Abstract
Fetal Alcohol Spectrum Disorder (FASD) is a common neurodevelopmental disorder that affects an estimated 2–5% of North Americans. FASD is induced by prenatal alcohol exposure (PAE) during pregnancy and while there is a clear genetic contribution, few genetic factors are currently identified or understood. In this study, using a candidate gene approach, we performed a genetic variant analysis of retinoic acid (RA) metabolic and developmental signaling pathway genes on whole exome sequencing data of 23 FASD-diagnosed individuals. We found risk and resilience alleles in ADH and ALDH genes known to normally be involved in alcohol detoxification at the expense of RA production, causing RA deficiency, following PAE. Risk and resilience variants were also identified in RA-regulated developmental pathway genes, especially in SHH and WNT pathways. Notably, we also identified significant variants in the causative genes of rare neurodevelopmental disorders sharing comorbidities with FASD, including STRA6 (Matthew–Wood), SOX9 (Campomelic Dysplasia), FDG1 (Aarskog), and 22q11.2 deletion syndrome (TBX1). Although this is a small exploratory study, the findings support PAE-induced RA deficiency as a major etiology underlying FASD and suggest risk and resilience variants may be suitable biomarkers to determine the risk of FASD outcomes following PAE.
1. Introduction
Fetal Alcohol Spectrum Disorder (FASD) is a common neurodevelopmental disorder that affects an estimated 2–5% of the population [1]. Individuals across the spectrum show behavioural, adaptive, and cognitive abnormalities, which can be as serious in those without any physical features as in those with the most severe form of FASD, Fetal Alcohol Syndrome (FAS) [2]. Diagnosing individuals with FASD remains difficult as most do not present with FASD sentinel facial features [3], and it requires assessments involving many medical disciplines. In the absence of FASD physical hallmarks, confirmation of PAE and a psychological profile that meets diagnostic guideline criteria are necessary for diagnosis [2]. As such, additional diagnostic tools are needed to identify FASD for early intervention and better outcomes. Due to the complex nature of the disorder, FASD has many contributing risk factors including alcohol dosage [4,5], duration of exposure and gestational timing [6,7,8,9], maternal nutrition, metabolism, socioeconomic factors, stress, and genetic variant composition [10,11,12,13,14].
Genetic composition is a well-established risk factor in FASD. PAE animal models have shown that different strains of the same animal species have different susceptibilities to PAE [15]. Human twin studies on individuals with FASD have shown that monozygotic twins share 100% concordance for their FASD diagnoses, while fraternal twins have around 56% and non-twin siblings only have 46% [16]. Importantly, the majority of individuals with PAE do not develop FASD [17], suggesting that there is a genetic vulnerability that sensitizes an individual to develop FASD in the presence of PAE. One can hypothesize that allelic variants found in such genes can act as risk or resilience factors that could either drive or protect against FASD outcomes.
Research in PAE animal models has revealed many potential gene targets for investigation within PAE-impacted pathways, such as Shh and Wnt, that could host FASD risk variants [15]. Retinoic acid (RA) is a developmental signaling pathway shown to regulate many PAE-impacted pathways [18]. Moreover, Vitamin A (retinol) supplementation rescues PAE phenotypes in animal studies, while RA-deficiency phenocopies many of the craniofacial and neurological malformations found in PAE animal models and individuals with FASD [18,19,20,21]. Enzymes involved in RA biosynthesis have been shown to participate in ethanol detoxification at the expense of RA production. These findings indicate that variants in RA metabolic genes are likely candidates in determining whether an individual prenatally exposed to alcohol will develop FASD [18,21].
Many rare neurodevelopmental disorders (NDDs), such as CHARGE and Smith–Lemli–Opitz syndrome (SLOS), phenocopy FASD in terms of their specific craniofacial, neurodevelopmental, or behavioural manifestations [20]. Interestingly, many of the causative genes of these rare NDDs are regulated by RA signaling and crosstalk with well-established PAE targets such as SHH and WNT. Therefore, it is likely that allelic variants within the causative genes of these NDDs may also sensitize individuals to develop FASD following PAE, given their shared co-morbidities and signaling pathways. Haploinsufficiency in causative genes may be enough to disrupt the signaling pathway [15] and produce a similar phenotype seen in both NDDs and FASD.
Taken together, RA serves as a master regulator of this signaling network, which encompasses well-known PAE-implicated pathways and the causative genes of rare NDDs directing embryonic development [18,20]. Variants found within the RA metabolic and signaling network may sensitize or protect individuals against developing FASD following PAE. Furthermore, such variants may serve as genetic biomarkers of risk or resilience of FASD outcomes following PAE. The goal of this exploratory study was to investigate the variants found in the genes comprising the RA signaling network in 23 individuals with FASD and to discover novel FASD risk and resilience alleles to further our understanding of this disorder. To examine the variants found in our FASD cohort, the RA signaling network was divided into the following three candidate gene lists: (1) RA and alcohol (ethanol) metabolism genes, (2) RA-regulated developmental pathway genes (such as SHH and WNT), and (3) the causative genes of rare NDDs and their direct signaling targets.
2. Methods
2.1. Participants
The 23 children in this study are part of the Canadian NeuroDevNet/Kids Brain Health Network FASD Study Cohort [22,23]. Participants were diagnosed at the Manitoba FASD Centre where the assessment and diagnosis of FASD are based on an experienced multidisciplinary team approach including medical evaluation of developmental pediatricians (MD) and medical geneticists (MD) with dysmorphology training using the revised Canadian FASD Diagnostic Guidelines [2]. These guidelines incorporate the use of the Washington four-digit code. Differential diagnosis carefully considers other genetic, neurodevelopmental, medical, and environmental contributors to an individual presentation. The dedicated diagnostic team also includes qualified and certified speech and language pathologists, Clinical Psychologists (PhD), social workers (MSW), and clinic coordinators. Relevant birth records were obtained, and a comprehensive review of prenatal alcohol and substance exposure history, medical record review, and complete social history was conducted as part of the assessment for FASD-related diagnoses. Eleven children were male and 12 were female. In our FASD cohort, there were 3 pairs of siblings, each pair had matched diagnoses, 2 pairs were both diagnosed with partial FAS (pFAS), and the other pair with ARND. Ten children were of European ancestry, and the remaining 13 were of Indigenous ancestry, either First Nations (n = 10), Inuit (n = 1), or Metis (n = 2). Six individuals were diagnosed with FAS, 12 with pFAS, and 5 with alcohol-related neurodevelopmental disorder (ARND). Research ethics board approvals were obtained at collection centres and either written or verbal assent, as well as written consent, was given by all participants and their respective caregivers or legal guardians, respectively. Saliva samples were then obtained from the children; the details of sample collection and storage can be found elsewhere [23]. Approval of this study was granted by the Health Research Ethics Board at the University of Manitoba, (REB Study Number: H2010:206).
2.2. Whole Exome Sequencing
DNA was extracted from the 23 participants’ saliva samples and sent for whole exome sequencing at the Center of Applied Genomics located at The Hospital for Sick Children in Toronto, Canada. DNA samples were first ligated with SOLiD A1 and P1 adaptors to undergo PCR amplification (Thermo Fisher Scientific, Waltham, MA, USA). Exome capture was then completed according to the Agilent SureSelect Protocol (Agilent Technologies, Inc., Santa Clara, CA, USA). Upon completion, the samples underwent a PCR amplification to apply barcodes for multiplex sequencing. The captured library was then purified using AMPure XP beads following the Agilent SureSelect Protocol. Paired-end sequencing was completed with ABI SOLiD 5500xl (Thermo Fisher Scientific, Waltham, MA, USA) on the purified library and then analyzed using the Applied Biosystems corona pipeline to generate sequencing data. These paired end-reads generated by the ABI SOLiD were mapped to the human reference genome (hg19) using BFAST [24] for 75 bp forward reads and BFAST implemented BWA version 0.6.5a [25] for 35 bp reverse reads. MarkDuplicates was used to remove any duplicate paired-end reads that were found [26]. The duplicate-free alignments were then refined using local realignment with SRMA version 0.1.15 [27]. GATK version 1.1.28 was then used for variant calling [28] (for QC see Section 2.4 below). The resulting variant analysis had a sequencing depth of 30×.
2.3. Candidate Gene Lists
All candidate gene lists found in this paper were manually curated based on a review of literature sources, excluding papers studying cancer. Literature sources for the RA and alcohol metabolism candidate gene list were compiled based on the genes that encompass the RA and alcohol metabolism pathways, as well as a selection of variants previously associated with alcohol consumption, dependence, and clearance (Supplementary Table S1). To assemble the list of candidate developmental genes found in RA-regulated PAE-impacted pathways, the literature was searched for known RA-controlled genes active during development affected by PAE, as well as their direct interaction partners (Supplementary Materials, Table S2). Literature sources examining the causative genes of rare NDDs with shared specific craniofacial, neurological, as well as behavioural co-morbidities with FASD and their directly interacting genes were used to compile the third candidate gene list (Supplementary Table S3), as discussed in our recent review [20]. A fourth candidate gene list was also compiled that includes synthesizing enzymes and receptors for neurotransmitters, forebrain development genes, and collagen genes (Supplementary Table S4; the results are not discussed in this report but can be found in Supplementary Table S8).
2.4. Variant Analysis of Whole Exome Sequencing Data Using Candidate Gene Approaches
Aligned variants found in the whole exome sequencing data of the FASD cohort were imported into R, and all variants found in the genes from our gene lists of interest were compiled; the frequencies of variant alleles in our FASD cohort were then determined. The corresponding 1000 Genomes Project global allelic frequencies were used as controls (NCBI) (Figure 1). Variants further analyzed included those found in exons, introns, 3′ UTR, 5′ UTR, and upstream regions. As the smallest detected frequency in our cohort was 1/46 (2.17%), a minor allele frequency (MAF) threshold filter of ≥2.17% (QC filtering parameters) was applied to all alleles in the 1000 Genomes Project dataset for variants presented in the results tables. The allele frequencies for the FASD cohort and the 1000 Genomes Project control dataset were then compared to each other using a chi-squared test. A Benjamini–Hochberg (BH) multiple test correction with a 0.05 significance threshold was then applied to the chi-square test p-values (number of tests (n) = 1100) with a resulting p-value threshold of 0.024. To account for variants with known alcohol associations, such as alcohol consumption and dependence that may have low frequencies, as well as any known rare pathogenic variants that may be found within developmental genes in our FASD cohort, known variants with frequencies below 2.17% in the 1000 Genomes Project dataset were included in the total number of tests. All statistically significant genetic variants were then sequentially examined in the literature and annotated with PolyPhen2 and SIFT scores [29,30] based on their impact on gene function and association to clinically relevant phenotypes to determine risk and resilience variants (Figure 1). Discovered risk and resilience variants were then annotated with CADD scores [31]. For predictive tools, we considered deleterious SIFT and possibly damaging or damaging PolyPhen2 predictions, as well as CADD scores > 10, to be potentially damaging to gene function [29,30,31]. It should be noted that the majority of these presumptive risk and resilience alleles require further experimentation to confirm their true impact on the severity of PAE teratogenesis.
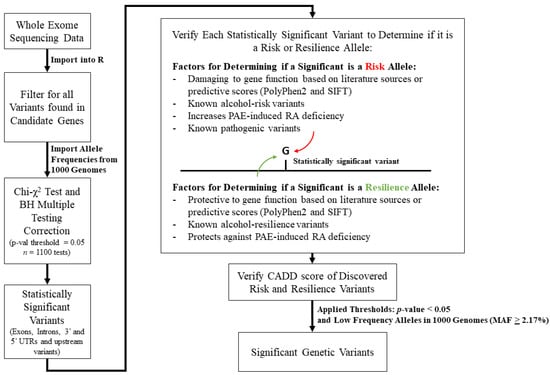
Figure 1.
