiotic stress. Phytochemistry 2008, 69, 894–910. [Google Scholar] [CrossRef]
- Toussaint, J.P.; Smith, F.A.; Smith, S.E. Arbuscular mycorrhizal fungi can induce the production of phytochemicals in sweet basil irrespective of phosphorus nutrition. Mycorrhiza 2007, 17, 291–297. [Google Scholar] [CrossRef]
- Araim, G.; Saleem, A.; Arnason John, T.; Charest, C. Root colonization by an Arbuscular Mycorrhizal (AM) fungus increases growth and secondary metabolism of purple coneflower, Echinacea purpurea (L.) Moench. J. Agric. Food Chem. 2009, 57, 2255–2258. [Google Scholar] [CrossRef]
- Ramos-Solano, B.; Algar, E.; Gutierrez-Mañero, F.J.; Bonilla, A.; Lucas, J.A.; García-Seco, D. Bacterial bioeffectors delay postharvest fungal growth and modify total phenolics, flavonoids and anthocyanins in blackberries. LWT Food Sci. Technol. 2015, 61, 437–443. [Google Scholar] [CrossRef]
- van de Mortel, J.E.; Vos, R.C.H.; De Dekkers, E.; Pineda, A.; Guillod, L.; Bouwmeester, K.; van Loon, J.J.A.; Dicke, M.; Raaijmakers, J.M. Metabolic and transcriptomic changes induced in Arabidopsis by the rhizobacterium Pseudomonas. Plant Physiol. 2012, 160, 2173–2188. [Google Scholar] [CrossRef]
- Planchamp, C.; Glauser, G.; Mauch-Mani, B. Root inoculation with Pseudomonas putida KT2440 induces transcriptional and metabolic changes and systemic resistance in maize plants. Front. Plant Sci. 2014, 5, 719. [Google Scholar] [CrossRef] [PubMed]
- Ongena, M.; Jacques, P.; Touré, Y.; Destain, J.; Jabrane, A. Involvement of fengycin-type lipopeptides in the multifaceted biocontrol potential of Bacillus subtilis. Appl. Microbiol. Biotechnol. 2005, 69, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Schenk, S.T.; Hernández-reyes, C.; Samans, B.; Stein, E.; Neumann, C.; Schikora, M.; Reichelt, M.; Mithöfer, A.; Becker, A.; Kogel, K.H.; et al. N-acyl-homoserine lactone primes plants for cell wall reinforcement and induces resistance to bacterial pathogens via the salicylic acid / oxylipin pathway. Plant Cell 2014, 26, 2708–2723. [Google Scholar] [CrossRef] [PubMed]
- Schikora, A. Beneficial effects of bacteria-plant communication based on quorum sensing molecules of the N-acyl homoserine lactone group. Plant Methods 2016, 90, 605–612. [Google Scholar] [CrossRef] [PubMed]
- Priti, V.; Ramesha, B.T.; Singh, S.; Ravikanth, G.; Ganeshaiah, K.N.; Suryanarayanan, T.S.; Uma Shaanker, R. How promising are endophytic fungi as alternative sources of plant secondary metabolites. Curr. Sci. 2009, 97, 477–478. [Google Scholar]
- Torres, M.S.; White, J.F., Jr.; Zhang, X.; Hinton, D.M.; Bacon, C.W. Endophyte mediated adjustments in host morphology and physiology and effects on host fitness traits in grasses. Fungal Ecol. 2012, 5, 322–330. [Google Scholar] [CrossRef]
- Zhou, F.; Gao, Y.B.; Ma, W.J. Effects of phosphorus deficiency on growth of perennial rye grass fungal endophyte symbiont and phenolic content in root. Plant Physiol. Commun. 2003, 39, 321–325. [Google Scholar]
- Taghavi, S.; van der Lelie, D.; Hoffman, A.; Zhang, Y.B.; Walla, M.D.; Vangronsveld, J.L.; Newman, S. Monchy. Genome sequence of the plant growth promoting endophytic bacterium Enterobacter sp. 638. PLoS Genet. 2010, 6, e1000943. [Google Scholar] [CrossRef]
- Khan, A.R.; Park, G.S.; Asaf, S.; Hong, S.J.; Jung, B.K.; Shin, J.H. Complete genome analysis of Serratia marcescens RSC-14: A plant growth-promoting bacterium that alleviates cadmium stress in host plants. PLoS ONE 2017, 12, e0171534. [Google Scholar] [CrossRef]
- Pasternak, T.; Potters, G.; Caubergs, R.; Jansen, M.A. Complementary interactions between oxidative stress and auxins control plant growth responses at plant, organ, and cellular level. J. Exp. Bot. 2005, 56, 1991–2001. [Google Scholar] [CrossRef]
