Looking for Hidden Enemies of Metabarcoding: Species Composition, Habitat and Management Can Strongly Influence DNA Extraction while Examining Grassland Communities
Abstract
1. Introduction
1.1. Assessing Belowground Biodiversity
1.2. Metabarcoding as a Useful Tool for Identifying Belowground Species Richness
1.3. Factors Affecting the Quality and Quantity of Extracted DNA (Including Environmental and Biochemical)
2. Materials and Methods
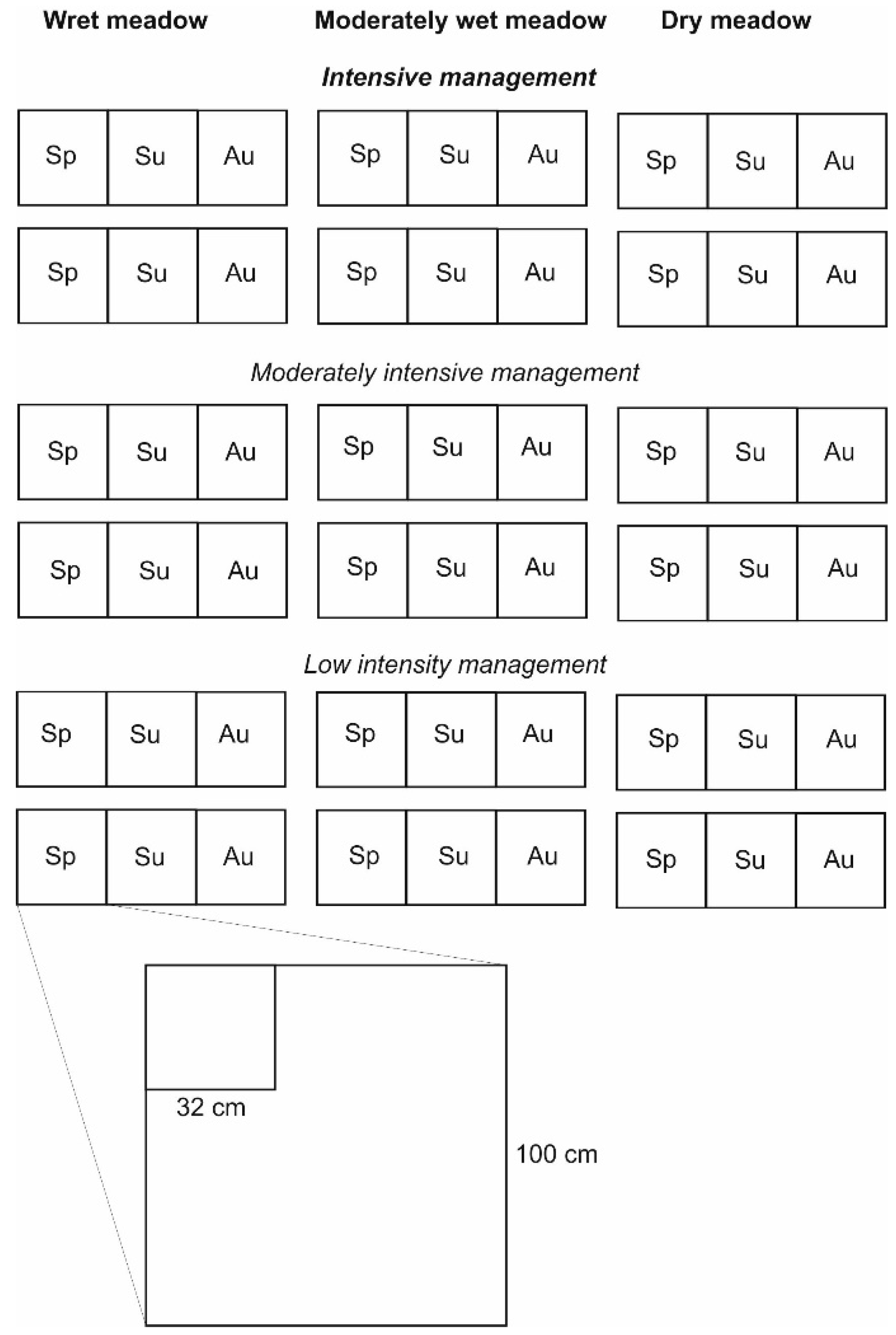
2.1. Study Area and Sampling Design
2.2. Collection of Floristic and Environmental Data
2.3. Extraction, Isolation and Determination of DNA Quality and Quantity
2.3.1. Soil and Plant Samples for DNA Extraction
2.3.2. DNA Extraction
- (i)
- DNA extraction
- (ii)
- DNA purification
2.3.3. Quantity and Quality Measurement of Plant DNA
2.3.4. PCR Amplification of DNA
2.4. Statistical Analysis
3. Results
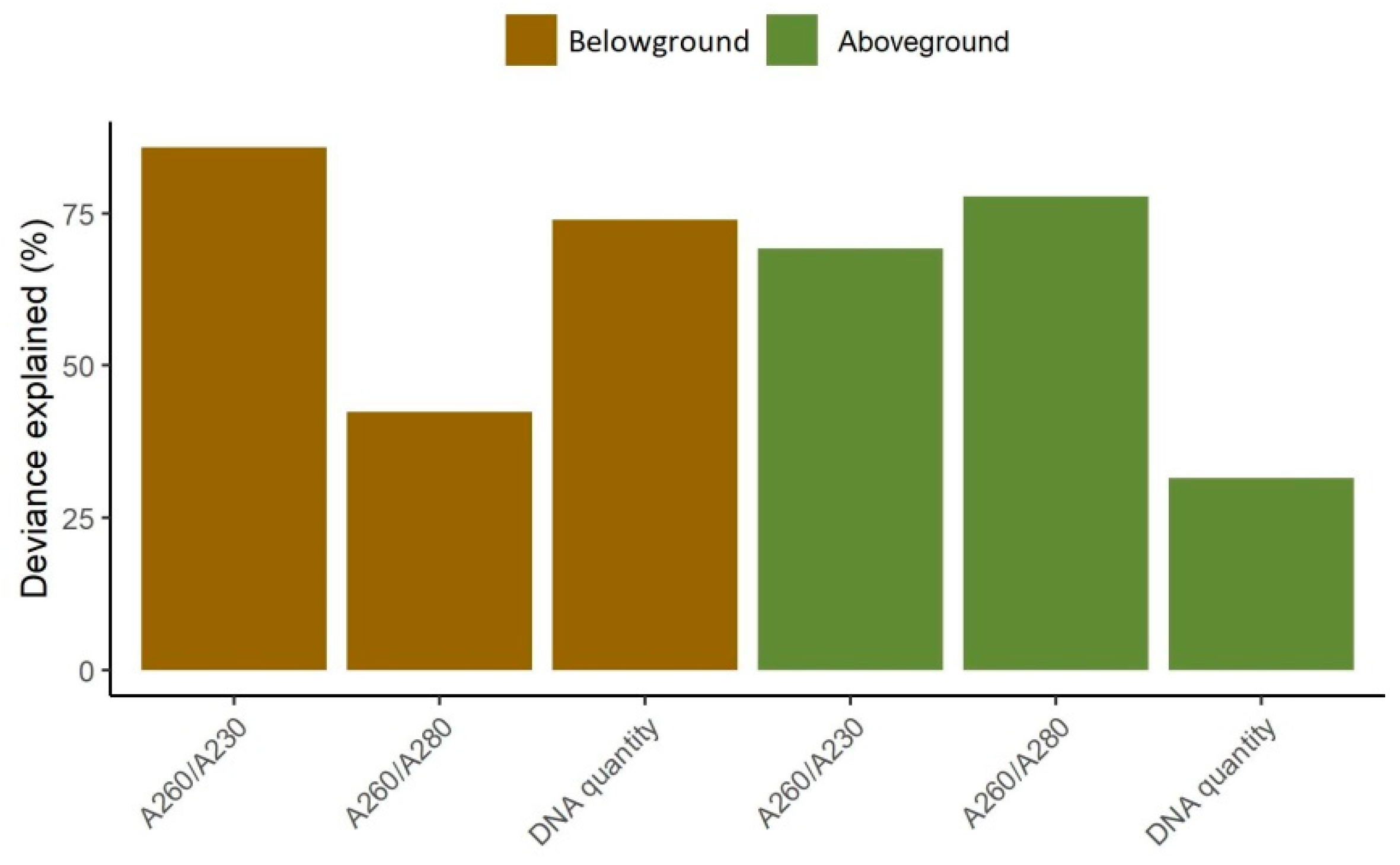
3.1. Model Performance
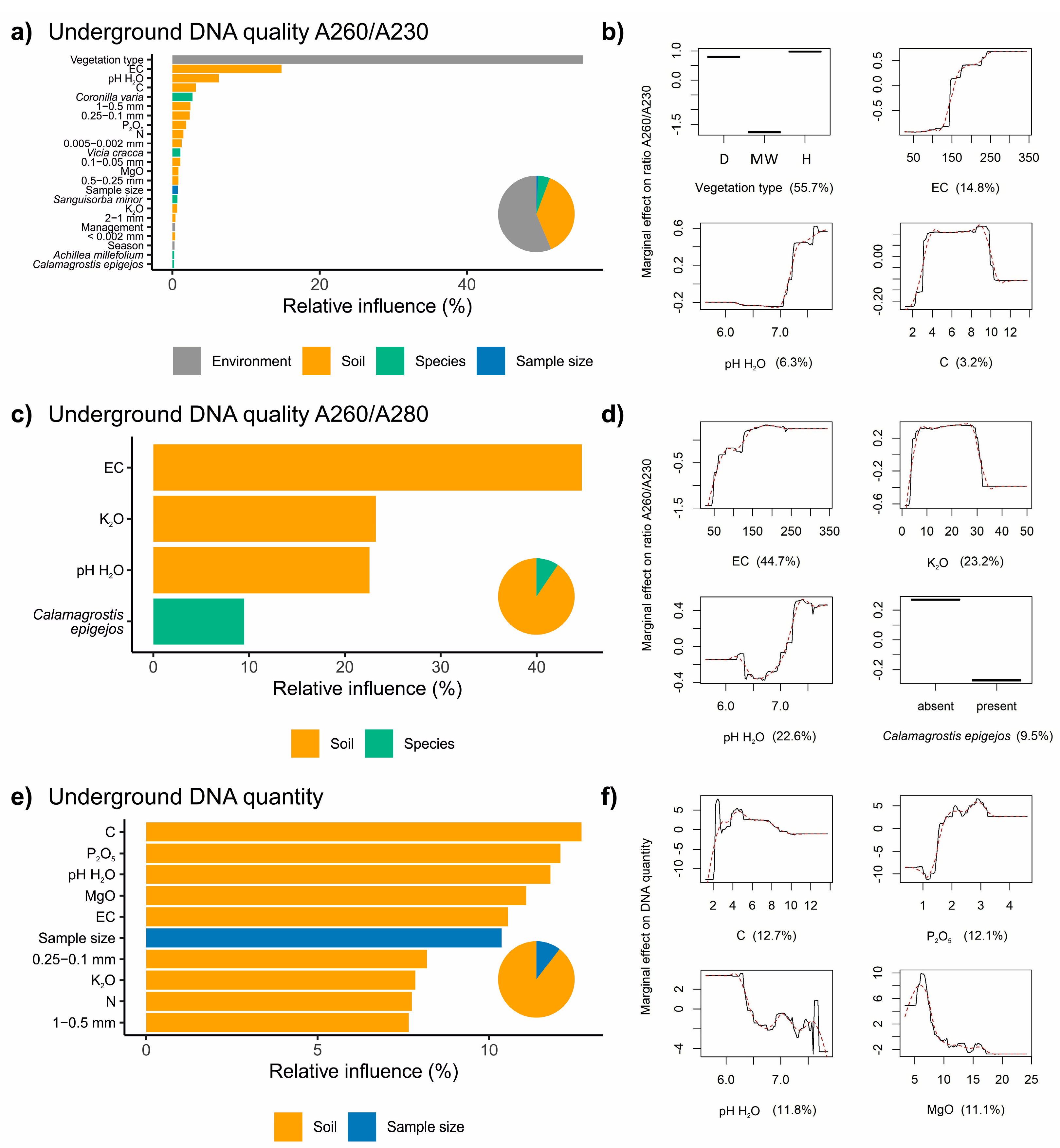
3.2. Factors Influencing the Extraction of DNA from Belowground Biomass
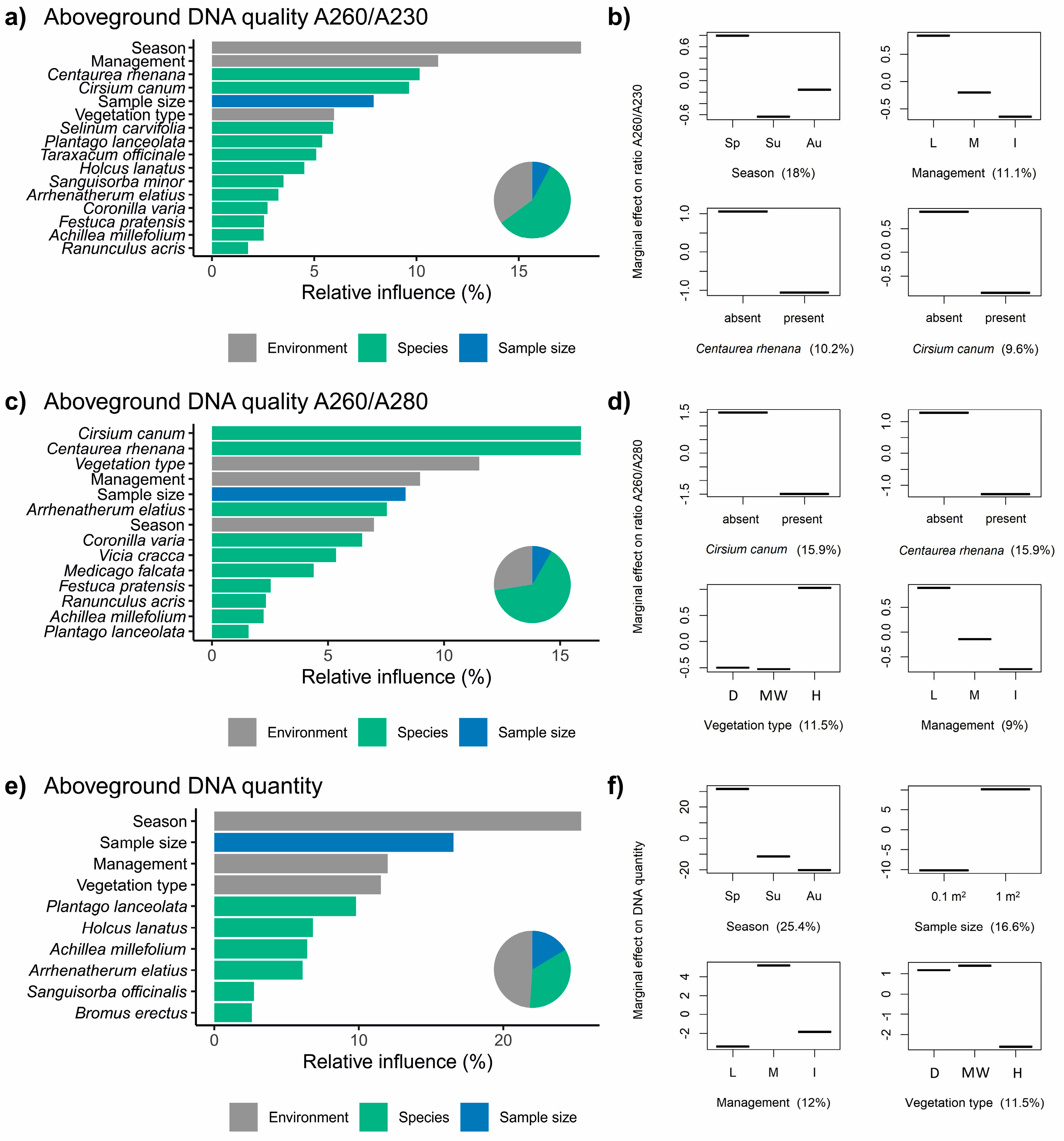
3.3. Factors Influencing the Extraction of DNA from Aboveground Biomass
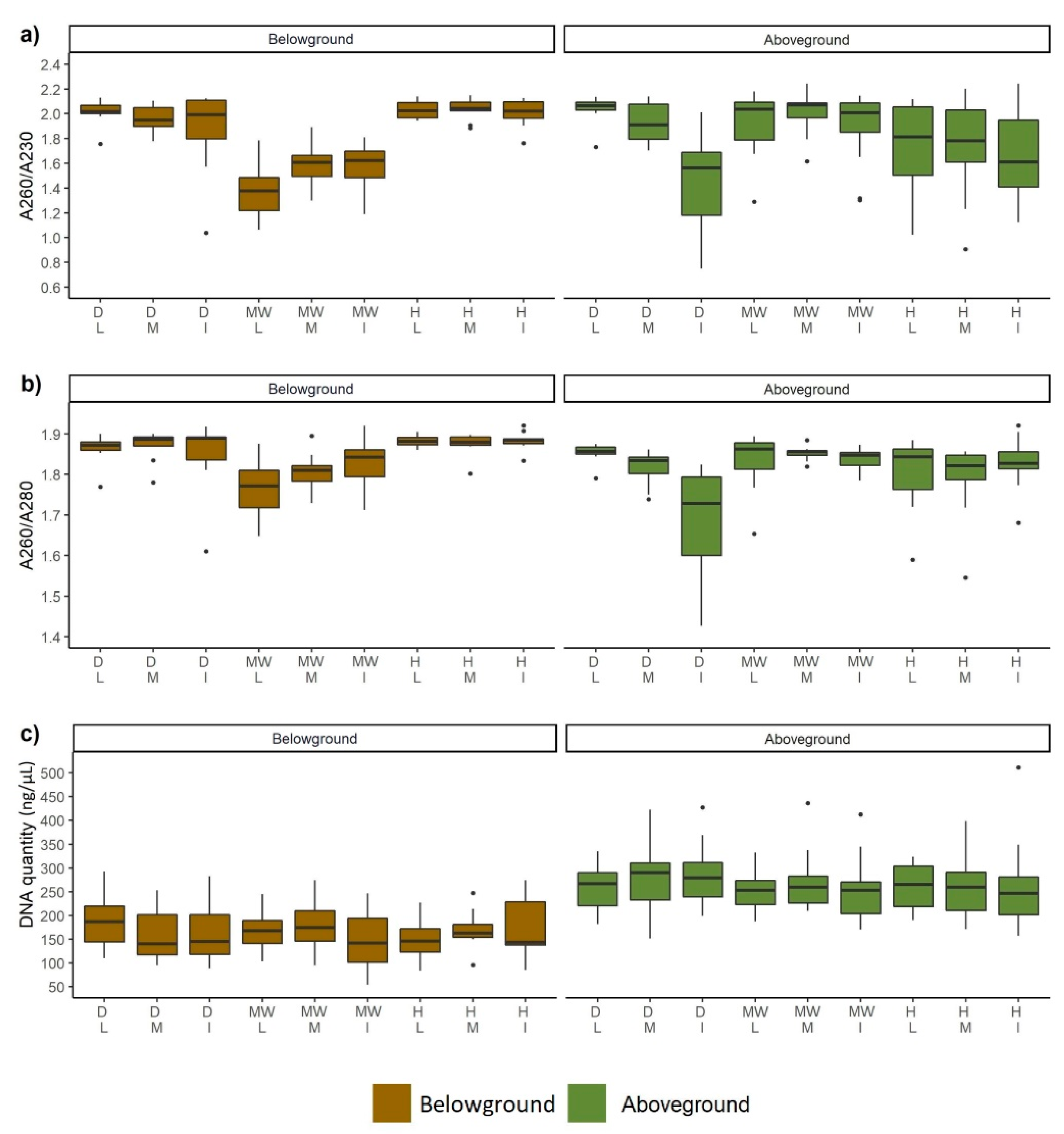
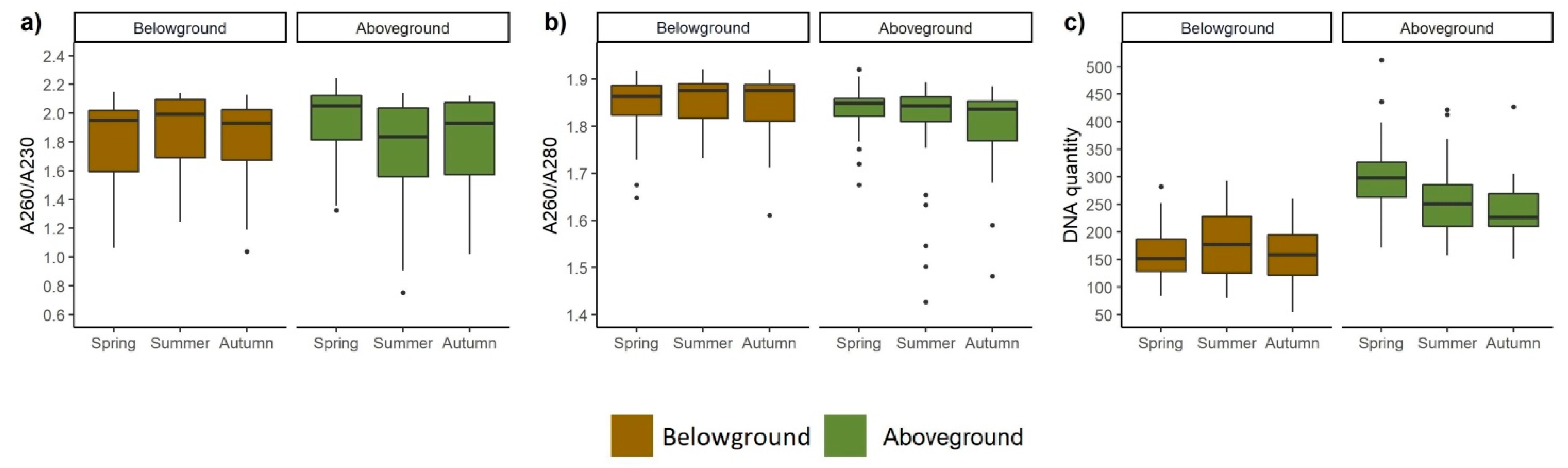
3.4. DNA Quality and Quantity
4. Discussion
4.1. Plant Chemical Manufacture (Species Composition) Plays a Dominant Role in the Extraction Effectiveness
4.2. Soil Properties also Play an Important Role in the Extraction Effectiveness
4.3. Aboveground, the Species Composition Is the Most Influential
4.4. Season and Stress Play an Important Role in Extraction of High Quality DNA from the Aboveground Biomass
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chytrý, M.; Chiarucci, A.; Pillar, V.D.; Pärtel, M. Plant communities: Their conservation assessment and surveys across continents and in the tropics. Appl. Veg. Sci. 2015, 18, 1–2. [Google Scholar] [CrossRef][Green Version]
- Dengler, J.; Boch, S. Sampling-design effects on properties of species-area relationships—A case study from Estonian dry grassland communities. Folia Geobot. 2008, 43, 289–304. [Google Scholar] [CrossRef]
- HilleRisLambers, J.; Adler, P.B.; Harpole, W.S.; Levine, J.M.; Mayfield, M.M. Rethinking Community Assembly through the Lens of Coexistence Theory. Annu. Rev. Ecol. Evol. Syst. 2012, 43, 227–248. [Google Scholar] [CrossRef]
- Wagner, V.; Chytrý, M.; Zelený, D.; von Wehrden, H.; Brinkert, A.; Danihelka, J.; Hölzel, N.; Jansen, F.; Kamp, J.; Lustyk, P.; et al. Regional differences in soil pH niche among dry grassland plants in Eurasia. Oikos 2017, 126, 660–670. [Google Scholar] [CrossRef]
- Chytrý, M.; Lososová, Z.; Horsák, M.; Uher, B.; Čejka, T.; Danihelka, J.; Fajmon, K.; Hájek, O.; Juřičková, L.; Kintrová, K.; et al. Dispersal limitation is stronger in communities of microorganisms than macroorganisms across Central European cities. J. Biogeogr. 2012, 39, 1101–1111. [Google Scholar] [CrossRef]
- Träger, S.; Öpik, M.; Vasar, M.; Wilson, S.D. Belowground plant parts are crucial for comprehensively estimating total plant richness in herbaceous and woody habitats. Ecology 2019, 100, e02575. [Google Scholar] [CrossRef] [PubMed]
- Klimešová, J.; Klimeš, L. Bud banks and their role in vegetative regeneration—A literature review and proposal for simple classification and assessment. Perspect. Plant Ecol. Evol. Syst. 2007, 8, 115–129. [Google Scholar] [CrossRef]
