Recognition of ATT Triplex and DNA:RNA Hybrid Structures by Benzothiazole Ligands
Abstract
1. Introduction
2. Materials and Methods
2.1. Spectroscopy Measurements
2.2. ITC Measurements
2.3. Molecular Modeling
3. Results and Discussion
3.1. Characterization of Compounds in Aqueous Medium
3.2. Study of Interactions of Benzothiazoles with Nucleic Acids in Aqueous Medium
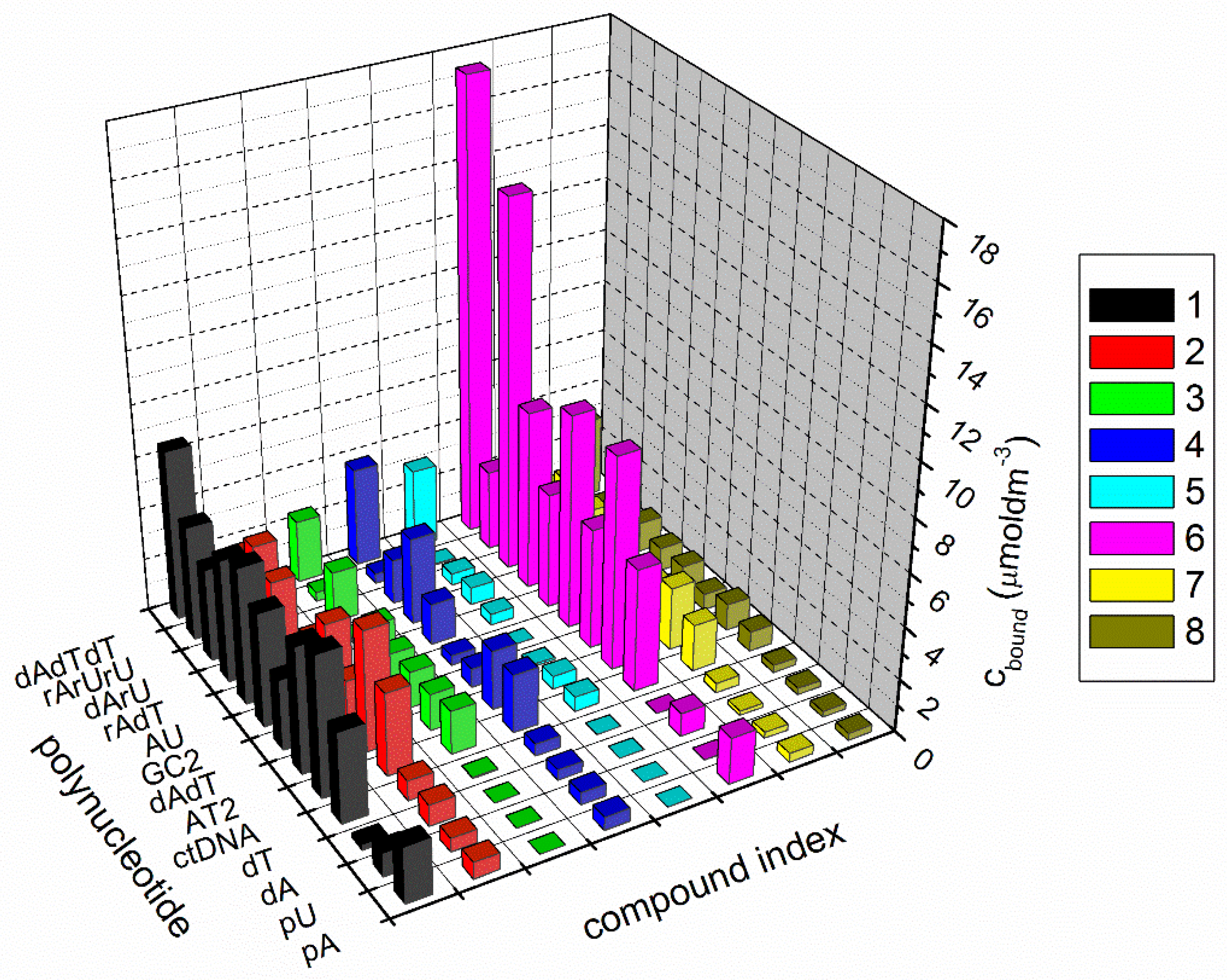
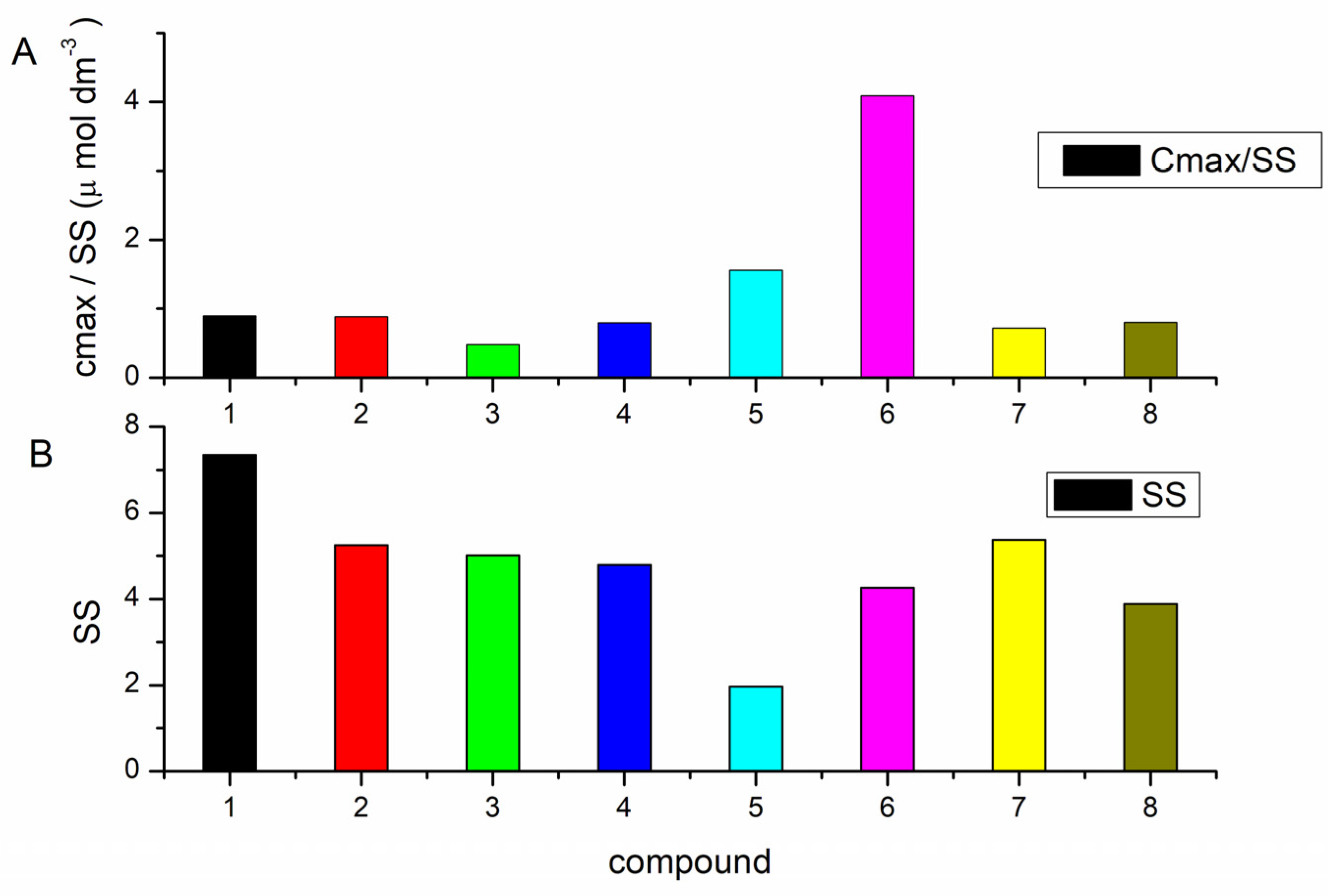
3.2.1. Competition Dialysis Assay with 1–9
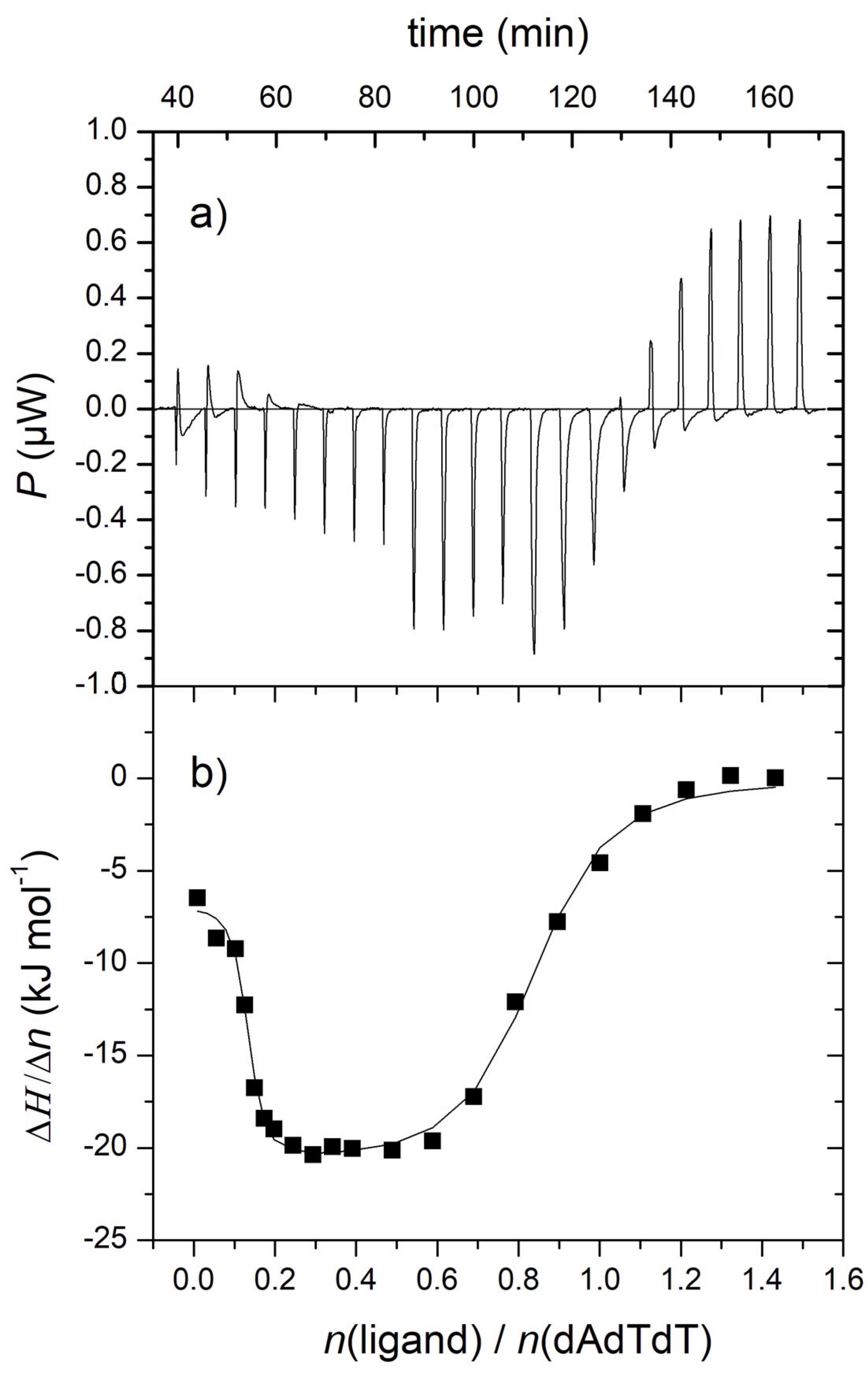
3.2.2. Fluorescence Spectroscopy and Isothermal Titration Calorimetry
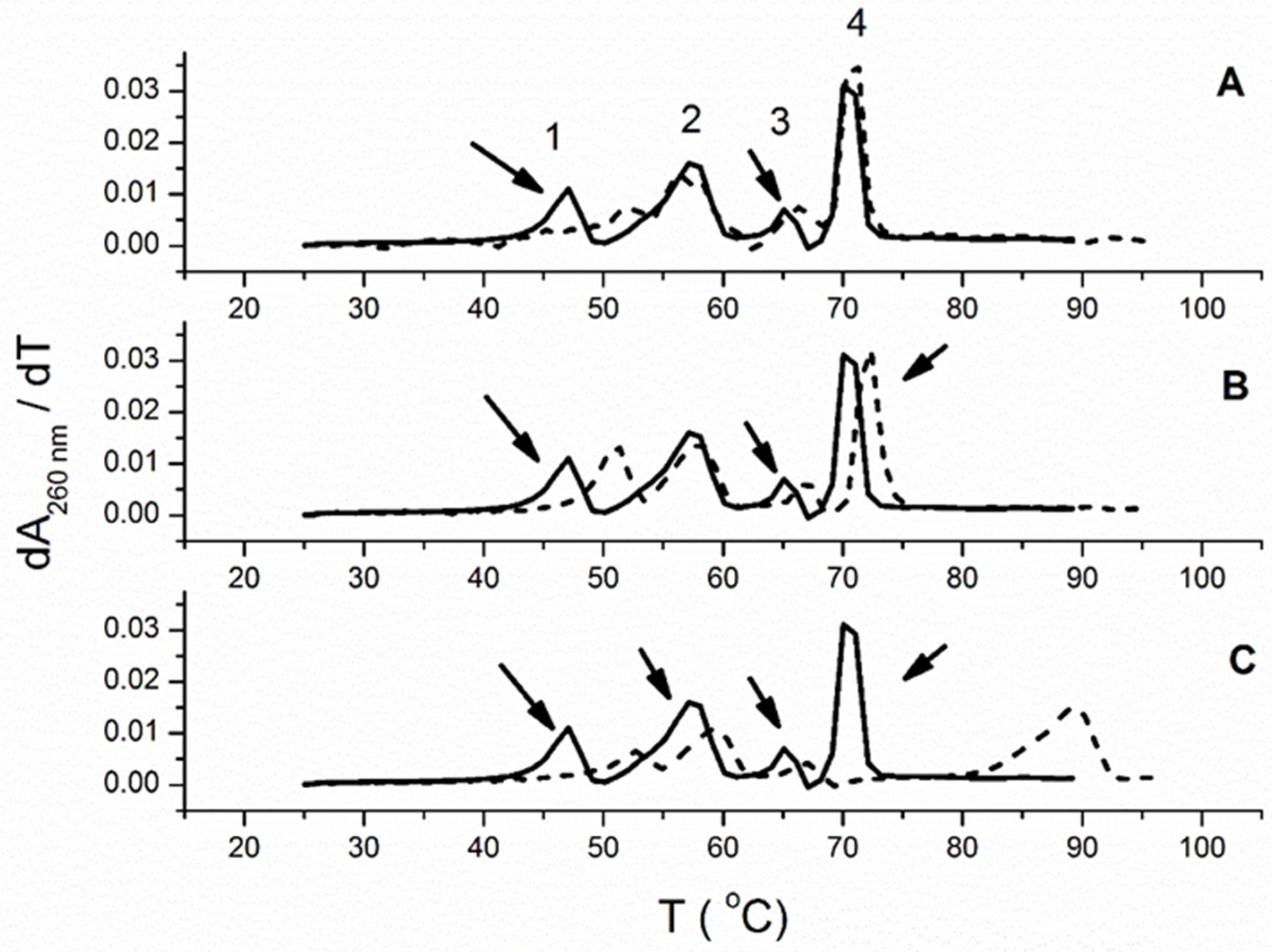
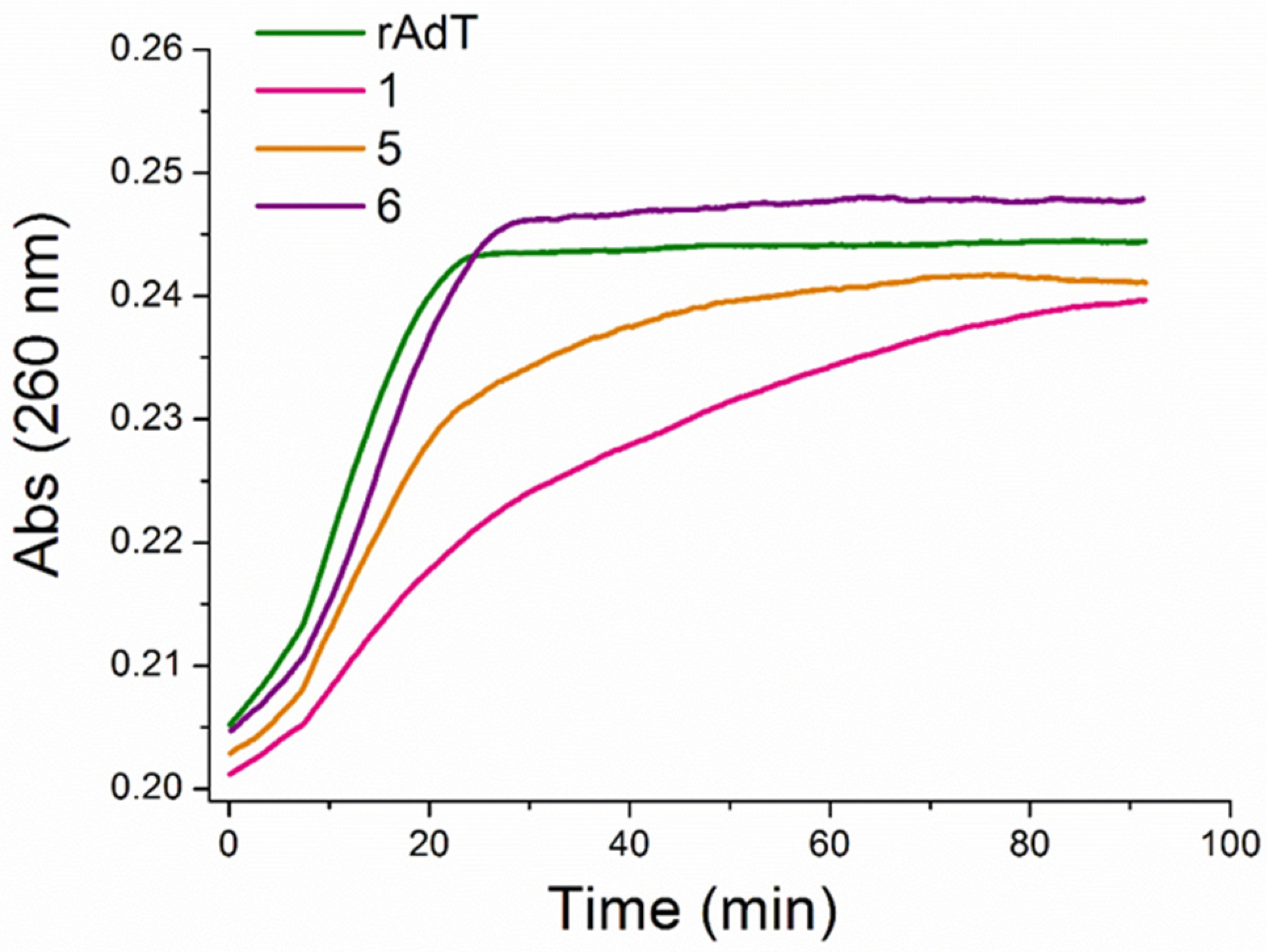
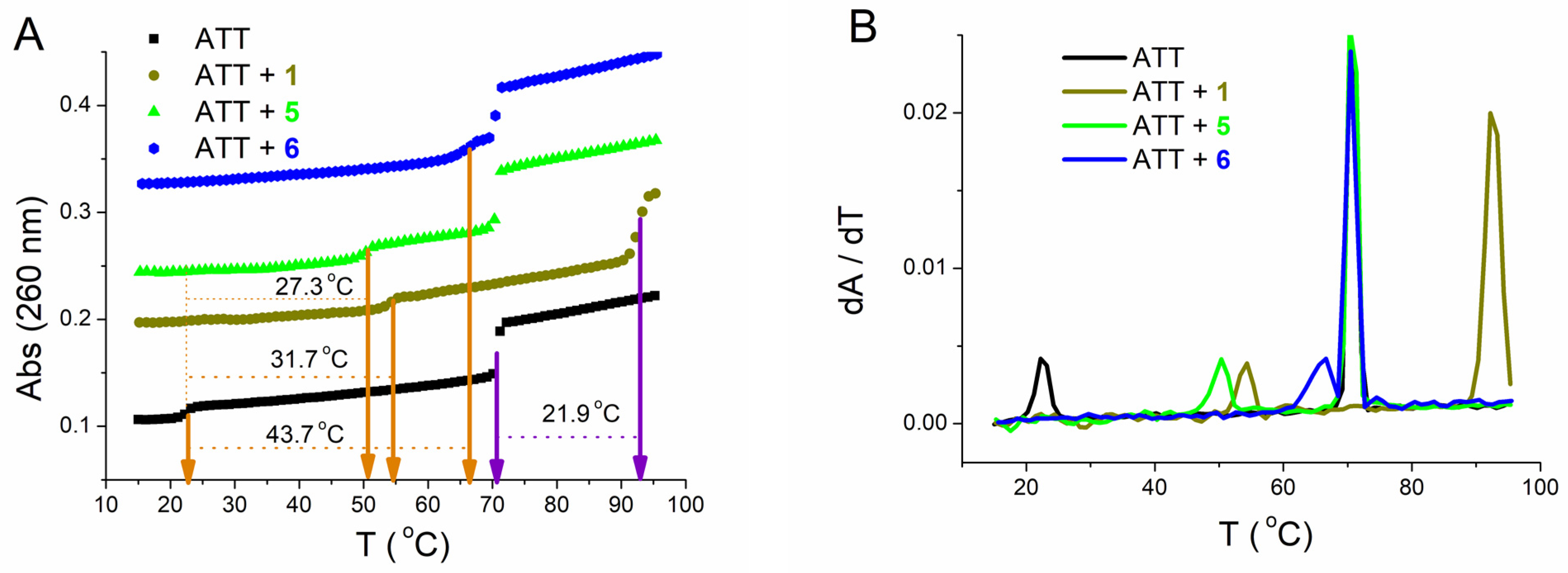
3.2.3. Thermal Melting Experiments and RNase H Assay
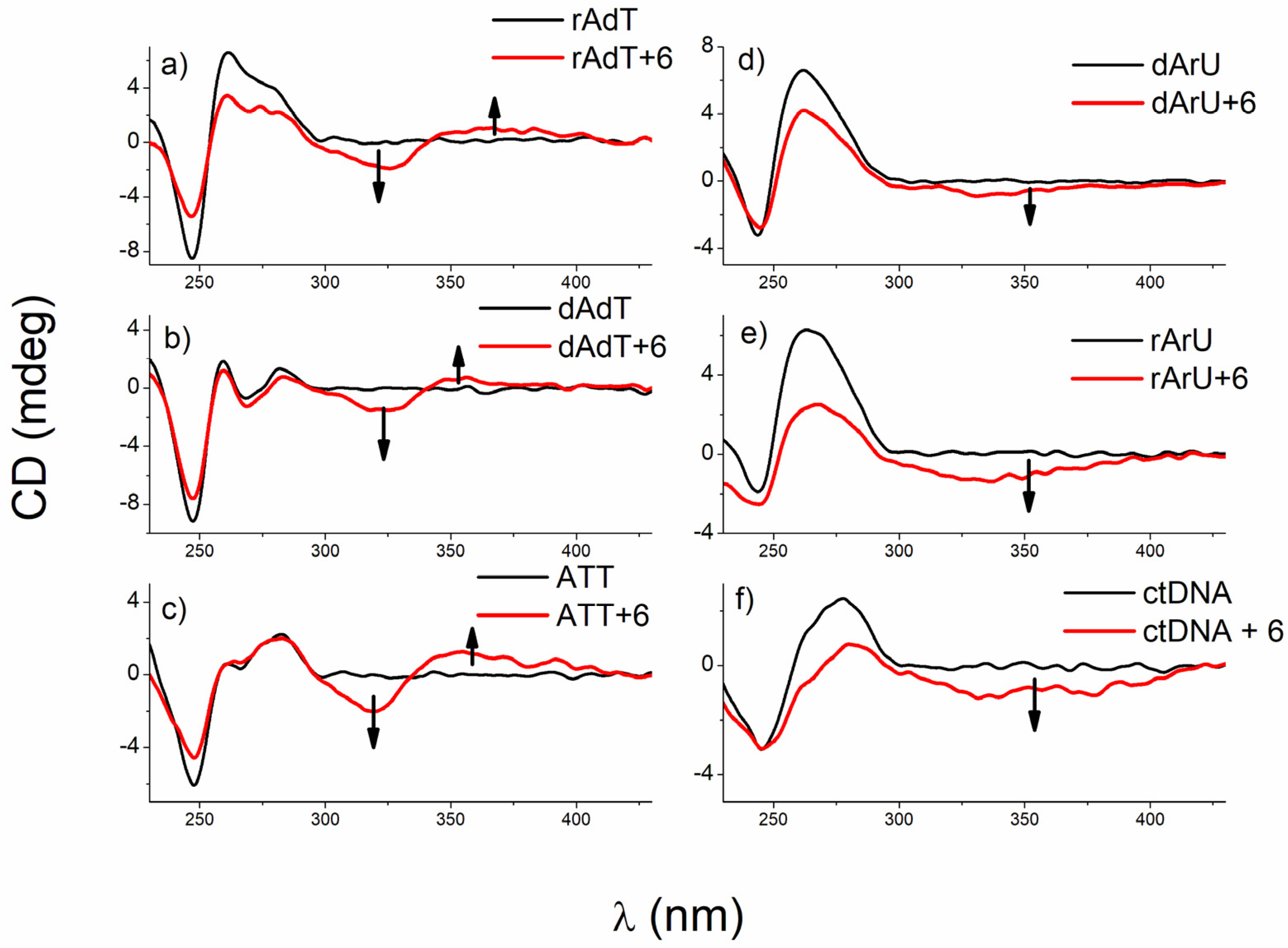
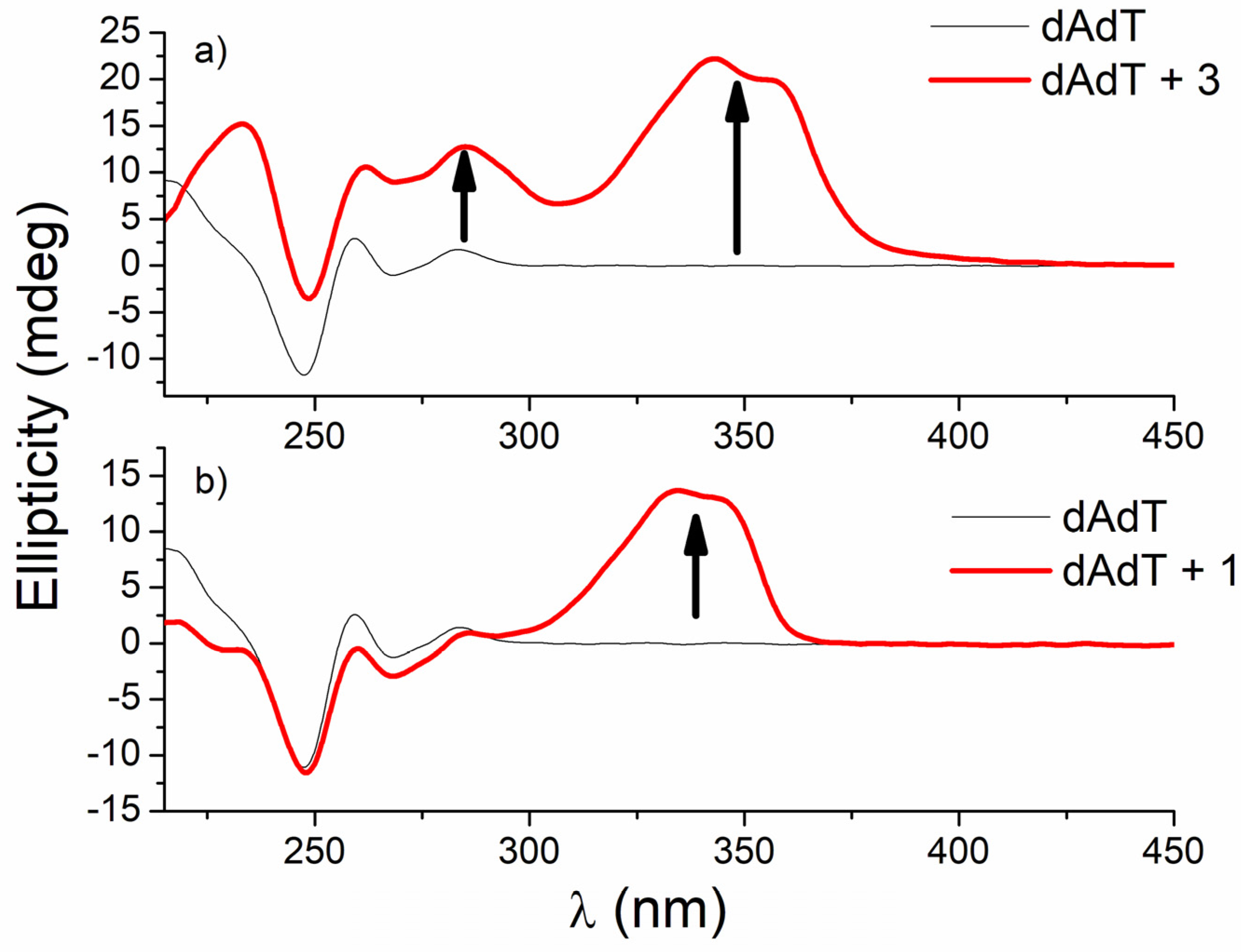
3.2.4. Circular Dichroism (CD) Experiments
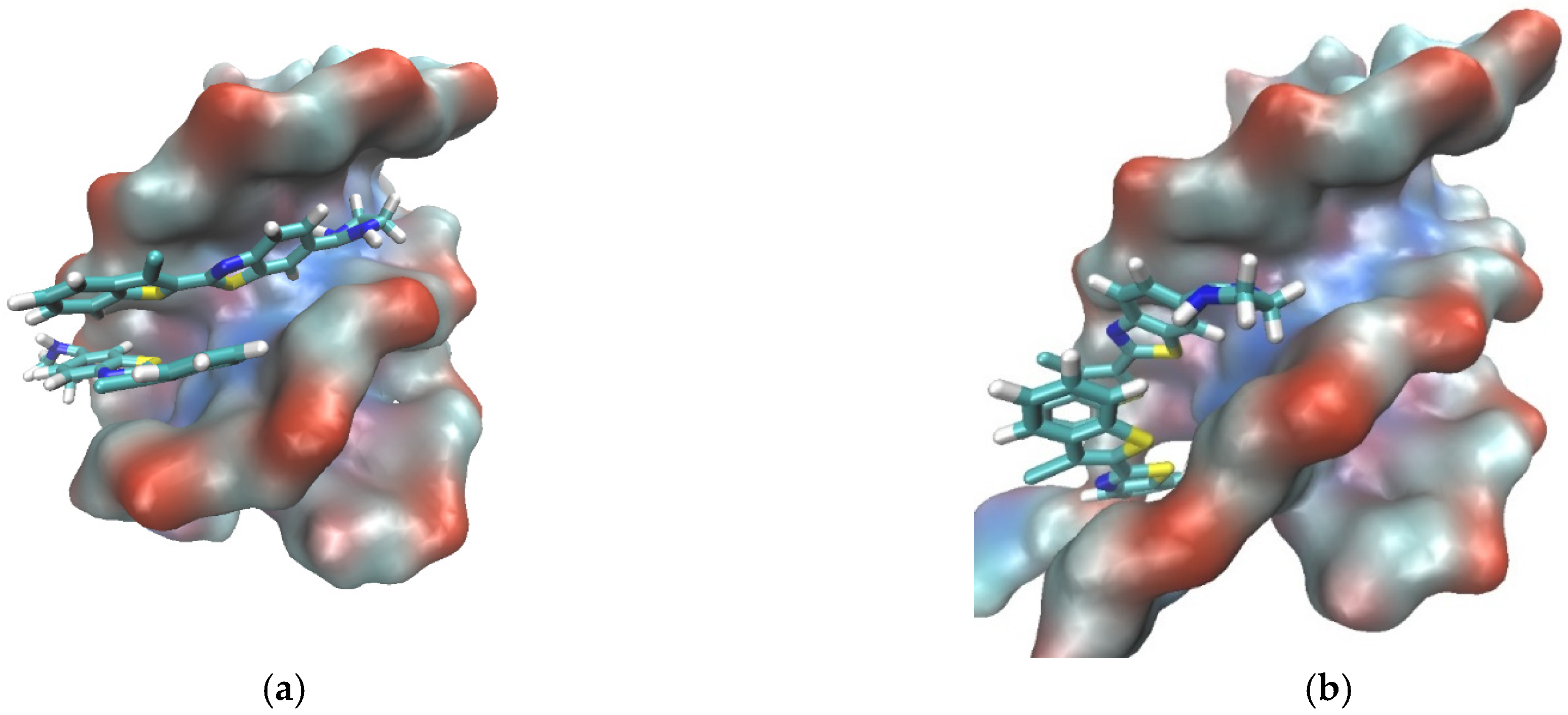
3.2.5. Molecular Modeling
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hurley, L.H. DNA and its associated processes as targets for cancer therapy. Nat. Cancer 2002, 2, 188–200. [Google Scholar] [CrossRef] [PubMed]
- Demeunynck, M.; Bailly, C.; Wilson, W.D. Small Molecule DNA and RNA Binders: From Synthesis to Nucleic Acid Complexes; Demeunynck, M., Bailly, C., Wilson, W.D., Eds.; Wiley-VCH: Weinheim, Germany, 2003. [Google Scholar]
- Escudé, C.; Nguyen, C.H.; Kukreti, S.; Janin, Y.; Sun, J.-S.; Bisagni, E.; Garestier, T.; Hélène, C. Rational design of a triple helix-specific intercalating ligand. Proc. Natl. Acad. Sci. USA 1998, 95, 3591–3596. [Google Scholar] [CrossRef] [PubMed]
- Arya, D.P. New Approaches Toward Recognition of Nucleic Acid Triple Helices. Acc. Chem. Res. 2011, 44, 134–146. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.Q.N.; Lim, K.W.; Phan, A.T. A Dual-Specific Targeting Approach Based on the Simultaneous Recognition of Duplex and Quadruplex Motifs. Sci. Rep. 2017, 7, 11969. [Google Scholar] [CrossRef]
- Le, D.D.; Di Antonio, M.; Chan, L.K.M.; Balasubramanian, S. G-quadruplex ligands exhibit differential G-tetrad selectivity. Chem. Commun. 2015, 51, 8048–8050. [Google Scholar] [CrossRef]
- Ranjan, N.; Davis, E.; Xue, L.; Arya, D.P. Dual recognition of the human telomeric G-quadruplex by a neomycin–anthraquinone conjugate. Chem. Commun. 2013, 49, 5796–5798. [Google Scholar] [CrossRef]
- Van Dyke, M.W.; Ohyama, T. Do DNA Triple Helices or Quadruplexes Have a Role in Transcription? In DNA Conformation and Transcription; Springer Science and Business Media LLC: Berlin/Heidelberg, Germany, 2005; pp. 105–126. [Google Scholar]