- Yue, C.C.; Miller, J.; White, J.; Richardson, M. Isolation and characterization of fungal inhibitors from Epichloe festucae. J. Agric. Food Chem. 2000, 48, 4687–4692. [Google Scholar] [CrossRef]
- Piccoli, P.; Travaglia, C.; Cohen, A.; Sosa, L.; Cornejo, P.; Masuelli, R.; Bottini, R. An endophytic bacterium isolated from roots of the halophyte Prosopis strombulifera produces ABA, IAA, gibberellins A1 and A3 and jasmonic acid in chemically-defined culture medium. Plant Growth Regul. 2011, 64, 207–210. [Google Scholar] [CrossRef]
- de Santi Ferrara, F.I.; Oliveira, Z.M.; Gonzales, H.H.S.; Floh, E.I.S.; Barbosa, H.R. Endophytic and rhizospheric enterobacteria isolated from sugar cane have different potentials for producing plant growth-promoting substances. Plant Soil. 2012, 353, 409–417. [Google Scholar] [CrossRef]
- Mishra, S.K.; Khan, M.H.; Misra, S.; Dixit, K.V.; Khare, P.; Srivastava, S.; Chauhan, P.S. Characterisation of Pseudomonas spp. and Ochrobactrum sp. isolated from volcanic soil. Antonie Van Leeuwenhoek 2017, 110, 253–270. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Yin, Z.H.; Yin, C.Y. Biotransformation of ginsenoside Rb1 to ginsenoside Rg3 by endophytic bacterium Burkholderia sp. GE 17_7 isolated from Panax ginseng. J. Appl. Microbiol. 2017, 122, 1579–1585. [Google Scholar] [CrossRef]
- Li, J.; Zhao, G.Z.; Varma, A.; Qin, S.; Xiong, Z.; Huang, H.Y.; Li, W.J. An endophytic Pseudonocardia species induces the production of artemisinin in Artemisia annua. PLoS ONE 2012, 7, e51410. [Google Scholar] [CrossRef] [PubMed]
- Arc, E.; Sechet, J.; Corbineau, F.; Rajjou, L.; Marion-Poll, A. ABA crosstalk with ethylene and nitric oxide in seed dormancy and germination. Front. Plant Sci. 2013, 4, 63. [Google Scholar] [CrossRef]
- Bakshi, A.; Shemansky, J.M.; Chang, C.; Binder, B.M. History of research on the plant hormone ethylene. J. Plant Growth Regul. 2015, 34, 809–827. [Google Scholar] [CrossRef]
- Wei, L.; Deng, X.G.; Zhu, T.; Zheng, T.; Li, P.X.; Wu, J.Q.; Lin, H.H. Ethylene is involved in brassinosteroids induced alternative respiratory pathway in cucumber (Cucumis sativus L.) seedlings response to abiotic stress. Front. Plant Sci. 2015, 6, 982. [Google Scholar] [CrossRef] [PubMed]
- Hardoim, P.R.; van Overbeek, L.S.; Berg, G.; Pirttilä, A.M.; Compant, S.; Campisano, A.; Döring, M.; Sessitsch, A. The hidden world within Plants: Ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiol. Mol. Biol. Rev. 2015, 79, 293–320. [Google Scholar] [CrossRef] [PubMed]
- Bargabus, R.L.; Zidack, N.K.; Shewood, J.E.; Jacobsen, B.J. C haracterization of systemic resistance in sugar beet elicited by a non-pathogenic, phyllosphere-colonizing Bacillus mycoides, biological control agent. Physiol. Mol. Plant Path. 2002, 61, 289–298. [Google Scholar] [CrossRef]
- Li, S.B.; Fang, M.; Zhou, R.C.; Huang, J. Characterization and evaluation of the endophyte Bacillus B014 as a potential biocontrol agent for the control of Xanthomonas axonopodis pv. dieffenbachiae—Induced blight of Anthurium. Biol. Control 2012, 63, 9–16. [Google Scholar] [CrossRef]
- Brock, A.K.; Berger, B.; Mewis, I.; Ruppel, S. Impact of the PGPB Enterobacter radicincitans DSM 16656 on Growth, Glucosinolate Profile, and Immune Responses of Arabidopsis thaliana. Microb. Ecol. 2013, 65, 661–670. [Google Scholar] [CrossRef]
- Cawoy, H.; Mariutto, M.; Henry, G.; Fisher, C.; Vasilyeva, N.; Thonart, P.; Dommes, J.; Ongena, M. Plant defense stimulation by natural isolates of Bacillus depends on efficient surfactin production. Mol. Plant Microbe Interact. 2014, 27, 87–100. [Google Scholar] [CrossRef]
