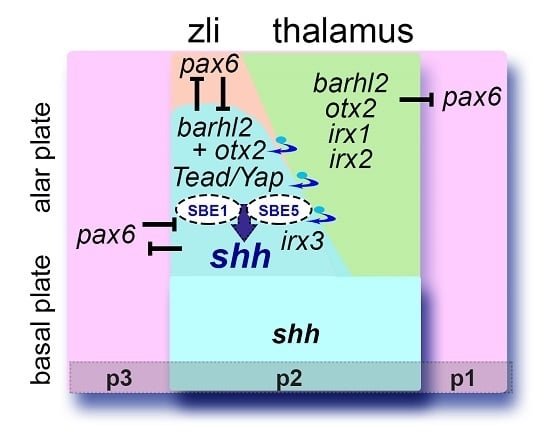
An Evolutionarily Conserved Network Mediates Development of the zona limitans intrathalamica, a Sonic Hedgehog-Secreting Caudal Forebrain Signaling Center
Abstract
:1. Introduction
2. The Zli: A Unique Shh-Expressing Compartment in the Caudal Forebrain
2.1. Forebrain Morphology and Zli Positioning
2.2. Shh Expression Initiates and Demarcates Zli Development
2.3. Physical Separation: The Zli as a Tissue Compartment
3. A Role for Wnt Ligands in Generating a Zli-Permissive Compartment
4. Laying the Ground for the zli: Otx2 and Barhl2 Binding to cis-Regulatory Sequences Confers Competence for zli Formation
5. Hedgehog Sequential Induction Process in zli Formation: Shh Is a Secreted Signal Read out through a Primary Cilium
5.1. Shh Maturation and Secretion
5.2. The Shh Signal Is Read through a Primary Cilium
6. Building Compartments: The Iroquois Genes Refine the Zli Borders
7. Sticking Together: GRNs and Signaling Pathways Modulate Zli Cell Differential Adhesive Properties and Proliferation Rate
7.1. Canonical Wnt Signaling Regulates Thalamic Cell Adhesiveness and Segregation
7.2. The Notch/Delta Pathway Contributes to Correct Separation of the Zli and Thalamic Fields
7.3. The TFs Barhl2 and Irx Facilitate Acquisition of Zli Compartment Properties
8. The Programming of A “Zli-Like” Structure in X. laevis Animal Cap Explants Confirms Otx2, Barhl2 and Iroquois Cell Autonomous and Non-Autonomous Activities in Zli Formation
9. Inductive Cues and Zli Positioning: A Role for the p2/p3 Border and for Fezf/Fez TFs
10. The GRN Involved in Zli Formation is Evolutionarily Conserved
11. Forebrain Malformations Associated with Defects in the GRN and Signaling Pathways Supporting Zli Formation
12. Conclusions
Acknowledgments
Conflicts of Interest
References
- Jessell, T.M.; Sanes, J.R. Development. The decade of the developing brain. Curr. Opin. Neurobiol. 2000, 10, 599–611. [Google Scholar] [CrossRef]
- Hebert, J.M.; Fishell, G. The genetics of early telencephalon patterning: Some assembly required. Nat. Rev. Neurosci. 2008, 9, 678–685. [Google Scholar] [CrossRef] [PubMed]
- Scholpp, S.; Lumsden, A. Review: Building a bridal chamber: Development of the thalamus. Trends Neurosci. 2010, 33, 373–380. [Google Scholar] [CrossRef] [PubMed]
- Cavodeassi, F.; Houart, C. Brain regionalization: Of signaling centers and boundaries. Dev. Neurobiol. 2012, 72, 218–233. [Google Scholar] [CrossRef] [PubMed]
- Kiecker, C.; Lumsden, A. The role of organizers in patterning the nervous system. Annu. Rev. Neurosci. 2012, 35, 347–367. [Google Scholar] [CrossRef] [PubMed]
- Echevarría, D.; Vieira, C.; Gimeno, L.; Martinez, S. Neuroepithelial secondary organizers and cell fate specification in the developing brain. Brain Res. Brain Res. Rev. 2003, 43, 179–191. [Google Scholar] [CrossRef] [PubMed]
- Anderson, C.; Stern, C.D. Organizers in development. Curr. Top. Dev. Biol. 2016, 117, 435–454. [Google Scholar] [PubMed]
- Scholpp, S.; Wolf, O.; Brand, M.; Lumsden, A. Hedgehog signalling from the zona limitans intrathalamica orchestrates patterning of the zebrafish diencephalon. Development 2006, 133, 855–864. [Google Scholar] [CrossRef] [PubMed]
- Vieira, C.; Garda, A.L.; Shimamura, K.; Martinez, S. Thalamic development induced by shh in the chick embryo. Dev. Biol. 2005, 284, 351–363. [Google Scholar] [CrossRef] [PubMed]
- Kiecker, C.; Lumsden, A. Hedgehog signaling from the zli regulates diencephalic regional identity. Nat. Neurosci. 2004, 7, 1242–1249. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto-Torii, K.; Motoyama, J.; Hui, C.C.; Kuroiwa, A.; Nakafuku, M.; Shimamura, K. Differential activities of sonic hedgehog mediated by gli transcription factors define distinct neuronal subtypes in the dorsal thalamus. Mech. Dev. 2003, 120, 1097–1111. [Google Scholar] [CrossRef] [PubMed]
- Juraver-Geslin, H.A.; Gomez-Skarmeta, J.L.; Durand, B.C. The conserved barh-like homeobox-2 gene barhl2 acts downstream of orthodentricle-2 and together with iroquois-3 in establishment of the caudal forebrain signaling center induced by sonic hedgehog. Dev. Biol. 2014, 396, 107–120. [Google Scholar] [CrossRef] [PubMed]
- Epstein, D.J. Regulation of thalamic development by sonic hedgehog. Front. Neurosci. 2012, 6, 57. [Google Scholar] [CrossRef] [PubMed]