Flow chart of bioinformatic pipeline and determination factors considered for assigning risk and resilience alleles of genetic variants in the whole exome sequencing data of the FASD cohort. Once variants were imported into R, only variants found in our candidate genes were taken for analysis. This was followed by a chi-squared test using the 1000 Genomes Project allele frequencies as controls, which was followed by Benjamini–Hochberg multiple testing correction. Each variant was then sequentially determined to be an FASD risk or resilience allele based on defined criteria (literature sources and predictive scores) as seen in the Supplementary Tables. The CADD scores of discovered risk and resilience alleles were then investigated. Lastly, thresholds were applied at p-values < 0.05 and for low-frequency alleles in 1000 Genomes, and only significant genetic variants passing thresholds are shown in the results tables.
2.5. TaqMan SNP Genotyping Assays
Following variant analysis of the FASD whole exome sequencing data, 16 genetic variants of interest were selected for validation by TaqMan probes ordered from Applied Biosystems (Thermo Fisher Scientific, Waltham, MA, USA). As the RA and alcohol metabolism pathways are the primary focus of this paper, variants in these pathways were prioritized if they were presumed pathogenic variants and/or planned to be assessed in future Xenopus and mouse model studies. TaqMan genotyping assays were conducted following the manufacturer’s guidelines. TaqMan reactions were performed in a 20 μL reaction volume in a Bio-Rad Hard-Shell 96-well plate and run using a Bio-Rad CFX96 Real-Time PCR Detection System (Bio-Rad, Hercules, CA, USA). The TaqMan genotyping results were then analyzed using Bio-Rad CFX Maestro Software (version 4.1.2433.1219) (Bio-Rad, Hercules, CA, USA) to determine the genotypes for each individual in the FASD cohort. All but one probe (rs7978237, NCOR2) amplified sufficiently to validate the WES genotype in all individuals.
2.6. Determining Differences between FASD Diagnostic Groups and Pathway Analysis
Statistically significant variants were grouped as follows: those unique to individuals with FAS, pFAS, or ARND and those significant variants that were common to both FAS and pFAS individuals (the two diagnostic categories that share clinical sentinel facial features) (see Supplementary Table S9). For genetic variants shared by both FAS and pFAS, the number of risk alleles in a given gene per person was determined, and this number was included in the represented major PAE-implicated pathway (RA, SHH, WNT, FGF, TGF- β, RAS-MAPK, RHO-RAC, and mTOR). The following operations were then applied: (1) If one variant in a gene was common to pFAS and FAS, other variants within that common gene were included. (2) Variants found in common to all three diagnostic groups were not included. (3) If a gene was found to not interact with any of the developmental pathways, then variants within that common gene were also not considered. The total number of risk alleles per pathway was compiled and then plotted using principal component analysis (PCA) using R software (version 4.1.2) to determine the degree of separation between FAS and pFAS diagnostic groups.
3. Results
3.1. The FASD-Diagnosed Cohort Has Increased Frequencies of RA and Alcohol Metabolism Gene Variants Known to Be Associated with FASD and Alcoholism
Alcohol dehydrogenases (ADHs) and acetaldehyde dehydrogenases (ALDHs) comprise a two-step sequential oxidation pathway where these enzyme families convert ethanol (alcohol) to acetic acid. Some members of these families also perform the two sequential oxidations of retinol (vitamin A) to produce RA (Figure 2). Many variants of these two enzyme families have been associated with increased alcohol consumption, dependence, and altered enzyme activity, but only a few have thus far been associated with FASD [32]. Our group and others have shown in animal models that alcohol-induced RA deficiency during gastrulation can result in most known FASD developmental phenotypes [33,34]. Accordingly, certain genetic variants of RA metabolism may predispose to RA deficiency following PAE, and these same variants would be found with increased allele frequencies in children diagnosed with FASD in this cohort. Using a candidate gene approach, all detected variants in the 44 genes of RA metabolism were analyzed (Figure 2 and Supplementary Table S1) in the FASD cohort whole exome sequencing data.
As hypothesized, significant variants were found in the genes coding for alcohol metabolic enzymes and the broader RA metabolic pathway associated with FASD. Specifically, 15 variants were more frequently observed in individuals with FASD, while five alleles were less frequently observed in individuals with FASD when compared to the 1000 Genomes Project dataset (Table 1); moreover, over half of these variants were previously shown to be associated with increased alcohol consumption, dependence, or altered enzyme activity (Table 1 and Figure 2; known alcohol risk variants enriched in our study are denoted by a red dot and resilience variants by a green dot). Known FASD protective variants in ADH1B were found to be significantly reduced in our FASD cohort (rs1229984 and rs2006702) [32,35,36]. These two variants modify the kinetic speed and binding efficiency of ADH1B, changing how fast alcohol is oxidized and, consequently, are associated with alcohol consumption and dependency. Noteworthy variants also associated with alcohol dependence and consumption found to be enriched in the cohort included the risk alleles rs1614972 and rs698 in ADH1C, along with rs3762894 and rs671 in ADH4 and ALDH2, respectively [37,38,39,40].

Table 1.
Variant analysis results of retinoic acid and alcohol metabolism candidate genes.
Other significant variants in other RA metabolism genes that could impact the severity of alcohol teratogenicity in a PAE individual were also found (Figure 1 and Table 1). One risk variant in RXRG, rs2134095, which affects the splicing of RXRG [41] was found to be enriched in the FASD cohort. RXRG is necessary to mediate RA-regulated transcription; therefore, rs2134095 may potentially hinder RA signaling efficiency in impacted individuals. Two risk variants, rs971756 and rs11857410, were found in STRA6, which is an essential transporter of retinol into the cell [42]. Interestingly, both STRA6 variants have been associated with Matthew–Wood syndrome, a genetic syndrome that has many comorbidities of FASD such as craniofacial, ocular, and cardiac defects. Both rs971756 and rs11857410 have a CADD score of over 10, making them likely damaging variants that reduce the amount of retinol entering the cell to be converted to RA, thereby reducing RA signaling, and likely sensitizing a PAE individual to develop FASD. New potential FASD risk variants in RA metabolism with damaging PolyPhen, SIFT, and/or CADD scores were also found in BCO2, CYP26C1, DGAT1, FABP4, LIPC, and NCOR2; these variants could influence RA levels and signaling, potentially increasing the risk that a PAE individual will develop FASD.
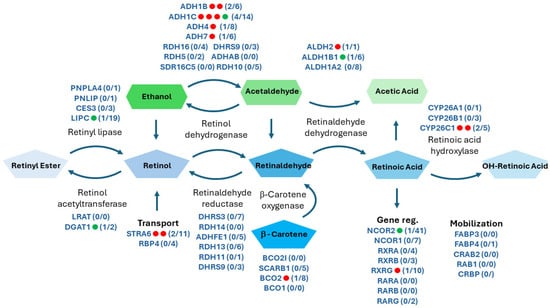
Figure 2.
The FASD cohort is enriched in FASD risk alleles throughout the retinoic acid and alcohol (ethanol metabolic pathways and deficient in resilience alleles). Ethanol (green highlight) and retinoic acid (blue highlight) metabolic pathways are shown. All major participating genes involved in specific metabolic steps are indicated across their respective enzymatic reaction. Red and green dots listed to the side of gene names represent enriched FASD risk and resilience alleles found in that gene, respectively, that were found to be significant in the FASD cohort when compared to the allele frequencies cited in the 1000 Genomes Project dataset. The numbers next to gene names show the number of risk and resilience alleles over the total number of detected variants for that gene that passed our threshold assessment (Figure adapted from Parihar M. et al., 2021 [43]).
3.2. The FASD Cohort Is Enriched in Both Risk and Resilience Variants in Retinoic Acid-Regulated Developmental Pathways
Research in animal models of PAE has shown that many important developmental pathways are impacted by alcohol including SHH, WNT, and TGF-β among others [15,20]. Many of these pathways are either directly or indirectly regulated by RA signaling during development [18,20]. As such, genes within PAE-impacted developmental pathways that are known to be regulated by RA were chosen to be examined (Supplementary Table S2). Interestingly, more significant frequency changes in resilience variants (16 reduced) were detected, compared to frequency changes in risk variants (11 enhanced).
When examining variants found in the genes in RA-controlled developmental pathways, 66% (18/24) were found to affect craniofacial, heart, and/or bone development. Eight significant variants associated with craniofacial malformations such as non-syndromic cleft lip and palate, mandibular prognathism, ocular abnormalities, and tooth defects are found in AXIN2, BMP2, BMP4, NOTCH3, PTCH1, SOX10, and WNT10A (Table 2 and Supplementary Table S6) [44,45,46,47,48,49,50,51]. The variants in these genes could increase the risk of developing FASD in a PAE individual given their potential to cause similar facial features as seen in children with FASD. Six significant variants within AXIN1, AXIN2, CAV1, GLI1, RYR2, and SFRP4 genes were discovered that have an impact on heart development and function [52,53,54,55,56], which could worsen cardiac malformations in FASD individuals. Four significant variants in the BMP2, WNT10B, and WNT16 genes were found and could have an impact on bone mineral density, ossification defects, or increased risk of bone fracture [57,58,59,60].

Table 2.
Variant analysis results of genes in retinoic acid-regulated pathways.
Additional significant variants were discovered in RA-regulated pathways that impact different aspects of development or tissue function that could adversely affect FASD individuals (Table 2 and Supplementary Table S6). Two variants with damaging PolyPhen and SIFT predictions were found in genes of the SHH pathway: rs2592595, associated with ectrodactyly, and rs11573590, found in GLI2 and PTCH3, respectively [61]. One significant variant was found in the WNT pathway, rs34072914 in WNT9B. This variant is associated with Mayer–Rokitansky–Küster–Hauser syndrome [62] and has a CADD score of 12.75, making it potentially pathogenic. Three significant variants were also detected in NOTCH genes: rs35769976 in NOTCH3, which has been found to lead to a disruption of its protein structure [63], and rs1048672 and rs8192591 in NOTCH3 and NOTCH4, respectively, which were both found to have damaging PolyPhen and SIFT predictions. rs1140475 in EGFR is another significant variant that is located in the protein kinase domain and associated with temporomandibular disorder [64,65].
3.3. The FASD Cohort Is Enriched in Risk Variants in the Causative Genes of Rare Neurodevelopmental Disorders
Many rare neurodevelopmental disorders (NDDs), such as 22q11.2 deletion syndrome (22q11.2DS) and Smith–Lemli–Opitz syndrome (SLOS), share FASD comorbidities and physical and neurological features in addition to their behavioural outcomes [20]. Given the comorbidity between FASD and these rare NDDs, we wondered whether certain variants in these rare disease genes could sensitize FASD risk outcomes in children with PAE. Accordingly, variants found in the FASD cohort within the causative genes of NDDs and their downstream gene targets were investigated (Supplementary Table S3). The FASD cohort was found to be statistically significant in 15 risk and 5 resilience alleles when compared to the 1000 Genomes Project dataset (Table 3 and Supplementary Table S7).

Table 3.
Variant analysis of causative genes of rare neurodevelopmental disorders.
Several potential FASD significant variants in the causative genes and/or their interaction partners associated with Campomelic Dysplasia, Apert, SLOS, 22q11.2DS, Three M Syndrome 2, and Aarskog syndromes were detected in our FASD cohort (Table 3 and Supplementary Table S7). Eleven statistically significant variants with the potential to play a role in the development of FASD in genes found in the 22q11.2 region were discovered in our FASD cohort. Notable among these were two significant variants found in TBX1, rs5746826, and rs41298840, the most well-characterized gene in 22q11.2DS. rs55975541 in CDC42BPG, a variant with damaging PolyPhen and SIFT predictions, was found to be enriched; CDC42BPG is downstream of CDC42, the direct downstream target of the causative gene of Aarskog syndrome, FGD1 [20]. The T variant of rs2229989 in SOX9, the causative gene of Campomelic Dysplasia [66], was also found to be more prevalent in our FASD cohort when compared to controls. The A and G alleles of rs2229989 result in an H169Q substitution that has only 46% of the activity of the WT enzyme and was found in a patient with mild Campomelic Dysplasia [66]. While the T allele results in a synonymous mutation, it did have a CADD score of 11.83, making it potentially pathogenic [15,21]. When examining variants within the FGF and FGFR genes, the FASD cohort was found to be enriched in the A allele of rs147057 in FGFR2, which has been found in a patient with Apert syndrome [67]. Cholesterol metabolism genes were also examined as SLOS—another NDD with shared FASD comorbidities—characterized by cholesterol deficiency due to mutations in the cholesterol-synthesizing enzyme DHCR7 [68]. A significant allele, rs2230808 in ABCA1, which plays an important role in cholesterol homeostasis by its function in the reverse cholesterol transport pathway, was detected. The FASD cohort was found to be enriched in the C allele of rs2230808, which has been associated with more severe phenotypes in SLOS patients [69]. Lastly, we found a significant allele, rs10932816, in OBSL1, one of the causative genes of Three M Syndrome 2 [70] that has a damaging SIFT prediction.