- Gómez-Lama Cabanás, C.; Schilirò, E.; Valverde-Corredor, A.; Mercado-Blanco, J. The biocontrol endophytic bacterium Pseudomonas fluorescens PICF7 induces systemic defense responses in aerial tissues upon colonization of olive roots. Front. Microbiol. 2014, 5, 427. [Google Scholar] [CrossRef]
- Chowdhury, S.P.; Uhl, J.; Grosch, R.; Alquéres, S.; Pittroff, S.; Dietel, K.; Schmitt-Kopplin, P.; Borriss, R.; Hartmann, A. Cyclic lipopeptides of Bacillus amyloliquefaciens subsp. Plantarum colonizing the lettuce rhizosphere enhance plant defense responses toward the bottom rot pathogen Rhizoctonia solani. Mol. Plant Microbe Interact. 2015, 28, 984–995. [Google Scholar] [CrossRef]
- Pangesti, N.; Pineda, A.; Dicke, M.; Van Loon, J.J.A. Variation in plant-mediated interactions between rhizobacteria and caterpillars: Potential role of soil composition. Plant Biol. 2015, 17, 474–483. [Google Scholar] [CrossRef]
- Pangesti, N.; Reichelt, M.; van de Mortel, J.E.; Kapsomenou, E.; Gershenzon, J.; van Loon, J.J.; Dicke, M.; Pineda, A. Jasmonic acid and ethylene signaling pathways regulate glucosinolate levels in plants during rhizobacteria-induced systemic resistance against a leaf-chewing herbivore. J. Chem. Ecol. 2016, 42, 1212–1225. [Google Scholar] [CrossRef] [PubMed]
- Zebelo, S.; Song, Y.; Kloepper, J.W.; Fadamiro, H. Rhizobacteria activates (C)-d-cadinene synthase genes and induces systemic resistance in cotton against beet armyworm (Spodoptera exigua). Plant Cell Environ. 2016, 39, 935–943. [Google Scholar] [CrossRef]
- Haidar, R.; Roudet, J.; Bonnard, O.; Dufour, M.C.; Corio-Costet, M.F.; Fert, M.; Gautier, T.; Deschamps, A.; Fermaud, M. Screening and modes of action of antagonistic bacteria to control the fungal pathogen Phaeomoniella chlamydospora involved in grapevine trunk diseases. Microbiol. Res. 2016, 192, 172–184. [Google Scholar] [CrossRef] [PubMed]
- Patel, J.K.; Madaan, S.; Archana, G. Antibiotic producing endophytic Streptomyces spp. colonize above-ground plant parts and promote shoot growth in multiple healthy and pathogen challenged cereal crops. Microbiol. Res. 2018, 215, 36–45. [Google Scholar] [CrossRef] [PubMed]
- Yanti, Y.; Warnita; Reflin. Involvement of Jasmonic Acid in the Induced Systemic Resistance of Tomato against Ralstonia syzigiisub sp. indonesiensis by Indigenous Endophyte Bacteria. In Proceedings of the 6th International Conference on Sustainable Agriculture, Food and Energy, Earth and Environmental Science, Manila, Philippines, 18–21 October 2018; IOP Publishing: Bristol, UK, 2019; Volume 347, p. 012024. [Google Scholar] [CrossRef]
| Endophytes | Plant Host Class | Activity | Compounds | Chemical Classes | References | |
|---|---|---|---|---|---|---|
| Antibiotics | Pseudomonas viridiflava | Grass | Antifungal | Ecomycin | Peptide | [89] |
| B. subtilis 168 | Antifungal | Bacilysocin | Phospholipid | [105] | ||
| B. thuringiensis | Insecticidal | β-exotoxin | Polypeptide | [106] | ||
| Streptomyces sp. strain NRRL 30562 | Kennedia nigriscans | Antifungal/Antibacterial | Munumbicins A, B, C, and D, | Peptide | [107] | |
| Streptomyces sp. strain NRRL 30566 | Grevillea pteridifolia | Antibacterial | Kakadumycin A | Peptide | [108] | |
| Verrucosispora maris AB-18-032 | Sonchus oleraceus | Antibacterial | Proximicin | Peptide | [109] | |
| Streptomyces sp. HK 10595 | Kandelia candel | Antibacterial | Xiamycin B, Indosespine and Sespenine | Entacyclic indolosesquiterpine | [110] | |
| Streptomyces sp. | marine mudflat-derived actinomycete | Antibacterial | Harmaomycin | Peptide derivatives | [111] | |
| B. substilis | Antibacterial | Subtilin | Peptides | [112] | ||