- Nadal, A.; Eritja, R.; Esteve, T.; Pla, M. “Parallel” and “antiparallel tail-clamps” increase the efficiency of triplex formation with structured DNA and RNA targets. ChemBioChem 2005, 6, 1034–1042. [Google Scholar] [CrossRef]
- Wang, G.; Seidman, M.M.; Glazer, P.M. Mutagenesis in Mammalian Cells Induced by Triple Helix Formation and Transcription-Coupled Repair. Science 1996, 271, 802–805. [Google Scholar] [CrossRef]
- Jain, A.; Wang, G.; Vasquez, K.M. DNA triple helices: Biological consequences and therapeutic potential. Biochimie 2008, 90, 1117–1130. [Google Scholar] [CrossRef]
- Xi, H.; Kumar, S.; Dosen-Micovic, L.; Arya, D.P. Calorimetric and spectroscopic studies of aminoglycoside binding to AT-rich DNA triple helices. Biochimie 2010, 92, 514–529. [Google Scholar] [CrossRef]
- Sharma, S.K.; Frase, W. Selectivity of a bromoacridine containing fuorophore for triplex DNA. Mon. Chem. 2021, 152, 1013–1016. [Google Scholar] [CrossRef]
- Xue, L.; Xi, H.; Kumar, S.; Gray, D.; Davis, E.; Hamilton, P.; Skriba, M.; Arya, D.P. Probing the Recognition Surface of a DNA Triplex: Binding Studies with Intercalator−Neomycin Conjugates. Biochemistry 2010, 49, 5540–5552. [Google Scholar] [CrossRef] [PubMed]
- Mergny, J.L.; Duval-Valentin, G.; Nguyen, C.H.; Perrouault, L.; Faucon, B.; Rougée, M.; Montenay-Garestier, T.; Bisagni, E.; Hélène, C. Triple Helix-Specific Ligands. Science 1992, 256, 1681–1684. [Google Scholar] [CrossRef] [PubMed]
- Houlard, M.; Artus, J.; Leguillier, T.; Vandormael-Pournin, S.; Cohen-Tannoudji, M. DNA-RNA hybrids contribute to the replication dependent genomic instability induced byOmcg1deficiency. Cell Cycle 2011, 10, 108–117. [Google Scholar] [CrossRef] [PubMed]
- Balk, B.; Dees, M.; Bender, K.; Luke, B. The differential processing of telomeres in response to increased telomeric transcription and RNA–DNA hybrid accumulation. RNA Biol. 2014, 11, 95–100. [Google Scholar] [CrossRef] [PubMed]
- Stuckey, R.; García-Rodríguez, N.; Aguilera, A.; Wellinger, R.E. Role for RNA:DNA hybrids in origin-independent replication priming in a eukaryotic system. Proc. Natl. Acad. Sci. USA 2015, 112, 5779–5784. [Google Scholar] [CrossRef]
- Balk, B.; Maicher, A.; Dees, M.; Klermund, J.; Luke-Glaser, S.; Bender, K.; Luke, B. Telomeric RNA-DNA hybrids affect telomere-length dynamics and senescence. Nat. Struct. Mol. Biol. 2013, 20, 1199–1205. [Google Scholar] [CrossRef]