- Reintal, M.; Tali, K.; Haldna, M.; Kull, T. Habitat preferences as related to the prolonged dormancy of perennial herbs and ferns. Plant Ecol. 2010, 210, 111–123. [Google Scholar] [CrossRef]
- Schenk, H.J.; Jackson, R.B. The global biography of roots. Ecol. Monogr. 2002, 72, 311–328. [Google Scholar] [CrossRef]
- Zobel, M.; Moora, M.; Herben, T. Clonal mobility and its implications for spatio-temporal patterns of plant communities: What do we need to know next? Oikos 2010, 119, 802–806. [Google Scholar] [CrossRef]
- Pärtel, M.; Hiiesalu, I.; Öpik, M.; Wilson, S.D. Below-ground plant species richness: New insights from DNA-based methods. Funct. Ecol. 2012, 26, 775–782. [Google Scholar] [CrossRef]
- Dexter, K.G.; Pennington, T.D.; Cunningham, C.W. Using DNA to assess errors in tropical tree identifications: How often are ecologists wrong and when does it matter? Ecol. Monogr. 2010, 80, 267–286. [Google Scholar] [CrossRef]
- Lecroq, B.; Lejzerowicz, F.; Bachar, D.; Christen, R.; Esling, P.; Baerlocher, L.; Østerås, M.; Farinelli, L.; Pawlowski, J. Ultra-deep sequencing of foraminiferal microbarcodes unveils hidden richness of early monothalamous lineages in deep-sea sediments. Proc. Natl. Acad. Sci. USA 2011, 108, 13177–13182. [Google Scholar] [CrossRef]
- Shefferson, R.P. The evolutionary ecology of vegetative dormancy in mature herbaceous perennial plants. J. Ecol. 2009, 97, 1000–1009. [Google Scholar] [CrossRef]
- Pärtel, M.; Zobel, M. Small-scale dynamics and species richness in successional alvar plant communities. Ecography 1995, 18, 83–90. [Google Scholar] [CrossRef]
- Wilson, S.D.; Tilman, D. Quadratic variation in old-field species richness along gradients of disturbance and nitrogen. Ecology 2002, 83, 492–504. [Google Scholar] [CrossRef]
- Price, J.N.; Hiiesalu, I.; Gerhold, P.; Pärtel, M. Small-scale grassland assembly patterns differ above and below the soil surface. Ecology 2012, 93, 1290–1296. [Google Scholar] [CrossRef]
- Wilson, S.D. Competition, resources, and vegetation during 10 years in native grassland. Ecology 2007, 88, 2951–2958. [Google Scholar] [CrossRef]
- Hiiesalu, I.; Öpik, M.; Metsis, M.; Lilje, L.; Davison, J.; Vasar, M.; Moora, M.; Zobel, M.; Wilson, S.D.; Pärtel, M. Plant species richness belowground: Higher richness and new patterns revealed by next-generation sequencing. Mol. Ecol. 2012, 21, 2004–2016. [Google Scholar] [CrossRef]
- Herben, T.; Suda, J.; Klimešová, J. Polyploid species rely on vegetative reproduction more than diploids: A re-examination of the old hypothesis. Ann. Bot. 2017, 120, 341–349. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Lamb, E.G.; Piper, C.L.; Siciliano, S.D. Plant belowground diversity and species segregation by depth in a semi-arid grassland. Ecoscience 2017, 25, 1–7. [Google Scholar] [CrossRef]
- Parducci, L.; Matetovici, I.; Fontana, S.L.; Bennett, K.D.; Suyama, Y.; Haile, J.; Kjær, K.H.; Larsen, N.K.; Drouzas, A.D.; Willerslev, E. Molecular- and pollen-based vegetation analysis in lake sediments from central Scandinavia. Mol. Ecol. 2013, 22, 3511–3524. [Google Scholar] [CrossRef] [PubMed]
- De Barba, M.; Miquel, C.; Boyer, F.; Mercier, C.; Rioux, D.; Coissac, E.; Taberlet, P. DNA metabarcoding multiplexing and validation of data accuracy for diet assessment: Application to omnivorous diet. Mol. Ecol. Resour. 2014, 14, 306–323. [Google Scholar] [CrossRef]
- Alsos, I.G.; Lammers, Y.; Yoccoz, N.G.; Jørgensen, T.; Sjögren, P.; Gielly, L.; Edwards, M.E. Plant DNA metabarcoding of lake sediments: How does it represent the contemporary vegetation. PLoS ONE 2018, 13, e0195403. [Google Scholar] [CrossRef]
