Functional Annotation of a Full-Length Transcriptome and Identification of Genes Associated with Flower Development in Rhododendron simsii (Ericaceae)
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. RNA Extraction
2.3. Library Construction and SMRT Sequencing
2.4. Error Correction and Quality Filtering
2.5. Functional Annotation of Transcripts
2.6. Identification of TFs, lncRNAs and SSR
2.7. Development of SSR Markers
2.8. qRT-PCR Analysis
3. Results and Analysis
3.1. Single Molecule Real Time Sequencing (SMRT)
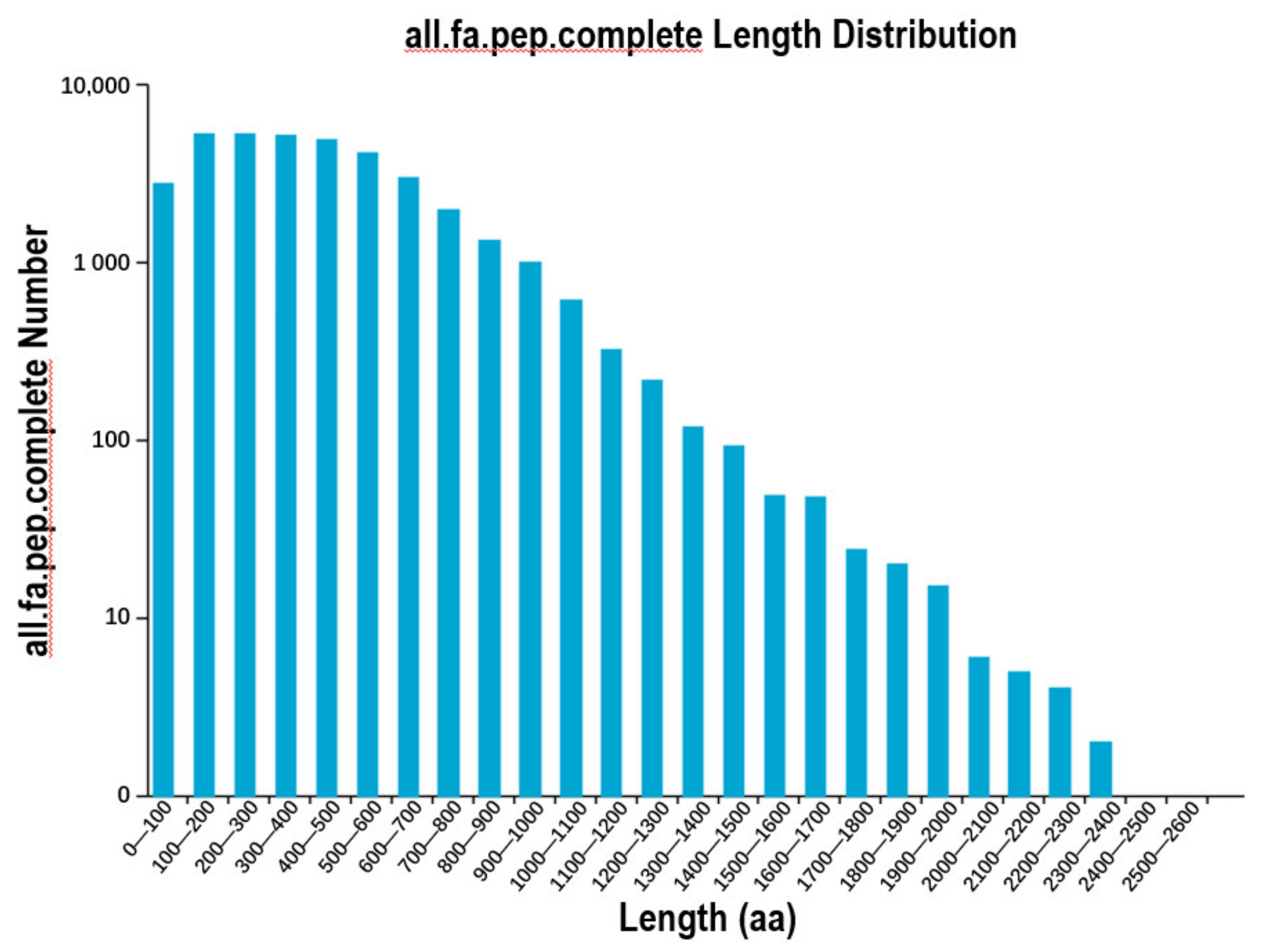
3.2. Open Reading Frame and Alternative Splicing Event Prediction
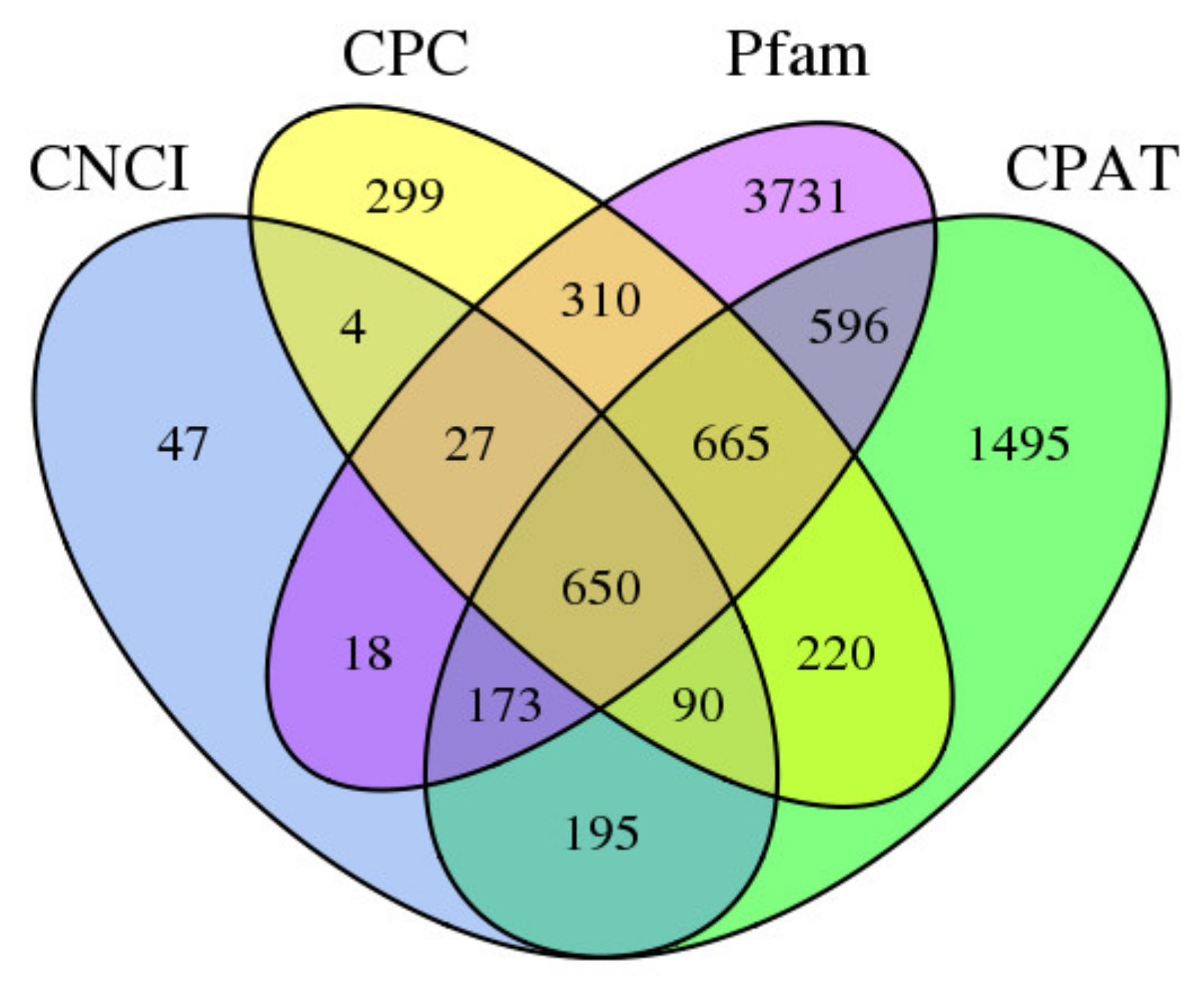
3.3. Long Non-Coding RNA Identification
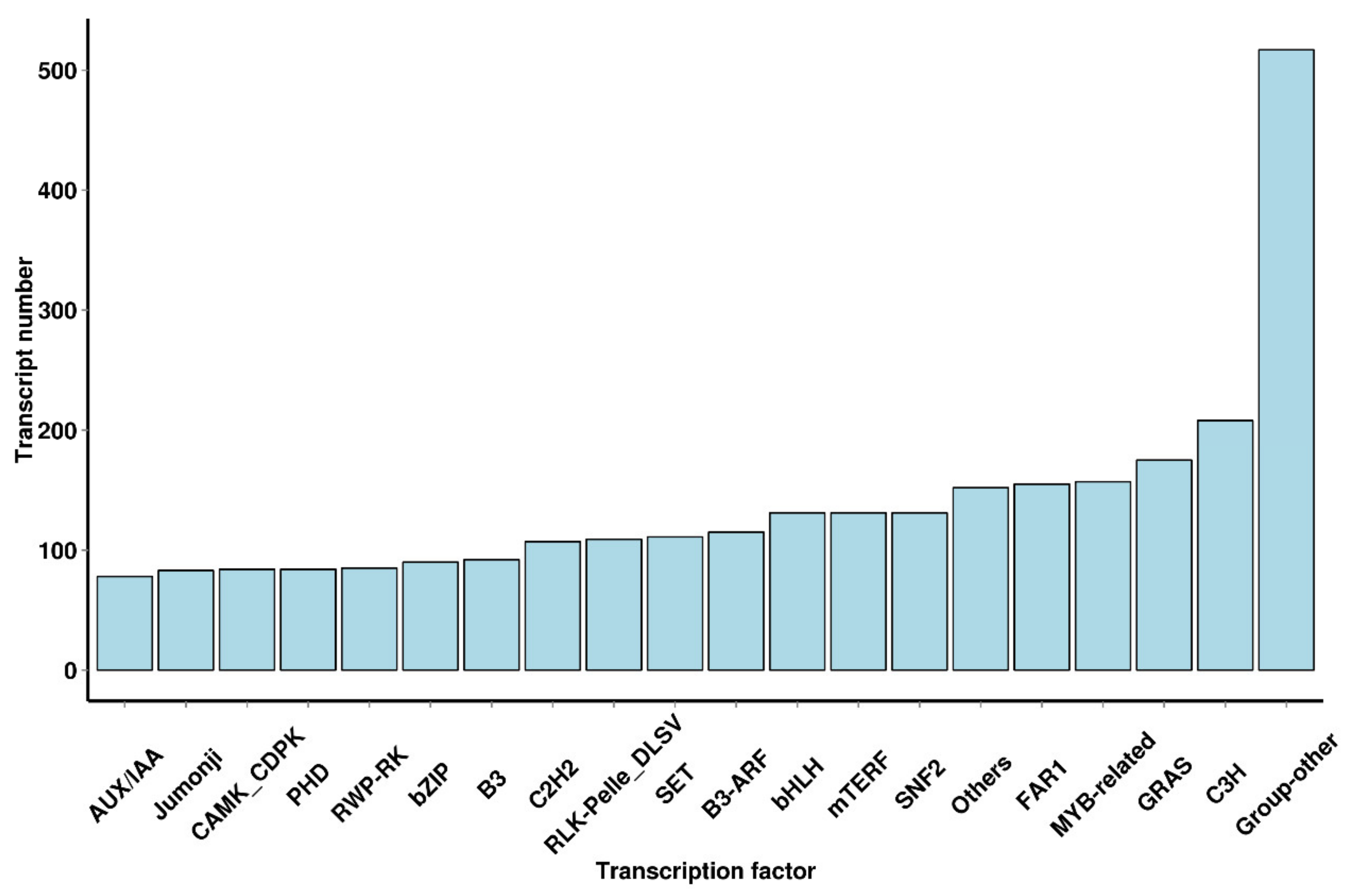
3.4. Transcription Factor Prediction
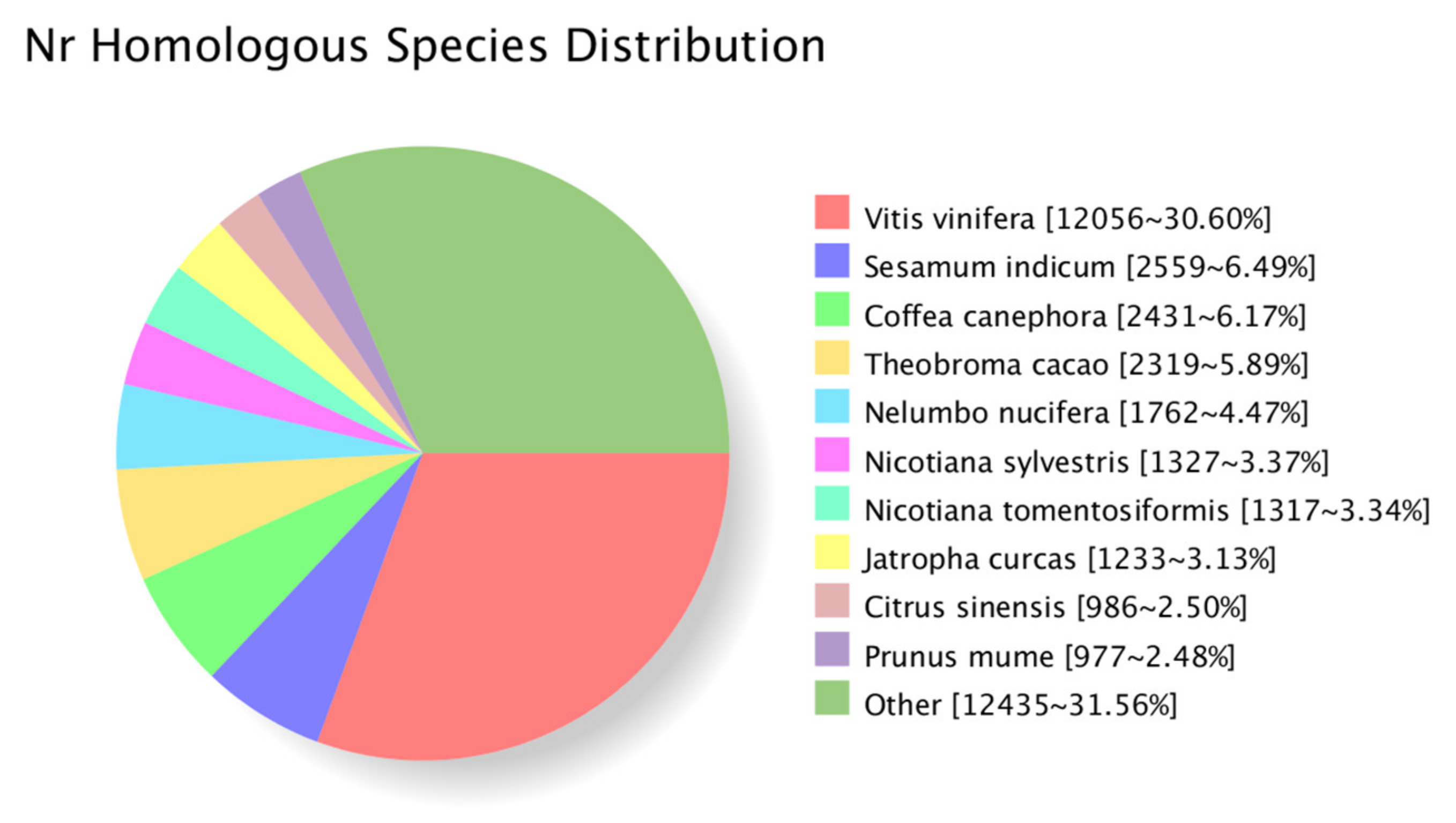
3.5. Functional Annotation of Transcripts
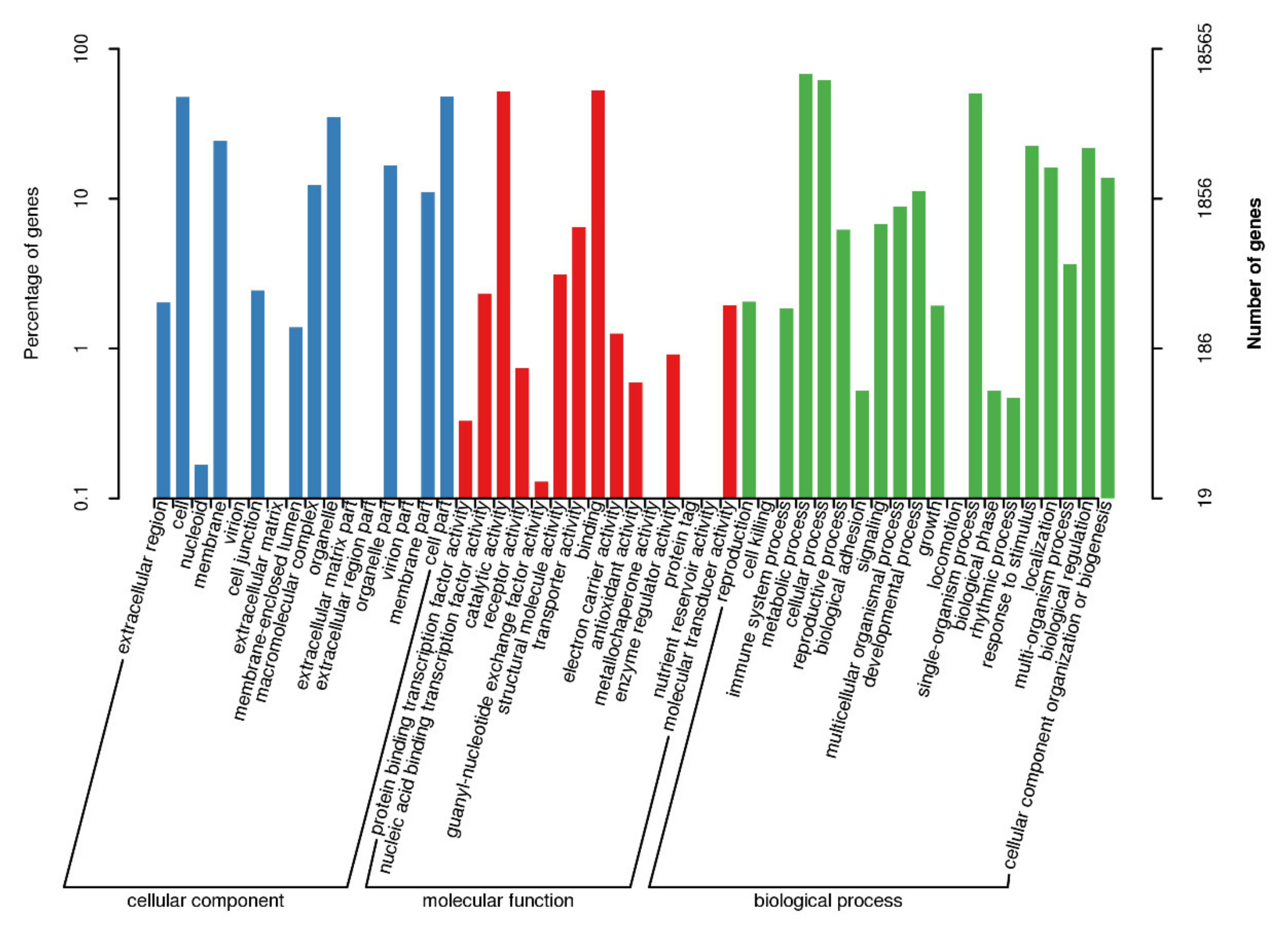
3.6. GO Classified Transcripts
3.7. COG Function Classification
3.8. KEGG Annotated Transcripts
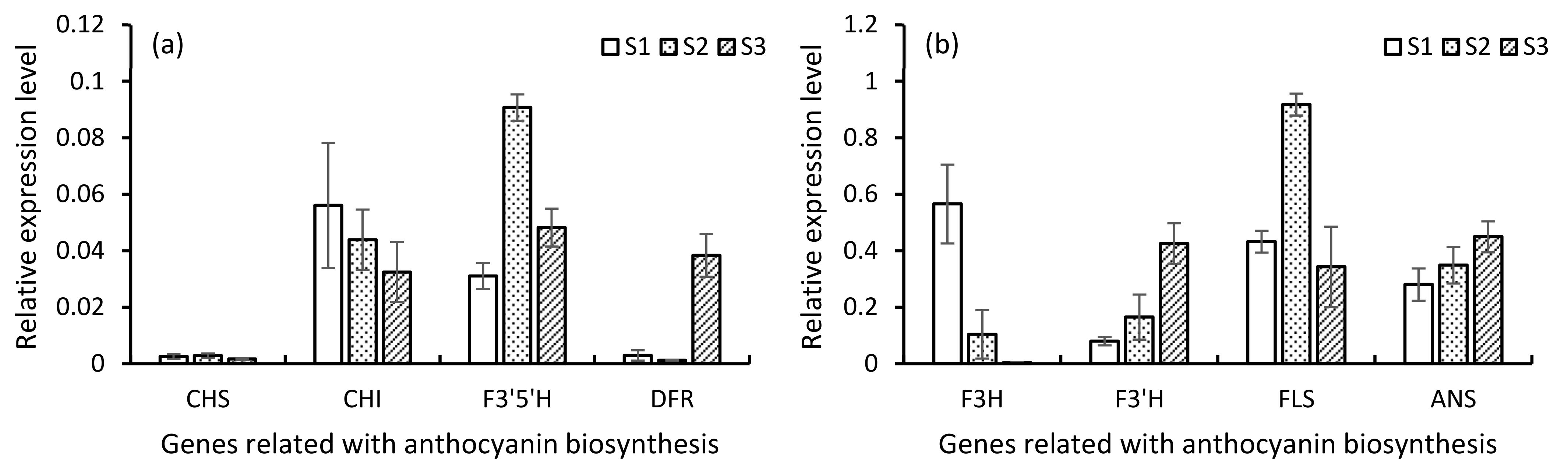
3.9. Expression Analysis in qRT PCR
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Cai, L.; Neilsen, J.; Dao, Z.L.; Ma, Y.P. Rhododendron longipedicellatum (Ericaceae), a new species from Southeastern Yunnan, China. Phytotaxa 2016, 282, 296–300. [Google Scholar] [CrossRef]
- Proestos, C.; Boziaris, I.S.; Nychas, G.J.; Komaitis, M. Analysis of flavonoids and phenolic acids in Greek aromatic plants: Investigation of their antioxidant capacity and antimicrobial activity. Food Chem. 2006, 95, 664–671. [Google Scholar] [CrossRef]
- Castañeda-Ovando, A.; Sedo, O.; Havel, J.; Pacheco, L.; Galán-Vidal, C.A.; López, E.C. Identification of anthocyanins in red grape, plum and capulin by MALDI-ToF MS. J. Mex. Chem. Soc. 2012, 56, 378–383. [Google Scholar] [CrossRef]
