Plants Metabolome Study: Emerging Tools and Techniques
Abstract
:1. Metabolomics: Plant Biology Perspective
1.1. Primary Metabolites
| Plant Species | Class | Analytical Tools | Key Metabolites | Reference |
|---|---|---|---|---|
| Primary metabolites | ||||
| Plantago ovata | Fatty acids | GC-MS | α-linolenic acid, linoleic acid and palmitic acid | [10] |
| P. ovata | Fatty acids | GC-MS | Pentadecanoic acid, palmitic acid, heptadecanoic acid, stearic acid, oleic acid, linoleic acid, γ-linolenic acid and arachidic acid | [9] |
| Jatropha curcas | Fatty acids | GC | Oleic acid, palmitic acid and linolenic acid | [29] |
| Paeonia rockii, P. potaninii, and P. lutea | Fatty acids | GC-MS | α-linolenic acid, oleic acid and linoleic acid | [27] |
| Cicer arietinum | Fatty acids | GC-MS | Pentadecanoic acid, palmitic acid, palmitoleic acid, stearic acid, oleic acid, linoleic acid, α-linolenic acid and arachidic acid | [33] |
| P. ovata | Amino acids | HPLC | Isoleucine, threonine, leucine, histidine and lysine | [10] |
| P. ovata | Amino acids | HPLC | Aspartate, glutamine, glycine, alanine, arginine, serine, proline, isoleucine and methionine | [9] |
| Fritillaria thunbergii | Amino acids | GC-MS | Tryptophan, phenylalanine and histidine | [31] |
| C. arietinum | Amino acids | GC-MS | L-glutamic acid, L-tryptophan, phenylalanine, glycine, serine, L-threonine, L-valine, L-ornithine and L-proline | [33] |
| C. arietinum | Sugars and Sugar alcohols | GC-MS | Sucrose, cellobiose, galactose, methylgalactoside, myo-inositol | [33] |
| C. arietinum | Sugar alcohols | GC-QqQ-MS | Galactitol, erythritol, arabitol, xylitol, mannitol and inositol | [32] |
| Secondary metabolites | ||||
| Beta vulgaris | Terpenes | HPLC-MS | Oleanolic acid, hederagenin, akebonoic acid and gypsogenin | [34] |
| Ocimum gratissimum | Terpenes | GC-MS | m-chavicol, t-anethole, germacrene-D, naphthalene, ledene, eucalyptol, azulene and comphore | [35] |
| Mentha piperita | Terpenes | GC-MS | Menthone, menthol, pulegone and menthofuran | [36] |
| M. arvensis | Terpenes | GLC | Menthol, isomenthone, L-methone and menthyl acetate | [37] |
| Achyranthes bidentata | Terpenes | HPLC | Oleanolic acid and ecdysterone | [38] |
| Arabidopsis thaliana | Phenolics | UHPLC-MS | Scopoletin, umbelliferone and esculetin, scopolin, skimmin and esculin | [39] |
| P. ovata | Phenolics | LC-MS | Luteolin, quercetagetin, syringetin, kaempferol, limocitrin, helilupolone and catechin | [10] |
| P. ovata | Phenolics | LC-MS | Kaempferol 3-(2″,3″-diacetylrhamnoside)-7-rhamnoside and apigenin 7-rhamnoside | [9] |
| P. ovata | Alkaloids | LC-MS | Lunamarine, hordatine B and pinidine | [10] |
| Dendrobium Snowflake ‘Red Star’ | Alkaloids | 1H and 2D NMR | Dendrobine and nobilonine | [40] |
1.2. Secondary Metabolites
2. Involvement of Metabolomics in Genetically Modified (GM) Crops
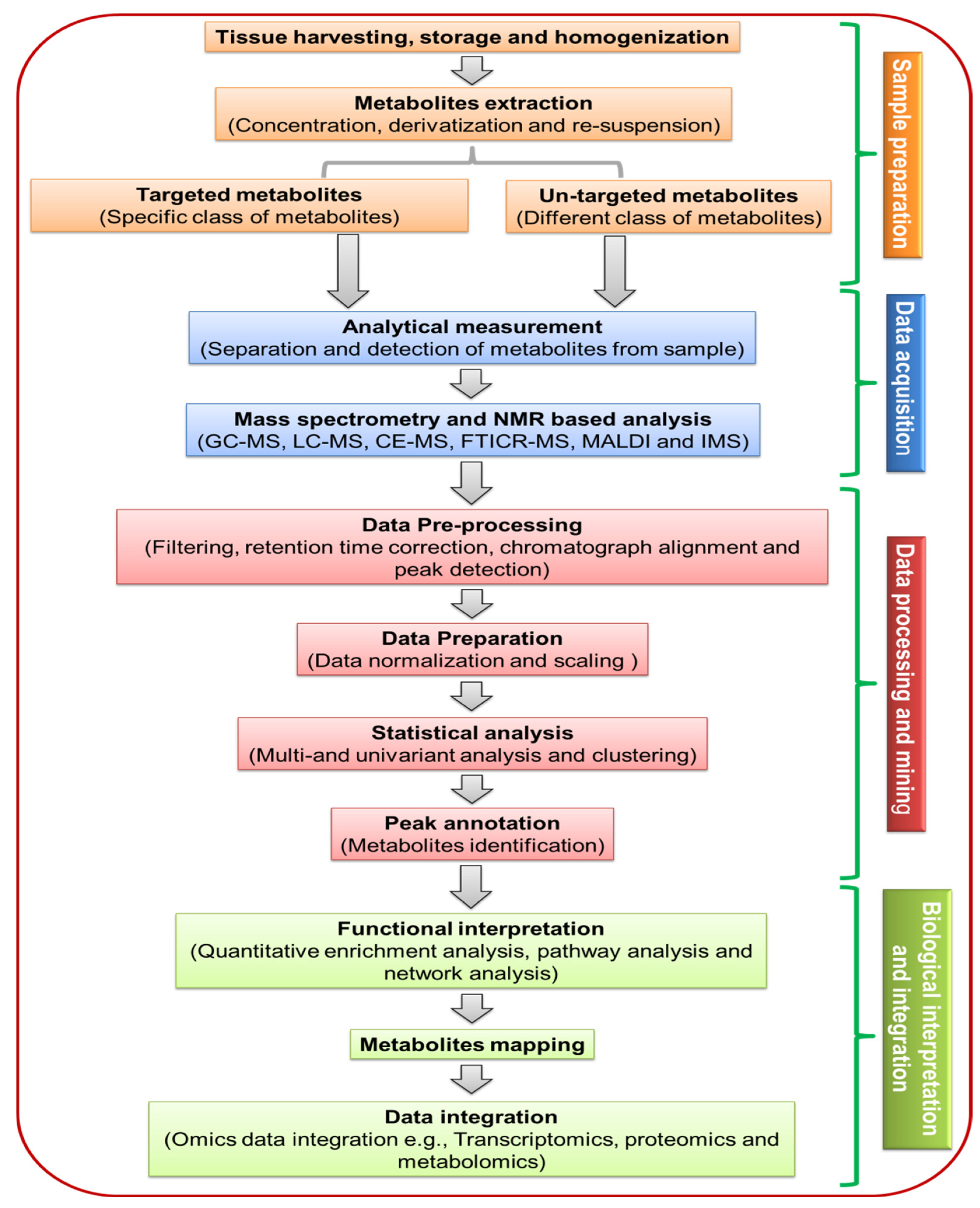
3. Significance of Sample Preparation in Plant Metabolites
3.1. Sample Harvesting and Storage
3.2. Sample Preparation
4. Analytical Techniques Used for Plant Metabolome
4.1. Gas Chromatography-Mass Spectrometry (GC-MS)
4.2. Liquid Chromatography-Mass Spectrometry (LC-MS)
4.3. Capillary Electrophoresis-Mass Spectrometry (CE-MS)
4.4. Fourier Transform ion Cyclotron Resonance-Mass Spectrometry (FTICR-MS)
4.5. Matrix-Assisted Laser Desorption/Ionization (MALDI)
4.6. Ion Mobility Spectrometry (IMS)
4.7. Nuclear Magnetic Resonance (NMR)
5. Metabolomic Data Processing, Annotation, Database and Bioinformatics Tools for Plants METABOLOME Analysis
5.1. Data Processing and Annotation
5.2. Network Analysis
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Razzaq, A.; Sadia, B.; Raza, A.; Khalid Hameed, M.; Saleem, F. Metabolomics: A way forward for crop improvement. Metabolites 2019, 9, 303. [Google Scholar] [CrossRef] [Green Version]
- Peters, K.; Worrich, A.; Weinhold, A.; Alka, O.; Balcke, G.; Birkemeyer, C.; Bruelheide, H.; Calf, O.W.; Dietz, S.; Dührkop, K.; et al. Current challenges in plant eco-metabolomics. Int. J. Mol. Sci. 2018, 19, 1385. [Google Scholar] [CrossRef] [Green Version]
- Hong, J.; Yang, L.; Zhang, D.; Shi, J. Plant Metabolomics: An indispensable system biology tool for plant science. Int. J. Mol. Sci. 2016, 17, 767. [Google Scholar] [CrossRef] [PubMed]
- Shulaev, V.; Cortes, D.; Miller, G.; Mittler, R. Metabolomics for plant stress response. Physiol Plant. 2008, 132, 199–208. [Google Scholar] [CrossRef] [PubMed]
- Piasecka, A.; Kachlicki, P.; Stobiecki, M. Analytical methods for detection of plant metabolomes changes in response to biotic and abiotic stresses. Int. J. Mol. Sci. 2019, 20, 379. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Patel, M.K.; Mishra, A.; Jha, B. Untargeted metabolomics of halophytes. In Marine Omics: Principles and Applications; Kim, S., Ed.; CRC Press: Boca Raton, FL, USA, 2016; pp. 309–325. [Google Scholar]
- Mishra, A.; Patel, M.K.; Jha, B. Non–targeted metabolomics and scavenging activity of reactive oxygen species reveal the potential of Salicornia brachiata as a functional food. J. Funct. Foods 2015, 13, 21–31. [Google Scholar] [CrossRef]
- Pandey, S.; Patel, M.K.; Mishra, A.; Jha, B. Physio-biochemical composition and untargeted metabolomics of cumin (Cuminum cyminum L.) make it promising functional food and help in mitigating salinity stress. PLoS ONE 2015, 10, e0144469. [Google Scholar] [CrossRef] [Green Version]
- Patel, M.K.; Mishra, A.; Jaiswar, S.; Jha, B. Metabolic profiling and scavenging activities of developing circumscissile fruit of psyllium (Plantago ovata Forssk.) reveal variation in primary and secondary metabolites. BMC Plant Biol. 2020, 20, 116. [Google Scholar] [CrossRef]
- Patel, M.K.; Mishra, A.; Jha, B. Non-targeted metabolite profiling and scavenging activity unveil the nutraceutical potential of psyllium (Plantago ovata Forsk). Front. Plant Sci. 2016, 7, 431. [Google Scholar] [CrossRef] [Green Version]
- Bénard, C.; Bernillon, S.; Biais, B.; Osorio, S.; Maucourt, M.; Ballias, P.; Deborde, C.; Colombié, S.; Cabasson, C.; Jacob, D. Metabolomic profiling in tomato reveals diel compositional changes in fruit affected by source–sink relationships. J. Exp. Bot. 2015, 66, 3391–3404. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, Z.; Wang, C.; Zhu, S.; Wang, W.; Xu, J.; Zhao, X. Characterizing the metabolites related to rice salt tolerance with introgression lines exhibiting contrasting performances in response to saline conditions. Plant Growth Regul. 2020, 92, 157–167. [Google Scholar] [CrossRef]
- Francki, M.G.; Hayton, S.; Gummer, J.; Rawlinson, C.; Trengove, R.D. Metabolomic profiling and genomic analysis of wheat aneuploid lines to identify genes controlling biochemical pathways in mature grain. Plant Biotechnol. J. 2016, 14, 649–660. [Google Scholar] [CrossRef] [PubMed]
