Ribonucleoprotein Granules: Between Stress and Transposable Elements
Abstract
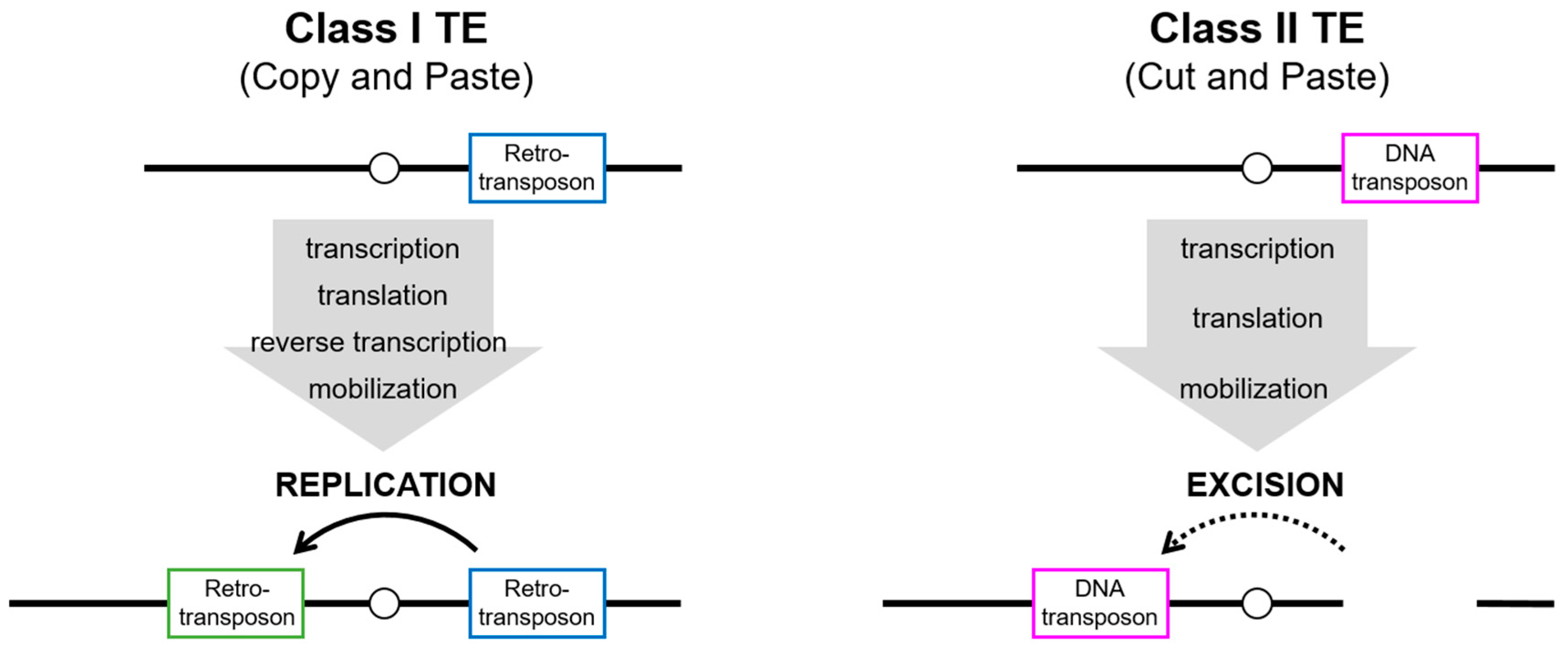
1. Invasion of Transposable Elements (TEs) and Host Defense System
2. Cytoplasmic RNP Granules
2.1. Stress Granule (SG)
2.1.1. Sequestration of RNAs into SG
2.1.2. Cellular Signaling and Apoptosis
2.1.3. Antiviral Function
2.2. Processing Body (P-body)
2.2.1. Composition of P-body
2.2.2. Function of P-body
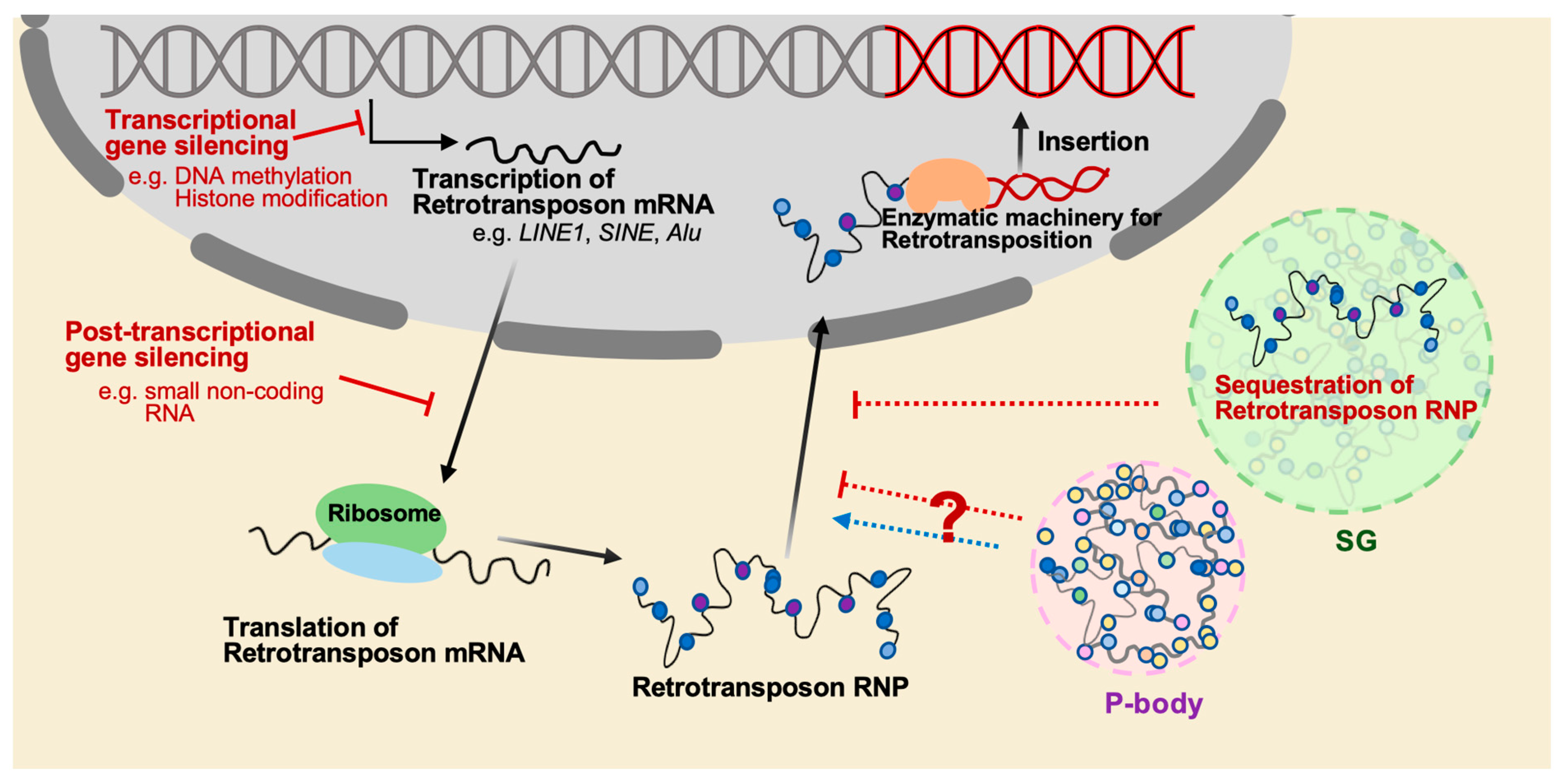
3. Current Understanding of TEs Related to Cytoplasmic RNP Granules
3.1. Stress Granule (SG) and TE Regulation
3.2. Processing Body (P-body) and TE Regulation
| Type of RNP Granule | TE | TE Component | Co-Localized with RNP Granule Components | References | Effect on Retrotransposition |
|---|---|---|---|---|---|
| Stress granule | LINE1 | ORF1p (engineered) ORF2p (engineered) ORF1p (Antibody) ORF2p (Antibody) 3′ UTR (MS2 fusion) L1 5′ UTR of LINE1 | TIA1, G3BP, HuR, eIF3η G3BP, eIF3η TIA-1, G3BP, eIF3η Co-localized with ORF1p TIA-1 Co-localized with ORF1p | [99,100,101,104,105,109,129] [100] [99,101,103,104,105,109,110] [101] [100,101,103,105] [104] | Inhibitory: [103,104,105,109,110,152] Supportive: [146,147] |
| SINE1 (Alu) | Alu-like 7SL RNA (MS2 fusion) Transgene with inverted Alu repeat | Co-expressed with ORF1p eIF3η | [101] [112] | ||
| ERV | HERVK(HML-2) Reverse transcriptase | TIA-1 G3BP1 | [120] [121] | ||
| P-body | Ty | Ty1 GAG Ty3 POL3/GAG Ty1 mRNA | Dhh1, Dcp2 Xrn1, Dhh1, Pat1, Dcp2, Ded1, Lsm1 | [146,147] [145,148] [146] | |
| IAP | IAP gag mRNA | RCK | [152,153] | ||
| Het-A I-element | Het-A and I-element (MS2 fusion) | PCM | [158] | ||
| Alu | Transgene containing inverted Alu repeat at 3′ UTR | GW182 | [112] |
4. Concluding Remarks and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lander, E.S.; Linton, L.M.; Birren, B.; Nusbaum, C.; Zody, M.C.; Baldwin, J.; Devon, K.; Dewar, K.; Doyle, M.; FitzHugh, W.; et al. Initial sequencing and analysis of the human genome. Nature 2001, 409, 860–921. [Google Scholar] [CrossRef] [PubMed]
- Wicker, T.; Sabot, F.; Hua-Van, A.; Bennetzen, J.L.; Capy, P.; Chalhoub, B.; Flavell, A.; Leroy, P.; Morgante, M.; Panaud, O.; et al. A unified classification system for eukaryotic transposable elements. Nat. Rev. Genet. 2007, 8, 973–982. [Google Scholar] [CrossRef] [PubMed]
- Boeke, J.D. LINEs and Alus—The polyA connection. Nat. Genet. 1997, 16, 6–7. [Google Scholar] [CrossRef]
- Ostertag, E.M.; Goodier, J.L.; Zhang, Y.; Kazazian, H.H., Jr. SVA Elements Are Nonautonomous Retrotransposons that Cause Disease in Humans. Am. J. Hum. Genet. 2003, 73, 1444–1451. [Google Scholar] [CrossRef] [PubMed]
- Wells, J.N.; Feschotte, C. A Field Guide to Eukaryotic Transposable Elements. Annu. Rev. Genet. 2020, 54, 539–561. [Google Scholar] [CrossRef]
- Montgomery, E.A.; Huang, S.M.; Langley, C.H.; Judd, B.H. Chromosome rearrangement by ectopic recombination in Drosophila melanogaster: Genome structure and evolution. Genetics 1991, 129, 1085–1098. [Google Scholar] [CrossRef]
- Ade, C.; Roy-Engel, A.M.; Deininger, P.L. Alu elements: An intrinsic source of human genome instability. Curr. Opin. Virol. 2013, 3, 639–645. [Google Scholar] [CrossRef]
- Sundaram, V.; Wysocka, J. Transposable elements as a potent source of diverse cis-regulatory sequences in mammalian genomes. Philos. Trans. R. Soc. B Biol. Sci. 2020, 375, 20190347. [Google Scholar] [CrossRef]
- Moschetti, R.; Palazzo, A.; Lorusso, P.; Viggiano, L.; Marsano, R.M. “What You Need, Baby, I Got it”: Transposable Elements as Suppliers of Cis-Operating Sequences in Drosophila. Biology 2020, 9, 25. [Google Scholar] [CrossRef]
- Chuong, E.B.; Elde, N.C.; Feschotte, C. Regulatory activities of transposable elements: From conflicts to benefits. Nat. Rev. Genet. 2016, 18, 71–86. [Google Scholar] [CrossRef]
- Garcia-Perez, J.L.; Widmann, T.J.; Adams, I.R. The impact of transposable elements on mammalian development. Development 2016, 143, 4101–4114. [Google Scholar] [CrossRef] [PubMed]
- Hancks, D.C.; Kazazian, H.H., Jr. Roles for retrotransposon insertions in human disease. Mob. DNA 2016, 7, 9. [Google Scholar] [CrossRef]
- Nagirnaja, L.; Mørup, N.; Nielsen, J.E.; Stakaitis, R.; Golubickaite, I.; Oud, M.S.; Winge, S.B.; Carvalho, F.; Aston, K.I.; Khani, F.; et al. Variant PNLDC1, Defective piRNA Processing, and Azoospermia. N. Engl. J. Med. 2021, 385, 707–719. [Google Scholar] [CrossRef] [PubMed]
- Aravin, A.A.; Naumova, N.M.; Tulin, A.V.; Vagin, V.V.; Rozovsky, Y.M.; Gvozdev, V.A. Double-stranded RNA-mediated silencing of genomic tandem repeats and transposable elements in the D. melanogaster germline. Curr. Biol. 2001, 11, 1017–1027. [Google Scholar] [CrossRef] [PubMed]
