Transcriptomics and Metabonomics Identify Essential Metabolic Signatures in Calorie Restriction (CR) Regulation across Multiple Mouse Strains
Abstract
:1. Introduction
2. Results
2.1. CR Led in a Significant Modulation of Body Weight Gain across the Animal Strains
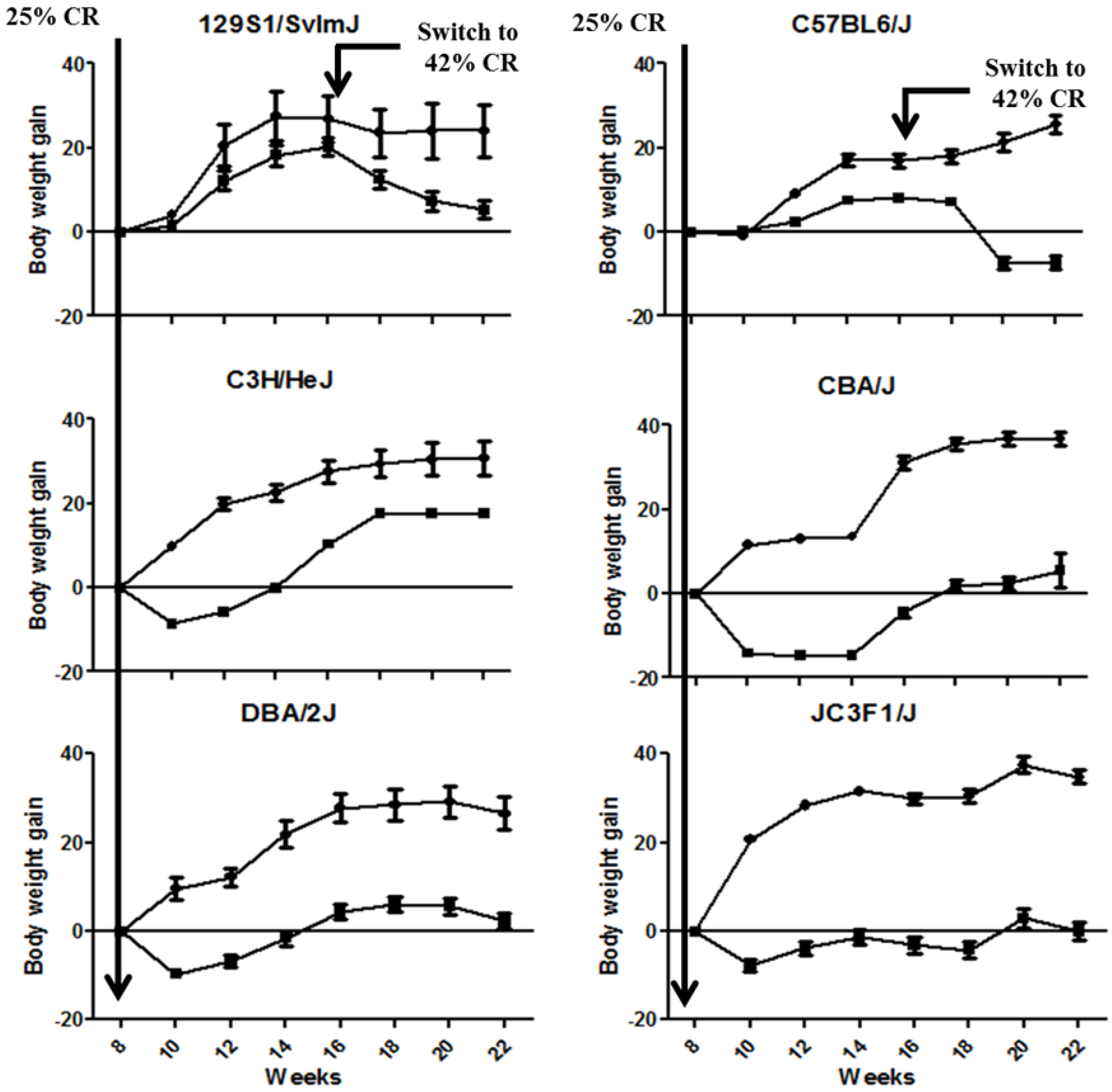
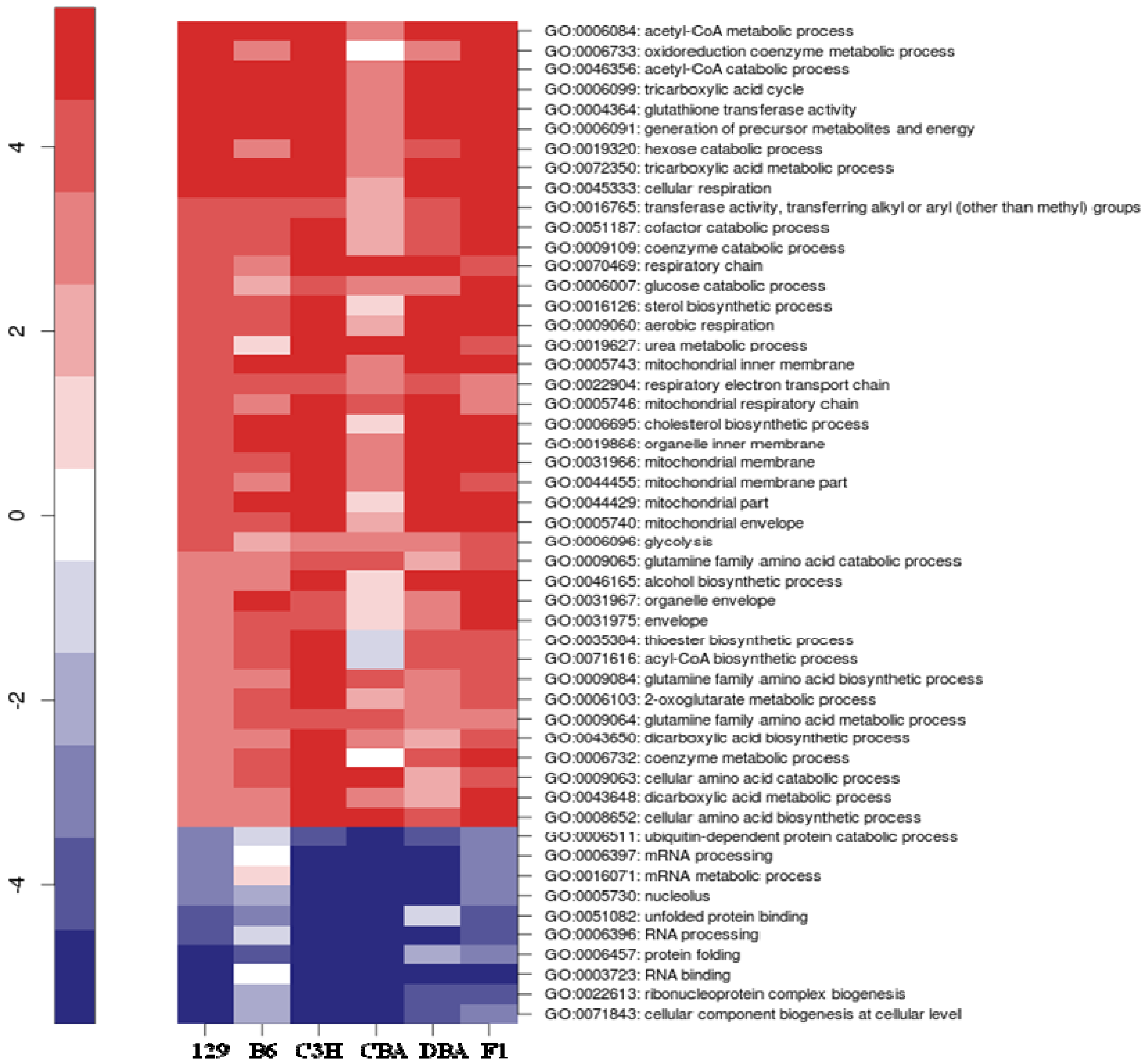
2.2. Identification of Transcriptional Biomarkers of CR in Mouse Liver

2.3. Identification of Common Pathways Affected by CR in Liver of Multiple Mice Strains
2.4. 1H NMR Spectroscopy of Blood Plasma on the Six Strains Highlighted a Very Heterogeneous Response in Circulating Lipids Following CR
| Metabolites | Chemical shift | 129S1/SvlmJ | C57BL6/J | C3H/HeJ | CBA/J | DBA/2J | JC3F1/J | |
|---|---|---|---|---|---|---|---|---|
| 1 | Methyl signal of fatty acids in HDL | 0.83 (m) | −0.60 | −0.64 | 0.51 | 0.09 | 0.52 | −0.37 |
| 2 | Methyl signal of fatty acids in LDL | 0.85 (m) | −0.64 | −0.92 | 0.68 | −0.30 | 0.79 | −0.84 |
| 3 | Methyl signal of fatty acids in VLDL | 0.87 (m) | −0.15 | −0.49 | −0.69 | −0.85 | 0.46 | −0.62 |
| 5 | N-acetyl glycoprotein (NAG) associated resonances | 2.04 (s) | −0.83 | −0.75 | −0.91 | −0.67 | 0.41 | −0.55 |
| 6 | Lipids (FA chain) | 2.23 (m) | −0.64 | −0.57 | −0.68 | −0.85 | 0.37 | −0.47 |
| 7 | Lipids (Polyunsaturated FAs) | 2.75 (m) | −0.85 | −0.85 | −0.63 | −0.81 | 0.51 | −0.91 |
| 8 | Glycerophosphocholine | 3.22 (s) | −0.33 | −0.73 | 0.81 | 0.22 | 0.81 | −0.66 |
| 9 | Lipids (Unsaturated FAs) | 5.30 (m) | −0.88 | −0.88 | −0.85 | 0.82 | 0.71 | −0.96 |
2.5. Metabolic Profiling of Urine and Liver Tissues in C57BL6/J Mice
| Metabolites | Chemical shift | Control Mean ± SD | CR Mean ± SD | OPLS-DA correlation coefficients (p corr) | p-Value | |||
|---|---|---|---|---|---|---|---|---|
| Liver hydrophilic fraction | Valine | 1.04 (d) | 4.54 ± 1.26 | 7.11 ± 2.12 | 0.68 | <0.001 | ||
| Isoleucine | 1.02 (d) | 1.22 ± 2.90 | 1.84 ± 5.75 | 0.74 | 0.001 | |||
| GSH | 2.55(m) | 3.55 ± 1.04 | 6.60 ± 1.94 | 0.87 | <0.001 | |||
| Carnosine | 4.46(m) | 2.86 ± 0.68 | 4.38 ± 1.09 | 0.73 | <0.001 | |||
| Uridine | 5.9 (d) | 2.11 ± 0.5 | 2.03 ± 0.72 | −0.79 | 0.01 | |||
| Phenylalanine | 7.36 (m) | 1.26 ± 0.26 | 1.02 ± 0.34 | −0.73 | <0.01 | |||
| Nicotinurate | 8.93 (d) | 1.34 ± 0.37 | 2.44 ± 0.52 | 0.77 | <0.001 | |||
| N,N-dimethylglycine | 2.92 (s) | 3.91 ± 1.1 | 6.14 ± 1.37 | 0.81 | <0.001 | |||
| Liver Organic Phase | Cholesterol (free) | 0.93, 1.01, 1.04 (m) | 1.26 ± 0.92 | 3.14 ± 0.33 | 0.84 | <0.001 | ||
| Cholesterol (total) | 0.69 (s), 0.86–0.88 (dd) | 1.54 ± 0.31 | 3.67 ± 0.31 | 0.68 | <0.001 | |||
| Sphingomyeline | 5.61 (m), 5.35 (m)3.29 (s) | 5.98 ± 3.26 | 18.7 ± 3.36 | 0.97 | <0.001 | |||
| -3 FA | 0.97 (t) | 3.50 ± 0.16 | 5.61 ± 0.13 | 0.68 | <0.001 | |||
| UFA | 5.31 (m), | 2.63 ± 0.28 | 2.05 ± 0.24 | −0.69 | <0.001 | |||
| Retyl-Conjugate | 6.18 (d) | 2.57 ± 1.07 | 9.37 ± 1.11 | 0.87 | <0.001 | |||
| Fatty acyl chain | 2.34 (m) | 4.05 ± 0.23 | 3.25 ± 0.15 | −0.97 | <0.001 | |||
| Triacylglycerides (TAG) | 5.09 (m) | 5.14 ± 0.56 | 2.19 ± 0.53 | −0.68 | <0.001 | |||
| Urine | L-Fucose | 4.56 (d) | 2.90 ± 0.21 | 3.87 ± 0.42 | 0.83 | <0.001 | ||
| Allantoin | 5.41 (s) | 5.10 ± 1.35 | 9.28 ± 2.30 | 0.84 | <0.001 | |||
| alpha-ketogluterate | 3.01 (t) | 1.16 ± 0.11 | 1.74 ± 0.22 | 0.88 | 0.01 | |||
| Taurine | 3.26 (t) | 4.63 ± 0.59 | 7.06± 2.14 | 0.84 | 0.001 | |||
| Uracil | 7.54 (d) | 5.96 ± 1.15 | 8.25 ± 1.25 | 0.72 | 0.001 | |||
| Pseudouridine | 7.69 (s) | 5.01 ± 0.51 | 6.75 ± 0.96 | 0.88 | 0.001 | |||
| N-Methylnicotinamide | 9.28 (s) | 2.22 ± 0.49 | 3.45 ± 0.52 | 0.76 | 0.001 | |||
| Phenylacetylglycine | 7.43 (m) | 1.47 ± 0.33 | 2.00 ± 0.24 | 0.77 | 0.001 | |||
| Deoxycytidine | 7.83 (d), 6.28 (t), 6.06 (d) | 3.95 ± 0.56 | 6.67 ± 1.72 | 0.77 | <0.001 | |||
2.6. Targeted MS Profiles of Blood Plasma from C57BL6/J Mice
| Metabolite (μM/L) | Control Mean ± SD | CR Mean ± SD | p-Value | OPLS-DA correlation coefficients (p corr) |
|---|---|---|---|---|
| PC-O 34:1 | 188.5 ± 11.05 | 62.6 ± 10.5 | <0.001 | −0.81 |
| C18:1 | 0.44 ± 0.07 | 0.22 ± 0.06 | <0.001 | −0.90 |
| PC 36:1 | 24.73 ± 3.65 | 16.4 ± 1.58 | <0.001 | −0.86 |
| PC 34:3 | 6.73 ± 1.13 | 3.49 ± 1.15 | 0.0001 | −0.88 |
| C14:1 | 0.22 ± 0.04 | 0.14 ± 0.03 | 0.0002 | −0.82 |
| PC-O 38:0 | 3.91 ± 0.92 | 2.31 ± 0.57 | 0.0002 | −0.89 |
| PC-O 40:5 | 3.48 ± 1.04 | 2.90 ± 0.53 | 0.01 | −0.80 |
| PC 40:4 | 1.61 ± 0.30 | 1.24 ± 0.24 | 0.007 | −0.75 |
| PC 36:5 | 3.51 ± 0.92 | 2.08 ± 0.61 | <0.001 | −0.83 |
| Sugars-Hexose | 5996 ± 1.303 | 3534 ± 1397 | 0.001 | −0.68 |
3. Discussion