- Rao, J.; Cheng, F.; Hu, C.; Quan, S.; Lin, H.; Wang, J.; Chen, G.; Zhao, X.; Alexander, D.; Guo, L. Metabolic map of mature maize kernels. Metabolomics 2014, 10, 775–787. [Google Scholar] [CrossRef]
- Kumar, R.; Bohra, A.; Pandey, A.K.; Pandey, M.K.; Kumar, A. Metabolomics for plant improvement: Status and prospects. Front. Plant Sci. 2017, 8, 1302. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tohge, T.; De Souza, L.P.; Fernie, A.R. Genome-enabled plant metabolomics. J. Chromatogr. B 2014, 966, 7–20. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Chen, Z.; Wang, F.; Jia, W.; Xu, Z. Combined transcriptomic and metabolomic analyses uncover rearranged gene expression and metabolite metabolism in tobacco during cold acclimation. Sci. Rep. 2020, 10, 1–13. [Google Scholar]
- Hamany Djande, C.Y.; Pretorius, C.; Tugizimana, F.; Piater, L.A.; Dubery, I.A. Metabolomics: A tool for cultivar phenotyping and investigation of grain crops. Agronomy 2020, 10, 831. [Google Scholar] [CrossRef]
- Patel, M.K.; Kumar, M.; Li, W.; Luo, Y.; Burritt, D.J.; Alkan, N.; Tran, L.-S.P. Enhancing salt tolerance of plants: From metabolic reprogramming to exogenous chemical treatments and molecular approaches. Cells 2020, 9, 2492. [Google Scholar] [CrossRef]
- Rupasinghe, T.W.; Roessner, U. Extraction of plant lipids for LC-MS-based untargeted plant lipidomics. Plant Metab. 2018, 1778, 125–135. [Google Scholar]
- Shulaev, V.; Chapman, K.D. Plant lipidomics at the crossroads: From technology to biology driven science. BBA––Mol. Cell. Biol. Lipids 2017, 1862, 786–791. [Google Scholar] [CrossRef]
- Kofeler, H.C.; Fauland, A.; Rechberger, G.N.; Trötzmüller, M. Mass spectrometry based lipidomics: An overview of technological platforms. Metabolites 2012, 2, 19–38. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ni, Z.; Milic, I.; Fedorova, M. Identification of carbonylated lipids from different phospholipid classes by shotgun and LC-MS lipidomics. Anal. Bioanal. Chem. 2015, 407, 5161–5173. [Google Scholar] [CrossRef]
- Okazaki, Y.; Kamide, Y.; Hirai, M.Y.; Saito, K. Plant lipidomics based on hydrophilic interaction chromatography coupled to ion trap time-of-flight mass spectrometry. Metabolomics 2013, 9, 121–131. [Google Scholar] [CrossRef] [Green Version]
- Abbadi, A.; Domergue, F.; Bauer, J.; Napier, J.A.; Welti, R.; Zähringer, U.; Cirpus, P.; Heinz, E. Biosynthesis of very-long-chain polyunsaturated fatty acids in transgenic oilseeds: Constraints on their accumulation. Plant Cell 2004, 16, 2734–2748. [Google Scholar] [CrossRef] [Green Version]
- Yu, D.; Boughton, B.A.; Hill, C.B.; Feussner, I.; Roessner, U.; Rupasinghe, T.W. Insights into oxidized lipid modification in barley roots as an adaptation mechanism to salinity stress. Front. Plant Sci. 2020, 11, 1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Q.Y.; Yu, R.; Xie, L.H.; Rahman, M.M.; Kilaru, A.; Niu, L.X.; Zhang, Y.L. Fatty acid and associated gene expression analyses of three tree peony species reveal key genes for α-linolenic acid synthesis in seeds. Front. Plant Sci. 2018, 9, 106. [Google Scholar] [CrossRef] [Green Version]
- Patel, M.K.; Pandey, S.; Brahmbhatt, H.R.; Mishra, A.; Jha, B. Lipid content and fatty acid profile of selected halophytic plants reveal a promising source of renewable energy. Biomass Bioenergy 2019, 124, 25–32. [Google Scholar] [CrossRef]
- Sinha, P.; Islam, M.A.; Negi, M.S.; Tripathi, S.B. Changes in oil content and fatty acid composition in Jatropha curcas during seed development. Ind. Crops. Prod. 2015, 77, 508–510. [Google Scholar] [CrossRef]
- Nimbalkar, M.S.; Pai, S.R.; Pawar, N.V.; Oulkar, D.; Dixit, G.B. Free amino acid profiling in grain Amaranth using LC–MS/MS. Food Chem. 2012, 134, 2565–2569. [Google Scholar] [CrossRef]
- Cui, M.C.; Chen, S.J.; Wang, H.H.; Li, Z.H.; Chen, H.J.; Chen, Y.; Zhou, H.B.; Li, X.; Chen, J.W. Metabolic profiling investigation of Fritillaria thunbergii Miq. by gas chromatography–mass spectrometry. J. Food Drug Anal. 2018, 26, 337–347. [Google Scholar] [CrossRef] [Green Version]
- Dias, D.A.; Hill, C.B.; Jayasinghe, N.S.; Atieno, J.; Sutton, T.; Roessner, U. Quantitative profiling of polar primary metabolites of two chickpea cultivars with contrasting responses to salinity. J. Chromatogr. B. 2015, 1000, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Pandey, S.; Kumari, A.; Shree, M.; Kumar, V.; Singh, P.; Bharadwaj, C.; Loake, G.J.; Parida, S.K.; Masakapalli, S.K.; Gupta, K.J. Nitric oxide accelerates germination via the regulation of respiration in chickpea. J. Exp. Bot. 2019, 70, 4539–4555. [Google Scholar] [CrossRef]
- Mikołajczyk-Bator, K.; Błaszczyk, A.; Czyżniejewski, M.; Kachlicki, P. Characterization and identification of triterpene saponins in the roots of red beets (Beta vulgaris L.) using two HPLC–MS systems. Food Chem. 2016, 192, 979–990. [Google Scholar] [CrossRef] [PubMed]
- Hazzoumi, Z.; Moustakime, Y.; Joutei, K.A. Effect of gibberellic acid (GA), indole acetic acid (IAA) and benzylaminopurine (BAP) on the synthesis of essential oils and the isomerization of methyl chavicol and trans-anethole in Ocimum gratissimum L. SpringerPlus 2014, 3, 321–327. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Santoro, M.V.; Nievas, F.; Zygadlo, J.; Giordano, W.; Banchio, E. Effects of growth regulators on biomass and the production of secondary metabolites in peppermint (Mentha piperita) micropropagated in vitro. Am. J. Plant Sci. 2013, 4, 49. [Google Scholar] [CrossRef] [Green Version]
- Naeem, M.; Khan, M.M.A.; Idrees, M.; Aftab, T. Triacontanol-mediated regulation of growth yield, physiological activities and active constituents of Mentha arvensis L. Plant Growth Regul. 2011, 65, 195–206. [Google Scholar] [CrossRef]
- Li, J.T.; Hu, Z.H. Accumulation and dynamic trends of triterpenoid saponin in vegetative organ of Achyranthus bidentata. J. Integr. Plant Biol. 2009, 51, 122–129. [Google Scholar] [CrossRef]
- Perkowska, I.; Siwinska, J.; Olry, A.; Grosjean, J.; Hehn, A.; Bourgaud, F.; Lojkowska, E.; Ihnatowicz, A. Identification and quantification of coumarins by UHPLC-MS in Arabidopsis thaliana natural populations. Molecules 2021, 26, 1804. [Google Scholar] [CrossRef]
- Morita, H.; Fujiwara, M.; Yoshida, N.; Kobayashi, J. New picrotoxin-type and dendrobine-type sesquiterpenoids from Dendrobium snowflake ‘Red Star’. Tetrahedron 2000, 56, 5801–5805. [Google Scholar] [CrossRef]
- Zagorchev, L.; Seal, C.E.; Kranner, I.; Odjakova, M. A central role for thiols in plant tolerance to abiotic stress. Int. J. Mol. Sci. 2013, 14, 7405–7432. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, J.; Yu, Y.; Shi, R.; Xie, G.; Zhu, Y.; Wu, G.; Qin, M. Organ-specific metabolic shifts of flavonoids in Scutellaria baicalensis at different growth and development stages. Molecules 2018, 23, 428. [Google Scholar] [CrossRef] [Green Version]