- Fang, Z.Z.; Zhou, D.R.; Ye, X.F.; Jiang, C.C.; Pan, S.L. Identification of candidate anthocyanin-related genes by transcriptomic analysis of ‘Furongli’plum (Prunus salicina Lindl.) during fruit ripening using RNA-seq. Front. Plant Sci. 2016, 7, 1338. [Google Scholar] [CrossRef] [PubMed]
- Mizuta, D.; Nakatsuka, A.; Ban, T.; Miyajima, I.; Kobayashi, N. Pigment composition patterns and expression of anthocyanin biosynthesis genes in Rhododendron kiusianum, R. kaempferi, and their natural hybrids on Kirishima Mountain Mass, Japan. J. Jpn. Soc. Hortic. Sci. 2013, 83, 156–162. [Google Scholar] [CrossRef]
- Khoo, H.E.; Azlan, A.; Tang, S.T.; Lim, S.M. Anthocyanidins and anthocyanins: Colored pigments as food, pharmaceutical ingredients, and the potential health benefits. Food Nutr. Res. 2017, 61, 1361779. [Google Scholar] [CrossRef] [PubMed]
- Zeng, J.; Chen, J.; Kou, Y.; Wang, Y. Application of EST-SSR markers developed from the transcriptome of Torreya grandis (Taxaceae), a threatened nut-yielding conifer tree. PeerJ 2018, 6, e5606. [Google Scholar] [CrossRef]
- Peng, Z.; Hu, Y.; Xie, J.; Potnis, N.; Akhunova, A.; Jones, J.; Liu, S. Long read and single molecule DNA sequencing simplifies genome assembly and TAL effector gene analysis of Xanthomonas translucens. BMC Genom. 2016, 17, 1–19. [Google Scholar] [CrossRef]
- Zhang, J.; Liu, C.; He, M.; Xiang, Z.; Yin, Y.; Liu, S.; Zhuang, Z. A full-length transcriptome of Sepia esculenta using a combination of single-molecule long-read (SMRT) and Illumina sequencing. Mar. Genom. 2019, 43, 54–57. [Google Scholar] [CrossRef]
