Integrated Transcriptomic, Proteomic, and Metabolomics Analysis Reveals Peel Ripening of Harvested Banana under Natural Condition
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Treatment
2.2. Determination of Physiological Parameters during Fruit Ripening
2.3. RNA-Seq (Quantification) Analysis
2.4. Quantitative Real-Time Polymerase Chain Reaction Analysis
2.5. Two-Dimensional Gel Electrophoresis (2-DE) and Protein Identification
2.6. Metabolic Profiling
2.7. Bioinformatics Analysis
2.8. Statistical Analysis
3. Results
3.1. Ripening and Physiological Characteristics of Harvested Banana Fruit
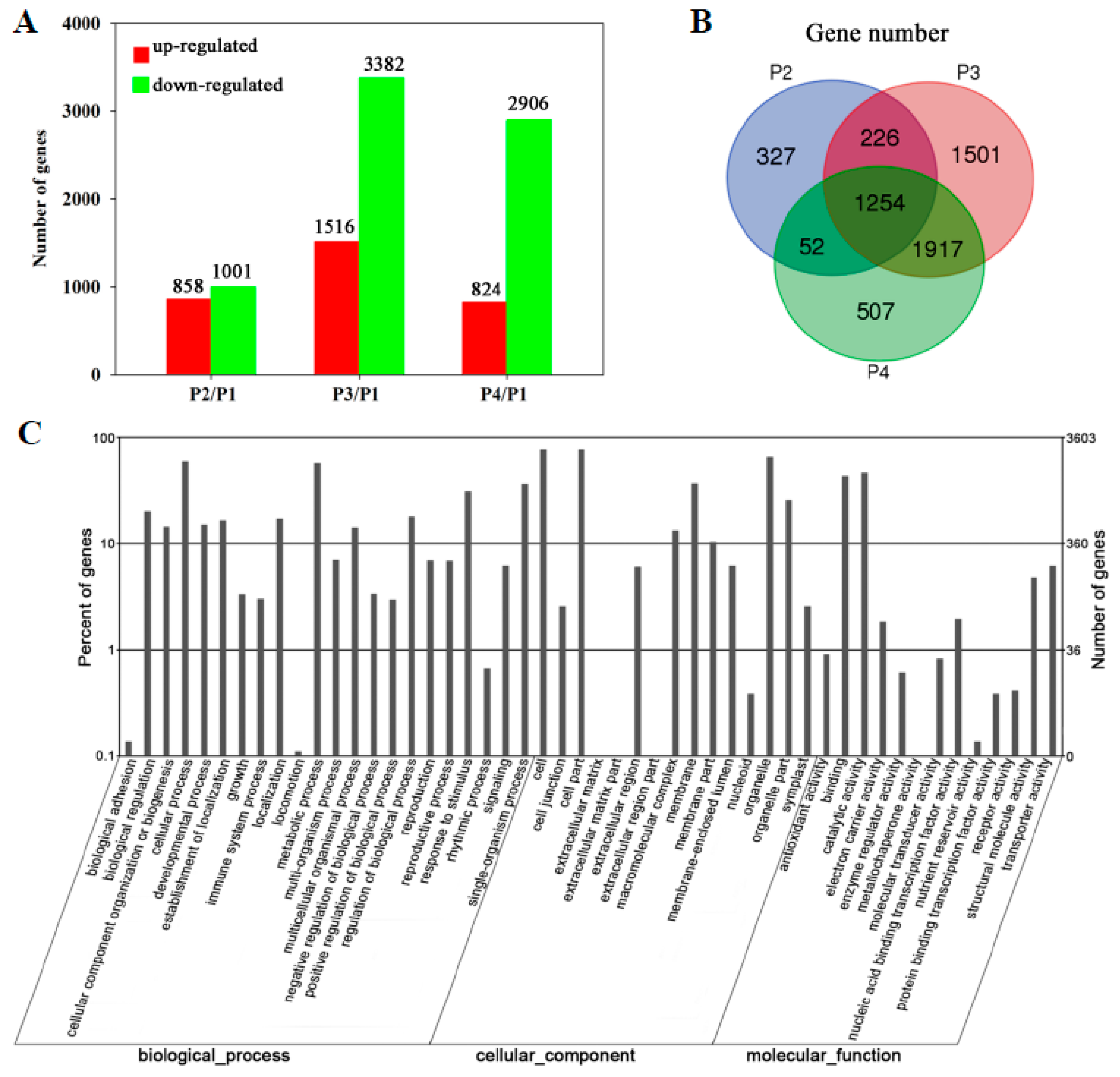
3.2. Transcriptomic Analysis of Banana Fruit during Ripening
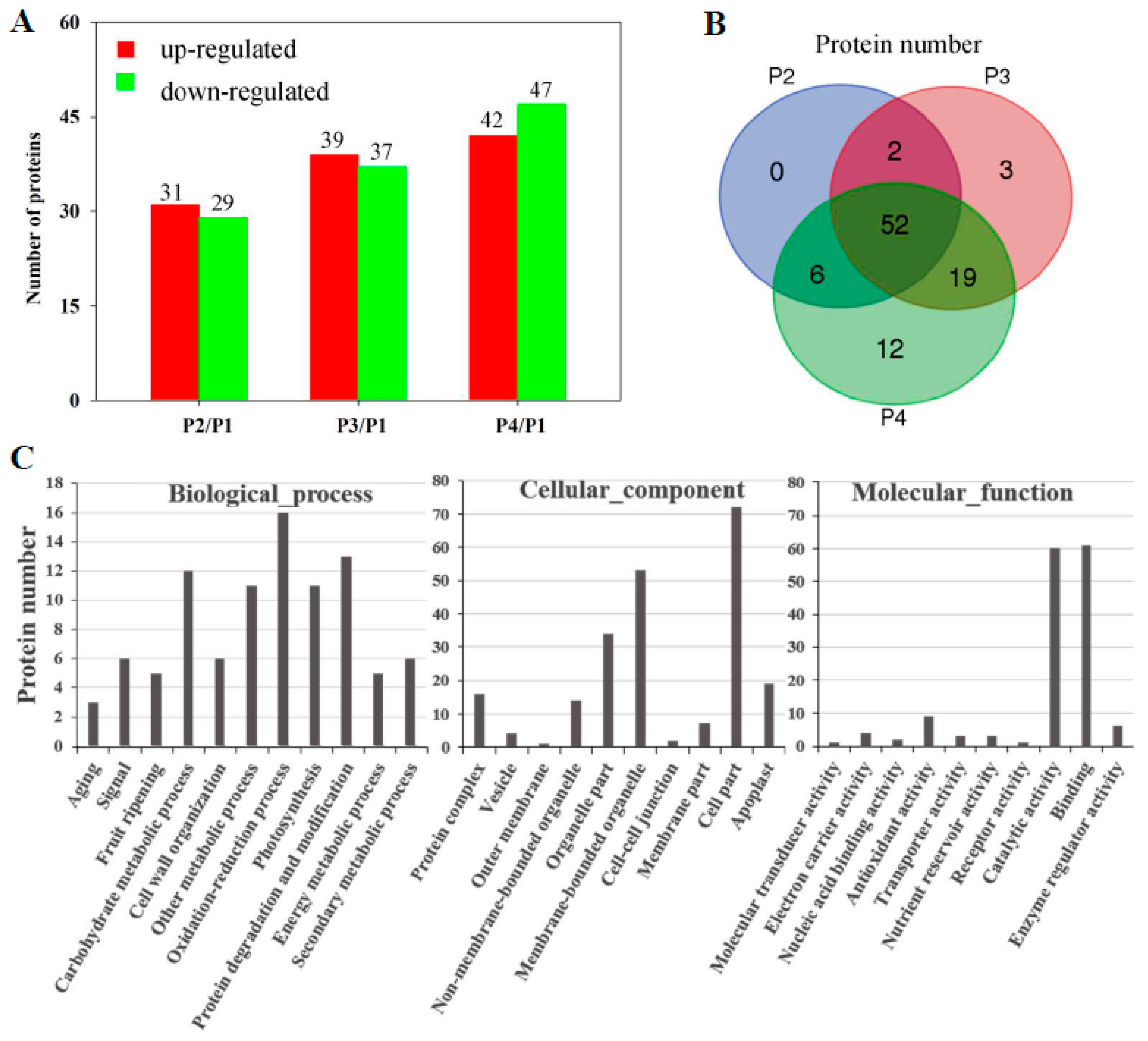
3.3. Proteomic Analysis of Banana Fruit during Ripening
3.4. Metabolomics Profiling of Banana Fruit during Ripening
3.5. Correlation Analysis of Transcriptomic and Proteomic Data
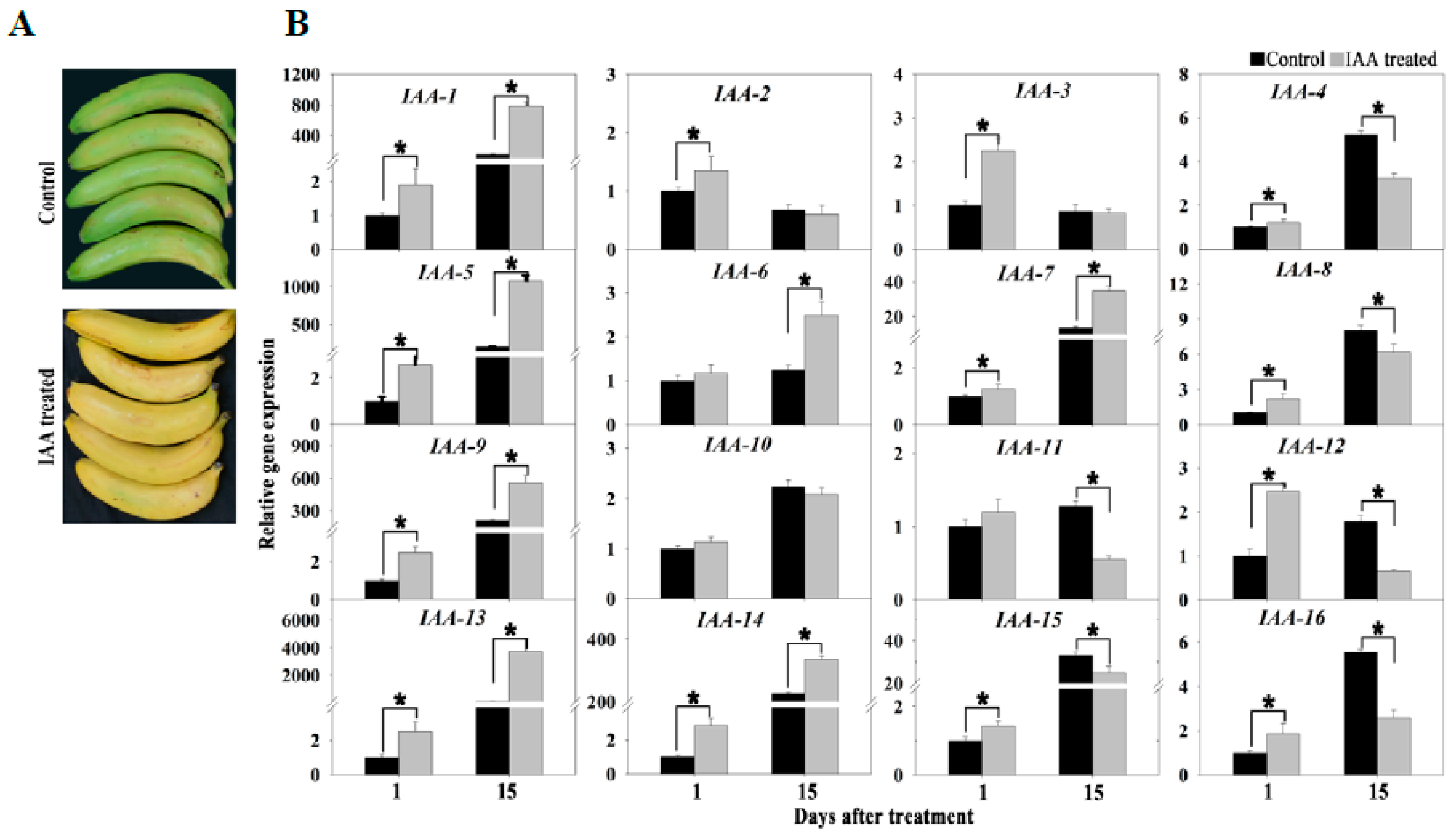
3.6. Specifically Expressed Signal Transduction-Related Genes and Proteins during Banana Ripening
3.7. Specifically Expressed Transcription Factor Genes and Proteins during Banana Ripening
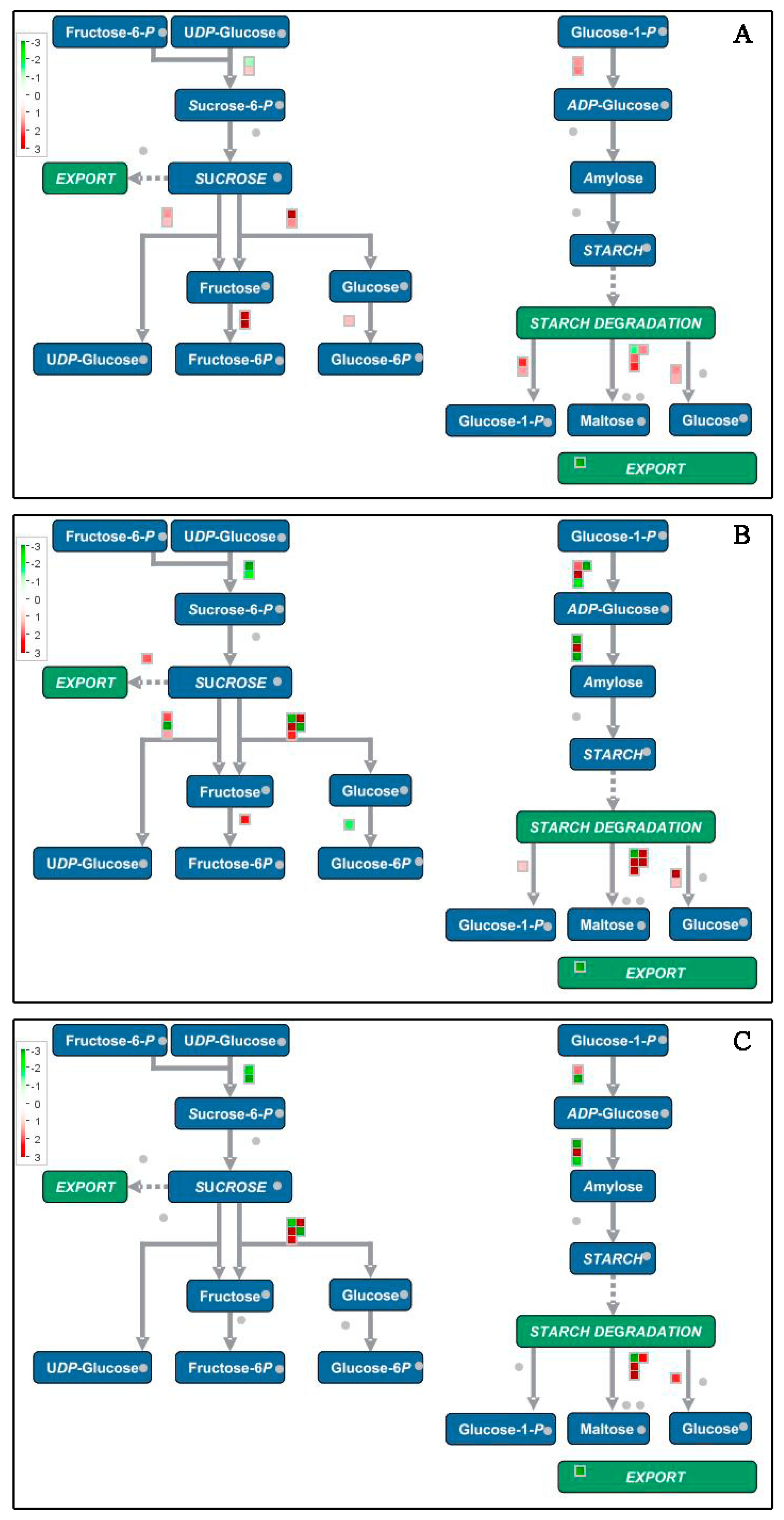
3.8. Specifically Expressed Metabolic Process-Related Genes and Proteins during Banana Ripening
3.9. Specific Genes and Proteins Related to Protein Modification and Degradation during Banana Ripening
4. Discussion
4.1. Hormone Signaling
4.2. Transcription Factors
4.3. Cell Wall Degradation
4.4. Synthesis of Volatile Compounds
4.5. Protein Modification
4.6. Energy Metabolism
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Clendennen, S.K.; May, G.D. Differential gene expression in ripening banana fruit. Plant Physiol. 1997, 115, 463–469. [Google Scholar] [CrossRef]
- Giovannoni, J. Molecular biology of fruit maturation and ripening. Annu. Rev. Plant Phys. Plant Mol. Biol. 2001, 52, 725–749. [Google Scholar] [CrossRef]
- Cherian, S.; Figueroa, C.R.; Nair, H. ‘Movers and shakers’ in the regulation of fruit ripening: A cross-dissection of climacteric versus non-climacteric fruit. J. Exp. Bot. 2014, 65, 4705–4722. [Google Scholar] [CrossRef]
