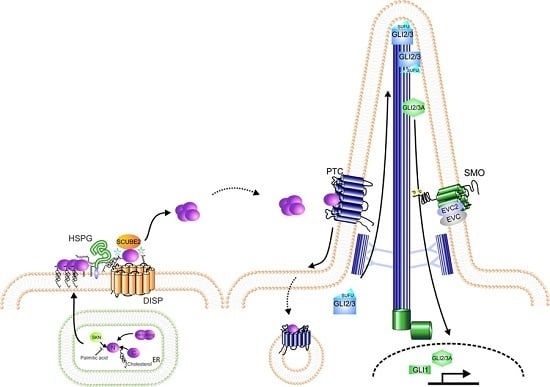
Regulation of Hedgehog Signalling Inside and Outside the Cell
Abstract
:1. Introduction
2. Hedgehog Expression
3. Ligand Synthesis
4. Ligand Release and Dispersal
5. Heparan Sulfate Proteoglycans
6. Regulation by SULF1
7. Regulation at the Membrane
8. Downstream of Smo
9. The Primary Cilium
10. Perspectives
Acknowledgments
Conflicts of Interest
References
- Mohler, J. Requirements for hedgehog, a segmental polarity gene, in patterning larval and adult cuticle of drosophila. Genetics 1988, 120, 1061–1072. [Google Scholar] [PubMed]
- Krauss, S.; Concordet, J.P.; Ingham, P.W. A functionally conserved homolog of the drosophila segment polarity gene hh is expressed in tissues with polarizing activity in zebrafish embryos. Cell 1993, 75, 1431–1444. [Google Scholar] [CrossRef]
- Riddle, R.D.; Johnson, R.L.; Laufer, E.; Tabin, C. Sonic hedgehog mediates the polarizing activity of the ZPA. Cell 1993, 75, 1401–1416. [Google Scholar] [CrossRef]
- Roelink, H.; Augsburger, A.; Heemskerk, J.; Korzh, V.; Norlin, S.; Ruiz i Altaba, A.; Tanabe, Y.; Placzek, M.; Edlund, T.; Jessell, T.M.; et al. Floor plate and motor neuron induction by vhh-1, a vertebrate homolog of hedgehog expressed by the notochord. Cell 1994, 76, 761–775. [Google Scholar] [CrossRef]
- Briscoe, J.; Small, S. Morphogen rules: Design principles of gradient-mediated embryo patterning. Development 2015, 142, 3996–4009. [Google Scholar] [CrossRef] [PubMed]
- Huangfu, D.; Liu, A.; Rakeman, A.S.; Murcia, N.S.; Niswander, L.; Anderson, K.V. Hedgehog signalling in the mouse requires intraflagellar transport proteins. Nature 2003, 426, 83–87. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.J.; Ekker, S.C.; von Kessler, D.P.; Porter, J.A.; Sun, B.I.; Beachy, P.A. Autoproteolysis in hedgehog protein biogenesis. Science 1994, 266, 1528–1537. [Google Scholar] [CrossRef] [PubMed]
- Porter, J.A.; Young, K.E.; Beachy, P.A. Cholesterol modification of hedgehog signaling proteins in animal development. Science 1996, 274, 255–259. [Google Scholar] [CrossRef] [PubMed]
- Pepinsky, R.B.; Zeng, C.; Wen, D.; Rayhorn, P.; Baker, D.P.; Williams, K.P.; Bixler, S.A.; Ambrose, C.M.; Garber, E.A.; Miatkowski, K.; et al. Identification of a palmitic acid-modified form of human sonic hedgehog. J. Biol. Chem. 1998, 273, 14037–14045. [Google Scholar] [CrossRef] [PubMed]
- Ingham, P.W.; Nakano, Y.; Seger, C. Mechanisms and functions of hedgehog signalling across the metazoa. Nat. Rev. Genet. 2011, 12, 393–406. [Google Scholar] [CrossRef] [PubMed]
- Jenkins, D. Hedgehog signalling: Emerging evidence for non-canonical pathways. Cell. Signal. 2009, 21, 1023–1034. [Google Scholar] [CrossRef] [PubMed]
- Robbins, D.J.; Fei, D.L.; Riobo, N.A. The hedgehog signal transduction network. Sci. Signal. 2012, 5, re6. [Google Scholar] [CrossRef] [PubMed]
- Tabata, T.; Eaton, S.; Kornberg, T.B. The drosophila hedgehog gene is expressed specifically in posterior compartment cells and is a target of engrailed regulation. Genes Dev. 1992, 6, 2635–2645. [Google Scholar] [CrossRef] [PubMed]
- Strutt, D.I.; Mlodzik, M. The regulation of hedgehog and decapentaplegic during drosophila eye imaginal disc development. Mech. Dev. 1996, 58, 39–50. [Google Scholar] [CrossRef]
- Bejarano, F.; Perez, L.; Apidianakis, Y.; Delidakis, C.; Milan, M. Hedgehog restricts its expression domain in the drosophila wing. EMBO Rep. 2007, 8, 778–783. [Google Scholar] [CrossRef] [PubMed]
- Tokusumi, Y.; Tokusumi, T.; Stoller-Conrad, J.; Schulz, R.A. Serpent, suppressor of hairless and u-shaped are crucial regulators of hedgehog niche expression and prohemocyte maintenance during drosophila larval hematopoiesis. Development 2010, 137, 3561–3568. [Google Scholar] [CrossRef] [PubMed]
- Epstein, D.J.; McMahon, A.P.; Joyner, A.L. Regionalization of sonic hedgehog transcription along the anteroposterior axis of the mouse central nervous system is regulated by hnf3-dependent and -independent mechanisms. Development 1999, 126, 281–292. [Google Scholar] [PubMed]
- Briscoe, J.; Pierani, A.; Jessell, T.M.; Ericson, J. A homeodomain protein code specifies progenitor cell identity and neuronal fate in the ventral neural tube. Cell 2000, 101, 435–445. [Google Scholar] [CrossRef]
- Cho, G.; Lim, Y.; Cho, I.T.; Simonet, J.C.; Golden, J.A. Arx together with foxa2, regulates shh floor plate expression. Dev. Biol. 2014, 393, 137–148. [Google Scholar] [CrossRef] [PubMed]
- Ruiz i Altaba, A.; Jessell, T.M.; Roelink, H. Restrictions to floor plate induction by hedgehog and winged-helix genes in the neural tube of frog embryos. Mol. Cell. Neurosci. 1995, 6, 106–121. [Google Scholar] [CrossRef] [PubMed]
- Lettice, L.A.; Heaney, S.J.; Purdie, L.A.; Li, L.; de Beer, P.; Oostra, B.A.; Goode, D.; Elgar, G.; Hill, R.E.; de Graaff, E. A long-range shh enhancer regulates expression in the developing limb and fin and is associated with preaxial polydactyly. Hum. Mol. Genet. 2003, 12, 1725–1735. [Google Scholar] [CrossRef] [PubMed]
- Lettice, L.A.; Williamson, I.; Wiltshire, J.H.; Peluso, S.; Devenney, P.S.; Hill, A.E.; Essafi, A.; Hagman, J.; Mort, R.; Grimes, G.; et al. Opposing functions of the ets factor family define shh spatial expression in limb buds and underlie polydactyly. Dev. Cell 2012, 22, 459–467. [Google Scholar] [CrossRef] [PubMed]
- Kitamura, K.; Yanazawa, M.; Sugiyama, N.; Miura, H.; Iizuka-Kogo, A.; Kusaka, M.; Omichi, K.; Suzuki, R.; Kato-Fukui, Y.; Kamiirisa, K.; et al. Mutation of arx causes abnormal development of forebrain and testes in mice and x-linked lissencephaly with abnormal genitalia in humans. Nat. Genet. 2002, 32, 359–369. [Google Scholar] [CrossRef] [PubMed]
