Fungal Infection Induces Anthocyanin Biosynthesis and Changes in DNA Methylation Configuration of Blood Orange [Citrus sinensis L. (Osbeck)]
Abstract
1. Introduction
2. Results
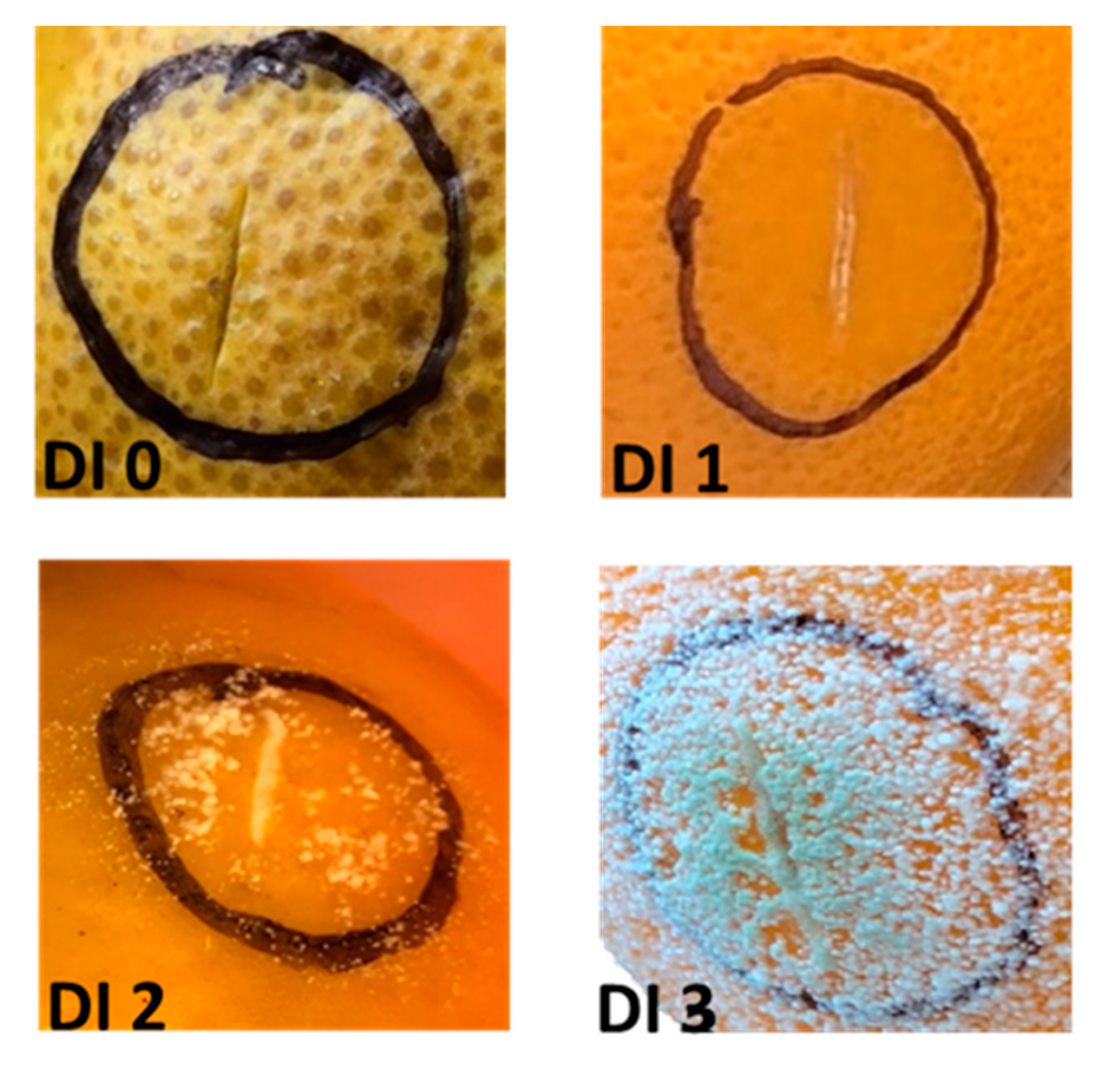
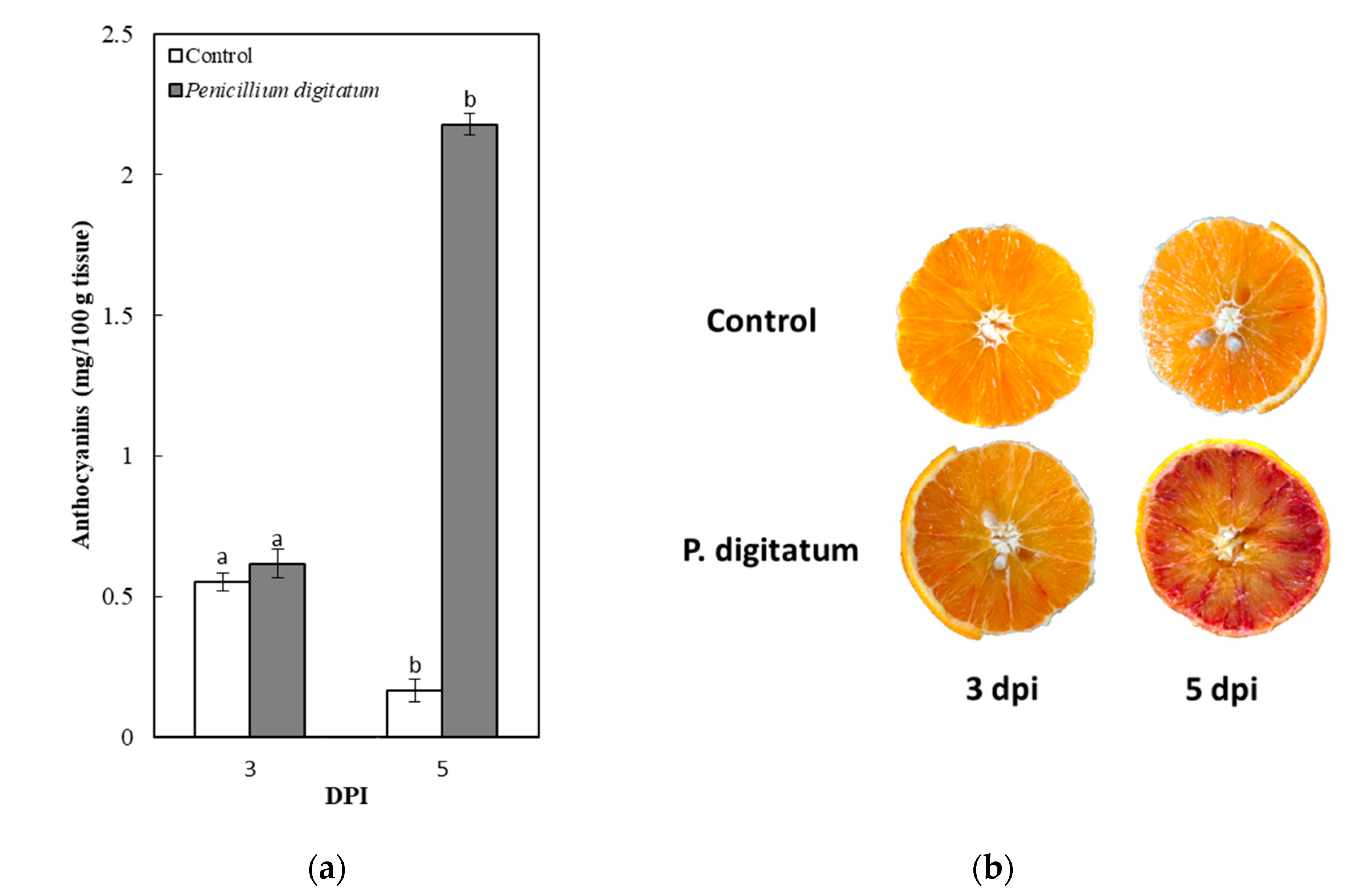
2.1. Fruit Appearance
2.2. Effect of Fungal Inoculation upon the Anthocyanin Content
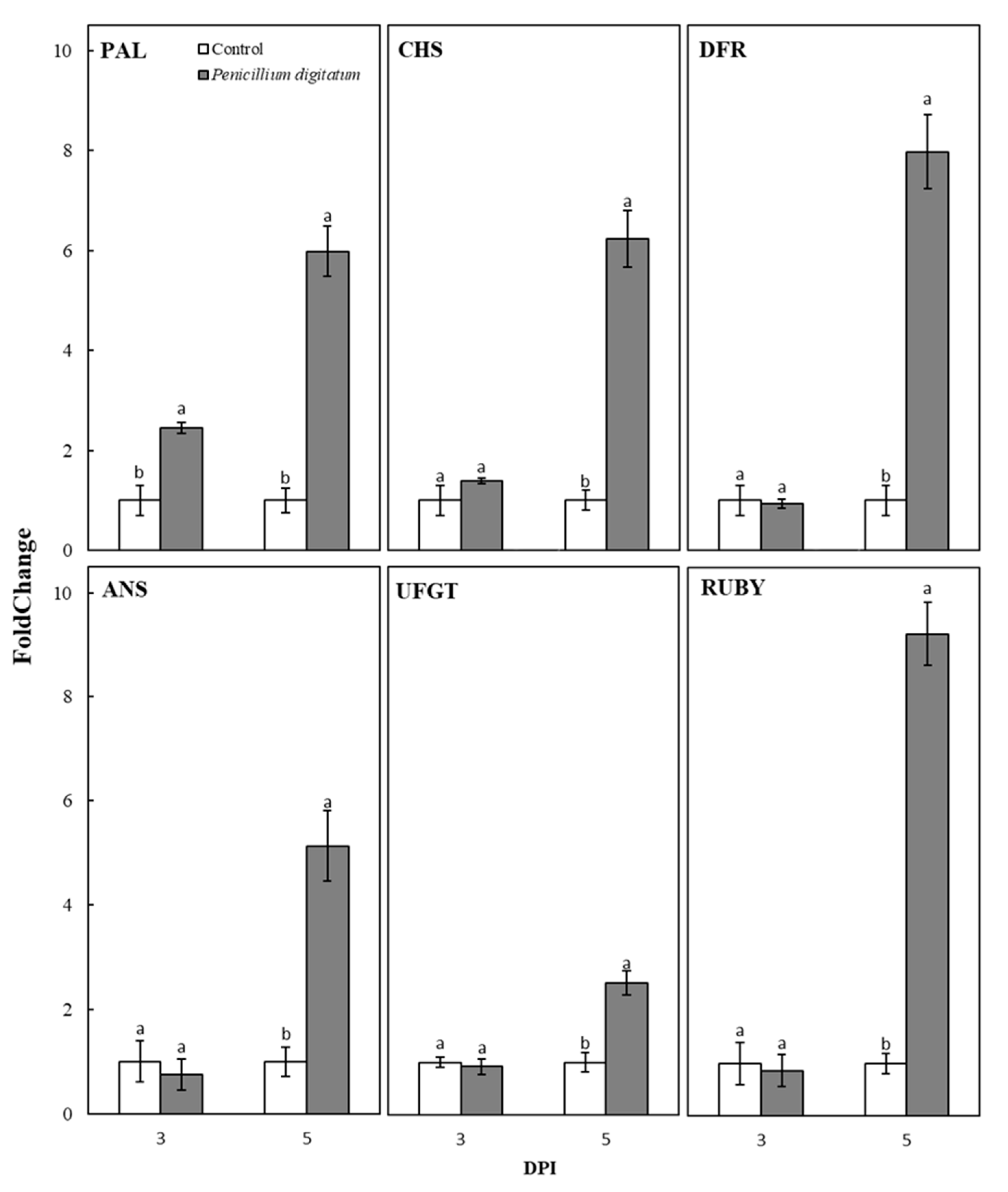
2.3. Analysis of the Expression of Genes Involved in Anthocyanin Biosynthesis
2.4. DNA Methylation Level of Dfr and Ruby Promoter Regions
2.5. Analysis of the Expression of Genes Involved in DNA Demethylation
3. Discussion
4. Materials and Methods
4.1. Fungal Inoculum Preparation
4.2. Fruit Material and Inoculation
4.3. Extraction of Total RNA and cDNA Synthesis
4.4. Measurement of Gene Expression
4.5. Total Anthocyanins Content
4.6. Methylation Sensitive Digestion
4.7. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bendokas, V.; Skemiene, K.; Trumbeckaite, S.; Stanys, V.; Passamonti, S.; Borutaite, V.; Liobikas, J. Anthocyanins: From plant pigments to health benefits at mitochondrial level. Crit. Rev. Food Sci. Nutr. 2019, 60, 3352–3365. [Google Scholar] [CrossRef] [PubMed]
- Lo Piero, A.R. The state of the art in biosynthesis of anthocyanins and its regulation in pigmented sweet oranges [(Citrus sinensis) L. Osbeck]. J. Agric. Food Chem. 2015, 63, 4031–4041. [Google Scholar] [CrossRef] [PubMed]
- Lo Piero, A.R.; Consoli, A.; Puglisi, I.; Orestano, G.; Reforgiato Recupero, G.; Petrone, G. Anthocyaninless cultivars of sweet orange lack to express the UDP-glucose flavonoid 3-O-Glucosyl transferase. J. Plant Biochem. Biotechnol. 2005, 14, 1–6. [Google Scholar] [CrossRef]