- Li, T.-K.; Barbieri, C.M.; Lin, H.-C.; Rabson, A.B.; Yang, G.; Fan, Y.; Gaffney, B.L.; Jones, R.A.; Pilch, D.S. Drug Targeting of HIV-1 RNA·DNA Hybrid Structures: Thermodynamics of Recognition and Impact on Reverse Transcriptase-Mediated Ribonuclease H Activity and Viral Replication. Biochemistry 2004, 43, 9732–9742. [Google Scholar] [CrossRef]
- Lombraña, R.; Almeida, R.; Alvarez, A.; Gómez, M. R-loops and initiation of DNA replication in human cells: A missing link? Front. Genet. 2015, 6, 158. [Google Scholar] [CrossRef]
- Saretzki, G. Telomerase inhibition as cancer therapy. Cancer Lett. 2003, 194, 209–219. [Google Scholar] [CrossRef]
- Mergny, J.-L.; Lacroix, L.; Teulade-Fichou, M.-P.; Hounsou, C.; Guittat, L.; Hoarau, M.; Arimondo, P.B.; Vigneron, J.-P.; Lehn, J.-M.; Riou, J.-F.; et al. Telomerase inhibitors based on quadruplex ligands selected by a fluorescence assay. Proc. Natl. Acad. Sci. USA 2001, 98, 3062–3067. [Google Scholar] [CrossRef] [PubMed]
- Su, H.-P.; Yan, Y.; Prasad, G.S.; Smith, R.F.; Daniels, C.L.; Abeywickrema, P.D.; Reid, J.C.; Loughran, H.M.; Kornienko, M.; Sharma, S.; et al. Structural Basis for the Inhibition of RNase H Activity of HIV-1 Reverse Transcriptase by RNase H Active Site-Directed Inhibitors. J. Virol. 2010, 84, 7625–7633. [Google Scholar] [CrossRef] [PubMed]
- Tramontano, E.; Di Santo, R. HIV-1 RT-associated RNase H function inhibitors: Recent advances in drug development. Curr. Med. Chem. 2010, 17, 2837–2853. [Google Scholar] [CrossRef]
- Ren, J.; Qu, X.; Dattagupta, N.; Chaires, J. Molecular Recognition of a RNA:DNA Hybrid Structure. J. Am. Chem. Soc. 2001, 123, 6742–6743. [Google Scholar] [CrossRef]
- Shaw, N.N.; Arya, D.P. Recognition of the unique structure of DNA:RNA hybrids. Biochimie 2008, 90, 1026–1039. [Google Scholar] [CrossRef] [PubMed]
- West, C.; Francis, R.; Friedman, S.H. Small molecule/Nucleic acid affinity chromatography: Application for the identification of telomerase inhibitors which target its key RNA/DNA heteroduplex. Bioorganic Med. Chem. Lett. 2001, 11, 2727–2730. [Google Scholar] [CrossRef]
- Wheelhouse, R.T.; Chaires, J.B. Drug binding to DNA x RNA hybrid structures. Methods Mol. Biol. 2010, 613, 55–70. [Google Scholar]
- Shi, X.; Chaires, J.B. Sequence- and structural-selective nucleic acid binding revealed by the melting of mixtures. Nucleic Acids Res. 2006, 34, e14. [Google Scholar] [CrossRef]
- Shaw, N.N.; Xi, H.; Arya, D.P. Molecular recognition of a DNA:RNA hybrid: Sub-nanomolar binding by a neomycin–methidium conjugate. Bioorganic Med. Chem. Lett. 2008, 18, 4142–4145. [Google Scholar] [CrossRef]
- Xi, H.; Davis, E.; Ranjan, N.; Xue, L.; Hyde-Volpe, D.; Arya, D.P. Thermodynamics of Nucleic Acid “Shape Readout” by an Aminosugar. Biochemistry 2011, 50, 9088–9113. [Google Scholar] [CrossRef]
