Conversion of Germ Cells to Somatic Cell Types in C. elegans
Abstract
1. Loss of Germline Totipotency and Teratoma Formation
1.1. Translational Regulators Maintain Germline Totipotency and Prevent Teratomas
1.2. P Granules Safeguard Germline Identity
1.3. Epigenetic Regulators SPR-5/LSD1 and LET-418/Mi2 Protect Germline Fate
1.4. Epigenetic Regulators SET-2/SETD1A and WDR-5.1/WDR5 Maintain Germ Cell Pluripotency
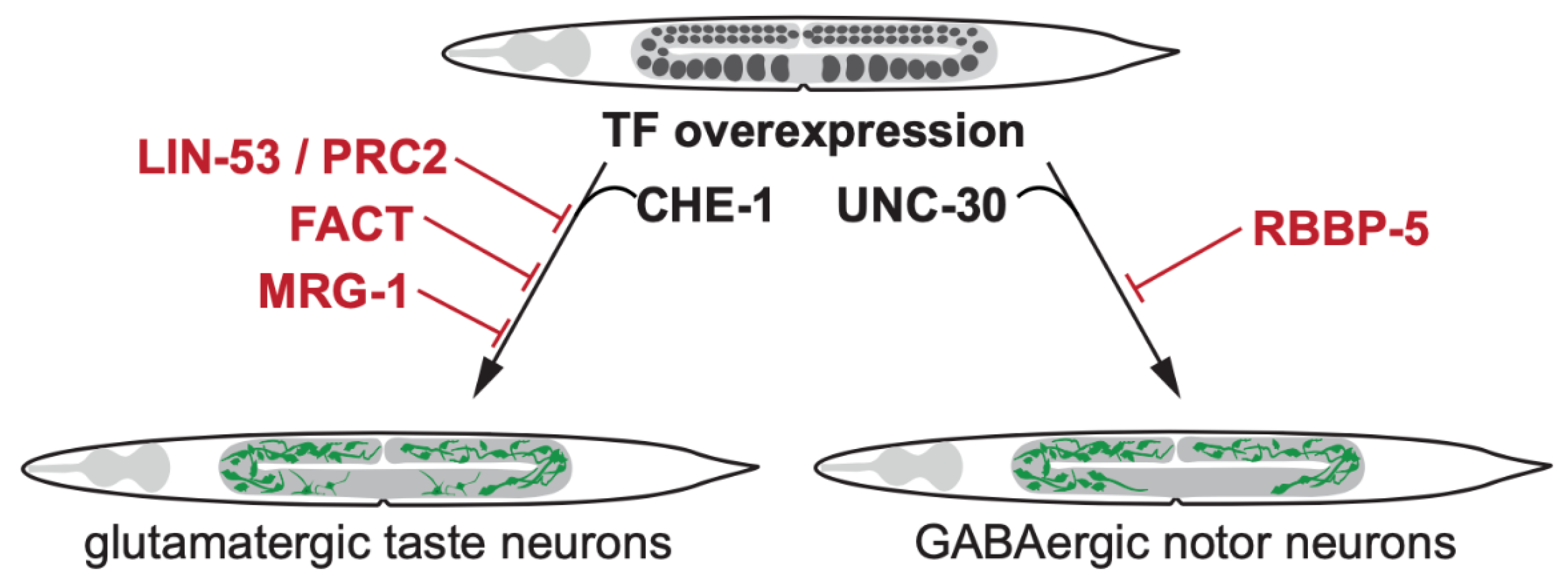
2. Transcription Factor-Induced Germ Cell Reprogramming to Somatic Fates
2.1. Loss of Histone Chaperone LIN-53 Permits Germ Cell Reprogramming by Transcription Factors
2.2. LIN-53 Cooperates with PRC2 to Prevent Permits Germ Cell Reprogramming by TFs
2.3. LIN-53 and PRC2 Antagonize Notch Signaling-Enhanced Germ Cell Reprogramming
2.4. The Heterodimeric Histone Chaperone FACT Prevents TF-Induced Germ Cell Reprogramming
2.5. The Chromodomain Protein MRG-1 Counteracts TF-Mediated Germ Cell Reprogramming
2.6. The Methyltransferase Complex Member RBBP-5 Blocks TF-Mediated Germ Cell Conversion
3. Outlook
Author Contributions
Funding
Conflicts of Interest
References
- Ciosk, R.; DePalma, M.; Priess, J.R. Translational regulators maintain totipotency in the Caenorhabditis elegans germline. Science 2006, 311, 851. [Google Scholar] [CrossRef] [PubMed]
- Updike, D.L.; Knutson, A.K.K.A.; Egelhofer, T.A.; Campbell, A.C.; Strome, S. Germ-granule components prevent somatic development in the C. Elegans germline. Curr. Biol. 2014, 24, 970–975. [Google Scholar] [CrossRef] [PubMed]
- Lei, H.; Liu, J.; Fukushige, T.; Fire, A.; Krause, M. Caudal-like PAL-1 directly activates the bodywall muscle module regulator hlh-1 in C. elegans to initiate the embryonic muscle gene regulatory network. Development 2009, 136, 1241–1249. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.T.; Seydoux, G. P granules. Curr. Biol. 2014, 24, 637–638. [Google Scholar] [CrossRef]
- Miller, D.M.; Stockdale, F.E.; Karn, J. Immunological identification of the genes encoding the four myosin heavy chain isoforms of Caenorhabditis elegans. Proc. Natl. Acad. Sci. USA 1986, 83, 2305–2309. [Google Scholar] [CrossRef]
- Etchberger, J.F.; Lorch, A.; Sleumer, M.C.; Zapf, R.; Jones, S.J.; Marra, M.A.; Holt, R.A.; Moerman, D.G.; Hobert, O. The molecular signature and cis-regulatory architecture of a C. elegans gustatory neuron. Genes Dev. 2007, 21, 1653–1674. [Google Scholar] [CrossRef]
- Tursun, B.; Patel, T.; Kratsios, P.; Hobert, O. Direct conversion of C. elegans germ cells into specific neuron types. Science 2011, 331, 304–308. [Google Scholar] [CrossRef]
