AG-9, an Elastin-Derived Peptide, Increases In Vitro Oral Tongue Carcinoma Cell Invasion, through an Increase in MMP-2 Secretion and MT1-MMP Expression, in a RPSA-Dependent Manner
Abstract
1. Introduction
2. Materials and Methods
2.1. Peptide Synthesis
2.2. Cell Culture
2.3. Cytotoxicity Assay
2.4. Proliferation Assay
2.5. In Vitro Invasion Assays
2.6. Zymography Analyses
2.6.1. Cell Incubation with Effectors
2.6.2. Gelatin Zymography
2.6.3. Gelatin-Plasminogen Zymography
2.7. Western-Blot Analyses
2.8. Immunocytochemistry
2.9. RNA Isolation and Real-Time PCR Analysis
2.10. SiRNA Transfection
2.11. Docking Experiments
2.12. Statistical Analysis
3. Results
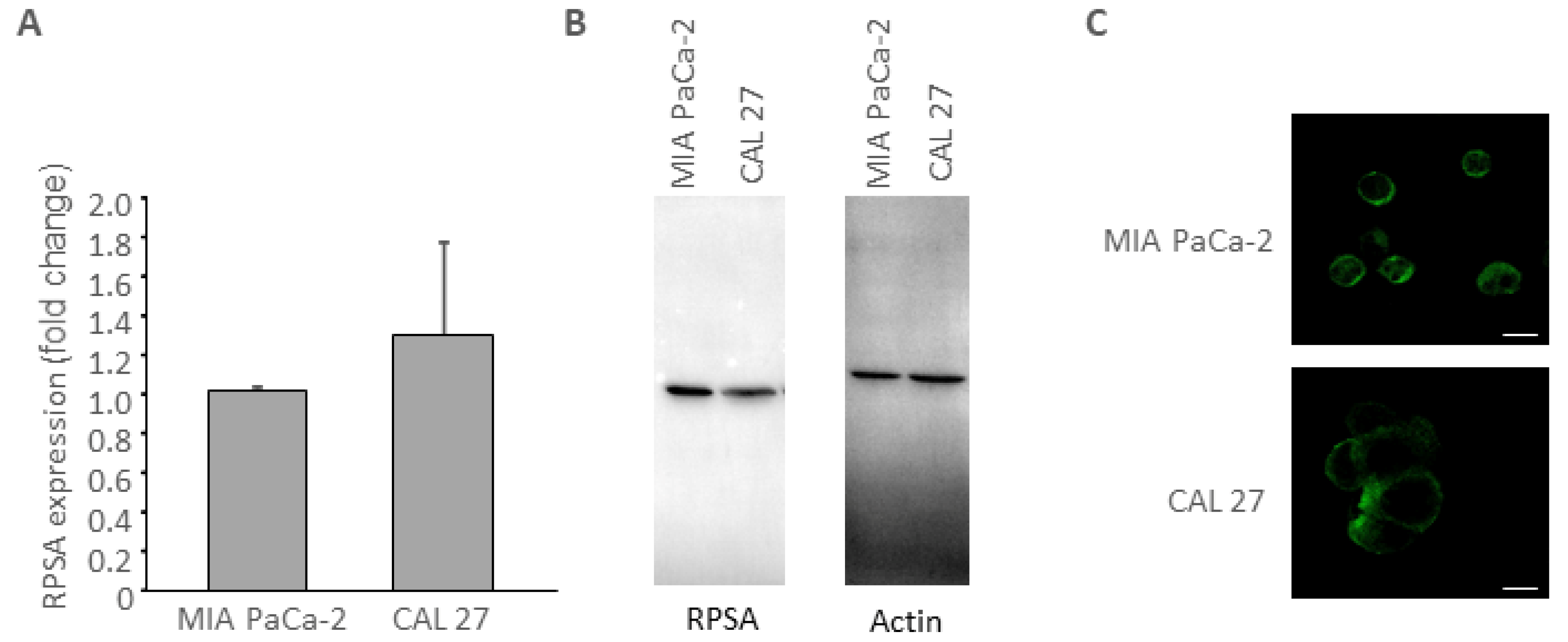
3.1. CAL 27 Cells Express RPSA Receptor
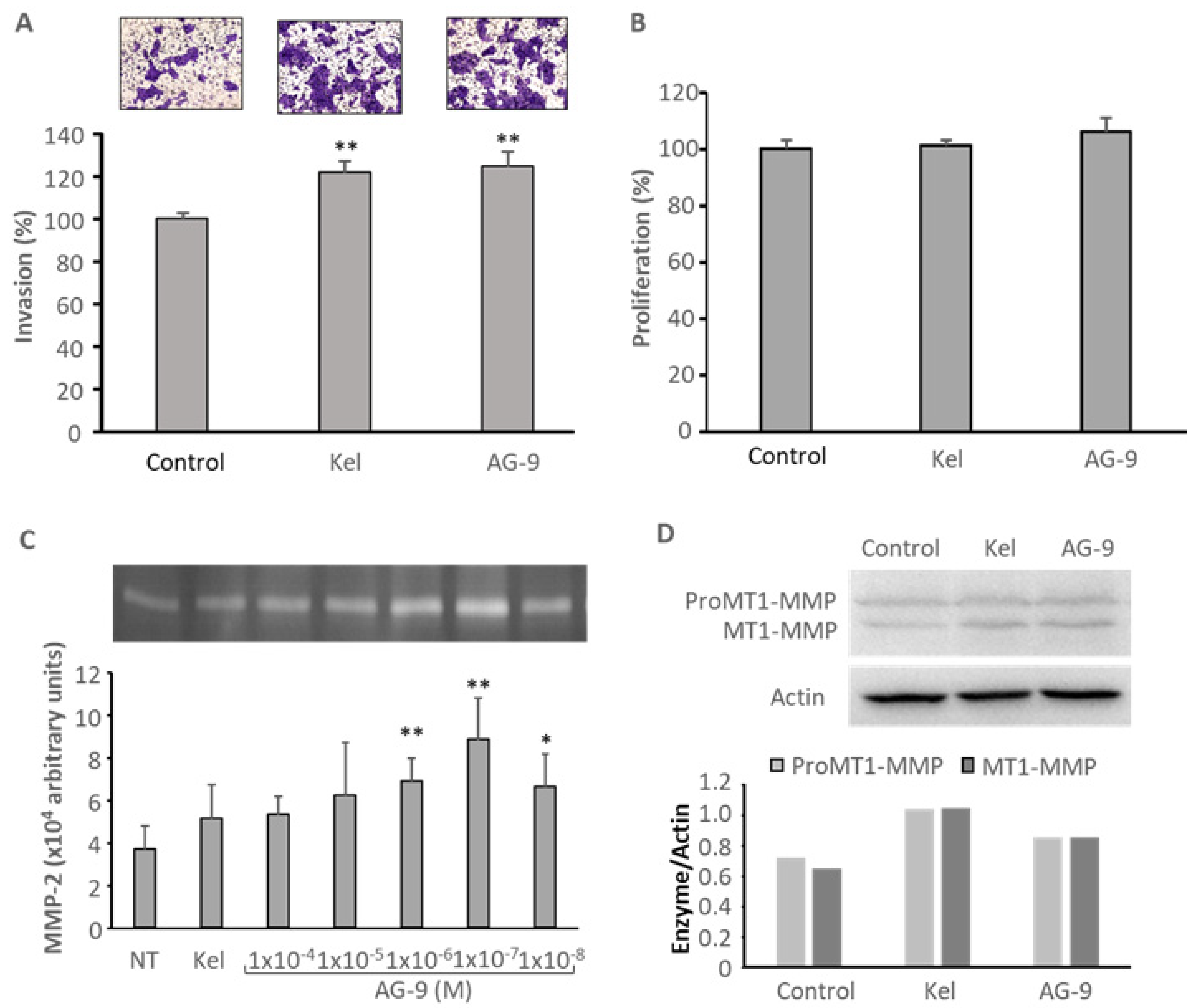
3.2. AG-9 Increases CAL 27 Invasion through Matrigel®, MMP-2 Secretion, and MT1-MMP Expression
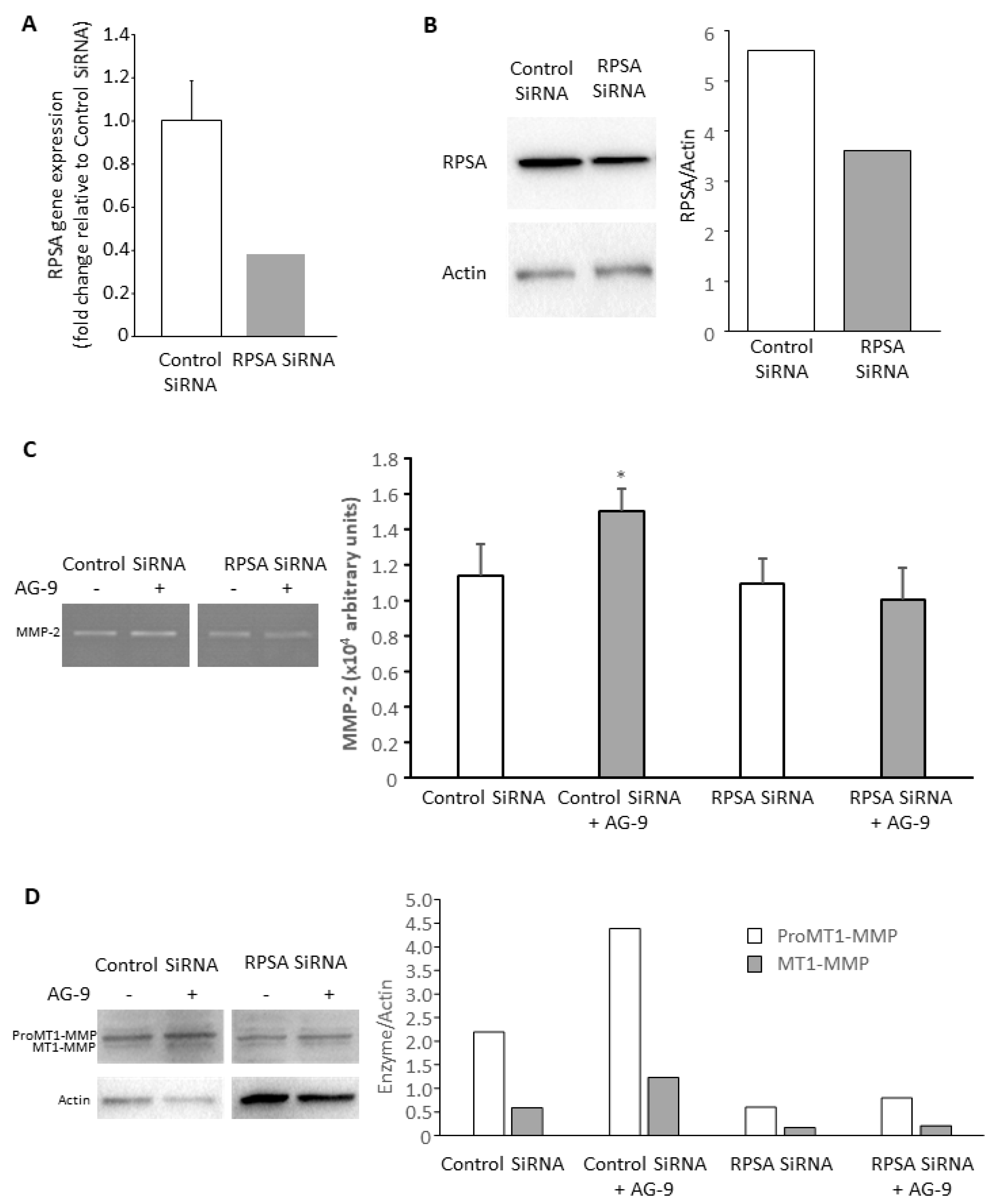
3.3. AG-9 Proinvasive Effects Are Mediated through RPSA Receptor
3.4. In Silico Study of AG-9 and EGCG Binding on RPSA
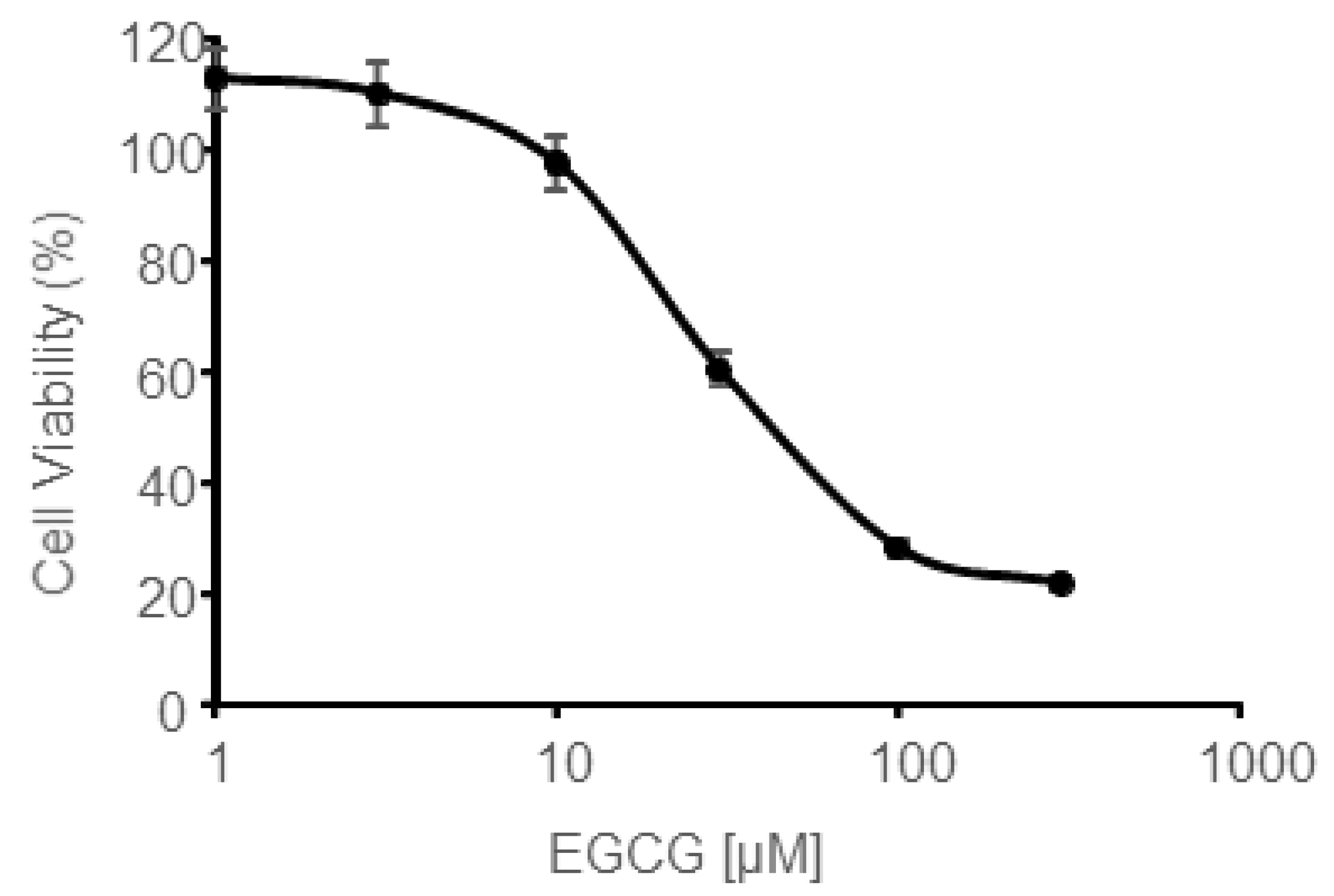
3.5. In Vitro EGCG Cytotoxicity on CAL 27 Cells
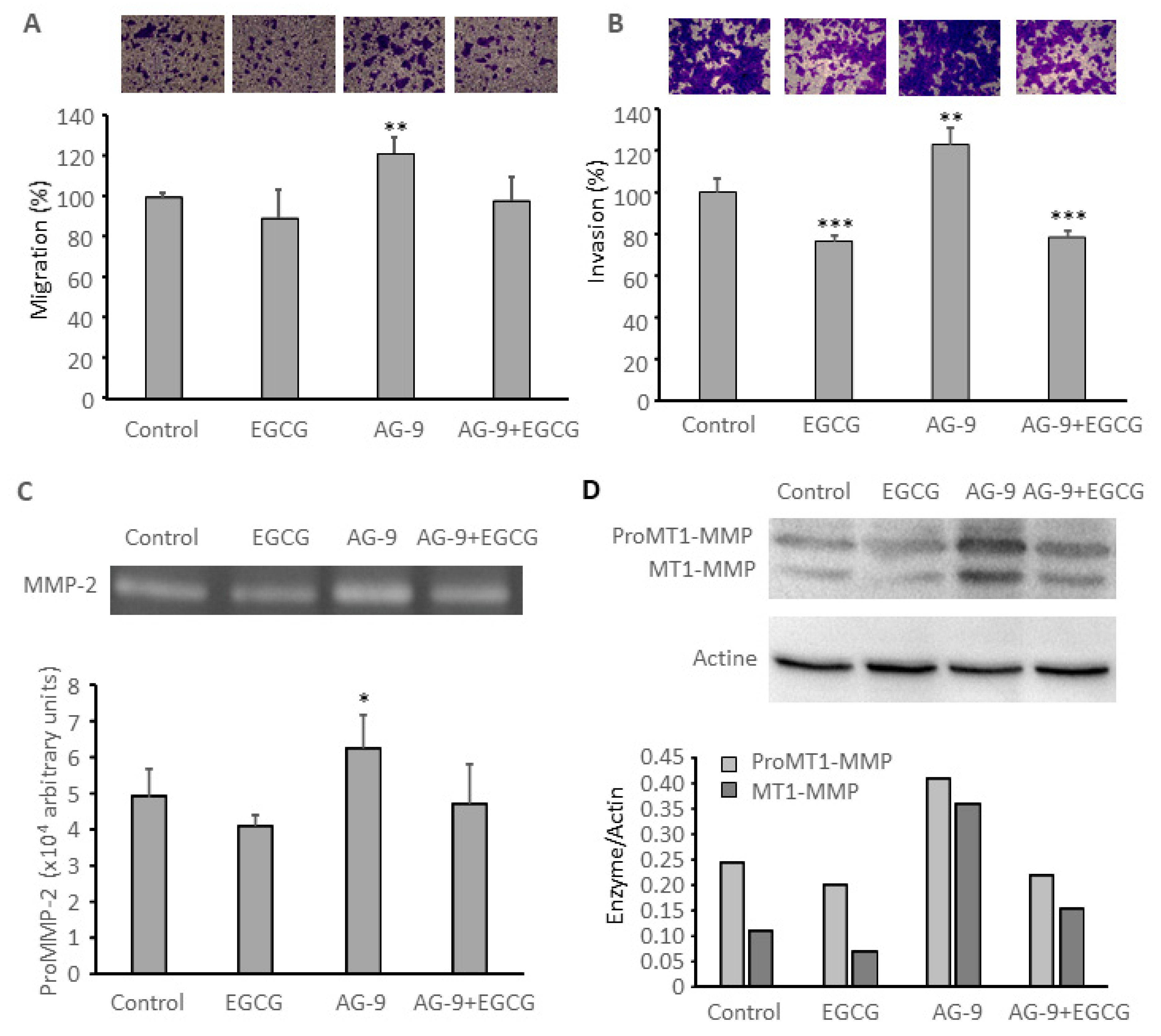
3.6. EGCG Prevents AG-9 Stimulatory Effect on CAL 27 Migration, Invasion, MMP-2 Secretion, and MT1-MMP Expression