- Hill, R.E.; Lettice, L.A. Alterations to the remote control of shh gene expression cause congenital abnormalities. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2013, 368, 20120357. [Google Scholar] [CrossRef] [PubMed]
- Johnson, E.J.; Neely, D.M.; Dunn, I.C.; Davey, M.G. Direct functional consequences of zrs enhancer mutation combine with secondary long range shh signalling effects to cause preaxial polydactyly. Dev. Biol. 2014, 392, 209–220. [Google Scholar] [CrossRef] [PubMed]
- Marsh, E.; Fulp, C.; Gomez, E.; Nasrallah, I.; Minarcik, J.; Sudi, J.; Christian, S.L.; Mancini, G.; Labosky, P.; Dobyns, W.; et al. Targeted loss of arx results in a developmental epilepsy mouse model and recapitulates the human phenotype in heterozygous females. Brain 2009, 132, 1563–1576. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Tukachinsky, H.; Huang, C.H.; Jao, C.; Chu, Y.R.; Tang, H.Y.; Mueller, B.; Schulman, S.; Rapoport, T.A.; Salic, A. Processing and turnover of the hedgehog protein in the endoplasmic reticulum. J. Cell Biol. 2011, 192, 825–838. [Google Scholar] [CrossRef] [PubMed]
- Briscoe, J.; Therond, P.P. The mechanisms of hedgehog signalling and its roles in development and disease. Nat. Rev. Mol. Cell. Biol. 2013, 14, 416–429. [Google Scholar] [CrossRef] [PubMed]
- Chamoun, Z.; Mann, R.K.; Nellen, D.; von Kessler, D.P.; Bellotto, M.; Beachy, P.A.; Basler, K. Skinny hedgehog, an acyltransferase required for palmitoylation and activity of the hedgehog signal. Science 2001, 293, 2080–2084. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.D.; Treisman, J.E. Sightless has homology to transmembrane acyltransferases and is required to generate active hedgehog protein. Curr. Biol. 2001, 11, 1147–1152. [Google Scholar] [CrossRef]
- Micchelli, C.A.; The, I.; Selva, E.; Mogila, V.; Perrimon, N. Rasp, a putative transmembrane acyltransferase, is required for hedgehog signaling. Development 2002, 129, 843–851. [Google Scholar] [PubMed]
- Buglino, J.A.; Resh, M.D. Hhat is a palmitoylacyltransferase with specificity for n-palmitoylation of sonic hedgehog. J. Biol. Chem. 2008, 283, 22076–22088. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.H.; Li, Y.J.; Kawakami, T.; Xu, S.M.; Chuang, P.T. Palmitoylation is required for the production of a soluble multimeric hedgehog protein complex and long-range signaling in vertebrates. Genes Dev. 2004, 18, 641–659. [Google Scholar] [CrossRef] [PubMed]
- Zeng, X.; Goetz, J.A.; Suber, L.M.; Scott, W.J., Jr.; Schreiner, C.M.; Robbins, D.J. A freely diffusible form of sonic hedgehog mediates long-range signalling. Nature 2001, 411, 716–720. [Google Scholar] [CrossRef] [PubMed]
- Taylor, F.R.; Wen, D.; Garber, E.A.; Carmillo, A.N.; Baker, D.P.; Arduini, R.M.; Williams, K.P.; Weinreb, P.H.; Rayhorn, P.; Hronowski, X.; et al. Enhanced potency of human sonic hedgehog by hydrophobic modification. Biochemistry 2001, 40, 4359–4371. [Google Scholar] [CrossRef] [PubMed]
- Bumcrot, D.A.; Takada, R.; McMahon, A.P. Proteolytic processing yields two secreted forms of sonic hedgehog. Mol. Cell. Biol. 1995, 15, 2294–2303. [Google Scholar] [CrossRef] [PubMed]
- Porter, J.A.; von Kessler, D.P.; Ekker, S.C.; Young, K.E.; Lee, J.J.; Moses, K.; Beachy, P.A. The product of hedgehog autoproteolytic cleavage active in local and long-range signalling. Nature 1995, 374, 363–366. [Google Scholar] [CrossRef] [PubMed]
- Porter, J.A.; Ekker, S.C.; Park, W.J.; von Kessler, D.P.; Young, K.E.; Chen, C.H.; Ma, Y.; Woods, A.S.; Cotter, R.J.; Koonin, E.V.; et al. Hedgehog patterning activity: Role of a lipophilic modification mediated by the carboxy-terminal autoprocessing domain. Cell 1996, 86, 21–34. [Google Scholar] [CrossRef]
- Briscoe, J.; Chen, Y.; Jessell, T.M.; Struhl, G. A hedgehog-insensitive form of patched provides evidence for direct long-range morphogen activity of sonic hedgehog in the neural tube. Mol. Cell 2001, 7, 1279–1291. [Google Scholar] [CrossRef]
- Burke, R.; Nellen, D.; Bellotto, M.; Hafen, E.; Senti, K.A.; Dickson, B.J.; Basler, K. Dispatched, a novel sterol-sensing domain protein dedicated to the release of cholesterol-modified hedgehog from signaling cells. Cell 1999, 99, 803–815. [Google Scholar] [CrossRef]
- Radhakrishnan, A.; Sun, L.P.; Kwon, H.J.; Brown, M.S.; Goldstein, J.L. Direct binding of cholesterol to the purified membrane region of scap: Mechanism for a sterol-sensing domain. Mol. Cell 2004, 15, 259–268. [Google Scholar] [CrossRef] [PubMed]
- Tukachinsky, H.; Kuzmickas, R.P.; Jao, C.Y.; Liu, J.; Salic, A. Dispatched and scube mediate the efficient secretion of the cholesterol-modified hedgehog ligand. Cell Rep. 2012, 2, 308–320. [Google Scholar] [CrossRef] [PubMed]
- Kawakami, T.; Kawcak, T.; Li, Y.J.; Zhang, W.; Hu, Y.; Chuang, P.T. Mouse dispatched mutants fail to distribute hedgehog proteins and are defective in hedgehog signaling. Development 2002, 129, 5753–5765. [Google Scholar] [CrossRef] [PubMed]
- Jakobs, P.; Exner, S.; Schurmann, S.; Pickhinke, U.; Bandari, S.; Ortmann, C.; Kupich, S.; Schulz, P.; Hansen, U.; Seidler, D.G.; et al. Scube2 enhances proteolytic shh processing from the surface of shh-producing cells. J. Cell Sci. 2014, 127, 1726–1737. [Google Scholar] [CrossRef] [PubMed]
- Ortmann, C.; Pickhinke, U.; Exner, S.; Ohlig, S.; Lawrence, R.; Jboor, H.; Dreier, R.; Grobe, K. Sonic hedgehog processing and release are regulated by glypican heparan sulfate proteoglycans. J. Cell Sci. 2015, 128, 2374–2385. [Google Scholar] [CrossRef] [PubMed]