- Elitzur, T.; Vrebalov, J.; Giovannoni, J.J.; Goldschmidt, E.E.; Friedman, H. The regulation of MADS-box gene expression during ripening of banana and their regulatory interaction with ethylene. J. Exp. Bot. 2010, 61, 1523–1535. [Google Scholar] [CrossRef] [PubMed]
- Shan, W.; Kuang, J.-F.; Chen, L.; Xie, H.; Peng, H.-H.; Xiao, Y.-Y.; Li, X.-P.; Chen, W.-X.; He, Q.-G.; Chen, J.-Y.; et al. Molecular characterization of banana NAC transcription factors and their interactions with ethylene signalling component EIL during fruit ripening. J. Exp. Bot. 2012, 63, 5171–5187. [Google Scholar] [CrossRef] [PubMed]
- Nham, N.T.; de Freitas, S.T.; Macnish, A.J.; Carr, K.M.; Kietikul, T.; Guilatco, A.J.; Jiang, C.-Z.; Zakharov, F.; Mitcham, E.J. A transcriptome approach towards understanding the development of ripening capacity in ‘Bartlett’ pears (Pyrus communis L.). BMC Genomics 2015, 16. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Q.; Song, J.; Campbell-Palmer, L.; Thompson, K.; Li, L.; Walker, B.; Cui, Y.; Li, X. A proteomic investigation of apple fruit during ripening and in response to ethylene treatment. J. Proteomics 2013, 93, 276–294. [Google Scholar] [CrossRef]
- Pan, L.; Zeng, W.; Niu, L.; Lu, Z.; Liu, H.; Cui, G.; Zhu, Y.; Chu, J.; Li, W.; Fang, W.; et al. PpYUC11, a strong candidate gene for the stony hard phenotype in peach (Prunus persica L. Batsch), participates in IAA biosynthesis during fruit ripening. J. Exp. Bot. 2015, 66, 7031–7044. [Google Scholar] [CrossRef]
- D’Hont, A.; Denoeud, F.; Aury, J.-M.; Baurens, F.-C.; Carreel, F.; Garsmeur, O.; Noel, B.; Bocs, S.; Droc, G.; Rouard, M.; et al. The banana (Musa acuminata) genome and the evolution of monocotyledonous plants. Nature 2012, 488, 213. [Google Scholar] [CrossRef]
- Asif, M.H.; Lakhwani, D.; Pathak, S.; Gupta, P.; Bag, S.K.; Nath, P.; Trivedi, P.K. Transcriptome analysis of ripe and unripe fruit tissue of banana identifies major metabolic networks involved in fruit ripening process. BMC Plant Biol. 2014, 14. [Google Scholar] [CrossRef]
- Toledo, T.T.; Nogueira, S.B.; Cordenunsi, B.R.; Gozzo, F.C.; Pilau, E.J.; Lajolo, F.M.; Oliveira do Nascimento, J.R. Proteomic analysis of banana fruit reveals proteins that are differentially accumulated during ripening. Postharvest Biol. Tec. 2012, 70, 51–58. [Google Scholar] [CrossRef]