- Wheelhouse, R.T.; Garbett, N.C.; Buurma, N.J.; Chaires, J. Probing the Molecular Recognition of a DNA⋅RNA Hybrid Duplex. Angew. Chem. Int. Ed. 2010, 49, 3207–3210. [Google Scholar] [CrossRef] [PubMed]
- Racané, L.; Kraljević Pavelić, S.; Ratkaj, I.; Stepanić, V.; Pavelić, K.; Tralić-Kulenović, V.; Karminski-Zamola, G. Synthesis and antiproliferative evaluation of some new amidino-substituted bis-benzothiazolyl-pyridines and pyrazine. Eur. J. Med. Chem. 2012, 55, 108–116. [Google Scholar] [CrossRef] [PubMed]
- Racané, L.; Sedić, M.; Ilić, N.; Aleksić, M.; Pavelić, S.K.; Karminski-Zamola, G. Novel 2-Thienyl- and 2-Benzothienyl-Substituted 6-(2-Imidazolinyl)Benzothiazoles: Synthesis; in vitro Evaluation of Antitumor Effects and Assessment of Mitochondrial Toxicity. Anti-Cancer Agents Med. Chem. 2017, 17, 57–66. [Google Scholar]
- Racané, L.; Ptiček, L.; Sedić, M.; Grbčić, P.; Pavelić, S.K.; Bertosa, B.; Sović, I.; Karminski-Zamola, G. Eco-friendly synthesis, in vitro anti-proliferative evaluation, and 3D-QSAR analysis of a novel series of monocationic 2-aryl/heteroaryl-substituted 6-(2-imidazolinyl)benzothiazole mesylates. Mol. Divers. 2018, 22, 723–741. [Google Scholar] [CrossRef]
- Chaires, J.; Dattagupta, N.; Crothers, D.M. Studies on interaction of anthracycline antibiotics and deoxyribonucleic acid: Equilibrium binding studies on the interaction of daunomycin with deoxyribonucleic acid. Biochemistry 1982, 21, 3933–3940. [Google Scholar] [CrossRef]
- Chalikian, T.V.; Volker, J.; Plum, G.E.; Breslauer, K.J. A more unified picture for the thermodynamics of nucleic acid duplex melting: A characterization by calorimetric and volumetric techniques. Proc. Natl. Acad. Sci. USA 1999, 96, 7853–7858. [Google Scholar] [CrossRef]
- Scatchard, G. The Attractions of Proteins for Small Molecules and Ions. Ann. N. Y. Acad. Sci. 1949, 51, 660–672. [Google Scholar] [CrossRef]
- Ragazzon, P.; Chaires, J.B. Use of competition dialysis in the discovery of G-quadruplex selective ligands. Methods 2007, 43, 313–323. [Google Scholar] [CrossRef]
- Ivani, I.; Dans, P.D.; Noy, A.; Pérez, A.; Faustino, I.; Hospital, A.; Walther, J.; Andrio, P.; Goñi, R.; Balaceanu, A.; et al. Parmbsc1: A Refined ForceField for DNA Simulations. Nat. Methods 2015, 13, 55–58. [Google Scholar] [CrossRef]
