1. Introduction
Lipidomic profiling methods reflect the lipid status of a phenotype at a particular time point [
1,
2,
3] and are therefore valuable tools to improve the understanding the biological roles of lipids. Unlike genes or proteins, the lipid composition can rapidly be influenced by external factors like nutrition or environmental conditions and is alterable even within seconds [
2,
3,
4,
5]. It is not sufficient enough to consider these topics solely on the transcriptional level, because many genes involved in lipid biosynthesis or transport exist in multiple copies, which can compensate a dysfunction or lack in one gene and thus, no effects on the lipid profile can be observed. Moreover, several studies have shown that organisms like the yeast
S. cerevisiae can tolerate great changes in their lipid composition, compensating for example for the absence of one lipid by overproduction of another, without notable effects on their viability [
5,
6].
Despite many mass spectrometry based lipidomics methods developed today [
7], the current knowledge of the lipidome of eukaryotic organisms is still limited. As the lipidome of higher eukaryotic organisms consists of hundreds to thousands of individual molecular species, a model organism is needed, which possesses a relatively simple lipidome, but still reflects the main biosynthetic and metabolic pathways of higher eukaryotic organisms. It should be easy to handle and also, if necessary, easy to manipulate. Another important criterion is a detailed knowledge on gene, protein and also lipid biosynthesis, which enables to fill gaps in the understanding of a complex biological network. Such a suitable eukaryotic organism is yeast, as it fulfills all the requirements listed above [
8,
9].
One of the yeasts investigated best is the common bakers’ yeast,
S. cerevisiae, for which complete genome, as well as detailed protein data, are available. Therefore, many studies have used this model organism for lipidomics studies. One of the major lipid categories of eukaryotic organisms are glycerophospholipids (GPs), which cover diverse biological roles like cell compartmentalization, energy storage and multiple signaling functions. Consequently, they are the subject of many studies, because their biosynthesis and metabolism is very similar to those of higher eukaryotes, with three main exceptions. Firstly, yeast phosphatidylserine (PS) is mainly synthesized by the CDP-DAG pathway and not by PS synthase from phosphatidylethanolamine (PE). Secondly, for phosphatidylcholine (PC) synthesis, an alternative route exists besides the Kennedy-Pathway (CDP-choline), which is the exclusive pathway in mammals. In yeast, the successive methylation of PE to
mono-methyl-phosphatidylethanolamine (MMPE),
di-methyl-phosphatidylethanolamine (DMPE) and finally PC occurs, catalyzed by
N-methyl-transferases [
2,
9]. Thirdly, the difference to mammals is the relatively low abundance of polyunsaturated fatty acids (PUFAs), or rather the complete absence of PUFAs like in
S. cerevisiae.
Numerous studies have been dedicated to understand the role of GPs in
S. cerevisiae. It has been shown that the faultless biosynthesis and metabolism of particular GPs appear to be essential for cell vitality. For instance, mutations in the gene encoding the phosphatidylinositol (PI) synthase are lethal for the organism [
6,
9]. On the other hand, not all GP classes seem to be essential for the survivability of the organism, as defects in genes encoding enzymes involved in PC, PE or PS biosynthesis result in phenotypic changes but are not lethal under laboratory conditions. Adaptive changes of GP composition after applying varying environmental conditions (e.g., elevated growth temperatures) or by supplementation of cell culture media (e.g., with fatty acids) are also investigated in several studies along with changes in lipid profile when comparing mutants with defects in lipid biosynthesis [
10,
11,
12].
The question addressed in this paper is if phylogenetically different yeast strains possess characteristic GP profiles, and if genetically closely related strains show analogies in their GP composition. The profiling was carried out by the previously described HPLC/LIT-FTICRMS-method [
13] including automated data processing by the Profiler-Merger-Viewer software [
14], of which the applicability for different complex GP samples was demonstrated recently [
15,
16].
2. Results and Discussion
Four yeast strains with few genetic analogies were selected to investigate possible characteristic GP profiles based on genetic differences.
Saccharomyces cerevisiae (common baker’s yeast) was selected as the first candidate, because this organism has been investigated in detail on protein, gene as well as on lipid levels.