- Inaba, A.; Liu, X.; Yokotani, N.; Yamane, M.; Lu, W.-J.; Nakano, R.; Kubo, Y. Differential feedback regulation of ethylene biosynthesis in pulp and peel tissues of banana fruit. J. Exp. Bot. 2007, 58, 1047–1057. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Qian, Z.; Ma, S.; Zhou, Y.; Patrick, J.W.; Duan, X.; Jiang, Y.; Qu, H. Energy status of ripening and postharvest senescent fruit of litchi (Litchi chinensis Sonn.). BMC Plant Biol. 2013, 13. [Google Scholar] [CrossRef]
- Gao, H.; Huang, S.; Dong, T.; Yang, Q.; Yi, G. Analysis of resistant starch degradation in postharvest ripening of two banana cultivars: Focus on starch structure and amylases. Postharvest Biol. Technol. 2016, 119, 1–8. [Google Scholar] [CrossRef]
- Zhao, Y.; Lin, H.; Wang, J.; Lin, Y.; Chen, Y. Inhibiting aril breakdown and degradation of cell wall material in pulp of harvested longan fruits by heat treatment. Trans. Chin. Soc. Agric. Eng. 2014, 30, 268–275. [Google Scholar]
- Li, R.; Yu, C.; Li, Y.; Lam, T.-W.; Yiu, S.-M.; Kristiansen, K.; Wang, J. SOAP2: An improved ultrafast tool for short read alignment. Bioinformatics 2009, 25, 1966–1967. [Google Scholar] [CrossRef]
- Mortazavi, A.; Williams, B.A.; McCue, K.; Schaeffer, L.; Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 2008, 5, 621–628. [Google Scholar] [CrossRef]
- Yun, Z.; Jin, S.; Ding, Y.; Wang, Z.; Gao, H.; Pan, Z.; Xu, J.; Cheng, Y.; Deng, X. Comparative transcriptomics and proteomics analysis of citrus fruit, to improve understanding of the effect of low temperature on maintaining fruit quality during lengthy post-harvest storage. J. Exp. Bot. 2012, 63, 2873–2893. [Google Scholar] [CrossRef]
- Yun, Z.; Li, W.; Pan, Z.; Xu, J.; Cheng, Y.; Deng, X. Comparative proteomics analysis of differentially accumulated proteins in juice sacs of ponkan (Citrus reticulata) fruit during postharvest cold storage. Postharvest Biol. Technol. 2010, 56, 189–201. [Google Scholar] [CrossRef]