3.4. Determining Differences between FASD Diagnosis and Pathway Polygenic Risk Score Analysis
There is extensive evidence in human and animal studies that there is a clear underlying genetic component to FASD. Therefore, it should be possible to separate individuals with a FAS and pFAS diagnosis by their genetic differences in PCA. Additionally, more severe FASD clinical outcomes, including sharing sentinel facial features, may have increased genetic variants in the developmental pathways known to regulate the phenotypes observed. As an explorative exercise to test the hypothesis of genetic variants sorting by diagnosis, all candidate genetic variants found in common within both FAS and pFAS diagnostic groups were identified and then grouped by gene (Supplementary Table S9). The genes were then grouped by their interactions with developmental pathways—RA, SHH, WNT, FGF, TGF-β, RAS-MAPK, RHO-RAC, and mTOR [15,18,20] (eight pathways). Perhaps not surprisingly, a global PCA analysis of all genetic variants detected in the FASD cohort identified ethnicity as the largest principal component (Supplementary Figure S1a); however, PCA analysis of the significant variants in this study did not (Supplementary Figure S1b). Surprisingly, PCA analysis of genetic variants significantly enriched in the aforementioned developmental pathways sorted by diagnosis and, furthermore, with little bias towards ethnicity (Figure 3). RA was the most highly represented pathway with 44 significant variants (14%), followed by SHH at 29 variants (9%), WNT at 22 variants (7%), TGF-β at 18 variants (6%), RHO/RAC and mTOR at 13 variants (4%), and RAS/MAPK and FGF each at 11 variants (4%). Interestingly, 44% of the genes found to be common or unique to FAS and pFAS groups are causative genes of NDDs such as CHARGE, SLOS, and 22q11.2DS. Potentially pathogenic variants were discovered in the FAS and pFAS groups, including those known to cause craniofacial malformations, such as SOX9 (Campomelic Dysplasia), STRA6 (Matthew–Wood), CHD7 (CHARGE), MID1 and MED15 (Opitz–Kaveggia syndrome), and several 22q11.2DS genes [20]. Unexpectedly, two individuals were found to be heterozygous for a known pathogenic variant, rs138659167, in the DCHR7 gene for SLOS [71]. pFAS had the largest number of unique genes at 26, FAS had nine, and the ARND group had one (Supplementary Table S9).
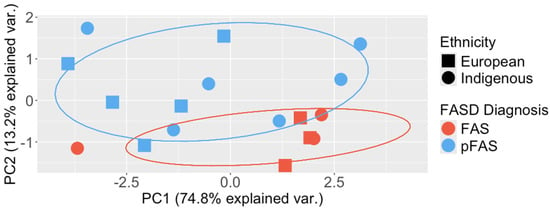
Figure 3.
The FASD cohort clusters by diagnosis when using pathway−based polygenic risk scores. When examining pathway-based polygenic risk scores for each patient using only variants in genes common to individuals with a FAS and pFAS diagnosis, they were found to cluster by diagnosis, and not by ethnicity.
4. Discussion
Herein we report 15 new risk variants and five new resilience variants in RA metabolism that have significant allele frequency changes in a cohort of 23 children diagnosed with FASD. Many impacted genes were found to have multiple associated risk or resilience variants that may independently affect gene and protein function. Allele frequencies of two ADH1B variants previously associated with FASD, rs2066702 and rs1229984, which are both protective variants associated with increased alcohol oxidation and less consumption, were found to be reduced [32,35,36]. Finding a significant reduction in the protective alleles of both rs2066702 and rs1229984 supports the employed candidate gene approach and the methodology used in this study. Moreover, eight new FASD-associated variants were found in alcohol metabolism genes [37,38,39,40,72,73,74,75]. Six of the eight alcohol metabolism variants are risk alleles; the other two variants are resilience alleles (Table 1 and Figure 2). Outside of the shared alcohol oxidizing enzymes, we identified nine additional new FASD variants in the remaining RA metabolism genes (seven risk and two resilience variants).
It is important to note that RA variants previously found associated with alcohol dependency and addiction are not functionally associated. These risks are not directly relevant to FASD outcomes in the embryo itself; rather, they are associated with the risk of embryo/developmental outcomes within the context of our hypothesis that PAE results in an RA deficiency within the embryo proper. Under conditions of PAE, the 15 new risk variants may enzymatically sensitize individuals to poorer maternal alcohol detoxification and, in turn, metabolically result in a state of RA deficiency in the embryo. When undergoing PAE, the accumulating acetaldehyde acts as a toxin in the embryo and its elimination is thus the priority [76,77]. As the alcohol and RA metabolic pathways share familial enzymes, both retinol and retinaldehyde dehydrogenases will help to detoxify alcohol and acetaldehyde during PAE [33,76,77]. Presumably, this is why many of the shared alcohol metabolism variants were originally associated with alcohol behaviour risks in the mother and father [32], and it would be interesting to determine which are inherited maternally or paternally. Here, their direct role in alcohol detoxification can explain enrichment in frequencies in a high number of risk alleles in these alcohol metabolism genes. Usurping retinol and retinaldehyde dehydrogenase for alcohol detoxification also comes at the cost of reduced synthesis of RA, particularly when it is essential to guide cellular spatiotemporal patterning [33]. Most individuals who are prenatally exposed to alcohol do not develop FASD [17], which is likely due to the robustness of the RA signaling network being able to adjust to acute PAE accordingly [43]. However, when damaging variants are within the RA and alcohol metabolism genes and already perturb the RA pathway, PAE as an additional factor might disrupt the pathway so severely that FASD develops. Similarly, the reduced frequency of five resilience alleles in our FASD cohort also favours alcohol detoxification at the expense of RA production, leading to RA deficiency.
When viewed through this lens, the FASD cohort appears to be particularly sensitized to alcohol teratogenesis, as individuals harbour multiple damaging variants throughout the shared RA and alcohol metabolism pathways. The rs971756 and rs11857410 variants in STRA6, which are associated with Matthew–Wood syndrome and were enriched in the FASD cohort, are a good example of this [42]. These STRA6 variants were also found uniquely in pFAS and FAS individuals, the more severe forms of FASD, demonstrating the importance of RA signaling when determining an individual’s risk of FASD outcomes following PAE.
When taken together, these findings support the proposed hypothesis, wherein PAE-induced RA deficiency is a potential underlying etiology of FASD, particularly during early gastrulation. Animal models have supported this hypothesis, both in studies conducted by our group and others, and now here, with the enrichment of risk alleles of RA metabolism in a human FASD cohort [33,34]. Our FASD cohort is small, and while it supports the hypothesis, these results need to be validated in a larger multiethnic cohort. To our surprise, there were not many damaging variants found in the ALDH1A2 gene (Supplementary Table S5), which has been shown to be competitively inhibited by ethanol during embryogenesis, thereby causing PAE-induced RA deficiency [33]. It may be that damaging variants in the ALDH1A2 gene may result in loss of pregnancy following PAE, as observed in animal models targeting the ALDH1A2 gene [78]. It is well known that PAE is associated with higher numbers of stillbirths and miscarriages [79]. FASD cohort studies that include parent genotyping, and dosage and duration of PAE insult will be able to explore this avenue of research.
Extending the candidate gene approach analysis to RA-interacting major developmental pathways known to be impacted by PAE, such as SHH, WNT, and TGF-β, identified many new potential FASD risk and resilience alleles (Table 2 and Supplementary Table S3) [15,20]. Interestingly, a greater number of resilience than risk alleles, 16 vs 11, were found in these pathways, compared to those found in the RA metabolism pathway. This was a surprising result given that animal models have shown these pathways are directly impacted by PAE, and when perturbed can exacerbate PAE teratogenesis [80,81]. Although these PAE-impacted pathways can be viewed as independent of each other, they are co-dependent and function through defined signaling gradients [82]. If one is disrupted, the others are consequently perturbed, perhaps amplifying disruptive effects following PAE and/or RA deficiency.
This study identified 15 risk and five resilience variants in causative genes of rare NDDs that share clinical comorbidity with FASD. Our group has previously identified several rare NDDs that share specific comorbidities with children with FASD, including the causative genes and/or their interaction partners in Campomelic Dysplasia, Apert, SLOS, 22q11.2DS, Three M Syndrome 2, and Aarskog syndromes (Table 3) [20]. Rare NDDs that share many co-morbidities with FASD may offer unique insights into the underlying genetics of FASD. Most of these NDDs are rare disorders requiring recessive homozygosity within a single gene. As such, these causative variants are likely to highlight a single genetic developmental pathway that plays a key role in the development of NDD phenotypes. Therefore, genetic variants may exist in the shared NDD gene pathway that could be sensitized following acute PAE and result in similar clinical NDD comorbidities. Finding 20 new genetic variants in NDD rare disease genes with the potential to influence PAE teratogenesis provides initial evidence that this may be the case. It is also important to highlight that we also found many low-frequency alleles that did not pass the filtering threshold in some known causative genes of NDDs, CHD7, BRD4, and a known SLOS pathogenic variant in DHCR7 [20,71], which could be considered FASD risk alleles. The SLOS pathogenic variant requires recessive homozygosity to develop the disorder; therefore, these children were not misdiagnosed, but are rather potentially sensitized to PAE by being heterozygous for this variant. If these findings hold true in larger cohort studies, this could increase the number of candidate genetic biomarkers for PAE teratogenesis, and the number of new targets of potential therapeutics for both NDD and FASD children.
Another unexpected finding was that newly identified risk variants are able to separate FAS and pFAS individuals within the FASD cohort by assessing variants found in genes shared between these two groups. However, this was only possible when we categorized shared variants using pathway-based polygenic risk score analysis, which is a higher-level approach that shows the impact of variants on major PAE-impacted signaling pathways (such as RA, SHH, and WNT). This assessment also required that we account for low-frequency alleles present in the cohort that were found in the shared genes of FAS and pFAS individuals (Supplementary Table S9). However, it is interesting to speculate that the different clinical diagnoses may align with high principal components associated with major developmental pathways. This may be due to an underlying connection between all of these pathways, as they are all involved in the regulation of neural crest cell development [20]. FASD and NDDs share co-morbidities and could be considered neurocristopathies due to their defects in neural crest cell lineage development that explain many of the shared craniofacial, neurological, and behavioural deficits seen in these disorders. Variants that identify risk alleles associated developmentally with FASD sentinel facial features may not greatly increase the diagnosis of FAS and pFAS in children with PAE. However, these results need to be validated in larger multiethnic FASD studies.
It is important to recognize this exploratory study has limitations. First, due to the small study cohort size, these potential genetic biomarkers need to be validated in a larger multiethnic and multi-diagnosis FASD cohort. Second, the discovered variants in this paper are only probable FASD risk and resilience alleles and require additional independent replication and validation to confirm their impact on protecting and sensitizing individuals to FASD. Third, the small sample size and the ability to segregate the FASD diagnosis groups from one another using pathway polygenic risk scores based on shared genetic variants found in the individuals with FAS and pFAS must also be replicated in larger multiethnic FASD studies to validate the findings of this study. Additionally, including trios (child FASD diagnosis, maternal and paternal genomes) in larger FASD studies would be insightful for many FASD-related analyses, such as PAE- and NDD-sensitizing variants. Lastly, this study was unable to have matched controls, nor is there an ideal reference population from which to draw control allelic frequencies. Currently, there is no public Canadian Indigenous Peoples database available to serve as a control for these kinds of genetic studies, although one is currently being assembled. While genetic differences are likely influencing the results of this study, the fact that individuals were segregated based on diagnosis potentially supports the methodology to use the 1000 Genomes Project dataset as a control cohort, despite the FASD cohort being heterogeneous. This suggests that ethnicity may not be the driving factor in the genetic differences found between the diagnostic groups and supports this study’s results to investigate a more diverse population in a similar methodology.