Kluyveromyces thermotolerans,
Pichia angusta and
Yarrowia lipolytica were chosen for the comparative study, as they are not closely related to each other (see
Figure 1a). A further aim of the study was to reveal analogies in the lipid profile of closely related yeast strains. Accordingly,
Saccharomyces bayanus was chosen as a close relative of
Saccharomyces cerevisiae. To avoid variations in the GP profiles due to changing external conditions, all strains were grown and extracted in parallel, using the same medium batch. Hence, observed characteristics are likely based on genetic differences.
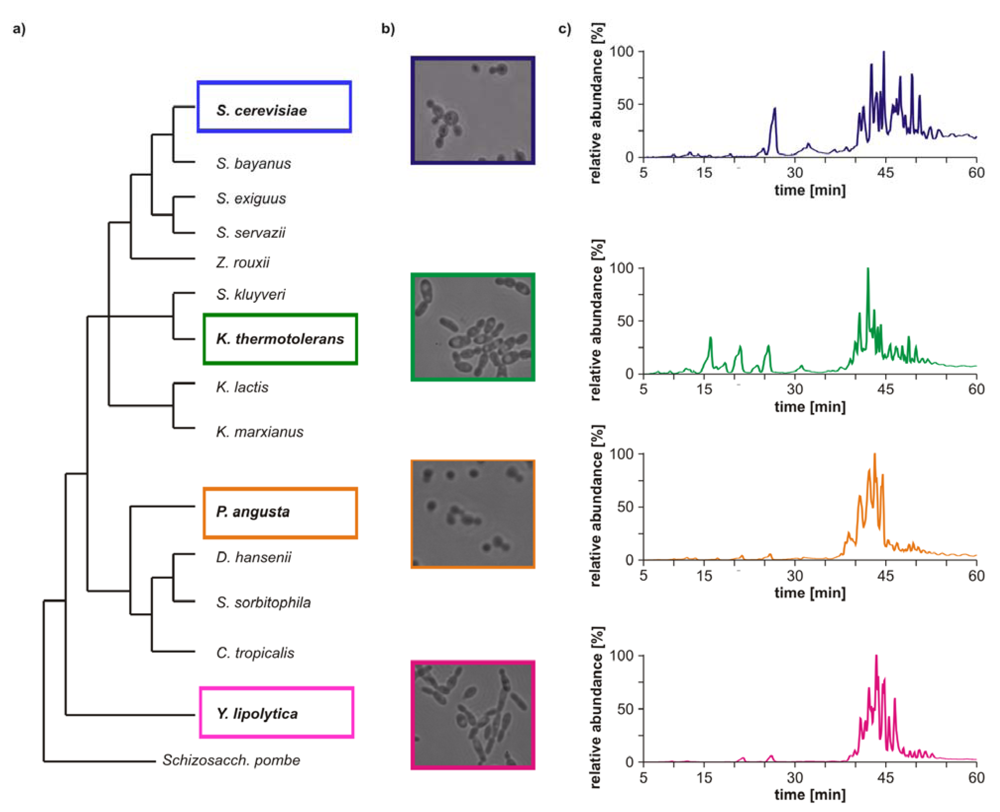
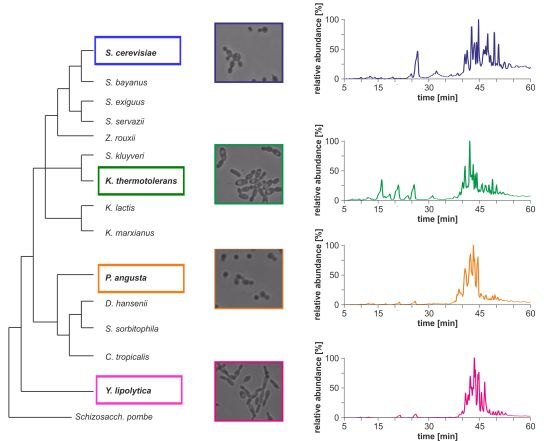
Morphological differences between the four yeast strains can already be observed on a macroscopic level (
Figure 1b) and even without detailed lipid profiling the total ion currents (TICs) of the HPLC/ESI-LIT-FTICRMS-measurements showed different patterns (
Figure 1c). Within a yeast strain, the profile was reproducible in all biological replicates (n = 3; data not shown).