- Yamaguchi, M.; Valliyodan, B.; Zhang, J.; Lenoble, M.E.; Yu, O.; Rogers, E.E.; Nguyen, H.T.; Sharp, R.E. Regulation of growth response to water stress in the soybean primary root. I. Proteomic analysis reveals region-specific regulation of phenylpropanoid metabolism and control of free iron in the elongation zone. Plant Cell Environ. 2010, 33, 223–243. [Google Scholar] [CrossRef]
- Yun, Z.; Gao, H.; Liu, P.; Liu, S.; Luo, T.; Jin, S.; Xu, Q.; Xu, J.; Cheng, Y.; Deng, X. Comparative proteomic and metabolomic profiling of citrus fruit with enhancement of disease resistance by postharvest heat treatment. BMC Plant Biol. 2013, 13. [Google Scholar] [CrossRef] [PubMed]
- Jing, G.; Li, T.; Qu, H.; Yun, Z.; Jia, Y.; Zheng, X.; Jiang, Y. Carotenoids and volatile profiles of yellow- and red-fleshed papaya fruit in relation to the expression of carotenoid cleavage dioxygenase genes. Postharvest Biol. Technol. 2015, 109, 114–119. [Google Scholar] [CrossRef]
- Prasanna, V.; Prabha, T.N.; Tharanathan, R.N. Fruit ripening phenomena - An overview. Crit. Rev. Food Sci. Nutr. 2007, 47, 1–19. [Google Scholar] [CrossRef]
- Jiang, Y.M.; Joyce, D.C.; Macnish, A.J. Responses of banana fruit to treatment with 1-methylcyclopropene. Plant Growth Regul. 1999, 28, 77–82. [Google Scholar] [CrossRef]
- Adams-Phillips, L.; Barry, C.; Giovannoni, J. Signal transduction systems regulating fruit ripening. Trends Plant Sci. 2004, 9, 331–338. [Google Scholar] [CrossRef] [PubMed]
- Seymour, G.B.; Ostergaard, L.; Chapman, N.H.; Knapp, S.; Martin, C. Fruit development and ripening. Annu. Rev. Plant Biol. 2013, 64, 219–241. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; Zhong, S.; Grierson, D. Recent advances in ethylene research. J. Exp. Bot. 2009, 60, 3311–3336. [Google Scholar] [CrossRef]
- An, F.; Zhao, Q.; Ji, Y.; Li, W.; Jiang, Z.; Yu, X.; Zhang, C.; Han, Y.; He, W.; Liu, Y.; et al. Ethylene-induced stabilization of ETHYLENE INSENSITIVE3 and EIN3-LIKE1 is mediated by proteasomal degradation of EIN3 binding F-Box 1 and 2 that requires EIN2 in Arabidopsis. Plant Cell 2010, 22, 2384–2401. [Google Scholar] [CrossRef] [PubMed]
- Hagen, G. Auxin signal transduction. Essays Biochem. 2015, 58, 1–12. [Google Scholar] [CrossRef]
- El-Sharkawy, I.; Sherif, S.M.; Jones, B.; Mila, I.; Kumar, P.P.; Bouzayen, M.; Jayasankar, S. TIR1-like auxin-receptors are involved in the regulation of plum fruit development. J. Exp. Bot. 2014, 65, 5205–5215. [Google Scholar] [CrossRef]