- Aravin, A.; Gaidatzis, D.; Pfeffer, S.; Lagos-Quintana, M.; Landgraf, P.; Iovino, N.; Morris, P.; Brownstein, M.J.; Kuramochi-Miyagawa, S.; Nakano, T.; et al. A novel class of small RNAs bind to MILI protein in mouse testes. Nature 2006, 442, 203–207. [Google Scholar] [CrossRef]
- Lau, N.C.; Seto, A.G.; Kim, J.; Kuramochi-Miyagawa, S.; Nakano, T.; Bartel, D.P.; Kingston, R.E. Characterization of the piRNA Complex from Rat Testes. Science 2006, 313, 363–367. [Google Scholar] [CrossRef]
- Houwing, S.; Kamminga, L.M.; Berezikov, E.; Cronembold, D.; Girard, A.; van den Elst, H.; Filippov, D.V.; Blaser, H.; Raz, E.; Moens, C.B.; et al. A Role for Piwi and piRNAs in Germ Cell Maintenance and Transposon Silencing in Zebrafish. Cell 2007, 129, 69–82. [Google Scholar] [CrossRef]
- Gainetdinov, I.; Colpan, C.; Arif, A.; Cecchini, K.; Zamore, P.D. A Single Mechanism of Biogenesis, Initiated and Directed by PIWI Proteins, Explains piRNA Production in Most Animals. Mol. Cell 2018, 71, 775–790.e775. [Google Scholar] [CrossRef]
- Girard, A.; Sachidanandam, R.; Hannon, G.J.; Carmell, M.A. A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 2006, 442, 199–202. [Google Scholar] [CrossRef]
- Brennecke, J.; Aravin, A.A.; Stark, A.; Dus, M.; Kellis, M.; Sachidanandam, R.; Hannon, G.J. Discrete Small RNA-Generating Loci as Master Regulators of Transposon Activity in Drosophila. Cell 2007, 128, 1089–1103. [Google Scholar] [CrossRef]
- Ozata, D.M.; Gainetdinov, I.; Zoch, A.; O’Carroll, D.; Zamore, P.D. PIWI-interacting RNAs: Small RNAs with big functions. Nat. Rev. Genet. 2019, 20, 89–108. [Google Scholar] [CrossRef] [PubMed]
- Ecco, G.; Cassano, M.; Kauzlaric, A.; Duc, J.; Coluccio, A.; Offner, S.; Imbeault, M.; Rowe, H.M.; Turelli, P.; Trono, D. Transposable Elements and Their KRAB-ZFP Controllers Regulate Gene Expression in Adult Tissues. Dev. Cell 2016, 36, 611–623. [Google Scholar] [CrossRef] [PubMed]
- Playfoot, C.J.; Adams, I.R. KRABs RegulaTE Gene Expression beyond the Embryo. Dev. Cell 2016, 36, 591–592. [Google Scholar] [CrossRef] [PubMed]
- Ecco, G.; Imbeault, M.; Trono, D. KRAB zinc finger proteins. Development 2017, 144, 2719–2729. [Google Scholar] [CrossRef]
- Schultz, D.C.; Friedman, J.R.; Rauscher, F.J., 3rd. Targeting histone deacetylase complexes via KRAB-zinc finger proteins: The PHD and bromodomains of KAP-1 form a cooperative unit that recruits a novel isoform of the Mi-2α subunit of NuRD. Genes Dev. 2001, 15, 428–443. [Google Scholar] [CrossRef] [PubMed]
- Schultz, D.C.; Ayyanathan, K.; Negorev, D.; Maul, G.G.; Rauscher, F.J., 3rd. SETDB1: A novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev. 2002, 16, 919–932. [Google Scholar] [CrossRef]
- Sripathy, S.P.; Stevens, J.; Schultz, D.C. The KAP1 Corepressor Functions to Coordinate the Assembly of De Novo HP1-Demarcated Microenvironments of Heterochromatin Required for KRAB Zinc Finger Protein-Mediated Transcriptional Repression. Mol. Cell. Biol. 2006, 26, 8623–8638. [Google Scholar] [CrossRef]
- Quenneville, S.; Turelli, P.; Bojkowska, K.; Raclot, C.; Offner, S.; Kapopoulou, A.; Trono, D. The KRAB-ZFP/KAP1 System Contributes to the Early Embryonic Establishment of Site-Specific DNA Methylation Patterns Maintained during Development. Cell Rep. 2012, 2, 766–773. [Google Scholar] [CrossRef]
- Wiznerowicz, M.; Jakobsson, J.; Szulc, J.; Liao, S.; Quazzola, A.; Beermann, F.; Aebischer, P.; Trono, D. The Krüppel-associated Box Repressor Domain Can Trigger de Novo Promoter Methylation during Mouse Early Embryogenesis. J. Biol. Chem. 2007, 282, 34535–34541. [Google Scholar] [CrossRef]
- Quenneville, S.; Verde, G.; Corsinotti, A.; Kapopoulou, A.; Jakobsson, J.; Offner, S.; Baglivo, I.; Pedone, P.V.; Grimaldi, G.; Riccio, A.; et al. In Embryonic Stem Cells, ZFP57/KAP1 Recognize a Methylated Hexanucleotide to Affect Chromatin and DNA Methylation of Imprinting Control Regions. Mol. Cell 2011, 44, 361–372. [Google Scholar] [CrossRef]
- Wolf, D.; Goff, S.P. TRIM28 Mediates Primer Binding Site-Targeted Silencing of Murine Leukemia Virus in Embryonic Cells. Cell 2007, 131, 46–57. [Google Scholar] [CrossRef] [PubMed]
- Wolf, D.; Goff, S.P. Embryonic stem cells use ZFP809 to silence retroviral DNAs. Nature 2009, 458, 1201–1204. [Google Scholar] [CrossRef]
- Turelli, P.; Castro-Diaz, N.; Marzetta, F.; Kapopoulou, A.; Raclot, C.; Duc, J.; Tieng, V.; Quenneville, S.; Trono, D. Interplay of TRIM28 and DNA methylation in controlling human endogenous retroelements. Genome Res. 2014, 24, 1260–1270. [Google Scholar] [CrossRef] [PubMed]
- Najafabadi, H.S.; Mnaimneh, S.; Schmitges, F.W.; Garton, M.; Lam, K.N.; Yang, A.; Albu, M.; Weirauch, M.T.; Radovani, E.; Kim, P.M.; et al. C2H2 zinc finger proteins greatly expand the human regulatory lexicon. Nat. Biotechnol. 2015, 33, 555–562. [Google Scholar] [CrossRef] [PubMed]
- Imbeault, M.; Helleboid, P.-Y.; Trono, D. KRAB zinc-finger proteins contribute to the evolution of gene regulatory networks. Nature 2017, 543, 550–554. [Google Scholar] [CrossRef] [PubMed]