- Jeong, Y.; Dolson, D.K.; Waclaw, R.R.; Matise, M.P.; Sussel, L.; Campbell, K.; Kaestner, K.H.; Epstein, D.J. Spatial and temporal requirements for sonic hedgehog in the regulation of thalamic interneuron identity. Development 2011, 138, 531–541. [Google Scholar] [CrossRef] [PubMed]
- Szabo, N.E.; Zhao, T.; Zhou, X.; Alvarez-Bolado, G. The role of sonic hedgehog of neural origin in thalamic differentiation in the mouse. J. Neurosci. 2009, 29, 2453–2466. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, M.; Li, J.Y. Patterning and compartment formation in the diencephalon. Front. Neurosci. 2012, 6, 66. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Ferre, A.; Martinez, S. Molecular regionalization of the diencephalon. Front. Neurosci. 2012, 6, 73. [Google Scholar] [CrossRef] [PubMed]
- Bergquist, H.; Kallen, B. Notes on the early histogenesis and morphogenesis of the central nervous system in vertebrates. J. Comp. Neurol. 1954, 100, 627–659. [Google Scholar] [CrossRef] [PubMed]
- Coggeshall, R.E. A study of diencephalic development in the albino rat. J. Comp. Neurol. 1964, 122, 241–269. [Google Scholar] [CrossRef] [PubMed]
- Keyser, A. The development of the diencephalon of the chinese hamster. An investigation of the validity of the criteria of subdivision of the brain. Acta Anat. Suppl. 1972, 59, 1–178. [Google Scholar] [CrossRef]
- Vaage, S. The segmentation of the primitive neural tube in chick embryos (gallus domesticus): A morphological, histochemical and autoradiographical investigation. Ergeb. Anat. Entwicklungsgesch. 1969, 41, 3–87. [Google Scholar] [PubMed]
- Bulfone, A.; Puelles, L.; Porteus, M.H.; Frohman, M.A.; Martin, G.R.; Rubenstein, J.L. Spatially restricted expression of dlx-1, dlx-2 (tes-1), gbx-2, and wnt-3 in the embryonic day 12.5 mouse forebrain defines potential transverse and longitudinal segmental boundaries. J. Neurosci. 1993, 13, 3155–3172. [Google Scholar] [PubMed]
- Rubenstein, J.L.; Martinez, S.; Shimamura, K.; Puelles, L. The embryonic vertebrate forebrain: The prosomeric model. Science 1994, 266, 578–580. [Google Scholar] [CrossRef] [PubMed]
- Figdor, M.C.; Stern, C.D. Segmental organization of embryonic diencephalon. Nature 1993, 363, 630–634. [Google Scholar] [CrossRef] [PubMed]
- Puelles, L.; Rubenstein, J.L.R. Forebrain gene expression domains and the evolving prosomeric model. Trends Neurosci. 2003, 26, 469–476. [Google Scholar] [CrossRef]
- Thompson, C.L.; Ng, L.; Menon, V.; Martinez, S.; Lee, C.K.; Glattfelder, K.; Sunkin, S.M.; Henry, A.; Lau, C.; Dang, C.; et al. A high-resolution spatiotemporal atlas of gene expression of the developing mouse brain. Neuron 2014, 83, 309–323. [Google Scholar] [CrossRef] [PubMed]
- Ferran, J.L.; Puelles, L.; Rubenstein, J.L. Molecular codes defining rostrocaudal domains in the embryonic mouse hypothalamus. Front. Neuroanat. 2015, 9, 46. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Lopez, R.; Vieira, C.; Echevarria, D.; Martinez, S. Fate map of the diencephalon and the zona limitans at the 10-somites stage in chick embryos. Dev. Biol. 2004, 268, 514–530. [Google Scholar] [CrossRef] [PubMed]
- Larsen, C.W.; Zeltser, L.M.; Lumsden, A. Boundary formation and compartition in the avian diencephalon. J. Neurosci. 2001, 21, 4699–4711. [Google Scholar] [PubMed]
- Shimamura, K.; Hartigan, D.J.; Martinez, S.; Puelles, L.; Rubenstein, J.L. Longitudinal organization of the anterior neural plate and neural tube. Development 1995, 121, 3923–3933. [Google Scholar] [PubMed]
- Zeltser, L.M.; Larsen, C.W.; Lumsden, A. A new developmental compartment in the forebrain regulated by lunatic fringe. Nat. Neurosci. 2001, 4, 683–684. [Google Scholar] [CrossRef] [PubMed]
- Scholpp, S.; Foucher, I.; Staudt, N.; Peukert, D.; Lumsden, A.; Houart, C. Otx1l, otx2 and irx1b establish and position the zli in the diencephalon. Development 2007, 134, 3167–3176. [Google Scholar] [CrossRef] [PubMed]
- Ericson, J.; Morton, S.; Kawakami, A.; Roelink, H.; Jessell, T.M. Two critical periods of sonic hedgehog signaling required for the specification of motor neuron identity. Cell 1996, 87, 661–673. [Google Scholar] [CrossRef]
- Watanabe, Y.; Nakamura, H. Control of chick tectum territory along dorsoventral axis by sonic hedgehog. Development 2000, 127, 1131–1140. [Google Scholar] [PubMed]
- Placzek, M.; Tessier-Lavigne, M.; Yamada, T.; Jessell, T.; Dodd, J. Mesodermal control of neural cell identity: Floor plate induction by the notochord. Science 1990, 250, 985–988. [Google Scholar] [CrossRef] [PubMed]