5. Conclusions
In this exploratory study, the goal was to investigate the variants found in the genes comprising the RA signaling network in 23 individuals with FASD. Novel FASD risk and resilience alleles in each of our three candidate gene lists were discovered: (1) RA and alcohol (ethanol) metabolism genes, (2) RA-interacting developmental pathway genes, and (3) the causative genes of rare NDDs and their direct signaling targets. Two previously known FASD protective variants were identified to be reduced in our cohort, and an additional 67 new potential FASD risk and resilience genetic variants were found. That these new variants were found in genes throughout the RA and alcohol metabolic pathways, in developmental pathways, and in known causative genes of NDDs that share co-morbidities with FASD suggests that there is a genetic vulnerability that sensitizes an individual to develop FASD in the presence of PAE. These results support the possibility that genetic biomarkers for the risk of FASD outcomes may help in the much earlier assessment of children with PAE and early signs of neurodevelopmental deficits. Surprisingly, the results of this study have shown that specific genetic variants of FASD risk and resilience may be able to be used to distinguish FASD outcomes in individuals of different FASD diagnoses and, moreover, possibly identify the risk of FASD outcomes in the larger majority of currently undiagnosed children with FASD. Ideally, this work will put us one step closer to developing a clinical genetic assessment tool to assess the risk of PAE teratogenesis and FASD diagnosis at a much younger age. This will allow for earlier intervention for FASD individuals that would significantly mitigate secondary disabilities associated with FASD and have a profound impact on the life course trajectory of individuals with this disorder and their families.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/biom14050569/s1, Figure S1: PCA analysis of all variants segregate by ethnicity; however, analysis of all variants found significant in this study largely do not; Table S1: Retinoic Acid and Ethanol Metabolism Gene Candidate List; Table S2: Retinoic Acid-Dependent Signaling Network Candidate Genes; Table S3: Candidate Genes in the Causative Genes of Rare Neurodevelopmental Disorders and their Interactive Partners; Table S4: Candidate Genes in Neurotransmitter, Forebrain Development and Collagen Genes; Table S5: Variant Analysis Results of Retinoic Acid and Alcohol Metabolism Candidate Genes; Table S6: Variant Analysis Results of Genes in Retinoic Acid-Regulated Pathways; Table S7: Variant Analysis Results of the Causative Genes of Rare Neurodevelopmental Disorders; Table S8: Variant Analysis Results of Neurotransmitter, Forebrain Development and Collagen Genes; Table S9: Variants in Genes Found by FASD Diagnosis; Supplemental Specific References [83,84,85,86,87,88,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116,117,118,119,120,121,122,123,124,125,126,127,128,129,130,131,132,133,134,135,136,137,138,139,140,141,142,143,144,145,146,147,148,149,150,151,152].
Author Contributions
The authors confirm contributions to the paper as follows: Conceptualization, A.F. and G.G.H.; methodology, G.G.H. and L.M.; validation, L.M. and M.P.; formal analysis, L.M.; investigation, L.M.; resources, J.N.R., S.W.S., R.F.W., A.E.C. and A.H.-D.; data curation, L.M.; writing—original draft preparation, L.M. and B.P.; writing—review and editing, G.G.H., L.M., B.P., A.F., M.P. and B.D.; visualization, L.M.; supervision, G.G.H.; funding acquisition, G.G.H. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by grants from Manitoba Liquor and Lotteries (55380) to G.G.H., Manitoba Medical Service Foundation (313152) to G.G.H., Canadian Institutes of Health Research (51270) to G.G.H., and Research Manitoba/Children’s Hospital Research Institute of Manitoba MSc Studentship and Kid’s Brain Health Network (54525) stipend support to L.M.
Institutional Review Board Statement
Health Research Ethics Board, Bannatyne Campus, University of Manitoba, Approved REB Ethics Reference Number: H2010:206.
Informed Consent Statement
Informed consent was obtained from all subjects involved in the study. Written informed consent has been obtained from the patients and respective caregivers to publish this paper.
Data Availability Statement
The 1000 Genomes Project frequencies employed in this study can be found on NCBI (https://www.ncbi.nlm.nih.gov/snp/, accessed on 2 March 2022). The original data and contributions presented in this study are included in the article/Supplementary Material; further inquiries can be directed to the corresponding author.
Acknowledgments
We thank Richard LeDuc, CHRIM, for assistance with statistical approaches and Emily Turner-Brannen, CHRIM, for assistance with TaqMan probe experiments.
Conflicts of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as potential conflicts of interest.
References
- Popova, S.; Lange, S.; Poznyak, V.; Chudley, A.E.; Shield, K.D.; Reynolds, J.N.; Murray, M.; Rehm, J. Population-Based Prevalence of Fetal Alcohol Spectrum Disorder in Canada. BMC Public Health 2019, 19, 845. [Google Scholar] [CrossRef] [PubMed]
- Cook, J.L.; Green, C.R.; Lilley, C.M.; Anderson, S.M.; Baldwin, M.E.; Chudley, A.E.; Conry, J.L.; Leblanc, N.; Loock, C.A.; Lutke, J.; et al. Fetal Alcohol Spectrum Disorder: A Guideline for Diagnosis across the Lifespan. Can. Med Assoc. J. 2016, 188, 191–197. [Google Scholar] [CrossRef] [PubMed]
- Chasnoff, I.J.; Wells, A.M.; King, L. Misdiagnosis and Missed Diagnoses in Foster and Adopted Children with Prenatal Alcohol Exposure. Pediatrics 2015, 135, 264–270. [Google Scholar] [CrossRef] [PubMed]
- Bonthius, D.J.; Goodlett, C.R.; West, J.R. Blood Alcohol Concentration and Severity of Microencephaly in Neonatal Rats Depend on the Pattern of Alcohol Administration. Alcohol 1988, 5, 209–214. [Google Scholar] [CrossRef] [PubMed]
- Pierce, D.R.; West, J.R. Alcohol-Induced Microencephaly during the Third Trimester Equivalent: Relationship to Dose and Blood Alcohol Concentration. Alcohol 1986, 3, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Parnell, S.; Holloway, H.T.; O’Leary-Moore, S.K.; Dehart, D.B.; Paniaqua, B.; Oguz, I.; Budin, F.; Styner, M.A.; Johnson, G.A.; Sulik, K.K. Magnetic Resonance Microscopy-Based Analyses of the Neuroanatomical Effects of Gestational Day 9 Ethanol Exposure in Mice. Neurotoxicol. Teratol. 2013, 39, 77–83. [Google Scholar] [CrossRef]
- Godin, E.A.; O’Leary-Moore, S.K.; Khan, A.A.; Parnell, S.E.; Ament, J.J.; Dehart, D.B.; Johnson, B.W.; Allan Johnson, G.; Styner, M.A.; Sulik, K.K. Magnetic Resonance Microscopy Defines Ethanol-Induced Brain Abnormalities in Prenatal Mice: Effects of Acute Insult on Gestational Day 7. Alcohol. Clin. Exp. Res. 2010, 34, 98–111. [Google Scholar] [CrossRef]
- Parnell, S.E.; O’Leary-Moore, S.K.; Godin, E.A.; Dehart, D.B.; Johnson, B.W.; Allan Johnson, G.; Styner, M.A.; Sulik, K.K. Magnetic Resonance Microscopy Defines Ethanol-Induced Brain Abnormalities in Prenatal Mice: Effects of Acute Insult on Gestational Day 8. Alcohol. Clin. Exp. Res. 2009, 33, 1001–1011. [Google Scholar] [CrossRef] [PubMed]
- Anthony, B.; Vinci-Booher, S.; Wetherill, L.; Ward, R.; Goodlett, C.; Zhou, F.C. Alcohol-Induced Facial Dysmorphology in C57BL/6 Mouse Models of Fetal Alcohol Spectrum Disorder. Alcohol 2010, 44, 659–671. [Google Scholar] [CrossRef]
- Kaminen-ahola, N.; Fahey, P.; Cox, T.C.; Kaminen-ahola, N.; Ahola, A.; Maga, M.; Mallitt, K.; Fahey, P.; Cox, T.C. Maternal Ethanol Consumption Alters the Epigenotype and the Phenotype of Offspring in a Mouse Model Maternal Ethanol Consumption Alters the Epigenotype and the Phenotype of Offspring in a Mouse Model. PLoS ONE 2010, 6, e1000811. [Google Scholar] [CrossRef]
- Keen, C.; Uriu-Adams, J.Y.; Skalny, A.; Grabeklis, A.; Grabeklis, S.; Green, K.; Yevtushok, L.; Wertelecki, W.W.; Chambers, C.D. The Plausibility of Maternal Nutritional Status Being a Contributing Factor to the Risk for Fetal Alcohol Spectrum Disorders: The Potential Influence of Zinc Status as an Example. Biofactors 2010, 36, 125–135. [Google Scholar] [CrossRef]
- Burd, L.; Blair, J.; Dropps, K. Prenatal Alcohol Exposure, Blood Alcohol Concentrations and Alcohol Elimination Rates for the Mother, Fetus and Newborn. J. Perinatol. 2012, 32, 652–659. [Google Scholar] [CrossRef]
- Alberry, B.; Singh, S.M. Developmental and Behavioral Consequences of Early Life Maternal Separation Stress in a Mouse Model of Fetal Alcohol Spectrum Disorder. Behav. Brain Res. 2016, 308, 94–103. [Google Scholar] [CrossRef]
- Weinberg, J. Prenatal Ethanol Exposure Alters Adrenocortical Development of Offspring. Alcohol. Clin. Exp. Res. 1989, 13, 73–83. [Google Scholar] [CrossRef]
- Smith, S.M.; Garic, A.; Berres, M.E.; Flentke, G.R. Genomic Factors That Shape Craniofacial Outcome and Neural Crest Vulnerability in FASD. Front. Genet. 2014, 5, 224. [Google Scholar] [CrossRef] [PubMed]
- Hemingway, S.J.A.; Bledsoe, J.M.; Davies, J.K.; Brooks, A.; Jirikowic, T.; Olson, E.M.; Thorne, J.C. Twin Study Confirms Virtually Identical Prenatal Alcohol Exposures Can Lead to Markedly Different Fetal Alcohol Spectrum Disorder Outcomes- Fetal Genetics Influences Fetal Vulnerability. Adv. Pediatr. Res. 2018, 5, 23. [Google Scholar] [CrossRef]
- Bandoli, G.; Kable, J.A.; Coles, C.D.; del Campo, M.; Suttie, M.; Chambers, C.D. Trajectories of Prenatal Alcohol Exposure and Behavioral Outcomes: Findings from a Community-Based Sample. Drug Alcohol Depend. 2022, 233, 109351. [Google Scholar] [CrossRef] [PubMed]
- Petrelli, B.; Bendelac, L.; Hicks, G.G.; Fainsod, A. Insights into Retinoic Acid Deficiency and the Induction of Craniofacial Malformations and Microcephaly in Fetal Alcohol Spectrum Disorder. Genesis 2019, 57, e23278. [Google Scholar] [CrossRef] [PubMed]
- Marrs, J.A.; Clendenon, S.G.; Ratcliffe, D.R.; Fielding, S.M.; Liu, Q.; Bosron, W.F. Zebrafish Fetal Alcohol Syndrome Model: Effects of Ethanol Are Rescued by Retinoic Acid Supplement. Alcohol 2010, 44, 707–715. [Google Scholar] [CrossRef]
- McKay, L.; Petrelli, B.; Chudley, A.E.; Hicks, G.G. Genetics of FASD: Confounding Rare Craniofacial and Neurodevelopmental Disorders May Identify Ethanol-Sensitizing Genetic Variants of FASD. In Neuromethods; Chudley, A., Geoffrey, H., Eds.; Humana Press Inc.: Totowa, NJ, USA, 2022; Volume 188, pp. 77–117. [Google Scholar]
- Yelin, R.; Schyr, R.B.H.; Kot, H.; Zins, S.; Frumkin, A.; Pillemer, G.; Fainsod, A. Ethanol Exposure Affects Gene Expression in the Embryonic Organizer and Reduces Retinoic Acid Levels. Dev. Biol. 2005, 279, 193–204. [Google Scholar] [CrossRef]
- Reynolds, J.N.; Weinberg, J.; Clarren, S.; Beaulieu, C.; Rasmussen, C.; Kobor, M.; Dube, M.P.; Goldowitz, D. Fetal Alcohol Spectrum Disorders: Gene-Environment Interactions, Predictive Biomarkers, and the Relationship Between Structural Alterations in the Brain and Functional Outcomes. Semin. Pediatr. Neurol. 2011, 18, 49–55. [Google Scholar] [CrossRef] [PubMed]
- Portales-Casamar, E.; Lussier, A.A.; Jones, M.J.; MacIsaac, J.L.; Edgar, R.D.; Mah, S.M.; Barhdadi, A.; Provost, S.; Lemieux-Perreault, L.P.; Cynader, M.S.; et al. DNA Methylation Signature of Human Fetal Alcohol Spectrum Disorder. Epigenet. Chromatin 2016, 9, 25. [Google Scholar] [CrossRef] [PubMed]
- Homer, N.; Merriman, B.; Nelson, S.F. BFAST: An Alignment Tool for Large Scale Genome Resequencing. PLoS ONE 2009, 4, e7767. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and Accurate Short Read Alignment with Burrows-Wheeler Transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Broad Institute. Available online: https://gatk.broadinstitute.org/hc/en-us/articles/360046222751-MarkDuplicates-Picard (accessed on 15 January 2021).