3.1. Complementary Transcriptomics and Metabonomics Reveal CR Effects on Fatty Acid Oxidation, Gluconeogenesis and Cholesterol Metabolism
3.2. Complementary Transcriptomics and Metabonomics Reveal Impact of CR on Insulin Signaling and Stress Response
4. Experimental Section
4.1. Animals and Dietary Manipulation
4.2. Sample Collection
4.3. Transcriptomic Analyses
4.4. Targeted LC-MS/MS Metabolite Profiling
4.5. Annotation of Lipid Species
4.6. Metabolite Profiling-Sample Preparation and 1H NMR Spectroscopic Analysis
4.7. Chemometrics
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- Sansoni, P.; Vescovini, R.; Fagnoni, F.; Biasini, C.; Zanni, F.; Zanlari, L.; Telera, A.; Lucchini, G.; Passeri, G.; Monti, D.; et al. The immune system in extreme longevity. Exp. Gerontol. 2008, 43, 61–65. [Google Scholar] [CrossRef]
- Franceschi, C. Inflammaging as a major characteristic of old people: Can it be prevented or cured? Nutr. Rev. 2007, 65, 173–176. [Google Scholar] [CrossRef]
- Burkle, A.; Caselli, G.; Franceschi, C.; Mariani, E.; Sansoni, P.; Santoni, A.; Vecchio, G.; Witkowski, J.M.; Caruso, C. Pathophysiology of ageing, longevity and age related diseases. Immun. Ageing 2007, 4, 4–17. [Google Scholar] [CrossRef]
- Franceschi, C.; Capri, M.; Monti, D.; Giunta, S.; Olivieri, F.; Sevini, F.; Panourgia, M.P.; Invidia, L.; Celani, L.; Scurti, M.; et al. Inflammaging and anti-inflammaging: A systemic perspective on aging and longevity emerged from studies in humans. Mech. Ageing Dev. 2007, 128, 92–105. [Google Scholar] [CrossRef]
- Weindruch, R. Caloric restriction and aging. Sci. Am. 1996, 274, 46–52. [Google Scholar] [CrossRef]
- Roth, G.S.; Lane, M.A.; Ingram, D.K. Caloric restriction mimetics: The next phase. Ann. NY Acad. Sci. 2005, 1057, 365–371. [Google Scholar] [CrossRef]
- Anderson, R.M.; Weindruch, R. Metabolic reprogramming, caloric restriction and aging. Trends Endocrin. Metab 2010, 21, 134–141. [Google Scholar]
- Weindruch, R.; Naylor, P.H.; Goldstein, A.L.; Walford, R.L. Influences of aging and dietary restriction on serum thymosin alpha 1 levels in mice. J. Gerontol. 1988, 43, B40–B42. [Google Scholar] [CrossRef]
- Selman, C.; Withers, D.J. Mammalian models of extended healthy lifespan. Philos. T. R. Soc. B. 2011, 366, 99–107. [Google Scholar] [CrossRef]
- Fontana, L.; Partridge, L.; Longo, V.D. Extending healthy life span—from yeast to humans. Science 2010, 328, 321–326. [Google Scholar] [CrossRef]
- Fontana, L.; Villareal, D.T.; Weiss, E P.; Racette, S.B.; Steger-May, K.; Klein, S.; Holloszy, J.O. Calorie restriction or exercise: Effects on coronary heart disease risk factors. A randomized, controlled trial. Am. J. Physiol—Endoc. M. 2007, 293, 197–202. [Google Scholar]
- Weindruch, R.; Walford, R.L.; Fligiel, S.; Guthrie, D. The retardation of aging in mice by dietary restriction: Longevity, cancer, immunity and lifetime energy intake. J. Nutr. 1986, 116, 641–654. [Google Scholar]
- Kealy, R.D.; Lawler, D.F.; Ballam, J.M.; Mantz, S.L.; Biery, D.N.; Greeley, E.H. Effects of diet restriction on life span and age-related changes in dogs. J. Am. Vet. Med. A. 2001, 220, 1315–1320. [Google Scholar]
- Forster, M.J.; Morris, P.; Sohal, R.S. Genotype and age influence the effect of caloric intake on mortality in mice. FASEB J. 2003, 17, 690–692. [Google Scholar]
- De Grey, A.D. The unfortunate influence of the weather on the rate of ageing: Why human caloric restriction or its emulation may only extend life expectancy by 2–3 years. Gerontology 2005, 51, 73–82. [Google Scholar] [CrossRef]
- Heilbronn, L.K.; Ravussin, E. Calorie restriction and aging: Review of the literature and implications for studies in humans. Am. J. Clin. Nutr. 2003, 78, 361–369. [Google Scholar]
- Suzuki, M.; Akisaka, M.; Ashitomi, I.; Higa, K.; Nozaki, H. Chronological study concerning ADL among Okinawan centenarians. Nippon Ronen Igakkai Zasshi 1995, 32, 416–423. [Google Scholar] [CrossRef]
- Willcox, D.C.; Willcox, B.J.; Shimajiri, S.; Kurechi, S.; Suzuki, M. Aging gracefully: A retrospective analysis of functional status in Okinawan centenarians. Am. J. Geriat. Psychiat. 2007, 15, 252–256. [Google Scholar] [CrossRef]
- Walford, R.L.; Mock, D.; Verdery, R.; MacCallum, T. Calorie restriction in biosphere 2: Alterations in physiologic, hematologic, hormonal, and biochemical parameters in humans restricted for a 2-year period. J. Gerontol. A—Biol. 2002, 57, B211–B224. [Google Scholar] [CrossRef]
- Ingram, D.K.; Zhu, M.; Mamczarz, J.; Zou, S.; Lane, M.A.; Roth, G.S.; de Cabo, R. Calorie restriction mimetics: An emerging research field. Aging Cell 2006, 5, 97–108. [Google Scholar] [CrossRef]
- Masoro, E.J. Dietary restriction and aging. J. Am. Geriatr. Soc. 1993, 41, 994–999. [Google Scholar]