- Lin, L.; Huang, X.; Lv, Z. Isolation and identification of flavonoids components from Pteris vittata L. SpringerPlus 2016, 5, 1649. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, J.; Ma, C.; Xu, H.; Yuan, K.; Lu, X.; Zhu, Z.; Wu, Y.; Xu, G. Metabolic profiling of transgenic rice with cryIAc and sck genes: An evaluation of unintended effects at metabolic level by using GC-FID and GC–MS. J. Chromatogr. B. 2009, 877, 725–732. [Google Scholar] [CrossRef]
- Iwaki, T.; Guo, L.; Ryals, J.A.; Yasuda, S.; Shimazaki, T.; Kikuchi, A.; Watanabe, K.N.; Kasuga, M.; Yamaguchi-Shinozaki, K.; Ogawa, T.; et al. Metabolic profiling of transgenic potato tubers expressing Arabidopsis dehydration response element-binding protein 1A (DREB1A). J. Agric. Food Chem. 2013, 61, 893–900. [Google Scholar] [CrossRef]
- Ma, C.; Wang, H.; Lu, X.; Wang, H.; Xu, G.; Liu, B. Terpenoid metabolic profiling analysis of transgenic Artemisia annua L. by comprehensive two-dimensional gas chromatography time-of-flight mass spectrometry. Metabolomics 2009, 5, 497–506. [Google Scholar] [CrossRef]
- Sobolev, A.P.; Testone, G.; Santoro, F.; Nicolodi, C.; Iannelli, M.A.; Amato, M.E.; Ianniello, A.; Brosio, E.; Giannino, D.; Mannina, L. Quality traits of conventional and transgenic lettuce (Lactuca sativa L.) at harvesting by NMR metabolic profiling. J. Agric. Food Chem. 2010, 58, 6928–6936. [Google Scholar] [CrossRef]
- Roessner-Tunali, U.; Hegemann, B.; Lytovchenko, A.; Carrari, F.; Bruedigam, C.; Granot, D.; Fernie, A.R. Metabolic profiling of transgenic tomato plants overexpressing hexokinase reveals that the influence of hexose phosphorylation diminishes during fruit development. Plant Physiol. 2003, 133, 84–99. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Choi, H.K.; Choi, Y.H.; Verberne, M.; Lefeber, A.W.; Erkelens, C.; Verpoorte, R. Metabolic fingerprinting of wild type and transgenic tobacco plants by 1H NMR and multivariate analysis technique. Phytochemistry 2004, 65, 857–864. [Google Scholar] [CrossRef]
- Jha, R.K.; Patel, J.; Patel, M.K.; Mishra, A.; Jha, B. Introgression of a novel cold and drought regulatory-protein encoding CORA-like gene, SbCDR, induced osmotic tolerance in transgenic tobacco. Physiol. Plant 2021, 172, 1170–1188. [Google Scholar] [CrossRef]
- Chang, Y.; Zhao, C.; Zhu, Z.; Wu, Z.; Zhou, J.; Zhao, Y.; Lu, X.; Xu, G. Metabolic profiling based on LC/MS to evaluate unintended effects of transgenic rice with cry1Ac and sck genes. Plant Mol. Biol. 2012, 78, 477–487. [Google Scholar] [CrossRef] [PubMed]
- Payyavula, R.S.; Tschaplinski, T.J.; Jawdy, S.S.; Sykes, R.W.; Tuskan, G.A.; Kalluri, U.C. Metabolic profiling reveals altered sugar and secondary metabolism in response to UGPase overexpression in Populus. BMC Plant Boil. 2014, 14, 1–14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stamova, B.S.; Roessner, U.; Suren, S.; Laudencia-Chingcuanco, D.; Bacic, A.; Beckles, D.M. Metabolic profiling of transgenic wheat over-expressing the high-molecular-weight Dx5 glutenin subunit. Metabolomics 2009, 5, 239–252. [Google Scholar] [CrossRef]
- Niu, F.; Jiang, Q.; Sun, X.; Hu, Z.; Wang, L.; Zhang, H. Metabolic profiling of DREB-overexpressing transgenic wheat seeds by liquid chromatography–mass spectrometry. Crop J. 2020, 8, 1025–1036. [Google Scholar] [CrossRef]
- Piccioni, F.; Capitani, D.; Zolla, L.; Mannina, L. NMR metabolic profiling of transgenic maize with the Cry1A(b) gene. J. Agric. Food Chem. 2020, 57, 6041–6049. [Google Scholar] [CrossRef]
- Tanna, B.; Mishra, A. Metabolomics of seaweeds: Tools and techniques. In Plant Metabolites and Regulation Under Environmental Stress; Ahmad, P., Ahanger, M.A., Singh, V.P., Tripathi, D.K., Alam, P., Alyemeni, M.N., Eds.; Academic Press: Cambridge, MA, USA, 2018; pp. 37–52. [Google Scholar]
- Salem, M.A.; Perez de Souza, L.; Serag, A.; Fernie, A.R.; Farag, M.A.; Ezzat, S.M.; Alseekh, S. Metabolomics in the context of plant natural products research: From sample preparation to metabolite analysis. Metabolites 2020, 10, 37. [Google Scholar] [CrossRef] [Green Version]
- Sumner, L.W.; Amberg, A.; Barrett, D.; Beale, M.H.; Beger, R.; Daykin, C.A.; Fan, T.W.M.; Fiehn, O.; Goodacre, R.; Griffin, J.L.; et al. Proposed minimum reporting standards for chemical analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics 2007, 3, 211–221. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harbourne, N.; Marete, E.; Jacquier, J.C.; O’Riordan, D. Effect of drying methods on the phenolic constituents of meadowsweet (Filipendula ulmaria) and willow (Salix alba). LWT––Food Sci. Technol. 2009, 42, 1468–1473. [Google Scholar] [CrossRef]
- Parida, A.K.; Panda, A.; Rangani, J. Metabolomics-guided elucidation of abiotic stress tolerance mechanisms in plants. In Plant Metabolites and Regulation Under Environmental Stress; Ahmad, P., Ahanger, M.A., Singh, V.P., Tripathi, D.K., Alam, P., Alyemeni, M.N., Eds.; Academic Press: Cambridge, MA, USA, 2018; pp. 89–131. [Google Scholar]
- Gong, Z.G.; Hu, J.; Wu, X.; Xu, Y.J. The recent developments in sample preparation for mass spectrometry-based metabolomics. Crit. Rev. Anal. Chem. 2017, 8347, 1–7. [Google Scholar] [CrossRef]
- Silva-Navas, J.; Moreno-Risueno, M.A.; Manzano, C. Flavonols mediate root phototropism and growth through regulation of proliferation-to differentiation transition. Plant Cell 2016, 28, 1372–1387. [Google Scholar] [CrossRef] [Green Version]
- Corrales, A.R.; Carrillo, L.; Lasierra, P. Multifaceted role of cycling DOF factor 3 (CDF3) in the regulation of flowering time and abiotic stress responses in Arabidopsis. Plant Cell Environ. 2017, 40, 748–764. [Google Scholar] [CrossRef]
- Sánchez-Parra, B.; Frerigmann, H.; Pérez Alonso, M.-M. Characterization of four bifunctional plant IAM/PAM-amidohydrolases capable of contributing to auxin biosynthesis. Plants 2014, 3, 324–347. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lehmann, T.; Janowitz, T.; Sánchez-Parra, B. Arabidopsis NITRILASE 1 contributes to the regulation of root growth and development through modulation of auxin biosynthesis in seedlings. Front. Plant Sci. 2017, 8, 36. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- T’Kindt, R.; Morreel, K.; Deforce, D. Joint GC–MS and LC–MS platforms for comprehensive plant metabolomics: Repeatability and sample pre-treatment. J. Chromatogr. B 2009, 877, 3572–3580. [Google Scholar] [CrossRef]
- Giavalisco, P.; Li, Y.; Matthes, A. Elemental formula annotation of polar and lipophilic metabolites using (13) C, (15) N and (34) S isotope labelling, in combination with high-resolution mass spectrometry. Plant J. 2011, 68, 364–376. [Google Scholar] [CrossRef]