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chi, A.C.; Day, T.A.; Neville, B.W. Oral cavity and oropharyngeal squamous cell carcinoma-an update. CA. Cancer J. Clin. 2015. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.J.; Kim, J.H. Increasing incidence and improving survival of oral tongue squamous cell carcinoma. Sci. Rep. 2020. [Google Scholar] [CrossRef]
- Ellington, T.D.; Henley, S.J.; Senkomago, V.; O’Neil, M.E.; Wilson, R.J.; Singh, S.; Thomas, C.C.; Wu, M.; Richardson, L.C. Trends in Incidence of Cancers of the Oral Cavity and Pharynx—United States 2007–2016. MMWR Morb. Mortal. Wkly. Rep. 2020. [Google Scholar] [CrossRef] [PubMed]
- Paderno, A.; Morello, R.; Piazza, C. Tongue carcinoma in young adults: A review of the literature. Acta Otorhinolaryngol. Ital. 2018, 38, 175–180. [Google Scholar] [PubMed]
- Lenze, N.R.; Farquhar, D.R.; Dorismond, C.; Sheth, S.; Zevallos, J.P.; Blumberg, J.; Lumley, C.; Patel, S.; Hackman, T.; Weissler, M.C.; et al. Age and risk of recurrence in oral tongue squamous cell carcinoma: Systematic review. Head Neck 2020, 42, 3755–3768. [Google Scholar] [CrossRef] [PubMed]
- Mukdad, L.; Heineman, T.E.; Alonso, J.; Badran, K.W.; Kuan, E.C.; St. John, M.A. Oral tongue squamous cell carcinoma survival as stratified by age and sex: A surveillance, epidemiology, and end results analysis. Laryngoscope 2019, 129. [Google Scholar] [CrossRef] [PubMed]
- Tota, J.E.; Anderson, W.F.; Coffey, C.; Califano, J.; Cozen, W.; Ferris, R.L.; St. John, M.; Cohen, E.E.W.; Chaturvedi, A.K. Rising incidence of oral tongue cancer among white men and women in the United States, 1973–2012. Oral Oncol. 2017. [Google Scholar] [CrossRef]
- Gonzalez, M.; Riera, A. Tongue Cancer. Available online: https://www.ncbi.nlm.nih.gov/books/NBK562324/ (accessed on 30 December 2020).