- Chen, J.; Li, R.; Xia, Y.; Bai, G.; Guo, P.; Wang, Z.; Siddique, K.H. Development of EST-SSR markers in flowering Chinese cabbage (Brassica campestris L. ssp. chinensis var. utilis Tsen et Lee) based on de novo transcriptomic assemblies. PLoS ONE 2017, 12, e0184736. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Wang, X.; He, Q.; Liu, X.; Xu, W.; Li, L.; Wang, F. Transcriptome analysis of the roots at early and late seedling stages using Illumina paired-end sequencing and development of EST-SSR markers in radish. Plant Cell Rep. 2012, 31, 1437–1447. [Google Scholar] [CrossRef] [PubMed]
- Yagi, M.; Yamamoto, T.; Isobe, S.; Hirakawa, H.; Tabata, S.; Tanase, K.; Onozaki, T. Construction of a reference genetic linkage map for carnation (Dianthus caryophyllus L.). BMC Genom. 2013, 14, 734. [Google Scholar] [CrossRef] [PubMed]
- Chaisson, M.J.; Huddleston, J.; Dennis, M.Y.; Sudmant, P.H.; Malig, M.; Hormozdiari, F.; Eichler, E.E. Resolving the complexity of the human genome using single-molecule sequencing. Nature 2015, 517, 608–611. [Google Scholar] [CrossRef]
- Levy, S.E.; Myers, R.M. Advancements in next-generation sequencing. Annu. Rev. Genom. Hum. Genet. 2016, 17, 95–115. [Google Scholar] [CrossRef] [PubMed]
- Jiao, W.B.; Schneeberger, K. The impact of third generation genomic technologies on plant genome assembly. Curr. Opin. Plant Biol. 2017, 36, 64–70. [Google Scholar] [CrossRef]
- Slatko, B.E.; Gardner, A.F.; Ausubel, F.M. Overview of next-generation sequencing technologies. Curr. Protoc. Mol. Biol. 2018, 122, e59. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Sun, X.; Zhao, Y.; Guo, X.; Jiang, H.; Li, H.; Gu, Z. Evolution of gene regulation during transcription and translation. Genome Biol. Evol. 2015, 7, 1155–1167. [Google Scholar] [CrossRef]