- Wolf, G.; de Iaco, A.; Sun, M.-A.; Bruno, M.; Tinkham, M.; Hoang, D.; Mitra, A.; Ralls, S.; Trono, D.; Macfarlan, T.S. KRAB-zinc finger protein gene expansion in response to active retrotransposons in the murine lineage. Elife 2020, 9, e56337. [Google Scholar] [CrossRef]
- Rowe, H.M.; Jakobsson, J.; Mesnard, D.; Rougemont, J.; Reynard, S.; Aktas, T.; Maillard, P.V.; Layard-Liesching, H.; Verp, S.; Marquis, J.; et al. KAP1 controls endogenous retroviruses in embryonic stem cells. Nature 2010, 463, 237–240. [Google Scholar] [CrossRef]
- Matsui, T.; Leung, D.; Miyashita, H.; Maksakova, I.A.; Miyachi, H.; Kimura, H.; Tachibana, M.; Lorincz, M.C.; Shinkai, Y. Proviral silencing in embryonic stem cells requires the histone methyltransferase ESET. Nature 2010, 464, 927–931. [Google Scholar] [CrossRef]
- Buchan, J.R. mRNP granules. Assembly, function, and connections with disease. RNA Biol. 2014, 11, 1019–1030. [Google Scholar] [CrossRef]
- Kedersha, N.; Anderson, P. Mammalian Stress Granules and Processing Bodies. Methods Enzymol. 2007, 431, 61–81. [Google Scholar] [CrossRef]
- Mahboubi, H.; Stochaj, U. Cytoplasmic stress granules: Dynamic modulators of cell signaling and disease. Biochim. Biophys. Acta Mol. Basis Dis. 2017, 1863, 884–895. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.; Jin, X.; Liu, B. The involvement of stress granules in aging and aging-associated diseases. Aging Cell 2020, 19, e13136. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.I.; Namkoong, S. Stress granules dynamics: Benefits in cancer. BMB Rep. 2022, 55, 577–586. [Google Scholar] [CrossRef] [PubMed]
- Anderson, P.; Kedersha, N. Stress granules. Curr. Biol. 2009, 19, R397–R398. [Google Scholar] [CrossRef]
- Ivanov, P.; Kedersha, N.; Anderson, P. Stress Granules and Processing Bodies in Translational Control. Cold Spring Harb. Perspect. Biol. 2019, 11, a032813. [Google Scholar] [CrossRef]
- Marcelo, A.; Koppenol, R.; de Almeida, L.P.; Matos, C.A.; Nóbrega, C. Stress granules, RNA-binding proteins and polyglutamine diseases: Too much aggregation? Cell Death Dis. 2021, 12, 592. [Google Scholar] [CrossRef]
- Kedersha, N.L.; Gupta, M.; Li, W.; Miller, I.; Anderson, P. RNA-Binding Proteins Tia-1 and Tiar Link the Phosphorylation of Eif-2α to the Assembly of Mammalian Stress Granules. J. Cell Biol. 1999, 147, 1431–1442. [Google Scholar] [CrossRef]
- Mazroui, R.; Sukarieh, R.; Bordeleau, M.-E.; Kaufman, R.J.; Northcote, P.; Tanaka, J.; Gallouzi, I.-E.; Pelletier, J.; Ravel-Chapuis, A.; Gunnewiek, A.K.; et al. Inhibition of Ribosome Recruitment Induces Stress Granule Formation Independently of Eukaryotic Initiation Factor 2α Phosphorylation. Mol. Biol. Cell 2006, 17, 4212–4219. [Google Scholar] [CrossRef]
- Mateju, D.; Eichenberger, B.; Voigt, F.; Eglinger, J.; Roth, G.; Chao, J.A. Single-Molecule Imaging Reveals Translation of mRNAs Localized to Stress Granules. Cell 2020, 183, P1801–P1812.e13. [Google Scholar] [CrossRef]
- Namkoong, S.; Ho, A.; Woo, Y.M.; Kwak, H.; Lee, J.H. Systematic Characterization of Stress-Induced RNA Granulation. Mol. Cell 2018, 70, 175–187.e8. [Google Scholar] [CrossRef]
- Vanderweyde, T.; Youmans, K.; Liu-Yesucevitz, L.; Wolozin, B. Role of Stress Granules and RNA-Binding Proteins in Neurodegeneration: A Mini-Review. Gerontology 2013, 59, 524–533. [Google Scholar] [CrossRef] [PubMed]
- Alberti, S. Phase separation in biology. Curr. Biol. 2017, 27, R1097–R1102. [Google Scholar] [CrossRef]
- Wheeler, J.R.; Matheny, T.; Jain, S.; Abrisch, R.; Parker, R. Distinct stages in stress granule assembly and disassembly. Elife 2016, 5, e18413. [Google Scholar] [CrossRef] [PubMed]
- Protter, D.S.W.; Parker, R. Principles and Properties of Stress Granules. Trends Cell Biol. 2016, 26, 668–679. [Google Scholar] [CrossRef] [PubMed]
- Reineke, L.C.; Tsai, W.-C.; Jain, A.; Kaelber, J.T.; Jung, S.Y.; Lloyd, R.E. Casein Kinase 2 Is Linked to Stress Granule Dynamics through Phosphorylation of the Stress Granule Nucleating Protein G3BP1. Mol. Cell. Biol. 2017, 37, e00596-16. [Google Scholar] [CrossRef]
- Tanikawa, C.; Ueda, K.; Suzuki, A.; Iida, A.; Nakamura, R.; Atsuta, N.; Tohnai, G.; Sobue, G.; Saichi, N.; Momozawa, Y.; et al. Citrullination of RGG Motifs in FET Proteins by PAD4 Regulates Protein Aggregation and ALS Susceptibility. Cell Rep. 2018, 22, 1473–1483. [Google Scholar] [CrossRef]
- Gwon, Y.; Maxwell, B.A.; Kolaitis, R.-M.; Zhang, P.; Kim, H.J.; Taylor, J.P. Ubiquitination of G3BP1 mediates stress granule disassembly in a context-specific manner. Science 2021, 372, eabf6548. [Google Scholar] [CrossRef]
- Marmor-Kollet, H.; Siany, A.; Kedersha, N.; Knafo, N.; Rivkin, N.; Danino, Y.M.; Moens, T.G.; Olender, T.; Sheban, D.; Cohen, N.; et al. Spatiotemporal Proteomic Analysis of Stress Granule Disassembly Using APEX Reveals Regulation by SUMOylation and Links to ALS Pathogenesis. Mol. Cell 2020, 80, 876–891.e876. [Google Scholar] [CrossRef]