- Lo Piero, A.R.; Puglisi, I.; Petrone, G. Gene characterization, analysis of expression and in vitro synthesis of dihydroflavonol 4-reductase from [Citrus sinensis (L.) Osbeck]. Phytochemistry 2006, 67, 684–695. [Google Scholar] [CrossRef] [PubMed]
- Lo Piero, A.R.; Puglisi, I.; Petrone, G. Gene isolation, analysis of expression and in vitro synthesis of a glutathione S-transferase from orange fruit. [Citrus sinensis L. (Osbeck)]. J. Agric. Food Chem. 2006, 54, 9227–9233. [Google Scholar] [CrossRef] [PubMed]
- Ramsay, N.A.; Glover, B.J. MYB-bHLH-WD40 protein complex and the evolution of cellular diversity. Trends Plant Sci. 2005, 10, 63–70. [Google Scholar] [CrossRef]
- Butelli, E.; Licciardello, C.; Zhang, Y.; Liu, J.; Mackay, S.; Bailey, P.; Reforgiato Recupero, G.; Martin, C. Retrotransposons control fruit-specific, cold-dependent accumulation of anthocyanins in blood oranges. Plant Cell 2012, 24, 1242–1255. [Google Scholar] [CrossRef]
- de Pascual-Teresa, S.; Sanchez-Ballesta, M.T. Anthocyanins: From plant to health. Phytochem. Rev. 2008, 7, 281–299. [Google Scholar] [CrossRef]
- Aminifard, M.H.; Mohammadi, S.; Fatemi, H. Inhibition of green mould in blood orange (Citrus sinensis var. Moro) with salicylic acid treatment. Arch. Phytopathol. Plant Prot. 2013, 6, 695–703. [Google Scholar] [CrossRef]
- Lo Piero, A.R.; Puglisi, I.; Rapisarda, P.; Petrone, G. Anthocyanins accumulation and related gene expression in red orange fruit induced by low temperature storage. J. Agric. Food Chem. 2005, 53, 9083–9088. [Google Scholar] [CrossRef]
- Crifò’, T.; Puglisi, I.; Petrone, G.; Reforgiato Recupero, G.; Lo Piero, A.R. Expression analysis in response to low temperature stress in blood oranges: Implication of the flavonoid biosynthetic pathway. Gene 2011, 476, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Crifò, T.; Petrone, G.; Lo Cicero, L.; Lo Piero, A.R. Storage temperature and time influence the anthocyanin content and the level of transcript related to their biosynthesis in blood oranges. J. Agric. Food Chem. 2012, 60, 476–481. [Google Scholar] [CrossRef] [PubMed]