- Xie, J.; Zuo, J.; Huang, Y.; Li, C.; Chen, Y. The origin and germplasm collection for cultivated Dendrobium officinale K. Kimura & Migo individuals revealed by EST-SSR markers. Genet. Resour. Crop Evol. 2020, 67, 1209–1219. [Google Scholar]
- Wang, J.; Zhu, J.; Zhang, Y.; Fan, F.; Li, W.; Wang, F.; Yang, J. Comparative transcriptome analysis reveals molecular response to salinity stress of salt-tolerant and sensitive genotypes of indica rice at seedling stage. Sci. Rep. 2018, 8, 1–13. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S.; Kawashima, S.; Okuno, Y.; Hattori, M. The KEGG resource for deciphering the genome. Nucleic Acids Res. 2004, 32, D277–D280. [Google Scholar] [CrossRef]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Harris, M.A. Gene ontology: Tool for the unification of biology. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Jiao, W.; Song, H.; Qu, H.; Li, D.; Mei, H.; Tong, Q. miRNA-558 promotes gastric cancer progression through attenuating Smad4-mediated repression of heparanase expression. Cell Death Dis. 2016, 7, e2382. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Luo, H.; Bu, D.; Zhao, G.; Yu, K.; Zhang, C.; Zhao, Y. Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res. 2013, 41, e166. [Google Scholar] [CrossRef]
- Kong, L.; Zhang, Y.; Ye, Z.Q.; Liu, X.Q.; Zhao, S.Q.; Wei, L.; Gao, G. CPC: Assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 2007, 35, W345–W349. [Google Scholar] [CrossRef]