- Schrödinger, L.; DeLano, W. The PyMOL Molecular Graphics System, Version 1.5.0.1.; Schrödinger, Inc.: New York, NY, USA, 2010. [Google Scholar]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- Li, P.; Roberts, B.P.; Chakravorty, D.K.; Merz, J.K.M. Rational Design of Particle Mesh Ewald Compatible Lennard-Jones Parameters for +2 Metal Cations in Explicit Solvent. J. Chem. Theory Comput. 2013, 9, 2733–2748. [Google Scholar] [CrossRef] [PubMed]
- Loncharich, R.J.; Brooks, B.R.; Pastor, R.W. Langevin dynamics of peptides: The frictional dependence of isomerization rates ofN-acetylalanyl-N?-methylamide. Biopolymers 1992, 32, 523–535. [Google Scholar] [CrossRef] [PubMed]
- Berendsen, H.J.C.; Postma, J.P.M.; Van Gunsteren, W.F.; DiNola, A.; Haak, J.R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 1984, 81, 3684. [Google Scholar] [CrossRef]
- Košćak, M.; Krošl, I.; Žinić, B.; Piantanida, I. Fluorescent Analogues of FRH Peptide: Cu(II) Binding and Interactions with ds-DNA/RNA. Chemosensors 2022, 10, 34. [Google Scholar] [CrossRef]
- Bronowska, A.K. Thermodynamics of Ligand-Protein Interactions: Implications for Molecular Design. In Thermodynamics—Interaction Studies—Solids, Liquids and Gases; IntechOpen: London, UK, 2011. [Google Scholar]
- Perozzo, R.; Folkers, G.; Scapozza, L. Thermodynamics of Protein–Ligand Interactions: History, Presence, and Future Aspects. J. Recept. Signal Transduct. 2004, 24, 1–52. [Google Scholar] [CrossRef]
- McGhee, J.D.; von Hippel, P.H. Theoretical aspects of DNA-protein interactions: Co-operative and non-co-operative binding of large ligands to a one-dimensional homogeneous lattice. J. Mol. Biol. 1974, 86, 469–489. [Google Scholar] [CrossRef]
- Chaires, J.B. A thermodynamic signature for drug–DNA binding mode. Arch. Biochem. Biophys. 2006, 453, 26–31. [Google Scholar] [CrossRef]
- Mergny, J.-L.; Lacroix, L. Analysis of Thermal Melting Curves. Oligonucleotides 2003, 13, 515–537. [Google Scholar] [CrossRef]
- Lesnik, E.A.; Freier, S.M. Relative Thermodynamic Stability of DNA, RNA, and DNA:RNA Hybrid Duplexes: Relationship with Base Composition and Structure. Biochemistry 1995, 34, 10807–10815. [Google Scholar] [CrossRef]
- Choi, B.-H.; Yeo, G.; Jung, J.; Lee, B.W.; Han, S.; Cho, T. A Thermodynamic Investigation into the Stabilization of Poly(dA)·[poly(dT)] 2 Triple Helical DNA by Various Divalent Metal Ions. Bull. Korean Chem. Soc. 2009, 30, 2691–2696. [Google Scholar]
- Arya, D.P.; Coffee, R.L.; Willis, B.; Abramovitch, A.I. Aminoglycoside−Nucleic Acid Interactions: Remarkable Stabilization of DNA and RNA Triple Helices by Neomycin. J. Am. Chem. Soc. 2001, 123, 5385–5395. [Google Scholar] [CrossRef] [PubMed]
- Berova, N.; Nakanishi, K.; Woody, R. Circular Dichroism Principles and Applications, 2nd ed.; Wiley-VCH: New York, NY, USA, 2000. [Google Scholar]