A detailed insight into the complexity of the GP profiles was realized by HPLC/MS(/MS) investigation of the lipid extracts. Using the recently developed aforementioned HPLC/ESI-LIT-FTICRMS method, 106–120 molecular GP species from nine GP classes (cardiolipins (CA), phosphatidic acids (PA), phophatidylethanolamines (PE),
mono-methyl-phosphatidylethanolamines (MMPE),
di-methyl-phosphatidylethanolamines (DMPE), phosphatidylcholines (PC), phosphatidylinositoles (PI), phosphatidylserines (PS), and phosphatidylglycerols (PG)) were relatively quantified in
S. cerevisiae,
S. bayanus,
K. thermotolerans,
P. angusta, and
Y. lipolytica. A comparison of the GP species belonging to the six most abundant GP classes of the five yeast strains is given in
Figure 2. Please note that fatty acid chains are abbreviated (xx:y), with xx the total number of carbon atoms and y the sum of double bonds in the fatty acid chains. The relative amount of one species is calculated in relation to the sum of all species contributing to the same GP class.
Figure 1.
(
a) Phylogeny of hemiascomycetous yeasts based on 15S-rRNA data (adapted from [
17]); (
b) light microscopic pictures of the chosen yeast strains and (
c) sections of the total ion currents (TICs) of the HPLC/ESI-LIT-FTICRMS-measurements of four yeast strains.
These comparative GP profiles show that significant differences in number, distribution and relative amount of the identified GP species exist among the phylogenetically different yeast strains. In general, the number of identified species is less in
S. cerevisiae and
S. bayanus, whereas
K. thermotolerans,
P. angusta and
Y. lipolytica possess a larger variety of GP species. In addition, the number, as well as the distribution, of major GP species is significantly different in the genetically diverse yeasts, whereas the patterns of the related yeasts strains show analogies. In particular,
S. cerevisiae and
S. bayanus possess in general four major species, with rather short acyl chains and a lower number of double bonds. The latter are PE(32:2), PE(34:2), PC(32:2) and PC(34:2), respectively. The yeast
Y. lipolytica possesses also only a few major species in each GP class, but unlike the
Saccharomyces strains, the chain length and degree of unsaturation is considerably higher. In contrast, the lipid profiles of
K. thermotolerans and
P. angusta show a larger variety of GP species in each class. Compared to the three previous yeast strains, the fatty acid chains are longer and have an increased number of double bonds. Compared to each other, this trend is stronger in
P. angusta. All these differences are especially pronounced in the class of CAs (
Figure 2). An exception from this divergence seems to be in the GP classes PS, PI and PG. The major species identified in these classes are very similar for all investigated yeast strains (
Figure 2).
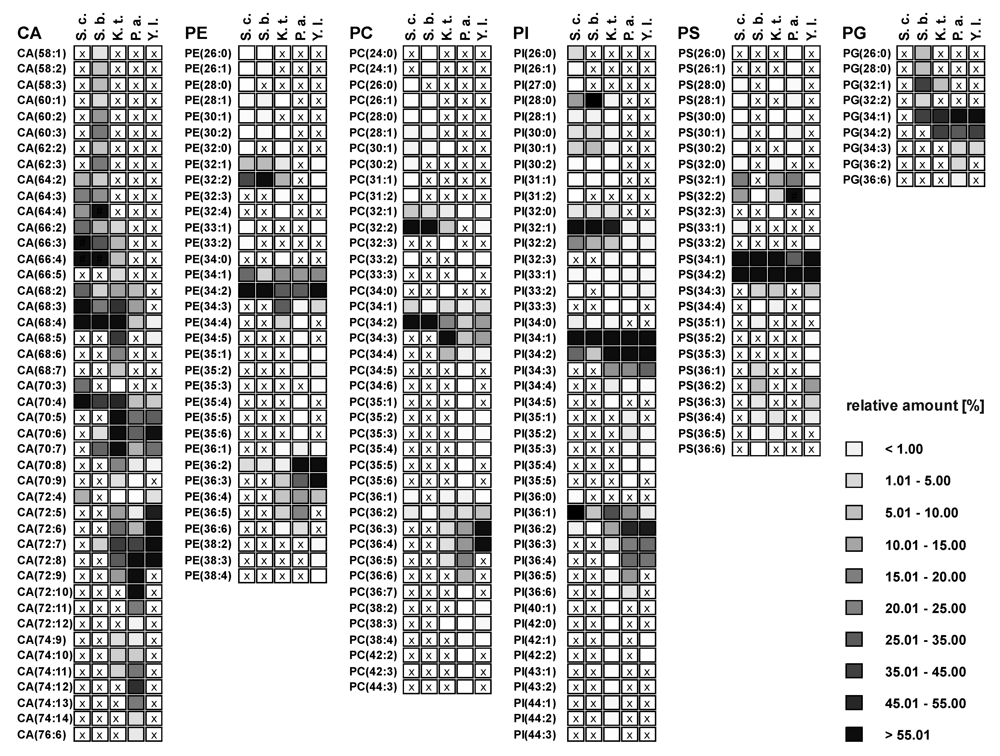
Figure 2.