- Busatto, N.; Tadiello, A.; Trainotti, L.; Costa, F. Climacteric ripening of apple fruit is regulated by transcriptional circuits stimulated by cross-talks between ethylene and auxin. Plant Signal. Behav. 2017, 12. [Google Scholar] [CrossRef]
- Lakhwani, D.; Pandey, A.; Dhar, Y.V.; Bag, S.K.; Trivedi, P.K.; Asif, M.H. Genome-wide analysis of the AP2/ERF family in Musa species reveals divergence and neofunctionalisation during evolution. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef]
- Li, T.; Jiang, Z.; Zhang, L.; Tan, D.; Wei, Y.; Yuan, H.; Li, T.; Wang, A. Apple (Malus domestica) MdERF2 negatively affects ethylene biosynthesis during fruit ripening by suppressing MdACS1 transcription. Plant J. 2016, 88, 735–748. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.-C.; Kuang, J.-F.; Chen, J.-Y.; Liu, X.-C.; Xiao, Y.-Y.; Fu, C.-C.; Wang, J.-N.; Wu, K.-Q.; Lu, W.-J. Banana transcription factor MaERF11 recruits histone deacetylase MaHDA1 and represses the expression of MaACO1 and expansins during fruit ripening. Plant Physiol. 2016, 171, 1070–1084. [Google Scholar] [CrossRef] [PubMed]
- Kuang, J.-F.; Chen, J.-Y.; Liu, X.-C.; Han, Y.-C.; Xiao, Y.-Y.; Shan, W.; Tang, Y.; Wu, K.-Q.; He, J.-X.; Lu, W.-J. The transcriptional regulatory network mediated by banana (Musa acuminata) dehydration-responsive element binding (MaDREB) transcription factors in fruit ripening. New Phytol. 2017, 214, 762–781. [Google Scholar] [CrossRef] [PubMed]
- Endo, T.; Fujii, H.; Sugiyama, A.; Nakano, M.; Nakajima, N.; Ikoma, Y.; Omura, M.; Shimada, T. Overexpression of a citrus basic helix-loop-helix transcription factor (CubHLH1), which is homologous to Arabidopsis activation-tagged bri1 suppressor 1 interacting factor genes, modulates carotenoid metabolism in transgenic tomato. Plant Sci. 2016, 243, 35–48. [Google Scholar] [CrossRef]
- Wei, H.; Chen, X.; Zong, X.; Shu, H.; Gao, D.; Liu, Q. Comparative transcriptome analysis of genes involved in anthocyanin biosynthesis in the red and yellow fruits of sweet cherry (Prunus avium L.). PLoS ONE 2015, 10. [Google Scholar] [CrossRef]