- Ries, R.J.; Zaccara, S.; Klein, P.; Olarerin-George, A.; Namkoong, S.; Pickering, B.F.; Patil, D.P.; Kwak, H.; Lee, J.H.; Jaffrey, S.R. m(6)A enhances the phase separation potential of mRNA. Nature 2019, 571, 424–428. [Google Scholar] [CrossRef]
- Fu, Y.; Zhuang, X. m6A-binding YTHDF proteins promote stress granule formation. Nat. Chem. Biol. 2020, 16, 955–963. [Google Scholar] [CrossRef]
- Deigendesch, N.; Koch-Nolte, F.; Rothenburg, S. ZBP1 subcellular localization and association with stress granules is controlled by its Z-DNA binding domains. Nucleic Acids Res. 2006, 34, 5007–5020. [Google Scholar] [CrossRef]
- Park, C.; Choi, S.; Kim, Y.-E.; Lee, S.; Park, S.-H.; Adelstein, R.S.; Kawamoto, S.; Kim, K.K. Stress Granules Contain Rbfox2 with Cell Cycle-related mRNAs. Sci. Rep. 2017, 7, 1211. [Google Scholar] [CrossRef]
- Wilbertz, J.H.; Voigt, F.; Horvathova, I.; Roth, G.; Zhan, Y.; Chao, J.A. Single-Molecule Imaging of mRNA Localization and Regulation during the Integrated Stress Response. Mol. Cell 2019, 73, 946–958.e947. [Google Scholar] [CrossRef]
- Arimoto, K.; Fukuda, H.; Imajoh-Ohmi, S.; Saito, H.; Takekawa, M. Formation of stress granules inhibits apoptosis by suppressing stress-responsive MAPK pathways. Nat. Cell Biol. 2008, 10, 1324–1332. [Google Scholar] [CrossRef] [PubMed]
- Thedieck, K.; Holzwarth, B.; Prentzell, M.T.; Boehlke, C.; Kläsener, K.; Ruf, S.; Sonntag, A.G.; Maerz, L.; Grellscheid, S.-N.; Kremmer, E.; et al. Inhibition of mTORC1 by Astrin and Stress Granules Prevents Apoptosis in Cancer Cells. Cell 2013, 154, 859–874. [Google Scholar] [CrossRef] [PubMed]
- Wippich, F.; Bodenmiller, B.; Trajkovska, M.G.; Wanka, S.; Aebersold, R.; Pelkmans, L. Dual Specificity Kinase DYRK3 Couples Stress Granule Condensation/Dissolution to mTORC1 Signaling. Cell 2013, 152, 791–805. [Google Scholar] [CrossRef]
- Mediani, L.; Antoniani, F.; Galli, V.; Vinet, J.; Carrà, A.D.; Bigi, I.; Tripathy, V.; Tiago, T.; Cimino, M.; Leo, G.; et al. Hsp90-mediated regulation of DYRK3 couples stress granule disassembly and growth via mTORC1 signaling. EMBO Rep. 2021, 22, e51740. [Google Scholar] [CrossRef] [PubMed]
- Omer, A.; Patel, D.; Lian, X.J.; Sadek, J.; Di Marco, S.; Pause, A.; Gorospe, M.; Gallouzi, I.E. Stress granules counteract senescence by sequestration of PAI-1. EMBO Rep. 2018, 19, e44722. [Google Scholar] [CrossRef] [PubMed]
- Miller, C.L. Stress granules and virus replication. Future Virol. 2011, 6, 1329–1338. [Google Scholar] [CrossRef]
- Wu, Y.; Zhang, Z.; Li, Y.; Li, Y. The Regulation of Integrated Stress Response Signaling Pathway on Viral Infection and Viral Antagonism. Front. Microbiol. 2021, 12, 814635. [Google Scholar] [CrossRef]
- Mateju, D.; Chao, J.A. Stress granules: Regulators or by-products? FEBS J. 2022, 289, 363–373. [Google Scholar] [CrossRef] [PubMed]
- McCormick, C.; Khaperskyy, D.A. Translation inhibition and stress granules in the antiviral immune response. Nat. Rev. Immunol. 2017, 17, 647–660. [Google Scholar] [CrossRef] [PubMed]
- Reineke, L.C.; Lloyd, R.E. The Stress Granule Protein G3BP1 Recruits Protein Kinase R to Promote Multiple Innate Immune Antiviral Responses. J. Virol. 2015, 89, 2575–2589. [Google Scholar] [CrossRef] [PubMed]
- Ivashkiv, L.B.; Donlin, L.T. Regulation of type I interferon responses. Nat. Rev. Immunol. 2014, 14, 36–49. [Google Scholar] [CrossRef] [PubMed]
- Onomoto, K.; Jogi, M.; Yoo, J.-S.; Narita, R.; Morimoto, S.; Takemura, A.; Sambhara, S.; Kawaguchi, A.; Osari, S.; Nagata, K.; et al. Critical Role of an Antiviral Stress Granule Containing RIG-I and PKR in Viral Detection and Innate Immunity. PLoS ONE 2012, 7, e43031. [Google Scholar] [CrossRef]
- Langereis, M.A.; Feng, Q.; van Kuppeveld, F.J. MDA5 Localizes to Stress Granules, but This Localization Is Not Required for the Induction of Type I Interferon. J. Virol. 2013, 87, 6314–6325. [Google Scholar] [CrossRef]
- Yi, Z.; Pan, T.; Wu, X.; Song, W.; Wang, S.; Xu, Y.; Rice, C.M.; MacDonald, M.R.; Yuan, Z. Hepatitis C Virus Co-Opts Ras-GTPase-Activating Protein-Binding Protein 1 for Its Genome Replication. J. Virol. 2011, 85, 6996–7004. [Google Scholar] [CrossRef]
- Garaigorta, U.; Heim, M.H.; Boyd, B.; Wieland, S.; Chisari, F.V. Hepatitis C Virus (HCV) Induces Formation of Stress Granules Whose Proteins Regulate HCV RNA Replication and Virus Assembly and Egress. J. Virol. 2012, 86, 11043–11056. [Google Scholar] [CrossRef]
- Zheng, Z.-Q.; Wang, S.-Y.; Xu, Z.-S.; Fu, Y.-Z.; Wang, Y.-Y. SARS-CoV-2 nucleocapsid protein impairs stress granule formation to promote viral replication. Cell Discov. 2021, 7, 38. [Google Scholar] [CrossRef]