- Fontana, L.; Meyer, T.E.; Klein, S.; Holloszy, J.O. Long-term calorie restriction is highly effective in reducing the risk for atherosclerosis in humans. Proc. Natl. Acad. Sci. USA 2004, 101, 6659–6663. [Google Scholar]
- Mattison, J.A.; Roth, G.S.; Beasley, T.M.; Tilmont, E.M.; Handy, A.M.; Herbert, R.L.; Longo, D.L.; Allison, D.B.; Young, J.E.; Bryant, M.; et al. Impact of caloric restriction on health and survival in rhesus monkeys from the NIA study. Nature 2012, 489, 318–321. [Google Scholar] [CrossRef]
- Liao, C.Y.; Rikke, B.A.; Johnson, T.E.; Diaz, V.; Nelson, J.F. Genetic variation in the murine lifespan response to dietary restriction: From life extension to life shortening. Aging Cell 2010, 9, 92–95. [Google Scholar] [CrossRef]
- Wijeyesekera, A.; Selman, C.; Barton, R.H.; Holmes, E.; Nicholson, J.K.; Withers, D.J. Metabotyping of long-lived mice using 1H NMR spectroscopy. J. Proteome. Res. 2012, 11, 2224–2235. [Google Scholar] [CrossRef]
- Selman, C.; Kerrison, N.D.; Cooray, A.; Piper, M.D.; Lingard, S.J.; Barton, R.H.; Schuster, E.F.; Blanc, E.; Gems, D.; Nicholson, J.K.; et al. Coordinated multi-tissue transcriptional and plasma metabonomic profiles following acute caloric restriction in mice. Physiol. Genomics 2006, 27, 187–200. [Google Scholar] [CrossRef]
- Wang, Y.; Lawler, D.; Larson, B.; Ramadan, Z.; Kochhar, S.; Holmes, E.; Nicholson, J.K. Metabonomic investigations of aging and caloric restriction in a life-long dog study. J. Proteome Res. 2007, 6, 1846–1854. [Google Scholar] [CrossRef]
- Rezzi, S.; Martin, F.P.; Shanmuganayagam, D.; Colman, R.J.; Nicholson, J.K.; Weindruch, R. Metabolic shifts due to long-term caloric restriction revealed in nonhuman primates. Exp. Gerontol. 2009, 44, 356–362. [Google Scholar] [CrossRef]
- Boulange, C.L.; Claus, S.P.; Chou, C.J.; Collino, S.; Montoliu, I.; Kochhar, S.; Holmes, E.; Rezzi, S.; Nicholson, J.K.; Dumas, M.E.; et al. Early metabolic adaptation in C57BL/6 mice resistant to high fat diet induced weight gain involves an activation of mitochondrial oxidative pathways. J. Proteome Res. 2013, 12, 1956–1986. [Google Scholar] [CrossRef]
- West, D.B.; Boozer, C.N.; Moody, D.L.; Atkinson, R.L. Dietary obesity in nine inbred mouse strains. Am. J. Physiol. 1992, 262, R1025–R1032. [Google Scholar]
- Champy, M.F.; Selloum, M.; Zeitler, V.; Caradec, C.; Jung, B.; Rousseau, S.; Pouilly, L.; Sorg, T.; Auwerx, J. Genetic background determines metabolic phenotypes in the mouse. Mamm. Genome 2008, 19, 318–331. [Google Scholar] [CrossRef]
- Ni, Y.; Su, M.; Lin, J.; Wang, X.; Qiu, Y.; Zhao, A.; Chen, T.; Jia, W. Metabolic profiling reveals disorder of amino acid metabolism in four brain regions from a rat model of chronic unpredictable mild stress. FEBS Lett. 2008, 582, 2627–2636. [Google Scholar] [CrossRef]
- Coleman, E.A.; Eilertsen, T.B.; Magid, D.J.; Conner, D.A.; Beck, A.; Kramer, A.M. The association between care co-ordination and emergency department use in older managed care enrollees. Int. J. Integr. Care 2002, 2, e03. [Google Scholar]
- Adams, S.H.; Hoppel, C.L.; Lok, K.H.; Zhao, L.; Wong, S.W.; Minkler, P.E.; Hwang, D.H.; Newman, J.W.; Garvey, W.T. Plasma acylcarnitine profiles suggest incomplete long-chain fatty acid beta-oxidation and altered tricarboxylic acid cycle activity in type 2 diabetic African-American women. J. Nutr. 2009, 139, 1073–1081. [Google Scholar] [CrossRef]
- Noland, R.C.; Koves, T.R.; Seiler, S.E.; Lum, H.; Lust, R.M.; Ilkayeva, O.; Stevens, R.D.; Hegardt, F.G.; Muoio, D.M. Carnitine insufficiency caused by aging and overnutrition compromises mitochondrial performance and metabolic control. J. Biol. Chem. 2009, 284, 22840–22852. [Google Scholar] [CrossRef]
- Bauer, M.; Hamm, A.C.; Bonaus, M.; Jacob, A.; Jaekel, J.; Schorle, H.; Pankratz, M.J.; Katzenberger, J.D. Starvation response in mouse liver shows strong correlation with life-span-prolonging processes. Physiol. Genomics 2004, 17, 230–244. [Google Scholar] [CrossRef]
- Dhahbi, J.M.; Mote, P.L.; Wingo, J.; Tillman, J.B.; Walford, R.L.; Spindler, S.R. Calories and aging alter gene expression for gluconeogenic, glycolytic, and nitrogen-metabolizing enzymes. Am. J. Physiol. 1999, 277, E352–E360. [Google Scholar]
- Kabe, Y.; Ohmori, M.; Shinouchi, K.; Tsuboi, Y.; Hirao, S.; Azuma, M.; Watanabe, H.; Okura, I.; Handa, H. Porphyrin accumulation in mitochondria is mediated by 2-oxoglutarate carrier. J. Biol. Chem. 2006, 281, 31729–31735. [Google Scholar] [CrossRef]