- Peng, H.-H.; Shan, W.; Kuang, J.-F.; Lu, W.-J.; Chen, J.-Y. Molecular characterization of cold-responsive basic helix-loop-helix transcription factors MabHLHs that interact with MaICE1 in banana fruit. Planta 2013, 238, 937–953. [Google Scholar] [CrossRef]
- Wang, D.; Zhang, H.; Wu, F.; Li, T.; Liang, Y.; Duan, X. Modification of pectin and hemicellulose polysaccharides in relation to aril breakdown of harvested longan fruit. Int. J. Mol. Sci. 2013, 14, 23356–23368. [Google Scholar] [CrossRef]
- Bennett, A.B.; Labavitch, J.M. Ethylene and ripening-regulated expression and function of fruit cell wall modifying proteins. Plant Sci. 2008, 175, 130–136. [Google Scholar] [CrossRef]
- Eriksson, E.M.; Bovy, A.; Manning, K.; Harrison, L.; Andrews, J.; De Silva, J.; Tucker, G.A.; Seymour, G.B. Effect of the Colorless non-ripening mutation on cell wall biochemistry and gene expression during tomato fruit development and ripening. Plant Physiol. 2004, 136, 4184–4197. [Google Scholar] [CrossRef] [PubMed]
- Marin-Rodriguez, M.C.; Orchard, J.; Seymour, G.B. Pectate lyases, cell wall degradation and fruit softening. J. Exp. Bot. 2002, 53, 2115–2119. [Google Scholar] [CrossRef] [PubMed]
- Yokoyama, R.; Uwagaki, Y.; Sasaki, H.; Harada, T.; Hiwatashi, Y.; Hasebe, M.; Nishitani, K. Biological implications of the occurrence of 32 members of the XTH (xyloglucan endotransglucosylase/hydrolase) family of proteins in the bryophyte Physcomitrella patens. Plant J. 2010, 64, 645–656. [Google Scholar] [CrossRef] [PubMed]
- Miedes, E.; Suslov, D.; Vandenbussche, F.; Kenobi, K.; Ivakov, A.; Van Der Straeten, D.; Lorences, E.P.; Mellerowicz, E.J.; Verbelen, J.-P.; Vissenberg, K. Xyloglucan endotransglucosylase/hydrolase (XTH) overexpression affects growth and cell wall mechanics in etiolated Arabidopsis hypocotyls. J. Exp. Bot. 2013, 64, 2481–2497. [Google Scholar] [CrossRef]
- Peled-Zehavi, H.; Oliva, M.; Xie, Q.; Tzin, V.; Oren-Shamir, M.; Aharoni, A.; Galili, G. Metabolic engineering of the phenylpropanoid and its primary, precursor pathway to enhance the flavor of fruits and the aroma of flowers. Bioengineering 2015, 2, 204–212. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Man, C.; Li, D.; Tan, H.; Xie, Y.; Huang, J. Arogenate dehydratase isoforms differentially regulate anthocyanin biosynthesis in Arabidopsis thaliana. Mol. Plant 2016, 9, 1609–1619. [Google Scholar] [CrossRef] [PubMed]