Overview of species distribution in the six most common GP classes: cardiolipins (CA), phophatidylethanolamines (PE), phosphatidylcholines (PC), phosphatidylinositoles (PI), phosphatidylserines (PS), and phosphatidylglycerols (PG) for S. cerevisiae (S.c.), S. bayanus (S.b.), K. thermotolerans (K.t.), P. angusta (P.a.) and Y. lipolytica (Y.l.). (x indicate not detected GP species).
For a more detailed interpretation, the yeast strains were divided into two groups based on the overall GP pattern. The first group comprises
K. thermotolerans,
P. angusta and
Y. lipolytica, the second group contains
S. cerevisiae and
S. bayanus. The relative amounts of the identified species from the first group are depicted in
Figure 3. For better representation, only species which contributed at least to 5% to the GP profile of a single class are represented (an overview of all identified GPs and their relative amounts is given in
Table S1 of the Supporting Information).
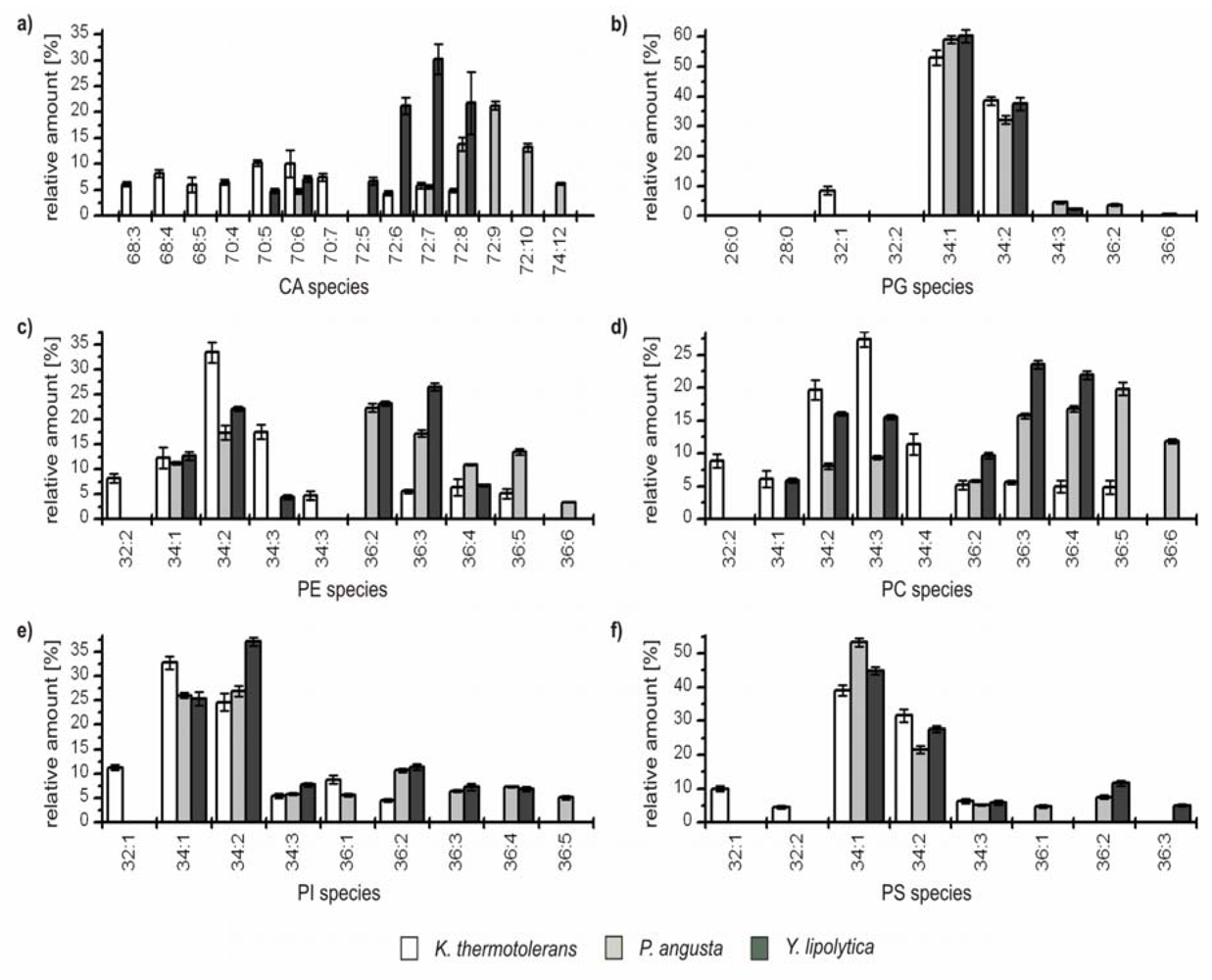
Figure 3.