- Coghlan, S.A.; Shafer, A.B.A.; Freeland, J.R. Development of an environmental DNA metabarcoding assay for aquatic vascular plant communities. Environ. DNA 2020, 00, 1–16. [Google Scholar] [CrossRef]
- Banchi, E.; Ametrano, C.G.; Tordoni, E.; Stanković, D.; Ongaro, S.; Tretiach, M.; Pallavicini, A.; Muggia, L.; Verardo, P.; Tassan, F.; et al. Environmental DNA assessment of airborne plant and fungal seasonal diversity. Sci. Total Environ. 2020, 738, 140249. [Google Scholar] [CrossRef] [PubMed]
- Cheng, T.; Xu, C.; Lei, L.; Li, C.; Zhang, Y.; Zhou, S. Barcoding the kingdom Plantae: New PCR primers for ITS regions of plants with improved universality and specificity. Mol. Ecol. Resour. 2016, 16, 138–149. [Google Scholar] [CrossRef]
- Fahner, N.A.; Shokralla, S.; Baird, D.J.; Hajibabaei, M. Large-scale monitoring of plants through environmental DNA metabarcoding of soil: Recovery, resolution, and annotation of four DNA markers. PLoS ONE 2016, 11, 1–16. [Google Scholar] [CrossRef]
- Soininen, E.M.; Valentini, A.; Coissac, E.; Miquel, C.; Gielly, L.; Brochmann, C.; Brysting, A.K.; Sønstebø, J.H.; Ims, R.A.; Yoccoz, N.G.; et al. Analysing diet of small herbivores: The efficiency of DNA barcoding coupled with high-throughput pyrosequencing for deciphering the composition of complex plant mixtures. Front. Zool. 2009, 6, 16. [Google Scholar] [CrossRef]
- Pegard, A.; Miquel, C.; Valentini, A.; Coissac, E.; Bouvier, F.; François, D.; Taberlet, P.; Engel, E.; Pompanon, F. Universal DNA-based methods for assessing the diet of grazing livestock and wildlife from feces. J. Agric. Food Chem. 2009, 57, 5700–5706. [Google Scholar] [CrossRef] [PubMed]
- Valentini, A.; Miquel, C.; Nawaz, M.A.; Bellemain, E.; Coissac, E.; Pompanon, F.; Gielly, L.; Cruaud, C.; Nascetti, G.; Wincker, P.; et al. New perspectives in diet analysis based on DNA barcoding and parallel pyrosequencing: The trnL approach. Mol. Ecol. Resour. 2009, 9, 51–60. [Google Scholar] [CrossRef] [PubMed]
- Sønstebø, J.H.; Gielly, L.; Brysting, A.K.; Elven, R.; Edwards, M.; Haile, J.; Willerslev, E.; Coissac, E.; Rioux, D.; Sannier, J.; et al. Using next-generation sequencing for molecular reconstruction of past Arctic vegetation and climate. Mol. Ecol. Resour. 2010, 10, 1009–1018. [Google Scholar] [CrossRef]
- Taberlet, P.; Coissac, E.; Hajibabaei, M.; Rieseberg, L.H. Environmental DNA. Mol. Ecol. 2012, 21, 1789–1793. [Google Scholar] [CrossRef] [PubMed]
- Bertrand, H.; Poly, F.; Van, V.T.; Lombard, N.; Nalin, R.; Vogel, T.M.; Simonet, P. High molecular weight DNA recovery from soils prerequisite for biotechnological metagenomic library construction. J. Microbiol. Methods 2005, 62, 1–11. [Google Scholar] [CrossRef]
- Lakay, F.M.; Botha, A.; Prior, B.A. Comparative analysis of environmental DNA extraction and purification methods from different humic acid-rich soils. J. Appl. Microbiol. 2007, 102, 265–273. [Google Scholar] [CrossRef] [PubMed]
- Delmont, T.O.; Robe, P.; Clark, I.; Simonet, P.; Vogel, T.M. Metagenomic comparison of direct and indirect soil DNA extraction approaches. J. Microbiol. Methods 2011, 86, 397–400. [Google Scholar] [CrossRef]
- Kotchoni, S.O.; Gachomo, E.W.; Jimenez-Lopez, J.C. A plant cocktail amenable for PCR-based genetic analysis in Arabidopsis thaliana. Mol. Biol. Rep. 2011, 38, 5281–5284. [Google Scholar] [CrossRef]