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Salazar, G.A. The Pfam protein family’s database: Towards a more sustainable future. Nucleic Acids Res. 2016, 44, D279–D285. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Zhang, L.; Wang, Y.; Li, H.; Ren, X.; Wei, F.; Hao, X. Long noncoding RNA HOTAIR involvement in cancer. Tumor Biol. 2014, 35, 9531–9538. [Google Scholar] [CrossRef]
- Beier, S.; Thiel, T.; Münch, T.; Scholz, U.; Mascher, M. MISA-web: A web server for microsatellite prediction. Bioinformatics 2017, 33, 2583–2585. [Google Scholar] [CrossRef]
- Byrne, A.; Cole, C.; Volden, R.; Vollmers, C. Realizing the potential of full-length transcriptome sequencing. Philos. Trans. R. Soc. B 2019, 374, 20190097. [Google Scholar] [CrossRef]
- Wu, L.; Zhang, X.; Zhao, Z.; Wang, L.; Li, B.; Li, G.; Rao, W. Full-length single-cell RNA-seq applied to a viral human cancer: Applications to HPV expression and splicing analysis in HeLa S3 cells. Gigascience 2015, 4, 51. [Google Scholar] [CrossRef]
- Ozsolak, F. Third-generation sequencing techniques and applications to drug discovery. Expert Opin. Drug Discov. 2012, 7, 231–243. [Google Scholar] [CrossRef]
- Wei, C.; Yang, H.; Wang, S.; Zhao, J.; Liu, C.; Gao, L.; Sun, J. Draft genome sequence of Camellia sinensis var. sinensis provides insights into the evolution of the tea genome and tea quality. Proc. Natl. Acad. Sci. USA 2018, 115, E4151–E4158. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Zhang, H.; Kohnen, M.V.; Prasad, K.V.; Gu, L.; Reddy, A.S. Analysis of transcriptome and epitranscriptome in plants using PacBio Iso-Seq and nanopore-based direct RNA sequencing. Front. Genet. 2019, 10, 253. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Zhai, Y.; Luo, X.; Zhang, Y.; Shi, Q. Comparative transcriptome analyses reveal genes related to pigmentation in the petals of red and white Primula vulgaris cultivars. Physiol. Mol. Biol. Plants 2019, 25, 1029–1041. [Google Scholar] [CrossRef]