- Chamberlain, C.E.; Jeong, J.; Guo, C.; Allen, B.L.; McMahon, A.P. Notochord-derived shh concentrates in close association with the apically positioned basal body in neural target cells and forms a dynamic gradient during neural patterning. Development 2008, 135, 1097–1106. [Google Scholar] [CrossRef] [PubMed]
- Placzek, M.; Briscoe, J. The floor plate: Multiple cells, multiple signals. Nat. Rev. Neurosci. 2005, 6, 230–240. [Google Scholar] [CrossRef] [PubMed]
- Zeltser, L.M. Shh-dependent formation of the zli is opposed by signals from the dorsal diencephalon. Development 2005, 132, 2023–2033. [Google Scholar] [CrossRef] [PubMed]
- Vieira, C.; Martinez, S. Sonic hedgehog from the basal plate and the zona limitans intrathalamica exhibits differential activity on diencephalic molecular regionalization and nuclear structure. Neuroscience 2006, 143, 129–140. [Google Scholar] [CrossRef] [PubMed]
- Mathieu, J.; Barth, A.; Rosa, F.M.; Wilson, S.W.; Peyrieras, N. Distinct and cooperative roles for nodal and hedgehog signals during hypothalamic development. Development 2002, 129, 3055–3065. [Google Scholar] [PubMed]
- Holzschuh, J.; Hauptmann, G.; Driever, W. Genetic analysis of the roles of hh, fgf8, and nodal signaling during catecholaminergic system development in the zebrafish brain. J. Neurosci. 2003, 23, 5507–5519. [Google Scholar] [PubMed]
- Sampath, K.; Rubinstein, A.L.; Cheng, A.M.; Liang, J.O.; Fekany, K.; Solnica-Krezel, L.; Korzh, V.; Halpern, M.E.; Wright, C.V. Induction of the zebrafish ventral brain and floorplate requires cyclops/nodal signalling. Nature 1998, 395, 185–189. [Google Scholar] [PubMed]
- Martinez, S.; Puelles, L. Neurogenetic compartments of the mouse diencephalon and some characteristic gene expression patterns. Results Probl. Cell Differ. 2000, 30, 91–106. [Google Scholar] [PubMed]
- Kiecker, C.; Lumsden, A. Compartments and their boundaries in vertebrate brain development. Nat. Rev. Neurosci. 2005, 6, 553–564. [Google Scholar] [CrossRef] [PubMed]
- Crossley, P.H.; Martinez, S.; Martin, G.R. Midbrain development induced by fgf8 in the chick embryo. Nature 1996, 380, 66–68. [Google Scholar] [CrossRef] [PubMed]
- Robertshaw, E.; Matsumoto, K.; Lumsden, A.; Kiecker, C. Irx3 and pax6 establish differential competence for shh-mediated induction of gabaergic and glutamatergic neurons of the thalamus. Proc. Natl. Acad. Sci. USA 2013, 110, E3919–E3926. [Google Scholar] [CrossRef] [PubMed]
- Rubenstein, J.L.; Shimamura, K.; Martinez, S.; Puelles, L. Regionalization of the prosencephalic neural plate. Annu. Rev. Neurosci. 1998, 21, 445–477. [Google Scholar] [CrossRef] [PubMed]
- Mattes, B.; Weber, S.; Peres, J.; Chen, Q.; Davidson, G.; Houart, C.; Scholpp, S. Wnt3 and wnt3a are required for induction of the mid-diencephalic organizer in the caudal forebrain. Neural Dev. 2012, 7, 12. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Ferre, A.; Navarro-Garberi, M.; Bueno, C.; Martinez, S. Wnt signal specifies the intrathalamic limit and its organizer properties by regulating shh induction in the alar plate. J. Neurosci. 2013, 33, 3967–3980. [Google Scholar] [CrossRef] [PubMed]
- Houart, C.; Westerfield, M.; Wilson, S.W. A small population of anterior cells patterns the forebrain during zebrafish gastrulation. Nature 1998, 391, 788–792. [Google Scholar] [CrossRef] [PubMed]
- Wilson, S.W.; Houart, C. Review: Early steps in the development of the forebrain. Dev. Cell 2004, 6, 167–181. [Google Scholar] [CrossRef]
- Ruiz i Altaba, A.; Nguyen, V.; Palma, V. The emergent design of the neural tube: Prepattern, shh morphogen and gli code. Curr. Opin. Genet. Dev. 2003, 13, 513–521. [Google Scholar] [CrossRef] [PubMed]
- Jacob, J.; Briscoe, J.; Britto, J.; Tannahill, D.; Keynes, R. Gli proteins and the control of spinal-cord patterning. EMBO Rep. 2003, 4, 761–765. [Google Scholar] [CrossRef] [PubMed]
- Jessell, T.M. Neuronal specification in the spinal cord: Inductive signals and transcriptional codes. Nat. Rev. Genet. 2000, 1, 20–29. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, D.; Kobayashi, M.; Matsumoto, K.; Ogura, T.; Nakafuku, M.; Shimamura, K. Early subdivisions in the neural plate define distinct competence for inductive signals. Development 2002, 129, 83–93. [Google Scholar] [PubMed]