- Kalua, C.M.; Boss, P.K. Evolution of volatile compounds during the development of cabernet sauvignon grapes (Vitis vinifera L.). J. Agric. Food Chem. 2009, 57, 3818–3830. [Google Scholar] [CrossRef]
- D’Ambrosio, C.; Arena, S.; Rocco, M.; Verrillo, F.; Novi, G.; Viscosi, V.; Marra, M.; Scaloni, A. Proteomic analysis of apricot fruit during ripening. J. Proteomics 2013, 78, 39–57. [Google Scholar] [CrossRef]
- Nieuwenhuizen, N.J.; Chen, X.; Wang, M.Y.; Matich, A.J.; Perez, R.L.; Allan, A.C.; Green, S.A.; Atkinson, R.G. Natural variation in monoterpene synthesis in kiwifruit: transcriptional regulation of terpene synthases by NAC and ETHYLENE-INSENSITIVE3-like transcription factors. Plant Physiol. 2015, 167, 1243–1258. [Google Scholar] [CrossRef] [PubMed]
- Kaur, G.; Lieu, K.G.; Jans, D.A. 70-kDa heat shock cognate protein hsc70 mediates calmodulin-dependent nuclear import of the sex-determining factor SRY. J. Biol. Chem. 2013, 288, 4148–4157. [Google Scholar] [CrossRef]
- Aghdam, M.S.; Bodbodak, S. Physiological and biochemical mechanisms regulating chilling tolerance in fruits and vegetables under postharvest salicylates and jasmonates treatments. Sci. Hortic. 2013, 156, 73–85. [Google Scholar] [CrossRef]
- Zou, J.; Liu, C.; Liu, A.; Zou, D.; Chen, X. Overexpression of OsHsp17.0 and OsHsp23.7 enhances drought and salt tolerance in rice. J. Plant Physiol. 2012, 169, 628–635. [Google Scholar] [CrossRef]
- Ramadan, A.; Nemoto, K.; Seki, M.; Shinozaki, K.; Takeda, H.; Takahashi, H.; Sawasaki, T. Wheat germ-based protein libraries for the functional characterisation of the Arabidopsis E2 ubiquitin conjugating enzymes and the RING-type E3 ubiquitin ligase enzymes. BMC Plant Biol. 2015, 15. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, W.; Cai, J.; Zhang, Y.; Qin, G.; Tian, S. Tomato nuclear proteome reveals the involvement of specific E2 ubiquitin-conjugating enzymes in fruit ripening. Genome Biol. 2014, 15. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.-H.; Zhang, J.; Jia, C.-H.; Zhang, J.-B.; Wang, J.-S.; Yang, Z.-X.; Xu, B.-Y.; Jin, Z.-Q. The interaction of banana MADS-box protein MuMADS1 and ubiquitin-activating enzyme E-MuUBA in post-harvest banana fruit. Plant Cell Rep. 2013, 32, 129–137. [Google Scholar] [CrossRef] [PubMed]