- Gallet, A.; Ruel, L.; Staccini-Lavenant, L.; Therond, P.P. Cholesterol modification is necessary for controlled planar long-range activity of hedgehog in drosophila epithelia. Development 2006, 133, 407–418. [Google Scholar] [CrossRef] [PubMed]
- Goetz, J.A.; Singh, S.; Suber, L.M.; Kull, F.J.; Robbins, D.J. A highly conserved amino-terminal region of sonic hedgehog is required for the formation of its freely diffusible multimeric form. J. Biol. Chem. 2006, 281, 4087–4093. [Google Scholar] [CrossRef] [PubMed]
- Lewis, P.M.; Dunn, M.P.; McMahon, J.A.; Logan, M.; Martin, J.F.; St-Jacques, B.; McMahon, A.P. Cholesterol modification of sonic hedgehog is required for long-range signaling activity and effective modulation of signaling by ptc1. Cell 2001, 105, 599–612. [Google Scholar] [CrossRef]
- Matusek, T.; Wendler, F.; Poles, S.; Pizette, S.; D’Angelo, G.; Furthauer, M.; Therond, P.P. The escrt machinery regulates the secretion and long-range activity of hedgehog. Nature 2014, 516, 99–103. [Google Scholar] [CrossRef] [PubMed]
- Parchure, A.; Vyas, N.; Ferguson, C.; Parton, R.G.; Mayor, S. Oligomerization and endocytosis of hedgehog is necessary for its efficient exovesicular secretion. Mol. Biol. Cell 2015, 26, 4700–4717. [Google Scholar] [CrossRef] [PubMed]
- D’Angelo, G.; Matusek, T.; Pizette, S.; Therond, P.P. Endocytosis of hedgehog through dispatched regulates long-range signaling. Dev. Cell 2015, 32, 290–303. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, Y.; Okada, Y.; Hirokawa, N. Fgf-induced vesicular release of sonic hedgehog and retinoic acid in leftward nodal flow is critical for left-right determination. Nature 2005, 435, 172–177. [Google Scholar] [CrossRef] [PubMed]
- Liegeois, S.; Benedetto, A.; Garnier, J.M.; Schwab, Y.; Labouesse, M. The v0-atpase mediates apical secretion of exosomes containing hedgehog-related proteins in caenorhabditis elegans. J. Cell Biol. 2006, 173, 949–961. [Google Scholar] [CrossRef] [PubMed]
- Jakobs, P.; Schulz, P.; Ortmann, C.; Schurmann, S.; Exner, S.; Rebollido-Rios, R.; Dreier, R.; Seidler, D.G.; Grobe, K. Bridging the gap: Heparan sulfate and scube2 assemble sonic hedgehog release complexes at the surface of producing cells. Sci. Rep. 2016, 6, 26435. [Google Scholar] [CrossRef] [PubMed]
- Bernfield, M.; Gotte, M.; Park, P.W.; Reizes, O.; Fitzgerald, M.L.; Lincecum, J.; Zako, M. Functions of cell surface heparan sulfate proteoglycans. Annu. Rev. Biochem. 1999, 68, 729–777. [Google Scholar] [CrossRef] [PubMed]
- Lamanna, W.C.; Kalus, I.; Padva, M.; Baldwin, R.J.; Merry, C.L.; Dierks, T. The heparanome—The enigma of encoding and decoding heparan sulfate sulfation. J. Biotechnol. 2007, 129, 290–307. [Google Scholar] [CrossRef] [PubMed]
- Turnbull, J.; Powell, A.; Guimond, S. Heparan sulfate: Decoding a dynamic multifunctional cell regulator. Trends Cell Biol. 2001, 11, 75–82. [Google Scholar] [CrossRef]
- Lin, X. Functions of heparan sulfate proteoglycans in cell signaling during development. Development 2004, 131, 6009–6021. [Google Scholar] [CrossRef] [PubMed]
- Eugster, C.; Panakova, D.; Mahmoud, A.; Eaton, S. Lipoprotein-heparan sulfate interactions in the hh pathway. Dev. Cell 2007, 13, 57–71. [Google Scholar] [CrossRef] [PubMed]
- Panakova, D.; Sprong, H.; Marois, E.; Thiele, C.; Eaton, S. Lipoprotein particles are required for hedgehog and wingless signalling. Nature 2005, 435, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Vyas, N.; Goswami, D.; Manonmani, A.; Sharma, P.; Ranganath, H.A.; VijayRaghavan, K.; Shashidhara, L.S.; Sowdhamini, R.; Mayor, S. Nanoscale organization of hedgehog is essential for long-range signaling. Cell 2008, 133, 1214–1227. [Google Scholar] [CrossRef] [PubMed]
- Chan, J.A.; Balasubramanian, S.; Witt, R.M.; Nazemi, K.J.; Choi, Y.; Pazyra-Murphy, M.F.; Walsh, C.O.; Thompson, M.; Segal, R.A. Proteoglycan interactions with sonic hedgehog specify mitogenic responses. Nat. Neurosci. 2009, 12, 409–417. [Google Scholar] [CrossRef] [PubMed]
- Whalen, D.M.; Malinauskas, T.; Gilbert, R.J.; Siebold, C. Structural insights into proteoglycan-shaped hedgehog signaling. Proc. Natl. Acad. Sci. USA 2013, 110, 16420–16425. [Google Scholar] [CrossRef] [PubMed]
- Pellegrini, L. Role of heparan sulfate in fibroblast growth factor signalling: A structural view. Curr. Opin. Struct. Biol. 2001, 11, 629–634. [Google Scholar] [CrossRef]
- Filmus, J.; Capurro, M. The role of glypicans in hedgehog signaling. Matrix Biol. 2014, 35, 248–252. [Google Scholar] [CrossRef] [PubMed]
- Capurro, M.I.; Xu, P.; Shi, W.; Li, F.; Jia, A.; Filmus, J. Glypican-3 inhibits hedgehog signaling during development by competing with patched for hedgehog binding. Dev. Cell 2008, 14, 700–711. [Google Scholar] [CrossRef] [PubMed]
- Gallet, A.; Staccini-Lavenant, L.; Therond, P.P. Cellular trafficking of the glypican dally-like is required for full-strength hedgehog signaling and wingless transcytosis. Dev. Cell 2008, 14, 712–725. [Google Scholar] [CrossRef] [PubMed]
- Witt, R.M.; Hecht, M.L.; Pazyra-Murphy, M.F.; Cohen, S.M.; Noti, C.; van Kuppevelt, T.H.; Fuller, M.; Chan, J.A.; Hopwood, J.J.; Seeberger, P.H.; et al. Heparan sulfate proteoglycans containing a glypican 5 core and 2-o-sulfo-iduronic acid function as sonic hedgehog co-receptors to promote proliferation. J. Biol. Chem. 2013, 288, 26275–26288. [Google Scholar] [CrossRef] [PubMed]