- Eriksson, M.; Nordén, B. Linear and circular dichroism of drug-nucleic acid complexes. In Methods in Enzymology; Elsevier BV: Amsterdam, The Netherlands, 2001; Volume 340, pp. 68–98. [Google Scholar]
- Egli, M.; Saenger, W. Principles of Nucleic Acid Structure; Springer: Berlin/Heidelberg, Germany, 1983. [Google Scholar]
- Cantor, C.R.; Schimmel, P.R. Biophysical Chemistry; W.H. Freeman: San Francisco, CA, USA, 1980. [Google Scholar]
- Neidle, S. Oxford Handbook of Nucleic Acid Structure; Oxford University Press: Oxford, UK, 1999. [Google Scholar]
- Fedoroff, O.Y.; Salazar, M.; Reid, B.R. Structure of a DNA: RNA Hybrid Duplex: Why RNase H Does Not Cleave Pure RNA. J. Mol. Biol. 1993, 233, 509–523. [Google Scholar] [CrossRef] [PubMed]
- Radhakrishnan, I.; Patel, D.J. Solution structure of a pyrimidine·purine·pyrimidine DNA triplex containing T·AT, C+ GC and G·TA triples. Structure 1994, 2, 17–32. [Google Scholar] [CrossRef]
- Raghunathan, G.; Miles, H.T.; Sasisekharan, V. Symmetry and molecular structure of a DNA triple helix: D(T)n.cntdot.d(A)n.cntdot.d(T)n. Biochemistry 1993, 32, 455–462. [Google Scholar] [CrossRef]
- Esguerra, M.; Nilsson, L.; Villa, A. Triple helical DNA in a duplex context and base pair opening. Nucleic Acids Res. 2014, 42, 11329–11338. [Google Scholar] [CrossRef]
- Šmidlehner, T.; Piantanida, I.; Pescitelli, G. Polarization spectroscopy methods in the determination of interactions of small molecules with nucleic acids—Tutorial. Beilstein J. Org. Chem. 2018, 14, 84–105. [Google Scholar] [CrossRef]
- Stojković, M.R.; González-García, J.; Šupljika, F.; Galiana-Roselló, C.; Guijarro, L.; Gazzè, S.A.; Francis, L.W.; Piantanida, I.; García-España, E. Specific and highly efficient condensation of GC and IC DNA by polyaza pyridinophane derivatives. Int. J. Biol. Macromol. 2017, 109, 143–151. [Google Scholar] [CrossRef]
- Wang, J.; Wang, W.; Kollman, P.A.; Case, D.A. Automatic atom type and bond type perception in molecular mechanical calculations. J. Mol. Graph. Model. 2006, 25, 247–260. [Google Scholar] [CrossRef]
- Case, D.; Betz, R.; Cerutti, D.S.; Cheatham, T.; Darden, T.; Duke, R.; Giese, T.J.; Gohlke, H.; Götz, A.; Homeyer, N.; et al. Amber 16; University of California: San Francisco, CA, USA, 2016. [Google Scholar]
- Zgarbová, M.; Šponer, J.; Otyepka, M.; Cheatham, I.T.E.; Galindo-Murillo, R.; Jurečka, P. Refinement of the Sugar–Phosphate Backbone Torsion Beta for AMBER Force Fields Improves the Description of Z- and B-DNA. J. Chem. Theory Comput. 2015, 11, 5723–5736. [Google Scholar] [CrossRef]