- Gomez-Skarmeta, J.L.; Campuzano, S.; Modolell, J. Half a century of neural prepatterning: The story of a few bristles and many genes. Nat. Rev. Neurosci. 2003, 4, 587–598. [Google Scholar] [CrossRef] [PubMed]
- Acampora, D.; Postiglione, M.P.; Avantaggiato, V.; di Bonito, M.; Simeone, A. The role of otx and otp genes in brain development. Int. J. Dev. Biol. 2000, 44, 669–677. [Google Scholar] [PubMed]
- Beby, F.; Lamonerie, T. The homeobox gene otx2 in development and disease. Exp. Eye Res. 2013, 111, 9–16. [Google Scholar] [CrossRef] [PubMed]
- Pannese, M.; Polo, C.; Andreazzoli, M.; Vignali, R.; Kablar, B.; Barsacchi, G.; Boncinelli, E. The xenopus homologue of otx2 is a maternal homeobox gene that demarcates and specifies anterior body regions. Development 1995, 121, 707–720. [Google Scholar] [PubMed]
- Schuhmacher, L.N.; Albadri, S.; Ramialison, M.; Poggi, L. Evolutionary relationships and diversification of barhl genes within retinal cell lineages. BMC Evol. Biol. 2011, 11, 340. [Google Scholar] [CrossRef] [PubMed]
- Reig, G.; Cabrejos, M.E.; Concha, M.L. Functions of barh transcription factors during embryonic development. Dev. Biol. 2007, 302, 367–375. [Google Scholar] [CrossRef] [PubMed]
- Juraver-Geslin, H.A.; Durand, B.C. Early development of the neural plate: New roles for apoptosis and for one of its main effectors caspase-3. Genesis 2015, 53, 203–224. [Google Scholar] [CrossRef] [PubMed]
- Offner, N.; Duval, N.; Jamrich, M.; Durand, B. The pro-apoptotic activity of a vertebrate bar-like homeobox gene plays a key role in patterning the xenopus neural plate by limiting the number of chordin- and shh-expressing cells. Development 2005, 132, 1807–1818. [Google Scholar] [CrossRef] [PubMed]
- Juraver-Geslin, H.A.; Ausseil, J.J.; Wassef, M.; Durand, B.C. Barhl2 limits growth of the diencephalic primordium through caspase3 inhibition of beta-catenin activation. Proc. Natl. Acad. Sci. USA 2011, 108, 2288–2293. [Google Scholar] [CrossRef] [PubMed]
- Staudt, N.; Houart, C. The prethalamus is established during gastrulation and influences diencephalic regionalization. PLoS Biol. 2007, 5, e69. [Google Scholar] [CrossRef] [PubMed]
- Mo, Z.; Li, S.; Yang, X.; Xiang, M. Role of the barhl2 homeobox gene in the specification of glycinergic amacrine cells. Development 2004, 131, 1607–1618. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Minor, P.J.; Zhao, Y.T.; Jeong, Y.; Pani, A.M.; King, A.N.; Symmons, O.; Gan, L.; Cardoso, W.V.; Spitz, F.; et al. cis-Regulatory architecture of a brain signaling center predates the origin of chordates. Nat. Genet. 2016, 48, 575–580. [Google Scholar] [CrossRef] [PubMed]
- Patterson, K.D.; Cleaver, O.; Gerber, W.V.; White, F.G.; Krieg, P.A. Distinct expression patterns for two xenopus bar homeobox genes. Dev. Genes Evol. 2000, 210, 140–144. [Google Scholar] [CrossRef] [PubMed]
- Colombo, A.; Reig, G.; Mione, M.; Concha, M.L. Zebrafish barh-like genes define discrete neural domains in the early embryo. Gene Expr. Patterns 2006, 6, 347–352. [Google Scholar] [CrossRef] [PubMed]
- Hirata, T.; Nakazawa, M.; Muraoka, O.; Nakayama, R.; Suda, Y.; Hibi, M. Zinc-finger genes fez and fez-like function in the establishment of diencephalon subdivisions. Development 2006, 133, 3993–4004. [Google Scholar] [CrossRef] [PubMed]
- Ding, Q.; Balasubramanian, R.; Zheng, D.; Liang, G.; Gan, L. Barhl2 determines the early patterning of the diencephalon by regulating shh. Mol. Neurobiol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Ramsbottom, S.A.; Pownall, M.E. Regulation of hedgehog signalling inside and outside the cell. J. Dev. Biol. 2016, 4, 23. [Google Scholar] [CrossRef] [PubMed]
- Porter, J.A.; Ekker, S.C.; Park, W.J.; von Kessler, D.P.; Young, K.E.; Chen, C.H.; Ma, Y.; Woods, A.S.; Cotter, R.J.; Koonin, E.V.; et al. Hedgehog patterning activity: Role of a lipophilic modification mediated by the carboxy-terminal autoprocessing domain. Cell 1996, 86, 21–34. [Google Scholar] [CrossRef]
- Porter, J.A.; von Kessler, D.P.; Ekker, S.C.; Young, K.E.; Lee, J.J.; Moses, K.; Beachy, P.A. The product of hedgehog autoproteolytic cleavage active in local and long-range signalling. Nature 1995, 374, 363–366. [Google Scholar] [CrossRef] [PubMed]