- Wagenseil, J.E.; Mecham, R.P. Vascular extracellular matrix and arterial mechanics. Physiol. Rev. 2009, 89, 957–989. [Google Scholar] [CrossRef]
- Marettova, E.; Maretta, M. Distribution of elastic fibers in the cat tongue. Rev. Med. Vet. (Toulouse). 2012, 163, 577–580. [Google Scholar]
- Maquart, F.X.; Pasco, S.; Ramont, L.; Hornebeck, W.; Monboisse, J.C. An introduction to matrikines: Extracellular matrix-derived peptides which regulate cell activity - Implication in tumor invasion. Crit. Rev. Oncol. Hematol. 2004, 49, 199–202. [Google Scholar] [CrossRef]
- Duca, L.; Floquet, N.; Alix, A.J.P.; Haye, B.; Debelle, L. Elastin as a matrikine. Crit. Rev. Oncol. Hematol. 2004, 49, 235–244. [Google Scholar] [CrossRef]
- Heinz, A. Elastases and elastokines: Elastin degradation and its significance in health and disease. Crit. Rev. Biochem. Mol. Biol. 2020, 55, 252–273. [Google Scholar] [CrossRef]
- Brassart, B.; Randoux, A.; Hornebeck, W.; Emonard, H. Regulation of matrix metalloproteinase-2 (gelatinase A, MMP-2), membrane-type matrix metalloproteinase-1 (MT1-MMP) and tissue inhibitor of metalloproteinases-2 (TIMP-2) expression by elastin-derived peptides in human HT-1080 fibrosarcoma cell line. Clin. Exp. Metastasis 1998, 16, 489–500. [Google Scholar] [CrossRef]
- Huet, E.; Brassart, B.; Cauchard, J.-H.; Debelle, L.; Birembaut, P.; Wallach, J.; Emonard, H.; Polette, M.; Hornebeck, W. Cumulative influence of elastin peptides and plasminogen on matrix metalloproteinase activation and type I collagen invasion by HT-1080 fibrosarcoma cells. Clin. Exp. Metastasis 2002, 19. [Google Scholar] [CrossRef]
- Robinet, A.; Fahem, A.; Cauchard, J.-H.; Huet, E.; Vincent, L.; Lorimier, S.; Antonicelli, F.; Soria, C.; Crepin, M.; Hornebeck, W.; et al. Elastin-derived peptides enhance angiogenesis by promoting endothelial cell migration and tubulogenesis through upregulation of MT1-MMP. J. Cell Sci. 2005, 118, 343–356. [Google Scholar] [CrossRef]
- Donet, M.; Brassart-Pasco, S.; Salesse, S.; Maquart, F.-X.; Brassart, B. Elastin peptides regulate HT-1080 fibrosarcoma cell migration and invasion through an Hsp90-dependent mechanism. Br. J. Cancer 2014, 111. [Google Scholar] [CrossRef]
- Da Silva, J.; Lameiras, P.; Beljebbar, A.; Berquand, A.; Villemin, M.; Ramont, L.; Dukic, S.; Nuzillard, J.-M.; Molinari, M.; Gautier, M.; et al. Structural characterization and in vivo pro-tumor properties of a highly conserved matrikine. Oncotarget 2018, 9. [Google Scholar] [CrossRef]
- Brassart, B.; Da Silva, J.; Donet, M.; Seurat, E.; Hague, F.; Terryn, C.; Velard, F.; Michel, J.; Ouadid-Ahidouch, H.; Monboisse, J.C.; et al. Tumour cell blebbing and extracellular vesicle shedding: Key role of matrikines and ribosomal protein SA. Br. J. Cancer 2019, 120, 453–465. [Google Scholar] [CrossRef]
- Mecham, R.P.; Hinek, A.; Griffin, G.L.; Senior, R.M.; Liotta, L.A. The elastin receptor shows structural and functional similarities to the 67-kDa tumor cell laminin receptor. J. Biol. Chem. 1989, 264, 16652–16657. [Google Scholar]
- Digiacomo, V.; Meruelo, D. Looking into laminin receptor: Critical discussion regarding the non-integrin 37/67-kDa laminin receptor/RPSA protein. Biol. Rev. 2016, 91, 288–310. [Google Scholar] [CrossRef]
- Vania, L.; Morris, G.; Otgaar, T.C.; Bignoux, M.J.; Bernert, M.; Burns, J.; Gabathuse, A.; Singh, E.; Ferreira, E.; Weiss, S.F.T. Patented therapeutic approaches targeting LRP/LR for cancer treatment. Expert Opin Ther Pat. 2019, 29, 987–1009. [Google Scholar] [CrossRef]
- Gan, R.Y.; Li, H.B.; Sui, Z.Q.; Corke, H. Absorption, metabolism, anti-cancer effect and molecular targets of epigallocatechin gallate (EGCG): An updated review. Crit. Rev. Food Sci. Nutr. 2018, 58, 924–941. [Google Scholar] [CrossRef]