- Touahri, Y.; Escalas, N.; Benazeraf, B.; Cochard, P.; Danesin, C.; Soula, C. Sulfatase 1 promotes the motor neuron-to-oligodendrocyte fate switch by activating shh signaling in olig2 progenitors of the embryonic ventral spinal cord. J. Neurosci. 2012, 32, 18018–18034. [Google Scholar] [CrossRef] [PubMed]
- Danesin, C.; Agius, E.; Escalas, N.; Ai, X.; Emerson, C.; Cochard, P.; Soula, C. Ventral neural progenitors switch toward an oligodendroglial fate in response to increased sonic hedgehog (shh) activity: Involvement of sulfatase 1 in modulating shh signaling in the ventral spinal cord. J. Neurosci. 2006, 26, 5037–5048. [Google Scholar] [CrossRef] [PubMed]
- Al Oustah, A.; Danesin, C.; Khouri-Farah, N.; Farreny, M.A.; Escalas, N.; Cochard, P.; Glise, B.; Soula, C. Dynamics of sonic hedgehog signaling in the ventral spinal cord are controlled by intrinsic changes in source cells requiring sulfatase 1. Development 2014, 141, 1392–1403. [Google Scholar] [CrossRef] [PubMed]
- Ramsbottom, S.A.; Maguire, R.J.; Fellgett, S.W.; Pownall, M.E. Sulf1 influences the shh morphogen gradient during the dorsal ventral patterning of the neural tube in xenopus tropicalis. Dev. Biol. 2014, 391, 207–218. [Google Scholar] [CrossRef] [PubMed]
- Ai, X.; Do, A.T.; Kusche-Gullberg, M.; Lindahl, U.; Lu, K.; Emerson, C.P., Jr. Substrate specificity and domain functions of extracellular heparan sulfate 6-o-endosulfatases, qsulf1 and qsulf2. J. Biol. Chem. 2006, 281, 4969–4976. [Google Scholar] [CrossRef] [PubMed]
- Wojcinski, A.; Nakato, H.; Soula, C.; Glise, B. Dsulfatase-1 fine-tunes hedgehog patterning activity through a novel regulatory feedback loop. Dev. Biol. 2011, 358, 168–180. [Google Scholar] [CrossRef] [PubMed]
- Bellaiche, Y.; The, I.; Perrimon, N. Tout-velu is a drosophila homologue of the putative tumour suppressor ext-1 and is needed for hh diffusion. Nature 1998, 394, 85–88. [Google Scholar] [PubMed]
- Han, C.; Belenkaya, T.Y.; Khodoun, M.; Tauchi, M.; Lin, X.; Lin, X. Distinct and collaborative roles of drosophila ext family proteins in morphogen signalling and gradient formation. Development 2004, 131, 1563–1575. [Google Scholar] [CrossRef] [PubMed]
- Ohlig, S.; Pickhinke, U.; Sirko, S.; Bandari, S.; Hoffmann, D.; Dreier, R.; Farshi, P.; Gotz, M.; Grobe, K. An emerging role of sonic hedgehog shedding as a modulator of heparan sulfate interactions. J. Biol. Chem. 2012, 287, 43708–43719. [Google Scholar] [CrossRef] [PubMed]
- Ingham, P.W.; Taylor, A.M.; Nakano, Y. Role of the drosophila patched gene in positional signalling. Nature 1991, 353, 184–187. [Google Scholar] [CrossRef] [PubMed]
- Carpenter, D.; Stone, D.M.; Brush, J.; Ryan, A.; Armanini, M.; Frantz, G.; Rosenthal, A.; de Sauvage, F.J. Characterization of two patched receptors for the vertebrate hedgehog protein family. Proc. Natl. Acad. Sci. USA 1998, 95, 13630–13634. [Google Scholar] [CrossRef] [PubMed]
- Nieuwenhuis, E.; Motoyama, J.; Barnfield, P.C.; Yoshikawa, Y.; Zhang, X.; Mo, R.; Crackower, M.A.; Hui, C.C. Mice with a targeted mutation of patched2 are viable but develop alopecia and epidermal hyperplasia. Mol. Cell. Biol. 2006, 26, 6609–6622. [Google Scholar] [CrossRef] [PubMed]
- Beachy, P.A.; Hymowitz, S.G.; Lazarus, R.A.; Leahy, D.J.; Siebold, C. Interactions between hedgehog proteins and their binding partners come into view. Genes Dev. 2010, 24, 2001–2012. [Google Scholar] [CrossRef] [PubMed]
- Allen, B.L.; Song, J.Y.; Izzi, L.; Althaus, I.W.; Kang, J.S.; Charron, F.; Krauss, R.S.; McMahon, A.P. Overlapping roles and collective requirement for the coreceptors gas1, cdo, and boc in shh pathway function. Dev. Cell 2011, 20, 775–787. [Google Scholar] [CrossRef] [PubMed]
- Chuang, P.T.; McMahon, A.P. Vertebrate hedgehog signalling modulated by induction of a hedgehog-binding protein. Nature 1999, 397, 617–621. [Google Scholar] [CrossRef] [PubMed]
- Brazil, D.P.; Church, R.H.; Surae, S.; Godson, C.; Martin, F. Bmp signalling: Agony and antagony in the family. Trends Cell Biol. 2015, 25, 249–264. [Google Scholar] [CrossRef] [PubMed]
- Holtz, A.M.; Griffiths, S.C.; Davis, S.J.; Bishop, B.; Siebold, C.; Allen, B.L. Secreted hhip1 interacts with heparan sulfate and regulates hedgehog ligand localization and function. J. Cell Biol. 2015, 209, 739–757. [Google Scholar] [CrossRef] [PubMed]
- Kwong, L.; Bijlsma, M.F.; Roelink, H. Shh-mediated degradation of hhip allows cell autonomous and non-cell autonomous shh signalling. Nat. Commun. 2014, 5, 4849. [Google Scholar] [CrossRef] [PubMed]
- Bourikas, D.; Pekarik, V.; Baeriswyl, T.; Grunditz, A.; Sadhu, R.; Nardo, M.; Stoeckli, E.T. Sonic hedgehog guides commissural axons along the longitudinal axis of the spinal cord. Nat. Neurosci. 2005, 8, 297–304. [Google Scholar] [CrossRef] [PubMed]
- Wilson, N.H.; Stoeckli, E.T. Sonic hedgehog regulates its own receptor on postcrossing commissural axons in a glypican1-dependent manner. Neuron 2013, 79, 478–491. [Google Scholar] [CrossRef] [PubMed]
- Alcedo, J.; Ayzenzon, M.; von Ohlen, T.; Noll, M.; Hooper, J.E. The drosophila smoothened gene encodes a seven-pass membrane protein, a putative receptor for the hedgehog signal. Cell 1996, 86, 221–232. [Google Scholar] [CrossRef]
- Van den Heuvel, M.; Ingham, P.W. Smoothened encodes a receptor-like serpentine protein required for hedgehog signalling. Nature 1996, 382, 547–551. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Burgess, S.; Hopkins, N. Analysis of the zebrafish smoothened mutant reveals conserved and divergent functions of hedgehog activity. Development 2001, 128, 2385–2396. [Google Scholar] [PubMed]