- El-Sharkawy, I.; Sherif, S.; El Kayal, W.; Jones, B.; Li, Z.; Sullivan, A.J.; Jayasankar, S. Overexpression of plum auxin receptor PslTIR1 in tomato alters plant growth, fruit development and fruit shelf-life characteristics. BMC Plant Biol. 2016, 16. [Google Scholar] [CrossRef] [PubMed]
- Collet, J.-F.; Messens, J. Structure, function, and mechanism of thioredoxin proteins. Antioxid. Redox Signal. 2010, 13, 1205–1216. [Google Scholar] [CrossRef]
- Borsani, J.; Budde, C.O.; Porrini, L.; Lauxmann, M.A.; Lombardo, V.A.; Murray, R.; Andreo, C.S.; Drincovich, M.F.; Lara, M.V. Carbon metabolism of peach fruit after harvest: Changes in enzymes involved in organic acid and sugar level modifications. J. Exp. Bot. 2009, 60, 1823–1837. [Google Scholar] [CrossRef]





© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yun, Z.; Li, T.; Gao, H.; Zhu, H.; Gupta, V.K.; Jiang, Y.; Duan, X. Integrated Transcriptomic, Proteomic, and Metabolomics Analysis Reveals Peel Ripening of Harvested Banana under Natural Condition. Biomolecules 2019, 9, 167. https://doi.org/10.3390/biom9050167
Yun Z, Li T, Gao H, Zhu H, Gupta VK, Jiang Y, Duan X. Integrated Transcriptomic, Proteomic, and Metabolomics Analysis Reveals Peel Ripening of Harvested Banana under Natural Condition. Biomolecules. 2019; 9(5):167. https://doi.org/10.3390/biom9050167
Chicago/Turabian StyleYun, Ze, Taotao Li, Huijun Gao, Hong Zhu, Vijai Kumar Gupta, Yueming Jiang, and Xuewu Duan. 2019. "Integrated Transcriptomic, Proteomic, and Metabolomics Analysis Reveals Peel Ripening of Harvested Banana under Natural Condition" Biomolecules 9, no. 5: 167. https://doi.org/10.3390/biom9050167
APA StyleYun, Z., Li, T., Gao, H., Zhu, H., Gupta, V. K., Jiang, Y., & Duan, X. (2019). Integrated Transcriptomic, Proteomic, and Metabolomics Analysis Reveals Peel Ripening of Harvested Banana under Natural Condition. Biomolecules, 9(5), 167. https://doi.org/10.3390/biom9050167