- Sharma, A.D.; Gill, P.K.; Singh, P. DNA isolation from dry and fresh samples of polysaccharide-rich plants. Plant Mol. Biol. Report. 2002, 20, 415a–415f. [Google Scholar] [CrossRef]
- Shepherd, L.D.; McLay, T.G.B. Two micro-scale protocols for the isolation of DNA from polysaccharide-rich plant tissue. J. Plant Res. 2011, 124, 311–314. [Google Scholar] [CrossRef] [PubMed]
- Afshar-Mohammadian, M.; Rezadoost, M.H.; Fallah, S.F. Comparative analysis and innovation of a simple and rapid method for high-quality RNA and DNA extraction of kiwifruit. MethodsX 2018, 5, 352–361. [Google Scholar] [CrossRef] [PubMed]
- Arruda, S.R.; Pereira, D.G.; Silva-Castro, M.M.; Brito, M.G.; Waldschmidt, A.M. An optimized protocol for DNA extraction in plants with a high content of secondary metabolites, based on leaves of Mimosa tenuiflora (Willd.) Poir. (Leguminosae). Genet. Mol. Res. 2017, 16, gmr16039063. [Google Scholar] [CrossRef]
- Lear, G.; Dickie, I.; Banks, J.; Boyer, S.; Buckley, H.; Buckley, T.; Cruickshank, R.; Dopheide, A.; Handley, K.; Hermans, S.; et al. Methods for the extraction, storage, amplification and sequencing of DNA from environmental samples. N. Z. J. Ecol. 2018, 42, 10. [Google Scholar] [CrossRef]
- Kreader, C.A. Relief of amplification inhibition in PCR with bovine serum albumin or T4 gene 32 protein. Appl. Environ. Microbiol. 1996, 62, 1102–1106. [Google Scholar] [CrossRef]
- Schrader, C.; Schielke, A.; Ellerbroek, L.; Johne, R. PCR inhibitors—Occurrence, properties and removal. J. Appl. Microbiol. 2012, 113, 1014–1026. [Google Scholar] [CrossRef]
- Nowak, A.; Spałek, K. Ochrona szaty roślinnej Śląska Opolskiego; Wydawnictwo Uniwersytetu Opolskiego: Opole, Poland, 2004; p. 391. [Google Scholar]
- Jürgens, N.; Schmiedel, U.; Haarmeyer, D.H.; Dengler, J.; Finckh, M.; Goetze, D.; Gröngröft, A.; Hahn, K.; Koulibaly, A.; Luther-Mosebach, J.; et al. The BIOTA Biodiversity Observatories in Africa-a standardized framework for large-scale environmental monitoring. Environ. Monit. Assess. 2012, 184, 655–678. [Google Scholar] [CrossRef] [PubMed]
- Dengler, J. A flexible multi-scale approach for standardised recording of plant species richness patterns. Ecol. Indic. 2009, 9, 1169–1178. [Google Scholar] [CrossRef]
- Woś, A. Klimat Polski; Wydawnictwo Naukowe PWN: Warszawa, Poland, 1999; p. 301. [Google Scholar]
- Rutkowski, L. Klucz do oznaczania roślin naczyniowych Polski niżowej; Wydawnictwo Naukowe PWN: Warszawa, Poland, 2007; p. 814. [Google Scholar]
- Mirek, Z.; Piękoś-Mirkowa, H.; Zając, A.; Zając, M. Flowering plants and pteridiophytes in Poland. A Checklist. Biodiversity of Poland. Vol. 1; Instytut Botaniki im. W. Szafera, PAN: Kraków, Poland, 2002; p. 440. [Google Scholar]
- Elith, J.; Leathwick, J.R.; Hastie, T. A working guide to boosted regression trees. J. Anim. Ecol. 2008, 77, 802–813. [Google Scholar] [CrossRef] [PubMed]
- Greenwell, B.; Boehmke, B.; Cunningham, J. GBM Developers gbm: Generalized Boosted Regression Models. R Package Version 2.1.5. 2019. Available online: https://www.researchgate.net/publication/240311335_Gbm_Generalized_Boosted_Regression_Models_R_Package_15 (accessed on 18 February 2021).
- Hijmans, R.J.; Phillips, S.; Leathwick, J.; Elith, J. dismo: Species Distribution Modeling. R package version 1.1-4. 2017. Available online: http://www2.uaem.mx/r-mirror/web/packages/dismo/vignettes/sdm.pdf (accessed on 18 February 2021).
- R Core Team R: A Language and Environment for Statistical Computing. 2020. Available online: http://softlibre.unizar.es/manuales/aplicaciones/r/fullrefman.pdf (accessed on 18 February 2021).