- Homer, N.; Nelson, S.F. Improved Variant Discovery through Local Re-Alignment of Short-Read next-Generation Sequencing Data Using SRMA. Genome Biol. 2010, 11, R99. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Depristo, M.A.; Banks, E.; Poplin, R.; Garimella, K.V.; Maguire, J.R.; Hartl, C.; Philippakis, A.A.; Del Angel, G.; Rivas, M.A.; Hanna, M.; et al. A Framework for Variation Discovery and Genotyping Using Next-Generation DNA Sequencing Data. Nat. Genet. 2011, 43, 491–501. [Google Scholar] [CrossRef] [PubMed]
- Adzhubei, I.A.; Schmidt, S.; Peshkin, L.; Ramensky, V.E.; Gerasimova, A.; Bork, P.; Kondrashov, A.S.; Sunyaev, S.R. A Method and Server for Predicting Damaging Missense Mutations. Nat. Methods 2010, 7, 248–249. [Google Scholar] [CrossRef]
- Ng, P.C.; Henikoff, S. Predicting Deleterious Amino Acid Substitutions. Genome Res. 2001, 11, 863–874. [Google Scholar] [CrossRef] [PubMed]
- Schubach, M.; Maass, T.; Nazaretyan, L.; Röner, S.; Kircher, M. CADD v1.7: Using Protein Language Models, Regulatory CNNs and Other Nucleotide-Level Scores to Improve Genome-Wide Variant Predictions. Nucleic Acids Res. 2024, 52, D1143–D1154. [Google Scholar] [CrossRef]
- Sambo, D.; Goldman, D. Genetic Influences on Fetal Alcohol Spectrum Disorder. Genes 2023, 14, 195. [Google Scholar] [CrossRef]
- Kot-Leibovich, H.; Fainsod, A. Ethanol Induces Embryonic Malformations by Competing for Retinaldehyde Dehydrogenase Activity during Vertebrate Gastrulation. Dis. Model. Mech. 2009, 2, 295–305. [Google Scholar] [CrossRef] [PubMed]
- Petrelli, B.; Oztürk, A.; Pind, M.; Ayele, H.; Fainsod, A.; Hicks, G.G. Genetically Programmed Retinoic Acid Deficiency during Gastrulation Phenocopies Most Known Developmental Defects Due to Acute Prenatal Alcohol Exposure in FASD. Front. Cell Dev. Biol. 2023, 11, 1208279. [Google Scholar] [CrossRef] [PubMed]
- McCarver, D.G.; Thomasson, H.R.; Martier, S.S.; Sokol, R.J.; Li, T. Alcohol Dehydrogenase-2*3 Allele Protects against Alcohol-Related Birth Defects among African Americans. J. Pharmacol. Exp. Ther. 1997, 283, 1095–1101. [Google Scholar] [PubMed]
- Viljoen, D.L.; Carr, L.G.; Foroud, T.M.; Brook, L.; Ramsay, M.; Li, T.K. Alcohol Dehydrogenase-2*2 Allele Is Associated with Decreased Prevalence of Fetal Alcohol Syndrome in the Mixed-Ancestry Population of the Western Cape Province, South Africa. Alcohol. Clin. Exp. Res. 2001, 25, 1719–1722. [Google Scholar] [CrossRef] [PubMed]
- Cleveland, H.H.; Schlomer, G.L.; Vandenbergh, D.J.; Wolf, P.S.A.; Feinberg, M.E.; Greenberg, M.T.; Spoth, R.L.; Redmond, C. Associations between Alcohol Dehydrogenase Genes and Alcohol Use across Early and Middle Adolescence: Moderation × Preventive Intervention. Dev. Psychopathol. 2018, 30, 297–313. [Google Scholar] [CrossRef] [PubMed]
- Meyers, J.L.; Shmulewitz, D.; Aharonovich, E.; Waxman, R.; Frisch, A.; Weizman, A.; Spivak, B.; Edenberg, H.J.; Gelernter, J.; Hasin, D.S. Alcohol-Metabolizing Genes and Alcohol Phenotypes in an Israeli Household Sample. Alcohol. Clin. Exp. Res. 2013, 37, 1872–1881. [Google Scholar] [CrossRef][Green Version]
- Edenberg, H.J.; Xuei, X.; Chen, H.J.; Tian, H.; Wetherill, L.F.; Dick, D.M.; Almasy, L.; Bierut, L.; Bucholz, K.K.; Goate, A.; et al. Association of Alcohol Dehydrogenase Genes with Alcohol Dependence: A Comprehensive Analysis. Hum. Mol. Genet. 2006, 15, 1539–1549. [Google Scholar] [CrossRef] [PubMed]
- Edenberg, H.J.; McClintick, J.N. Alcohol Dehydrogenases, Aldehyde Dehydrogenases, and Alcohol Use Disorders: A Critical Review. Alcohol. Clin. Exp. Res. 2018, 42, 2281–2297. [Google Scholar] [CrossRef]
- Yu, X.; Song, L.; Zheng, H.; Wei, S.; Wen, X.; Huang, B.; Liu, D. Association between Functional Genetic Variants in Retinoid X Receptor—α/γ and the Risk of Gestational Diabetes Mellitus in a Southern Chinese Population. Biosci. Rep. 2021, 41, BSR20211338. [Google Scholar] [CrossRef]
- Golzio, C.; Martinovic-Bouriel, J.; Thomas, S.; Mougou-Zrelli, S.; Grattagliano-Bessières, B.; Bonnière, M.; Delahaye, S.; Munnich, A.; Encha-Razavi, F.; Lyonnet, S.; et al. Matthew-Wood Syndrome Is Caused by Truncating Mutations in the Retinol-Binding Protein Receptor Gene STRA6. Am. J. Hum. Genet. 2007, 80, 1179–1187. [Google Scholar] [CrossRef]
- Parihar, M.; Bendelac-Kapon, L.; Gur, M.; Abbou, T.; Belorkar, A.; Achanta, S.; Kinberg, K.; Vadigepalli, R.; Fainsod, A. Retinoic Acid Fluctuation Activates an Uneven, Direction-Dependent Network-Wide Robustness Response in Early Embryogenesis. Front. Cell Dev. Biol. 2021, 9, 747969. [Google Scholar] [CrossRef]
- Noroozi, N.; Dastgheib, S.A.; Lookzadeh, M.H.; Mirjalili, S.R.; Noorishadkam, M.; Akbarian-Bafghi, M.J.; Neamatzadeh, H. Association of Axis Inhibition Protein 2 Polymorphisms with Non-Syndromic Cleft Lip with or without Cleft Palate in Iranian Children. Fetal Pediatr. Pathol. 2020, 39, 29–37. [Google Scholar] [CrossRef]
- Calvano, E.; Caio, K.; Bitencourt, L.; Julia, R.; Scariot, R.; Ricardo, P.N.; Olimpio, A.; Mírian, P.; Nakane, A.; Peter, M.; et al. Potential Interactions among Single Nucleotide Polymorphisms in Bone- and Cartilage-Related Genes in Skeletal Malocclusions. Orthod. Craniofacial Res. 2021, 24, 277–287. [Google Scholar] [CrossRef] [PubMed]
- Hao, J.; Gao, R.; Wu, W.; Hua, L.; Chen, Y.; Li, F.; Liu, J. Association between BMP4 Gene Polymorphisms and Cleft Lip with or without Cleft Palate in a Population from South China. Arch. Oral. Biol. 2018, 93, 95–99. [Google Scholar] [CrossRef]
- Rafighdoost, H.; Hashemi, M.; Danesh, H.; Bizhani, F.; Bahari, G.; Taheri, M. Association of Single Nucleotide and IRF6 with Non-Syndromic Cleft Lip with or without Cleft Palate in a Sample of the Southeast Iranian Population. J. Appl. Oral. Sci. 2017, 25, 650–656. [Google Scholar] [CrossRef][Green Version]
- Han, X.; Xiong, X.; Shi, X.; Chen, F.; Li, Y. Targeted Sequencing of NOTCH Signaling Pathway Genes and Association Analysis of Variants Correlated with Mandibular Prognathism. Head Face Med. 2021, 17, 17. [Google Scholar] [CrossRef] [PubMed]
- Carter, T.C.; Molloy, A.M.; Pangilinan, F.; Troendle, J.F.; Kirke, P.N.; Conley, M.R.; Orr, D.J.A.; Earley, M.; Mckiernan, E.; Lynn, E.C.; et al. Testing Reported Associations of Genetic Risk Factors for Oral Clefts in a Large Irish Study Population. Birth Defects Res. (Part A) 2010, 93, 84–93. [Google Scholar] [CrossRef]
- Ma, J.; Lin, K. A Novel Mutation of the PAX3 Gene in a Chinese Family with Waardenburg Syndrome Type I. Mol. Genet. Genom. Med. 2019, 7, e00798. [Google Scholar] [CrossRef] [PubMed]
- Saket, M.; Saliminejad, K.; Kamali, K.; Moghadam, F.A.; Anvar, N.E.; Khorram Khorshid, H.R.; Aghakhani, F.; Esmaeili, N.; Reza, H.; Khorshid, K. BMP2 and BMP4 Variations and Risk of Non-Syndromic Cleft Lip and Palate. Arch. Oral. Biol. 2016, 72, 134–137. [Google Scholar] [CrossRef]
- Crauciuc, G.A.; Iancu, M.; Olah, P.; Tripon, F.; Anciuc, M.; Gozar, L.; Togănel, R.; Bănescu, C. Significant Associations between AXIN1 Rs1805105, Rs12921862, Rs370681 Haplotypes and Variant Genotypes of AXIN2 Rs2240308 with Risk of Congenital Heart Defects. Int. J. Environ. Res. Public Health 2020, 17, 7671. [Google Scholar] [CrossRef]
- Lin, H.; Sinner, M.F.; Brody, J.A.; Arking, D.E.; Lunetta, K.L.; Rienstra, M.; Lubitz, S.A.; Magnani, J.W.; Sotoodehnia, N.; Mcknight, B.; et al. Targeted Sequencing in Candidate Genes for Atrial Fibrillation: The Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Targeted Sequencing Study. Heart Rhythm. 2014, 11, 452–457. [Google Scholar] [CrossRef] [PubMed]
- Milting, H.; Lukas, N.; Osterziel, K.; Peters, S.; Thieleczek, R.; Varsa, M. Composite Polymorphisms in the Ryanodine Receptor 2 Gene Associated with Arrhythmogenic Right Ventricular Cardiomyopathy. Cardiovasc. Res. 2006, 71, 496–505. [Google Scholar] [CrossRef] [PubMed]
- Hu, D.; Xiao, L.; Li, S.; Hu, S.; Sun, Y.; Wang, Y. Prediction of HF-Related Mortality Risk Using Genetic Risk Score Alone and in Combination With Traditional Risk Factors. Front. Cardiovasc. Med. 2021, 8, 634966. [Google Scholar] [CrossRef] [PubMed]
- Guan, W.; Zhang, J.; Chen, J. Connection of GLI1 Variants to Congenital Heart Disease Susceptibility. Medicine 2020, 99, e19868. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Gao, R.; Liu, Y.; He, B.; Lv, S.; Hao, D. The Pathogenesis of Ossification of the Posterior Longitudinal Ligament. Aging Dis. 2017, 8, 570–582. [Google Scholar] [CrossRef] [PubMed]