- Bumcrot, D.A.; Takada, R.; McMahon, A.P. Proteolytic processing yields two secreted forms of sonic hedgehog. Mol. Cell. Biol. 1995, 15, 2294–2303. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Tukachinsky, H.; Huang, C.H.; Jao, C.; Chu, Y.R.; Tang, H.Y.; Mueller, B.; Schulman, S.; Rapoport, T.A.; Salic, A. Processing and turnover of the hedgehog protein in the endoplasmic reticulum. J. Cell Biol. 2011, 192, 825–838. [Google Scholar] [CrossRef] [PubMed]
- Porter, J.A.; Young, K.E.; Beachy, P.A. Cholesterol modification of hedgehog signaling proteins in animal development. Science 1996, 274, 255–259. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhang, H.; Litingtung, Y.; Chiang, C. Cholesterol modification restricts the spread of shh gradient in the limb bud. Proc. Natl. Acad. Sci. USA 2006, 103, 6548–6553. [Google Scholar] [CrossRef] [PubMed]
- Pepinsky, R.B.; Zeng, C.; Wen, D.; Rayhorn, P.; Baker, D.P.; Williams, K.P.; Bixler, S.A.; Ambrose, C.M.; Garber, E.A.; Miatkowski, K.; et al. Identification of a palmitic acid-modified form of human sonic hedgehog. J. Biol. Chem. 1998, 273, 14037–14045. [Google Scholar] [CrossRef] [PubMed]
- Buglino, J.A.; Resh, M.D. Hhat is a palmitoylacyltransferase with specificity for N-palmitoylation of sonic hedgehog. J. Biol. Chem. 2008, 283, 22076–22088. [Google Scholar] [CrossRef] [PubMed]
- Chamoun, Z.; Mann, R.K.; Nellen, D.; von Kessler, D.P.; Bellotto, M.; Beachy, P.A.; Basler, K. Skinny hedgehog, an acyltransferase required for palmitoylation and activity of the hedgehog signal. Science 2001, 293, 2080–2084. [Google Scholar] [CrossRef] [PubMed]
- Hardy, R.Y.; Resh, M.D. Identification of n-terminal residues of sonic hedgehog important for palmitoylation by hedgehog acyltransferase. J. Biol. Chem. 2012, 287, 42881–42889. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.H.; Li, Y.J.; Kawakami, T.; Xu, S.M.; Chuang, P.T. Palmitoylation is required for the production of a soluble multimeric hedgehog protein complex and long-range signaling in vertebrates. Genes Dev. 2004, 18, 641–659. [Google Scholar] [CrossRef] [PubMed]
- Kohtz, J.D.; Lee, H.Y.; Gaiano, N.; Segal, J.; Ng, E.; Larson, T.; Baker, D.P.; Garber, E.A.; Williams, K.P.; Fishell, G. N-Terminal fatty-acylation of sonic hedgehog enhances the induction of rodent ventral forebrain neurons. Development 2001, 128, 2351–2363. [Google Scholar] [PubMed]
- Burke, R.; Nellen, D.; Bellotto, M.; Hafen, E.; Senti, K.A.; Dickson, B.J.; Basler, K. Dispatched, a novel sterol-sensing domain protein dedicated to the release of cholesterol-modified hedgehog from signaling cells. Cell 1999, 99, 803–815. [Google Scholar] [CrossRef]
- Creanga, A.; Glenn, T.D.; Mann, R.K.; Saunders, A.M.; Talbot, W.S.; Beachy, P.A. Scube/you activity mediates release of dually lipid-modified hedgehog signal in soluble form. Genes Dev. 2012, 26, 1312–1325. [Google Scholar] [CrossRef] [PubMed]
- Tukachinsky, H.; Kuzmickas, R.P.; Jao, C.Y.; Liu, J.; Salic, A. Dispatched and scube mediate the efficient secretion of the cholesterol-modified hedgehog ligand. Cell Rep. 2012, 2, 308–320. [Google Scholar] [CrossRef] [PubMed]
- Eugster, C.; Panakova, D.; Mahmoud, A.; Eaton, S. Lipoprotein-heparan sulfate interactions in the hh pathway. Dev. Cell 2007, 13, 57–71. [Google Scholar] [CrossRef] [PubMed]
- Panakova, D.; Sprong, H.; Marois, E.; Thiele, C.; Eaton, S. Lipoprotein particles are required for hedgehog and wingless signalling. Nature 2005, 435, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Liegeois, S.; Benedetto, A.; Garnier, J.M.; Schwab, Y.; Labouesse, M. The v0-atpase mediates apical secretion of exosomes containing hedgehog-related proteins in caenorhabditis elegans. J. Cell Biol. 2006, 173, 949–961. [Google Scholar] [CrossRef] [PubMed]
- Callejo, A.; Bilioni, A.; Mollica, E.; Gorfinkiel, N.; Andres, G.; Ibanez, C.; Torroja, C.; Doglio, L.; Sierra, J.; Guerrero, I. Dispatched mediates hedgehog basolateral release to form the long-range morphogenetic gradient in the drosophila wing disk epithelium. Proc. Natl. Acad. Sci. USA 2011, 108, 12591–12598. [Google Scholar] [CrossRef] [PubMed]
- Vyas, N.; Walvekar, A.; Tate, D.; Lakshmanan, V.; Bansal, D.; Lo Cicero, A.; Raposo, G.; Palakodeti, D.; Dhawan, J. Vertebrate hedgehog is secreted on two types of extracellular vesicles with different signaling properties. Sci. Rep. 2014, 4, 7357. [Google Scholar] [CrossRef] [PubMed]
- Huangfu, D.; Liu, A.; Rakeman, A.S.; Murcia, N.S.; Niswander, L.; Anderson, K.V. Hedgehog signalling in the mouse requires intraflagellar transport proteins. Nature 2003, 426, 83–87. [Google Scholar] [CrossRef] [PubMed]