- Yuliana, N.D.; Khatib, A.; Verpoorte, R.; Choi, Y.H. Comprehensive extraction method integrated with NMR metabolomics: A new bioactivity screening method for plants, adenosine a1 receptor binding compounds in Orthosiphon stamineus, Benth. Anal. Chem. 2011, 83, 6902–6906. [Google Scholar] [CrossRef]
- Gratacós-Cubarsí, M.; Ribas-Agustí, A.; García-Regueiro, J.A.; Castellari, M. Simultaneous evaluation of intact glucosinolates and phenolic compounds by UPLC-DAD-MS/MS in Brassica oleracea L. var. botrytis. Food Chem. 2010, 121, 257–263. [Google Scholar] [CrossRef]
- Teo, C.C.; Chong, W.P.K.; Ho, Y.S. Development and application of microwave-assisted extraction technique in biological sample preparation for small molecule analysis. Metabolomics 2013, 9, 1109–1128. [Google Scholar] [CrossRef]
- Altemimi, A.; Watson, D.G.; Choudhary, R.; Dasari, M.R.; Lightfoot, D.A. Ultrasound assisted extraction of phenolic compounds from peaches and pumpkins. PLoS ONE 2016, 11, e0148758. [Google Scholar] [CrossRef] [Green Version]
- Velickovic, D.; Chu, R.K.; Myers, G.L.; Ahkami, A.H.; Anderton, C.R. An approach for visualizing the spatial metabolome of an entire plant root system inspired by the swiss-rolling technique. J. Mass Spectrom. 2020, 55, 4363. [Google Scholar] [CrossRef]
- Zuorro, A.; Lavecchia, R.; Medici, F.; Piga, L. Enzyme-assisted production of tomato seed oil enriched with lycopene from tomato pomace. Food Bioprocess Tech. 2013, 6, 3499–3509. [Google Scholar] [CrossRef]
- Zhou, J.; Yin, Y. Strategies for large-scale targeted metabolomics quantification by liquid chromatography-mass spectrometry. Analyst 2016, 141, 6362–6373. [Google Scholar] [CrossRef]
- Cajka, T.; Fiehn, O. Toward merging untargeted and targeted methods in mass spectrometry-based metabolomics and lipidomics. Anal. Chem. 2016, 88, 524–545. [Google Scholar] [CrossRef] [PubMed]
- Bojko, B.; Reyes-Garcés, N.; Bessonneau, V.; Goryński, K.; Mousavi, F.; Silva, E.A.S.; Pawliszyn, J. Solid-phase microextraction in metabolomics. Trends Analyt. Chem. 2014, 61, 168–180. [Google Scholar] [CrossRef]
- Ciccimaro, E.; Blair, I.A. Stable-isotope dilution LC–MS for quantitative biomarker analysis. Bioanalysis 2010, 2, 311–341. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, J.; Liu, H.; Liu, Y.; Liu, J.; Zhao, X.; Yin, Y. Development and evaluation of a parallel reaction monitoring strategy for large-scale targeted metabolomics quantification. Anal. Chem. 2016, 88, 4478. [Google Scholar] [CrossRef] [PubMed]
- Guo, B.; Chen, B.; Liu, A.; Zhu, W.; Yao, S. Liquid chromatography-mass spectrometric multiple reaction monitoring-based strategies for expanding targeted profiling towards quantitative metabolomics. Curr. Drug Metab. 2012, 13, 1226–1243. [Google Scholar] [CrossRef]
- Bianchi, F.; Ilag, L.; Termopoli, V.; Mendez, L. Advances in MS-based analytical methods: Innovations and future trends. J. Anal. Methods Chem. 2018, 2018, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Fiehn, O. Metabolomics by gas chromatography–mass spectrometry: Combined targeted and untargeted profiling. Curr. Protoc. Mol. Biol. 2016, 114, 1–32. [Google Scholar] [CrossRef]
- Kopka, J. Current challenges and developments in GC–MS based metabolite profiling technology. J. Biotechnol. 2006, 124, 312–322. [Google Scholar] [CrossRef]
- Harvey, D.J.; Vouros, P. Mass spectrometric fragmentation of trimethylsilyl and related alkylsilyl derivatives. Mass Spectrom. Rev. 2020, 39, 105–211. [Google Scholar] [CrossRef]
- Jorge, T.F.; Rodrigues, J.A.; Caldana, C.; Schmidt, R.; Van Dongen, J.T.; Thomas-Oates, J.; António, C. Mass spectrometry-based plant metabolomics: Metabolite responses to abiotic stress. Mass Spectrom. Rev. 2016, 35, 620–649. [Google Scholar] [CrossRef]
- Hall, R.D. Plant metabolomics: From holistic hope, to hype, to hot topic. New Phytol. 2006, 169, 453–468. [Google Scholar] [CrossRef]
- Kumar, M.; Kuzhiumparambil, U.; Pernice, M.; Jiang, Z.; Ralph, P.J. Metabolomics: An emerging frontier of systems biology in marine macrophytes. Algal Res. 2016, 16, 76–92. [Google Scholar] [CrossRef]
- Tsugawa, H.; Tsujimoto, Y.; Arita, M.; Bamba, T.; Fukusaki, E.; Fiehn, O. GC/MS based metabolomics: Development of a data mining system for metabolite identification by using soft independent modeling of class analogy (simca). BMC Bioinform. 2011, 12, 131. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koek, M.M.; Jellema, R.H.; Van der Greef, J.; Tas, A.C.; Hankemeier, T. Quantitative metabolomics based on gas chromatography mass spectrometry: Status and perspectives. Metabolomics 2011, 7, 307–328. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mastrangelo, A.; Ferrarini, A.; Rey-Stolle, F.; García, A.; Barbas, C. From sample treatment to biomarker discovery: A tutorial for untargeted metabolomics based on GC-(EI)-Q-MS. Anal. Chim. Acta 2015, 900, 21–35. [Google Scholar] [CrossRef] [PubMed]
- Beale, D.J.; Pinu, F.R.; Kouremenos, K.A.; Poojary, M.M.; Narayana, V.K.; Boughton, B.A.; Dias, D.A. Review of recent developments in GC–MS approaches to metabolomics-based research. Metabolomics 2018, 14, 1–31. [Google Scholar] [CrossRef] [PubMed]
- Sissener, N.H.; Hemre, G.-I.; Lall, S.P.; Sagstad, A.; Petersen, K.; Williams, J.; Rohloff, J.; Sanden, M. Are apparent negative effects of feeding genetically modified MON810 maize to Atlantic salmon, Salmo salar caused by confounding factors? Br. J. Nutr. 2011, 106, 42–56. [Google Scholar] [CrossRef]
- Matsuda, F.; Hirai, M.Y.; Sasaki, E.; Akiyama, K.; Yonekura-Sakakibara, K.; Provart, N.J.; Sakurai, T.; Shimada, Y.; Saito, K. AtMetExpress development: A phytochemical atlas of Arabidopsis development. Plant Physiol. 2010, 152, 566–578. [Google Scholar] [CrossRef] [Green Version]
- Obata, T.; Fernie, A.R. The use of metabolomics to dissect plant responses to abiotic stresses. Cell Mol. Life Sci. 2012, 69, 3225–3243. [Google Scholar] [CrossRef] [Green Version]
- Okazaki, Y.; Saito, K. Recent advances of metabolomics in plant biotechnology. Plant Biotechnol. Rep. 2012, 6, 1–15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Holčapek, M.; Jirásko, R.; Lísa, M. Recent developments in liquid chromatography–mass spectrometry and related techniques. J. Chromatogr. 2012, 1259, 3–15. [Google Scholar] [CrossRef] [PubMed]
- Allwood, J.W.; Goodacre, R. An introduction to liquid chromatography–mass spectrometry instrumentation applied in plant metabolomic analyses. Phytochem. Anal. 2010, 21, 33–47. [Google Scholar] [CrossRef]
- Schiffmann, C.; Hansen, R.; Baumann, S.; Kublik, A.; Nielsen, P.H.; Adrian, L.; Von Bergen, M.; Jehmlich, N.; Seifert, J. Comparison of targeted peptide quantification assays for reductive dehalogenases by selective reaction monitoring (SRM) and precursor reaction monitoring (PRM). Anal. Bioanal. Chem. 2014, 406, 283–291. [Google Scholar] [CrossRef]