- Lo Piero, A.R.; Lo Cicero, L.; Puglisi, I. The metabolic fate of citric acid as affected by cold storage in blood oranges. J. Plant Biochem. Biotechnol. 2014, 23, 161–166. [Google Scholar] [CrossRef]
- Carmona, L.; Alquézar, B.; Tárraga, S.; Peña, L. Protein analysis of Moro blood orange pulp during storage at low temperatures. Food Chem. 2019, 277, 75–83. [Google Scholar] [CrossRef]
- Iriti, M.; Rossoni, M.; Borgo, M.; Faoro, F. Benzothiadiazole enhances resveratrol and anthocyanin biosynthesis in grapevine, meanwhile improving resistance to Botrytis cinerea. J. Agric. Food Chem. 2004, 52, 4406–4413. [Google Scholar] [CrossRef]
- Zhang, Y.; Butelli, E.; De Stefano, R.; Schoonbeek, H.J.; Magusin, A.; Pagliarani, C.; Wellner, N.; Hill, L.; Orzaez, D.; Granell, A.; et al. Anthocyanins double the shelf life of tomatoes by delaying overripening and reducing susceptibility to gray mold. Curr. Biol. 2013, 23, 1094–1100. [Google Scholar] [CrossRef]
- Sivankalyani, V.; Feygenberg, O.; Diskin, S.; Wright, B.; Alkan, N. Increased anthocyanin and flavonoids in mango fruit peel are associated with cold and pathogen resistance. Postharvest Biol. Technol. 2016, 111, 132–139. [Google Scholar] [CrossRef]
- Zhang, H.; Lang, Z.; Zhu, J.-K. Dynamics and function of DNA methylation in plants. Nat. Rev. Mol. Cell Biol. 2018, 19, 489–506. [Google Scholar] [CrossRef]
- Tirnaz, S.; Batley, J. DNA Methylation: Toward Crop Disease Resistance Improvement. Trends Plant Sci. 2019, 24, 1137–1150. [Google Scholar] [CrossRef]