- Goodrich, L.V.; Milenkovic, L.; Higgins, K.M.; Scott, M.P. Altered neural cell fates and medulloblastoma in mouse patched mutants. Science 1997, 277, 1109–1113. [Google Scholar] [CrossRef] [PubMed]
- Denef, N.; Neubuser, D.; Perez, L.; Cohen, S.M. Hedgehog induces opposite changes in turnover and subcellular localization of patched and smoothened. Cell 2000, 102, 521–531. [Google Scholar] [CrossRef]
- Incardona, J.P.; Gruenberg, J.; Roelink, H. Sonic hedgehog induces the segregation of patched and smoothened in endosomes. Curr. Biol. 2002, 12, 983–995. [Google Scholar] [CrossRef]
- Taipale, J.; Cooper, M.K.; Maiti, T.; Beachy, P.A. Patched acts catalytically to suppress the activity of smoothened. Nature 2002, 418, 892–897. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.K.; Taipale, J.; Cooper, M.K.; Beachy, P.A. Inhibition of hedgehog signaling by direct binding of cyclopamine to smoothened. Genes Dev. 2002, 16, 2743–2748. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.K.; Taipale, J.; Young, K.E.; Maiti, T.; Beachy, P.A. Small molecule modulation of smoothened activity. Proc. Natl. Acad. Sci. USA 2002, 99, 14071–14076. [Google Scholar] [CrossRef] [PubMed]
- Rohatgi, R.; Scott, M.P. Patching the gaps in hedgehog signalling. Nat. Cell. Biol. 2007, 9, 1005–1009. [Google Scholar] [CrossRef] [PubMed]
- Bijlsma, M.F.; Peppelenbosch, M.P.; Spek, C.A. A dual role for 7-dehydrocholesterol reductase in regulating hedgehog signalling? Development 2006, 133, 3951; author reply 3952–3953. [Google Scholar] [CrossRef] [PubMed]
- Nedelcu, D.; Liu, J.; Xu, Y.; Jao, C.; Salic, A. Oxysterol binding to the extracellular domain of smoothened in hedgehog signaling. Nat. Chem. Biol. 2013, 9, 557–564. [Google Scholar] [CrossRef] [PubMed]
- Dwyer, J.R.; Sever, N.; Carlson, M.; Nelson, S.F.; Beachy, P.A.; Parhami, F. Oxysterols are novel activators of the hedgehog signaling pathway in pluripotent mesenchymal cells. J. Biol. Chem. 2007, 282, 8959–8968. [Google Scholar] [CrossRef] [PubMed]
- Khaliullina, H.; Panakova, D.; Eugster, C.; Riedel, F.; Carvalho, M.; Eaton, S. Patched regulates smoothened trafficking using lipoprotein-derived lipids. Development 2009, 136, 4111–4121. [Google Scholar] [CrossRef] [PubMed]
- Yavari, A.; Nagaraj, R.; Owusu-Ansah, E.; Folick, A.; Ngo, K.; Hillman, T.; Call, G.; Rohatgi, R.; Scott, M.P.; Banerjee, U. Role of lipid metabolism in smoothened derepression in hedgehog signaling. Dev. Cell 2010, 19, 54–65. [Google Scholar] [CrossRef] [PubMed]
- Khaliullina, H.; Bilgin, M.; Sampaio, J.L.; Shevchenko, A.; Eaton, S. Endocannabinoids are conserved inhibitors of the hedgehog pathway. Proc. Natl. Acad. Sci. USA 2015, 112, 3415–3420. [Google Scholar] [CrossRef] [PubMed]
- Jia, J.; Tong, C.; Wang, B.; Luo, L.; Jiang, J. Hedgehog signalling activity of smoothened requires phosphorylation by protein kinase a and casein kinase i. Nature 2004, 432, 1045–1050. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Tong, C.; Jiang, J. Hedgehog regulates smoothened activity by inducing a conformational switch. Nature 2007, 450, 252–258. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Sasai, N.; Ma, G.; Yue, T.; Jia, J.; Briscoe, J.; Jiang, J. Sonic hedgehog dependent phosphorylation by ck1alpha and grk2 is required for ciliary accumulation and activation of smoothened. PLoS Biol. 2011, 9, e1001083. [Google Scholar] [CrossRef] [PubMed]
- Apionishev, S.; Katanayeva, N.M.; Marks, S.A.; Kalderon, D.; Tomlinson, A. Drosophila smoothened phosphorylation sites essential for hedgehog signal transduction. Nat. Cell Biol. 2005, 7, 86–92. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Ospina, J.K.; Zhang, J.; Michelson, A.P.; Schoen, A.M.; Zhu, A.J. Sequential phosphorylation of smoothened transduces graded hedgehog signaling. Sci. Signal. 2011, 4, ra43. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Ma, G.; Wang, B.; Jiang, J. Hedgehog induces formation of pka-smoothened complexes to promote smoothened phosphorylation and pathway activation. Sci. Signal. 2014, 7, ra62. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Li, S.; Tong, C.; Zhao, Y.; Wang, B.; Liu, Y.; Jia, J.; Jiang, J. G protein-coupled receptor kinase 2 promotes high-level hedgehog signaling by regulating the active state of smo through kinase-dependent and kinase-independent mechanisms in drosophila. Genes Dev. 2010, 24, 2054–2067. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.; Liu, Y.; Xia, R.; Tong, C.; Yue, T.; Jiang, J.; Jia, J. Casein kinase 2 promotes hedgehog signaling by regulating both smoothened and cubitus interruptus. J. Biol. Chem. 2010, 285, 37218–37226. [Google Scholar] [CrossRef] [PubMed]
- Maier, D.; Cheng, S.; Faubert, D.; Hipfner, D.R. A broadly conserved g-protein-coupled receptor kinase phosphorylation mechanism controls drosophila smoothened activity. PLoS Genet. 2014, 10, e1004399. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Lee, R.T.; Pusapati, G.V.; Iyu, A.; Rohatgi, R.; Ingham, P.W. An essential role for grk2 in hedgehog signalling downstream of smoothened. EMBO Rep. 2016, 17, 739–752. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Li, S.; Han, Y.; Tong, C.; Wang, B.; Chen, Y.; Jiang, J. Regulation of smoothened phosphorylation and high-level hedgehog signaling activity by a plasma membrane associated kinase. PLoS Biol. 2016, 14, e1002481. [Google Scholar] [CrossRef] [PubMed]
- Forbes, A.J.; Nakano, Y.; Taylor, A.M.; Ingham, P.W. Genetic analysis of hedgehog signalling in the drosophila embryo. Dev. Suppl. 1993, 115–124. [Google Scholar]