| Bacillus atrophaeus, Bacillus mojavensis | Glycyrrhiza uralensis | Antifungal | 1,2-bezenedicarboxyl acid, Methyl ester, Decanodioic acid, bis(2-ehtylhexyl) ester | Polyketides | [113] | |
| Lysinibacillus, Staphylococcus, Enterobacter, Pseudomonas, and Bacillus species | Combretum molle | Antibacterial | [114] | |||
| Bacillus subtilis strain 1-L-29 | Camellia oleifera | Antifungal | [115] | |||
| VOCS | Nodulisporium sp. | Myroxylon balsamum | Antifungal | phenylethyl alcohol. alkyl alcohols alkyl alcohols (1-butanol-3-methyl, 1-propanol-2-methyl, 1-pentanol, 1-hexanol, 1-heptanol, 1-octanol, 1-nonanol | esters, ketones, benzene derivatives, a terpenoids, hydrocarbons. | [116] |
| Enterobacter aerogenes | Maize | Antifungal | 2,3-butanediol | [117] | ||
| Pseudomonas putida BP25 | Antifungal/antiparasitic | [96] | ||||
| Bacillus velezensis ZSY-1, | Chinese catalpa | Antifungal | 2-tridecanone, pyrazine (2,5-dimethyl), benzothiazole, and phenol (4-chloro-3-methyl) | ketones, alcohols, and alkanes | [118] | |
| Bacillus spp. | Antifungal | 1-Hexadecanol, Hexacosyl acetate, Tryphenylphosphine oxide, 1,3-Propanediol, 2-methyl, dipropanoate, 1,4-Pentadiene, Hydroxyurea, Decyl trifluoroacetate, Pentadecane, 4-Ethyl-1-octyn-3-ol, Tridecane Benzothiazole, N,N-Dimethyldodecylamine, 1,3-Butadiene, Dodecane, 2-Undecanone IR-(+)-a-pinene | [119] | |||
| Bacillus subtilis DZSY21 | Eucommia ulmoides | Antifungal | 2-Methylbutyric acid, 2-heptanone, and isopentyl acetate | [120] | ||
| Enzymes | B. cereus 65 | mustard | Antifungal | chitinase | [93] | |
| Lysobacter enzymogenes | Sugar Beet | Antifungal | 1,3-glucanase | [121] | ||
| Streptomyces hygroscopicus | chitinase | [94] | ||||
| Bacillus pumilus JK-SX001 | Populus | Antifungal | cellulases and protease | [122] | ||
| Bacillus sp., Micrococcus sp., and P. polymyxa | Panax ginseng- and Plectranthus tenuiflorus | Antibacterial | amylase, esterase, lipase, protease, pectinase, xylanase, and cellulase | [123] | ||
| Paenibacillus sp. and Bacillus sp. | Lonicera japonica | cellulase and pectinase | [124] |
| Endophyte | Host Plant | Application of Endophytes | Plant Pathogens | Signaling Pathway | Induced Defenses | Reference |
|---|---|---|---|---|---|---|
| B. pumilus SE34 | Tomato | Germinated seeds | Fusarium oxysporum f. sp. radicis lycopersici | Elaboration of structural barriers, production phenolic compounds and β-1,3-glucanases. | [159] | |
| Serratia plymuthica | Cucumber | Seeds | Pythium ultimum | Callose-enriched wall appositions at sites of pathogen penetration. Accumulation of an osmiophilic material in the colonized areas. | [142] | |
| B. mycoides | Sugar beet | Foliar application | Cercosporonbeticola Sacc. | Increased activity of B-1,3-glucunase, chitinase, peroxidase and PR proteins. | [192] | |
| Bacillus subtilis GB03 and Bacillus amyloliquefaciens IN937 | A. thaliana | Exposition to the VOCs produced by the isolates | Erwinia carotovora subsp. carotovora | ET pathway | [139] | |
| Actinobacteria | A. thaliana | Roots | E. carotovora subsp. Carotovora | JA/ET pathway | Upregulation of, PR-1, PR-4 as well as PDF1.2 and HEL genes. | [160] |
| B. amyloliquefaciens strain TB2 | Lychee | Fruit | Peronophthora litchi | Increase PPRs production including 1,3-glucanase and chitinase. | [156] | |
| Bacillus cereus AR156 | Tomato | Roots | Pseudomonas syringae pv. tomato | Simultaneous activation of SA- and JA/Et-dependent signaling pathways | Activation of some defense-related genes including PR1, PR2, PR5, and PDF1.2. | [132] |
| Bacillus B014 | Anthurium | Foliar application | Xanthomonas axonopodis pv. dieffenbachiae | Activation of defense-related enzymes PAL, POD and PPO after pathogen attack. | [193] | |