- Hilgendorf, K.I.; Johnson, C.T.; Jackson, P.K. The primary cilium as a cellular receiver: Organizing ciliary gpcr signaling. Curr. Opin. Cell Biol. 2016, 39, 84–92. [Google Scholar] [CrossRef] [PubMed]
- Bai, C.B.; Stephen, D.; Joyner, A.L. All mouse ventral spinal cord patterning by hedgehog is gli dependent and involves an activator function of gli3. Dev. Cell 2004, 6, 103–115. [Google Scholar] [CrossRef]
- Briscoe, J. Making a grade: Sonic hedgehog signalling and the control of neural cell fate. EMBO J. 2009, 28, 457–465. [Google Scholar] [CrossRef] [PubMed]
- Fuccillo, M.; Joyner, A.L.; Fishell, G. Morphogen to mitogen: The multiple roles of hedgehog signalling in vertebrate neural development. Nat. Rev. Neurosci. 2006, 7, 772–783. [Google Scholar] [CrossRef] [PubMed]
- Mukhopadhyay, S.; Rohatgi, R. G-Protein-coupled receptors, hedgehog signaling and primary cilia. Semin. Cell. Dev. Biol. 2014, 33, 63–72. [Google Scholar] [CrossRef] [PubMed]
- Willaredt, M.A.; Hasenpusch-Theil, K.; Gardner, H.A.; Kitanovic, I.; Hirschfeld-Warneken, V.C.; Gojak, C.P.; Gorgas, K.; Bradford, C.L.; Spatz, J.; Wolfl, S.; et al. A crucial role for primary cilia in cortical morphogenesis. J. Neurosci. 2008, 28, 12887–12900. [Google Scholar] [CrossRef] [PubMed]
- Cruz, C.; Ribes, V.; Kutejova, E.; Cayuso, J.; Lawson, V.; Norris, D.; Stevens, J.; Davey, M.; Blight, K.; Bangs, F.; et al. Foxj1 regulates floor plate cilia architecture and modifies the response of cells to sonic hedgehog signalling. Development 2010, 137, 4271–4282. [Google Scholar] [CrossRef] [PubMed]
- Chung, M.I.; Peyrot, S.M.; LeBoeuf, S.; Park, T.J.; McGary, K.L.; Marcotte, E.M.; Wallingford, J.B. Rfx2 is broadly required for ciliogenesis during vertebrate development. Dev. Biol. 2012, 363, 155–165. [Google Scholar] [CrossRef] [PubMed]
- Kramer-Zucker, A.G.; Olale, F.; Haycraft, C.J.; Yoder, B.K.; Schier, A.F.; Drummond, I.A. Cilia-driven fluid flow in the zebrafish pronephros, brain and kupffer’s vesicle is required for normal organogenesis. Development 2005, 132, 1907–1921. [Google Scholar] [CrossRef] [PubMed]
- Christ, A.; Christa, A.; Kur, E.; Lioubinski, O.; Bachmann, S.; Willnow, T.E.; Hammes, A. Lrp2 is an auxiliary shh receptor required to condition the forebrain ventral midline for inductive signals. Dev. Cell. 2012, 22, 268–278. [Google Scholar] [CrossRef] [PubMed]
- Hagenlocher, C.; Walentek, P.; Müller, C.; Thumberger, T.; Feistel, K. Ciliogenesis and cerebrospinal fluid flow in the developing xenopus brain are regulated by foxj1. Cilia 2013, 2, 12. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Skarmeta, J.L.; Modolell, J. Iroquois genes: Genomic organization and function in vertebrate neural development. Curr. Opin. Genet. Dev. 2002, 12, 403–408. [Google Scholar] [CrossRef]
- Cavodeassi, F.; Modolell, J.; Gomez-Skarmeta, J.L. The iroquois family of genes: From body building to neural patterning. Development 2001, 128, 2847–2855. [Google Scholar] [PubMed]
- Peters, T.; Dildrop, R.; Ausmeier, K.; Ruther, U. Organization of mouse iroquois homeobox genes in two clusters suggests a conserved regulation and function in vertebrate development. Genome Res. 2000, 10, 1453–1462. [Google Scholar] [CrossRef] [PubMed]
- De la Calle-Mustienes, E.; Modolell, J.; Gomez-Skarmeta, J.L. The xiro-repressed gene corest is expressed in xenopus neural territories. Mech. Dev. 2002, 110, 209–211. [Google Scholar] [CrossRef]
- Rodriguez-Seguel, E.; Alarcon, P.; Gomez-Skarmeta, J.L. The xenopus irx genes are essential for neural patterning and define the border between prethalamus and thalamus through mutual antagonism with the anterior repressors fezf and arx. Dev. Biol. 2009, 329, 258–268. [Google Scholar] [CrossRef] [PubMed]
- Lecaudey, V.; Anselme, I.; Rosa, F.; Schneider-Maunoury, S. The zebrafish iroquois gene iro7 positions the r4/r5 boundary and controls neurogenesis in the rostral hindbrain. Development 2004, 131, 3121–3131. [Google Scholar] [CrossRef] [PubMed]
- Dahmann, C.; Oates, A.C.; Brand, M. Boundary formation and maintenance in tissue development. Nat. Rev. Genet. 2011, 12, 43–55. [Google Scholar] [CrossRef] [PubMed]
- Baek, J.H.; Hatakeyama, J.; Sakamoto, S.; Ohtsuka, T.; Kageyama, R. Persistent and high levels of hes1 expression regulate boundary formation in the developing central nervous system. Development 2006, 133, 2467–2476. [Google Scholar] [CrossRef] [PubMed]