- Zmuda, J.M.; Yerges, L.M.; Kammerer, C.M.; Cauley, J.A.; Wang, X.; Nestlerode, C.S.; Wheeler, V.W.; Patrick, A.L.; Bunker, C.H.; Moffett, S.P.; et al. Association Analysis of WNT10B With Bone Mass and Structure Among Individuals of African Ancestry. J. Bone Miner. Res. 2009, 24, 437–447. [Google Scholar] [CrossRef]
- Zheng, H.; Tobias, J.H.; Duncan, E.; Evans, D.M.; Eriksson, J.; Bergstro, U.; Paternoster, L.; Yerges-armstrong, L.M.; Lehtima, T.; Goltzman, D.; et al. WNT16 Influences Bone Mineral Density, Cortical Bone, Bone Strength, and Osteoporotic Fracture Risk. PLoS ONE 2012, 8, e1002745. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Mori, S.; Ishizaki, T.; Takahashi, A.; Matsuda, K.; Koretsune, Y. Bone Reports Genetic Risk Score Based on the Prevalence of Vertebral Fracture in Japanese Women with Osteoporosis. Bone Rep. 2016, 5, 168–172. [Google Scholar] [CrossRef] [PubMed]
- David, D.; Marques, B.; Ferreira, C.; Vieira, P.; Corona-Rivera, A.; Ferreira, J.C.; Van Bokhoven, H. Characterization of Two Ectrodactyly-Associated Breakpoints Separated by 2.5Mb on chromosome 2q14.1–Q14.2. Eur. J. Hum. Genet. 2009, 17, 1024–1033. [Google Scholar] [CrossRef]
- Ma, W.; Li, Y.; Wang, M.; Li, H.; Su, T.; Li, Y.; Wang, S. Associations of Polymorphisms in WNT9B and PBX1 with Mayer-Rokitansky-Küster- Hauser Syndrome in Chinese Han. PLoS ONE 2015, 10, e0130202. [Google Scholar] [CrossRef][Green Version]
- Vlachakis, D.; Tsaniras, S.C.; Ioannidou, K.; Papageorgiou, L.; Baumann, M.; Kossida, S. A Series of Notch3 Mutations in CADASIL.; Insights from 3D Molecular Modelling and Evolutionary Analyses. J. Mol. Biochem. 2019, 3, 134. [Google Scholar]
- Toomey, S.; Madden, S.F.; Furney, S.J.; Fan, Y.; Mccormack, M.; Stapleton, C.; Cremona, M.; Cavalleri, G.L.; Milewska, M.; Elster, N.; et al. The Impact of ERBB-Family Germline Single Nucleotide Polymorphisms on Survival Response to Adjuvant Trastuzumab Treatment in HER2-Positive Breast Cancer. Oncotarget 2016, 7, 75518–75525. [Google Scholar] [CrossRef]
- Martin, L.J.; Smith, S.B.; Khoutorsky, A.; Magnussen, C.A.; Samoshkin, A.; Sorge, R.E.; Cho, C.; Yosefpour, N.; Sivaselvachandran, S.; Tohyama, S.; et al. Epiregulin and EGFR Interactions Are Involved in Pain Processing. J. Clin. Investig. 2017, 127, 3353–3366. [Google Scholar] [CrossRef] [PubMed]
- Matsushita, M.; Kitoh, H.; Kaneko, H.; Mishima, K.; Kadono, I.; Ishiguro, N.; Nishimura, G. A Novel SOX9 H169Q Mutation in a Family With Overlapping Phenotype of Mild Campomelic Dysplasia and Small Patella Syndrome. Am. J. Med. Genet. Part A 2013, 161, 2528–2534. [Google Scholar] [CrossRef]
- Munshi, A.; Khetarpal, P.; Das, S. Apert’s Syndrome: Study by Whole Exome Sequencing. Genes. Dis. 2018, 5, 119–122. [Google Scholar] [CrossRef] [PubMed]
- Nowaczyk, M.; Wassif, C. Smith-Lemli-Opitz Syndrome; Adam, M., Ardinger, H., Pagon, R.A., Eds.; GeneReviews: Seattle, WA, USA, 2024. [Google Scholar]
- Barboza-Cerda, M.C.; Barboza-Quintana, O.; Martínez-Aldape, G.; Garza-Guajardo, R.; Déctor, M.A. Phenotypic Severity in a Family with MEND Syndrome Is Directly Associated with the Accumulation of Potentially Functional Variants of Cholesterol Homeostasis Genes. Mol. Genet. Genomic Med. 2019, 7, e931. [Google Scholar] [CrossRef] [PubMed]
- Irving, M.; Holder-Espinasse, M. Three M Syndrome; Adam, M., Feldman, J., Mirzaa, G., Eds.; GeneReviews: Seattle, WA, USA, 2019. [Google Scholar]
- Park, J.E.; Lee, T.; Ha, K.; Ki, C.S. Carrier Frequency and Incidence Estimation of Smith—Lemli—Opitz Syndrome in East Asian Populations by Genome Aggregation Database (GnomAD) Based Analysis. Orphanet. J. Rare Dis. 2021, 16, 166. [Google Scholar] [CrossRef] [PubMed]
- Kranzler, H.R.; Zhou, H.; Kember, R.L.; Smith, R.V.; Justice, A.C.; Damrauer, S.; Tsao, P.S.; Klarin, D.; Baras, A.; Reid, J.; et al. Genome-Wide Association Study of Alcohol Consumption and Use Disorder in 274,424 Individuals from Multiple Populations. Nat. Commun. 2019, 10, 1499. [Google Scholar] [CrossRef] [PubMed]
- Gelernter, J.; Kranzler, H.R.; Sherva, R.; Almasy, L.; Koesterer, R.; Smith, A.H.; Anton, R.; Preuss, U.W.; Ridinger, M.; Rujescu, D. Genome-Wide Association Study of Alcohol Dependence: Significant Findings in African- and European-Americans Including Novel Risk Loci. Mol. Psychiatry 2014, 19, 41–49. [Google Scholar] [CrossRef]
- Luo, X.; Kranzler, H.R.; Zuo, L.; Wang, S.; Schork, N.J.; Gelernter, J. Multiple ADH Genes Modulate Risk for Drug Dependence in Both African- and European-Americans. Hum. Mol. Genet. 2007, 16, 380–390. [Google Scholar] [CrossRef]
- Bjerregaard, P.; Mikkelsen, S.S.; Becker, U.; Hansen, T.; Tolstrup, J.S. Genetic Variation in Alcohol Metabolizing Enzymes among Inuit and Its Relation to Drinking Patterns. Drug Alcohol Depend. 2014, 144, 239–244. [Google Scholar] [CrossRef] [PubMed]
- Shabtai, Y.; Bendelac, L.; Jubran, H.; Hirschberg, J.; Fainsod, A. Acetaldehyde Inhibits Retinoic Acid Biosynthesis to Mediate Alcohol Teratogenicity. Sci. Rep. 2018, 8, 347. [Google Scholar] [CrossRef] [PubMed]
- Shabtai, Y.; Fainsod, A. Competition between Ethanol Clearance and Retinoic Acid Biosynthesis in the Induction of Fetal Alcohol Syndrome. Biochem. Cell Biol. 2018, 96, 148–160. [Google Scholar] [CrossRef] [PubMed]
- Ribes, V.; Wang, Z.; Dollé, P.; Niederreither, K. Retinaldehyde Dehydrogenase 2 (RALDH2)-Mediated Retinoic Acid Synthesis Regulates Early Mouse Embryonic Forebrain Development by Controlling FGF and Sonic Hedgehog Signaling. Development 2006, 133, 351–361. [Google Scholar] [CrossRef] [PubMed]
- Bailey, B.A.; Sokol, R.J. Preterm Delivery, and Sudden Infant Death Syndrome. Alcohol Res. Health 2011, 34, 86–91. [Google Scholar] [PubMed]
- Kietzman, H.W.; Everson, J.L.; Sulik, K.K.; Lipinski, R.J. The Teratogenic Effects of Prenatal Ethanol Exposure Are Exacerbated by Sonic Hedgehog or Gli2 Haploinsufficiency in the Mouse. PLoS ONE 2014, 9, e89448. [Google Scholar] [CrossRef] [PubMed]
- Sambo, D.; Gohel, C.; Yuan, Q.; Sukumar, G.; Alba, C.; Dalgard, C.L.; Goldman, D. Cell Type-Specific Changes in Wnt Signaling and Neuronal Differentiation in the Developing Mouse Cortex after Prenatal Alcohol Exposure during Neurogenesis. Front. Cell Dev. Biol. 2022, 10, 1011974. [Google Scholar] [CrossRef] [PubMed]
- Niederreither, K.; Dollé, P. Retinoic Acid in Development: Towards an Integrated View. Nat. Rev. Genet. 2008, 9, 541–553. [Google Scholar] [CrossRef] [PubMed]
- Edenberg, H.J. The Genetics of Alcohol Metabolism: Role of Alcohol Dehydrogenase and Aldehyde Dehydrogenase Variants. Alcohol Res. Health 2007, 30, 5–13. [Google Scholar]
- Bierut, L.J.; Goate, A.M.; Breslau, N.; Johnson, E.O.; Bertelsen, S.; Fox, L.; Agrawal, A.; Bucholz, K.K.; Grucza, R.; Hesselbrock, V.; et al. ADH1B Is Associated with Alcohol Dependence and Alcohol Consumption in Populations of European and African Ancestry. Mol. Psychiatry 2012, 5, 445–450. [Google Scholar] [CrossRef]
- Warren, K.R.; Li, T.K. Genetic Polymorphisms: Impact on the Risk of Fetal Alcohol Spectrum Disorders. Birth Defects Res. A Clin. Mol. Teratol. 2005, 73, 195–203. [Google Scholar] [CrossRef]
- Hoang, Y.T.T.; Nguyen, Y.T.; Nguyen, H.D.; Le, A.T.P.; Bui, H.T.T.; Vu, N.P.; Nguyen, H.H. Single Nucleotide Polymorphisms of ADH1B, ADH1C and ALDH2 Genes in 235 People Living in Thai Nguyen Province of Vietnam. Asian Pac. J. Cancer Prev. 2022, 23, 4243–4251. [Google Scholar] [CrossRef] [PubMed]
- Luo, X.; Kranzler, H.R.; Zuo, L.; Yang, B.; Lappalainen, J.; Gelernter, J. ADH4 Gene Variation Is Associated with Alcohol and Drug Dependence: Results from Family Controlled and Population-Structured Association Studies. Pharmacogenet. Genom. 2005, 15, 755–768. [Google Scholar] [CrossRef]
- Luo, X.; Kranzler, H.R.; Zuo, L.; Wang, S.; Gelernter, J. Personality Traits of Agreeableness and Extraversion Are Associated with ADH4 Variation. Biol. Psychiatry 2007, 61, 599–608. [Google Scholar] [CrossRef]
- Pavan, M.; Ruiz, V.F.; Silva, F.A.; Sobreira, T.J.; Cravo, R.M.; Vasconcelos, M.; Marques, L.P.; Mesquita, S.M.F.; Krieger, J.E.; Lopes, A.A.B.; et al. ALDH1A2 (RALDH2) Genetic Variation in Human Congenital Heart Disease. BMC Med. Genet. 2009, 10, 113. [Google Scholar] [CrossRef] [PubMed]
- Deak, K.L.; Dickerson, M.E.; Linney, E.; Enterline, D.S.; George, T.M.; Melvin, E.C.; Graham, F.L.; Siegel, D.G.; Hammock, P.; Mehltretter, L.; et al. Analysis of ALDH1A2, CYP26A1, CYP26B1, CRABP1, and CRABP2 in Human Neural Tube Defects Suggests a Possible Association with Alleles in ALDH1A2. Birth Defects Res. A Clin. Mol. Teratol. 2005, 73, 868–875. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Zhong, Y.; Peng, Y.; Zhou, B.; Wang, Y.; Li, Q.; Zhang, Y.; Song, H.; Rao, L. Association Between AXIN1 Gene Polymorphisms and Dilated Cardiomyopathy in a Chinese Han Population. DNA Cell Biol. 2019, 38, 436–442. [Google Scholar] [CrossRef] [PubMed]