- Aza-Blanc, P.; Ramirez-Weber, F.A.; Laget, M.P.; Schwartz, C.; Kornberg, T.B. Proteolysis that is inhibited by hedgehog targets cubitus interruptus protein to the nucleus and converts it to a repressor. Cell 1997, 89, 1043–1053. [Google Scholar] [CrossRef]
- Hooper, J.E.; Scott, M.P. Communicating with hedgehogs. Nat. Rev. Mol. Cell. Biol. 2005, 6, 306–317. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Amanai, K.; Wang, B.; Jiang, J. Interactions with costal2 and suppressor of fused regulate nuclear translocation and activity of cubitus interruptus. Genes Dev. 2000, 14, 2893–2905. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.T.; Holmgren, R.A. The subcellular localization and activity of drosophila cubitus interruptus are regulated at multiple levels. Development 1999, 126, 5097–5106. [Google Scholar] [PubMed]
- Wang, G.; Jiang, J. Multiple cos2/ci interactions regulate ci subcellular localization through microtubule dependent and independent mechanisms. Dev. Biol. 2004, 268, 493–505. [Google Scholar] [CrossRef] [PubMed]
- Ogden, S.K.; Ascano, M., Jr.; Stegman, M.A.; Suber, L.M.; Hooper, J.E.; Robbins, D.J. Identification of a functional interaction between the transmembrane protein smoothened and the kinesin-related protein costal2. Curr. Biol. 2003, 13, 1998–2003. [Google Scholar] [CrossRef] [PubMed]
- Sisson, J.C.; Ho, K.S.; Suyama, K.; Scott, M.P. Costal2, a novel kinesin-related protein in the hedgehog signaling pathway. Cell 1997, 90, 235–245. [Google Scholar] [CrossRef]
- Robbins, D.J.; Nybakken, K.E.; Kobayashi, R.; Sisson, J.C.; Bishop, J.M.; Therond, P.P. Hedgehog elicits signal transduction by means of a large complex containing the kinesin-related protein costal2. Cell 1997, 90, 225–234. [Google Scholar] [CrossRef]
- Zhang, W.; Zhao, Y.; Tong, C.; Wang, G.; Wang, B.; Jia, J.; Jiang, J. Hedgehog-regulated costal2-kinase complexes control phosphorylation and proteolytic processing of cubitus interruptus. Dev. Cell 2005, 8, 267–278. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Gallaher, N.; Goodman, R.H.; Smolik, S.M. Protein kinase a directly regulates the activity and proteolysis of cubitus interruptus. Proc. Natl. Acad. Sci. USA 1998, 95, 2349–2354. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Wang, B.; Jiang, J. Protein kinase a antagonizes hedgehog signaling by regulating both the activator and repressor forms of cubitus interruptus. Genes Dev. 1999, 13, 2828–2837. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.; Struhl, G. Protein kinase A and hedgehog signaling in drosophila limb development. Cell 1995, 80, 563–572. [Google Scholar] [CrossRef]
- Jiang, J.; Struhl, G. Regulation of the hedgehog and wingless signalling pathways by the f-box/wd40-repeat protein slimb. Nature 1998, 391, 493–496. [Google Scholar] [CrossRef] [PubMed]
- Methot, N.; Basler, K. Suppressor of fused opposes hedgehog signal transduction by impeding nuclear accumulation of the activator form of cubitus interruptus. Development 2000, 127, 4001–4010. [Google Scholar] [PubMed]
- Han, Y.; Shi, Q.; Jiang, J. Multisite interaction with sufu regulates Ci/Gli activity through distinct mechanisms in hh signal transduction. Proc. Natl. Acad. Sci. USA 2015, 112, 6383–6388. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Kalderon, D. Hedgehog activates fused through phosphorylation to elicit a full spectrum of pathway responses. Dev. Cell 2011, 20, 802–814. [Google Scholar] [CrossRef] [PubMed]
- Akimaru, H.; Chen, Y.; Dai, P.; Hou, D.X.; Nonaka, M.; Smolik, S.M.; Armstrong, S.; Goodman, R.H.; Ishii, S. Drosophila CBP is a co-activator of cubitus interruptus in hedgehog signalling. Nature 1997, 386, 735–738. [Google Scholar] [CrossRef] [PubMed]
- Ruiz i Altaba, A. Combinatorial gli gene function in floor plate and neuronal inductions by sonic hedgehog. Development 1998, 125, 2203–2212. [Google Scholar] [PubMed]
- Pan, Y.; Bai, C.B.; Joyner, A.L.; Wang, B. Sonic hedgehog signaling regulates gli2 transcriptional activity by suppressing its processing and degradation. Mol. Cell. Biol. 2006, 26, 3365–3377. [Google Scholar] [CrossRef] [PubMed]
- Dai, P.; Akimaru, H.; Tanaka, Y.; Maekawa, T.; Nakafuku, M.; Ishii, S. Sonic hedgehog-induced activation of the gli1 promoter is mediated by gli3. J. Biol. Chem. 1999, 274, 8143–8152. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, H.; Nishizaki, Y.; Hui, C.; Nakafuku, M.; Kondoh, H. Regulation of gli2 and gli3 activities by an amino-terminal repression domain: Implication of gli2 and gli3 as primary mediators of shh signaling. Development 1999, 126, 3915–3924. [Google Scholar] [PubMed]
- Wang, B.; Li, Y. Evidence for the direct involvement of {beta}trcp in gli3 protein processing. Proc. Natl. Acad. Sci. USA 2006, 103, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Bhatia, N.; Thiyagarajan, S.; Elcheva, I.; Saleem, M.; Dlugosz, A.; Mukhtar, H.; Spiegelman, V.S. Gli2 is targeted for ubiquitination and degradation by beta-trcp ubiquitin ligase. J. Biol. Chem. 2006, 281, 19320–19326. [Google Scholar] [CrossRef] [PubMed]
- Tempé, D.; Casas, M.; Karaz, S.; Blanchet-Tournier, M.F.; Concordet, J.P. Multisite protein kinase a and glycogen synthase kinase 3beta phosphorylation leads to gli3 ubiquitination by scfbetatrcp. Mol. Cell. Biol. 2006, 26, 4316–4326. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Wang, B. A novel protein-processing domain in gli2 and gli3 differentially blocks complete protein degradation by the proteasome. J. Biol. Chem. 2007, 282, 10846–10852. [Google Scholar] [CrossRef] [PubMed]