- Last, R.L.; Jones, A.D.; Shachar-Hill, Y. Towards the plant metabolome and beyond. Nat. Rev. Mol. Cell Biol. 2007, 8, 167–174. [Google Scholar] [CrossRef] [PubMed]
- Alseekh, S.; Fernie, A.R. Metabolomics 20 years on: What have we learned and what hurdles remain? Plant J. 2018, 94, 933–942. [Google Scholar] [CrossRef] [PubMed]
- Soga, T.; Imaizumi, M. Capillary electrophoresis method for the analysis of inorganic anions, organic acids, amino acids, nucleotides, carbohydrates and other anionic compounds. Electrophoresis 2001, 22, 3418–3425. [Google Scholar] [CrossRef]
- Williams, B.J.; Cameron, C.J.; Workman, R.; Broeckling, C.D.; Sumner, L.W.; Smith, J.T. Amino acid profiling in plant cell cultures: An inter-laboratory comparison of CE-MS and GC-MS. Electrophoresis 2007, 28, 1371–1379. [Google Scholar] [CrossRef]
- Ren, J.L.; Zhang, A.H.; Kong, L.; Wang, X.J. Advances in mass spectrometry-based metabolomics for investigation of metabolites. RSC Adv. 2018, 8, 22335–22350. [Google Scholar] [CrossRef] [Green Version]
- Nikolaev, E.N.; Kostyukevich, Y.I.; Vladimirov, G.N. Fourier transform ion cyclotron resonance (FTICR) mass spectrometry: Theory and simulations. Mass Spectrom. Rev. 2016, 35, 219–258. [Google Scholar] [CrossRef] [PubMed]
- Baker, T.C.; Han, J.; Borchers, C.H. Recent advancements in matrix-assisted laser desorption/ionization mass spectrometry imaging. Curr. Opin. Biotechnol. 2017, 43, 62–69. [Google Scholar] [CrossRef]
- Cha, S.; Zhang, H.; Ilarslan, H.I.; Wurtele, E.S.; Brachova, L.; Nikolau, B.J.; Yeung, E.S. Direct profiling and imaging of plant metabolites in intact tissues by using colloidal graphite-assisted laser desorption ionization mass spectrometry. Plant J. 2008, 55, 348–360. [Google Scholar] [CrossRef] [PubMed]
- Jun, J.H.; Song, Z.; Liu, Z.; Nikolau, B.J.; Yeung, E.S.; Lee, Y.J. High-spatial and high-mass resolution imaging of surface metabolites of Arabidopsis thaliana by laser desorption-ionization mass spectrometry using colloidal silver. Anal. Chem. 2010, 82, 3255–3265. [Google Scholar] [CrossRef]
- Goodwin, R.J.; Pennington, S.R.; Pitt, A.R. Protein and peptides in pictures: Imaging with MALDI mass spectrometry. Proteomics 2008, 8, 3785–3800. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.J.; Perdian, D.C.; Song, Z.; Yeung, E.S.; Nikolau, B.J. Use of mass spectrometry for imaging metabolites in plants. Plant J. 2012, 70, 81–95. [Google Scholar] [CrossRef] [Green Version]
- Kathirvel, S.; Gayatri Ramya, M.; Rajesh, A. An overview on the benefits and applications of high performance ion mobility spectrometer in pharmaceutical arena-focus on current research. World J. Pharm. Pharm. Sci. 2017, 6, 402–406. [Google Scholar]
- Hernandez-Mesa, M.; Escourrou, A.; Monteau, F.; Le Bizec, B.; Dervilly-Pinel, G. Current applications and perspectives of ion mobility spectrometry to answer chemical food safety issues. Trends Anal. Chem. 2017, 94, 39–53. [Google Scholar] [CrossRef]
- Campuzano, I.D.G.; Lippens, J.L. Ion mobility in the pharmaceutical industry: An established biophysical technique or still niche? Curr. Opin. Chem. Biol. 2018, 42, 147–159. [Google Scholar] [CrossRef]
- Burnum-Johnson, K.E.; Zheng, X.; Dodds, J.N.; Ash, J.; Fourches, D.; Nicora, C.D.; Wendler, J.P.; Metz, T.O.; Waters, K.M.; Jansson, J.K.; et al. Ion mobility spectrometry and the omics: Distinguishing isomers, molecular classes and contaminant ions in complex samples. Trends Anal. Chem. 2019, 116, 292–299. [Google Scholar] [CrossRef]
- Odenkirk, M.T.; Baker, E.S. Utilizing drift tube ion mobility spectrometry for the evaluation of metabolites and xenobiotics. Methods Mol. Biol. 2020, 2084, 35–54. [Google Scholar]
- Armenta, S.; Esteve-Turrillas, F.A.; Alcala, M. Analysis of hazardous chemicals by “stand alone” drift tube ion mobility spectrometry: A review. Anal. Methods 2020, 12, 1163–1181. [Google Scholar] [CrossRef]
- Garcia, X.; Sabaté, M.D.M.; Aubets, J.; Jansat, J.M.; Sentellas, S. Ion mobility–mass spectrometry for bioanalysis. Separations 2021, 8, 33. [Google Scholar] [CrossRef]
- May, J.C.; Morris, C.B.; McLean, J.A. Ion mobility collision cross section compendium. Anal. Chem. 2017, 89, 1032–1044. [Google Scholar] [CrossRef] [Green Version]
- Mu, Y.; Schulz, B.L.; Ferro, V. Applications of ion mobility-mass spectrometry in carbohydrate chemistry and glycobiology. Molecules 2018, 23, 2557. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hofmann, J.; Pagel, K. Glycan analysis by ion mobility-mass spectrometry. Angew. Chem. Int. Ed. 2017, 56, 8342–8349. [Google Scholar] [CrossRef]
- Li, H.; Bendiak, B.; Kaplan, K.; Davis, E.; Siems, W.F.; Hill, H.H. Evaluation of ion mobility-mass spectrometry for determining the isomeric heterogeneity of oligosaccharide-alditols derived from bovine submaxillary mucin. Int. J. Mass Spectrom. 2013, 352, 9–18. [Google Scholar] [CrossRef] [Green Version]
- Ahonen, L.; Fasciotti, M.; Gennäs, G.B.A.; Kotiaho, T.; Daroda, R.J.; Eberlin, M.; Kostiainen, R. Separation of steroid isomers by ion mobility mass spectrometry. J. Chromatogr. A. 2013, 1310, 133–137. [Google Scholar] [CrossRef] [PubMed]
- Clowers, B.H.; Dwivedi, P.; Steiner, W.E.; Hill, H.H.; Bendiak, B. Separation of sodiated isobaric disaccharides and trisaccharides using electrospray ionization-atmospheric pressure ion mobility-time of flight mass spectrometry. J. Am. Soc. Mass Spectrom. 2005, 16, 660–669. [Google Scholar] [CrossRef] [Green Version]
- Struwe, W.B.; Benesch, J.L.; Harvey, D.J.; Pagel, K. Collision cross sections of high-mannose N-glycans in commonly observed adduct states–identification of gas-phase conformers unique to [M-H]- ions. Analyst 2015, 140, 6799–6803. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lapthorn, C.; Pullen, F.; Chowdhry, B.Z. Ion mobility spectrometry-mass spectrometry (IMS-MS) of small molecules: Separating and assigning structures to ions. Mass Spectrom Rev. 2013, 32, 43–71. [Google Scholar] [CrossRef] [Green Version]
- Lanucara, F.; Holman, S.W.; Gray, C.J.; Eyers, C.E. The power of ion mobility-mass spectrometry for structural characterization and the study of conformational dynamics. Nat. Chem. 2014, 6, 281. [Google Scholar] [CrossRef]
- Luo, M.D.; Zhou, Z.W.; Zhu, Z.J. The application of ion mobility-mass spectrometry in untargeted metabolomics: From separation to identification. J. Anal. Test. 2020, 4, 163–174. [Google Scholar] [CrossRef]
- Zhou, Z.; Tu, J.; Xiong, X.; Shen, X.; Zhu, Z.J. LipidCCS: Prediction of collision cross-section values for lipids with high precision to support ion mobility–mass spectrometry-based lipidomics. Anal. Chem. 2017, 89, 9559–9566. [Google Scholar] [CrossRef]
- Zhou, Z.; Xiong, X.; Zhu, Z.J. MetCCS Predictor: A web server for predicting collision cross-section values of metabolite in metabolomics. Bioinformatics 2017, 33, 2235–2237. [Google Scholar] [CrossRef] [Green Version]