- Käser-Pébernard, S.; Müller, F.; Wicky, C. LET-418/Mi2 and SPR-5/LSD1 cooperatively prevent somatic reprogramming of C. Elegans germline stem cells. Stem Cell Reports 2014, 2, 547–559. [Google Scholar] [CrossRef][Green Version]
- Shilatifard, A. The COMPASS family of histone H3K4 methylases: Mechanisms of regulation in development and disease pathogenesis. Annu. Rev. Biochem. 2012, 81, 65–95. [Google Scholar] [CrossRef]
- Unhavaithaya, Y.; Shin, T.H.; Miliaras, N.; Lee, J.; Oyama, T.; Mello, C.C. MEP-1 and a homolog of the NURD complex component Mi-2 act together to maintain germline-soma distinctions in C. elegans. Cell 2002, 111, 991–1002. [Google Scholar] [CrossRef]
- Li, T.; Kelly, W.G. A Role for Set1/MLL-Related Components in Epigenetic Regulation of the Caenorhabditis elegans Germ Line. PLoS Genet. 2011, 7, e1001349. [Google Scholar] [CrossRef] [PubMed]
- Robert, V.J.; Mercier, M.G.; Bedet, C.; Janczarski, S.; Merlet, J.; Garvis, S.; Ciosk, R.; Palladino, F. The SET-2/SET1 Histone H3K4 Methyltransferase Maintains Pluripotency in the Caenorhabditis elegans Germline. Cell Rep. 2014, 2, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Bedet, C.; Robert, V.J.P.; Simonet, T.; Dunkelbarger, S.; Rakotomalala, C.; Soete, G.; Korswagen, H.C.; Strome, S.; Palladino, F. Caenorhabditis elegans chromatin-associated proteins SET-2 and ASH-2 are differentially required for histone H3 Lys 4 methylation in embryos and adult germ cells. Proc. Natl. Acad. Sci. USA 2011, 108, 8305–8310. [Google Scholar] [CrossRef] [PubMed]
- Buckley, B.A.; Burkhart, K.B.; Gu, S.G.; Spracklin, G.; Kershner, A.; Fritz, H.; Kimble, J.; Fire, A.; Kennedy, S. A nuclear Argonaute promotes multigenerational epigenetic inheritance and germline immortality. Nature 2012, 489, 447–451. [Google Scholar] [CrossRef]
- Yuzyuk, T.; Fakhouri, T.H.I.; Kiefer, J.; Mango, S.E. The Polycomb Complex Protein mes-2/E(z) Promotes the Transition from Developmental Plasticity to Differentiation in C. elegans Embryos. Dev. Cell 2009, 16, 699–710. [Google Scholar] [CrossRef]
- Harrison, M.M.; Ceol, C.J.; Lu, X.; Horvitz, H.R. Some C. elegans class B synthetic multivulva proteins encode a conserved LIN-35 Rb-containing complex distinct from a NuRD-like complex. Proc. Natl. Acad. Sci. USA 2006, 103, 16782–16787. [Google Scholar] [CrossRef]
- Patel, T.; Tursun, B.; Rahe, D.P.; Hobert, O. Removal of Polycomb Repressive Complex 2 Makes C. elegans Germ Cells Susceptible to Direct Conversion into Specific Somatic Cell Types. Cell Rep. 2012, 2, 1178–1186. [Google Scholar] [CrossRef]