- Hastie, T.; Tibshirani, R.; Friedman, J. The Elements of Statistical Learning. Data Mining, Inference, and Prediction, 2nd ed.; Springer: New York, NY, USA, 2009. [Google Scholar]
- Lucas, M.S.; Carvalho, C.S.; Hypolito, G.B.; Côrtes, M.C. Optimized protocol to isolate high quality genomic DNA from different tissues of a palm species. Hoehnea 2019, 46, e942018. [Google Scholar] [CrossRef]
- Northup, R.R.; Dahlgren, R.A.; McColl, J.G. Polyphenols as regulators of plant-litter-soil interactions in northern California’s pygmy forest: A positive feedback? Biogeochemistry 1998, 42, 189–220. [Google Scholar] [CrossRef]
- Tiley, G.E.D. Biological Flora of the British Isles: Cirsium arvense (L.) Scop. J. Ecol. 2010, 98, 938–983. [Google Scholar] [CrossRef]
- Tolrà, R.P.; Poschenrieder, C.; Luppi, B.; Barceló, J. Aluminium-induced changes in the profiles of both organic acids and phenolic substances underlie Al tolerance in Rumex acetosa L. Environ. Exp. Bot. 2005, 54, 231–238. [Google Scholar] [CrossRef]
- Díaz, K.; Espinoza, L.; Madrid, A.; Pizarro, L.; Chamy, R. Isolation and identification of compounds from bioactive extracts of Taraxacum officinale Weber ex F. H. Wigg. (Dandelion) as a potential source of antibacterial agents. Evidence-based Complement. Altern. Med. 2018, 2018, 2706417. [Google Scholar]
- Bar-On, Y.M.; Phillips, R.; Milo, R. The biomass distribution on Earth. Proc. Natl. Acad. Sci. USA 2018, 115, 6506–6511. [Google Scholar] [CrossRef]
- Somers, E.; Vanderleyden, J.; Srinivasan, M. Rhizosphere bacterial signalling: A love parade beneath our feet. Crit. Rev. Microbiol. 2004, 30, 205–240. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.C.; Yokosho, K.; Kashino, M.; Zhao, F.J.; Yamaji, N.; Ma, J.F. Adaptation to acidic soil is achieved by increased numbers of cis-acting elements regulating ALMT1 expression in Holcus lanatus. Plant J. 2013, 76, 10–23. [Google Scholar]
- Gloser, V. Seasonal changes of nitrogen storage compounds in a rhizomatous grass Calamagrostis epigeios. Biol. Plant. 2002, 45, 563–568. [Google Scholar] [CrossRef]
- Dušek, J. Seasonal dynamic of nonstructutral saccharides in a rhizomatous grass Calamagrostis epigeios. Biol. Plant. 2002, 45, 383–387. [Google Scholar] [CrossRef]
- Dehpour, A.A.; Eslami, B.; Rezaie, S.; Hashemian, S.F.; Shafie, F.; Kiaie, M. Chemical composition of essential oil and in vitro antibacterial and anticancer activity of the hydroalcolic extract from Coronilla varia. Int. J. Biol. Biomol. Agric. Food Biotechnol. Eng. 2014, 8, 1414–1417. [Google Scholar]
- Matasyoh, L.G.; Wachira, F.N.; Kinyua, M.G.; Thairu Muigai, A.W.; Mukiama, T.K. Leaf storage conditions and genomic DNA isolation efficiency in Ocimum gratissimum L. from Kenya. Afr. J. Biotechnol. 2008, 7, 557–564. [Google Scholar]
- Khanuja, S.; Shasany, A.; Darokar, M.P.; Kumar, S. Rapid Isolation of DNA from Dry and Fresh Samples of Plants Producing Large Amounts of Secondary Metabolites and Essential Oils. Plant Mol. Biol. Report. 1999, 17, 74. [Google Scholar] [CrossRef]
- Al-Snafi, A.E. The pharmacological and toxicological effects of Coronilla varia and Coronilla scorpioides: A review. Pharm. Chem. J. 2016, 3, 105–114. [Google Scholar]
- Pierret, A.; Moran, C.J.; Doussan, C. Conventional detection methodology is limiting our ability to understand the roles and functions of fine roots. New Phytol. 2005, 166, 967–980. [Google Scholar] [CrossRef] [PubMed]