- Barger, J.L.; Kayo, T.; Vann, J.M.; Arias, E.B.; Wang, J.; Hacker, T.A.; Wang, Y.; Raederstorff, D.; Morrow, J.D.; Leeuwenburgh, C.; et al. A low dose of dietary resveratrol partially mimics caloric restriction and retards aging parameters in mice. PLoS One 2008, 3, 2264–2272. [Google Scholar] [CrossRef]
- Rider, M.H.; Bertrand, L.; Vertommen, D.; Michels, P.A.; Rousseau, G.G.; Hue, L. 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase: head-to-head with a bifunctional enzyme that controls glycolysis. Biochem. J. 2004, 381, 561–579. [Google Scholar] [CrossRef]
- Pilloff, D.; Dabovic, K.; Romanowski, M.J.; Bonanno, J.B.; Doherty, M.; Burley, S.K.; Leyh, T.S. The kinetic mechanism of phosphomevalonate kinase. J. Biol. Chem. 2003, 278, 4510–4515. [Google Scholar]
- Richards, S.E.; Wang, Y.; Lawler, D.; Kochhar, S.; Holmes, E.; Lindon, J.C.; Nicholson, J.K. Self-modeling curve resolution recovery of temporal metabolite signal modulation in NMR spectroscopic data sets: Application to a life-long caloric restriction study in dogs. Anal. Chem. 2008, 80, 4876–4885. [Google Scholar] [CrossRef]
- Kochhar, S.; Jacobs, D.M.; Ramadan, Z.; Berruex, F.; Fuerholz, A.; Fay, L.B. Probing gender-specific metabolism differences in humans by nuclear magnetic resonance-based metabonomics. Anal. Biochem. 2006, 352, 274–281. [Google Scholar] [CrossRef]
- Pereira, S.P.; Cassell, T.B.; Engelman, J.L.; Sladen, G.E.; Murphy, G.M.; Dowling, R.H. Plasma arachidonic acid-rich phospholipids in Crohn’s disease: Response to treatment. Clin. Sci. 1996, 91, 509–512. [Google Scholar]
- Bartke, A. Insulin resistance and cognitive aging in long-lived and short-lived mice. J. Gerontol. A—Biol. 2005, 60, 133–134. [Google Scholar] [CrossRef]
- Barger, J.L.; Kayo, T.; Pugh, T.D.; Prolla, T.A.; Weindruch, R. Short-term consumption of a resveratrol-containing nutraceutical mixture mimics gene expression of long-term caloric restriction in mouse heart. Exp. Gerontol. 2008, 43, 859–866. [Google Scholar] [CrossRef]
- Newgard, C.B.; An, J.; Bain, J.R.; Muehlbauer, M.J.; Stevens, R.D.; Lien, L.F.; Haqq, A.M.; Shah, S.H.; Arlotto, M.; Slentz, C.A.; et al. A branched-chain amino acid-related metabolic signature that differentiates obese and lean humans and contributes to insulin resistance. Cell Metab. 2009, 9, 311–326. [Google Scholar] [CrossRef]
- Pass, G.J.; Becker, W.; Kluge, R.; Linnartz, K.; Plum, L.; Giesen, K.; Joost, H.G. Effect of hyperinsulinemia and type 2 diabetes-like hyperglycemia on expression of hepatic cytochrome p450 and glutathione s-transferase isoforms in a New Zealand obese-derived mouse backcross population. J. Pharmacol. Exp. Ther. 2002, 302, 442–450. [Google Scholar] [CrossRef]
- Wu, G.; Fang, Y.Z.; Yang, S.; Lupton, J.R.; Turner, N.D. Glutathione metabolism and its implications for health. J. Nutr. 2004, 134, 489–492. [Google Scholar]
- Yu, Z.; Zhai, G.; Singmann, P.; He, Y.; Xu, T.; Prehn, C.; Romisch-Margl, W.; Lattka, E.; Gieger, C.; Soranzo, N.; et al. Human serum metabolic profiles are age dependent. Aging Cell 2012, 11, 960–967. [Google Scholar] [CrossRef]
- Biesalski, H.K.; Nohr, D. New aspects in vitamin a metabolism: The role of retinyl esters as systemic and local sources for retinol in mucous epithelia. J. Nutr. 2004, 134, 3453S–3457S. [Google Scholar]
- Zoeller, R.A.; Lake, A.C.; Nagan, N.; Gaposchkin, D.P.; Legner, M.A.; Lieberthal, W. Plasmalogens as endogenous antioxidants: Somatic cell mutants reveal the importance of the vinyl ether. Biochem. J. 1999, 338, 769–776. [Google Scholar] [CrossRef]
- Wallner, S.; Schmitz, G. Plasmalogens the neglected regulatory and scavenging lipid species. Chem. Phys. Lipids 2011, 164, 573–589. [Google Scholar] [CrossRef]
- Scott, B.L.; Bazan, N.G. Membrane docosahexaenoate is supplied to the developing brain and retina by the liver. Proc. Natl. Acad. Sci. USA 1989, 86, 2903–2907. [Google Scholar] [CrossRef]
- Pugh, T.D.; Klopp, R.G.; Weindruch, R. Controlling caloric consumption: Protocols for rodents and rhesus monkeys. Neurobiol. Aging 1999, 20, 157–165. [Google Scholar] [CrossRef]
- Lee, C.K.; Allison, D.B.; Brand, J.; Weindruch, R.; Prolla, T.A. Transcriptional profiles associated with aging and middle age-onset caloric restriction in mouse hearts. Proc. Natl. Acad. Sci. USA 2002, 99, 14988–14993. [Google Scholar] [CrossRef]
- Benjamini, Y.; Yekutieli, D. The control of the false discovery rate in multiple hypotheses testing under dependency. Ann. Stat. 2001, 29, 1165–1188. [Google Scholar] [CrossRef]
- Kim, S.Y.; Volsky, D.J. PAGE: parametric analysis of gene set enrichment. BMC Bioinformatics 2005, 8, 144–156. [Google Scholar]
- NCBI-GEO. Available online: http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE50789 (accessed on 28 June 2013).