- Liu, H.; Bai, Y.; Zhang, X.; Gao, T.; Liu, Y.; Li, E.; Wang, X.; Cao, Z.; Zhu, L.; Dong, Q.; et al. SARS-CoV-2 N Protein Antagonizes Stress Granule Assembly and IFN Production by Interacting with G3BPs to Facilitate Viral Replication. J. Virol. 2022, 96, e0041222. [Google Scholar] [CrossRef]
- Loll-Krippleber, R.; Brown, G.W. P-body proteins regulate transcriptional rewiring to promote DNA replication stress resistance. Nat. Commun. 2017, 8, 558. [Google Scholar] [CrossRef] [PubMed]
- Banani, S.F.; Lee, H.O.; Hyman, A.A.; Rosen, M.K. Biomolecular condensates: Organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 2017, 18, 285–298. [Google Scholar] [CrossRef]
- Hubstenberger, A.; Courel, M.; Bénard, M.; Souquere, S.; Ernoult-Lange, M.; Chouaib, R.; Yi, Z.; Morlot, J.-B.; Munier, A.; Fradet, M.; et al. P-Body Purification Reveals the Condensation of Repressed mRNA Regulons. Mol. Cell 2017, 68, 144–157.e5. [Google Scholar] [CrossRef]
- Pitchiaya, S.; Mourao, M.D.A.; Jalihal, A.P.; Xiao, L.; Jiang, X.; Chinnaiyan, A.M.; Schnell, S.; Walter, N.G. Dynamic Recruitment of Single RNAs to Processing Bodies Depends on RNA Functionality. Mol. Cell 2019, 74, 521–533.e526. [Google Scholar] [CrossRef] [PubMed]
- Chalabi Hagkarim, N.; Grand, R.J. The Regulatory Properties of the Ccr4–Not Complex. Cells 2020, 9, 2379. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Na, Z.; Slavoff, S.A. P-Bodies: Composition, Properties, and Functions. Biochemistry 2018, 57, 2424–2431. [Google Scholar] [CrossRef]
- Chang, C.-T.; Bercovich, N.; Loh, B.; Jonas, S.; Izaurralde, E. The activation of the decapping enzyme DCP2 by DCP1 occurs on the EDC4 scaffold and involves a conserved loop in DCP1. Nucleic Acids Res. 2014, 42, 5217–5233. [Google Scholar] [CrossRef]
- Decker, C.J.; Parker, R. P-Bodies and Stress Granules: Possible Roles in the Control of Translation and mRNA Degradation. Cold Spring Harb. Perspect. Biol. 2012, 4, a012286. [Google Scholar] [CrossRef]
- Kuzuoğlu-Öztürk, D.; Bhandari, D.; Huntzinger, E.; Fauser, M.; Helms, S.; Izaurralde, E. miRISC and the CCR4-NOT complex silence mRNA targets independently of 43S ribosomal scanning. EMBO J. 2016, 35, 1186–1203. [Google Scholar] [CrossRef]
- Chang, W.-L.; Tarn, W.-Y. A role for transportin in deposition of TTP to cytoplasmic RNA granules and mRNA decay. Nucleic Acids Res. 2009, 37, 6600–6612. [Google Scholar] [CrossRef]
- Wang, C.; Schmich, F.; Srivatsa, S.; Weidner, J.; Beerenwinkel, N.; Spang, A. Context-dependent deposition and regulation of mRNAs in P-bodies. Elife 2018, 7, e29815. [Google Scholar] [CrossRef] [PubMed]
- Xing, W.; Muhlrad, D.; Parker, R.; Rosen, M.K. A quantitative inventory of yeast P body proteins reveals principles of composition and specificity. Elife 2020, 9, e56525. [Google Scholar] [CrossRef] [PubMed]
- Eulalio, A.; Behm-Ansmant, I.; Schweizer, D.; Izaurralde, E. P-Body Formation Is a Consequence, Not the Cause, of RNA-Mediated Gene Silencing. Mol. Cell. Biol. 2007, 27, 3970–3981. [Google Scholar] [CrossRef]
- Standart, N.; Weil, D. P-Bodies: Cytosolic Droplets for Coordinated mRNA Storage. Trends Genet. 2018, 34, 612–626. [Google Scholar] [CrossRef] [PubMed]
- Tkach, J.M.; Yimit, A.; Lee, A.Y.; Riffle, M.; Costanzo, M.; Jaschob, D.; Hendry, J.A.; Ou, J.; Moffat, J.; Boone, C.; et al. Dissecting DNA damage response pathways by analysing protein localization and abundance changes during DNA replication stress. Nat. Cell Biol. 2012, 14, 966–976. [Google Scholar] [CrossRef]
- Beck, C.R.; Garcia-Perez, J.L.; Badge, R.M.; Moran, J.V. LINE-1 Elements in Structural Variation and Disease. Annu. Rev. Genom. Hum. Genet. 2011, 12, 187–215. [Google Scholar] [CrossRef]
- Babushok, D.V.; Kazazian, H.H., Jr. Progress in understanding the biology of the human mutagen LINE-1. Hum. Mutat. 2007, 28, 527–539. [Google Scholar] [CrossRef] [PubMed]
- Martin, S.L.; Branciforte, D. Synchronous expression of LINE-1 RNA and protein in mouse embryonal carcinoma cells. Mol. Cell. Biol. 1993, 13, 5383–5392. [Google Scholar] [CrossRef]
- Goodier, J.L.; Zhang, L.; Vetter, M.R.; Kazazian, H.H., Jr. LINE-1 ORF1 Protein Localizes in Stress Granules with other RNA-Binding Proteins, Including Components of RNA Interference RNA-Induced Silencing Complex. Mol. Cell. Biol. 2007, 27, 6469–6483. [Google Scholar] [CrossRef]
- Doucet, A.J.; Hulme, A.E.; Sahinovic, E.; Kulpa, D.A.; Moldovan, J.B.; Kopera, H.C.; Athanikar, J.N.; Hasnaoui, M.; Bucheton, A.; Moran, J.V.; et al. Characterization of LINE-1 Ribonucleoprotein Particles. PLOS Genet. 2010, 6, e1001150. [Google Scholar] [CrossRef]
- Goodier, J.L.; Mandal, P.K.; Zhang, L.; Kazazian, H.H., Jr. Discrete subcellular partitioning of human retrotransposon RNAs despite a common mechanism of genome insertion. Hum. Mol. Genet. 2010, 19, 1712–1725. [Google Scholar] [CrossRef]