- Schrader, E.K.; Harstad, K.G.; Holmgren, R.A.; Matouschek, A. A three-part signal governs differential processing of gli1 and gli3 proteins by the proteasome. J. Biol. Chem. 2011, 286, 39051–39058. [Google Scholar] [CrossRef] [PubMed]
- Kogerman, P.; Grimm, T.; Kogerman, L.; Krause, D.; Unden, A.B.; Sandstedt, B.; Toftgard, R.; Zaphiropoulos, P.G. Mammalian suppressor-of-fused modulates nuclear-cytoplasmic shuttling of gli-1. Nat. Cell Biol. 1999, 1, 312–319. [Google Scholar] [PubMed]
- Liu, X.; Wang, X.; Du, W.; Chen, L.; Wang, G.; Cui, Y.; Liu, Y.; Dou, Z.; Wang, H.; Zhang, P.; et al. Suppressor of fused (sufu) represses gli1 transcription and nuclear accumulation, inhibits glioma cell proliferation, invasion and vasculogenic mimicry, improving glioma chemo-sensitivity and prognosis. Oncotarget 2014, 5, 11681–11694. [Google Scholar] [CrossRef] [PubMed]
- Ding, Q.; Fukami, S.; Meng, X.; Nishizaki, Y.; Zhang, X.; Sasaki, H.; Dlugosz, A.; Nakafuku, M.; Hui, C. Mouse suppressor of fused is a negative regulator of sonic hedgehog signaling and alters the subcellular distribution of gli1. Curr. Biol. 1999, 9, 1119–1122. [Google Scholar] [CrossRef]
- Tukachinsky, H.; Lopez, L.V.; Salic, A. A mechanism for vertebrate hedgehog signaling: Recruitment to cilia and dissociation of sufu-gli protein complexes. J. Cell Biol. 2010, 191, 415–428. [Google Scholar] [CrossRef] [PubMed]
- Shimokawa, T.; Tostar, U.; Lauth, M.; Palaniswamy, R.; Kasper, M.; Toftgard, R.; Zaphiropoulos, P.G. Novel human glioma-associated oncogene 1 (gli1) splice variants reveal distinct mechanisms in the terminal transduction of the hedgehog signal. J. Biol. Chem. 2008, 283, 14345–14354. [Google Scholar] [CrossRef] [PubMed]
- Lo, H.W.; Zhu, H.; Cao, X.; Aldrich, A.; Ali-Osman, F. A novel splice variant of gli1 that promotes glioblastoma cell migration and invasion. Cancer Res. 2009, 69, 6790–6798. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.; Geradts, J.; Dewhirst, M.W.; Lo, H.W. Upregulation of VEGF-A and CD24 gene expression by the tgli1 transcription factor contributes to the aggressive behavior of breast cancer cells. Oncogene 2012, 31, 104–115. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Platt, K.A.; Censullo, P.; Ruiz i Altaba, A. Gli1 is a target of sonic hedgehog that induces ventral neural tube development. Development 1997, 124, 2537–2552. [Google Scholar] [PubMed]
- Carpenter, R.L.; Lo, H.W. Hedgehog pathway and gli1 isoforms in human cancer. Discov. Med. 2012, 13, 105–113. [Google Scholar] [PubMed]
- Chen, M.H.; Gao, N.; Kawakami, T.; Chuang, P.T. Mice deficient in the fused homolog do not exhibit phenotypes indicative of perturbed hedgehog signaling during embryonic development. Mol. Cell. Biol. 2005, 25, 7042–7053. [Google Scholar] [CrossRef] [PubMed]
- Preat, T. Characterization of suppressor of fused, a complete suppressor of the fused segment polarity gene of drosophila melanogaster. Genetics 1992, 132, 725–736. [Google Scholar] [PubMed]
- Cooper, A.F.; Yu, K.P.; Brueckner, M.; Brailey, L.L.; Johnson, L.; McGrath, J.M.; Bale, A.E. Cardiac and cns defects in a mouse with targeted disruption of suppressor of fused. Development 2005, 132, 4407–4417. [Google Scholar] [CrossRef] [PubMed]
- Svard, J.; Heby-Henricson, K.; Persson-Lek, M.; Rozell, B.; Lauth, M.; Bergstrom, A.; Ericson, J.; Toftgard, R.; Teglund, S. Genetic elimination of suppressor of fused reveals an essential repressor function in the mammalian hedgehog signaling pathway. Dev. Cell 2006, 10, 187–197. [Google Scholar] [CrossRef] [PubMed]
- Varjosalo, M.; Li, S.P.; Taipale, J. Divergence of hedgehog signal transduction mechanism between drosophila and mammals. Dev. Cell 2006, 10, 177–186. [Google Scholar] [CrossRef] [PubMed]
- Goetz, S.C.; Anderson, K.V. The primary cilium: A signalling centre during vertebrate development. Nat. Rev. Genet. 2010, 11, 331–344. [Google Scholar] [CrossRef] [PubMed]
- Szymanska, K.; Johnson, C.A. The transition zone: An essential functional compartment of cilia. Cilia 2012, 1, 10. [Google Scholar] [CrossRef] [PubMed]
- Chih, B.; Liu, P.; Chinn, Y.; Chalouni, C.; Komuves, L.G.; Hass, P.E.; Sandoval, W.; Peterson, A.S. A ciliopathy complex at the transition zone protects the cilia as a privileged membrane domain. Nat. Cell Biol. 2012, 14, 61–72. [Google Scholar] [CrossRef] [PubMed]
- Coppieters, F.; Lefever, S.; Leroy, B.P.; de Baere, E. Cep290, a gene with many faces: Mutation overview and presentation of cep290base. Hum. Mutat. 2010, 31, 1097–1108. [Google Scholar] [CrossRef] [PubMed]
- Ramsbottom, S.; Miles, C.; Sayer, J. Murine cep290 phenotypes are modified by genetic backgrounds and provide an impetus for investigating disease modifier alleles. F1000Res 2015, 4, 590. [Google Scholar] [CrossRef] [PubMed]
- Wen, X.; Lai, C.K.; Evangelista, M.; Hongo, J.A.; de Sauvage, F.J.; Scales, S.J. Kinetics of hedgehog-dependent full-length gli3 accumulation in primary cilia and subsequent degradation. Mol. Cell. Biol. 2010, 30, 1910–1922. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Kato, M.; Beachy, P.A. Gli2 trafficking links hedgehog-dependent activation of smoothened in the primary cilium to transcriptional activation in the nucleus. Proc. Natl. Acad. Sci. USA 2009, 106, 21666–21671. [Google Scholar] [CrossRef] [PubMed]
- Humke, E.W.; Dorn, K.V.; Milenkovic, L.; Scott, M.P.; Rohatgi, R. The output of hedgehog signaling is controlled by the dynamic association between suppressor of fused and the gli proteins. Genes Dev. 2010, 24, 670–682. [Google Scholar] [CrossRef] [PubMed]