- Colby, S.M.; Thomas, D.G.; Nuñez, J.R.; Baxter, D.J.; Glaesemann, K.R.; Brown, J.M.; Pirrung, M.A.; Govind, N.; Teeguarden, J.G.; Metz, T.O.; et al. ISiCLE: A quantum chemistry pipeline for establishing in silico collision cross section libraries. Anal. Chem. 2019, 91, 4346–4356. [Google Scholar] [CrossRef]
- Zhou, Z.; Luo, M.; Chen, X.; Yin, Y.; Xiong, X.; Wang, R.; Zhu, Z.J. Ion mobility collision cross-section atlas for known and unknown metabolite annotation in untargeted metabolomics. Nat. Commun. 2020, 11, 1–13. [Google Scholar] [CrossRef] [PubMed]
- McCullagh, M.; Douce, D.; Van Hoeck, E.; Goscinny, S. Exploring the complexity of steviol glycosides analysis using ion mobility mass spectrometry. Anal. Chem. 2018, 90, 4585–4595. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, M.; Meyer, S.W.; Heyman, H.M.; Barsch, A.; Sumner, L.W. Generation of a collision cross section library for multi-dimensional plant metabolomics using UHPLC-trapped ion mobility-MS/MS. Metabolites 2019, 10, 13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, H.K.; Choi, Y.H.; Verpoorte, R. NMR-based metabolomic analysis of plants. Nat. Proto. 2010, 5, 536–549. [Google Scholar] [CrossRef]
- Foroutan, A.; Goldansaz, S.A.; Lipfert, M.; Wishart, D.S. Protocols for NMR analysis in livestock metabolomics. In Metabolomics; Humana: New York, NY, USA, 2019; pp. 311–324. [Google Scholar]
- Ward, J.L.; Baker, J.M.; Beale, M.H. Recent applications of NMR spectroscopy in plant metabolomics. FEBS J. 2007, 274, 1126–1131. [Google Scholar] [CrossRef] [Green Version]
- Emwas, A.H.M. The strengths and weaknesses of NMR spectroscopy and mass spectrometry with particular focus on metabolomics research. Methods Mol. Biol. 2015, 1277, 161–193. [Google Scholar] [PubMed]
- Deborde, C.; Moing, A.; Roch, L.; Jacob, D.; Rolin, D.; Giraudeau, P. Plant metabolism as studied by NMR spectroscopy. Prog. Nucl. Magn. Reson. Spectrosc. 2017, 102, 61–97. [Google Scholar] [CrossRef]
- Kim, H.K.; Choi, Y.H.; Verpoorte, R. NMR-based plant metabolomics: Where do we stand, where do we go? Trends Biotechnol. 2011, 29, 267–275. [Google Scholar] [CrossRef]
- Viant, M.R. Improved methods for the acquisition and interpretation of NMR metabolomic data. Biochem. Biophys. Res. Commun. 2003, 310, 943–948. [Google Scholar] [CrossRef]
- Tautenhahn, R.; Patti, G.J.; Rinehart, D.; Siuzdak, G. XCMS Online: A web-based platform to process untargeted metabolomic data. Anal. Chem. 2012, 84, 5035–5039. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Montenegro-Burke, J.R.; Aisporna, A.E.; Benton, H.P.; Rinehart, D.; Fang, M.; Huan, T.; Warth, B.; Forsberg, E.; Abe, B.T.; Ivanisevic, J. Data streaming for metabolomics: Accelerating data processing and analysis from days to minutes. Anal. Chem. 2017, 89, 1254–1259. [Google Scholar] [CrossRef]
- Smith, C.A.; O’Maille, G.; Want, E.J.; Qin, C.; Trauger, S.A.; Brandon, T.R.; Custodio, D.E.; Abagyan, R.; Siuzdak, G. METLIN: A metabolite mass spectral database. Ther. Drug Monit. 2005, 27, 747–751. [Google Scholar] [CrossRef] [PubMed]
- Kessler, N.; Neuweger, H.; Bonte, A.; Langenkämper, G.; Niehaus, K.; Nattkemper, T.W.; Goesmann, A. MeltDB 2.0–advances of the metabolomics software system. Bioinformatics 2013, 29, 2452–2459. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xia, J.; Wishart, D.S. Using MetaboAnalyst 3.0 for comprehensive metabolomics data analysis. Curr. Protoc. Bioinform. 2016, 55, 14.10.1–14.10.91. [Google Scholar] [CrossRef]
- Wang, M.; Carver, J.J.; Phelan, V.V.; Sanchez, L.M.; Garg, N.; Peng, Y.; Nguyen, D.D.; Watrous, J.; Kapono, C.A.; Luzzatto-Knaan, T.; et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 2016, 34, 828–837. [Google Scholar] [CrossRef] [Green Version]
- Pluskal, T.; Castillo, S.; Villar-Briones, A.; Orešič, M. MZmine 2: Modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinform. 2010, 11, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Jiang, W.; Qiu, Y.; Ni, Y.; Su, M.; Jia, W.; Du, X. An automated data analysis pipeline for GC−TOF−MS metabonomics studies. J. Proteome Res. 2010, 9, 5974–5981. [Google Scholar] [CrossRef] [PubMed]
- Behrends, V.; Tredwell, G.D.; Bundy, J.G. A software complement to AMDIS for processing GC-MS metabolomic data. Anal. Biochem. 2011, 415, 206–208. [Google Scholar] [CrossRef] [PubMed]
- Skogerson, K.; Wohlgemuth, G.; Barupal, D.K.; Fiehn, O. The volatile compound BinBase mass spectral database. BMC Bioinform. 2011, 12, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Kind, T.; Wohlgemuth, G.; Lee, D.Y.; Lu, Y.; Palazoglu, M.; Shahbaz, S.; Fiehn, O. FiehnLib: Mass spectral and retention index libraries for metabolomics based on quadrupole and time-of-flight gas chromatography/mass spectrometry. Anal. Chem. 2009, 81, 10038–10048. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kameyama, A.; Kikuchi, N.; Nakaya, S.; Ito, H.; Sato, T.; Shikanai, T.; Takahashi, Y.; Takahashi, K.; Narimatsu, H. A strategy for identification of oligosaccharide structures using observational multistage mass spectral library. Anal. Chem. 2005, 77, 4719–4725. [Google Scholar] [CrossRef]
- Aoki, K.F.; Kanehisa, M. Using the KEGG database resource. Curr. Protoc. Bioinform. 2005, 11, 1–12. [Google Scholar] [CrossRef]
- Shinbo, Y.; Nakamura, Y.; Altaf-Ul-Amin, M.; Asahi, H.; Kurokawa, K.; Arita, M.; Saito, K.; Ohta, D.; Shibata, D.; Kanaya, S. KNApSAcK: A comprehensive species-metabolite relationship database. In Plant Metabolomics; Saito, K., Richard, A.D., Willmitzer, L., Eds.; Springer: Berlin/Heidelberg, Germany, 2006; pp. 165–181. [Google Scholar]
- Kaever, A.; Landesfeind, M.; Feussner, K.; Mosblech, A.; Heilmann, I.; Morgenstern, B.; Feussner, I.; Meinicke, P. MarVis-Pathway: Integrative and exploratory pathway analysis of non-targeted metabolomics data. Metabolomics 2015, 11, 764–777. [Google Scholar] [CrossRef] [Green Version]
- Ara, T.; Sakurai, N.; Suzuki, H.; Aoki, K.; Saito, K.; Shibata, D. MassBase: A large-scaled depository of mass spectrometry datasets for metabolome analysis. Plant Biotechnol. 2021, 38, 167–171. [Google Scholar] [CrossRef]
- Clasquin, M.F.; Melamud, E.; Rabinowitz, J.D. LC-MS data processing with MAVEN: A metabolomic analysis and visualization engine. Curr. Protoc. Bioinform. 2012, 37, 1–23. [Google Scholar]
- Xia, J.; Psychogios, N.; Young, N.; Wishart, D.S. MetaboAnalyst: A web server for metabolomic data analysis and interpretation. Nucleic Acids Res. 2009, 37, 652–660. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carroll, A.J.; Badger, M.R.; Millar, A.H. The Metabolome Express Project: Enabling web-based processing, analysis and transparent dissemination of GC/MS metabolomics datasets. BMC Bioinform. 2010, 11, 1–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, B.; Wang, J.; Ressom, H.W. MetaboSearch: Tool for mass-based metabolite identification using multiple databases. PLoS ONE 2012, 7, e40096. [Google Scholar] [CrossRef] [Green Version]