- Schulten, H.R.; Leinweber, P. Characterization of humic and soil particles by analytical pyrolysis and computer modeling. J. Anal. Appl. Pyrolysis 1996, 38, 1–53. [Google Scholar] [CrossRef]
- Chiodi, C.; Moro, M.; Squartini, A.; Concheri, G.; Occhi, F.; Fornasier, F.; Cagnin, M.; Bertoldo, G.; Broccanello, C.; Stevanato, P. High-Throughput Isolation of Nucleic Acids from Soil. Soil Syst. 2019, 4, 3. [Google Scholar] [CrossRef]
- Nardi, S.; Pizzeghello, D.; Muscolo, A.; Vianello, A. Physiological effects of humic substances on higher plants. Soil Biol. Biochem. 2002, 34, 1527–1536. [Google Scholar] [CrossRef]
- Zhou, J.; Bruns, M.A.; Tiedje, J.M. DNA recovery from soils of diverse composition. Appl. Environ. Microbiol. 1996, 62, 316–322. [Google Scholar] [CrossRef]
- Trevisan, S.; Francioso, O.; Quaggiotti, S.; Nardi, S. Humic substances biological activity at the plant-soil interface: From environmental aspects to molecular factors. Plant Signal. Behav. 2010, 5, 635–643. [Google Scholar] [CrossRef] [PubMed]
- Corwin, D.L.; Lesch, S.M. Apparent soil electrical conductivity measurements in agriculture. Comput. Electron. Agric. 2005, 46, 11–43. [Google Scholar] [CrossRef]
- Hebda, L.M.; Foran, D.R. Assessing the utility of soil DNA extraction kits for increasing DNA yields and eliminating PCR inhibitors from buried skeletal remains. J. Forensic Sci. 2015, 60, 1322–1330. [Google Scholar] [CrossRef]
- Saeki, K.; Sakai, M.; Wada, S.I. DNA adsorption on synthetic and natural allophanes. Appl. Clay Sci. 2010, 50, 493–497. [Google Scholar] [CrossRef]
- Shen, Y.; Kim, H.; Tong, M.; Li, Q. Influence of solution chemistry on the deposition and detachment kinetics of RNA on silica surfaces. Colloids Surfaces B Biointerfaces 2011, 82, 443–449. [Google Scholar] [CrossRef]
- Kasterova, E.; Zibareva, L.; Revushkin, A. Secondary metabolites of some Siberian species of plants tribe Cynareae (Asteraceae). S. Afr. J. Bot. 2019, 125, 24–29. [Google Scholar] [CrossRef]
- Jordon-Thaden, I.E.; Louda, S.M. Chemistry of Cirsium and Carduus: A role in ecological risk assessment for biological control of weeds? Biochem. Syst. Ecol. 2003, 31, 1353–1396. [Google Scholar] [CrossRef]
- Kozyra, M.; Biernasiuk, A.; Malm, A.; Chowaniec, M. Chemical compositions and antibacterial activity of extracts obtained from the inflorescences of Cirsium canum (L.) all. Nat. Prod. Res. 2015, 29, 2059–2063. [Google Scholar] [CrossRef]
- Olson, B.E.; Kelsey, R.G. Effect of Centaurea maculosa on sheep rumen microbial activity and mass in vitro. J. Chem. Ecol. 1997, 23, 1131–1144. [Google Scholar] [CrossRef]
- Tharayil, N. To survive or to slay: Resource-foraging role of metabolites implicated in allelopathy. Plant Signal. Behav. 2009, 4, 580–583. [Google Scholar] [CrossRef] [PubMed]
- Gfeller, V.; Huber, M.; Förster, C.; Huang, W.; Köllner, T.G.; Erb, M. Root volatiles in plant–plant interactions I: High root sesquiterpene release is associated with increased germination and growth of plant neighbours. Plant Cell Environ. 2019, 42, 1950–1963. [Google Scholar] [CrossRef]
- Molloy, L.F.; Richards, E.L. Complexing of calcium and magnesium by the organic constituents of yorkshire fog (Holcus lanatus): I—The organic acids, lignin and cell wall polysaccharides of Yorkshire Fog. J. Sci. Food Agric. 1971, 22, 393–396. [Google Scholar] [CrossRef]
- Varma, A.; Padh, H.; Shrivastava, N. Plant genomic DNA isolation: An art or a science. Biotechnol. J. 2007, 2, 386–392. [Google Scholar] [CrossRef] [PubMed]