- Grigoryan, T.; Wend, P.; Klaus, A.; Birchmeier, W. Deciphering the function of canonical wnt signals in development and disease: Conditional loss- and gain-of-function mutations of beta-catenin in mice. Genes Dev. 2008, 22, 2308–2341. [Google Scholar] [CrossRef] [PubMed]
- Van Amerongen, R.; Nusse, R. Towards an integrated view of wnt signaling in development. Development 2009, 136, 3205–3214. [Google Scholar] [CrossRef] [PubMed]
- Noelanders, R.; Vleminckx, K. How wnt signaling builds the brain: Bridging development and disease. Neuroscientist 2016. [Google Scholar] [CrossRef] [PubMed]
- Redies, C. Cadherins in the central nervous system. Prog. Neurobiol. 2000, 61, 611–648. [Google Scholar] [CrossRef]
- Puelles, E. Genetic control of basal midbrain development. J. Neurosci. Res. 2007, 85, 3530–3534. [Google Scholar] [CrossRef] [PubMed]
- Peukert, D.; Weber, S.; Lumsden, A.; Scholpp, S. Lhx2 and lhx9 determine neuronal differentiation and compartition in the caudal forebrain by regulating wnt signaling. PLoS Biol. 2011, 9, e1001218. [Google Scholar] [CrossRef] [PubMed]
- Pinto-Teixeira, F.; Desplan, C. Notch activity in neural progenitors coordinates cytokinesis and asymmetric differentiation. Sci. Signal. 2014, 7. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.C.; Amoyel, M.; Qiu, X.; Jiang, Y.J.; Xu, Q.; Wilkinson, D.G. Notch activation regulates the segregation and differentiation of rhombomere boundary cells in the zebrafish hindbrain. Dev. Cell 2004, 6, 539–550. [Google Scholar] [CrossRef]
- Tossell, K.; Kiecker, C.; Wizenmann, A.; Lang, E.; Irving, C. Notch signalling stabilises boundary formation at the midbrain-hindbrain organiser. Development 2011, 138, 3745–3757. [Google Scholar] [CrossRef] [PubMed]
- Barrios, N.; Gonzalez-Perez, E.; Hernandez, R.; Campuzano, S. The homeodomain iroquois proteins control cell cycle progression and regulate the size of developmental fields. PLoS Genet. 2015, 11, e1005463. [Google Scholar] [CrossRef] [PubMed]
- Villa-Cuesta, E.; Gonzalez-Perez, E.; Modolell, J. Apposition of iroquois expressing and non-expressing cells leads to cell sorting and fold formation in the drosophila imaginal wing disc. BMC Dev. Biol. 2007, 7, 106. [Google Scholar] [CrossRef] [PubMed]
- Jimenez-Guri, E.; Pujades, C. An ancient mechanism of hindbrain patterning has been conserved in vertebrate evolution. Evol. Dev. 2011, 13, 38–46. [Google Scholar] [CrossRef] [PubMed]
- Durand, B.C. Stem cell-like xenopus embryonic explants to study early neural developmental features in vitro and in vivo. J. Vis. Exp. 2016, e53474. [Google Scholar]
- Jeong, J.Y.; Einhorn, Z.; Mathur, P.; Chen, L.; Lee, S.; Kawakami, K.; Guo, S. Patterning the zebrafish diencephalon by the conserved zinc-finger protein fezl. Development 2007, 134, 127–136. [Google Scholar] [CrossRef] [PubMed]
- Hagemann, A.I.; Scholpp, S. The tale of the three brothers—Shh, wnt, and fgf during development of the thalamus. Front. Neurosci. 2012, 6, 76. [Google Scholar] [CrossRef] [PubMed]
- Caballero, I.M.; Manuel, M.N.; Molinek, M.; Quintana-Urzainqui, I.; Mi, D.; Shimogori, T.; Price, D.J. Cell-autonomous repression of shh by transcription factor pax6 regulates diencephalic patterning by controlling the central diencephalic organizer. Cell Rep. 2014, 8, 1405–1418. [Google Scholar] [CrossRef] [PubMed]
- Pani, A.M.; Mullarkey, E.E.; Aronowicz, J.; Assimacopoulos, S.; Grove, E.A.; Lowe, C.J. Ancient deuterostome origins of vertebrate brain signalling centres. Nature 2012, 483, 289–294. [Google Scholar] [CrossRef] [PubMed]
- Lowe, C.J.; Wu, M.; Salic, A.; Evans, L.; Lander, E.; Stange-Thomann, N.; Gruber, C.E.; Gerhart, J.; Kirschner, M. Anteroposterior patterning in hemichordates and the origins of the chordate nervous system. Cell 2003, 113, 853–865. [Google Scholar] [CrossRef]
- Llinas, R.R. Thalamo-cortical dysrhythmia syndrome: Neuropsychiatric features. Anales de la Real Academia Nacional de Medicina 2003, 120, 267–290; discussion 290–265. [Google Scholar] [PubMed]
- Marcorelles, P.; Laquerriere, A. Neuropathology of holoprosencephaly. Am. J. Med. Genet. Part C Semin. Med. Genet. 2010, 154, 109–119. [Google Scholar] [CrossRef] [PubMed]
- Willaredt, M.A.; Tasouri, E.; Tucker, K.L. Primary cilia and forebrain development. Mech. Dev. 2013, 130, 373–380. [Google Scholar] [CrossRef] [PubMed]