- Baur, P.; Martin, F.P.; Gruber, L.; Bosco, N.; Brahmbhatt, V.; Collino, S.; Guy, P.; Montoliu, I.; Rozman, J.; Klingenspor, M.; et al. Metabolic phenotyping of the Crohn’s disease-like IBD etiopathology in the TNF(DeltaARE/WT) mouse model. J. Proteome Res. 2011, 10, 5523–5535. [Google Scholar] [CrossRef]
- Folch, J.; Lees, M.; Sloane Stanley, G.H. A simple method for the isolation and purification of total lipides from animal tissues. J. Biol. Chem. 1957, 226, 497–509. [Google Scholar]
- Wiklund, S.; Johansson, E.; Sjostrom, L.; Mellerowicz, E.J.; Edlund, U.; Shockcor, J.P.; Gottfries, J.; Moritz, T.; Trygg, J. Visualization of GC/TOF-MS-based metabolomics data for identification of biochemically interesting compounds using OPLS class models. Anal. Chem. 2008, 80, 115–122. [Google Scholar] [CrossRef]
Appendix
| Weeks | Body weight | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 8 | 10 | 12 | 14 | 16 | 18 | 20 | 22 | ||
| 129S1/SvlmJ | Control | 22.8 ± 1.26 | 23.7 ± 1.41 | 27.4 ± 3.66 | 29.0 ± 4.17 | 28.8 ± 4.04 | 28.1 ± 3.85 | 28.2 ± 4.35 | 28.2 ± 4.11 |
| CR | 22.5 ± 0.43 | 22.8 ± 0.41 | 25.2 ± 1.33 | 26.5 ± 1.40 | 27.0 ± 1.28 | 25.3 ± 1.32 | 24.1 *** ± 1.35 | 23.7 *** ± 1.37 | |
| C57BL6/J | Control | 24.4 ± 2.11 | 24.1 ± 1.47 | 26.6 ± 1.98 | 28.5 ± 2.24 | 28.4 ± 2.39 | 28.7 ± 2.39 | 29.5 ± 2.37 | 30.5 ± 2.55 |
| CR | 25.6± 1.53 | 25.6 ± 1.05 | 26.1 ± 1.01 | 27.4± 0.96 | 27.6± 1.13 | 27.3± 1.01 | 23.6 *** ± 0.71 | 23.6 *** ± 0.77 | |
| C3H/HeJ | Control | 25.8± 1.03 | 28.3 ± 1.14 | 30.1 ± 1.59 | 31.6 ± 1.99 | 32.9 ± 2.83 | 33.4 ± 3.14 | 33.6 ± 3.77 | 33.7 ± 3.73 |
| CR | 24.2 ***± 0.82 | 22.1 *** ± 0.72 | 22.8 *** ± 0.89 | 24.2 *** ± 0.76 | 26.7 *** ± 0.73 | 28.4 *** ± 0.83 | 28.4 *** ± 0.73 | 28.4 *** ± 0.78 | |
| CBA/J | Control | 26.3 ± 0.90 | 29.3 ± 0.95 | 29.7 ± 0.99 | 29.8 ± 1.17 | 34.5 ± 1.54 | 35.6 ± 1.41 | 35.9 ± 1.21 | 35.9 ± 1.36 |
| CR | 25.1 *** ± 1.27 | 21.6 *** ± 1.04 | 21.4 *** ± 0.99 | 21.4 *** ± 0.96 | 24.0 *** ± 0.59 | 25.6 *** ± 1.04 | 25.7 *** ± 1.45 | 26.5 *** ± 3.67 | |
| DBA/2J | Control | 24.9 ± 0.79 | 27.2 ± 1.54 | 27.8 ± 1.11 | 30.2 ± 1.85 | 31.7 ± 2.09 | 31.9 ± 2.27 | 32.1 ± 2.24 | 31.4 ± 2.48 |
| CR | 24.4 ± 1.05 | 22.1 *** ± 1.11 | 22.8 *** ± 1.22 | 24.1 *** ± 1.36 | 25.5 *** ± 1.41 | 25.9 *** ± 1.53 | 25.8 *** ± 1.55 | 25.0 *** ± 1.54 | |
| JC3F1/J | Control | 27.8 ± 0.79 | 33.6 ± 0.55 | 35.7 ± 0.60 | 36.6 ± 1.03 | 36.1 ± 0.99 | 36.3 ± 0.94 | 38.3 ± 1.22 | 37.5 ± 1.04 |
| CR | 28.4 ± 0.94 | 26.2 *** ± 0.67 | 27.3 *** ± 1.05 | 28.0 *** ± 1.11 | 27.5 *** ± 1.24 | 27.2 *** ± 1.45 | 29.2 *** ± 1.59 | 28.4 *** ± 1.47 | |
| Gene Symbol | Gene Title | p-value (129) | FDR (129) | FC (129) | p-value (B6) | FDR (B6) | FC (B6) | p-value (C3H) | FDR (C3H) | FC (C3H) | p-value (CBA) | FDR (CBA) | FC (CBA) | p-value (DBA) | FDR (DBA) | FC (DBA) | p-value (F1) | FDR (F1) | FC (F1) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 4931406C07Rik | RIKEN cDNA 4931406C07 gene | 0.000 | 0.003 | −1.380 | 0.001 | 0.030 | −1.290 | 0.004 | 0.071 | −1.290 | 0.000 | 0.000 | −1.900 | 0.000 | 0.003 | −1.680 | 0.000 | 0.009 | −1.400 |
| 9530008L14Rik | RIKEN cDNA 9530008L14 gene | 0.000 | 0.002 | −2.010 | 0.000 | 0.013 | −1.430 | 0.001 | 0.031 | −1.310 | 0.000 | 0.000 | −1.840 | 0.000 | 0.008 | −1.890 | 0.000 | 0.001 | −1.390 |
| Acadm | Acyl-Coenzyme A dehydrogenase, medium chain | 0.000 | 0.000 | −1.850 | 0.000 | 0.003 | −1.430 | 0.001 | 0.030 | −1.270 | 0.009 | 0.033 | −1.210 | 0.002 | 0.088 | −1.510 | 0.000 | 0.007 | −1.430 |
| Acss2 | acyl-CoA synthetase short-chain family member 2 | 0.006 | 0.099 | 4.230 | 0.000 | 0.014 | 4.340 | 0.000 | 0.002 | 3.600 | 0.000 | 0.002 | 2.400 | 0.000 | 0.001 | 5.640 | 0.000 | 0.001 | 6.270 |
| Adh4 | alcohol dehydrogenase 4 (class II), pi polypeptide | 0.000 | 0.011 | −1.570 | 0.002 | 0.035 | −1.460 | 0.007 | 0.106 | −1.310 | 0.000 | 0.002 | −2.250 | 0.000 | 0.034 | −1.800 | 0.000 | 0.001 | −1.640 |
| Aldoc | aldolase C, fructose-bisphosphate | 0.000 | 0.000 | 4.140 | 0.000 | 0.001 | 2.230 | 0.004 | 0.074 | 1.590 | 0.000 | 0.000 | 2.190 | 0.000 | 0.001 | 2.080 | 0.002 | 0.062 | 1.780 |
| Asl | argininosuccinate lyase | 0.000 | 0.003 | 1.700 | 0.003 | 0.048 | 1.450 | 0.000 | 0.001 | 1.670 | 0.000 | 0.000 | 2.740 | 0.000 | 0.004 | 1.920 | 0.002 | 0.059 | 1.660 |
| Ass1 | argininosuccinate synthetase 1 | 0.000 | 0.005 | 2.330 | 0.001 | 0.026 | 1.670 | 0.000 | 0.003 | 2.010 | 0.000 | 0.000 | 5.340 | 0.000 | 0.000 | 2.950 | 0.001 | 0.032 | 2.010 |
| Atp5g1 | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit c1 (subunit 9) | 0.001 | 0.022 | 1.600 | 0.003 | 0.043 | 1.510 | 0.000 | 0.000 | 1.410 | 0.000 | 0.000 | 1.770 | 0.000 | 0.017 | 1.830 | 0.000 | 0.006 | 1.520 |
| Chrna2 | cholinergic receptor, nicotinic, alpha polypeptide 2 (neuronal) | 0.000 | 0.001 | −5.230 | 0.000 | 0.002 | −3.170 | 0.000 | 0.003 | −2.670 | 0.000 | 0.000 | −4.260 | 0.002 | 0.078 | −4.090 | 0.002 | 0.052 | −1.960 |