- Almatrood, S.A.; Almatroudi, A.; Khan, A.A.; Alhumaydh, F.A.; Alsahl, M.A.; Rahmani, A.H. Potential therapeutic targets of epigallocatechin gallate (EGCG), the most abundant catechin in green tea, and its role in the therapy of various types of cancer. Molecules 2020, 25, 3146. [Google Scholar] [CrossRef]
- Aggarwal, V.; Tuli, H.S.; Tania, M.; Srivastava, S.; Ritzer, E.E.; Pandey, A.; Aggarwal, D.; Barwal, T.S.; Jain, A.; Kaur, G.; et al. Molecular mechanisms of action of epigallocatechin gallate in cancer: Recent trends and advancement. Semin. Cancer Biol. 2020, 32461153. [Google Scholar] [CrossRef]
- Ho, Y.C.; Yang, S.F.; Peng, C.Y.; Chou, M.Y.; Chang, Y.C. Epigallocatechin-3-gallate inhibits the invasion of human oral cancer cells and decreases the productions of matrix metalloproteinases and urokinase-plasminogen activator. J. Oral Pathol. Med. 2007. [Google Scholar] [CrossRef] [PubMed]
- Chiang, W.C.; Wong, Y.K.; Lin, S.C.; Chang, K.W.; Liu, C.J. Increase of MMP-13 expression in multi-stage oral carcinogenesis and epigallocatechin-3-gallate suppress MMP-13 expression. Oral Dis. 2006. [Google Scholar] [CrossRef]
- Chen, P.N.; Chu, S.C.; Kuo, W.H.; Chou, M.Y.; Lin, J.K.; Hsieh, Y.S. Epigallocatechin-3 gallate inhibits invasion, epithelial-mesenchymal transition, and tumor growth in oral cancer cells. J. Agric. Food Chem. 2011. [Google Scholar] [CrossRef]
- Irimie, A.I.; Braicu, C.; Zanoaga, O.; Pileczki, V.; Gherman, C.; Berindan-Neagoe, I.; Campian, R.S. Epigallocatechin-3-gallate suppresses cell proliferation and promotes apoptosis and autophagy in oral cancer SSC-4 cells. Onco. Targets. Ther. 2015. [Google Scholar] [CrossRef]
- Li, A.; Gu, K.; Wang, Q.; Chen, X.; Fu, X.; Wang, Y.; Wen, Y. Epigallocatechin-3-gallate affects the proliferation, apoptosis, migration and invasion of tongue squamous cell carcinoma through the Hippo-TAZ signaling pathway. Int. J. Mol. Med. 2018. [Google Scholar] [CrossRef]
- Fujimura, Y.; Sumida, M.; Sugihara, K.; Tsukamoto, S.; Yamada, K.; Tachibana, H. Green tea polyphenol EGCG sensing motif on the 67-kDa laminin receptor. PLoS ONE 2012. [Google Scholar] [CrossRef]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef]
- Fülöp, T.; Larbi, A. Putative role of 67 kDa elastin-laminin receptor in tumor invasion. Semin. Cancer Biol. 2002. [Google Scholar] [CrossRef]
- Lefebvre, T.; Rybarczyk, P.; Bretaudeau, C.; Vanlaeys, A.; Cousin, R.; Brassart-Pasco, S.; Chatelain, D.; Dhennin-Duthille, I.; Ouadid-Ahidouch, H.; Brassart, B.; et al. TRPM7/RPSA Complex Regulates Pancreatic Cancer Cell Migration. Front. Cell Dev. Biol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Tachibana, H.; Koga, K.; Fujimura, Y.; Yamada, K. A receptor for green tea polyphenol EGCG. Nat. Struct. Mol. Biol. 2004. [Google Scholar] [CrossRef] [PubMed]
- Weisburg, J.H.; Weissman, D.B.; Sedaghat, T.; Babich, H. In vitro cytotoxicity of epigallocatechin gallate and tea extracts to cancerous and normal cells from the human oral cavity. Basic Clin. Pharmacol. Toxicol. 2004. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.M.; Chang, P.Y.; Tu, M.G.; Lu, C.C.; Kuo, S.C.; Amagaya, S.; Lee, C.Y.; Jao, H.Y.; Chen, M.Y.; Yang, J.S. Epigallocatechin gallate sensitizes CAL-27 human oral squamous cell carcinoma cells to the anti-metastatic effects of gefitinib (Iressa) via synergistic suppression of epidermal growth factor receptor and matrix metalloproteinase-2. Oncol. Rep. 2012. [Google Scholar] [CrossRef]
- Zanoni, D.K.; Montero, P.H.; Migliacci, J.C.; Shah, J.P.; Wong, R.J.; Ganly, I.; Patel, S.G. Survival outcomes after treatment of cancer of the oral cavity (1985–2015). Oral Oncol. 2019. [Google Scholar] [CrossRef]