- Zhou, B.; Tang, T.; Chen, P.; Pu, Y.; Ma, M.; Zhang, D.; Li, L.; Zhang, P.; Song, Y.; Zhang, L. The Variations in the AXIN1 Gene and Susceptibility to Cryptorchidism. J. Pediatr. Urol. 2015, 11, 132.e1–132.e5. [Google Scholar] [CrossRef] [PubMed]
- Styrkarsdottir, U.; Thorleifsson, G.; Gudjonsson, S.A.; Sigurdsson, A.; Center, J.R.; Lee, S.H.; Nguyen, T.V.; Kwok, T.C.Y.; Lee, J.S.W.; Ho, S.C.; et al. Sequence Variants in the PTCH1 Gene Associate with Spine Bone Mineral Density and Osteoporotic Fractures. Nat. Commun. 2016, 7, 10129. [Google Scholar] [CrossRef]
- Callahan, N.; Modesto, A.; Meira, R.; Seymen, F.; Patir, A.; Vieira, A.R. Axis Inhibition Protein 2 (AXIN2) Polymorphisms and Tooth Agenesis. Arch. Oral. Biol. 2009, 54, 45–49. [Google Scholar] [CrossRef]
- Tian, T.; Lei, Y.; Chen, Y.; Karki, M.; Jin, L.; Finnell, R.H.; Wang, L.; Ren, A. Somatic Mutations in Planar Cell Polarity Genes in Neural Tissue from Human Fetuses with Neural Tube Defects. Hum. Genet. 2020, 139, 1299–1314. [Google Scholar] [CrossRef] [PubMed]
- Parmalee, N.L.; Schubert, C.; Merriam, J.E.; Allikmets, K.; Bird, A.C.; Gillies, M.C.; Peto, T.; Figueroa, M.; Friedlander, M.; Fruttiger, M.; et al. Analysis of Candidate Genes for Macular Telangiectasia Type 2. Mol. Vis. 2010, 16, 2718–2726. [Google Scholar] [PubMed]
- Lasabova, Z.; Stanclova, A.; Grendar, M.; Mikolajcikova, S.; Calkovska, A.; Lenhartova, N.; Ziak, P.; Matasova, K.; Caprnda, M.; Kruzliak, P.; et al. Genetic Association of Single Nucleotide Polymorphisms of FZD4 and BDNF Genes with Retinopathy of Prematurity. Ophthalmic Genet. 2018, 39, 332–337. [Google Scholar] [CrossRef] [PubMed]
- Magic, M.; Zeljic, K.; Jovandic, S.; Stepic, J.; Pejovic, M.; Colic, S.; Magic, Z.; Supic, G. Hedgehog Signaling Pathway and Vitamin D Receptor Gene Variants as Potential Risk Factors in Odontogenic Cystic Lesions. Clin. Oral. Investig. 2018, 23, 2675–2684. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.; Wang, S.; Zheng, Z. Etiology of Hypospadias: A Comparative Review of Genetic Factors and Developmental Processes between Human and Animal Models. Res. Rep. Urol. 2020, 12, 673–686. [Google Scholar] [CrossRef] [PubMed]
- Renard, E.; Chéry, C.; Oussalah, A.; Josse, T.; Perrin, P.; Tramoy, D.; Voirin, J.; Klein, O.; Leheup, B.; Feillet, F.; et al. Exome Sequencing of Cases with Neural Tube Defects Identifies Candidate Genes Involved in One—Carbon/Vitamin B12 Metabolisms and Sonic Hedgehog Pathway. Hum. Genet. 2019, 138, 703–713. [Google Scholar] [CrossRef] [PubMed]
- Deneault, E.; Faheem, M.; White, S.H.; Rodrigues, D.C.; Sun, S.; Wei, W.; Piekna, A.; Thompson, T.; Howe, J.L.; Chalil, L.; et al. CNTN5-/+or EHMT2-/+human IPSC-Derived Neurons from Individuals with Autism Develop Hyperactive Neuronal Networks. Elife 2019, 8, e40092. [Google Scholar] [CrossRef] [PubMed]
- Pinard, A.; Eudes, N.; Mitchell, J.; Bajolle, F.; Grelet, M.; Okoronkwo, J.; Bonnet, D.; Collod, G.; Stéphane, B. Analysis of HOXB1 Gene in a Cohort of Patients with Sporadic Ventricular Septal Defect. Mol. Biol. Rep. 2018, 45, 1507–1513. [Google Scholar] [CrossRef] [PubMed]
- Dargis, N.; Lamontagne, M.; Gaudreault, N.; Sbarra, L.; Henry, C.; Pibarot, P.; Mathieu, P.; Bossé, Y. Identification of Gender-Specific Genetic Variants in Patients With Bicuspid Aortic Valve. Am. J. Cardiol. 2016, 117, 420–426. [Google Scholar] [CrossRef]
- Girdauskas, E.; Kaemmerer, H.; Kodolitsch, Y. Von Unravelling the Pathogenetic Mechanisms in Congenital Aortopathies: Need for an Integrative Translational Approach. J. Clin. Med. 2020, 9, 204. [Google Scholar] [CrossRef]
- Zhu, C.; Wang, Y.; Zeng, Q.; Qian, Y.; Li, H.; Yang, Z. Combined Effects of Age and Polymorphisms in Notch3 in the Pathogenesis of Cerebral Infarction Disease. Metab. Brain Dis. 2016, 31, 1157–1164. [Google Scholar] [CrossRef] [PubMed]
- Sowers, L.P.; Loo, L.; Wu, Y.; Campbell, E.; Ulrich, J.D.; Wu, S.; Paemka, L.; Wassink, T.; Meyer, K.; Bing, X.; et al. Disruption of the Non-Canonical Wnt Gene PRICKLE2 Leads to Autism-like Behaviors with Evidence for Hippocampal Synaptic Dysfunction. Mol. Psychiatry 2013, 18, 1077–1089. [Google Scholar] [CrossRef]
- Koop, A.; Goldmann, P.; Chen, S.R.W.; Thieleczek, R.; Varsa, M. ARVC-Related Mutations in Divergent Region 3 Alter Functional Properties of the Cardiac Ryanodine Receptor. Biophys. J. 2008, 94, 4668–4677. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Martin, P.; Yang, X.; Robin, N.; Lam, E.; Rabinowitz, J.S.; Erdman, C.A.; Quinn, J.; Weiss, L.A.; Hamilton, S.P.; Kwok, P.; et al. A Rare WNT1 Missense Variant Overrepresented in ASD Leads to Increased Wnt Signal Pathway Activation. Transl. Psychiatry 2013, 3, e301. [Google Scholar] [CrossRef]
- Christodoulides, C.; Scarda, A.; Granzotto, M.; Milan, G.; Dalla Nora, E.; Keogh, J.; De Pergola, G.; Stirling, H.; Pannacciulli, N.; Sethi, J.; et al. WNT10B Mutations in Human Obesity. Diabetologia 2006, 49, 678–684. [Google Scholar] [CrossRef]
- Mues, G.; Bonds, J.; Xiang, L.; Vieira, A.R.; Seymen, F.; Klein, O.; Souza, R.N.D. The WNT10A Gene in Ectodermal Dysplasias and Selective Tooth Agenesis. Am. J. Med. Genet. A 2014, 164, 2455–2460. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Jiang, L.; Yang, H.; Wu, L.; Xu, N.; Zhou, X.; Li, J. Variations of Wnt/Β-catenin Pathway-related Genes in Susceptibility to Knee Osteoarthritis: A Three-centre Case-control Study. J. Cell Mol. Med. 2019, 23, 8246–8257. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Liu, H.; Dong, Y. Significance of Neurexin and Neuroligin Polymorphisms in Regulating Risk of Hirschsprung’s Disease. J. Investig. Med. 2018, 66, 944–951. [Google Scholar] [CrossRef]
- Corsten-Janssen, N.; Saitta, S.C.; Hoefsloot, L.H.; McDonal-McGinn, D.M.; Driscoll, D.A.; Derks, R.; Dickinson, K.A.; Kerstjens-Frederiske, W.S.; Emanuel, B.S.; Zackai, E.H.; et al. More Clinical Overlap between 22q11 . 2 Deletion Syndrome and CHARGE Syndrome than Often Anticipated. Mol. Syndromol. 2013, 4, 235–245. [Google Scholar] [CrossRef]
- Cuola, D.; Li, Y.; Yuanyuan, C.; Jianghua, L.; Shuguang, W. Association Study of Catechol-o-Methyltransferase and Alpha-1-Adrenergic Receptor Gene Polymorphisms with Multiple Phenotypes of Heroin Use Disorder. Neurosci. Lett. 2021, 748, 135677. [Google Scholar] [CrossRef]
- Bender, H.-U.; Almashanu, S.; Steel, G.; Hu, C.-A.; Lin, W.-W.; Willis, A.; Pulver, A.; Valle, D. Functional Consequences of PRODH Missense Mutations. Am. J. Hum. Genet. 2005, 76, 409–420. [Google Scholar] [CrossRef]
- Clelland, C.L.; Drouet, V.; Rilett, K.C.; Smeed, J.A.; Nadrich, R.H.; Rajparia, A.; Read, L.L.; Clelland, J.D. Evidence That COMT Genotype and Proline Interact on Negative-Symptom Outcomes in Schizophrenia and Bipolar Disorder. Transl. Psychiatry 2016, 6, e891. [Google Scholar] [CrossRef] [PubMed]
- Mozzi, A.; Guerini, F.R.; Forni, D.; Costa, A.S.; Nemni, R.; Baglio, F.; Cabinio, M.; Riva, S.; Pontremoli, C.; Clerici, M.; et al. REST, a Master Regulator of Neurogenesis, Evolved under Strong Positive Selection in Humans and in Non Human Primates. Sci. Rep. 2017, 7, 9530. [Google Scholar] [CrossRef]
- Ravizza, T.; Onat, F.Y.; Brooks-Kayal, A.R.; Depaulis, A.; Galanopoulou, A.S.; Mazarati, A.; Numis, A.L.; Sankar, R.; Friedman, A. WONOEP Appraisal: Biomarkers of Epilepsy-Associated Comorbidities. Epilepsia 2017, 58, 331–342. [Google Scholar] [CrossRef]
- Xu, Y.; Wang, J.; Xu, R.; Zhao, P.; Wang, X.; Sun, H.; Bao, L.; Shen, J. Detecting 22q11.2 Deletion in Chinese Children with Conotruncal Heart Defects and Single Nucleotide Polymorphisms in the Haploid TBX1 Locus. BMC Med. Genet. 2011, 12, 169. [Google Scholar] [CrossRef] [PubMed]
- Akiyama, K.; Saito, A.; Saito, S.; Ozeki, Y.; Watanabe, T. Association of Genetic Variants at 22q11.2 Chromosomal Region with Cognitive Performance in Japanese Patients with Schizophrenia. Schizophr. Res. Cogn. 2019, 17, 100134. [Google Scholar] [CrossRef]
- Caputo, V.; Strafella, C.; Termine, A.; Campione, E. RNAseq-Based Prioritization Revealed COL6A5, COL8A1, COL10A1 and MIR146A as Common and Differential Susceptibility Biomarkers for Psoriasis and Psoriatic Arthritis: Confirmation from Genotyping Analysis of 1417 Italian Subjects. Int. J. Mol. Sci. 2020, 21, 2740. [Google Scholar] [CrossRef] [PubMed]