- Soza, V.L.; Lindsley, D.; Waalkes, A.; Ramage, E.; Patwardhan, R.P.; Burton, J.N.; Hall, B. The Rhododendron genome and chromosomal organization provide insight into shared whole-genome duplications across the heath family (Ericaceae). Genome Biol. Evol. 2019, 11, 3353–3371. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Mao, X.; Huang, P.; Fang, S. Morphological characterization of flower buds development and related gene expression profiling at bud break stage in heterodichogamous Cyclocarya paliurus (Batal.) lljinskaja. Genes 2019, 10, 818. [Google Scholar] [CrossRef]
- Han, M.; Yang, C.; Zhou, J.; Zhu, J.; Meng, J.; Shen, T.; Li, H. Analysis of flavonoids and anthocyanin biosynthesis-related genes expression reveals the mechanism of petal color fading of Malus hupehensis (Rosaceae). Braz. J. Bot. 2020, 43, 81–89. [Google Scholar] [CrossRef]


| Primer ID | Gene ID | Forward Primer | Reverse Primer |
|---|---|---|---|
| RhCHS | F01_transcript/61502 | GCTTACCGTCGAGACCGTGG | AACAATGGGCCTCACCAGCC |
| RhCHI | F01_transcript/68926 | GCCGCGTTGGAGCAAATTGT | CCCCCTGGTTCCACCCAAAT |
| RhF3H | F01_transcript/64242 | CTCACTCTCGGCCTCAAGCG | TGCCTCAGGGGCTGGATTCT |
| RhDFR | F01_transcript/64345 | GGCTTCATCGGCTCATGGCT | TCCGCTTTGGGCAACTCCAG |
| RhANS | F01_transcript/40426 | CTCCTTCTCCTCCACCGGCT | GGGGGTTGTTCCAGGTGGTG |
| RhF3’H | F01_transcript/54870 | CGATCCCCACCACTCCATC | AAGAACTGTGCGGCAACCGA |
| RhF3’5’H | F01_transcript/50526 | TGCTGACTACGGCCCGAGAT | GGTATCACCACTGCCTCGCC |
| RhMYB1 | F01_transcript/63169 | ACTCGAGCTGTAGCCCACCA | CGTTCGGAAGACGAGCCTCC |