- Wang, J.; Wolf, R.M.; Caldwell, J.W.; Kollman, P.A.; Case, D.A. Development and testing of a general amber force field. J. Comput. Chem. 2004, 25, 1157–1174. [Google Scholar] [CrossRef]
- Gmeiner, W.H.; Cui, W.; Sharma, S.; Soto, A.M.; Marky, L.A.; Lown, J.W. Shape-Selective Binding of Geometrically-Constrained bis-Distamycins to a DNA Duplex and a Model Okazaki Fragment of Identical Sequence. Nucleosides Nucleotides Nucleic Acids 2000, 19, 1365–1379. [Google Scholar] [CrossRef] [PubMed]
- Ren, J.; Chaires, J.B. Sequence and Structural Selectivity of Nucleic Acid Binding Ligands. Biochemistry 1999, 38, 16067–16075. [Google Scholar] [CrossRef] [PubMed]
- Yamayoshi, A.; Miyoshi, D.; Zouzumi, Y.-K.; Matsuyama, Y.; Ariyoshi, J.; Shimada, N.; Murakami, A.; Wada, T.; Maruyama, A. Selective and Robust Stabilization of Triplex DNA Structures Using Cationic Comb-type Copolymers. J. Phys. Chem. B 2017, 121, 4015–4022. [Google Scholar] [CrossRef] [PubMed]
- Kan, Y.; Armitage, B.; Schuster, G.B. Selective Stabilization of Triplex DNA by Anthraquinone Sulfonamide Derivatives. Biochemistry 1997, 36, 1461–1466. [Google Scholar] [CrossRef]
- Ren, J.; Chaires, J.B. Preferential Binding of 3,3‘-Diethyloxadicarbocyanine to Triplex DNA. J. Am. Chem. Soc. 1999, 122, 424–425. [Google Scholar] [CrossRef]
- Tam, V.K.; Liu, Q.; Tor, Y. Extended ethidium bromide analogue as a triple helix intercalator: Synthesis, photophysical properties and nucleic acids binding. Chem. Commun. 2006, 25, 2684–2686. [Google Scholar] [CrossRef]
- Latimer, L.J.P.; Payton, N.; Forsyth, G.; Lee, J.S. The binding of analogues of coralyne and related heterocyclics to DNA triplexes. Biochem. Cell Biol. 1995, 73, 11–18. [Google Scholar] [CrossRef]
- Rajewska, M.; Wegrzyn, K.; Konieczny, I. AT-rich region and repeated sequences—The essential elements of replication origins of bacterial replicons. FEMS Microbiol. Rev. 2012, 36, 408–434. [Google Scholar] [CrossRef]
- Vijayanathan, V.; Thomas, T. DNA Nanoparticles and Development of DNA Delivery Vehicles for Gene Therapy. Biochemistry 2002, 41, 14085–14094. [Google Scholar] [CrossRef]
- Kotterman, M.A.; Schaffer, D.V. Engineering adeno-associated viruses for clinical gene therapy. Nat. Rev. Genet. 2014, 15, 445–451. [Google Scholar] [CrossRef]
- Lee, A.; Karcz, A.P.; Akman, R.; Zheng, T.; Kwon, S.; Chou, S.-T.; Sucayan, S.; Tricoli, L.J.; Hustedt, J.M.; Leng, Q.; et al. Direct observation of dynamic mechanical regulation of DNA condensation by environmental stimuli. Angew. Chem. Int. Ed. 2014, 53, 10631–10635. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yevdokimov, Y.M. Double-stranded DNA liquid-crystalline dispersions as biosensing units. Biochem. Soc. Trans. 2000, 28, 77–81. [Google Scholar] [CrossRef] [PubMed]