- Xavier, G.M.; Seppala, M.; Barrell, W.; Birjandi, A.A.; Geoghegan, F.; Cobourne, M.T. Hedgehog receptor function during craniofacial development. Dev. Biol. 2016, 415, 198–215. [Google Scholar] [CrossRef] [PubMed]
- Guerrini, R.; Dobyns, W.B. Malformations of cortical development: Clinical features and genetic causes. Lancet Neurol. 2014, 13, 710–726. [Google Scholar] [CrossRef]
- Hu, W.F.; Chahrour, M.H.; Walsh, C.A. The diverse genetic landscape of neurodevelopmental disorders. Ann. Rev. Genom. Hum. Genet. 2014, 15, 195–213. [Google Scholar] [CrossRef] [PubMed]
- Shepherd, G.M. Corticostriatal connectivity and its role in disease. Nat. Rev. Neurosci. 2013, 14, 278–291. [Google Scholar] [CrossRef] [PubMed]
- Hahn, J.S.; Barnes, P.D. Neuroimaging advances in holoprosencephaly: Refining the spectrum of the midline malformation. Am. J. Med. Genet. Part. C Semin. Med. Genet. 2010, 154C, 120–132. [Google Scholar] [CrossRef] [PubMed]
- Munke, M. Clinical, cytogenetic, and molecular approaches to the genetic heterogeneity of holoprosencephaly. Am. J. Med. Genet. 1989, 34, 237–245. [Google Scholar] [CrossRef] [PubMed]
- Nanni, L.; Ming, J.E.; Bocian, M.; Steinhaus, K.; Bianchi, D.W.; Die-Smulders, C.; Giannotti, A.; Imaizumi, K.; Jones, K.L.; Campo, M.D.; et al. The mutational spectrum of the sonic hedgehog gene in holoprosencephaly: Shh mutations cause a significant proportion of autosomal dominant holoprosencephaly. Hum. Mol. Genet. 1999, 8, 2479–2488. [Google Scholar] [CrossRef] [PubMed]
- Gongal, P.A.; French, C.R.; Waskiewicz, A.J. Aberrant forebrain signaling during early development underlies the generation of holoprosencephaly and coloboma. Biochim. Biophys. Acta 2011, 1812, 390–401. [Google Scholar] [CrossRef] [PubMed]
- Petryk, A.; Graf, D.; Marcucio, R. Holoprosencephaly: Signaling interactions between the brain and the face, the environment and the genes, and the phenotypic variability in animal models and humans. Wiley Interdiscip. Rev. Dev. Biol. 2015, 4, 17–32. [Google Scholar] [CrossRef] [PubMed]
- Dubourg, C.; Carre, W.; Hamdi-Roze, H.; Mouden, C.; Roume, J.; Abdelmajid, B.; Amram, D.; Baumann, C.; Chassaing, N.; Coubes, C.; et al. Mutational spectrum in holoprosencephaly shows that fgf is a new major signaling pathway. Hum. Mutat. 2016. [Google Scholar] [CrossRef] [PubMed]
- Dupe, V.; Rochard, L.; Mercier, S.; Le Petillon, Y.; Gicquel, I.; Bendavid, C.; Bourrouillou, G.; Kini, U.; Thauvin-Robinet, C.; Bohan, T.P.; et al. Notch, a new signaling pathway implicated in holoprosencephaly. Hum. Mol. Genet. 2011, 20, 1122–1131. [Google Scholar] [CrossRef] [PubMed]
- Mercier, S.; David, V.; Ratie, L.; Gicquel, I.; Odent, S.; Dupe, V. Nodal and shh dose-dependent double inhibition promotes an hpe-like phenotype in chick embryos. Dis. Models Mech. 2013, 6, 537–543. [Google Scholar] [CrossRef] [PubMed]
- Gorivodsky, M.; Mukhopadhyay, M.; Wilsch-Braeuninger, M.; Phillips, M.; Teufel, A.; Kim, C.; Malik, N.; Huttner, W.; Westphal, H. Intraflagellar transport protein 172 is essential for primary cilia formation and plays a vital role in patterning the mammalian brain. Dev. Biol. 2009, 325, 24–32. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sena, E.; Feistel, K.; Durand, B.C. An Evolutionarily Conserved Network Mediates Development of the zona limitans intrathalamica, a Sonic Hedgehog-Secreting Caudal Forebrain Signaling Center. J. Dev. Biol. 2016, 4, 31. https://doi.org/10.3390/jdb4040031
Sena E, Feistel K, Durand BC. An Evolutionarily Conserved Network Mediates Development of the zona limitans intrathalamica, a Sonic Hedgehog-Secreting Caudal Forebrain Signaling Center. Journal of Developmental Biology. 2016; 4(4):31. https://doi.org/10.3390/jdb4040031
Chicago/Turabian StyleSena, Elena, Kerstin Feistel, and Béatrice C. Durand. 2016. "An Evolutionarily Conserved Network Mediates Development of the zona limitans intrathalamica, a Sonic Hedgehog-Secreting Caudal Forebrain Signaling Center" Journal of Developmental Biology 4, no. 4: 31. https://doi.org/10.3390/jdb4040031
APA StyleSena, E., Feistel, K., & Durand, B. C. (2016). An Evolutionarily Conserved Network Mediates Development of the zona limitans intrathalamica, a Sonic Hedgehog-Secreting Caudal Forebrain Signaling Center. Journal of Developmental Biology, 4(4), 31. https://doi.org/10.3390/jdb4040031