| RhMYB2 | F01_transcript/66405 | GGCAACTGGAGGTCTCTGCC | CCGGCAGCCTTCCTGCTATG |
| RhbHLH1 | F01_transcript/65059 | CCCTTTCGGCTATGCTGCGA | CCCTTTCGGCTATGCTGCGA |
| RhbHLH2 | F01_transcript/50357 | CTTGGTTGCTTCGGCCTCCA | ACCTCACTCCCTCTCGCCTC |
| Samples | cDNA Size | CCS Number | Read Bases of CCS | Mean Read Length of CCS | Mean Number of Passes |
|---|---|---|---|---|---|
| F01 | 1–6 k | 833,137 | 2,298,485,998 | 2758 | 19 |
| Annotated Databases | Isoform Number |
|---|---|
| COG | 16,102 |
| GO | 18,565 |
| KEGG | 17,450 |
| KOG | 25,787 |
| Pfam | 34,156 |
| Eggnog | 29,498 |
| Swiss-Prot | 38,933 |
| Nr | 39,411 |
| All | 39,521 |
| Databases | Unigene Number | 300~1000 bp | ≥1000 bp | ||
|---|---|---|---|---|---|
| Number | Percentage | Number | Percentage | ||
| NR | 39,411 | 2171 | 5.51% | 37,230 | 94.47% |
| GO | 18,565 | 1521 | 8.19% | 17,040 | 91.79% |
| COG | 16,102 | 800 | 4.97% | 15,300 | 95.02% |
| eggNOG | 38,933 | 2115 | 5.43% | 36,811 | 94.55% |
| KEGG | 17,450 | 1109 | 6.36% | 16,336 | 93.62% |
| KOG | 25,787 | 1249 | 4.84% | 24,534 | 95.14% |
| Pfam | 34,156 | 1699 | 4.97% | 32,454 | 95.02% |
| Swiss-Prot | 29,498 | 1521 | 5.16% | 27,971 | 94.82% |
| All | 39,521 | 2201 | 5.57% | 37,310 | 94.41% |
| No. | Name of Pathway | Pathway ID | No of Transcripts (%) | |
|---|---|---|---|---|
| 1. | Carbon metabolism | ko01200 | 605 | (3.47%) |
| 2. | Protein processing in endoplasmic reticulum | ko04141 | 608 | (3.48%) |
| 3. | Biosynthesis of amino acid | ko01230 | 525 | (3.01%) |
| 4. | Spliceosome | ko03040 | 700 | (4.01%) |
| 5. | Ribosome | ko03010 | 491 | (2.81%) |
| 6. | RNA transport | ko03013 | 558 | (3.20%) |
| 7. | Starch and sucrose metabolism | ko00500 | 377 | (2.16%) |