- Kelly, W.G.; Fire, A. Chromatin silencing and the maintenance of a functional germline in Caenorhabditis elegans. Development 1998, 125, 2451. [Google Scholar] [CrossRef]
- Tursun, B. PcG Proteins in Caenorhabditis elegans. In Polycomb Group Proteins; Academic Press: New York, NY, USA, 2017; pp. 289–315. [Google Scholar]
- Xu, L.; Fong, Y.; Strome, S. The Caenorhabditis elegans maternal-effect sterile proteins, MES-2, MES-3, and MES-6, are associated in a complex in embryos. Proc. Natl. Acad. Sci. USA 2001, 98, 5061–5066. [Google Scholar] [CrossRef]
- Gaydos, L.J.; Wang, W.; Strome, S. H3K27me and PRC2 transmit a memory of repression across generations and during development. Science 2014, 345, 1515–1518. [Google Scholar] [CrossRef]
- Kaneshiro, K.R.; Rechtsteiner, A.; Strome, S. Sperm-inherited H3K27me3 impacts offspring transcription and development in C. elegans. Nat. Commun. 2019, 10, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Seelk, S.; Adrian-Kalchhauser, I.; Hargitai, B.; Hajduskova, M.; Gutnik, S.; Tursun, B.; Ciosk, R. Increasing notch signaling antagonizes PRC2-mediated silencing to promote reprograming of germ cells into neurons. Elife 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Berry, L.W.; Westlund, B.; Schedl, T. Germ-line tumor formation caused by activation of glp-1, a Caenorhabditis elegans member of the Notch family of receptors. Development 1997, 124, 925. [Google Scholar]
- Orphanides, G.; Wu, W.H.; Lane, W.S.; Hampsey, M.; Reinberg, D. The chromatin-specific transcription elongation factor FACT comprises human SPT16 and SSRP1 proteins. Nature 1999, 400, 284–288. [Google Scholar] [CrossRef] [PubMed]
- Stuwe, T.; Hothorn, M.; Lejeune, E.; Rybin, V.; Bortfeld, M.; Scheffzek, K.; Ladurner, A.G. The FACT Spt16 “peptidase” domain is a histone H3-H4 binding module. Proc. Natl. Acad. Sci. USA 2008, 105, 8884–8889. [Google Scholar] [CrossRef] [PubMed]
- Kolundzic, E.; Seelk, S.; Tursun, B. Application of RNAi and heat-shock-induced transcription factor expression to reprogram germ cells to neurons in C. elegans. J. Vis. Exp. 2018, 2018, e56889. [Google Scholar] [CrossRef] [PubMed]
- Kolundzic, E.; Ofenbauer, A.; Bulut, S.I.; Uyar, B.; Baytek, G.; Sommermeier, A.; Seelk, S.; He, M.; Hirsekorn, A.; Vucicevic, D.; et al. FACT Sets a Barrier for Cell Fate Reprogramming in Caenorhabditis elegans and Human Cells. Dev. Cell 2018, 46, 611–626. [Google Scholar] [CrossRef]
- Buenrostro, J.D.; Wu, B.; Chang, H.Y.; Greenleaf, W.J. ATAC-seq: A method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 2015, 2015, 21–29. [Google Scholar] [CrossRef]