- Dowen, R.H.; Pelizzola, M.; Schmitz, R.J.; Lister, R.; Dowen, J.M.; Nery, J.R.; Dixon, J.E.; Ecker, J.R. Widespread dynamic DNA methylation in response to biotic stress. Proc. Natl. Acad. Sci. USA 2012, 109, E2183–E2191. [Google Scholar] [CrossRef]
- Deng, B.; Wang, W.; Deng, L.; Yao, S.; Ming, J.; Zeng, K. Comparative RNA-seq analysis of citrus fruit in response to infection with three major postharvest fungi. Postharvest Biol. Technol. 2018, 146, 134–146. [Google Scholar] [CrossRef]
- González-Candelas, L.; Alamar, S.; Sánchez-Torres, P.; Zacarías, L.; Marcos, J.F. A transcriptomic approach highlights induction of secondary metabolism in citrus fruit in response to Penicillium digitatum infection. BMC Plant Biol. 2010, 10, 194–210. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.K. Active DNA demethylation mediated by DNA glycosylases. Annu. Rev. Genet. 2009, 43, 143–166. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Liu, R.; Niu, Q.; Tang, K.; Zhang, B.; Zhang, H.; Chen, K.; Zhu, J.-K.; Lang, Z. Global increase in DNA methylation during orange fruit development and ripening. Proc. Natl. Acad. Sci. USA 2019, 116, 1430–1436. [Google Scholar] [CrossRef]
- Sicilia, A.; Scialò, E.; Puglisi, I.; Lo Piero, A.R. Anthocyanin biosynthesis and DNA methylation dynamics in sweet orange fruit [Citrus sinensis L. (Osbeck)] under cold stress. J. Agric. Food Chem. 2020, 68, 7024–7031. [Google Scholar] [CrossRef] [PubMed]
- Zhong, S.; Fei, Z.; Chen, Y.R.; Zheng, Y.; Huang, M.; Vrebalov, J.; McQuinn, R.; Gapper, N.; Liu, B.; Xiang, J.; et al. Single-base resolution methylomes of tomato fruit development reveal epigenome modifications associated with ripening. Nat. Biotechnol. 2013, 31, 154–159. [Google Scholar] [CrossRef] [PubMed]