| Compd | UV/Vis λmax (nm) | ε × 103 /mmol−1 cm2 | Emission (nm) | Stokes Shift × 10−18 J | Φf c |
|---|---|---|---|---|---|
| 1 | 330 | 32.1 | 379 | 4.1 | 0.06 |
| 2 | 285/320 b | 31.7/20.6 | 450 | 1.2/1.5 | 0.13 |
| 3 | 330 | 30.7 | 385 | 3.6 | 0.13 |
| 4 | 343 | 44.9 | 416 | 2.7 | 0.37 |
| 5 | 391 | 77.9 | 497 | 1.9 | 0.69 |
| 6 | 342 | 25.1 | 430 | 2.3 | 0.20 |
| 7 | 325 | 35.4 | 444 | 1.7 | 0.38 |
| 8 | 355 | 36.6 | 430 | 2.6 | 0.54 |
| 9 | 343 | 24.2 | 442 | 2.0 | 0.11 |
| Complex | n1/n2 | log Ks1/Ks2 | ΔrH1,2°/kJ mol−1 | TΔr1,2S°/kJ mol−1 | ΔrG1,2°/kJ mol−1 |
|---|---|---|---|---|---|
| 1-ATT | 0.7/0.1 | 6.5/8.7 | −21.1/−6.8 | 15.7/43.3 | −36.8/−50.1 |
| n | log Ks | ΔrH°/kJ mol−1 | TΔrS°/kJ mol−1 | ΔrG°/kJ mol−1 | |
| 1-rA-dT | 0.1 | 7.8 | −6.3 | 38.4 | −44.7 |
| 6-ATT b | 0.7 | 7.6 | - c | - c | - c |
| 6-dA-rU b | 0.8 | 8.0 | - c | - c | - c |
| ΔTm a | |||
|---|---|---|---|
| Polynucleotide | 1 | 5 | 6 |
| poly dA−poly rU | 5.6 | 4 | 5.5 |
| poly rA−poly rU | 2.2 | 1 | 0 |
| poly rA−poly dT | 1.7 | 1.7 | 1.5 |
| poly dA-poly dT | 19.1 | 2.1 | 0 |
| ΔTm a | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 5 | 6 | |||||||
| cr | 0.1 | 0.2 | 0.3 | 0.1 | 0.2 | 0.3 | 0.025 | 0.05 | 0.1 |
| 31.7/21.9 | 32.5/23.9 | - | 27.3/0 | 30.1/0 | 32.8/0 | 36.7/0 | 42.0/0 | 43.7/0 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zonjić, I.; Tumir, L.-M.; Crnolatac, I.; Šupljika, F.; Racané, L.; Tomić, S.; Radić Stojković, M. Recognition of ATT Triplex and DNA:RNA Hybrid Structures by Benzothiazole Ligands. Biomolecules 2022, 12, 374. https://doi.org/10.3390/biom12030374
Zonjić I, Tumir L-M, Crnolatac I, Šupljika F, Racané L, Tomić S, Radić Stojković M. Recognition of ATT Triplex and DNA:RNA Hybrid Structures by Benzothiazole Ligands. Biomolecules. 2022; 12(3):374. https://doi.org/10.3390/biom12030374
Chicago/Turabian StyleZonjić, Iva, Lidija-Marija Tumir, Ivo Crnolatac, Filip Šupljika, Livio Racané, Sanja Tomić, and Marijana Radić Stojković. 2022. "Recognition of ATT Triplex and DNA:RNA Hybrid Structures by Benzothiazole Ligands" Biomolecules 12, no. 3: 374. https://doi.org/10.3390/biom12030374
APA StyleZonjić, I., Tumir, L.-M., Crnolatac, I., Šupljika, F., Racané, L., Tomić, S., & Radić Stojković, M. (2022). Recognition of ATT Triplex and DNA:RNA Hybrid Structures by Benzothiazole Ligands. Biomolecules, 12(3), 374. https://doi.org/10.3390/biom12030374