- Gao, G.; Guo, X.; Goff, S.P. Inhibition of Retroviral RNA Production by ZAP, a CCCH-Type Zinc Finger Protein. Science 2002, 297, 1703–1706. [Google Scholar] [CrossRef]
- Goodier, J.L.; Pereira, G.C.; Cheung, L.E.; Rose, R.J.; Kazazian, H.H., Jr. The Broad-Spectrum Antiviral Protein ZAP Restricts Human Retrotransposition. PLoS Genet. 2015, 11, e1005252. [Google Scholar] [CrossRef] [PubMed]
- Moldovan, J.B.; Moran, J.V. The Zinc-Finger Antiviral Protein ZAP Inhibits LINE and Alu Retrotransposition. PLOS Genet. 2015, 11, e1005121. [Google Scholar] [CrossRef]
- Huang, Y.; Xu, F.; Mei, S.; Liu, X.; Zhao, F.; Wei, L.; Fan, Z.; Hu, Y.; Wang, L.; Ai, B.; et al. MxB inhibits long interspersed element type 1 retrotransposition. PLOS Genet. 2022, 18, e1010034. [Google Scholar] [CrossRef]
- Liu, Z.; Pan, Q.; Ding, S.; Qian, J.; Xu, F.; Zhou, J.; Cen, S.; Guo, F.; Liang, C. The Interferon-Inducible MxB Protein Inhibits HIV-1 Infection. Cell Host Microbe 2013, 14, 398–410. [Google Scholar] [CrossRef] [PubMed]
- Rice, G.I.; Bond, J.; Asipu, A.; Brunette, R.L.; Manfield, I.W.; Carr, I.M.; Fuller, J.C.; Jackson, R.M.; Lamb, T.; Briggs, T.A.; et al. Mutations involved in Aicardi-Goutières syndrome implicate SAMHD1 as regulator of the innate immune response. Nat. Genet. 2009, 41, 829–832. [Google Scholar] [CrossRef]
- Zhao, K.; Du, J.; Han, X.; Goodier, J.L.; Li, P.; Zhou, X.; Wei, W.; Evans, S.L.; Li, L.; Zhang, W.; et al. Modulation of LINE-1 and Alu/SVA Retrotransposition by Aicardi-Goutières Syndrome-Related SAMHD1. Cell Rep. 2013, 4, 1108–1115. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; Li, J.; Xu, F.; Mei, S.; Le Duff, Y.; Yin, L.; Pang, X.; Cen, S.; Jin, Q.; Liang, C.; et al. SAMHD1 Inhibits LINE-1 Retrotransposition by Promoting Stress Granule Formation. PLOS Genet. 2015, 11, e1005367. [Google Scholar] [CrossRef]
- Schöbel, A.; Nguyen-Dinh, V.; Schumann, G.G.; Herker, E. Hepatitis C virus infection restricts human LINE-1 retrotransposition in hepatoma cells. PLOS Pathog. 2021, 17, e1009496. [Google Scholar] [CrossRef]
- Deininger, P.L.; Batzer, M.A. Alu Repeats and Human Disease. Mol. Genet. Metab. 1999, 67, 183–193. [Google Scholar] [CrossRef] [PubMed]
- Fitzpatrick, T.; Huang, S. 3’-UTR-located inverted Alu repeats facilitate mRNA translational repression and stress granule accumulation. Nucleus 2012, 3, 359–369. [Google Scholar] [CrossRef] [PubMed]
- Johnson, W.E. Origins and evolutionary consequences of ancient endogenous retroviruses. Nat. Rev. Microbiol. 2019, 17, 355–370. [Google Scholar] [CrossRef] [PubMed]
- Grow, E.J.; Flynn, R.A.; Chavez, S.L.; Bayless, N.L.; Wossidlo, M.; Wesche, D.J.; Martin, L.; Ware, C.B.; Blish, C.A.; Chang, H.Y.; et al. Intrinsic retroviral reactivation in human preimplantation embryos and pluripotent cells. Nature 2015, 522, 221–225. [Google Scholar] [CrossRef] [PubMed]
- Maze, E.A.; Agit, B.; Reeves, S.; Hilton, D.A.; Parkinson, D.B.; Laraba, L.; Ercolano, E.; Kurian, K.M.; Hanemann, C.O.; Belshaw, R.D.; et al. Human Endogenous Retrovirus Type K Promotes Proliferation and Confers Sensitivity to Antiretroviral Drugs in Merlin-Negative Schwannoma and Meningioma. Cancer Res. 2022, 82, 235–247. [Google Scholar] [CrossRef] [PubMed]
- Macfarlan, T.S.; Gifford, W.D.; Driscoll, S.; Lettieri, K.; Rowe, H.M.; Bonanomi, D.; Firth, A.; Singer, O.; Trono, D.; Pfaff, S.L. Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 2012, 487, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Göke, J.; Lu, X.; Chan, Y.-S.; Ng, H.-H.; Ly, L.-H.; Sachs, F.; Szczerbinska, I. Dynamic Transcription of Distinct Classes of Endogenous Retroviral Elements Marks Specific Populations of Early Human Embryonic Cells. Cell Stem Cell 2015, 16, 135–141. [Google Scholar] [CrossRef]
- Schupbach, T.; Wieschaus, E. Germline autonomy of maternal-effect mutations altering the embryonic body pattern of Drosophila. Dev. Biol. 1986, 113, 443–448. [Google Scholar] [CrossRef]
- Thomas, M.G.; Martinez Tosar, L.J.; Desbats, M.A.; Leishman, C.C.; Boccaccio, G.L. Mammalian Staufen 1 is recruited to stress granules and impairs their assembly. J. Cell Sci. 2009, 122, 563–573. [Google Scholar] [CrossRef]
- Hanke, K.; Hohn, O.; Liedgens, L.; Fiddeke, K.; Wamara, J.; Kurth, R.; Bannert, N. Staufen-1 Interacts with the Human Endogenous Retrovirus Family HERV-K(HML-2) Rec and Gag Proteins and Increases Virion Production. J. Virol. 2013, 87, 11019–11030. [Google Scholar] [CrossRef]
- Manghera, M.; Ferguson-Parry, J.; Douville, R.N. TDP-43 regulates endogenous retrovirus-K viral protein accumulation. Neurobiol. Dis. 2016, 94, 226–236. [Google Scholar] [CrossRef]
- Kinomoto, M.; Kanno, T.; Shimura, M.; Ishizaka, Y.; Kojima, A.; Kurata, T.; Sata, T.; Tokunaga, K. All APOBEC3 family proteins differentially inhibit LINE-1 retrotransposition. Nucleic Acids Res. 2007, 35, 2955–2964. [Google Scholar] [CrossRef] [PubMed]