- Lang, Z.; Wang, Y.; Tang, K.; Tang, D.; Datsenk, T.; Cheng, J.; Zhang, Y.; Hand, A.K.; Zhu, J.-K. Critical roles of DNA demethylation in the activation of ripening- induced genes and inhibition of ripening- repressed genes in tomato fruit. Proc. Natl. Acad. Sci. USA 2017, 114, E4511–E4519. [Google Scholar] [CrossRef]
- Chen, Z.; Zhao, Y.; Jiang, D.; Xiang, S.; Li, J. Changes in DNA methylation patterns affect ripening time in Satsuma mandarin fruit. Plant Breed. 2019, 138, 967–978. [Google Scholar] [CrossRef]
- Yaish, M.W.; Al-Lawati, A.; Al-Harrasi, I.; Patankar, H.V. Genome-wide DNA methylation analysis in response to salinity in themodel plant caliph medic (Medicago truncatula). BMC Genom. 2018, 19, 78. [Google Scholar] [CrossRef]
- Wang, W.; Pan, Y.; Zhao, X.; Dwivedi, D.; Zhu, L.; Ali, J.; Fu, B.; Li, Z. Drought-induced sitespecific DNA methylation and its association with drought tolerance in rice (Oryza sativa L.). J. Exp. Bot. 2010, 62, 1951–1960. [Google Scholar] [CrossRef]
- Li, Y.; Xi, Q.; Kou, H.; Wang, D.; Lin, X.; Wu, Y.; Xu, C.; Xing, S.; Liu, B. Induced Pib expression and resistance to Magnaporthe grisea are compromised by cytosine demethylation at critical promoter regions in rice. J. Integr. Plant Biol. 2011, 53, 814–823. [Google Scholar] [CrossRef] [PubMed]
- Prall, W.; Sharma, B.; Gregory, B.D. Transcription is just the beginning of gene expression regulation: The functional significance of RNA-binding proteins to post-transcriptional processes in plants. Plant Cell Physiol. 2019, 60, 1939–1952. [Google Scholar] [CrossRef] [PubMed]
- Platania, C.; Restuccia, C.; Muccilli, S.; Cirvilleri, G. Efficacy of killer yeasts in the biological control of Penicillium digitatum on Tarocco orange fruits (Citrus sinensis). Food Microbiol. 2012, 30, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Lo Cicero, L.; Puglisi, I.; Nicolosi, E.; Gentile, A.; Ferlito, F.; Continella, A.; Lo Piero, A.R. Anthocyanin levels and expression analysis of biosynthesis related genes during ripening of sicilian and international grape berries subjected to leaf removal and water deficit. J. Agric. Sci. Technol. 2016, 18, 1333–1344. [Google Scholar]