- Iwamori, N.; Tominaga, K.; Sato, T.; Riehle, K.; Iwamori, T.; Ohkawa, Y.; Coarfa, C.; Ono, E.; Matzuk, M.M. MRG15 is required for pre-mRNA splicing and spermatogenesis. Proc. Natl. Acad. Sci. USA 2016, 113, E5408–E5415. [Google Scholar] [CrossRef]
- Olgun, A.; Aleksenko, T.; Pereira-Smith, O.M.; Vassilatis, D.K. Functional analysis of MRG-1: The ortholog of human MRG15 in Caenorhabditis elegans. J. Gerontol. Ser. A Biol. Sci. Med. Sci. 2005, 60, 543–548. [Google Scholar] [CrossRef][Green Version]
- Beurton, F.; Stempor, P.; Caron, M.; Appert, A.; Dong, Y.; Herbette, M.; Huang, N.; Chen, A.; Cluet, D.; Cout, Y.; et al. Physical and functional interaction between SET1/COMPASS complex component CFP-1 and a Sin3S HDAC complex in C. elegans. Nucleic Acids Res. 2019, 47, 11164–11180. [Google Scholar] [CrossRef] [PubMed]
- Fujita, M.; Takasaki, T.; Nakajima, N.; Kawano, T.; Shimura, Y.; Sakamoto, H. MRG-1, a mortality factor-related chromodomain protein, is required maternally for primordial germ cells to initiate mitotic proliferation in C. elegans. Mech. Dev. 2002, 114, 61–69. [Google Scholar] [CrossRef]
- Gupta, P.; Leahul, L.; Wang, X.; Wang, C.; Bakos, B.; Jasper, K.; Hansen, D. Proteasome regulation of the chromodomain protein MRG-1 controls the balance between proliferative fate and differentiation in the C. elegans germ line. Development 2015, 142, 291–302. [Google Scholar] [CrossRef] [PubMed]
- Hajduskova, M.; Baytek, G.; Kolundzic, E.; Gosdschan, A.; Kazmierczak, M.; Ofenbauer, A.; Del Rosal, M.L.B.; Herzog, S.; Ul Fatima, N.; Mertins, P.; et al. MRG-1/MRG15 is a barrier for germ cell to neuron reprogramming in Caenorhabditis elegans. Genetics 2019, 211, 121–139. [Google Scholar] [CrossRef]
- Gambetta, M.C.; Müller, J. A critical perspective of the diverse roles of O-GlcNAc transferase in chromatin. Chromosoma 2015, 124, 429–442. [Google Scholar] [CrossRef]
- Hanover, J.A.; Forsythe, M.E.; Hennessey, P.T.; Brodigan, T.M.; Love, D.C.; Ashwell, G.; Krause, M. A Caenorhabditis elegans model of insulin resistance: Altered macronutrient storage and dauer formation in an OGT-1 knockout. Proc. Natl. Acad. Sci. USA 2005, 102, 11266–11271. [Google Scholar] [CrossRef]
- Hoe, M.; Nicholas, H.R. Evidence of a MOF histone acetyltransferase-containing NSL complex in C. elegans. Worm 2014, 3, e982967. [Google Scholar] [CrossRef]
- Alvarez, F.; Muñoz, F.; Schilcher, P.; Imhof, A.; Almouzni, G.; Loyola, A. Sequential Establishment of Marks on Soluble Histones H3 and H4. J. Biol. Chem. 2011, 286, 17714–17721. [Google Scholar] [CrossRef]
- Kazmierczak, M.; Farré i Díaz, C.; Ofenbauer, A.; Tursun, B. The CONDOR pipeline for simultaneous knockdown of multiple genes identifies RBBP-5 as a germ cell reprogramming barrier in C. elegans. bioRx 2020, 311, 851. [Google Scholar] [CrossRef]