| D16H22S680E | DNA segment, Chr 16, human D22S680E, expressed | 0.000 | 0.016 | 1.640 | 0.000 | 0.000 | 2.140 | 0.000 | 0.012 | 1.570 | 0.000 | 0.000 | 1.760 | 0.000 | 0.029 | 1.700 | 0.001 | 0.029 | 1.310 |
| Dhcr7 | 7-dehydrocholesterol reductase | 0.000 | 0.018 | 2.970 | 0.001 | 0.023 | 2.430 | 0.000 | 0.013 | 2.950 | 0.003 | 0.014 | 1.920 | 0.000 | 0.029 | 2.870 | 0.000 | 0.005 | 3.160 |
| Dlat | dihydrolipoamide S-acetyltransferase (E2 component of pyruvate dehydrogenase complex) | 0.001 | 0.026 | 1.530 | 0.000 | 0.009 | 1.470 | 0.001 | 0.022 | 1.830 | 0.000 | 0.004 | −1.710 | 0.000 | 0.029 | 1.960 | 0.001 | 0.039 | 1.970 |
| Echdc1 | enoyl Coenzyme A hydratase domain containing 1 | 0.005 | 0.090 | 1.780 | 0.000 | 0.002 | 1.790 | 0.002 | 0.051 | 1.680 | 0.000 | 0.000 | −2.410 | 0.000 | 0.024 | 1.890 | 0.005 | 0.100 | 1.970 |
| Eci1 | enoyl-Coenzyme A delta isomerase 1 | 0.000 | 0.000 | −2.720 | 0.000 | 0.000 | −1.800 | 0.002 | 0.049 | −1.440 | 0.005 | 0.022 | 1.310 | 0.000 | 0.030 | −1.620 | 0.004 | 0.087 | −1.520 |
| Fbxo18 | F-box protein 18 | 0.000 | 0.009 | 1.590 | 0.000 | 0.013 | 1.410 | 0.000 | 0.010 | 1.400 | 0.000 | 0.000 | 2.160 | 0.004 | 0.121 | 1.260 | 0.000 | 0.011 | 1.430 |
| Fdps | farnesyl diphosphate synthetase | 0.001 | 0.038 | 6.470 | 0.001 | 0.023 | 5.310 | 0.000 | 0.011 | 3.510 | 0.009 | 0.033 | 1.920 | 0.000 | 0.000 | 7.660 | 0.000 | 0.009 | 4.870 |
| Fh1 | fumarate hydratase 1 | 0.000 | 0.021 | 1.740 | 0.000 | 0.011 | 1.440 | 0.000 | 0.001 | 1.910 | 0.002 | 0.011 | 1.430 | 0.000 | 0.004 | 2.080 | 0.000 | 0.003 | 2.010 |
| Foxa3 | forkhead box A3 | 0.002 | 0.042 | −1.940 | 0.000 | 0.001 | −1.980 | 0.000 | 0.005 | −2.580 | 0.000 | 0.002 | −1.820 | 0.000 | 0.011 | −2.020 | 0.000 | 0.001 | −2.440 |
| Gls2 | glutaminase 2 (liver, mitochondrial) | 0.000 | 0.001 | 3.080 | 0.000 | 0.000 | 2.430 | 0.000 | 0.000 | 2.210 | 0.000 | 0.000 | 2.700 | 0.000 | 0.025 | 2.050 | 0.000 | 0.003 | 2.080 |
| Got1 | glutamate oxaloacetate transaminase 1, soluble | 0.000 | 0.009 | 3.130 | 0.000 | 0.001 | 2.310 | 0.000 | 0.000 | 4.230 | 0.000 | 0.002 | 4.020 | 0.004 | 0.121 | 4.300 | 0.000 | 0.000 | 4.130 |
| Gsta4 | glutathione S-transferase, alpha 4 | 0.000 | 0.000 | 2.510 | 0.000 | 0.000 | 2.690 | 0.003 | 0.070 | 1.400 | 0.000 | 0.000 | 1.910 | 0.000 | 0.035 | 1.510 | 0.000 | 0.018 | 1.760 |
| Gstm2 | glutathione S-transferase, mu 2 | 0.001 | 0.040 | 2.110 | 0.000 | 0.007 | 1.680 | 0.002 | 0.054 | 1.800 | 0.001 | 0.006 | 1.730 | 0.000 | 0.004 | 3.020 | 0.000 | 0.001 | 2.680 |
| Gstm6 | glutathione S-transferase, mu 6 | 0.001 | 0.023 | 2.030 | 0.000 | 0.002 | 2.030 | 0.000 | 0.015 | 2.300 | 0.002 | 0.009 | 1.840 | 0.000 | 0.001 | 3.030 | 0.000 | 0.000 | 2.880 |
| Heatr1 | HEAT repeat containing 1 | 0.000 | 0.001 | 2.110 | 0.000 | 0.011 | 1.520 | 0.000 | 0.000 | 1.690 | 0.002 | 0.009 | 1.370 | 0.000 | 0.003 | 1.550 | 0.000 | 0.001 | 1.750 |
| Hgd | homogentisate 1, 2-dioxygenase | 0.006 | 0.101 | 1.340 | 0.000 | 0.011 | 1.280 | 0.000 | 0.000 | 1.630 | 0.000 | 0.000 | 2.140 | 0.000 | 0.022 | 1.820 | 0.000 | 0.002 | 1.760 |
| Hsd3b5 | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 5 | 0.009 | 0.129 | −2.140 | 0.003 | 0.043 | −13.12 | 0.000 | 0.002 | −3.790 | 0.000 | 0.002 | −2.400 | 0.000 | 0.003 | −5.730 | 0.000 | 0.019 | −12.07 |
| Hsp90aa1 | heat shock protein 90, alpha (cytosolic), class A member 1 | 0.002 | 0.044 | −1.790 | 0.000 | 0.001 | −2.030 | 0.000 | 0.016 | −1.740 | 0.000 | 0.000 | −3.640 | 0.006 | 0.149 | −2.390 | 0.000 | 0.008 | −2.220 |
| Ikbkg | inhibitor of kappaB kinase gamma | 0.000 | 0.002 | −1.480 | 0.000 | 0.000 | −1.640 | 0.005 | 0.085 | −1.370 | 0.000 | 0.000 | −1.750 | 0.002 | 0.080 | −1.690 | 0.002 | 0.050 | −1.380 |
| Klc4 | kinesin light chain 4 | 0.000 | 0.001 | 1.750 | 0.000 | 0.001 | 1.460 | 0.000 | 0.003 | 1.490 | 0.001 | 0.004 | 1.410 | 0.000 | 0.032 | 1.550 | 0.000 | 0.006 | 1.370 |
| Lonp2 | lon peptidase 2, peroxisomal | 0.000 | 0.000 | −2.040 | 0.000 | 0.000 | −1.950 | 0.001 | 0.028 | −1.450 | 0.001 | 0.005 | −1.380 | 0.000 | 0.004 | −1.790 | 0.000 | 0.002 | −1.550 |
| Mdh2 | malate dehydrogenase 2, NAD (mitochondrial) | 0.003 | 0.063 | 1.540 | 0.002 | 0.032 | 1.540 | 0.000 | 0.017 | 1.590 | 0.000 | 0.000 | 1.810 | 0.004 | 0.126 | 2.030 | 0.001 | 0.043 | 2.080 |
| Mgll | monoglyceride lipase | 0.000 | 0.000 | −2.260 | 0.000 | 0.000 | −2.730 | 0.000 | 0.007 | −2.350 | 0.000 | 0.000 | −2.870 | 0.000 | 0.010 | −2.000 | 0.000 | 0.001 | −2.210 |
| Mmd | monocyte to macrophage differentiation-associated | 0.000 | 0.000 | −1.760 | 0.001 | 0.023 | −1.270 | 0.004 | 0.075 | −1.320 | 0.000 | 0.002 | −1.780 | 0.001 | 0.064 | −1.300 | 0.000 | 0.001 | −1.570 |
| Mthfd1 | methylenetetrahydrofolate dehydrogenase (NADP+ dependent), methenyltetrahydrofolate cyclohydrolase, formyltetrahydrofolate synthase | 0.003 | 0.068 | 1.510 | 0.000 | 0.008 | 1.680 | 0.000 | 0.022 | 1.410 | 0.000 | 0.000 | 2.820 | 0.001 | 0.054 | 1.740 | 0.009 | 0.137 | 1.380 |
| Nadsyn1 | NAD synthetase 1 | 0.000 | 0.008 | 1.790 | 0.000 | 0.008 | 1.610 | 0.004 | 0.072 | 1.380 | 0.000 | 0.000 | 1.570 | 0.003 | 0.099 | 1.610 | 0.000 | 0.024 | 1.360 |