- Sutherland, E.; Coe, L.; Raleigh, E.A. McrBC: A multisubunit GTP-dependent restriction endonuclease. J. Mol. Biol. 1992, 225, 327–348. [Google Scholar] [CrossRef]
- Irizarry, R.A.; Ladd-Acosta, C.; Carvalho, B.; Wu, H.; Brandenburg, S.A.; Jeddeloh, J.A.; Wen, B.; Feinberg, A.P. Comprehensive high-throughput arrays for relative methylation (CHARM). Genome Res. 2008, 18, 780–790. [Google Scholar] [CrossRef]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sicilia, A.; Catara, V.; Scialò, E.; Lo Piero, A.R. Fungal Infection Induces Anthocyanin Biosynthesis and Changes in DNA Methylation Configuration of Blood Orange [Citrus sinensis L. (Osbeck)]. Plants 2021, 10, 244. https://doi.org/10.3390/plants10020244
Sicilia A, Catara V, Scialò E, Lo Piero AR. Fungal Infection Induces Anthocyanin Biosynthesis and Changes in DNA Methylation Configuration of Blood Orange [Citrus sinensis L. (Osbeck)]. Plants. 2021; 10(2):244. https://doi.org/10.3390/plants10020244
Chicago/Turabian StyleSicilia, Angelo, Vittoria Catara, Emanuele Scialò, and Angela Roberta Lo Piero. 2021. "Fungal Infection Induces Anthocyanin Biosynthesis and Changes in DNA Methylation Configuration of Blood Orange [Citrus sinensis L. (Osbeck)]" Plants 10, no. 2: 244. https://doi.org/10.3390/plants10020244
APA StyleSicilia, A., Catara, V., Scialò, E., & Lo Piero, A. R. (2021). Fungal Infection Induces Anthocyanin Biosynthesis and Changes in DNA Methylation Configuration of Blood Orange [Citrus sinensis L. (Osbeck)]. Plants, 10(2), 244. https://doi.org/10.3390/plants10020244