| Enterobacter radicincitans DSM 16656 | A. thaliana | Roots | Both SA- and JA/Et-dependent signaling pathways | Accumulation of PR1, PR2, PR5, and PDF1.2 transcript 24 h after treatment with the endophyte. | [194] | |
| Bacillus spp. | Maize | Roots | Upregulation of pathogenesis-related genes. | [137] | ||
| B. amyloliquefaciens S499 | Tomato | Seeds and soil | Botrytis cinerea | Induction of lipoxygenase pathway (accumulation of transcripts of genes corresponding to the two isoforms, Tom LOXD and Tom LOXF. | [195] | |
| Pseudomonas fluorescens PICF7 | Olive | Roots | Verticillium wilt | Activation of olive genes potentially coding for lipoxygenase 2, catalase, 1-aminocyclopropane-1-carboxylate oxidase, and phenylananine ammonia-lyase. | [196] | |
| B. amyloliquefaciens strain FZB42 | Lettuce | Roots | R. solani | Higher expression of PDF 1 and 2. | [197] | |
| B. amyloliquefaciens strain Blu-v2 | Hosta | Filiar application | Spodoptera fruigiperda | Production lipopeptides that elicits ISR in plants against fall armyworms. | [151] | |
| Micromonospora spp. | Tomato | Roots | B. cinerea | JA/ET pathway | Induction of JA-regulated defenses (PINII and LOX A). | [18] |
| P. fluorescens WCS417r | A.thaliana | Soil drench | Chewing herbivores | JA/ET | Increased expression of LOX2, PDF1.2 and HEL. | [198] |
| P. simiae WCS417r | A. thaliana | Roots | Leaf-chewing herbivores | JA/ET | Higher expression of ORA59-branch respect to the JA-dependent MYC2 branch, Enhanced synthesis of camalexin and aliphatic glucosinolates (GLS). | [199] |
| Bacillus sp. | Cotton | Soil drench | Beet armyworm Spodoptera exigua | JA | Accumulation of JA, and JA-related genes. | [200] |
| B. pumilus | Grapevine | Soil drench | P. chlamydospora | Enhanced expression of different genes PR 1, PR 10, chitinase class III, PAL, stilbene synthase, chalcone synthase, anthranilate synthase, callose synthase, Glutathione S-transferase, and b-1,3 glucanase. | [201] | |
| B. subtilis | Tomato | Soil drench | Xanthomonas campestris pv. vesicatoria | Accumulation of antioxidant enzymes SOD, CAT, POD, and PPO. | [146] | |
| Bacillus amyloliquefaciens strain MBI600 | Tomato | Drench or foliar application | Tomato spotted wilt virus and Potato virus Y | SA pathway | Induction of the SA signaling pathway in tomato after MBI600 treatment. | [148] |
| Azospirillum sp. B510 | Rice | Soil drench | ET signaling is required for endophyte-mediated ISR | Induces systemic disease resistance in rice without accompanying defense-related gene expression. | [76] | |
| Streptomyces spp. | Rice, sorghum and wheat | Seedlings | Upregulation of PR10, NPR1, PAL and LOX2. | [202] | ||
| Bacillus cereus Serratia nematodiphila TLE1.1 | Tomato | Seeds | Ralstonia syzigiisub sp. | JA | Increase JA contained in leaves and roots of tomato significantly until 12 days after pathogen inoculation. | [203] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oukala, N.; Aissat, K.; Pastor, V. Bacterial Endophytes: The Hidden Actor in Plant Immune Responses against Biotic Stress. Plants 2021, 10, 1012. https://doi.org/10.3390/plants10051012
Oukala N, Aissat K, Pastor V. Bacterial Endophytes: The Hidden Actor in Plant Immune Responses against Biotic Stress. Plants. 2021; 10(5):1012. https://doi.org/10.3390/plants10051012
Chicago/Turabian StyleOukala, Nadira, Kamel Aissat, and Victoria Pastor. 2021. "Bacterial Endophytes: The Hidden Actor in Plant Immune Responses against Biotic Stress" Plants 10, no. 5: 1012. https://doi.org/10.3390/plants10051012
APA StyleOukala, N., Aissat, K., & Pastor, V. (2021). Bacterial Endophytes: The Hidden Actor in Plant Immune Responses against Biotic Stress. Plants, 10(5), 1012. https://doi.org/10.3390/plants10051012