- Murray, M.G.; Thompson, W.F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 1980, 8, 4321–4326. [Google Scholar] [CrossRef]
- Doyle, J.J.; Doyle, J.L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 1987, 19, 11–15. [Google Scholar]
- Williams, C.E.; Ronald, P.C. PCR template-DNA isolated quickly from monocot and dicot leaves without tissue homogenization. Nucleic Acids Res. 1994, 22, 1917–1918. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Freschet, G.T.; Roumet, C. Sampling roots to capture plant and soil functions. Funct. Ecol. 2017, 31, 1506–1518. [Google Scholar] [CrossRef]
- Dietze, M.C.; Sala, A.; Carbone, M.S.; Czimczik, C.I.; Mantooth, J.A.; Richardson, A.D.; Vargas, R. Nonstructural carbon in woody plants. Annu. Rev. Plant Biol. 2014, 65, 667–687. [Google Scholar] [CrossRef]
- Huang, W.; Bont, Z.; Hervé, M.R.; Robert, C.A.M.; Erb, M. Impact of seasonal and temperature-dependent variation in root defense metabolites on herbivore preference in Taraxacum officinale. J. Chem. Ecol. 2020, 46, 63–75. [Google Scholar] [CrossRef] [PubMed]
- Mithöfer, A.; Boland, W. Plant Defense Against Herbivores: Chemical Aspects. Annu. Rev. Plant Biol. 2012, 63, 431–450. [Google Scholar] [CrossRef]
- War, A.R.; Paulraj, M.G.; Ahmad, T.; Buhroo, A.A.; Hussain, B.; Ignacimuthu, S.; Sharma, H.C. Mechanisms of plant defense against insect herbivores. Plant Signal. Behav. 2012, 7, 1306–1320. [Google Scholar] [CrossRef] [PubMed]
- Huitu, O.; Forbes, K.M.; Helander, M.; Julkunen-Tiitto, R.; Lambin, X.; Saikkonen, K.; Stuart, P.; Sulkama, S.; Hartley, S. Silicon, endophytes and secondary metabolites as grass defenses against mammalian herbivores. Front. Plant Sci. 2014, 5, 478. [Google Scholar] [CrossRef]
- Isah, T. Stress and defense responses in plant secondary metabolites production. Biol. Res. 2019, 52, 39. [Google Scholar] [CrossRef]
- Niemeyer, H.M. Hydroxamic acids (4-hydroxy-1,4-benzoxazin-3-ones), defence chemicals in the gramineae. Phytochemistry 1988, 27, 3349–3358. [Google Scholar] [CrossRef]
- Bernays, E.A.; Driver, G.C.; Bilgener, M. Herbivores and plant tannins. Adv. Ecol. Res. 1989, 19, 263–302. [Google Scholar]
- Jones, D.A. Cyanogenesis in animal-plant interactions. Ciba Found. Symp. 1988, 140, 151–170. [Google Scholar] [PubMed]
- Taiz, L.; Zeiger, E. Plant Physiology, 5th ed.; Sinauer Associates, Inc.: Sunderland, MA, USA, 2010. [Google Scholar]
- Nabity, P.D.; Zavala, J.A.; Delucia, E.H. Herbivore induction of jasmonic acid and chemical defences reduce photosynthesis in Nicotiana attenuata. J. Exp. Bot. 2013, 64, 685–694. [Google Scholar] [CrossRef]
- Olivoto, T.; Nardino, M.; Carvalho, I.R.; Follmann, D.N.; Szareski, V.J.; Ferrari, M.; de Pelegrin, A.J.; de Souza, V.Q. Plant secondary metabolites and its dynamical systems of induction in response to environmental factors: A review. Afr. J. Agric. Res. 2017, 12, 71–84. [Google Scholar]
- Veitch, G.E.; Boyer, A.; Ley, S.V. The azadirachtin story. Angew. Chem. Int. Ed. 2008, 47, 9402–9429. [Google Scholar] [CrossRef]
- Adamczyk, B.; Kitunen, V.; Smolander, A. Response of soil C and N transformations to condensed tannins and different organic N-condensed tannin complexes. Appl. Soil Ecol. 2013, 64, 163–170. [Google Scholar] [CrossRef]
- Chen, Q.; Lu, X.; Guo, X.; Liu, J.; Liu, Y.; Guo, Q.; Tang, Z. The specific responses to mechanical wound in leaves and roots of Catharanthus roseus seedlings by metabolomics. J. Plant Interact. 2018, 13, 450–460. [Google Scholar] [CrossRef]