- Hynes, A.M.; Giles, R.H.; Srivastava, S.; Eley, L.; Whitehead, J.; Danilenko, M.; Raman, S.; Slaats, G.G.; Colville, J.G.; Ajzenberg, H.; et al. Murine joubert syndrome reveals hedgehog signaling defects as a potential therapeutic target for nephronophthisis. Proc. Natl. Acad. Sci. USA 2014, 111, 9893–9898. [Google Scholar] [CrossRef] [PubMed]
- Barzi, M.; Berenguer, J.; Menendez, A.; Alvarez-Rodriguez, R.; Pons, S. Sonic-hedgehog-mediated proliferation requires the localization of pka to the cilium base. J. Cell Sci. 2010, 123, 62–69. [Google Scholar] [CrossRef] [PubMed]
- Fumoto, K.; Hoogenraad, C.C.; Kikuchi, A. Gsk-3beta-regulated interaction of bicd with dynein is involved in microtubule anchorage at centrosome. EMBO J. 2006, 25, 5670–5682. [Google Scholar] [CrossRef] [PubMed]
- Sillibourne, J.E.; Milne, D.M.; Takahashi, M.; Ono, Y.; Meek, D.W. Centrosomal anchoring of the protein kinase ck1delta mediated by attachment to the large, coiled-coil scaffolding protein cg-nap/akap450. J. Mol. Biol. 2002, 322, 785–797. [Google Scholar] [CrossRef]
- Tuson, M.; He, M.; Anderson, K.V. Protein kinase a acts at the basal body of the primary cilium to prevent gli2 activation and ventralization of the mouse neural tube. Development 2011, 138, 4921–4930. [Google Scholar] [CrossRef] [PubMed]
- Rohatgi, R.; Milenkovic, L.; Scott, M.P. Patched1 regulates hedgehog signaling at the primary cilium. Science 2007, 317, 372–376. [Google Scholar] [CrossRef] [PubMed]
- Corbit, K.C.; Aanstad, P.; Singla, V.; Norman, A.R.; Stainier, D.Y.; Reiter, J.F. Vertebrate smoothened functions at the primary cilium. Nature 2005, 437, 1018–1021. [Google Scholar] [CrossRef] [PubMed]
- Kovacs, J.J.; Whalen, E.J.; Liu, R.; Xiao, K.; Kim, J.; Chen, M.; Wang, J.; Chen, W.; Lefkowitz, R.J. Beta-arrestin-mediated localization of smoothened to the primary cilium. Science 2008, 320, 1777–1781. [Google Scholar] [CrossRef] [PubMed]
- Rohatgi, R.; Milenkovic, L.; Corcoran, R.B.; Scott, M.P. Hedgehog signal transduction by smoothened: Pharmacologic evidence for a 2-step activation process. Proc. Natl. Acad. Sci. USA 2009, 106, 3196–3201. [Google Scholar] [CrossRef] [PubMed]
- Pal, K.; Hwang, S.H.; Somatilaka, B.; Badgandi, H.; Jackson, P.K.; DeFea, K.; Mukhopadhyay, S. Smoothened determines beta-arrestin-mediated removal of the G protein-coupled receptor gpr161 from the primary cilium. J. Cell Biol. 2016, 212, 861–875. [Google Scholar] [CrossRef] [PubMed]
- Mukhopadhyay, S.; Wen, X.; Ratti, N.; Loktev, A.; Rangell, L.; Scales, S.J.; Jackson, P.K. The ciliary g-protein-coupled receptor gpr161 negatively regulates the sonic hedgehog pathway via camp signaling. Cell 2013, 152, 210–223. [Google Scholar] [CrossRef] [PubMed]
- Dorn, K.V.; Hughes, C.E.; Rohatgi, R. A smoothened-evc2 complex transduces the hedgehog signal at primary cilia. Dev. Cell 2012, 23, 823–835. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Chen, W.; Chen, Y.; Jiang, J. Smoothened transduces hedgehog signal by forming a complex with evc/evc2. Cell Res. 2012, 22, 1593–1604. [Google Scholar] [CrossRef] [PubMed]
- Haycraft, C.J.; Banizs, B.; Aydin-Son, Y.; Zhang, Q.; Michaud, E.J.; Yoder, B.K. Gli2 and gli3 localize to cilia and require the intraflagellar transport protein polaris for processing and function. PLoS Genet. 2005, 1, e53. [Google Scholar] [CrossRef] [PubMed]
- He, M.; Subramanian, R.; Bangs, F.; Omelchenko, T.; Liem, K.F., Jr.; Kapoor, T.M.; Anderson, K.V. The kinesin-4 protein kif7 regulates mammalian hedgehog signalling by organizing the cilium tip compartment. Nat. Cell Biol. 2014, 16, 663–672. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.; Wang, B.; Niswander, L.A. Mouse intraflagellar transport proteins regulate both the activator and repressor functions of gli transcription factors. Development 2005, 132, 3103–3111. [Google Scholar] [CrossRef] [PubMed]
- Stasiulewicz, M.; Gray, S.D.; Mastromina, I.; Silva, J.C.; Bjorklund, M.; Seymour, P.A.; Booth, D.; Thompson, C.; Green, R.J.; Hall, E.A.; et al. A conserved role for Notch signaling in priming the cellular response to Shh through ciliary localisation of the key Shh transducer Smo. Development 2015, 142, 2291–2303. [Google Scholar] [CrossRef] [PubMed]
- Kong, J.H.; Yang, L.; Dessaud, E.; Chuang, K.; Moore, D.M.; Rohatgi, R.; Briscoe, J.; Novitch, B.G. Notch activity modulates the responsiveness of neural progenitors to sonic hedgehog signaling. Dev. Cell 2015, 33, 373–387. [Google Scholar] [CrossRef] [PubMed]
- Sasai, N.; Kutejova, E.; Briscoe, J. Integration of signals along orthogonal axes of the vertebrate neural tube controls progenitor competence and increases cell diversity. PLoS Biol. 2014, 12, e1001907. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramsbottom, S.A.; Pownall, M.E. Regulation of Hedgehog Signalling Inside and Outside the Cell. J. Dev. Biol. 2016, 4, 23. https://doi.org/10.3390/jdb4030023
Ramsbottom SA, Pownall ME. Regulation of Hedgehog Signalling Inside and Outside the Cell. Journal of Developmental Biology. 2016; 4(3):23. https://doi.org/10.3390/jdb4030023
Chicago/Turabian StyleRamsbottom, Simon A., and Mary E. Pownall. 2016. "Regulation of Hedgehog Signalling Inside and Outside the Cell" Journal of Developmental Biology 4, no. 3: 23. https://doi.org/10.3390/jdb4030023
APA StyleRamsbottom, S. A., & Pownall, M. E. (2016). Regulation of Hedgehog Signalling Inside and Outside the Cell. Journal of Developmental Biology, 4(3), 23. https://doi.org/10.3390/jdb4030023