Relative amounts of the GP species within the six major GP classes (a) cardiolipins (CA); (b) phosphatidylglcerols (PG); (c) phophatidylethanolamines (PE); (d) phosphatidylcholines (PC); (e) phosphatidylinositoles (PI); and (f) phosphatidylserines (PS) for K. thermotolerans (□), P. angusta (■) and Y. lipolytica (■). The relative amount of respective species is referred to 100% peak area of all identified GPs within one class and yeast strain. Only species which contributed to more than 5% of the respective GP class are displayed.
From this perspective, characteristic differences within the group can be recognized. The major GP species of
K. thermotolerans have 34 carbon atoms in their fatty acid residues and one to three double bonds. Some less abundant species containing 32 carbon atoms were identified in all GP classes. In addition, species with 36 carbon atoms and up to five double bonds could be identified in the classes of PE, PC and PI. In contrast,
P. angusta shows a narrower distribution, exhibiting major species with 34 and 36 carbon atoms, and is thus shifted by one or two C
2-units compared to
K. thermotolerans. The number of double bonds varies from one to six. This distribution is also observed within the PEs. A slightly asymmetric shift to species with 72 carbon atoms for CA and 36 carbon atoms is observed for PC (
Figure 3a,d), whereas the classes of PG, PI and PS show a shift to species with 34 carbon atoms (
Figure 3b, e and f). The species distribution of
Y. lipolytica is similar to that of
P. angusta concerning the number and shift of carbon atoms, but in contrast, the maximum number of double bonds observed was four. In all three yeast strains, minor amounts of odd numbered GP species were identified, consisting mainly of 33 and 35 carbon atoms and more seldom of 31 carbon atoms, with a maximum relative amount of 1%. The impressions of
Figure 2 are confirmed, as the species distribution of CA, PE and PC is particularly characteristic for each of the three yeast strains, whereas the profiles of PG, PI and PS possess identical major species and just small differences in the less abundant species are observed.
Analysis of the compositions of GPs based on HPLC/ESI-LIT-FTICRMS in multistage mode showed that palmitic acid (16:0) and stearic acid (18:0) were the most prominent fatty acid residues, whereas palmitoleic acid (16:1) and oleic acid (18:1) were the major unsaturated species. Minor species comprised FAs 14:0, 14:1, 12:0 and 12:1 as well as the odd numbered FAs 15:0, 15:1, 17:0 and 17:1. These results are in good agreement with additional GC/MS measurements after hydrolysis and derivatization of the lipid extracts (data not shown).
The detailed comparative profiling of the second group reveals the similarity of the genetically related yeasts
S. cerevisiae and
S. bayanus, respectively. All identified species are considered due to the lower complexity of the profiles, and the results are depicted in
Figure 4.