- Cheloufi, S.; Elling, U.; Hopfgartner, B.; Jung, Y.L.; Murn, J.; Ninova, M.; Hubmann, M.; Badeaux, A.I.; Ang, C.E.; Tenen, D.; et al. The histone chaperone CAF-1 safeguards somatic cell identity. Nature 2015, 528, 218–224. [Google Scholar] [CrossRef]

| Loss of Germline Totipotency and Teratoma Formation | |||
|---|---|---|---|
| Factor | Function | Human Homolog | Reference |
| GLD-1 | Translational regulation | QKI | [1] Ciosk et al., 2006 |
| MEX-3 | Translational regulation | MEX3A | [1] Ciosk et al., 2006 |
| SPR-5 | Chromatin regulation: | LSD-1 | [8] Käser-Pébernard et al., 2014 |
| Histone demethylation | |||
| LET-418 | Chromatin regulation: | CHD3 | [8] Käser-Pébernard et al., 2014 |
| Nucleosome remodeling, | |||
| Histone deacetylation | |||
| SET-2 | Chromatin regulation: | SETD1A | [12] Robert et al., 2014 |
| H3K4 methylation | |||
| WDR-5.1 | Chromatin regulation: | WDR5 | [12] Robert et al., 2014 |
| H3K4 methylation | |||
| HRDE-1 | RNA interference | - | [12] Robert et al., 2014 |
| P granule proteins | RNA helicase activity | DDX4 | [2] Updike et al., 2014 |
| (PGL-1/-3, GLH-1-4) | |||
| Antagonizing TF-Induced Germ Cell Reprogramming to Somatic Fates | |||
| LIN-53 | Histone chaperone | RBBP4/7 | [7] Tursun et al., 2011 |
| PRC-2 (MES-2/-3) | Epigenetic regulation H3K27methylation | EZH2, EED | [17] Patel et al., 2012 |
| MES-4 | Chromatin regulation | NSD proteins | [21] Gaydos et al., 2012 |
| H3K36 methylation | |||
| FACT | Histone chaperone | SSRP1, SUPT16H | [28] Kolundzic et al., 2018 |
| (HMG-3/-4, SPT-16) | |||
| MRG-1 | Part of NuA4 histone acteyltransferase complex | MRG-15 | [35] Hadjuskova et al., 2019 |
| RBBP-5 | Set1/MLL methyltransferase complex member | RBBP5 | [40] Kazmierck et al., 2020 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
ul Fatima, N.; Tursun, B. Conversion of Germ Cells to Somatic Cell Types in C. elegans. J. Dev. Biol. 2020, 8, 24. https://doi.org/10.3390/jdb8040024
ul Fatima N, Tursun B. Conversion of Germ Cells to Somatic Cell Types in C. elegans. Journal of Developmental Biology. 2020; 8(4):24. https://doi.org/10.3390/jdb8040024
Chicago/Turabian Styleul Fatima, Nida, and Baris Tursun. 2020. "Conversion of Germ Cells to Somatic Cell Types in C. elegans" Journal of Developmental Biology 8, no. 4: 24. https://doi.org/10.3390/jdb8040024
APA Styleul Fatima, N., & Tursun, B. (2020). Conversion of Germ Cells to Somatic Cell Types in C. elegans. Journal of Developmental Biology, 8(4), 24. https://doi.org/10.3390/jdb8040024