- Esnault, C.; Heidmann, O.; Delebecque, F.; Dewannieux, M.; Ribet, D.; Hance, A.J.; Heidmann, T.; Schwartz, O. APO-BEC3G cytidine deaminase inhibits retrotransposition of endogenous retroviruses. Nature 2005, 433, 430–433. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Lilley, C.E.; Yu, Q.; Lee, D.V.; Chou, J.; Narvaiza, I.; Landau, N.R.; Weitzman, M.D. APOBEC3A Is a Potent Inhibitor of Adeno-Associated Virus and Retrotransposons. Curr. Biol. 2006, 16, 480–485. [Google Scholar] [CrossRef]
- Bogerd, H.P.; Wiegand, H.L.; Doehle, B.P.; Lueders, K.K.; Cullen, B.R. APOBEC3A and APOBEC3B are potent inhibitors of LTR-retrotransposon function in human cells. Nucleic Acids Res. 2006, 34, 89–95. [Google Scholar] [CrossRef] [PubMed]
- Bogerd, H.P.; Wiegand, H.L.; Hulme, A.E.; Garcia-Perez, J.L.; O’Shea, K.S.; Moran, J.V.; Cullen, B.R. Cellular inhibitors of long interspersed element 1 and Alu retrotransposition. Proc. Natl. Acad. Sci. USA 2006, 103, 8780–8785. [Google Scholar] [CrossRef]
- Gallois-Montbrun, S.; Kramer, B.; Swanson, C.M.; Byers, H.; Lynham, S.; Ward, M.; Malim, M.H. Antiviral Protein APOBEC3G Localizes to Ribonucleoprotein Complexes Found in P Bodies and Stress Granules. J. Virol. 2007, 81, 2165–2178. [Google Scholar] [CrossRef] [PubMed]
- Kozak, S.L.; Marin, M.; Rose, K.M.; Bystrom, C.; Kabat, D. The Anti-HIV-1 Editing Enzyme APOBEC3G Binds HIV-1 RNA and Messenger RNAs That Shuttle between Polysomes and Stress Granules. J. Biol. Chem. 2006, 281, 29105–29119. [Google Scholar] [CrossRef]
- Horn, A.V.; Klawitter, S.; Held, U.; Berger, A.; Vasudevan, A.A.; Bock, A.; Hofmann, H.; Hanschmann, K.M.; Trösemeier, J.-H.; Flory, E.; et al. Human LINE-1 restriction by APOBEC3C is deaminase independent and mediated by an ORF1p interaction that affects LINE reverse transcriptase activity. Nucleic Acids Res. 2013, 42, 396–416. [Google Scholar] [CrossRef]
- Manjunath, L.; Oh, S.; Ortega, P.; Bouin, A.; Bournique, E.; Sanchez, A.; Martensen, P.M.; Auerbach, A.A.; Becker, J.T.; Seldin, M.; et al. APOBEC3B drives PKR-mediated translation shutdown and protects stress granules in response to viral infection. Nat. Commun. 2023, 14, 820. [Google Scholar] [CrossRef]
- Stenglein, M.D.; Harris, R.S. APOBEC3B and APOBEC3F Inhibit L1 Retrotransposition by a DNA Deamination-independent Mechanism. J. Biol. Chem. 2006, 281, 16837–16841. [Google Scholar] [CrossRef]
- Feng, Y.; Goubran, M.H.; Follack, T.B.; Chelico, L. Deamination-independent restriction of LINE-1 retrotransposition by APOBEC3H. Sci. Rep. 2017, 7, 10881. [Google Scholar] [CrossRef]
- Richardson, S.R.; Narvaiza, I.; Planegger, R.A.; Weitzman, M.D.; Moran, J.V. APOBEC3A deaminates transiently exposed single-strand DNA during LINE-1 retrotransposition. Elife 2014, 3, e02008. [Google Scholar] [CrossRef] [PubMed]
- Frost, R.J.A.; Hamra, F.K.; Richardson, J.A.; Qi, X.; Bassel-Duby, R.; Olson, E.N. MOV10L1 is necessary for protection of spermatocytes against retrotransposons by Piwi-interacting RNAs. Proc. Natl. Acad. Sci. USA 2010, 107, 11847–11852. [Google Scholar] [CrossRef]
- Zheng, K.; Xiol, J.; Reuter, M.; Eckardt, S.; Leu, N.A.; McLaughlin, K.J.; Stark, A.; Sachidanandam, R.; Pillai, R.S.; Wang, P.J. Mouse MOV10L1 associates with Piwi proteins and is an essential component of the Piwi-interacting RNA (piRNA) pathway. Proc. Natl. Acad. Sci. USA 2010, 107, 11841–11846. [Google Scholar] [CrossRef]
- Goodier, J.L.; Cheung, L.E.; Kazazian, H.H., Jr. MOV10 RNA Helicase Is a Potent Inhibitor of Retrotransposition in Cells. PLoS Genet. 2012, 8, e1002941. [Google Scholar] [CrossRef]
- Li, X.; Zhang, J.; Jia, R.; Cheng, V.; Xu, X.; Qiao, W.; Guo, F.; Liang, C.; Cen, S. The MOV10 Helicase Inhibits LINE-1 Mobility. J. Biol. Chem. 2013, 288, 21148–21160. [Google Scholar] [CrossRef]
- Meister, G.; Landthaler, M.; Peters, L.; Chen, P.Y.; Urlaub, H.; Lührmann, R.; Tuschl, T. Identification of Novel Argonaute-Associated Proteins. Curr. Biol. 2005, 15, 2149–2155. [Google Scholar] [CrossRef] [PubMed]
- Warkocki, Z.; Krawczyk, P.S.; Adamska, D.; Bijata, K.; Garcia-Perez, J.L.; Dziembowski, A. Uridylation by TUT4/7 Restricts Retrotransposition of Human LINE-1s. Cell 2018, 174, 1537–1548.e1529. [Google Scholar] [CrossRef] [PubMed]
- Boeke, J.D.; Garfinkel, D.J.; Styles, C.A.; Fink, G.R. Ty elements transpose through an RNA intermediate. Cell 1985, 40, 491–500. [Google Scholar] [CrossRef] [PubMed]
- Hansen, L.J.; Chalker, D.L.; Sandmeyer, S.B. Ty3, a yeast retrotransposon associated with tRNA genes, has homology to animal retroviruses. Mol. Cell. Biol. 1988, 8, 5245–5256. [Google Scholar] [CrossRef] [PubMed]
- Griffith, J.L.; Coleman, L.E.; Raymond, A.S.; Goodson, S.G.; Pittard, W.S.; Tsui, C.; Devine, S.E. Functional Genomics Reveals Relationships between the Retrovirus-Like Ty1 Element and Its Host Saccharomyces cerevisiae. Genetics 2003, 164, 867–879. [Google Scholar] [CrossRef] [PubMed]