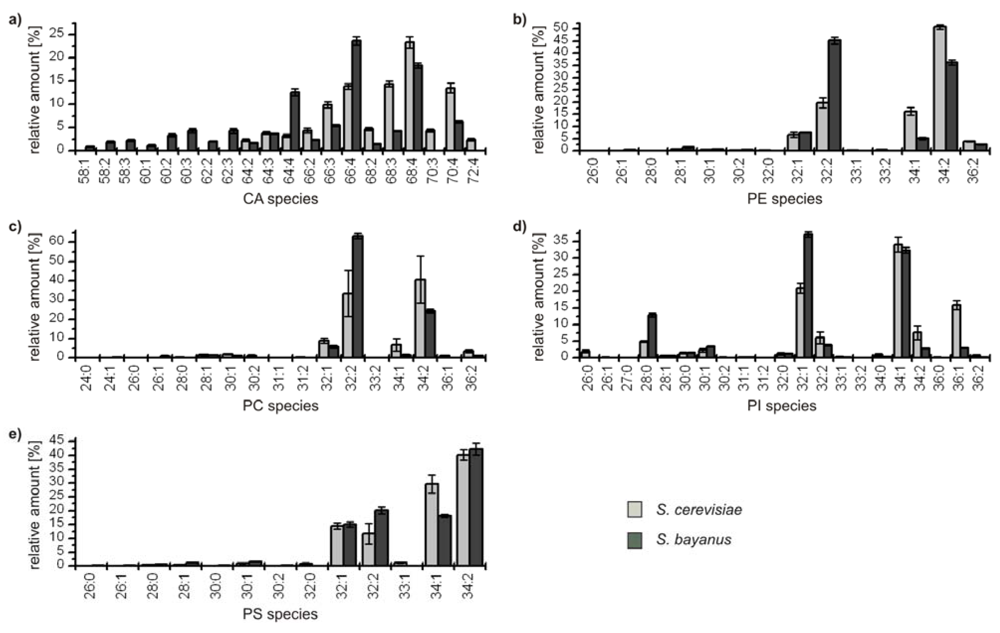
Figure 4.
Relative amounts of the GP species within the classes (a) cardiolipins (CA); (b) phophatidylethanolamines (PE); (c) phosphatidylcholines (PC); (d) phosphatidylinositoles (PI); and (e) phosphatidylserines (PS) for S. cerevisiae (■) and S. bayanus (■). The relative amount of respective species is referred to 100% peak area of all identified GPs within one class and yeast strain.
In all classes the distribution of the major compounds of S. cerevisiae and S. bayanus is similar, showing always four major GP species with 32 and 34 carbon atoms and one to two double bonds, except for the class of CA, which possesses a broader distribution due to its variation possibilities based on four bound fatty acids. It has to be noted that in the classes of PE and PC the most abundant species GP(34:2) of S. cerevisiae is the second most abundant species of S. bayanus, and vice versa for the species GP(32:2). Concerning the minor components, the lipid profiles of both yeast strains also show remarkable analogies. Only few species with 36 carbon atoms in the acyl chains could be identified and also some odd numbered species, bearing 31 and 33 carbon atoms. The maximum number of double bonds for a GP-species was two, except for CAs, where minor species with up to seven double bonds were identified. As PGs only were identified for S. bayanus, the diagram is not shown. The main species identified were PG(32:1), 40.8% ± 1.6% and PG(34:1), 43.1% ± 2.2%. Furthermore, PG(28:0), PG(26:0) and PG(32:2), contributing 7.7% ± 0.9%, 5.6% ± 0.8% and 2.8% ± 0.9% were identified.
The GP profiling data obtained for
S. cerevisiae within this work are in large parts in good agreement with previously published data by Ejsing
et al. [
11], who used a quantitative shotgun mass spectrometric approach. In both studies the same major GP species were identified, in particular PE(32:1), PE(32:2), PE(34:1), PE(34:2), PC(32:1), PC(32:2), PC(34:1), PC(34:2), PI(28:0), PI(32:1), PI(32:2), PI(34:1), PI(34:2), PS(32:1), PS(32:2), PS(34:1), and PS(34:2). Different results, referring to the study of Ejsing
et al. were for example obtained for the class of CAs. The major species identified in this study were CA(66:3), CA(66:4), CA(68:3), CA(68:4) and CA(70:4), whereas Ejsing
et al. found the species CA(64:4), CA(66:2) beside CA(66:4) and CA(68:4). In addition, no phosphatidic acid (PA) could be detected in
S. cerevisiae in contrast to the findings of Ejsing
et al., and each positive Profiler-Merger-Viewer software hit turned out to be an in-source fragmentation artifact of the corresponding PS species. However, the fact that PAs could be unambiguously identified in the other yeast strains, demonstrates that this class can also be detected by the applied method (see
Table S1 of the Supporting Information), but PAs may be below the limits of detection in the case of
S. cerevisiae. In addition to these predominant GP classes, in both studies minor amounts of PG could be identified, as well as the
lyso-forms of PE, PC, and PI. Moreover, the intermediates of PC biosynthesis via the PE-methylation pathway were identified,
i.e. MMPE and DMPE.