| 8. | Plant hormone signal transduction | ko04075 | 470 | (2.69%) |
| 9. | Oxidative phosphorylation | ko00190 | 280 | (1.60%) |
| 10. | Glycolysis/gluconeogenesis | ko00010 | 322 | (1.85%) |
| 11. | Plant pathogen interaction | ko04626 | 324 | (1.86%) |
| 12. | mRNA surveillance pathway | ko03015 | 469 | (2.69%) |
| 13. | Ubiquitin mediated proteolysis | ko04120 | 321 | (1.84%) |
| 14. | Amino sugar and nucleotide sugar metabolism | ko00520 | 271 | (1.55%) |
| 15. | Endocytosis | ko04144 | 251 | (1.44%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Q.; Liaquat, F.; He, Y.; Munis, M.F.H.; Zhang, C. Functional Annotation of a Full-Length Transcriptome and Identification of Genes Associated with Flower Development in Rhododendron simsii (Ericaceae). Plants 2021, 10, 649. https://doi.org/10.3390/plants10040649
Liu Q, Liaquat F, He Y, Munis MFH, Zhang C. Functional Annotation of a Full-Length Transcriptome and Identification of Genes Associated with Flower Development in Rhododendron simsii (Ericaceae). Plants. 2021; 10(4):649. https://doi.org/10.3390/plants10040649
Chicago/Turabian StyleLiu, Qunlu, Fiza Liaquat, Yefeng He, Muhammad Farooq Hussain Munis, and Chunying Zhang. 2021. "Functional Annotation of a Full-Length Transcriptome and Identification of Genes Associated with Flower Development in Rhododendron simsii (Ericaceae)" Plants 10, no. 4: 649. https://doi.org/10.3390/plants10040649
APA StyleLiu, Q., Liaquat, F., He, Y., Munis, M. F. H., & Zhang, C. (2021). Functional Annotation of a Full-Length Transcriptome and Identification of Genes Associated with Flower Development in Rhododendron simsii (Ericaceae). Plants, 10(4), 649. https://doi.org/10.3390/plants10040649