- Hollingsworth, P.M.; Forrest, L.L.; Spouge, J.L.; Hajibabaei, M.; Ratnasingham, S.; van der Bank, M.; Chase, M.W.; Cowan, R.S.; Erickson, D.L.; Fazekas, A.J.; et al. A DNA barcode for land plants. Proc. Natl. Acad. Sci. USA 2009, 106, 12794–12797. [Google Scholar]
- Taberlet, P.; Gielly, L.; Pautou, G.; Bouvet, J. Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol. Biol. 1991, 17, 1105–1109. [Google Scholar] [CrossRef]
| Predictor Category | Predictor | Type | Description |
|---|---|---|---|
| Environment | Vegetation type | categorical | dry grassland |
| moderately wet grassland | |||
| wet grassland | |||
| Season | categorical | spring | |
| summer | |||
| autumn | |||
| Management | categorical | low intensity | |
| medium intensity | |||
| high intensity | |||
| Area | Sample size | categorical | 0.01 m2 |
| 1 m2 | |||
| Soil parameters | pH H2O | continuous | pH measured in H2O |
| pH KCl | continuous | pH measured in KCl | |
| EC | continuous | electrical conductivity (μS·cm−1) | |
| CaCO3 | continuous | calcium (mg·kg−1 soil dry) | |
| C | continuous | total organic carbon (%) | |
| N | continuous | total nitrogen (% soil dry) | |
| MgO | continuous | magnesium (mg·kg−1 soil dry) | |
| P2O5 | continuous | phosphorus (mg·kg−1 soil dry) | |
| K2O | continuous | potassium (mg·kg−1 soil dry) | |
| 2–1 mm | continuous | soil fraction 2–1 mm (%) | |
| 1–0.5 mm | continuous | soil fraction 1–0.5 mm (%) | |
| 0.5–0.25 mm | continuous | soil fraction 0.5–0.25 mm (%) | |
| 0.25–0.1 mm | continuous | soil fraction 0.25–0.1 mm (%) | |
| 0.1–0.05 mm | continuous | soil fraction 0.1–0.05 mm (%) | |
| 0.05–0.02 mm | continuous | soil fraction 0.05–0.02 mm (%) | |
| 0.02–0.005 mm | continuous | soil fraction 0.02–0.005 mm (%) | |
| 0.005–0.002 mm | continuous | soil fraction 0.005–0.002 mm (%) | |
| < 0.002 mm | continuous | soil fraction < 0.002 mm (%) | |
| Species | Species occurrence | binary | species occurrence presence/absence (1/0) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rucińska, A.; Olszak, M.; Świerszcz, S.; Nobis, M.; Zubek, S.; Kusza, G.; Boczkowska, M.; Nowak, A. Looking for Hidden Enemies of Metabarcoding: Species Composition, Habitat and Management Can Strongly Influence DNA Extraction while Examining Grassland Communities. Biomolecules 2021, 11, 318. https://doi.org/10.3390/biom11020318
Rucińska A, Olszak M, Świerszcz S, Nobis M, Zubek S, Kusza G, Boczkowska M, Nowak A. Looking for Hidden Enemies of Metabarcoding: Species Composition, Habitat and Management Can Strongly Influence DNA Extraction while Examining Grassland Communities. Biomolecules. 2021; 11(2):318. https://doi.org/10.3390/biom11020318
Chicago/Turabian StyleRucińska, Anna, Marcin Olszak, Sebastian Świerszcz, Marcin Nobis, Szymon Zubek, Grzegorz Kusza, Maja Boczkowska, and Arkadiusz Nowak. 2021. "Looking for Hidden Enemies of Metabarcoding: Species Composition, Habitat and Management Can Strongly Influence DNA Extraction while Examining Grassland Communities" Biomolecules 11, no. 2: 318. https://doi.org/10.3390/biom11020318
APA StyleRucińska, A., Olszak, M., Świerszcz, S., Nobis, M., Zubek, S., Kusza, G., Boczkowska, M., & Nowak, A. (2021). Looking for Hidden Enemies of Metabarcoding: Species Composition, Habitat and Management Can Strongly Influence DNA Extraction while Examining Grassland Communities. Biomolecules, 11(2), 318. https://doi.org/10.3390/biom11020318