These results are also confirmed by other studies, where FA(16:1), FA(16:0) and FA(18:1) are described as most abundant fatty acids linked to the GPs. However, also FA(18:0) and FA(14:0) were frequently reported, which is in good accordance with the study of Ejsing
et al. and further former studies [
4,
9,
10,
11,
17,
18]. The instance that exclusively saturated and mono-unsaturated fatty acids were identified in
S. cerevsisae in the GP classes bearing two FAs is further corroborated by the genetic data of fatty acids synthesis, as
S. cerevisiae owns only a single fatty acid desaturase,
ole1p, which introduces a double bond in Δ9 position of FAs [
4,
19,
20]. Therefore, the restricted capacity of introducing double bonds is reflected in the GP profile of
S. cerevisisae.
Interestingly,
Y. lipolytica is known to possess an additional FA desaturase, which introduces a double bond in Δ12 position [
19]. This can explain the higher number of double bonds found in
Y. lipolytica’s GPs species. Unfortunately, no genetic data are available for the remaining yeast strains. However, due to the mentioned agreement of existing genetic data with the obtained lipid profiles, some of the lacking gene data can be extrapolated from the lipid profiles.
S. bayanus also seems to possess only the Δ9 desaturase, analogous to
S. cerevisiae.
K. thermotolerans and
P. angusta should own the additional Δ12 desaturase, like
Y. lipolytica. Moreover, they may contain further desaturases, introducing a third and fourth double bond in the FAs, as GPs with more than four double bonds in both fatty acid residues are identified for
K. thermotolerans and
P. angusta. For other fungi, the existence of those desaturases, which are likely Δ15 and Δ17 desaturases, is reported [
21,
22,
23,
24,
25,
26,
27,
28,
29,
30]. An external source, for example from the culture medium, can be excluded, as minimal medium was used [
4].
Besides the differences in the identity of the GP species, their relative amounts and distributions, we also studied the contribution of each the major GP classes to the whole GP lipidome. The relative amount was calculated as sum of all species belonging to a class in relation to the sum of all identified GPs. An overview is given in
Table 1.
Table 1.
Distribution of GP classes in the different yeasts. Relative amount is calculated as sum of all species constituting the same GP class.
| GP class | S. cerevisiae | S. bayanus | K. thermotolerans | P. angusta | Y. lipolytica |
|---|
| average [%] | SD [%] | average [%] | SD [%] | average [%] | SD [%] | average [%] | SD [%] | average [%] | SD [%] |
|---|
| CA | 3.6 | 1.6 | 6.1 | 0.6 | 3.1 | 1.2 | 2.4 | 0.4 | 2.8 | 0.3 |
| PG | 0.0 | 0.0 | 0.4 | 0.0 | 0.4 | 0.1 | 2.5 | 0.2 | 1.6 | 0.1 |
| PA | not identified | 0.3 | 0.0 | < 0.1 | 0.5 | 0.1 | 1.3 | 0.1 |
| PE | 17.4 | 1.1 | 24.7 | 2.2 | 14.9 | 2.3 | 27.8 | 0.8 | 31.9 | 0.4 |
| MMPE | 0.3 | 0.0 | 0.1 | 0.0 | < 0.1 | 2.5 | 0.2 | 0.9 | 0.0 |
| DMPE | 2.6 | 0.5 | 0.3 | 0.0 | 0.9 | 0.2 | not identified | 1.8 | 0.1 |
| PC | 31.5 | 8.7 | 12.5 | 3.0 | 26.3 | 7.9 | 40.0 | 0.6 | 32.6 | 0.8 |
| PI | 39.9 | 8.4 | 45.8 | 4.0 | 49.5 | 9.1 | 17.6 | 0.8 | 19.3 | 0.7 |
| PS | 4.8 | 0.5 | 9.7 | 0.7 | 4.9 | 0.7 | 6.7 | 0.3 | 7.7 | 0.3 |
The main GP class of
S. cerevisiae was determined to be PI. Equal observations are reported by Ejsing
et al [
11], but are deviating from former reports [
4,
9,
10,
17,
18]. PI is also the major GP class in
S. bayanus and
K. thermotolerans. The GP contents of
P. angusta and
Y. lipolytica mainly consist of PCs. It is remarkable that the closely related yeasts
S. cerevisiae and
S. bayanus differ significantly from each other concerning the relative amounts of PE, DMPE, PC and PS, unlike to their strong analogies in distribution of the species within a class. However, it is still obvious that the divergence in the GP class distribution is larger with increasing genetic differences.