| Ndufs8 | NADH dehydrogenase (ubiquinone) Fe-S protein 8 | 0.000 | 0.000 | 1.530 | 0.008 | 0.085 | 1.270 | 0.001 | 0.025 | 1.260 | 0.006 | 0.025 | −1.210 | 0.004 | 0.126 | 1.310 | 0.000 | 0.016 | 1.260 |
| Ngef | neuronal guanine nucleotide exchange factor | 0.000 | 0.002 | 2.140 | 0.000 | 0.000 | 2.100 | 0.000 | 0.003 | 1.470 | 0.000 | 0.000 | 1.930 | 0.003 | 0.094 | 1.360 | 0.001 | 0.027 | 1.340 |
| Nkiras2 | NFKB inhibitor interacting Ras-like protein 2 | 0.009 | 0.129 | 1.460 | 0.001 | 0.023 | 1.340 | 0.000 | 0.009 | 1.330 | 0.000 | 0.000 | 1.890 | 0.001 | 0.051 | 1.430 | 0.000 | 0.010 | 1.490 |
| Nnmt | nicotinamide N-methyltransferase | 0.000 | 0.001 | 12.500 | 0.000 | 0.003 | 2.890 | 0.000 | 0.020 | 20.330 | 0.000 | 0.001 | 4.000 | 0.000 | 0.002 | 18.800 | 0.000 | 0.011 | 5.630 |
| Nol8 | nucleolar protein 8 | 0.002 | 0.043 | −1.550 | 0.003 | 0.044 | −1.390 | 0.000 | 0.010 | −1.650 | 0.000 | 0.002 | −1.680 | 0.003 | 0.094 | −1.660 | 0.001 | 0.027 | −1.730 |
| Pa2g4 | proliferation-associated 2G4 | 0.000 | 0.003 | −1.400 | 0.000 | 0.001 | −2.010 | 0.000 | 0.011 | −1.620 | 0.000 | 0.000 | −1.610 | 0.001 | 0.065 | −1.450 | 0.000 | 0.005 | −1.520 |
| Paox | polyamine oxidase (exo-N4-amino) | 0.000 | 0.014 | 2.960 | 0.000 | 0.008 | 2.930 | 0.000 | 0.002 | 2.240 | 0.000 | 0.000 | 1.530 | 0.000 | 0.001 | 2.390 | 0.000 | 0.000 | 2.480 |
| Paqr9 | progestin and adipoQ receptor family member IX | 0.000 | 0.003 | −4.730 | 0.000 | 0.000 | −3.590 | 0.001 | 0.025 | −1.610 | 0.000 | 0.000 | −3.550 | 0.005 | 0.131 | −3.970 | 0.000 | 0.008 | −1.770 |
| Pcca | propionyl-Coenzyme A carboxylase, alpha polypeptide | 0.000 | 0.018 | 1.510 | 0.000 | 0.005 | 1.330 | 0.000 | 0.001 | 1.390 | 0.000 | 0.000 | 1.840 | 0.001 | 0.045 | 1.450 | 0.000 | 0.009 | 1.480 |
| Pctp | phosphatidylcholine transfer protein | 0.000 | 0.000 | −3.120 | 0.000 | 0.000 | −1.610 | 0.001 | 0.040 | −1.850 | 0.000 | 0.001 | −2.950 | 0.000 | 0.027 | −1.960 | 0.001 | 0.024 | −1.380 |
| Pdilt | protein disulfide isomerase-like, testis expressed | 0.000 | 0.003 | −3.470 | 0.001 | 0.023 | −2.850 | 0.000 | 0.018 | −2.100 | 0.000 | 0.000 | −3.690 | 0.003 | 0.102 | −2.180 | 0.009 | 0.135 | −1.580 |
| Pdzk1 | PDZ domain containing 1 | 0.000 | 0.002 | −1.550 | 0.001 | 0.028 | −1.410 | 0.000 | 0.002 | −1.470 | 0.004 | 0.018 | −1.300 | 0.001 | 0.061 | −1.480 | 0.000 | 0.009 | −1.310 |
| Pemt | phosphatidylethanolamine N-methyltransferase | 0.002 | 0.041 | 1.430 | 0.003 | 0.043 | 1.430 | 0.000 | 0.000 | 1.620 | 0.002 | 0.012 | 1.460 | 0.001 | 0.054 | 1.640 | 0.000 | 0.005 | 1.760 |
| Pfkfb1 | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 0.000 | 0.000 | 1.780 | 0.009 | 0.093 | 1.370 | 0.001 | 0.041 | 1.530 | 0.000 | 0.000 | 2.560 | 0.000 | 0.024 | 1.820 | 0.002 | 0.049 | 1.470 |
| Pgam1 | phosphoglycerate mutase 1 | 0.000 | 0.014 | 1.680 | 0.000 | 0.003 | 1.420 | 0.000 | 0.005 | 1.390 | 0.003 | 0.016 | 1.400 | 0.000 | 0.005 | 1.640 | 0.000 | 0.003 | 1.500 |
| Pgk1 | phosphoglycerate kinase 1 | 0.001 | 0.034 | 1.450 | 0.000 | 0.002 | 1.180 | 0.000 | 0.006 | 1.230 | 0.000 | 0.000 | 1.700 | 0.000 | 0.036 | 1.410 | 0.001 | 0.044 | 1.290 |
| Pla2g16 | phospholipase A2, group XVI | 0.000 | 0.006 | 2.180 | 0.002 | 0.033 | 1.840 | 0.000 | 0.003 | 1.590 | 0.000 | 0.000 | 2.210 | 0.000 | 0.038 | 2.060 | 0.000 | 0.005 | 1.930 |
| Pmvk | phosphomevalonate kinase | 0.002 | 0.044 | 2.380 | 0.001 | 0.019 | 4.340 | 0.000 | 0.010 | 3.110 | 0.001 | 0.005 | 1.960 | 0.000 | 0.001 | 4.520 | 0.000 | 0.003 | 4.000 |
| Ppif | peptidylprolyl isomerase F (cyclophilin F) | 0.000 | 0.004 | 1.790 | 0.001 | 0.024 | 1.390 | 0.001 | 0.027 | 1.320 | 0.000 | 0.000 | 2.080 | 0.001 | 0.048 | 1.580 | 0.000 | 0.006 | 1.330 |
| Ppp1r3g | protein phosphatase 1, regulatory (inhibitor) subunit 3G | 0.000 | 0.013 | 2.540 | 0.000 | 0.001 | 8.450 | 0.000 | 0.018 | 3.460 | 0.000 | 0.000 | 3.080 | 0.001 | 0.066 | 2.750 | 0.001 | 0.045 | 4.010 |
| Retsat | retinol saturase (all trans retinol 13,14 reductase) | 0.000 | 0.000 | −3.880 | 0.000 | 0.000 | −1.590 | 0.001 | 0.026 | −1.640 | 0.007 | 0.027 | −1.400 | 0.001 | 0.047 | −1.820 | 0.004 | 0.091 | −1.450 |
| Rilp | Rab interacting lysosomal protein | 0.009 | 0.128 | 1.410 | 0.000 | 0.010 | 1.350 | 0.007 | 0.105 | 1.420 | 0.000 | 0.000 | 2.060 | 0.000 | 0.008 | 1.630 | 0.002 | 0.049 | 1.310 |
| Ropn1l | ropporin 1-like | 0.000 | 0.004 | 2.030 | 0.000 | 0.012 | 1.590 | 0.007 | 0.108 | 1.520 | 0.003 | 0.013 | 1.650 | 0.000 | 0.014 | 1.950 | 0.006 | 0.109 | 1.420 |
| Sdhd | succinate dehydrogenase complex, subunit D, integral membrane protein | 0.001 | 0.039 | 1.570 | 0.003 | 0.049 | 1.250 | 0.000 | 0.002 | 1.270 | 0.000 | 0.000 | 1.990 | 0.000 | 0.004 | 1.710 | 0.001 | 0.027 | 1.340 |
| Sds | serine dehydratase | 0.000 | 0.008 | 2.080 | 0.000 | 0.000 | 2.620 | 0.000 | 0.009 | 2.650 | 0.001 | 0.009 | 2.210 | 0.002 | 0.089 | 2.190 | 0.000 | 0.002 | 2.850 |
| Selenbp1 | selenium binding protein 1 | 0.000 | 0.001 | 2.430 | 0.000 | 0.000 | 2.240 | 0.000 | 0.005 | 1.800 | 0.003 | 0.015 | 1.810 | 0.004 | 0.121 | 1.530 | 0.000 | 0.000 | 1.800 |