- Betre, H.; Ong, S.R.; Guilak, F.; Chilkoti, A.; Fermor, B.; Setton, L.A. Chondrocytic differentiation of human adipose-derived adult stem cells in elastin-like polypeptide. Biomaterials 2006, 27, 91–99. [Google Scholar] [CrossRef]
- Long, M.M.; King, V.J.; Prasad, K.U.; Freeman, B.A.; Urry, D.W. Elastin repeat peptides as chemoattractants for bovine aortic endothelial cells. J. Cell. Physiol. 1989, 140, 512–518. [Google Scholar] [CrossRef] [PubMed]
- Senior, R.M.; Griffin, G.L.; Mecham, R.P. Chemotactic activity of elastin-derived peptides. J. Clin. Invest. 1980, 66, 859–862. [Google Scholar] [CrossRef]
- Toupance, S.; Brassart, B.; Rabenoelina, F.; Ghoneim, C.; Vallar, L.; Polette, M.; Debelle, L.; Birembaut, P. Elastin-derived peptides increase invasive capacities of lung cancer cells by post-transcriptional regulation of MMP-2 and uPA. Clin. Exp. Metastasis 2012, 29. [Google Scholar] [CrossRef]
- Jiang, L.; Ji, N.; Zhou, Y.; Li, J.; Liu, X.; Wang, Z.; Chen, Q.; Zeng, X. CAL 27 is an oral adenosquamous carcinoma cell line. Oral Oncol. 2009. [Google Scholar] [CrossRef]
- Ntayi, C.; Labrousse, A.L.; Debret, R.; Birembaut, P.; Bellon, G.; Antonicelli, F.; Hornebeck, W.; Bernard, P. Elastin-Derived Peptides Upregulate Matrix Metalloproteinase-2-ediated Melanoma Cell Invasion Through Elastin-Binding Protein. J. Invest. Dermatol. 2004. [Google Scholar] [CrossRef] [PubMed]
- Szychowski, K.A.; Wójtowicz, A.K.; Gmiński, J. Impact of Elastin-Derived Peptide VGVAPG on Matrix Metalloprotease-2 and -9 and the Tissue Inhibitor of Metalloproteinase-1, -2, -3 and -4 mRNA Expression in Mouse Cortical Glial Cells In Vitro. Neurotox. Res. 2019. [Google Scholar] [CrossRef] [PubMed]
- Fang, C.Y.; Wu, C.C.; Hsu, H.Y.; Chuang, H.Y.; Huang, S.Y.; Tsai, C.H.; Chang, Y.; Tsao, G.S.W.; Chen, C.L.; Chen, J.Y. EGCG inhibits proliferation, invasiveness and tumor growth by up-regulation of adhesion molecules, suppression of gelatinases activity, and induction of apoptosis in nasopharyngeal carcinoma cells. Int. J. Mol. Sci. 2015, 16, 2530–2558. [Google Scholar] [CrossRef] [PubMed]
- Ho, H.C.; Huang, C.C.; Lu, Y.T.; Yeh, C.M.; Ho, Y.T.; Yang, S.F.; Hsin, C.H.; Lin, C.W. Epigallocatechin-3-gallate inhibits migration of human nasopharyngeal carcinoma cells by repressing MMP-2 expression. J. Cell. Physiol. 2019. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bretaudeau, C.; Baud, S.; Dupont-Deshorgue, A.; Cousin, R.; Brassart, B.; Brassart-Pasco, S. AG-9, an Elastin-Derived Peptide, Increases In Vitro Oral Tongue Carcinoma Cell Invasion, through an Increase in MMP-2 Secretion and MT1-MMP Expression, in a RPSA-Dependent Manner. Biomolecules 2021, 11, 39. https://doi.org/10.3390/biom11010039
Bretaudeau C, Baud S, Dupont-Deshorgue A, Cousin R, Brassart B, Brassart-Pasco S. AG-9, an Elastin-Derived Peptide, Increases In Vitro Oral Tongue Carcinoma Cell Invasion, through an Increase in MMP-2 Secretion and MT1-MMP Expression, in a RPSA-Dependent Manner. Biomolecules. 2021; 11(1):39. https://doi.org/10.3390/biom11010039
Chicago/Turabian StyleBretaudeau, Clara, Stéphanie Baud, Aurélie Dupont-Deshorgue, Rémi Cousin, Bertrand Brassart, and Sylvie Brassart-Pasco. 2021. "AG-9, an Elastin-Derived Peptide, Increases In Vitro Oral Tongue Carcinoma Cell Invasion, through an Increase in MMP-2 Secretion and MT1-MMP Expression, in a RPSA-Dependent Manner" Biomolecules 11, no. 1: 39. https://doi.org/10.3390/biom11010039
APA StyleBretaudeau, C., Baud, S., Dupont-Deshorgue, A., Cousin, R., Brassart, B., & Brassart-Pasco, S. (2021). AG-9, an Elastin-Derived Peptide, Increases In Vitro Oral Tongue Carcinoma Cell Invasion, through an Increase in MMP-2 Secretion and MT1-MMP Expression, in a RPSA-Dependent Manner. Biomolecules, 11(1), 39. https://doi.org/10.3390/biom11010039