- Den Hollander, W.; Pulyakhina, I.; Boer, C.; Bomer, N.; Van Der Breggen, R.; Arindrarto, W.; De Almeida, R.C.; Lakenberg, N.; Sentner, T.; Laros, J.F.J.; et al. Annotating Transcriptional Effects of Genetic Variants in Disease-Relevant Tissue: Transcriptome-Wide Allelic Imbalance in Osteoarthritic Cartilage. Arthritis Rheumatol. 2019, 71, 561–570. [Google Scholar] [CrossRef] [PubMed]
- Trefilova, V.V.; Shnayder, N.A.; Petrova, M.M.; Kaskaeva, D.S.; Tutynina, O.V.; Petrov, K.V.; Popova, T.E.; Balberova, O.V.; Medvedev, G.V.; Nasyrova, R.F. The Role of Polymorphisms in Collagen-Encoding Genes in Intervertebral Disc Degeneration. Biomolecules 2021, 11, 1279. [Google Scholar] [CrossRef]
- Jamieson, S.E.; De Roubaix, L.; Cortina-borja, M.; Tan, H.K.; Mui, E.J.; Cordell, H.J.; Kirisits, M.J.; Miller, E.N.; Peacock, C.S.; Hargrave, A.C.; et al. Genetic and Epigenetic Factors at COL2A1 and ABCA4 Influence Clinical Outcome in Congenital Toxoplasmosis. Investig. Ophthalmol. Vis. Sci. 2008, 3, e2285. [Google Scholar] [CrossRef]
- Tang, Y.; Epstein, M.P.; Anderson, G.M.; Zabetian, C.P.; Cubells, J.F. Genotypic and Haplotypic Associations of the DBH Gene with Plasma Dopamine b -Hydroxylase Activity in African Americans. Eur. J. Hum. Genet. 2007, 15, 878–883. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Lopez, E.; Kawasawa-imamura, Y.; Zhang, L.; Huang, X.; Koltun, W.A.; Coates, M.D.; Vrana, K.E. A Single Nucleotide Polymorphism in Dopamine Beta Hydroxylase (Rs6271 (C > T)) Is over-Represented in Inflammatory Bowel Disease Patients and Reduces Circulating Enzyme. PLoS ONE 2019, 14, e0210175. [Google Scholar] [CrossRef] [PubMed]
- Punchaichira, T.J.; Dey, S.K.; Mukhopadhyay, A. Characterization of SNPs in the Dopamine-β-Hydroxylase Gene Providing New Insights into Its Structure-Function Relationship. Neurogenetics 2017, 18, 155–168. [Google Scholar] [CrossRef] [PubMed]
- Ates, O.; Cam, F.; Erdogan, S.; Sezer, S.; Karakus, N. Association between 1603C > T Polymorphism of DBH Gene and Bipolar Disorder in a Turkish Population. Gene 2013, 519, 356–359. [Google Scholar] [CrossRef] [PubMed]
- Kukshal, P.; Chowdari, V.; Srivastava, V.; Wood, J.; Mcclain, L.; Bhatia, T.; Bhagwat, A.M.; Neelkanth, S.; Laxmikant, V.; Thelma, B.K. Dopaminergic Gene Polymorphisms and Cognitive Function in a North Indian Schizophrenia Cohort. J. Psychiatr. Res. 2013, 47, 1615–1622. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yeung, E.W.; Craggs, J.G.; Gizer, I.R. Comorbidity of Alcohol Use Disorder and Chronic Pain: Genetic Influences on Brain Reward and Stress Systems. Alcohol. Clin. Exp. Res. 2017, 41, 1831–1848. [Google Scholar] [CrossRef] [PubMed]
- Meyers, J.L.; Nyman, E.; Loukola, A.; Rose, R.J.; Kaprio, J.; Dick, D.M. The Association between DRD2/ANKK1 and Genetically Informed Measures of Alcohol Use and Problems. Addict. Biol. 2013, 18, 523–536. [Google Scholar] [CrossRef] [PubMed]
- Franco, G.B.; Mara, F.; Bertoli, D.P.; De Souza, J.F.; Scariot, R. Dopamine Receptor D2 and Ankyrin Repeat Domain Containing One in Temporomandibular Disorder of Adolescents. Int. J. Paediatr. Dent. 2019, 29, 748–755. [Google Scholar] [CrossRef]
- Popescu, A.; Marian, M.; Drăgoi, A.N.A.M.; Costea, R.V. Understanding the Genetics and Neurobiological Pathways behind Addiction (Review). Exp. Ther. Med. 2021, 21, 544. [Google Scholar] [CrossRef]
- Lundwall, R.A.; Guo, D.; Dannemiller, J.L. Exogenous Visual Orienting Is Associated with Specific Neurotransmitter Genetic Markers: A Population-Based Genetic Association Study. PLoS ONE 2012, 7, e30731. [Google Scholar] [CrossRef]
- Kirley, A.; Lowe, N.; Mullins, C.; Mccarron, M.; Daly, G.; Waldman, I.; Fitzgerald, M.; Gill, M.; Hawi, Z. Phenotype Studies of the DRD4 Gene Polymorphisms in ADHD: Association With Oppositional Defiant Disorder and Positive Family History Analysis of Association With. Am. J. Med. Genet. Part B 2004, 131, 38–42. [Google Scholar] [CrossRef] [PubMed]
- Nemoda, Z.; Szekely, A.; Sasvari-szekely, M. Neuroscience and Biobehavioral Reviews Psychopathological Aspects of Dopaminergic Gene Polymorphisms in Adolescence and Young Adulthood. Neurosci. Biobehav. Rev. 2011, 35, 1665–1686. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, M.; Hattori, E.; Yamada, K.; Iwayama, Y.; Toyota, T.; Iwata, Y.; Tsuchiya, K.; Sugihara, G.; Hashimoto, K.; Watanabe, H.; et al. Association and Synergistic Interaction between Promoter Variants of the DRD4 Gene in Japanese Schizophrenics. J. Hum. Genet. 2007, 52, 86–91. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wang, S.; Chen, Y.; Lee, C.; Cheng, C. Opioid Addiction, Genetic Susceptibility, and Medical Treatments: A Review. Int. J. Mol. Sci. 2019, 20, 4294. [Google Scholar] [CrossRef]
- Haenisch, B.; Linsel, K.; Br, M.; Gilsbach, R.; Propping, P.; Markus, M.N.; Rietschel, M.; Fimmers, R.; Maier, W.; Zobel, A.; et al. Association of Major Depression With Rare Functional Variants in Norepinephrine Transporter and Serotonin 1A Receptor Genes. Am. J. Med. Genet. Part B 2008, 150, 1013–1016. [Google Scholar] [CrossRef] [PubMed]
- Brüss, M.; Kostanian, A.; Bönisch, H.; Göthert, M. The Naturally Occurring Arg 219 Leu Variant of the Human 5-HT 1A Receptor: Impairment of Signal Transduction. Pharmacog. Genom. 2005, 15, 257–264. [Google Scholar] [CrossRef]
- Fanous, A.H.; Chen, X.; Wang, X.; Amdur, R.; O’Neill, A.; Walsh, D.; Kendler, K. Genetic Variation in the Serotonin 2A Receptor and Suicidal Ideation in a Sample of 270 Irish High-Density Schizophrenia Families. Am. J. Med. Genet. B Neuropsychiatr. Genet. 2009, 150, 411–417. [Google Scholar] [CrossRef]
- Tavares, G.A.; Torres, A.; Souza, J.A. De Early Life Stress and the Onset of Obesity: Proof of MicroRNAs ’ Involvement Through Modulation of Serotonin and Dopamine Systems’ Homeostasis. Front. Physiol. 2020, 11, 925. [Google Scholar] [CrossRef]
- Mccaffery, J.M.; Duan, Q.L.; Frasure-Smith, N.; Lespérance, F.; Théroux, P.; Rouleau, G.A.; Dubé, M.-P. Genetic Predictors of Depressive Symptoms in Cardiac Patients. Am. J. Med. Genet. B Neuropsychiatr. Genet. 2009, 150, 381–388. [Google Scholar] [CrossRef]
- Jakubczyk, A.; Klimkiewicz, A.; Kopera, M.; Krasowska, A.; Wrzosek, M.; Matsumoto, H.; Burmeister, M.; Brower, K.J.; Wojnar, M. The CC Genotype in the T102C HTR2A Polymorphism Predicts Relapse in Individuals after Alcohol Treatment. J. Psychiatr. Res. 2013, 47, 527–533. [Google Scholar] [CrossRef][Green Version]
- Jakubczyk, A.; Wrzosek, M.; Łukaszkiewicz, J.; Sadowska-Mazuryk, J.; Matsumoto, H.; Śliwerska, E.; Glass, J.; Burmeister, M.; Browser, K.J.; Wojnar, M. The CC Genotype in HTR2A T102C Polymorphism Is Associated with Behavioral Impulsivity in Alcohol-Dependent Patients. J. Psychiatr. Res. 2012, 46, 44–49. [Google Scholar] [CrossRef][Green Version]
- Cao, J.; Liu, X.; Han, S.; Zhang, C.K.; Liu, Z.; Li, D. Association of the HTR2A Gene with Alcohol and Heroin Abuse. Hum. Genet. 2014, 133, 357–365. [Google Scholar] [CrossRef]
- Sinopoli, V.M.; Burton, C.L.; Arnold, P.D. Neuroscience and Biobehavioral Reviews Review Article A Review of the Role of Serotonin System Genes in Obsessive-Compulsive Disorder. Neurosci. Biobehav. Rev. 2017, 80, 372–381. [Google Scholar] [CrossRef] [PubMed]
- Cheah, S.-Y.; Lawford, B.R.; Young, R.M.; Morris, C.P.; Voisey, J. MRNA Expression and DNA Methylation Analysis of Serotonin Receptor 2A (HTR2A) in the Human Schizophrenic Brain. Genes 2017, 8, 14. [Google Scholar] [CrossRef] [PubMed]
- Rappold, G.; Walstab, J.; Hammer, C.; Bo, H. Naturally Occurring Variants in the HTR3B Gene Significantly Alter Properties of Human Heteromeric 5-Hydroxytryptamine-3A/B Receptors. Pharmacog. Genom. 2008, 18, 793–802. [Google Scholar] [CrossRef]
- Yosifova, A.; Mushiroda, T.; Stoianov, D.; Vazharova, R.; Dimova, I.; Karachanak, S.; Zaharieva, I.; Milanova, V.; Madjirova, N.; Gerdjikov, I.; et al. Case-Control Association Study of 65 Candidate Genes Revealed a Possible Association of a SNP of HTR5A to Be a Factor Susceptible to Bipolar Disease in Bulgarian Population. J. Affect. Disord. 2009, 117, 87–97. [Google Scholar] [CrossRef] [PubMed]
- Stoychev, K.; Dilkov, D.; Naghavi, E.; Kamburova, Z. Genetic Basis of Dual Diagnosis: A Review of Genome-Wide Association Studies (GWAS) Focusing on Patients with Mood or Anxiety Disorders and Co-Occurring Alcohol-Use Disorders. Diagnostics 2021, 11, 1055. [Google Scholar] [CrossRef]
- Birkett, J.T.; Arranz, M.J.; Munro, J.; Osbourn, S.; Kerwin, R.W.; Collier, D.A. Association Analysis of the 5-HT5A Gene in Depression, Psychosis and Antipsychotic Response. Neuroreport 2000, 11, 2017–2020. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).