- Irwin, B.; Aye, M.; Baldi, P.; Beliakova-Bethell, N.; Cheng, H.; Dou, Y.; Liou, W.; Sandmeyer, S. Retroviruses and yeast retrotransposons use overlapping sets of host genes. Genome Res. 2005, 15, 641–654. [Google Scholar] [CrossRef]
- Aye, M.; Irwin, B.; Beliakova-Bethell, N.; Chen, E.; Garrus, J.; Sandmeyer, S. Host Factors That Affect Ty3 Retrotransposition in Saccharomyces cerevisiae. Genetics 2004, 168, 1159–1176. [Google Scholar] [CrossRef] [PubMed]
- Beliakova-Bethell, N.; Beckham, C.; Giddings, T.H., Jr.; Winey, M.; Parker, R.; Sandmeyer, S. Virus-like particles of the Ty3 retrotransposon assemble in association with P-body components. RNA 2006, 12, 94–101. [Google Scholar] [CrossRef] [PubMed]
- Checkley, M.A.; Nagashima, K.; Lockett, S.J.; Nyswaner, K.M.; Garfinkel, D.J. P-Body Components Are Required for Ty1 Retrotransposition during Assembly of Retrotransposition-Competent Virus-Like Particles. Mol. Cell. Biol. 2010, 30, 382–398. [Google Scholar] [CrossRef]
- Dutko, J.A.; Kenny, A.E.; Gamache, E.R.; Curcio, M.J. 5′ to 3′ mRNA Decay Factors Colocalize with Ty1 Gag and Human APOBEC3G and Promote Ty1 Retrotransposition. J. Virol. 2010, 84, 5052–5066. [Google Scholar] [CrossRef]
- Bilanchone, V.; Clemens, K.; Kaake, R.; Dawson, A.R.; Matheos, D.; Nagashima, K.; Sitlani, P.; Patterson, K.; Chang, I.; Huang, L.; et al. Ty3 Retrotransposon Hijacks Mating Yeast RNA Processing Bodies to Infect New Genomes. PLOS Genet. 2015, 11, e1005528. [Google Scholar] [CrossRef]
- Dewannieux, M.; Dupressoir, A.; Harper, F.; Pierron, G.; Heidmann, T. Identification of autonomous IAP LTR retrotransposons mobile in mammalian cells. Nat. Genet. 2004, 36, 534–539. [Google Scholar] [CrossRef]
- Kuff, E.L.; Lueders, K.K. The Intracisternal A-Particle Gene Family: Structure and Functional Aspects. Adv. Cancer Res. 1988, 51, 183–276. [Google Scholar] [CrossRef]
- Heldmann, O.; Heidmann, T. Retrotransposition of a mouse IAP sequence tagged with an indicator gene. Cell 1991, 64, 159–170. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.; Contreras, X.; Peterlin, B.M. P Bodies Inhibit Retrotransposition of Endogenous Intracisternal A Particles. J. Virol. 2011, 85, 6244–6251. [Google Scholar] [CrossRef]
- Lu, C.; Luo, Z.; Jäger, S.; Krogan, N.J.; Peterlin, B.M. Moloney Leukemia Virus Type 10 Inhibits Reverse Transcription and Retrotransposition of Intracisternal a Particles. J. Virol. 2012, 86, 10517–10523. [Google Scholar] [CrossRef]
- Benda, C. Neue mitteilungen über die entwickelung der genitaldrüsen und die metamorphose der samenzellen (histogenese der spermatozoen). Verhandlungen der berliner physiologischen gesellschaft. Arch. Anat. Physiol. 1891, 1891, 549–552. [Google Scholar]
- Eddy, E.M. Fine structural observations on the form and distribution of nuage in germ cells of the rat. Anat. Rec. 1974, 178, 731–757. [Google Scholar] [CrossRef] [PubMed]
- Kotaja, N.; Bhattacharyya, S.N.; Jaskiewicz, L.; Kimmins, S.; Parvinen, M.; Filipowicz, W.; Sassone-Corsi, P. The chromatoid body of male germ cells: Similarity with processing bodies and presence of Dicer and microRNA pathway components. Proc. Natl. Acad. Sci. USA 2006, 103, 2647–2652. [Google Scholar] [CrossRef] [PubMed]
- Aravin, A.A.; van der Heijden, G.W.; Castañeda, J.; Vagin, V.V.; Hannon, G.J.; Bortvin, A. Cytoplasmic Compartmentalization of the Fetal piRNA Pathway in Mice. PLOS Genet. 2009, 5, e1000764. [Google Scholar] [CrossRef] [PubMed]
- Lim, A.K.; Tao, L.; Kai, T. piRNAs mediate posttranscriptional retroelement silencing and localization to pi-bodies in the Drosophila germline. J. Cell Biol. 2009, 186, 333–342. [Google Scholar] [CrossRef]
- Liu, L.; Qi, H.; Wang, J.; Lin, H. PAPI, a novel TUDOR-domain protein, complexes with AGO3, ME31B and TRAL in the nuage to silence transposition. Development 2011, 138, 1863–1873. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions, and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions, or products referred to in the content. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Moon, S.; Namkoong, S. Ribonucleoprotein Granules: Between Stress and Transposable Elements. Biomolecules 2023, 13, 1027. https://doi.org/10.3390/biom13071027
Moon S, Namkoong S. Ribonucleoprotein Granules: Between Stress and Transposable Elements. Biomolecules. 2023; 13(7):1027. https://doi.org/10.3390/biom13071027
Chicago/Turabian StyleMoon, Sungjin, and Sim Namkoong. 2023. "Ribonucleoprotein Granules: Between Stress and Transposable Elements" Biomolecules 13, no. 7: 1027. https://doi.org/10.3390/biom13071027
APA StyleMoon, S., & Namkoong, S. (2023). Ribonucleoprotein Granules: Between Stress and Transposable Elements. Biomolecules, 13(7), 1027. https://doi.org/10.3390/biom13071027