| Slc25a11 | solute carrier family 25 (mitochondrial carrier oxoglutarate carrier), member 11 | 0.000 | 0.003 | 1.530 | 0.000 | 0.005 | 1.390 | 0.004 | 0.081 | 1.240 | 0.010 | 0.035 | 1.180 | 0.000 | 0.029 | 1.290 | 0.000 | 0.002 | 1.460 |
| Slc25a12 | solute carrier family 25 (mitochondrial carrier, Aralar), member 12 | 0.001 | 0.022 | 1.810 | 0.000 | 0.001 | 1.800 | 0.003 | 0.064 | 1.420 | 0.000 | 0.000 | 1.920 | 0.000 | 0.006 | 1.780 | 0.001 | 0.029 | 1.510 |
| Slc25a20 | solute carrier family 25 (mitochondrial carnitine/acylcarnitine translocase), member 20 | 0.000 | 0.001 | −2.090 | 0.000 | 0.003 | −1.590 | 0.002 | 0.052 | −1.500 | 0.000 | 0.001 | −2.070 | 0.001 | 0.053 | −1.310 | 0.000 | 0.003 | −1.620 |
| Slc2a4 | solute carrier family 2 (facilitated glucose transporter), member 4 | 0.006 | 0.102 | 2.460 | 0.000 | 0.002 | 3.060 | 0.007 | 0.111 | 1.750 | 0.004 | 0.018 | 2.000 | 0.000 | 0.018 | 2.410 | 0.000 | 0.003 | 2.820 |
| Slc3a1 | solute carrier family 3, member 1 | 0.000 | 0.002 | 3.170 | 0.000 | 0.001 | 1.740 | 0.001 | 0.032 | 2.740 | 0.000 | 0.000 | 3.020 | 0.000 | 0.009 | 2.100 | 0.000 | 0.001 | 2.630 |
| Stt3b | STT3, subunit of the oligosaccharyltransferase complex, homolog B (S. cerevisiae) | 0.000 | 0.002 | −1.310 | 0.000 | 0.000 | −1.410 | 0.001 | 0.027 | −1.220 | 0.000 | 0.000 | −1.440 | 0.003 | 0.106 | −1.330 | 0.000 | 0.006 | −1.260 |
| Tsc22d1 | TSC22 domain family, member 1 | 0.000 | 0.004 | −3.210 | 0.000 | 0.001 | −2.840 | 0.007 | 0.111 | −1.660 | 0.000 | 0.002 | −2.940 | 0.000 | 0.006 | −2.640 | 0.004 | 0.088 | −1.610 |
| Tubb4b | tubulin, beta 4B class IVB | 0.000 | 0.000 | −2.190 | 0.000 | 0.000 | −2.220 | 0.000 | 0.022 | −1.680 | 0.001 | 0.007 | −1.410 | 0.005 | 0.141 | −1.510 | 0.004 | 0.087 | −1.500 |
| Txndc15 | thioredoxin domain containing 15 | 0.001 | 0.024 | −1.350 | 0.001 | 0.018 | −1.280 | 0.000 | 0.017 | −1.170 | 0.000 | 0.000 | −1.750 | 0.003 | 0.096 | −1.380 | 0.000 | 0.006 | −1.490 |
| Ube2d2a | ubiquitin-conjugating enzyme E2D 2A | 0.002 | 0.048 | −1.260 | 0.000 | 0.001 | −1.310 | 0.001 | 0.033 | −1.250 | 0.000 | 0.000 | −1.930 | 0.003 | 0.094 | −1.210 | 0.002 | 0.047 | −1.350 |
| Ube2e2 | ubiquitin-conjugating enzyme E2E 2 | 0.000 | 0.000 | −2.060 | 0.000 | 0.002 | −1.780 | 0.003 | 0.066 | −1.370 | 0.000 | 0.000 | −2.310 | 0.000 | 0.009 | −1.860 | 0.000 | 0.023 | −1.420 |
| Ubxn4 | UBX domain protein 4 | 0.000 | 0.000 | −1.660 | 0.000 | 0.000 | −2.200 | 0.000 | 0.001 | −1.500 | 0.010 | 0.035 | −1.260 | 0.000 | 0.004 | −1.720 | 0.000 | 0.003 | −1.410 |
| Ugt2b35 | UDP glucuronosyltransferase 2 family, polypeptide B35 | 0.000 | 0.008 | −1.660 | 0.000 | 0.003 | −1.520 | 0.000 | 0.004 | −1.840 | 0.000 | 0.000 | −2.630 | 0.001 | 0.062 | −2.000 | 0.000 | 0.007 | −1.690 |
| Vnn1 | vanin 1 | 0.000 | 0.000 | −4.960 | 0.000 | 0.004 | −2.160 | 0.001 | 0.026 | −4.670 | 0.000 | 0.000 | −5.050 | 0.000 | 0.014 | −3.680 | 0.000 | 0.000 | −3.100 |
| Models | Model descriptor | 129S1/SvlmJ | C57BL6/J | C3H/HeJ | CBA/J | DBA/2J | JC3F1/J |
|---|---|---|---|---|---|---|---|
| NMR Diffusion edited | Q2Y | 0.22 | 0.60 | 0.33 | 0.50 | 0.19 | 0.64 |
| plasma | R2X | 0.17 | 0.22 | 0.11 | 0.12 | 0.13 | 0.22 |
| spectra | R2Y | 0.61 | 0.77 | 0.79 | 0.86 | 0.76 | 0.79 |
| Targeted MS Profiling | Q2Y R2X R2Y | - | 0.59 0.24 0.74 | - | - | - | - |
| Models | Model descriptor | C57BL6/J |
|---|---|---|
| NMR urine | Q2Y R2X R2Y | 0.910.91 |
| NMR water phase liver tissue | Q2Y R2X R2Y | 0.42 |
| NMR organic phase liver tissue | Q2Y R2X R2Y | 0.92 |
Supplementary Files
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Collino, S.; Martin, F.-P.J.; Montoliu, I.; Barger, J.L.; Da Silva, L.; Prolla, T.A.; Weindruch, R.; Kochhar, S. Transcriptomics and Metabonomics Identify Essential Metabolic Signatures in Calorie Restriction (CR) Regulation across Multiple Mouse Strains. Metabolites 2013, 3, 881-911. https://doi.org/10.3390/metabo3040881
Collino S, Martin F-PJ, Montoliu I, Barger JL, Da Silva L, Prolla TA, Weindruch R, Kochhar S. Transcriptomics and Metabonomics Identify Essential Metabolic Signatures in Calorie Restriction (CR) Regulation across Multiple Mouse Strains. Metabolites. 2013; 3(4):881-911. https://doi.org/10.3390/metabo3040881
Chicago/Turabian StyleCollino, Sebastiano, François-Pierre J. Martin, Ivan Montoliu, Jamie L. Barger, Laeticia Da Silva, Tomas A. Prolla, Richard Weindruch, and Sunil Kochhar. 2013. "Transcriptomics and Metabonomics Identify Essential Metabolic Signatures in Calorie Restriction (CR) Regulation across Multiple Mouse Strains" Metabolites 3, no. 4: 881-911. https://doi.org/10.3390/metabo3040881
APA StyleCollino, S., Martin, F.-P. J., Montoliu, I., Barger, J. L., Da Silva, L., Prolla, T. A., Weindruch, R., & Kochhar, S. (2013). Transcriptomics and Metabonomics Identify Essential Metabolic Signatures in Calorie Restriction (CR) Regulation across Multiple Mouse Strains. Metabolites, 3(4), 881-911. https://doi.org/10.3390/metabo3040881