- Wanichthanarak, K.; Fan, S.; Grapov, D.; Barupal, D.K.; Fiehn, O. Metabox: A toolbox for metabolomic data analysis, interpretation and integrative exploration. PLoS ONE 2017, 12, e0171046. [Google Scholar]
- Lommen, A.; Kools, H.J. MetAlign 3.0: Performance enhancement by efficient use of advances in computer hardware. Metabolomics 2012, 8, 719–726. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kastenmüller, G.; Römisch-Margl, W.; Wägele, B.; Altmaier, E.; Suhre, K. MetaP-server: A web-based metabolomics data analysis tool. BioMed Res. Int. 2010, 2011, 1–7. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Daly, R.; Rogers, S.; Wandy, J.; Jankevics, A.; Burgess, K.E.; Breitling, R. MetAssign: Probabilistic annotation of metabolites from LC–MS data using a Bayesian clustering approach. Bioinformatics 2014, 30, 2764–2771. [Google Scholar] [CrossRef] [Green Version]
- Ruttkies, C.; Schymanski, E.L.; Wolf, S.; Hollender, J.; Neumann, S. MetFrag relaunched: Incorporating strategies beyond in silico fragmentation. J. Cheminform. 2016, 8, 3. [Google Scholar] [CrossRef] [Green Version]
- Lei, Z.; Li, H.; Chang, J.; Zhao, P.X.; Sumner, L.W. MET-IDEA version 2.06; Improved efficiency and additional functions for mass spectrometry-based metabolomics data processing. Metabolomics 2012, 8, 105–110. [Google Scholar] [CrossRef]
- Rojas-Chertó, M.; Van Vliet, M.; Peironcely, J.E.; Van Doorn, R.; Kooyman, M.; Te Beek, T.; Van Driel, M.A.; Hankemeier, T.; Reijmers, T. MetiTree: A web application to organize and process high-resolution multi-stage mass spectrometry metabolomics data. Bioinformatics 2012, 28, 2707–2709. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cui, Q.; Lewis, I.A.; Hegeman, A.D.; Anderson, M.E.; Li, J.; Schulte, C.F.; Westler, W.M.; Eghbalnia, H.R.; Sussman, M.R.; Markley, J.L. Metabolite identification via the madison metabolomics consortium database. Nat. Biotechnol. 2008, 26, 162. [Google Scholar] [CrossRef]
- Menikarachchi, L.C.; Cawley, S.; Hill, D.W.; Hall, L.M.; Hall, L.; Lai, S.; Wilder, J.; Grant, D.F. MolFind: A software package enabling HPLC/MS-based identification of unknown chemical structures. Anal. Chem. 2012, 84, 9388–9394. [Google Scholar] [CrossRef] [Green Version]
- Mistrik, R.; Lutisan, J.; Huang, Y.; Suchy, M.; Wang, J.; Raab, M. mzCloud: A key conceptual shift to understand ’Who’s Who’ in untargeted metabolomics. In Proceedings of the Metabolomics Society 2013 Conference, Glasgow, UK, 1–13 July 2013; pp. 1–4. [Google Scholar]
- Draper, J.; Enot, D.P.; Parker, D.; Beckmann, M.; Snowdon, S.; Lin, W.; Zubair, H. Metabolite signal identification in accurate mass metabolomics data with MZedDB, an interactive m/z annotation tool utilising predicted ionisation behaviour ‘rules’. BMC Bioinform. 2009, 10, 1–16. [Google Scholar] [CrossRef] [Green Version]
- Rumble, J.R., Jr.; Bickham, D.M.; Powell, C.J. The NIST x-ray photoelectron spectroscopy database. Surf. Interface Anal. 1992, 19, 241–246. [Google Scholar] [CrossRef]
- Sakurai, T.; Yamada, Y.; Sawada, Y.; Matsuda, F.; Akiyama, K.; Shinozaki, K.; Hirai, M.Y.; Saito, K. PRIMe update: Innovative content for plant metabolomics and integration of gene expression and metabolite accumulation. Plant Cell Physiol. 2013, 54, e5. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Alonso, M.M.; Carrasco-Loba, V.; Pollmann, S. Advances in plant metabolomics. Annu. Plant. Rev. Online 2018, 1, 557–588. [Google Scholar]
- Kanehisa, M.; Furumichi, M.; Tanabe, M. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017, 45, 353–361. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Caspi, R.; Foerster, H.; Fulcher, C.A. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res. 2008, 36, 623–631. [Google Scholar] [CrossRef]
- Mueller, L.A.; Zhang, P.; Rhee, S.Y. AraCyc: A biochemical pathway database for Arabidopsis. Plant Physiol. 2003, 132, 453–460. [Google Scholar] [CrossRef] [Green Version]
- Jewison, T.; Su, Y.; Disfany, F.M. SMPDB 2.0: Big improvements to the small molecule pathway database. Nucleic Acids Res. 2014, 42, 478–484. [Google Scholar] [CrossRef] [Green Version]
- Xia, J.; Wishart, D.S. MSEA: A web-based tool to identify biologically meaningful patterns in quantitative metabolomic data. Nucleic Acids Res. 2010, 38, 71–77. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Karnovsky, A.; Weymouth, T.; Hull, T. Metscape 2 bioinformatics tool for the analysis and visualization of metabolomics and gene expression data. Bioinformatics 2012, 28, 373–380. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kohl, M.; Wiese, S.; Warscheid, B. Cytoscape: Software for visualization and analysis of biological networks. Methods Mol. Biol. 2011, 696, 291–303. [Google Scholar] [PubMed]
- Garcia-Alcalde, F.; Garcia-Lopez, F.; Dopazo, J. Paintomics: A web based tool for the joint visualization of transcriptomics and metabolomics data. Bioinformatics 2011, 27, 137–139. [Google Scholar] [CrossRef] [Green Version]
- Neuweger, H.; Persicke, M.; Albaum, S.P. Visualizing post genomics data-sets on customized pathway maps by ProMeTra-aeration-dependent gene expression and metabolism of Corynebacterium glutamicum as an example. BMC Syst. Biol. 2009, 3, 82. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grapov, D.; Wanichthanarak, K.; Fiehn, O. MetaMapR: Pathway independent metabolomic network analysis incorporating unknowns. Bioinformatics 2015, 31, 2757–2760. [Google Scholar] [CrossRef] [Green Version]
- Smoot, M.E.; Ono, K.; Ruscheinski, J.; Wang, P.L.; Ideker, T. Cytoscape 2.8: New features for data integration and network visualization. Bioinformatics 2011, 27, 431–432. [Google Scholar] [CrossRef] [Green Version]
- Kamburov, A.; Cavill, R.; Ebbels, T.M.; Herwig, R.; Keun, H.C. Integrated pathway-level analysis of transcriptomics and metabolomics data with IMPaLA. Bioinformatics 2011, 27, 2917–2918. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Yamada, T.; Kanehisa, M.; Bork, P. iPath: Interactive exploration of biochemical pathways and networks. Trends Biochem. Sci. 2008, 33, 101–103. [Google Scholar] [CrossRef]
- Thimm, O.; Bläsing, O.; Gibon, Y.; Nagel, A.; Meyer, S.; Krüger, P.; Selbig, J.; Müller, L.A.; Rhee, S.Y.; Stitt, M. MAPMAN: A user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J. 2004, 37, 914–939. [Google Scholar] [CrossRef]
- Chagoyen, M.; Pazos, F. MBRole: Enrichment analysis of metabolomic data. Bioinformatics 2011, 27, 730–731. [Google Scholar] [CrossRef] [Green Version]
- Ara, T.; Enomoto, M.; Arita, M.; Ikeda, C.; Kera, K.; Yamada, M.; Nishioka, T.; Ikeda, T.; Nihei, Y.; Shibata, D.; et al. Metabolonote: A wiki-based database for managing hierarchical metadata of metabolome analyses. Front. Bioeng. Biotechnol. 2015, 3, 38. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schreiber, F.; Colmsee, C.; Czauderna, T.; Grafahrend-Belau, E.; Hartmann, A.; Junker, A.; Junker, B.H.; Klapperstück, M.; Scholz, U.; Weise, S. MetaCrop 2.0: Managing and exploring information about crop plant metabolism. Nucleic Acids Res. 2012, 40, 1173–1177. [Google Scholar] [CrossRef]
- Xia, J.; Wishart, D.S. MetPA: A web-based metabolomics tool for pathway analysis and visualization. Bioinformatics 2010, 26, 2342–2344. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kankainen, M.; Gopalacharyulu, P.; Holm, L.; Orešič, M. MPEA—Metabolite pathway enrichment analysis. Bioinformatics 2011, 27, 1878–1879. [Google Scholar] [CrossRef] [PubMed]
- Elliott, B.; Kirac, M.; Cakmak, A.; Yavas, G.; Mayes, S.; Cheng, E.; Wang, Y.; Gupta, C.; Ozsoyoglu, G.; Meral Ozsoyoglu, Z. PathCase: Pathways database system. Bioinformatics 2008, 24, 2526–2533. [Google Scholar] [CrossRef]
- Mlecnik, B.; Scheideler, M.; Hackl, H.; Hartler, J.; Sanchez-Cabo, F.; Trajanoski, Z. PathwayExplorer: Web service for visualizing high-throughput expression data on biological pathways. Nucleic Acids Res. 2005, 33, 633–637. [Google Scholar] [CrossRef] [PubMed]
- Junker, B.H.; Klukas, C.; Schreiber, F. VANTED: A system for advanced data analysis and visualization in the context of biological networks. BMC Bioinform. 2006, 7, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Kelder, T.; Van Iersel, M.P.; Hanspers, K. WikiPathways: Building research communities on biological pathways. Nucleic Acids Res. 2012, 40, 1301–1307. [Google Scholar] [CrossRef] [Green Version]
- Razzaq, A.; Saleem, F.; Kanwal, M.; Mustafa, G.; Yousaf, S.; Imran Arshad, H.M.; Hameed, M.K.; Khan, M.S.; Joyia, F.A. Modern trends in plant genome editing: An inclusive review of the CRISPR/Cas9 toolbox. Int. J. Mol. Sci. 2019, 20, 4045. [Google Scholar] [CrossRef] [Green Version]
| Transgenic Plants | Analytical Techniques | Key Metabolites | References |
|---|---|---|---|
| Artemisia annua | GC-TOF-MS | Borneol, phytol, β-farnesene, germacrene D, artemisinic acid, dihydroartemisinic acid, and artemisinin | [46] |
| Lactuca sativa | NMR | Asparagine, glutamine, valine, isoleucine, α-chetoglutarate, succinate, fumarate, malate, sucrose, and fructose | [47] |
| Lycopersicon esculentum | GC-MS | γ-aminobutyric acid, histidine, proline, pyrrol-2-carboxylate, galactitiol/sorbitol, glycerol, maltitol, 3-phosphoglyceric acid, allantoin, homo-cystine, caffeate, gluconate, ribonate, lysine, threonine, homo-serine, tyrosine, tryptophan, leucine, arginine and valine | [48] |
| Nicotiana tabacum | NMR | Chlorogenic acid, 4-O-caffeoylquinic acid, malic acid, threonine, alanine, glycine, fructose, β-glucose, α-glucose, sucrose, fumaric acid and salicylic acid | [49] |
| N. tabacum | GC-MS | 4-Aminobutanoic acid, asparagine, glutamine, glycine, leucine, phenylalanine, proline, serine, threonine, tryptophan, chlorogenic acid, quininic acid, threonic acid, citric acid, malic acid and ethanolamine | [50] |
| Oryza sativa | GC-MS | Glycerol-3-phosphate, citric acid, linoleic acid, oleic acid, hexadecanoic acid, 2,3-dihydroxypropyl ester, sucrose, 9-octadecenoic acid, 2,3-dihydroxypropyl ester, sucrose, mannitol and glutamic acid | [44] |
| O. sativa | LC-MS | Tryptophan, phytosphingosine, palmitic acid, 5-hydroxy-2-octadenoic acid 9,10,13-trihydroxyoctadec-11-enoic acid and ethanolamine | [51] |
| Populus | GC-MS, HPLC | Caffeoyl and feruloyl conjugates, syringyl-to-guaiacyl ratio, asparagine, glutamine, aspartic acid, γ-amino-butyric acid, 5-oxo-proline, salicylic acid-2-O-glucoside, 2, 5-dihydroxybenzoic acid-5-O-glucoside, 2-methoxyhydroquinone-1-O-glucoside, 2-methoxyhydroquinone-4-O-glucoside, salicin, gallic acid, and dihydroxybenzoic acid | [52] |
| Solanum tuberosum | LC-TOF-MS | Glutathione, γ-aminobutyric acid, β-cyanoalanine, 5-oxoproline, sucrose, glucose-1-phosphate, glucose-6-phosphate, fructose-6-phosphate, ethanolamine, adenosine, and guanosine | [45] |
| Triticum aestivum | GC-MS | Guanine and 4-hydroxycinnamic acid | [53] |
| T. aestivum | LC-MS | Aminoacyl-tRNA biosynthesis, phenylalanine, tyrosine, tryptophan glyoxylic, tartaric acid, oxalic acids, sucrose, galactose, mannitol, leucine, valine, glutamate, proline, pyridoxamine, glutathione, arginine, citrulline, adenosine, hypoxanthine, allantoin, and adenosine monophosphate | [54] |
| Zea mays | 1H NMR | Lactic acid, citric acid, lysine, arginine, glycine-betaine, raffinose, trehalose, galactose, and adenine | [55] |
| Analytical Method | Advantage | Disadvantage |
|---|---|---|
| GC-MS |
|
|
| LC-MS |
|
|
| CE-MS |
|
|
| FTICR-MS |
|
|
| MALDI-MSI |
|
|
| IMS |
|
|
| NMR |
|
|
| Database | Website (URL, Accessed on 29 June 2021) | References |
|---|---|---|
| AraCyc | https://www.plantcyc.org/typeofpublication/aracyc | [175] |
| Cytoscape | http://www.cytoscape.org/ | [183] |
| IMPaLA | http://impala.molgen.mpg.de | [184] |
| iPath | http://pathways.embl.de/ | [185] |
| KEGG | http://www.genome.jp/kegg/ | [173] |
| MapMan | http://mapman.gabipd.org/web/guest/mapman | [186] |
| MBRole | http://csbg.cnb.csic.es/mbrole/ | [187] |
| Metabolonote | http://metabolonote.kazusa.or.jp/ | [188] |
| MetaCrop | http://metacrop.ipk-gatersleben.de | [189] |
| MetaCyc | http://www.metacyc.org | [174] |
| MetPA | http://metpa.metabolomics.ca/MetPA/ | [190] |
| MPEA | http://ekhidna.biocenter.helsinki.fi/poxo/mpea/ | [191] |
| MSEA | http://www.msea.ca. or http://www.metaboanalyst.ca | [177] |
| Pathcase | http://nashua.case.edu/PathwaysMAW/Web/ | [192] |
| PathwayExplorer | http://genome.tugraz.at/pathwayexplorer/pathwayexplorer_description.shtml | [193] |
| SMPDB | http://www.smpdb.ca | [176] |
| VANTED | https://immersive-nalytics.infotech.monash.edu/vanted/ | [194] |
| WikiPathways | http://wikipathways.org | [195] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Patel, M.K.; Pandey, S.; Kumar, M.; Haque, M.I.; Pal, S.; Yadav, N.S. Plants Metabolome Study: Emerging Tools and Techniques. Plants 2021, 10, 2409. https://doi.org/10.3390/plants10112409
Patel MK, Pandey S, Kumar M, Haque MI, Pal S, Yadav NS. Plants Metabolome Study: Emerging Tools and Techniques. Plants. 2021; 10(11):2409. https://doi.org/10.3390/plants10112409
Chicago/Turabian StylePatel, Manish Kumar, Sonika Pandey, Manoj Kumar, Md Intesaful Haque, Sikander Pal, and Narendra Singh Yadav. 2021. "Plants Metabolome Study: Emerging Tools and Techniques" Plants 10, no. 11: 2409. https://doi.org/10.3390/plants10112409
APA StylePatel, M. K., Pandey, S., Kumar, M., Haque, M. I., Pal, S., & Yadav, N. S. (2021). Plants Metabolome Study: Emerging Tools and Techniques. Plants, 10(11), 2409. https://doi.org/10.3390/plants10112409









